Please be patient as the page loads
|
RHG18_HUMAN
|
||||||
| SwissProt Accessions | Q8N392, Q58EZ3, Q6P679, Q6PJD7, Q96S64 | Gene names | ARHGAP18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho-GTPase-activating protein 18 (MacGAP). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
RHG18_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | Q8N392, Q58EZ3, Q6P679, Q6PJD7, Q96S64 | Gene names | ARHGAP18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho-GTPase-activating protein 18 (MacGAP). | |||||
|
RHG18_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.987326 (rank : 2) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q8K0Q5, Q8BP03, Q8R196 | Gene names | Arhgap18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho-GTPase-activating protein 18. | |||||
|
RHG07_MOUSE
|
||||||
| θ value | 1.09232e-21 (rank : 3) | NC score | 0.758744 (rank : 12) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9R0Z9, Q8R541 | Gene names | Dlc1, Arhgap7, Stard12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 7 (Rho-type GTPase-activating protein 7) (Deleted in liver cancer 1 protein homolog) (Dlc-1) (StAR-related lipid transfer protein 12) (StARD12) (START domain-containing protein 12). | |||||
|
RHG07_HUMAN
|
||||||
| θ value | 9.24701e-21 (rank : 4) | NC score | 0.753927 (rank : 13) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q96QB1, O14868, O43199, Q9C0E0 | Gene names | DLC1, ARHGAP7, KIAA1723, STARD12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 7 (Rho-type GTPase-activating protein 7) (Deleted in liver cancer 1 protein) (Dlc-1) (HP protein) (StAR-related lipid transfer protein 12) (StARD12) (START domain-containing protein 12). | |||||
|
MYO9B_HUMAN
|
||||||
| θ value | 8.10077e-17 (rank : 5) | NC score | 0.407466 (rank : 58) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q13459, O75314, Q9NUJ2, Q9UHN0 | Gene names | MYO9B, MYR5 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9B (Myosin IXb) (Unconventional myosin-9b). | |||||
|
MYO9B_MOUSE
|
||||||
| θ value | 1.38178e-16 (rank : 6) | NC score | 0.413054 (rank : 57) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9QY06, Q9QY07, Q9QY08, Q9QY09 | Gene names | Myo9b, Myr5 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9B (Myosin IXb) (Unconventional myosin-9b). | |||||
|
RBP1_MOUSE
|
||||||
| θ value | 1.38178e-16 (rank : 7) | NC score | 0.724453 (rank : 19) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 493 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q62172, Q9CRE5 | Gene names | Ralbp1, Rip1 | |||
|
Domain Architecture |
|
|||||
| Description | RalA-binding protein 1 (RalBP1) (Ral-interacting protein 1) (Dinitrophenyl S-glutathione ATPase) (DNP-SG ATPase). | |||||
|
RHG12_MOUSE
|
||||||
| θ value | 1.38178e-16 (rank : 8) | NC score | 0.767706 (rank : 7) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q8C0D4, Q8BVP8 | Gene names | Arhgap12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
RHG12_HUMAN
|
||||||
| θ value | 1.80466e-16 (rank : 9) | NC score | 0.771914 (rank : 6) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q8IWW6, Q86UB3, Q8IWW7, Q8N3L1, Q9NT76 | Gene names | ARHGAP12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
RBP1_HUMAN
|
||||||
| θ value | 3.07829e-16 (rank : 10) | NC score | 0.704603 (rank : 28) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q15311 | Gene names | RALBP1, RLIP1, RLIP76 | |||
|
Domain Architecture |
|
|||||
| Description | RalA-binding protein 1 (RalBP1) (Ral-interacting protein 1) (76 kDa Ral-interacting protein) (Dinitrophenyl S-glutathione ATPase) (DNP-SG ATPase). | |||||
|
STA13_HUMAN
|
||||||
| θ value | 8.95645e-16 (rank : 11) | NC score | 0.715308 (rank : 20) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9Y3M8, Q5HYH1, Q5TAE3, Q6UN61, Q86TP6, Q86WQ3, Q86XT1 | Gene names | STARD13, DLC2, GT650 | |||
|
Domain Architecture |

|
|||||
| Description | StAR-related lipid transfer protein 13 (StARD13) (START domain- containing protein 13) (46H23.2) (Deleted in liver cancer protein 2) (Rho GTPase-activating protein). | |||||
|
ABR_HUMAN
|
||||||
| θ value | 2.60593e-15 (rank : 12) | NC score | 0.693869 (rank : 30) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q12979, Q13693, Q13694 | Gene names | ABR | |||
|
Domain Architecture |

|
|||||
| Description | Active breakpoint cluster region-related protein. | |||||
|
3BP1_HUMAN
|
||||||
| θ value | 6.41864e-14 (rank : 13) | NC score | 0.753504 (rank : 14) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9Y3L3, Q6IBZ2, Q6ZVL9, Q96HQ5, Q9NSQ9 | Gene names | SH3BP1 | |||
|
Domain Architecture |
|
|||||
| Description | SH3 domain-binding protein 1 (3BP-1). | |||||
|
STA13_MOUSE
|
||||||
| θ value | 6.41864e-14 (rank : 14) | NC score | 0.704890 (rank : 27) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q923Q2, Q8K369 | Gene names | Stard13 | |||
|
Domain Architecture |

|
|||||
| Description | StAR-related lipid transfer protein 13 (StARD13) (START domain- containing protein 13). | |||||
|
GMIP_HUMAN
|
||||||
| θ value | 1.86753e-13 (rank : 15) | NC score | 0.763814 (rank : 10) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9P107 | Gene names | GMIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GEM-interacting protein (GMIP). | |||||
|
GMIP_MOUSE
|
||||||
| θ value | 4.16044e-13 (rank : 16) | NC score | 0.765847 (rank : 9) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q6PGG2, Q6P9S3 | Gene names | Gmip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GEM-interacting protein (GMIP). | |||||
|
RHG06_HUMAN
|
||||||
| θ value | 7.09661e-13 (rank : 17) | NC score | 0.778129 (rank : 4) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | O43182, O43437, Q9P1B3, Q9UK81, Q9UK82 | Gene names | ARHGAP6, RHOGAP6 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 6 (Rho-type GTPase-activating protein RhoGAPX-1). | |||||
|
RGAP1_HUMAN
|
||||||
| θ value | 9.26847e-13 (rank : 18) | NC score | 0.762627 (rank : 11) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9H0H5, Q6PJ26, Q9NWN2, Q9P250, Q9P2W2 | Gene names | RACGAP1, KIAA1478, MGCRACGAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rac GTPase-activating protein 1 (MgcRacGAP). | |||||
|
RHG26_HUMAN
|
||||||
| θ value | 9.26847e-13 (rank : 19) | NC score | 0.682144 (rank : 33) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9UNA1, O75117, Q9BYS6, Q9BYS7, Q9UJ00 | Gene names | ARHGAP26, GRAF, KIAA0621, OPHN1L | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 26 (Oligophrenin-1-like protein) (GTPase regulator associated with focal adhesion kinase). | |||||
|
STAR8_HUMAN
|
||||||
| θ value | 2.0648e-12 (rank : 20) | NC score | 0.697666 (rank : 29) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q92502, Q5JST0 | Gene names | STARD8, KIAA0189 | |||
|
Domain Architecture |

