Please be patient as the page loads
|
ITIH2_MOUSE
|
||||||
| SwissProt Accessions | Q61703 | Gene names | Itih2 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H2 precursor (ITI heavy chain H2) (Inter-alpha-inhibitor heavy chain 2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ITIH2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.995163 (rank : 2) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P19823, Q14659, Q15484 | Gene names | ITIH2, IGHEP2 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H2 precursor (ITI heavy chain H2) (Inter-alpha-inhibitor heavy chain 2) (Inter-alpha-trypsin inhibitor complex component II) (Serum-derived hyaluronan-associated protein) (SHAP). | |||||
|
ITIH2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q61703 | Gene names | Itih2 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H2 precursor (ITI heavy chain H2) (Inter-alpha-inhibitor heavy chain 2). | |||||
|
ITIH3_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.970738 (rank : 4) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q06033, Q99085 | Gene names | ITIH3 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H3 precursor (ITI heavy chain H3) (Inter-alpha-inhibitor heavy chain 3) (Serum-derived hyaluronan-associated protein) (SHAP). | |||||
|
ITIH3_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.971415 (rank : 3) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q61704 | Gene names | Itih3 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H3 precursor (ITI heavy chain H3) (Inter-alpha-inhibitor heavy chain 3). | |||||
|
ITIH1_MOUSE
|
||||||
| θ value | 3.47146e-177 (rank : 5) | NC score | 0.969596 (rank : 5) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q61702 | Gene names | Itih1 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H1 precursor (ITI heavy chain H1) (Inter-alpha-inhibitor heavy chain 1). | |||||
|
ITIH1_HUMAN
|
||||||
| θ value | 3.71755e-171 (rank : 6) | NC score | 0.965392 (rank : 6) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P19827, P78455, Q01746 | Gene names | ITIH1, IGHEP1 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H1 precursor (ITI heavy chain H1) (Inter-alpha-inhibitor heavy chain 1) (Inter-alpha-trypsin inhibitor complex component III) (Serum-derived hyaluronan-associated protein) (SHAP). | |||||
|
ITIH4_HUMAN
|
||||||
| θ value | 7.57797e-132 (rank : 7) | NC score | 0.955242 (rank : 7) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q14624, Q15135, Q9P190, Q9UQ54 | Gene names | ITIH4, IHRP, ITIHL1, PK120 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H4 precursor (ITI heavy chain H4) (Inter-alpha-inhibitor heavy chain 4) (Inter-alpha-trypsin inhibitor family heavy chain-related protein) (IHRP) (Plasma kallikrein sensitive glycoprotein 120) (PK-120) (GP120) [Contains: 70 kDa inter-alpha-trypsin inhibitor heavy chain H4; 35 kDa inter-alpha- trypsin inhibitor heavy chain H4]. | |||||
|
CAC2D_MOUSE
|
||||||
| θ value | 5.44631e-05 (rank : 8) | NC score | 0.312839 (rank : 8) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O08532, O08533, O08534, O08535, O08536 | Gene names | Cacna2d1, Cacna2 | |||
|
Domain Architecture |

|
|||||
| Description | Dihydropyridine-sensitive L-type calcium channel subunits alpha- 2/delta precursor [Contains: L-type calcium channel subunit alpha-2; L-type calcium channel subunit delta]. | |||||
|
CAC2D_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 9) | NC score | 0.311200 (rank : 9) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P54289 | Gene names | CACNA2D1, CACNL2A, CCHL2A | |||
|
Domain Architecture |

