Please be patient as the page loads
|
ECEL1_MOUSE
|
||||||
| SwissProt Accessions | Q9JMI0 | Gene names | Ecel1, Dine, Xce | |||
|
Domain Architecture |

|
|||||
| Description | Endothelin-converting enzyme-like 1 (EC 3.4.24.-) (Xce protein) (Damage-induced neuronal endopeptidase). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ECEL1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999131 (rank : 2) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O95672, Q45UD9, Q53RF9, Q6UW86, Q86TH4, Q9NY95 | Gene names | ECEL1, XCE | |||
|
Domain Architecture |

|
|||||
| Description | Endothelin-converting enzyme-like 1 (EC 3.4.24.-) (Xce protein). | |||||
|
ECEL1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9JMI0 | Gene names | Ecel1, Dine, Xce | |||
|
Domain Architecture |

|
|||||
| Description | Endothelin-converting enzyme-like 1 (EC 3.4.24.-) (Xce protein) (Damage-induced neuronal endopeptidase). | |||||
|
ECE2_HUMAN
|
||||||
| θ value | 6.82227e-149 (rank : 3) | NC score | 0.977721 (rank : 5) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O60344, Q96NX3, Q96NX4 | Gene names | ECE2, KIAA0604 | |||
|
Domain Architecture |

|
|||||
| Description | Endothelin-converting enzyme 2 (EC 3.4.24.71) (ECE-2). | |||||
|
ECE1_HUMAN
|
||||||
| θ value | 8.91014e-149 (rank : 4) | NC score | 0.978055 (rank : 4) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P42892, Q14217, Q58GE7, Q5THM5, Q5THM7, Q5THM8, Q9UJQ6, Q9UPF4, Q9UPM4, Q9Y501 | Gene names | ECE1 | |||
|
Domain Architecture |

|
|||||
| Description | Endothelin-converting enzyme 1 (EC 3.4.24.71) (ECE-1). | |||||
|
ECE1_MOUSE
|
||||||
| θ value | 2.19465e-147 (rank : 5) | NC score | 0.978215 (rank : 3) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q4PZA2, Q4PZ99, Q4PZA1, Q6P9Q9 | Gene names | Ece1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelin-converting enzyme 1 (EC 3.4.24.71) (ECE-1). | |||||
|
MMEL1_MOUSE
|
||||||
| θ value | 1.63136e-134 (rank : 6) | NC score | 0.972253 (rank : 7) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9JLI3, Q3U495, Q9ERK2, Q9ERK3, Q9QZV6, Q9QZV7 | Gene names | Mmel1, Nep2, Nl1, Sep | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane metallo-endopeptidase-like 1 (EC 3.4.24.11) (Neprilysin-2) (Neprilysin II) (NL2) (NEPII) (NEP2(m)) (Neprilysin-like peptidase) (NEPLP) (Neprilysin-like 1) (NL-1) (Soluble secreted endopeptidase) [Contains: Membrane metallo-endopeptidase-like 1, soluble form (Neprilysin-2 secreted) (NEP2(s))]. | |||||
|
MMEL1_HUMAN
|
||||||
| θ value | 1.29261e-131 (rank : 7) | NC score | 0.972548 (rank : 6) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q495T6, Q495T7, Q495T8, Q5SZS6, Q96PH9 | Gene names | MMEL1, MELL1, MMEL2, NEP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane metallo-endopeptidase-like 1 (EC 3.4.24.11) (Membrane metallo-endopeptidase-like 2) (Neprilysin-2) (Neprilysin II) (NL2) (NEPII) (NEP2(m)) [Contains: Membrane metallo-endopeptidase-like 1, soluble form (Neprilysin-2 secreted) (NEP2(s))]. | |||||
|
NEP_HUMAN
|
||||||
| θ value | 2.06847e-121 (rank : 8) | NC score | 0.969690 (rank : 8) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P08473 | Gene names | MME, EPN | |||
|
Domain Architecture |

|
|||||
| Description | Neprilysin (EC 3.4.24.11) (Neutral endopeptidase) (NEP) (Enkephalinase) (Neutral endopeptidase 24.11) (Atriopeptidase) (Common acute lymphocytic leukemia antigen) (CALLA) (CD10 antigen). | |||||
|
NEP_MOUSE
|
||||||
| θ value | 7.86017e-121 (rank : 9) | NC score | 0.969412 (rank : 9) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q61391, Q6NXX5, Q8K251 | Gene names | Mme | |||
|
Domain Architecture |