|
|||||
| Description | StAR-related lipid transfer protein 8 (StARD8) (START domain- containing protein 8). | |||||
|
CEND3_HUMAN
|
||||||
| θ value | 2.69671e-12 (rank : 21) | NC score | 0.578270 (rank : 54) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q8WWN8 | Gene names | CENTD3, ARAP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-delta 3 (Cnt-d3) (Arf-GAP, Rho-GAP, ankyrin repeat and pleckstrin homology domain-containing protein 3). | |||||
|
RHG06_MOUSE
|
||||||
| θ value | 2.69671e-12 (rank : 22) | NC score | 0.776791 (rank : 5) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | O54834, Q9QZL8 | Gene names | Arhgap6 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 6 (Rho-type GTPase-activating protein RhoGAPX-1). | |||||
|
CHIO_MOUSE
|
||||||
| θ value | 3.52202e-12 (rank : 23) | NC score | 0.714100 (rank : 21) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q80XD1, Q9D9W2, Q9ER57 | Gene names | Chn2, Arhgap3, Bch | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-chimaerin (Beta-chimerin) (Rho-GTPase-activating protein 3). | |||||
|
STAR8_MOUSE
|
||||||
| θ value | 3.52202e-12 (rank : 24) | NC score | 0.706411 (rank : 26) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8K031, Q3UZC7, Q6A0A8, Q6P5E0, Q8R3X8 | Gene names | Stard8, Kiaa0189 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | StAR-related lipid transfer protein 8 (StARD8) (START domain- containing protein 8). | |||||
|
CEND3_MOUSE
|
||||||
| θ value | 4.59992e-12 (rank : 25) | NC score | 0.586559 (rank : 52) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q8R5G7, Q5DTN4, Q6NVF1, Q8R5G6 | Gene names | Centd3, Arap3, Drag1, Kiaa4097 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-delta 3 (Cnt-d3) (Arf-GAP, Rho-GAP, ankyrin repeat and pleckstrin homology domain-containing protein 3) (Dual specificity Rho- and Arf-GTPase-activating protein 1). | |||||
|
CHIO_HUMAN
|
||||||
| θ value | 7.84624e-12 (rank : 26) | NC score | 0.680908 (rank : 35) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P52757, Q75MM2 | Gene names | CHN2, ARHGAP3, BCH | |||
|
Domain Architecture |
|
|||||
| Description | Beta-chimaerin (Beta-chimerin) (Rho-GTPase-activating protein 3). | |||||
|
RHG01_HUMAN
|
||||||
| θ value | 2.28291e-11 (rank : 27) | NC score | 0.744148 (rank : 16) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q07960 | Gene names | ARHGAP1, CDC42GAP, RHOGAP1 | |||
|
Domain Architecture |
|
|||||
| Description | Rho-GTPase-activating protein 1 (GTPase-activating protein rhoOGAP) (Rho-related small GTPase protein activator) (CDC42 GTPase-activating protein) (p50-RhoGAP). | |||||
|
RHG09_HUMAN
|
||||||
| θ value | 2.28291e-11 (rank : 28) | NC score | 0.779803 (rank : 3) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9BRR9, Q8TCJ3, Q8WYR0, Q96EZ2, Q96S74 | Gene names | ARHGAP9 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 9. | |||||
|
CHIN_HUMAN
|
||||||
| θ value | 2.98157e-11 (rank : 29) | NC score | 0.681554 (rank : 34) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P15882, Q53SD6, Q53SH5, Q96FB0 | Gene names | CHN1, ARHGAP2, CHN | |||
|
Domain Architecture |
|
|||||
| Description | N-chimaerin (NC) (N-chimerin) (Alpha chimerin) (A-chimaerin) (Rho- GTPase-activating protein 2). | |||||
|
3BP1_MOUSE
|
||||||
| θ value | 5.08577e-11 (rank : 30) | NC score | 0.767309 (rank : 8) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P55194, Q99KK8 | Gene names | Sh3bp1, 3bp1 | |||
|
Domain Architecture |
|
|||||
| Description | SH3 domain-binding protein 1 (3BP-1). | |||||
|
RHG25_HUMAN
|
||||||
| θ value | 8.67504e-11 (rank : 31) | NC score | 0.663949 (rank : 39) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 550 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | P42331, Q8IXQ2 | Gene names | ARHGAP25, KIAA0053 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
CHIN_MOUSE
|
||||||
| θ value | 1.133e-10 (rank : 32) | NC score | 0.680289 (rank : 36) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q91V57, Q3UGY3, Q7TQE5, Q8BWU6, Q9D9B3 | Gene names | Chn1, Arhgap2 | |||
|
Domain Architecture |
|
|||||
| Description | N-chimaerin (NC) (N-chimerin) (Alpha chimerin) (A-chimaerin) (Rho- GTPase-activating protein 2). | |||||
|
GRLF1_HUMAN
|
||||||
| θ value | 1.133e-10 (rank : 33) | NC score | 0.729330 (rank : 17) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9NRY4, Q14452, Q9C0E1 | Gene names | GRLF1, GRF1, KIAA1722 | |||
|
Domain Architecture |

|
|||||
| Description | Glucocorticoid receptor DNA-binding factor 1 (Glucocorticoid receptor repression factor 1) (GRF-1) (Rho GAP p190A) (p190-A). | |||||
|
K1688_MOUSE
|
||||||
| θ value | 1.133e-10 (rank : 34) | NC score | 0.593796 (rank : 51) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P59281, Q69ZD4, Q6P9R6 | Gene names | Kiaa1688, D15Wsu169e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1688. | |||||
|
OPHN1_MOUSE
|
||||||
| θ value | 1.133e-10 (rank : 35) | NC score | 0.709029 (rank : 24) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q99J31, Q544K7 | Gene names | Ophn1 | |||
|
Domain Architecture |

|
|||||
| Description | Oligophrenin 1. | |||||
|
TCGAP_HUMAN
|
||||||
| θ value | 1.133e-10 (rank : 36) | NC score | 0.630850 (rank : 44) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 872 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | O14559, O14552, O14560, Q6ZSP6, Q96CP3, Q9NT23 | Gene names | SNX26, TCGAP | |||
|
Domain Architecture |

|
|||||
| Description | TC10/CDC42 GTPase-activating protein (Sorting nexin-26). | |||||
|
TCGAP_MOUSE
|
||||||
| θ value | 1.133e-10 (rank : 37) | NC score | 0.710218 (rank : 23) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q80YF9, Q6P7W6 | Gene names | Snx26, Tcgap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TC10/CDC42 GTPase-activating protein (Sorting nexin-26). | |||||
|
K1688_HUMAN
|
||||||
| θ value | 1.47974e-10 (rank : 38) | NC score | 0.580052 (rank : 53) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9C0H5 | Gene names | KIAA1688 | |||
|
Domain Architecture |

|
|||||
| Description | Protein KIAA1688. | |||||
|
RHG08_MOUSE
|
||||||
| θ value | 1.47974e-10 (rank : 39) | NC score | 0.724505 (rank : 18) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9CXP4, Q99JY7 | Gene names | Arhgap8 | |||
|
Domain Architecture |
|
|||||
| Description | Rho-GTPase-activating protein 8. | |||||
|
CEND2_HUMAN
|
||||||
| θ value | 1.9326e-10 (rank : 40) | NC score | 0.604772 (rank : 50) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q96P48, O94879, Q59FI7, Q6PHS3, Q8WU51, Q96HP6, Q96L71 | Gene names | CENTD2, ARAP1, KIAA0782 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-delta 2 (Cnt-d2) (Arf-GAP, Rho-GAP, ankyrin repeat and pleckstrin homology domain-containing protein 1). | |||||
|
OPHN1_HUMAN
|
||||||
| θ value | 1.9326e-10 (rank : 41) | NC score | 0.708284 (rank : 25) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | O60890, Q5JQ81, Q8WX47 | Gene names | OPHN1 | |||
|
Domain Architecture |

|
|||||
| Description | Oligophrenin 1. | |||||
|
RHG25_MOUSE
|
||||||
| θ value | 1.9326e-10 (rank : 42) | NC score | 0.682555 (rank : 32) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q8BYW1, Q2VPR2, Q3TE04, Q3TVX0, Q3UVB7, Q6A0E0, Q8BX98 | Gene names | Arhgap25, Kiaa0053 | |||
|
Domain Architecture |
|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
RGAP1_MOUSE
|
||||||
| θ value | 4.30538e-10 (rank : 43) | NC score | 0.750594 (rank : 15) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9WVM1, Q3THR5, Q3TI41, Q3TM81 | Gene names | Racgap1, Mgcracgap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rac GTPase-activating protein 1 (MgcRacGAP). | |||||
|
RHG08_HUMAN
|
||||||
| θ value | 1.63604e-09 (rank : 44) | NC score | 0.714054 (rank : 22) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9NSG0, O75983, O95695, Q96RW1, Q96RW2, Q9HA49, Q9HC46, Q9NVX8, Q9NXL1, Q9UH20 | Gene names | ARHGAP8 | |||
|
Domain Architecture |
|
|||||
| Description | Rho-GTPase-activating protein 8. | |||||
|
BCR_HUMAN
|
||||||
| θ value | 6.21693e-09 (rank : 45) | NC score | 0.671531 (rank : 38) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P11274, P78501, Q12842 | Gene names | BCR, BCR1 | |||
|
Domain Architecture |