|
|||||
| Description | Dihydropyridine-sensitive L-type calcium channel subunits alpha- 2/delta precursor [Contains: L-type calcium channel subunit alpha-2; L-type calcium channel subunit delta]. | |||||
|
PARP4_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 10) | NC score | 0.247242 (rank : 10) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UKK3, O75903, Q14682, Q9H1M6 | Gene names | PARP4, ADPRTL1, KIAA0177, PARPL | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 4 (EC 2.4.2.30) (PARP-4) (Vault poly(ADP- ribose) polymerase) (VPARP) (193 kDa vault protein) (PARP- related/IalphaI-related H5/proline-rich) (PH5P). | |||||
|
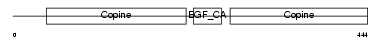
MATN1_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 11) | NC score | 0.066087 (rank : 13) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P51942 | Gene names | Matn1, Cmp, Crtm | |||
|
Domain Architecture |
|
|||||
| Description | Cartilage matrix protein precursor (Matrilin-1). | |||||
|
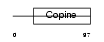
LHR2A_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 12) | NC score | 0.222215 (rank : 12) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99KC8, Q3TTU2, Q6UN22, Q8BHA8, Q9CTV9 | Gene names | Loh11cr2a | |||
|
Domain Architecture |
|
|||||
| Description | Loss of heterozygosity 11 chromosomal region 2 gene A protein homolog. | |||||
|
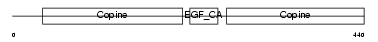
MATN1_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 13) | NC score | 0.059340 (rank : 14) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P21941 | Gene names | MATN1, CMP, CRTM | |||
|
Domain Architecture |
|
|||||
| Description | Cartilage matrix protein precursor (Matrilin-1). | |||||
|
COKA1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 14) | NC score | 0.031120 (rank : 15) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9P218, Q4VXQ4, Q8WUT2, Q96CY9, Q9BQU6, Q9BQU7 | Gene names | COL20A1, KIAA1510 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XX) chain precursor. | |||||
|
CO6A1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 15) | NC score | 0.030140 (rank : 16) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q04857 | Gene names | Col6a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VI) chain precursor. | |||||
|
TPH2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 16) | NC score | 0.013719 (rank : 19) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CGV2 | Gene names | Tph2, Ntph | |||
|
Domain Architecture |

|
|||||
| Description | Tryptophan 5-hydroxylase 2 (EC 1.14.16.4) (Tryptophan 5-monooxygenase 2) (Neuronal tryptophan hydroxylase). | |||||
|
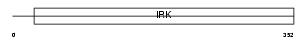
IRK15_HUMAN
|
||||||
| θ value | 1.81305 (rank : 17) | NC score | 0.011500 (rank : 20) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99712, O00564, Q96L28, Q99446 | Gene names | KCNJ15, KCNJ14 | |||
|
Domain Architecture |
|
|||||
| Description | ATP-sensitive inward rectifier potassium channel 15 (Potassium channel, inwardly rectifying subfamily J member 15) (Inward rectifier K(+) channel Kir4.2) (Kir1.3). | |||||
|
IRK15_MOUSE
|
||||||
| θ value | 1.81305 (rank : 18) | NC score | 0.011500 (rank : 21) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88932, Q9JK34 | Gene names | Kcnj15 | |||
|
Domain Architecture |
|
|||||
| Description | ATP-sensitive inward rectifier potassium channel 15 (Potassium channel, inwardly rectifying subfamily J member 15) (Inward rectifier K(+) channel Kir4.2). | |||||
|
ITAX_MOUSE
|
||||||
| θ value | 3.0926 (rank : 19) | NC score | 0.014363 (rank : 18) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9QXH4 | Gene names | Itgax | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-X precursor (Leukocyte adhesion glycoprotein p150,95 alpha chain) (Leukocyte adhesion receptor p150,95) (CD11c antigen). | |||||
|
AFF2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 20) | NC score | 0.008990 (rank : 22) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O55112 | Gene names | Aff2, Fmr2, Ox19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation protein 2 homolog) (Protein FMR-2) (FMR2P) (Protein Ox19). | |||||
|
KIF3C_HUMAN
|
||||||
| θ value | 6.88961 (rank : 21) | NC score | 0.006353 (rank : 28) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 478 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O14782, O43544 | Gene names | KIF3C | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin-like protein KIF3C. | |||||
|
KIF3C_MOUSE
|
||||||
| θ value | 6.88961 (rank : 22) | NC score | 0.006390 (rank : 27) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 468 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O35066, O35229 | Gene names | Kif3c | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin-like protein KIF3C. | |||||
|
UBP25_HUMAN
|
||||||
| θ value | 6.88961 (rank : 23) | NC score | 0.006666 (rank : 26) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UHP3, Q9H9W1 | Gene names | USP25, USP21 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 25 (EC 3.1.2.15) (Ubiquitin thioesterase 25) (Ubiquitin-specific-processing protease 25) (Deubiquitinating enzyme 25) (USP on chromosome 21). | |||||
|
ECEL1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 24) | NC score | 0.006885 (rank : 25) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95672, Q45UD9, Q53RF9, Q6UW86, Q86TH4, Q9NY95 | Gene names | ECEL1, XCE | |||
|
Domain Architecture |