|
|||||
| Description | Neprilysin (EC 3.4.24.11) (Neutral endopeptidase) (NEP) (Enkephalinase) (Neutral endopeptidase 24.11) (Atriopeptidase) (CD10 antigen). | |||||
|
PHEX_MOUSE
|
||||||
| θ value | 6.68496e-104 (rank : 10) | NC score | 0.963831 (rank : 10) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P70669, P97439 | Gene names | Phex, Hyp, Pex | |||
|
Domain Architecture |

|
|||||
| Description | Metalloendopeptidase homolog PEX (EC 3.4.24.-) (Phosphate regulating neutral endopeptidase) (X-linked hypophosphatemia protein) (HYP) (Vitamin D-resistant hypophosphatemic rickets protein). | |||||
|
PHEX_HUMAN
|
||||||
| θ value | 1.94503e-103 (rank : 11) | NC score | 0.963492 (rank : 11) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P78562, O00678, Q13646, Q93032, Q99827 | Gene names | PHEX, PEX | |||
|
Domain Architecture |

|
|||||
| Description | Phosphate-regulating neutral endopeptidase (EC 3.4.24.-) (Metalloendopeptidase homolog PEX) (X-linked hypophosphatemia protein) (HYP) (Vitamin D-resistant hypophosphatemic rickets protein). | |||||
|
KELL_HUMAN
|
||||||
| θ value | 2.09745e-73 (rank : 12) | NC score | 0.955184 (rank : 12) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P23276, Q96RS8, Q99885 | Gene names | KEL | |||
|
Domain Architecture |

|
|||||
| Description | Kell blood group glycoprotein (EC 3.4.24.-) (CD238 antigen). | |||||
|
SNX18_MOUSE
|
||||||
| θ value | 2.36792 (rank : 13) | NC score | 0.010163 (rank : 16) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91ZR2 | Gene names | Snag1, Snx18 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-18 (Sorting nexin-associated Golgi protein 1). | |||||
|
CHCH6_HUMAN
|
||||||
| θ value | 4.03905 (rank : 14) | NC score | 0.014395 (rank : 15) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BRQ6 | Gene names | CHCHD6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil-helix-coiled-coil-helix domain-containing protein 6. | |||||
|
IMA2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 15) | NC score | 0.006312 (rank : 23) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P52292, Q9BRU5 | Gene names | KPNA2, RCH1, SRP1 | |||
|
Domain Architecture |
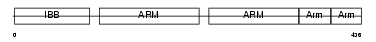
|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1). | |||||
|
PSA_HUMAN
|
||||||
| θ value | 4.03905 (rank : 16) | NC score | 0.014449 (rank : 14) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P55786, Q6P145, Q9NP16, Q9UEM2 | Gene names | NPEPPS, PSA | |||
|
Domain Architecture |

|
|||||
| Description | Puromycin-sensitive aminopeptidase (EC 3.4.11.-) (PSA). | |||||
|
PSA_MOUSE
|
||||||
| θ value | 4.03905 (rank : 17) | NC score | 0.014471 (rank : 13) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q11011, Q91VJ8 | Gene names | Npepps, Psa | |||
|
Domain Architecture |

|
|||||
| Description | Puromycin-sensitive aminopeptidase (EC 3.4.11.-) (PSA). | |||||
|
DESP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 18) | NC score | -0.000202 (rank : 26) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P15924, O75993, Q14189, Q9UHN4 | Gene names | DSP | |||
|
Domain Architecture |