|
|||||
| Description | Breakpoint cluster region protein (EC 2.7.11.1) (NY-REN-26 antigen). | |||||
|
CE005_MOUSE
|
||||||
| θ value | 6.21693e-09 (rank : 46) | NC score | 0.618616 (rank : 47) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q8K2H3 | Gene names | ||||
|
Domain Architecture |
|
|||||
| Description | Protein C5orf5 homolog. | |||||
|
RHG05_MOUSE
|
||||||
| θ value | 8.11959e-09 (rank : 47) | NC score | 0.686918 (rank : 31) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | P97393 | Gene names | Arhgap5, Rhogap5 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 5 (p190-B). | |||||
|
CE005_HUMAN
|
||||||
| θ value | 1.06045e-08 (rank : 48) | NC score | 0.620193 (rank : 46) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9NYF5, Q6PGQ2, Q9P0I7 | Gene names | C5orf5 | |||
|
Domain Architecture |
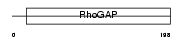
|
|||||
| Description | Protein C5orf5 (GAP-like protein N61). | |||||
|
RHG04_HUMAN
|
||||||
| θ value | 4.0297e-08 (rank : 49) | NC score | 0.651206 (rank : 42) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P98171, Q14144 | Gene names | ARHGAP4, KIAA0131, RGC1, RHOGAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 4 (Rho-GAP hematopoietic protein C1) (p115). | |||||
|
RHG05_HUMAN
|
||||||
| θ value | 4.0297e-08 (rank : 50) | NC score | 0.674161 (rank : 37) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q13017 | Gene names | ARHGAP5, RHOGAP5 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 5 (p190-B). | |||||
|
SRGP2_HUMAN
|
||||||
| θ value | 4.0297e-08 (rank : 51) | NC score | 0.632943 (rank : 43) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | O43295, Q8IX13, Q8IZV8 | Gene names | SRGAP3, ARHGAP14, KIAA0411, MEGAP, SRGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 3 (srGAP3) (srGAP2) (WAVE- associated Rac GTPase-activating protein) (WRP) (Mental disorder- associated GAP) (Rho-GTPase-activating protein 14). | |||||
|
SRGP2_MOUSE
|
||||||
| θ value | 8.97725e-08 (rank : 52) | NC score | 0.623985 (rank : 45) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q812A2, Q80U09, Q8BKP4, Q8BLD0, Q925I2 | Gene names | Srgap3, Arhgap14, Kiaa0411, Srgap2 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 3 (srGAP3) (srGAP2) (WAVE- associated Rac GTPase-activating protein) (WRP) (Rho-GTPase-activating protein 14). | |||||
|
FNBP2_HUMAN
|
||||||
| θ value | 1.99992e-07 (rank : 53) | NC score | 0.615601 (rank : 48) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | O75044 | Gene names | SRGAP2, FNBP2, KIAA0456 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 2 (srGAP2) (Formin-binding protein 2). | |||||
|
CEND1_MOUSE
|
||||||
| θ value | 3.41135e-07 (rank : 54) | NC score | 0.506620 (rank : 55) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q8BZ05, Q80TX2, Q8C3T2, Q8VEL6 | Gene names | Centd1, Kiaa0580 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-delta 1 (Cnt-d1). | |||||
|
CEND1_HUMAN
|
||||||
| θ value | 9.92553e-07 (rank : 55) | NC score | 0.486114 (rank : 56) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q8WZ64, Q4W5D2, Q7Z2L5, Q96L70, Q96P49, Q9Y4E4 | Gene names | CENTD1, ARAP2, KIAA0580 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-delta 1 (Cnt-d1) (Arf-GAP, Rho-GAP, ankyrin repeat and pleckstrin homology domain-containing protein 2) (PARX protein). | |||||
|
SRGP1_HUMAN
|
||||||
| θ value | 3.77169e-06 (rank : 56) | NC score | 0.610653 (rank : 49) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q7Z6B7, Q9H8A3, Q9P2P2 | Gene names | SRGAP1, ARHGAP13, KIAA1304 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 1 (srGAP1) (Rho-GTPase- activating protein 13). | |||||
|
TAGAP_MOUSE
|
||||||
| θ value | 9.29e-05 (rank : 57) | NC score | 0.656129 (rank : 40) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8K2L9, Q920D6 | Gene names | Tagap, Tagap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-cell activation Rho GTPase-activating protein (T-cell activation GTPase-activating protein). | |||||
|
TAGAP_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 58) | NC score | 0.651433 (rank : 41) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q8N103, Q2NKM8, Q8NI40, Q96KZ2, Q96QA2 | Gene names | TAGAP, TAGAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-cell activation Rho GTPase-activating protein (T-cell activation GTPase-activating protein). | |||||
|
P85A_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 59) | NC score | 0.319128 (rank : 59) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P27986 | Gene names | PIK3R1, GRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit alpha (PI3-kinase p85-subunit alpha) (PtdIns-3-kinase p85-alpha) (PI3K). | |||||
|
I5P2_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 60) | NC score | 0.305335 (rank : 61) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P32019, Q5VSG9, Q5VSH0, Q5VSH1, Q658Q5, Q6P6D4, Q6PD53, Q86YE1 | Gene names | INPP5B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Type II inositol-1,4,5-trisphosphate 5-phosphatase precursor (EC 3.1.3.36) (Phosphoinositide 5-phosphatase) (5PTase) (75 kDa inositol polyphosphate-5-phosphatase). | |||||
|
FACD2_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 61) | NC score | 0.066352 (rank : 70) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BXW9, Q2LA86, Q69YP9, Q6PJN7, Q9BQ06, Q9H9T9 | Gene names | FANCD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group D2 protein (Protein FACD2). | |||||
|
RABE1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 62) | NC score | 0.025276 (rank : 90) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1104 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q15276, O95369, Q8IVX3 | Gene names | RABEP1, RABPT5, RABPT5A | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 1 (Rabaptin-5) (Rabaptin-5alpha) (Rabaptin-4) (NY-REN-17 antigen). | |||||
|
ROCK1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 63) | NC score | 0.006144 (rank : 111) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 2000 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q13464 | Gene names | ROCK1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 1 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 1) (p160 ROCK-1) (p160ROCK) (NY-REN-35 antigen). | |||||
|
FLIP1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 64) | NC score | 0.016137 (rank : 97) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q7Z7B0, Q5VUL6, Q86TC3, Q8N8B9, Q96SK6, Q9NVI8, Q9ULE5 | Gene names | FILIP1, KIAA1275 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
5NT1B_MOUSE
|
||||||
| θ value | 1.06291 (rank : 65) | NC score | 0.022472 (rank : 92) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91YE9, Q91Y48 | Gene names | Nt5c1b, Airp | |||
|
Domain Architecture |

|
|||||
| Description | Cytosolic 5'-nucleotidase 1B (EC 3.1.3.5) (Cytosolic 5'-nucleotidase IB) (cN1B) (cN-IB) (Autoimmune infertility-related protein). | |||||
|
DAPK3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 66) | NC score | -0.002577 (rank : 117) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 945 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43293 | Gene names | DAPK3, ZIPK | |||
|
Domain Architecture |

|
|||||
| Description | Death-associated protein kinase 3 (EC 2.7.11.1) (DAP kinase 3) (DAP- like kinase) (Dlk) (ZIP-kinase). | |||||
|
IF4G1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 67) | NC score | 0.017646 (rank : 96) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q04637, O43177, O95066, Q5HYG0, Q6ZN21, Q8N102 | Gene names | EIF4G1, EIF4G, EIF4GI | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1) (p220). | |||||
|
DBLOH_MOUSE
|
||||||
| θ value | 1.81305 (rank : 68) | NC score | 0.030717 (rank : 89) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JIQ3, Q9CZD1, Q9DCD3 | Gene names | Diablo, Smac | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diablo homolog, mitochondrial precursor (Second mitochondria-derived activator of caspase) (Smac protein) (Direct IAP-binding protein with low pI). | |||||
|
GOGA4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 69) | NC score | 0.014202 (rank : 98) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
KI21B_HUMAN
|
||||||
| θ value | 1.81305 (rank : 70) | NC score | 0.007173 (rank : 108) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1017 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O75037, Q5T4J3 | Gene names | KIF21B, KIAA0449 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B. | |||||
|
MCM4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 71) | NC score | 0.018984 (rank : 95) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P49717, O89056 | Gene names | Mcm4, Cdc21, Mcmd4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM4 (CDC21 homolog) (P1-CDC21). | |||||
|
SMCA4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 72) | NC score | 0.008484 (rank : 106) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P51532, O95052, Q9HBD3 | Gene names | SMARCA4, BRG1, SNF2B, SNF2L4 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L4 (EC 3.6.1.-) (ATP- dependent helicase SMARCA4) (SNF2-beta) (BRG-1 protein) (Mitotic growth and transcription activator) (Brahma protein homolog 1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 4). | |||||
|
KPBB_MOUSE
|
||||||
| θ value | 4.03905 (rank : 73) | NC score | 0.010172 (rank : 104) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7TSH2 | Gene names | Phkb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphorylase b kinase regulatory subunit beta (Phosphorylase kinase subunit beta). | |||||
|
MCM4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 74) | NC score | 0.013549 (rank : 100) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P33991, Q8NEH1, Q99658 | Gene names | MCM4, CDC21 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM4 (CDC21 homolog) (P1-CDC21). | |||||
|
NIN_HUMAN
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | 0.022103 (rank : 93) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1416 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8N4C6, Q6P0P6, Q9BWU6, Q9C012, Q9C013, Q9C014, Q9H5I6, Q9HAT7, Q9HBY5, Q9HCK7, Q9UH61 | Gene names | NIN, KIAA1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein (hNinein) (Glycogen synthase kinase 3 beta-interacting protein) (GSK3B-interacting protein). | |||||
|
RAE2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 76) | NC score | 0.013703 (rank : 99) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P26374, Q9H1Y4 | Gene names | CHML, REP2 | |||
|
Domain Architecture |

|
|||||
| Description | Rab proteins geranylgeranyltransferase component A 2 (Rab escort protein 2) (REP-2) (Choroideraemia-like protein). | |||||
|
RBL1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 77) | NC score | 0.008603 (rank : 105) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q64701 | Gene names | Rbl1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoblastoma-like protein 1 (107 kDa retinoblastoma-associated protein) (PRB1) (P107). | |||||
|
SMRCD_MOUSE
|
||||||
| θ value | 5.27518 (rank : 78) | NC score | 0.006536 (rank : 109) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q04692 | Gene names | Smarcad1, Etl1, Kiaa1122 | |||
|
Domain Architecture |

|
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A containing DEAD/H box 1 (EC 3.6.1.-) (ATP- dependent helicase SMARCAD1) (Enhancer trap locus homolog 1) (Etl-1). | |||||
|
DBLOH_HUMAN
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.023731 (rank : 91) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NR28, Q96LV0, Q9BT11, Q9HAV6 | Gene names | DIABLO, SMAC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diablo homolog, mitochondrial precursor (Second mitochondria-derived activator of caspase) (Smac protein) (Direct IAP-binding protein with low pI). | |||||
|
PAR14_MOUSE
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.008333 (rank : 107) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q2EMV9, Q571C6, Q8BN35, Q8VDT6 | Gene names | Parp14, Kiaa1268 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 14 (EC 2.4.2.30) (PARP-14) (Collaborator of STAT6) (CoaSt6). | |||||
|
RABE1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.019800 (rank : 94) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O35551, Q99L08, Q9CRP3, Q9EQF9, Q9JI94 | Gene names | Rabep1, Rab5ep, Rabpt5, Rabpt5a | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 1 (Rabaptin-5) (Rabaptin-5alpha). | |||||
|
RPGF6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.005874 (rank : 112) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TEU7, Q8NI21, Q8TEU6, Q96PC1 | Gene names | RAPGEF6, PDZGEF2 | |||
|
Domain Architecture |