|
|||||
| Description | Endothelin-converting enzyme-like 1 (EC 3.4.24.-) (Xce protein). | |||||
|
ECEL1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 25) | NC score | 0.006895 (rank : 24) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JMI0 | Gene names | Ecel1, Dine, Xce | |||
|
Domain Architecture |

|
|||||
| Description | Endothelin-converting enzyme-like 1 (EC 3.4.24.-) (Xce protein) (Damage-induced neuronal endopeptidase). | |||||
|
GLGB_MOUSE
|
||||||
| θ value | 8.99809 (rank : 26) | NC score | 0.008974 (rank : 23) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D6Y9 | Gene names | Gbe1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 1,4-alpha-glucan branching enzyme (EC 2.4.1.18) (Glycogen branching enzyme) (Brancher enzyme). | |||||
|
MATN3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 27) | NC score | 0.022001 (rank : 17) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O35701, Q9JHM0 | Gene names | Matn3 | |||
|
Domain Architecture |
|
|||||
| Description | Matrilin-3 precursor. | |||||
|
LHR2A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 28) | NC score | 0.233689 (rank : 11) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O00534, Q6UN19, Q6UN20, Q9BVF8 | Gene names | LOH11CR2A, BCSC1 | |||
|
Domain Architecture |

|
|||||
| Description | Loss of heterozygosity 11 chromosomal region 2 gene A protein (Breast cancer suppressor candidate 1) (BCSC-1). | |||||
|
ITIH2_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q61703 | Gene names | Itih2 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H2 precursor (ITI heavy chain H2) (Inter-alpha-inhibitor heavy chain 2). | |||||
|
ITIH2_HUMAN
|
||||||
| NC score | 0.995163 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P19823, Q14659, Q15484 | Gene names | ITIH2, IGHEP2 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H2 precursor (ITI heavy chain H2) (Inter-alpha-inhibitor heavy chain 2) (Inter-alpha-trypsin inhibitor complex component II) (Serum-derived hyaluronan-associated protein) (SHAP). | |||||
|
ITIH3_MOUSE
|
||||||
| NC score | 0.971415 (rank : 3) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q61704 | Gene names | Itih3 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H3 precursor (ITI heavy chain H3) (Inter-alpha-inhibitor heavy chain 3). | |||||
|
ITIH3_HUMAN
|
||||||
| NC score | 0.970738 (rank : 4) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q06033, Q99085 | Gene names | ITIH3 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H3 precursor (ITI heavy chain H3) (Inter-alpha-inhibitor heavy chain 3) (Serum-derived hyaluronan-associated protein) (SHAP). | |||||
|
ITIH1_MOUSE
|
||||||
| NC score | 0.969596 (rank : 5) | θ value | 3.47146e-177 (rank : 5) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q61702 | Gene names | Itih1 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H1 precursor (ITI heavy chain H1) (Inter-alpha-inhibitor heavy chain 1). | |||||
|
ITIH1_HUMAN
|
||||||
| NC score | 0.965392 (rank : 6) | θ value | 3.71755e-171 (rank : 6) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P19827, P78455, Q01746 | Gene names | ITIH1, IGHEP1 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H1 precursor (ITI heavy chain H1) (Inter-alpha-inhibitor heavy chain 1) (Inter-alpha-trypsin inhibitor complex component III) (Serum-derived hyaluronan-associated protein) (SHAP). | |||||
|
ITIH4_HUMAN
|
||||||
| NC score | 0.955242 (rank : 7) | θ value | 7.57797e-132 (rank : 7) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q14624, Q15135, Q9P190, Q9UQ54 | Gene names | ITIH4, IHRP, ITIHL1, PK120 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H4 precursor (ITI heavy chain H4) (Inter-alpha-inhibitor heavy chain 4) (Inter-alpha-trypsin inhibitor family heavy chain-related protein) (IHRP) (Plasma kallikrein sensitive glycoprotein 120) (PK-120) (GP120) [Contains: 70 kDa inter-alpha-trypsin inhibitor heavy chain H4; 35 kDa inter-alpha- trypsin inhibitor heavy chain H4]. | |||||
|
CAC2D_MOUSE
|
||||||
| NC score | 0.312839 (rank : 8) | θ value | 5.44631e-05 (rank : 8) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O08532, O08533, O08534, O08535, O08536 | Gene names | Cacna2d1, Cacna2 | |||
|
Domain Architecture |