|
|||||
| Description | Desmoplakin (DP) (250/210 kDa paraneoplastic pemphigus antigen). | |||||
|
LUZP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 19) | NC score | 0.003112 (rank : 24) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 739 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86V48, Q5TH93, Q8N4X3, Q8TEH1 | Gene names | LUZP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper protein 1. | |||||
|
MX2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 20) | NC score | 0.006412 (rank : 22) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P20592 | Gene names | MX2 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced GTP-binding protein Mx2 (Interferon-regulated resistance GTP-binding protein MxB) (p78-related protein). | |||||
|
BRD8_MOUSE
|
||||||
| θ value | 6.88961 (rank : 21) | NC score | 0.008071 (rank : 18) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 422 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R3B7, Q8C049, Q8R583, Q8VDP0 | Gene names | Brd8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 8. | |||||
|
AMPE_HUMAN
|
||||||
| θ value | 8.99809 (rank : 22) | NC score | 0.009874 (rank : 17) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q07075 | Gene names | ENPEP | |||
|
Domain Architecture |
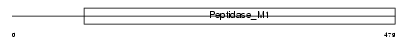
|
|||||
| Description | Glutamyl aminopeptidase (EC 3.4.11.7) (EAP) (Aminopeptidase A) (APA) (Differentiation antigen gp160) (CD249 antigen). | |||||
|
BRD8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 23) | NC score | 0.007918 (rank : 19) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H0E9, O43178, Q15355, Q59GN0, Q969M9 | Gene names | BRD8, SMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 8 (p120) (Skeletal muscle abundant protein) (Thyroid hormone receptor coactivating protein 120kDa) (TrCP120). | |||||
|
DDX56_MOUSE
|
||||||
| θ value | 8.99809 (rank : 24) | NC score | 0.001878 (rank : 25) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D0R4 | Gene names | Ddx56, D11Ertd619e, Noh61 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DDX56 (EC 3.6.1.-) (DEAD box protein 56) (ATP-dependent 61 kDa nucleolar RNA helicase). | |||||
|
ITIH2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 25) | NC score | 0.006895 (rank : 21) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61703 | Gene names | Itih2 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H2 precursor (ITI heavy chain H2) (Inter-alpha-inhibitor heavy chain 2). | |||||
|
ORC3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 26) | NC score | 0.007689 (rank : 20) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UBD5, Q13565, Q6IUY7, Q9UG44, Q9UNT6 | Gene names | ORC3L, LATHEO | |||
|
Domain Architecture |
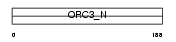
|
|||||
| Description | Origin recognition complex subunit 3 (Origin recognition complex subunit Latheo). | |||||
|
ECEL1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9JMI0 | Gene names | Ecel1, Dine, Xce | |||
|
Domain Architecture |

|
|||||
| Description | Endothelin-converting enzyme-like 1 (EC 3.4.24.-) (Xce protein) (Damage-induced neuronal endopeptidase). | |||||
|
ECEL1_HUMAN
|
||||||
| NC score | 0.999131 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O95672, Q45UD9, Q53RF9, Q6UW86, Q86TH4, Q9NY95 | Gene names | ECEL1, XCE | |||
|
Domain Architecture |

|
|||||
| Description | Endothelin-converting enzyme-like 1 (EC 3.4.24.-) (Xce protein). | |||||
|
ECE1_MOUSE
|
||||||
| NC score | 0.978215 (rank : 3) | θ value | 2.19465e-147 (rank : 5) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q4PZA2, Q4PZ99, Q4PZA1, Q6P9Q9 | Gene names | Ece1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelin-converting enzyme 1 (EC 3.4.24.71) (ECE-1). | |||||
|
ECE1_HUMAN
|
||||||
| NC score | 0.978055 (rank : 4) | θ value | 8.91014e-149 (rank : 4) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P42892, Q14217, Q58GE7, Q5THM5, Q5THM7, Q5THM8, Q9UJQ6, Q9UPF4, Q9UPM4, Q9Y501 | Gene names | ECE1 | |||
|
Domain Architecture |

|
|||||
| Description | Endothelin-converting enzyme 1 (EC 3.4.24.71) (ECE-1). | |||||
|
ECE2_HUMAN
|
||||||
| NC score | 0.977721 (rank : 5) | θ value | 6.82227e-149 (rank : 3) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O60344, Q96NX3, Q96NX4 | Gene names | ECE2, KIAA0604 | |||
|
Domain Architecture |