|
|||||
| Description | Rap guanine nucleotide exchange factor 6 (PDZ domain-containing guanine nucleotide exchange factor 2) (PDZ-GEF2) (RA-GEF-2). | |||||
|
WNK4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | -0.002275 (rank : 116) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1043 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96J92, Q8N8X3, Q8N8Z2, Q96DT8, Q9BYS5 | Gene names | WNK4, PRKWNK4 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK4 (EC 2.7.11.1) (Protein kinase with no lysine 4) (Protein kinase, lysine-deficient 4). | |||||
|
COG8_MOUSE
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.011591 (rank : 103) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JJA2 | Gene names | Cog8 | |||
|
Domain Architecture |
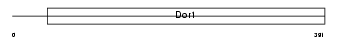
|
|||||
| Description | Conserved oligomeric Golgi complex component 8. | |||||
|
GOGA5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.013522 (rank : 101) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1434 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9QYE6, O88317, Q3TGE7, Q3U6S5, Q3UUF9 | Gene names | Golga5, Retii | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (Sumiko protein) (Ret-II protein). | |||||
|
ITB2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.001116 (rank : 115) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P05107, Q16418 | Gene names | ITGB2, CD18 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-2 precursor (Cell surface adhesion glycoproteins LFA- 1/CR3/p150,95 subunit beta) (Complement receptor C3 subunit beta) (CD18 antigen). | |||||
|
KI21B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.006229 (rank : 110) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 997 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9QXL1, P97424 | Gene names | Kif21b, Kif6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B (Kinesin-like protein KIF6). | |||||
|
KLHL6_MOUSE
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.001420 (rank : 114) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6V595 | Gene names | Klhl6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kelch-like protein 6. | |||||
|
PRAX_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.004814 (rank : 113) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BXM0, Q9BXL9, Q9HCF2 | Gene names | PRX, KIAA1620 | |||
|
Domain Architecture |
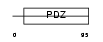
|
|||||
| Description | Periaxin. | |||||
|
TRIPB_HUMAN
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.012616 (rank : 102) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1587 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q15643, O14689, O15154, O95949 | Gene names | TRIP11, CEV14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid receptor-interacting protein 11 (TRIP-11) (Golgi-associated microtubule-binding protein 210) (GMAP-210) (Trip230) (Clonal evolution-related gene on chromosome 14). | |||||
|
ATCAY_HUMAN
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.050505 (rank : 88) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86WG3, Q8NAQ2, Q8TAQ3, Q96HC6, Q96JF5 | Gene names | ATCAY, KIAA1872 | |||
|
Domain Architecture |

|
|||||
| Description | Caytaxin (Ataxia Cayman type protein) (BNIP-H). | |||||
|
ATCAY_MOUSE
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.050965 (rank : 87) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BHE3, Q3TR94 | Gene names | Atcay | |||
|
Domain Architecture |
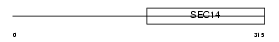
|
|||||
| Description | Caytaxin. | |||||
|
BNIP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.053614 (rank : 77) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q12982 | Gene names | BNIP2, NIP2 | |||
|
Domain Architecture |
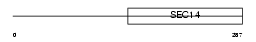
|
|||||
| Description | BCL2/adenovirus E1B 19 kDa protein-interacting protein 2. | |||||
|
BNIP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.053528 (rank : 78) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O54940 | Gene names | Bnip2, Nip2l | |||
|
Domain Architecture |
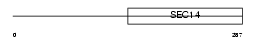
|
|||||
| Description | BCL2/adenovirus E1B 19 kDa protein-interacting protein 2. | |||||
|
CENA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.051319 (rank : 85) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75689 | Gene names | CENTA1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-alpha 1 (Putative MAPK-activating protein PM25). | |||||
|
CENA2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.054774 (rank : 73) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NPF8, Q8N4Q6, Q96SD5 | Gene names | CENTA2 | |||
|
Domain Architecture |
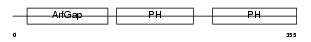
|
|||||
| Description | Centaurin-alpha 2. | |||||
|
CENB1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.070211 (rank : 68) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q15027 | Gene names | CENTB1, KIAA0050 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-beta 1 (Cnt-b1). | |||||
|
CENB2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.076308 (rank : 65) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q15057, Q9UQR3 | Gene names | CENTB2, KIAA0041 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-beta 2 (Cnt-b2). | |||||
|
CENB5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.070503 (rank : 67) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96P50, Q9BSR9, Q9C0E7 | Gene names | CENTB5, KIAA1716 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-beta 5 (Cnt-b5). | |||||
|
DDEF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.053901 (rank : 76) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 599 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O43150 | Gene names | DDEF2, KIAA0400 | |||
|
Domain Architecture |

|
|||||
| Description | Development and differentiation-enhancing factor 2 (Pyk2 C-terminus- associated protein) (PAP) (Paxillin-associated protein with ARFGAP activity 3) (PAG3). | |||||
|
DDEF2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.052664 (rank : 80) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 625 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7SIG6, Q501K1, Q66JN2 | Gene names | Ddef2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Development and differentiation-enhancing factor 2 (Pyk2 C-terminus- associated protein) (PAP) (Paxillin-associated protein with ARFGAP activity 3) (PAG3). | |||||
|
DDFL1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.052139 (rank : 82) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 353 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8TDY4, Q6P9F4, Q86UY1, Q9NXK2 | Gene names | DDEFL1, UPLC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Development and differentiation-enhancing factor-like 1 (Protein up- regulated in liver cancer 1). | |||||
|
FA13A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.065528 (rank : 72) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O94988, Q8NBA3 | Gene names | FAM13A1, KIAA0914 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13A1. | |||||
|
FA13A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.066038 (rank : 71) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BGI4, Q8BZ91 | Gene names | Fam13a1, Precm1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13A1 (Precm1 protein). | |||||
|
FA13C_HUMAN
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.051033 (rank : 86) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NE31, Q99787 | Gene names | FAM13C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13C1. | |||||
|
GRP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.068744 (rank : 69) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8IV61, O94931 | Gene names | RASGRP3, GRP3, KIAA0846 | |||
|
Domain Architecture |

|
|||||
| Description | RAS guanyl-releasing protein 3 (Calcium and DAG-regulated guanine nucleotide exchange factor III) (Guanine nucleotide exchange factor for Rap1). | |||||
|
MYO10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.052015 (rank : 84) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 920 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9HD67, Q9NYM7, Q9P110, Q9P111, Q9UHF6 | Gene names | MYO10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin X). | |||||
|
OCRL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.272443 (rank : 62) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q01968, O60800, Q15684, Q15774, Q5JQF1, Q5JQF2, Q9UJG5, Q9UMA5 | Gene names | OCRL, INPP5F, OCRL1 | |||
|
Domain Architecture |