|
|||||
| Description | Dihydropyridine-sensitive L-type calcium channel subunits alpha- 2/delta precursor [Contains: L-type calcium channel subunit alpha-2; L-type calcium channel subunit delta]. | |||||
|
CAC2D_HUMAN
|
||||||
| NC score | 0.311200 (rank : 9) | θ value | 0.000158464 (rank : 9) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P54289 | Gene names | CACNA2D1, CACNL2A, CCHL2A | |||
|
Domain Architecture |

|
|||||
| Description | Dihydropyridine-sensitive L-type calcium channel subunits alpha- 2/delta precursor [Contains: L-type calcium channel subunit alpha-2; L-type calcium channel subunit delta]. | |||||
|
PARP4_HUMAN
|
||||||
| NC score | 0.247242 (rank : 10) | θ value | 0.00175202 (rank : 10) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UKK3, O75903, Q14682, Q9H1M6 | Gene names | PARP4, ADPRTL1, KIAA0177, PARPL | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 4 (EC 2.4.2.30) (PARP-4) (Vault poly(ADP- ribose) polymerase) (VPARP) (193 kDa vault protein) (PARP- related/IalphaI-related H5/proline-rich) (PH5P). | |||||
|
LHR2A_HUMAN
|
||||||
| NC score | 0.233689 (rank : 11) | θ value | θ > 10 (rank : 28) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O00534, Q6UN19, Q6UN20, Q9BVF8 | Gene names | LOH11CR2A, BCSC1 | |||
|
Domain Architecture |

|
|||||
| Description | Loss of heterozygosity 11 chromosomal region 2 gene A protein (Breast cancer suppressor candidate 1) (BCSC-1). | |||||
|
LHR2A_MOUSE
|
||||||
| NC score | 0.222215 (rank : 12) | θ value | 0.00390308 (rank : 12) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99KC8, Q3TTU2, Q6UN22, Q8BHA8, Q9CTV9 | Gene names | Loh11cr2a | |||
|
Domain Architecture |

|
|||||
| Description | Loss of heterozygosity 11 chromosomal region 2 gene A protein homolog. | |||||
|
MATN1_MOUSE
|
||||||
| NC score | 0.066087 (rank : 13) | θ value | 0.00228821 (rank : 11) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P51942 | Gene names | Matn1, Cmp, Crtm | |||
|
Domain Architecture |

|
|||||
| Description | Cartilage matrix protein precursor (Matrilin-1). | |||||
|
MATN1_HUMAN
|
||||||
| NC score | 0.059340 (rank : 14) | θ value | 0.00390308 (rank : 13) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P21941 | Gene names | MATN1, CMP, CRTM | |||
|
Domain Architecture |