|
|||||
| Description | Endothelin-converting enzyme 2 (EC 3.4.24.71) (ECE-2). | |||||
|
MMEL1_HUMAN
|
||||||
| NC score | 0.972548 (rank : 6) | θ value | 1.29261e-131 (rank : 7) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q495T6, Q495T7, Q495T8, Q5SZS6, Q96PH9 | Gene names | MMEL1, MELL1, MMEL2, NEP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane metallo-endopeptidase-like 1 (EC 3.4.24.11) (Membrane metallo-endopeptidase-like 2) (Neprilysin-2) (Neprilysin II) (NL2) (NEPII) (NEP2(m)) [Contains: Membrane metallo-endopeptidase-like 1, soluble form (Neprilysin-2 secreted) (NEP2(s))]. | |||||
|
MMEL1_MOUSE
|
||||||
| NC score | 0.972253 (rank : 7) | θ value | 1.63136e-134 (rank : 6) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9JLI3, Q3U495, Q9ERK2, Q9ERK3, Q9QZV6, Q9QZV7 | Gene names | Mmel1, Nep2, Nl1, Sep | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane metallo-endopeptidase-like 1 (EC 3.4.24.11) (Neprilysin-2) (Neprilysin II) (NL2) (NEPII) (NEP2(m)) (Neprilysin-like peptidase) (NEPLP) (Neprilysin-like 1) (NL-1) (Soluble secreted endopeptidase) [Contains: Membrane metallo-endopeptidase-like 1, soluble form (Neprilysin-2 secreted) (NEP2(s))]. | |||||
|
NEP_HUMAN
|
||||||
| NC score | 0.969690 (rank : 8) | θ value | 2.06847e-121 (rank : 8) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P08473 | Gene names | MME, EPN | |||
|
Domain Architecture |

|
|||||
| Description | Neprilysin (EC 3.4.24.11) (Neutral endopeptidase) (NEP) (Enkephalinase) (Neutral endopeptidase 24.11) (Atriopeptidase) (Common acute lymphocytic leukemia antigen) (CALLA) (CD10 antigen). | |||||
|
NEP_MOUSE
|
||||||
| NC score | 0.969412 (rank : 9) | θ value | 7.86017e-121 (rank : 9) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q61391, Q6NXX5, Q8K251 | Gene names | Mme | |||
|
Domain Architecture |

|
|||||
| Description | Neprilysin (EC 3.4.24.11) (Neutral endopeptidase) (NEP) (Enkephalinase) (Neutral endopeptidase 24.11) (Atriopeptidase) (CD10 antigen). | |||||
|
PHEX_MOUSE
|
||||||
| NC score | 0.963831 (rank : 10) | θ value | 6.68496e-104 (rank : 10) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P70669, P97439 | Gene names | Phex, Hyp, Pex | |||
|
Domain Architecture |

|
|||||
| Description | Metalloendopeptidase homolog PEX (EC 3.4.24.-) (Phosphate regulating neutral endopeptidase) (X-linked hypophosphatemia protein) (HYP) (Vitamin D-resistant hypophosphatemic rickets protein). | |||||
|
PHEX_HUMAN
|
||||||
| NC score | 0.963492 (rank : 11) | θ value | 1.94503e-103 (rank : 11) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P78562, O00678, Q13646, Q93032, Q99827 | Gene names | PHEX, PEX | |||
|
Domain Architecture |

|
|||||
| Description | Phosphate-regulating neutral endopeptidase (EC 3.4.24.-) (Metalloendopeptidase homolog PEX) (X-linked hypophosphatemia protein) (HYP) (Vitamin D-resistant hypophosphatemic rickets protein). | |||||
|
KELL_HUMAN
|
||||||
| NC score | 0.955184 (rank : 12) | θ value | 2.09745e-73 (rank : 12) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P23276, Q96RS8, Q99885 | Gene names | KEL | |||
|
Domain Architecture |

|
|||||
| Description | Kell blood group glycoprotein (EC 3.4.24.-) (CD238 antigen). | |||||
|
PSA_MOUSE
|
||||||
| NC score | 0.014471 (rank : 13) | θ value | 4.03905 (rank : 17) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q11011, Q91VJ8 | Gene names | Npepps, Psa | |||
|
Domain Architecture |