|
|||||
| Description | Inositol polyphosphate 5-phosphatase OCRL-1 (EC 3.1.3.36) (Lowe oculocerebrorenal syndrome protein). | |||||
|
P85A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.306913 (rank : 60) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P26450 | Gene names | Pik3r1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit alpha (PI3-kinase p85-subunit alpha) (PtdIns-3-kinase p85-alpha) (PI3K). | |||||
|
P85B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.073961 (rank : 66) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O00459, Q9UPH9 | Gene names | PIK3R2 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit beta (PI3-kinase p85- subunit beta) (PtdIns-3-kinase p85-beta). | |||||
|
PCTL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.093774 (rank : 64) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y365, O60532 | Gene names | STARD10, SDCCAG28 | |||
|
Domain Architecture |
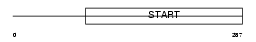
|
|||||
| Description | PCTP-like protein (PCTP-L) (StAR-related lipid transfer protein 10) (StARD10) (START domain-containing protein 10) (Serologically defined colon cancer antigen 28) (Antigen NY-CO-28). | |||||
|
PCTL_MOUSE
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.094861 (rank : 63) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9JMD3 | Gene names | Stard10, Pctpl, Sdccag28, Sdccagg28 | |||
|
Domain Architecture |
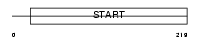
|
|||||
| Description | PCTP-like protein (PCTP-L) (StAR-related lipid transfer protein 10) (StARD10) (START domain-containing protein 10) (Serologically defined colon cancer antigen 28 homolog). | |||||
|
PKHA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.053131 (rank : 79) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9HB21, Q9BVK0 | Gene names | PLEKHA1, TAPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 1 (Tandem PH domain-containing protein 1) (TAPP-1). | |||||
|
SMAP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.052559 (rank : 81) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IYB5, Q53H70, Q5SYQ2, Q6PK24, Q8NDH4, Q96L38, Q96L39, Q9H8X4 | Gene names | SMAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stromal membrane-associated protein 1. | |||||
|
SMAP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.052022 (rank : 83) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q91VZ6, Q497I6, Q68EF3 | Gene names | Smap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stromal membrane-associated protein 1. | |||||
|
SMP1L_HUMAN
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.053941 (rank : 74) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8WU79, Q5QPL2, Q96C93, Q9NST2, Q9UJL8 | Gene names | SMAP1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stromal membrane-associated protein 1-like. | |||||
|
SMP1L_MOUSE
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.053908 (rank : 75) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7TN29, Q3U798 | Gene names | Smap1l, Smap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stromal membrane-associated protein 1-like (Stromal membrane- associated protein 2). | |||||
|
RHG18_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | Q8N392, Q58EZ3, Q6P679, Q6PJD7, Q96S64 | Gene names | ARHGAP18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho-GTPase-activating protein 18 (MacGAP). | |||||
|
RHG18_MOUSE
|
||||||
| NC score | 0.987326 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q8K0Q5, Q8BP03, Q8R196 | Gene names | Arhgap18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho-GTPase-activating protein 18. | |||||
|
RHG09_HUMAN
|
||||||
| NC score | 0.779803 (rank : 3) | θ value | 2.28291e-11 (rank : 28) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9BRR9, Q8TCJ3, Q8WYR0, Q96EZ2, Q96S74 | Gene names | ARHGAP9 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 9. | |||||
|
RHG06_HUMAN
|
||||||
| NC score | 0.778129 (rank : 4) | θ value | 7.09661e-13 (rank : 17) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | O43182, O43437, Q9P1B3, Q9UK81, Q9UK82 | Gene names | ARHGAP6, RHOGAP6 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 6 (Rho-type GTPase-activating protein RhoGAPX-1). | |||||
|
RHG06_MOUSE
|
||||||
| NC score | 0.776791 (rank : 5) | θ value | 2.69671e-12 (rank : 22) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | O54834, Q9QZL8 | Gene names | Arhgap6 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 6 (Rho-type GTPase-activating protein RhoGAPX-1). | |||||
|
RHG12_HUMAN
|
||||||
| NC score | 0.771914 (rank : 6) | θ value | 1.80466e-16 (rank : 9) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q8IWW6, Q86UB3, Q8IWW7, Q8N3L1, Q9NT76 | Gene names | ARHGAP12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
RHG12_MOUSE
|
||||||
| NC score | 0.767706 (rank : 7) | θ value | 1.38178e-16 (rank : 8) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q8C0D4, Q8BVP8 | Gene names | Arhgap12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
3BP1_MOUSE
|
||||||
| NC score | 0.767309 (rank : 8) | θ value | 5.08577e-11 (rank : 30) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P55194, Q99KK8 | Gene names | Sh3bp1, 3bp1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain-binding protein 1 (3BP-1). | |||||
|
GMIP_MOUSE
|
||||||
| NC score | 0.765847 (rank : 9) | θ value | 4.16044e-13 (rank : 16) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q6PGG2, Q6P9S3 | Gene names | Gmip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GEM-interacting protein (GMIP). | |||||
|
GMIP_HUMAN
|
||||||
| NC score | 0.763814 (rank : 10) | θ value | 1.86753e-13 (rank : 15) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9P107 | Gene names | GMIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GEM-interacting protein (GMIP). | |||||
|
RGAP1_HUMAN
|
||||||
| NC score | 0.762627 (rank : 11) | θ value | 9.26847e-13 (rank : 18) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9H0H5, Q6PJ26, Q9NWN2, Q9P250, Q9P2W2 | Gene names | RACGAP1, KIAA1478, MGCRACGAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rac GTPase-activating protein 1 (MgcRacGAP). | |||||
|
RHG07_MOUSE
|
||||||
| NC score | 0.758744 (rank : 12) | θ value | 1.09232e-21 (rank : 3) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9R0Z9, Q8R541 | Gene names | Dlc1, Arhgap7, Stard12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 7 (Rho-type GTPase-activating protein 7) (Deleted in liver cancer 1 protein homolog) (Dlc-1) (StAR-related lipid transfer protein 12) (StARD12) (START domain-containing protein 12). | |||||
|
RHG07_HUMAN
|
||||||
| NC score | 0.753927 (rank : 13) | θ value | 9.24701e-21 (rank : 4) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q96QB1, O14868, O43199, Q9C0E0 | Gene names | DLC1, ARHGAP7, KIAA1723, STARD12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 7 (Rho-type GTPase-activating protein 7) (Deleted in liver cancer 1 protein) (Dlc-1) (HP protein) (StAR-related lipid transfer protein 12) (StARD12) (START domain-containing protein 12). | |||||
|
3BP1_HUMAN
|
||||||
| NC score | 0.753504 (rank : 14) | θ value | 6.41864e-14 (rank : 13) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9Y3L3, Q6IBZ2, Q6ZVL9, Q96HQ5, Q9NSQ9 | Gene names | SH3BP1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain-binding protein 1 (3BP-1). | |||||
|
RGAP1_MOUSE
|
||||||
| NC score | 0.750594 (rank : 15) | θ value | 4.30538e-10 (rank : 43) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9WVM1, Q3THR5, Q3TI41, Q3TM81 | Gene names | Racgap1, Mgcracgap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rac GTPase-activating protein 1 (MgcRacGAP). | |||||
|
RHG01_HUMAN
|
||||||
| NC score | 0.744148 (rank : 16) | θ value | 2.28291e-11 (rank : 27) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q07960 | Gene names | ARHGAP1, CDC42GAP, RHOGAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 1 (GTPase-activating protein rhoOGAP) (Rho-related small GTPase protein activator) (CDC42 GTPase-activating protein) (p50-RhoGAP). | |||||
|
GRLF1_HUMAN
|
||||||
| NC score | 0.729330 (rank : 17) | θ value | 1.133e-10 (rank : 33) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9NRY4, Q14452, Q9C0E1 | Gene names | GRLF1, GRF1, KIAA1722 | |||
|
Domain Architecture |

|
|||||
| Description | Glucocorticoid receptor DNA-binding factor 1 (Glucocorticoid receptor repression factor 1) (GRF-1) (Rho GAP p190A) (p190-A). | |||||
|
RHG08_MOUSE
|
||||||
| NC score | 0.724505 (rank : 18) | θ value | 1.47974e-10 (rank : 39) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9CXP4, Q99JY7 | Gene names | Arhgap8 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 8. | |||||
|
RBP1_MOUSE
|
||||||
| NC score | 0.724453 (rank : 19) | θ value | 1.38178e-16 (rank : 7) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 493 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q62172, Q9CRE5 | Gene names | Ralbp1, Rip1 | |||
|
Domain Architecture |

|
|||||
| Description | RalA-binding protein 1 (RalBP1) (Ral-interacting protein 1) (Dinitrophenyl S-glutathione ATPase) (DNP-SG ATPase). | |||||
|
STA13_HUMAN
|
||||||
| NC score | 0.715308 (rank : 20) | θ value | 8.95645e-16 (rank : 11) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9Y3M8, Q5HYH1, Q5TAE3, Q6UN61, Q86TP6, Q86WQ3, Q86XT1 | Gene names | STARD13, DLC2, GT650 | |||
|
Domain Architecture |

|
|||||
| Description | StAR-related lipid transfer protein 13 (StARD13) (START domain- containing protein 13) (46H23.2) (Deleted in liver cancer protein 2) (Rho GTPase-activating protein). | |||||
|
CHIO_MOUSE
|
||||||
| NC score | 0.714100 (rank : 21) | θ value | 3.52202e-12 (rank : 23) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q80XD1, Q9D9W2, Q9ER57 | Gene names | Chn2, Arhgap3, Bch | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-chimaerin (Beta-chimerin) (Rho-GTPase-activating protein 3). | |||||
|
RHG08_HUMAN
|
||||||
| NC score | 0.714054 (rank : 22) | θ value | 1.63604e-09 (rank : 44) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9NSG0, O75983, O95695, Q96RW1, Q96RW2, Q9HA49, Q9HC46, Q9NVX8, Q9NXL1, Q9UH20 | Gene names | ARHGAP8 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 8. | |||||
|
TCGAP_MOUSE
|
||||||
| NC score | 0.710218 (rank : 23) | θ value | 1.133e-10 (rank : 37) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q80YF9, Q6P7W6 | Gene names | Snx26, Tcgap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TC10/CDC42 GTPase-activating protein (Sorting nexin-26). | |||||
|
OPHN1_MOUSE
|
||||||
| NC score | 0.709029 (rank : 24) | θ value | 1.133e-10 (rank : 35) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q99J31, Q544K7 | Gene names | Ophn1 | |||
|
Domain Architecture |

|
|||||
| Description | Oligophrenin 1. | |||||
|
OPHN1_HUMAN
|
||||||
| NC score | 0.708284 (rank : 25) | θ value | 1.9326e-10 (rank : 41) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | O60890, Q5JQ81, Q8WX47 | Gene names | OPHN1 | |||
|
Domain Architecture |

|
|||||
| Description | Oligophrenin 1. | |||||
|
STAR8_MOUSE
|
||||||
| NC score | 0.706411 (rank : 26) | θ value | 3.52202e-12 (rank : 24) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8K031, Q3UZC7, Q6A0A8, Q6P5E0, Q8R3X8 | Gene names | Stard8, Kiaa0189 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | StAR-related lipid transfer protein 8 (StARD8) (START domain- containing protein 8). | |||||
|
STA13_MOUSE
|
||||||
| NC score | 0.704890 (rank : 27) | θ value | 6.41864e-14 (rank : 14) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q923Q2, Q8K369 | Gene names | Stard13 | |||
|
Domain Architecture |

|
|||||
| Description | StAR-related lipid transfer protein 13 (StARD13) (START domain- containing protein 13). | |||||
|
RBP1_HUMAN
|
||||||
| NC score | 0.704603 (rank : 28) | θ value | 3.07829e-16 (rank : 10) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q15311 | Gene names | RALBP1, RLIP1, RLIP76 | |||
|
Domain Architecture |

|
|||||
| Description | RalA-binding protein 1 (RalBP1) (Ral-interacting protein 1) (76 kDa Ral-interacting protein) (Dinitrophenyl S-glutathione ATPase) (DNP-SG ATPase). | |||||
|
STAR8_HUMAN
|
||||||
| NC score | 0.697666 (rank : 29) | θ value | 2.0648e-12 (rank : 20) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q92502, Q5JST0 | Gene names | STARD8, KIAA0189 | |||
|
Domain Architecture |

|
|||||
| Description | StAR-related lipid transfer protein 8 (StARD8) (START domain- containing protein 8). | |||||
|
ABR_HUMAN
|
||||||
| NC score | 0.693869 (rank : 30) | θ value | 2.60593e-15 (rank : 12) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q12979, Q13693, Q13694 | Gene names | ABR | |||
|
Domain Architecture |