|
|||||
| Description | Cartilage matrix protein precursor (Matrilin-1). | |||||
|
COKA1_HUMAN
|
||||||
| NC score | 0.031120 (rank : 15) | θ value | 0.21417 (rank : 14) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9P218, Q4VXQ4, Q8WUT2, Q96CY9, Q9BQU6, Q9BQU7 | Gene names | COL20A1, KIAA1510 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XX) chain precursor. | |||||
|
CO6A1_MOUSE
|
||||||
| NC score | 0.030140 (rank : 16) | θ value | 1.38821 (rank : 15) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q04857 | Gene names | Col6a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VI) chain precursor. | |||||
|
MATN3_MOUSE
|
||||||
| NC score | 0.022001 (rank : 17) | θ value | 8.99809 (rank : 27) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O35701, Q9JHM0 | Gene names | Matn3 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilin-3 precursor. | |||||
|
ITAX_MOUSE
|
||||||
| NC score | 0.014363 (rank : 18) | θ value | 3.0926 (rank : 19) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9QXH4 | Gene names | Itgax | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-X precursor (Leukocyte adhesion glycoprotein p150,95 alpha chain) (Leukocyte adhesion receptor p150,95) (CD11c antigen). | |||||
|
TPH2_MOUSE
|
||||||
| NC score | 0.013719 (rank : 19) | θ value | 1.38821 (rank : 16) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CGV2 | Gene names | Tph2, Ntph | |||
|
Domain Architecture |

|
|||||
| Description | Tryptophan 5-hydroxylase 2 (EC 1.14.16.4) (Tryptophan 5-monooxygenase 2) (Neuronal tryptophan hydroxylase). | |||||
|
IRK15_HUMAN
|
||||||
| NC score | 0.011500 (rank : 20) | θ value | 1.81305 (rank : 17) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99712, O00564, Q96L28, Q99446 | Gene names | KCNJ15, KCNJ14 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-sensitive inward rectifier potassium channel 15 (Potassium channel, inwardly rectifying subfamily J member 15) (Inward rectifier K(+) channel Kir4.2) (Kir1.3). | |||||
|
IRK15_MOUSE
|
||||||
| NC score | 0.011500 (rank : 21) | θ value | 1.81305 (rank : 18) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88932, Q9JK34 | Gene names | Kcnj15 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-sensitive inward rectifier potassium channel 15 (Potassium channel, inwardly rectifying subfamily J member 15) (Inward rectifier K(+) channel Kir4.2). | |||||
|
AFF2_MOUSE
|
||||||
| NC score | 0.008990 (rank : 22) | θ value | 5.27518 (rank : 20) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O55112 | Gene names | Aff2, Fmr2, Ox19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation protein 2 homolog) (Protein FMR-2) (FMR2P) (Protein Ox19). | |||||
|
GLGB_MOUSE
|
||||||
| NC score | 0.008974 (rank : 23) | θ value | 8.99809 (rank : 26) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D6Y9 | Gene names | Gbe1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 1,4-alpha-glucan branching enzyme (EC 2.4.1.18) (Glycogen branching enzyme) (Brancher enzyme). | |||||
|
ECEL1_MOUSE
|
||||||
| NC score | 0.006895 (rank : 24) | θ value | 8.99809 (rank : 25) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JMI0 | Gene names | Ecel1, Dine, Xce | |||
|
Domain Architecture |

|
|||||
| Description | Endothelin-converting enzyme-like 1 (EC 3.4.24.-) (Xce protein) (Damage-induced neuronal endopeptidase). | |||||
|
ECEL1_HUMAN
|
||||||
| NC score | 0.006885 (rank : 25) | θ value | 8.99809 (rank : 24) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95672, Q45UD9, Q53RF9, Q6UW86, Q86TH4, Q9NY95 | Gene names | ECEL1, XCE | |||
|
Domain Architecture |

|
|||||
| Description | Endothelin-converting enzyme-like 1 (EC 3.4.24.-) (Xce protein). | |||||
|
UBP25_HUMAN
|
||||||
| NC score | 0.006666 (rank : 26) | θ value | 6.88961 (rank : 23) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UHP3, Q9H9W1 | Gene names | USP25, USP21 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 25 (EC 3.1.2.15) (Ubiquitin thioesterase 25) (Ubiquitin-specific-processing protease 25) (Deubiquitinating enzyme 25) (USP on chromosome 21). | |||||
|
KIF3C_MOUSE
|
||||||
| NC score | 0.006390 (rank : 27) | θ value | 6.88961 (rank : 22) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 468 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O35066, O35229 | Gene names | Kif3c | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF3C. | |||||
|
KIF3C_HUMAN
|
||||||
| NC score | 0.006353 (rank : 28) | θ value | 6.88961 (rank : 21) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 478 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O14782, O43544 | Gene names | KIF3C | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF3C. | |||||