|
|||||
| Description | Puromycin-sensitive aminopeptidase (EC 3.4.11.-) (PSA). | |||||
|
PSA_HUMAN
|
||||||
| NC score | 0.014449 (rank : 14) | θ value | 4.03905 (rank : 16) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P55786, Q6P145, Q9NP16, Q9UEM2 | Gene names | NPEPPS, PSA | |||
|
Domain Architecture |

|
|||||
| Description | Puromycin-sensitive aminopeptidase (EC 3.4.11.-) (PSA). | |||||
|
CHCH6_HUMAN
|
||||||
| NC score | 0.014395 (rank : 15) | θ value | 4.03905 (rank : 14) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BRQ6 | Gene names | CHCHD6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil-helix-coiled-coil-helix domain-containing protein 6. | |||||
|
SNX18_MOUSE
|
||||||
| NC score | 0.010163 (rank : 16) | θ value | 2.36792 (rank : 13) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91ZR2 | Gene names | Snag1, Snx18 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-18 (Sorting nexin-associated Golgi protein 1). | |||||
|
AMPE_HUMAN
|
||||||
| NC score | 0.009874 (rank : 17) | θ value | 8.99809 (rank : 22) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q07075 | Gene names | ENPEP | |||
|
Domain Architecture |

|
|||||
| Description | Glutamyl aminopeptidase (EC 3.4.11.7) (EAP) (Aminopeptidase A) (APA) (Differentiation antigen gp160) (CD249 antigen). | |||||
|
BRD8_MOUSE
|
||||||
| NC score | 0.008071 (rank : 18) | θ value | 6.88961 (rank : 21) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 422 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R3B7, Q8C049, Q8R583, Q8VDP0 | Gene names | Brd8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 8. | |||||
|
BRD8_HUMAN
|
||||||
| NC score | 0.007918 (rank : 19) | θ value | 8.99809 (rank : 23) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H0E9, O43178, Q15355, Q59GN0, Q969M9 | Gene names | BRD8, SMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 8 (p120) (Skeletal muscle abundant protein) (Thyroid hormone receptor coactivating protein 120kDa) (TrCP120). | |||||
|
ORC3_HUMAN
|
||||||
| NC score | 0.007689 (rank : 20) | θ value | 8.99809 (rank : 26) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UBD5, Q13565, Q6IUY7, Q9UG44, Q9UNT6 | Gene names | ORC3L, LATHEO | |||
|
Domain Architecture |

|
|||||
| Description | Origin recognition complex subunit 3 (Origin recognition complex subunit Latheo). | |||||
|
ITIH2_MOUSE
|
||||||
| NC score | 0.006895 (rank : 21) | θ value | 8.99809 (rank : 25) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61703 | Gene names | Itih2 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H2 precursor (ITI heavy chain H2) (Inter-alpha-inhibitor heavy chain 2). | |||||
|
MX2_HUMAN
|
||||||
| NC score | 0.006412 (rank : 22) | θ value | 5.27518 (rank : 20) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P20592 | Gene names | MX2 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced GTP-binding protein Mx2 (Interferon-regulated resistance GTP-binding protein MxB) (p78-related protein). | |||||
|
IMA2_HUMAN
|
||||||
| NC score | 0.006312 (rank : 23) | θ value | 4.03905 (rank : 15) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P52292, Q9BRU5 | Gene names | KPNA2, RCH1, SRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1). | |||||
|
LUZP1_HUMAN
|
||||||
| NC score | 0.003112 (rank : 24) | θ value | 5.27518 (rank : 19) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 739 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86V48, Q5TH93, Q8N4X3, Q8TEH1 | Gene names | LUZP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper protein 1. | |||||
|
DDX56_MOUSE
|
||||||
| NC score | 0.001878 (rank : 25) | θ value | 8.99809 (rank : 24) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D0R4 | Gene names | Ddx56, D11Ertd619e, Noh61 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DDX56 (EC 3.6.1.-) (DEAD box protein 56) (ATP-dependent 61 kDa nucleolar RNA helicase). | |||||
|
DESP_HUMAN
|
||||||
| NC score | -0.000202 (rank : 26) | θ value | 5.27518 (rank : 18) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P15924, O75993, Q14189, Q9UHN4 | Gene names | DSP | |||
|
Domain Architecture |

|
|||||
| Description | Desmoplakin (DP) (250/210 kDa paraneoplastic pemphigus antigen). | |||||