|
|||||
| Description | Active breakpoint cluster region-related protein. | |||||
|
RHG05_MOUSE
|
||||||
| NC score | 0.686918 (rank : 31) | θ value | 8.11959e-09 (rank : 47) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | P97393 | Gene names | Arhgap5, Rhogap5 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 5 (p190-B). | |||||
|
RHG25_MOUSE
|
||||||
| NC score | 0.682555 (rank : 32) | θ value | 1.9326e-10 (rank : 42) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q8BYW1, Q2VPR2, Q3TE04, Q3TVX0, Q3UVB7, Q6A0E0, Q8BX98 | Gene names | Arhgap25, Kiaa0053 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
RHG26_HUMAN
|
||||||
| NC score | 0.682144 (rank : 33) | θ value | 9.26847e-13 (rank : 19) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9UNA1, O75117, Q9BYS6, Q9BYS7, Q9UJ00 | Gene names | ARHGAP26, GRAF, KIAA0621, OPHN1L | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 26 (Oligophrenin-1-like protein) (GTPase regulator associated with focal adhesion kinase). | |||||
|
CHIN_HUMAN
|
||||||
| NC score | 0.681554 (rank : 34) | θ value | 2.98157e-11 (rank : 29) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P15882, Q53SD6, Q53SH5, Q96FB0 | Gene names | CHN1, ARHGAP2, CHN | |||
|
Domain Architecture |

|
|||||
| Description | N-chimaerin (NC) (N-chimerin) (Alpha chimerin) (A-chimaerin) (Rho- GTPase-activating protein 2). | |||||
|
CHIO_HUMAN
|
||||||
| NC score | 0.680908 (rank : 35) | θ value | 7.84624e-12 (rank : 26) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P52757, Q75MM2 | Gene names | CHN2, ARHGAP3, BCH | |||
|
Domain Architecture |
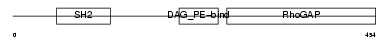
|
|||||
| Description | Beta-chimaerin (Beta-chimerin) (Rho-GTPase-activating protein 3). | |||||
|
CHIN_MOUSE
|
||||||
| NC score | 0.680289 (rank : 36) | θ value | 1.133e-10 (rank : 32) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q91V57, Q3UGY3, Q7TQE5, Q8BWU6, Q9D9B3 | Gene names | Chn1, Arhgap2 | |||
|
Domain Architecture |

|
|||||
| Description | N-chimaerin (NC) (N-chimerin) (Alpha chimerin) (A-chimaerin) (Rho- GTPase-activating protein 2). | |||||
|
RHG05_HUMAN
|
||||||
| NC score | 0.674161 (rank : 37) | θ value | 4.0297e-08 (rank : 50) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q13017 | Gene names | ARHGAP5, RHOGAP5 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 5 (p190-B). | |||||
|
BCR_HUMAN
|
||||||
| NC score | 0.671531 (rank : 38) | θ value | 6.21693e-09 (rank : 45) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P11274, P78501, Q12842 | Gene names | BCR, BCR1 | |||
|
Domain Architecture |

|
|||||
| Description | Breakpoint cluster region protein (EC 2.7.11.1) (NY-REN-26 antigen). | |||||
|
RHG25_HUMAN
|
||||||
| NC score | 0.663949 (rank : 39) | θ value | 8.67504e-11 (rank : 31) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 550 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | P42331, Q8IXQ2 | Gene names | ARHGAP25, KIAA0053 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
TAGAP_MOUSE
|
||||||
| NC score | 0.656129 (rank : 40) | θ value | 9.29e-05 (rank : 57) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8K2L9, Q920D6 | Gene names | Tagap, Tagap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-cell activation Rho GTPase-activating protein (T-cell activation GTPase-activating protein). | |||||
|
TAGAP_HUMAN
|
||||||
| NC score | 0.651433 (rank : 41) | θ value | 0.000786445 (rank : 58) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q8N103, Q2NKM8, Q8NI40, Q96KZ2, Q96QA2 | Gene names | TAGAP, TAGAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-cell activation Rho GTPase-activating protein (T-cell activation GTPase-activating protein). | |||||
|
RHG04_HUMAN
|
||||||
| NC score | 0.651206 (rank : 42) | θ value | 4.0297e-08 (rank : 49) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P98171, Q14144 | Gene names | ARHGAP4, KIAA0131, RGC1, RHOGAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 4 (Rho-GAP hematopoietic protein C1) (p115). | |||||
|
SRGP2_HUMAN
|
||||||
| NC score | 0.632943 (rank : 43) | θ value | 4.0297e-08 (rank : 51) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | O43295, Q8IX13, Q8IZV8 | Gene names | SRGAP3, ARHGAP14, KIAA0411, MEGAP, SRGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 3 (srGAP3) (srGAP2) (WAVE- associated Rac GTPase-activating protein) (WRP) (Mental disorder- associated GAP) (Rho-GTPase-activating protein 14). | |||||
|
TCGAP_HUMAN
|
||||||
| NC score | 0.630850 (rank : 44) | θ value | 1.133e-10 (rank : 36) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 872 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | O14559, O14552, O14560, Q6ZSP6, Q96CP3, Q9NT23 | Gene names | SNX26, TCGAP | |||
|
Domain Architecture |

|
|||||
| Description | TC10/CDC42 GTPase-activating protein (Sorting nexin-26). | |||||
|
SRGP2_MOUSE
|
||||||
| NC score | 0.623985 (rank : 45) | θ value | 8.97725e-08 (rank : 52) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q812A2, Q80U09, Q8BKP4, Q8BLD0, Q925I2 | Gene names | Srgap3, Arhgap14, Kiaa0411, Srgap2 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 3 (srGAP3) (srGAP2) (WAVE- associated Rac GTPase-activating protein) (WRP) (Rho-GTPase-activating protein 14). | |||||
|
CE005_HUMAN
|
||||||
| NC score | 0.620193 (rank : 46) | θ value | 1.06045e-08 (rank : 48) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9NYF5, Q6PGQ2, Q9P0I7 | Gene names | C5orf5 | |||
|
Domain Architecture |

|
|||||
| Description | Protein C5orf5 (GAP-like protein N61). | |||||
|
CE005_MOUSE
|
||||||
| NC score | 0.618616 (rank : 47) | θ value | 6.21693e-09 (rank : 46) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q8K2H3 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Protein C5orf5 homolog. | |||||
|
FNBP2_HUMAN
|
||||||
| NC score | 0.615601 (rank : 48) | θ value | 1.99992e-07 (rank : 53) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | O75044 | Gene names | SRGAP2, FNBP2, KIAA0456 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 2 (srGAP2) (Formin-binding protein 2). | |||||
|
SRGP1_HUMAN
|
||||||
| NC score | 0.610653 (rank : 49) | θ value | 3.77169e-06 (rank : 56) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q7Z6B7, Q9H8A3, Q9P2P2 | Gene names | SRGAP1, ARHGAP13, KIAA1304 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 1 (srGAP1) (Rho-GTPase- activating protein 13). | |||||
|
CEND2_HUMAN
|
||||||
| NC score | 0.604772 (rank : 50) | θ value | 1.9326e-10 (rank : 40) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q96P48, O94879, Q59FI7, Q6PHS3, Q8WU51, Q96HP6, Q96L71 | Gene names | CENTD2, ARAP1, KIAA0782 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-delta 2 (Cnt-d2) (Arf-GAP, Rho-GAP, ankyrin repeat and pleckstrin homology domain-containing protein 1). | |||||
|
K1688_MOUSE
|
||||||
| NC score | 0.593796 (rank : 51) | θ value | 1.133e-10 (rank : 34) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P59281, Q69ZD4, Q6P9R6 | Gene names | Kiaa1688, D15Wsu169e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1688. | |||||
|
CEND3_MOUSE
|
||||||
| NC score | 0.586559 (rank : 52) | θ value | 4.59992e-12 (rank : 25) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q8R5G7, Q5DTN4, Q6NVF1, Q8R5G6 | Gene names | Centd3, Arap3, Drag1, Kiaa4097 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-delta 3 (Cnt-d3) (Arf-GAP, Rho-GAP, ankyrin repeat and pleckstrin homology domain-containing protein 3) (Dual specificity Rho- and Arf-GTPase-activating protein 1). | |||||
|
K1688_HUMAN
|
||||||
| NC score | 0.580052 (rank : 53) | θ value | 1.47974e-10 (rank : 38) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9C0H5 | Gene names | KIAA1688 | |||
|
Domain Architecture |

|
|||||
| Description | Protein KIAA1688. | |||||
|
CEND3_HUMAN
|
||||||
| NC score | 0.578270 (rank : 54) | θ value | 2.69671e-12 (rank : 21) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q8WWN8 | Gene names | CENTD3, ARAP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-delta 3 (Cnt-d3) (Arf-GAP, Rho-GAP, ankyrin repeat and pleckstrin homology domain-containing protein 3). | |||||
|
CEND1_MOUSE
|
||||||
| NC score | 0.506620 (rank : 55) | θ value | 3.41135e-07 (rank : 54) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q8BZ05, Q80TX2, Q8C3T2, Q8VEL6 | Gene names | Centd1, Kiaa0580 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-delta 1 (Cnt-d1). | |||||
|
CEND1_HUMAN
|
||||||
| NC score | 0.486114 (rank : 56) | θ value | 9.92553e-07 (rank : 55) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q8WZ64, Q4W5D2, Q7Z2L5, Q96L70, Q96P49, Q9Y4E4 | Gene names | CENTD1, ARAP2, KIAA0580 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-delta 1 (Cnt-d1) (Arf-GAP, Rho-GAP, ankyrin repeat and pleckstrin homology domain-containing protein 2) (PARX protein). | |||||
|
MYO9B_MOUSE
|
||||||
| NC score | 0.413054 (rank : 57) | θ value | 1.38178e-16 (rank : 6) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9QY06, Q9QY07, Q9QY08, Q9QY09 | Gene names | Myo9b, Myr5 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9B (Myosin IXb) (Unconventional myosin-9b). | |||||
|
MYO9B_HUMAN
|
||||||
| NC score | 0.407466 (rank : 58) | θ value | 8.10077e-17 (rank : 5) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q13459, O75314, Q9NUJ2, Q9UHN0 | Gene names | MYO9B, MYR5 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9B (Myosin IXb) (Unconventional myosin-9b). | |||||
|
P85A_HUMAN
|
||||||
| NC score | 0.319128 (rank : 59) | θ value | 0.00102713 (rank : 59) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P27986 | Gene names | PIK3R1, GRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit alpha (PI3-kinase p85-subunit alpha) (PtdIns-3-kinase p85-alpha) (PI3K). | |||||
|
P85A_MOUSE
|
||||||
| NC score | 0.306913 (rank : 60) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P26450 | Gene names | Pik3r1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit alpha (PI3-kinase p85-subunit alpha) (PtdIns-3-kinase p85-alpha) (PI3K). | |||||
|
I5P2_HUMAN
|
||||||
| NC score | 0.305335 (rank : 61) | θ value | 0.00390308 (rank : 60) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P32019, Q5VSG9, Q5VSH0, Q5VSH1, Q658Q5, Q6P6D4, Q6PD53, Q86YE1 | Gene names | INPP5B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Type II inositol-1,4,5-trisphosphate 5-phosphatase precursor (EC 3.1.3.36) (Phosphoinositide 5-phosphatase) (5PTase) (75 kDa inositol polyphosphate-5-phosphatase). | |||||
|
OCRL_HUMAN
|
||||||
| NC score | 0.272443 (rank : 62) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q01968, O60800, Q15684, Q15774, Q5JQF1, Q5JQF2, Q9UJG5, Q9UMA5 | Gene names | OCRL, INPP5F, OCRL1 | |||
|
Domain Architecture |

|
|||||
| Description | Inositol polyphosphate 5-phosphatase OCRL-1 (EC 3.1.3.36) (Lowe oculocerebrorenal syndrome protein). | |||||
|
PCTL_MOUSE
|
||||||
| NC score | 0.094861 (rank : 63) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9JMD3 | Gene names | Stard10, Pctpl, Sdccag28, Sdccagg28 | |||
|
Domain Architecture |

|
|||||
| Description | PCTP-like protein (PCTP-L) (StAR-related lipid transfer protein 10) (StARD10) (START domain-containing protein 10) (Serologically defined colon cancer antigen 28 homolog). | |||||
|
PCTL_HUMAN
|
||||||
| NC score | 0.093774 (rank : 64) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y365, O60532 | Gene names | STARD10, SDCCAG28 | |||
|
Domain Architecture |

|
|||||
| Description | PCTP-like protein (PCTP-L) (StAR-related lipid transfer protein 10) (StARD10) (START domain-containing protein 10) (Serologically defined colon cancer antigen 28) (Antigen NY-CO-28). | |||||
|
CENB2_HUMAN
|
||||||
| NC score | 0.076308 (rank : 65) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q15057, Q9UQR3 | Gene names | CENTB2, KIAA0041 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-beta 2 (Cnt-b2). | |||||
|
P85B_HUMAN
|
||||||
| NC score | 0.073961 (rank : 66) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O00459, Q9UPH9 | Gene names | PIK3R2 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit beta (PI3-kinase p85- subunit beta) (PtdIns-3-kinase p85-beta). | |||||
|
CENB5_HUMAN
|
||||||
| NC score | 0.070503 (rank : 67) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96P50, Q9BSR9, Q9C0E7 | Gene names | CENTB5, KIAA1716 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-beta 5 (Cnt-b5). | |||||
|
CENB1_HUMAN
|
||||||
| NC score | 0.070211 (rank : 68) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q15027 | Gene names | CENTB1, KIAA0050 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-beta 1 (Cnt-b1). | |||||
|
GRP3_HUMAN
|
||||||
| NC score | 0.068744 (rank : 69) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8IV61, O94931 | Gene names | RASGRP3, GRP3, KIAA0846 | |||
|
Domain Architecture |

|
|||||
| Description | RAS guanyl-releasing protein 3 (Calcium and DAG-regulated guanine nucleotide exchange factor III) (Guanine nucleotide exchange factor for Rap1). | |||||
|
FACD2_HUMAN
|
||||||
| NC score | 0.066352 (rank : 70) | θ value | 0.0330416 (rank : 61) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BXW9, Q2LA86, Q69YP9, Q6PJN7, Q9BQ06, Q9H9T9 | Gene names | FANCD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group D2 protein (Protein FACD2). | |||||
|
FA13A_MOUSE
|
||||||
| NC score | 0.066038 (rank : 71) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BGI4, Q8BZ91 | Gene names | Fam13a1, Precm1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13A1 (Precm1 protein). | |||||
|
FA13A_HUMAN
|
||||||
| NC score | 0.065528 (rank : 72) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O94988, Q8NBA3 | Gene names | FAM13A1, KIAA0914 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13A1. | |||||
|
CENA2_HUMAN
|
||||||
| NC score | 0.054774 (rank : 73) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NPF8, Q8N4Q6, Q96SD5 | Gene names | CENTA2 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-alpha 2. | |||||
|
SMP1L_HUMAN
|
||||||
| NC score | 0.053941 (rank : 74) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8WU79, Q5QPL2, Q96C93, Q9NST2, Q9UJL8 | Gene names | SMAP1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stromal membrane-associated protein 1-like. | |||||
|
SMP1L_MOUSE
|
||||||
| NC score | 0.053908 (rank : 75) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7TN29, Q3U798 | Gene names | Smap1l, Smap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stromal membrane-associated protein 1-like (Stromal membrane- associated protein 2). | |||||
|
DDEF2_HUMAN
|
||||||
| NC score | 0.053901 (rank : 76) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 599 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O43150 | Gene names | DDEF2, KIAA0400 | |||
|
Domain Architecture |

|
|||||
| Description | Development and differentiation-enhancing factor 2 (Pyk2 C-terminus- associated protein) (PAP) (Paxillin-associated protein with ARFGAP activity 3) (PAG3). | |||||
|
BNIP2_HUMAN
|
||||||
| NC score | 0.053614 (rank : 77) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q12982 | Gene names | BNIP2, NIP2 | |||
|
Domain Architecture |

|
|||||
| Description | BCL2/adenovirus E1B 19 kDa protein-interacting protein 2. | |||||
|
BNIP2_MOUSE
|
||||||
| NC score | 0.053528 (rank : 78) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O54940 | Gene names | Bnip2, Nip2l | |||
|
Domain Architecture |

|
|||||
| Description | BCL2/adenovirus E1B 19 kDa protein-interacting protein 2. | |||||
|
PKHA1_HUMAN
|
||||||
| NC score | 0.053131 (rank : 79) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9HB21, Q9BVK0 | Gene names | PLEKHA1, TAPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 1 (Tandem PH domain-containing protein 1) (TAPP-1). | |||||
|
DDEF2_MOUSE
|
||||||
| NC score | 0.052664 (rank : 80) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 625 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7SIG6, Q501K1, Q66JN2 | Gene names | Ddef2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Development and differentiation-enhancing factor 2 (Pyk2 C-terminus- associated protein) (PAP) (Paxillin-associated protein with ARFGAP activity 3) (PAG3). | |||||
|
SMAP1_HUMAN
|
||||||
| NC score | 0.052559 (rank : 81) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IYB5, Q53H70, Q5SYQ2, Q6PK24, Q8NDH4, Q96L38, Q96L39, Q9H8X4 | Gene names | SMAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stromal membrane-associated protein 1. | |||||
|
DDFL1_HUMAN
|
||||||
| NC score | 0.052139 (rank : 82) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 353 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8TDY4, Q6P9F4, Q86UY1, Q9NXK2 | Gene names | DDEFL1, UPLC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Development and differentiation-enhancing factor-like 1 (Protein up- regulated in liver cancer 1). | |||||
|
SMAP1_MOUSE
|
||||||
| NC score | 0.052022 (rank : 83) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q91VZ6, Q497I6, Q68EF3 | Gene names | Smap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stromal membrane-associated protein 1. | |||||
|
MYO10_HUMAN
|
||||||
| NC score | 0.052015 (rank : 84) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 920 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9HD67, Q9NYM7, Q9P110, Q9P111, Q9UHF6 | Gene names | MYO10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin X). | |||||
|
CENA1_HUMAN
|
||||||
| NC score | 0.051319 (rank : 85) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75689 | Gene names | CENTA1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-alpha 1 (Putative MAPK-activating protein PM25). | |||||
|
FA13C_HUMAN
|
||||||
| NC score | 0.051033 (rank : 86) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NE31, Q99787 | Gene names | FAM13C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13C1. | |||||
|
ATCAY_MOUSE
|
||||||
| NC score | 0.050965 (rank : 87) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BHE3, Q3TR94 | Gene names | Atcay | |||
|
Domain Architecture |

|
|||||
| Description | Caytaxin. | |||||
|
ATCAY_HUMAN
|
||||||
| NC score | 0.050505 (rank : 88) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86WG3, Q8NAQ2, Q8TAQ3, Q96HC6, Q96JF5 | Gene names | ATCAY, KIAA1872 | |||
|
Domain Architecture |

|
|||||
| Description | Caytaxin (Ataxia Cayman type protein) (BNIP-H). | |||||
|
DBLOH_MOUSE
|
||||||
| NC score | 0.030717 (rank : 89) | θ value | 1.81305 (rank : 68) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JIQ3, Q9CZD1, Q9DCD3 | Gene names | Diablo, Smac | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diablo homolog, mitochondrial precursor (Second mitochondria-derived activator of caspase) (Smac protein) (Direct IAP-binding protein with low pI). | |||||
|
RABE1_HUMAN
|
||||||
| NC score | 0.025276 (rank : 90) | θ value | 0.0736092 (rank : 62) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1104 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q15276, O95369, Q8IVX3 | Gene names | RABEP1, RABPT5, RABPT5A | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 1 (Rabaptin-5) (Rabaptin-5alpha) (Rabaptin-4) (NY-REN-17 antigen). | |||||
|
DBLOH_HUMAN
|
||||||
| NC score | 0.023731 (rank : 91) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NR28, Q96LV0, Q9BT11, Q9HAV6 | Gene names | DIABLO, SMAC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diablo homolog, mitochondrial precursor (Second mitochondria-derived activator of caspase) (Smac protein) (Direct IAP-binding protein with low pI). | |||||
|
5NT1B_MOUSE
|
||||||
| NC score | 0.022472 (rank : 92) | θ value | 1.06291 (rank : 65) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91YE9, Q91Y48 | Gene names | Nt5c1b, Airp | |||
|
Domain Architecture |

|
|||||
| Description | Cytosolic 5'-nucleotidase 1B (EC 3.1.3.5) (Cytosolic 5'-nucleotidase IB) (cN1B) (cN-IB) (Autoimmune infertility-related protein). | |||||
|
NIN_HUMAN
|
||||||
| NC score | 0.022103 (rank : 93) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1416 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8N4C6, Q6P0P6, Q9BWU6, Q9C012, Q9C013, Q9C014, Q9H5I6, Q9HAT7, Q9HBY5, Q9HCK7, Q9UH61 | Gene names | NIN, KIAA1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein (hNinein) (Glycogen synthase kinase 3 beta-interacting protein) (GSK3B-interacting protein). | |||||
|
RABE1_MOUSE
|
||||||
| NC score | 0.019800 (rank : 94) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O35551, Q99L08, Q9CRP3, Q9EQF9, Q9JI94 | Gene names | Rabep1, Rab5ep, Rabpt5, Rabpt5a | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 1 (Rabaptin-5) (Rabaptin-5alpha). | |||||
|
MCM4_MOUSE
|
||||||
| NC score | 0.018984 (rank : 95) | θ value | 1.81305 (rank : 71) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P49717, O89056 | Gene names | Mcm4, Cdc21, Mcmd4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM4 (CDC21 homolog) (P1-CDC21). | |||||
|
IF4G1_HUMAN
|
||||||
| NC score | 0.017646 (rank : 96) | θ value | 1.06291 (rank : 67) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q04637, O43177, O95066, Q5HYG0, Q6ZN21, Q8N102 | Gene names | EIF4G1, EIF4G, EIF4GI | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1) (p220). | |||||
|
FLIP1_HUMAN
|
||||||
| NC score | 0.016137 (rank : 97) | θ value | 0.365318 (rank : 64) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q7Z7B0, Q5VUL6, Q86TC3, Q8N8B9, Q96SK6, Q9NVI8, Q9ULE5 | Gene names | FILIP1, KIAA1275 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
GOGA4_MOUSE
|
||||||
| NC score | 0.014202 (rank : 98) | θ value | 1.81305 (rank : 69) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
RAE2_HUMAN
|
||||||
| NC score | 0.013703 (rank : 99) | θ value | 5.27518 (rank : 76) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P26374, Q9H1Y4 | Gene names | CHML, REP2 | |||
|
Domain Architecture |

|
|||||
| Description | Rab proteins geranylgeranyltransferase component A 2 (Rab escort protein 2) (REP-2) (Choroideraemia-like protein). | |||||
|
MCM4_HUMAN
|
||||||
| NC score | 0.013549 (rank : 100) | θ value | 4.03905 (rank : 74) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P33991, Q8NEH1, Q99658 | Gene names | MCM4, CDC21 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM4 (CDC21 homolog) (P1-CDC21). | |||||
|
GOGA5_MOUSE
|
||||||
| NC score | 0.013522 (rank : 101) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1434 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9QYE6, O88317, Q3TGE7, Q3U6S5, Q3UUF9 | Gene names | Golga5, Retii | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (Sumiko protein) (Ret-II protein). | |||||
|
TRIPB_HUMAN
|
||||||
| NC score | 0.012616 (rank : 102) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1587 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q15643, O14689, O15154, O95949 | Gene names | TRIP11, CEV14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid receptor-interacting protein 11 (TRIP-11) (Golgi-associated microtubule-binding protein 210) (GMAP-210) (Trip230) (Clonal evolution-related gene on chromosome 14). | |||||
|
COG8_MOUSE
|
||||||
| NC score | 0.011591 (rank : 103) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JJA2 | Gene names | Cog8 | |||
|
Domain Architecture |

|
|||||
| Description | Conserved oligomeric Golgi complex component 8. | |||||
|
KPBB_MOUSE
|
||||||
| NC score | 0.010172 (rank : 104) | θ value | 4.03905 (rank : 73) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7TSH2 | Gene names | Phkb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphorylase b kinase regulatory subunit beta (Phosphorylase kinase subunit beta). | |||||
|
RBL1_MOUSE
|
||||||
| NC score | 0.008603 (rank : 105) | θ value | 5.27518 (rank : 77) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q64701 | Gene names | Rbl1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoblastoma-like protein 1 (107 kDa retinoblastoma-associated protein) (PRB1) (P107). | |||||
|
SMCA4_HUMAN
|
||||||
| NC score | 0.008484 (rank : 106) | θ value | 1.81305 (rank : 72) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P51532, O95052, Q9HBD3 | Gene names | SMARCA4, BRG1, SNF2B, SNF2L4 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L4 (EC 3.6.1.-) (ATP- dependent helicase SMARCA4) (SNF2-beta) (BRG-1 protein) (Mitotic growth and transcription activator) (Brahma protein homolog 1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 4). | |||||
|
PAR14_MOUSE
|
||||||
| NC score | 0.008333 (rank : 107) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q2EMV9, Q571C6, Q8BN35, Q8VDT6 | Gene names | Parp14, Kiaa1268 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 14 (EC 2.4.2.30) (PARP-14) (Collaborator of STAT6) (CoaSt6). | |||||
|
KI21B_HUMAN
|
||||||
| NC score | 0.007173 (rank : 108) | θ value | 1.81305 (rank : 70) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1017 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O75037, Q5T4J3 | Gene names | KIF21B, KIAA0449 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B. | |||||
|
SMRCD_MOUSE
|
||||||
| NC score | 0.006536 (rank : 109) | θ value | 5.27518 (rank : 78) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q04692 | Gene names | Smarcad1, Etl1, Kiaa1122 | |||
|
Domain Architecture |

|
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A containing DEAD/H box 1 (EC 3.6.1.-) (ATP- dependent helicase SMARCAD1) (Enhancer trap locus homolog 1) (Etl-1). | |||||
|
KI21B_MOUSE
|
||||||
| NC score | 0.006229 (rank : 110) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 997 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9QXL1, P97424 | Gene names | Kif21b, Kif6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B (Kinesin-like protein KIF6). | |||||
|
ROCK1_HUMAN
|
||||||
| NC score | 0.006144 (rank : 111) | θ value | 0.279714 (rank : 63) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 2000 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q13464 | Gene names | ROCK1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 1 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 1) (p160 ROCK-1) (p160ROCK) (NY-REN-35 antigen). | |||||
|
RPGF6_HUMAN
|
||||||
| NC score | 0.005874 (rank : 112) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TEU7, Q8NI21, Q8TEU6, Q96PC1 | Gene names | RAPGEF6, PDZGEF2 | |||
|
Domain Architecture |

|
|||||
| Description | Rap guanine nucleotide exchange factor 6 (PDZ domain-containing guanine nucleotide exchange factor 2) (PDZ-GEF2) (RA-GEF-2). | |||||
|
PRAX_HUMAN
|
||||||
| NC score | 0.004814 (rank : 113) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BXM0, Q9BXL9, Q9HCF2 | Gene names | PRX, KIAA1620 | |||
|
Domain Architecture |

|
|||||
| Description | Periaxin. | |||||
|
KLHL6_MOUSE
|
||||||
| NC score | 0.001420 (rank : 114) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6V595 | Gene names | Klhl6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kelch-like protein 6. | |||||
|
ITB2_HUMAN
|
||||||
| NC score | 0.001116 (rank : 115) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P05107, Q16418 | Gene names | ITGB2, CD18 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-2 precursor (Cell surface adhesion glycoproteins LFA- 1/CR3/p150,95 subunit beta) (Complement receptor C3 subunit beta) (CD18 antigen). | |||||
|
WNK4_HUMAN
|
||||||
| NC score | -0.002275 (rank : 116) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1043 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96J92, Q8N8X3, Q8N8Z2, Q96DT8, Q9BYS5 | Gene names | WNK4, PRKWNK4 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK4 (EC 2.7.11.1) (Protein kinase with no lysine 4) (Protein kinase, lysine-deficient 4). | |||||
|
DAPK3_HUMAN
|
||||||
| NC score | -0.002577 (rank : 117) | θ value | 1.06291 (rank : 66) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 945 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43293 | Gene names | DAPK3, ZIPK | |||
|
Domain Architecture |

|
|||||
| Description | Death-associated protein kinase 3 (EC 2.7.11.1) (DAP kinase 3) (DAP- like kinase) (Dlk) (ZIP-kinase). | |||||