Please be patient as the page loads
|
PAR14_MOUSE
|
||||||
| SwissProt Accessions | Q2EMV9, Q571C6, Q8BN35, Q8VDT6 | Gene names | Parp14, Kiaa1268 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 14 (EC 2.4.2.30) (PARP-14) (Collaborator of STAT6) (CoaSt6). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PAR14_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.983019 (rank : 2) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q460N5, Q460N4, Q8J027, Q9H9X9, Q9NV60, Q9ULF2 | Gene names | PARP14, BAL2, KIAA1268 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 14 (EC 2.4.2.30) (PARP-14) (B aggressive lymphoma protein 2). | |||||
|
PAR14_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 113 | |
| SwissProt Accessions | Q2EMV9, Q571C6, Q8BN35, Q8VDT6 | Gene names | Parp14, Kiaa1268 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 14 (EC 2.4.2.30) (PARP-14) (Collaborator of STAT6) (CoaSt6). | |||||
|
PAR15_HUMAN
|
||||||
| θ value | 8.73085e-104 (rank : 3) | NC score | 0.925679 (rank : 3) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q460N3, Q8N1K3 | Gene names | PARP15, BAL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 15 (EC 2.4.2.30) (PARP-15) (B-aggressive lymphoma protein 3). | |||||
|
PARP9_MOUSE
|
||||||
| θ value | 1.36585e-56 (rank : 4) | NC score | 0.807987 (rank : 5) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8CAS9, Q99LF9 | Gene names | Parp9, Bal | |||
|
Domain Architecture |
|
|||||
| Description | Poly [ADP-ribose] polymerase 9 (EC 2.4.2.30) (PARP-9) (B aggressive lymphoma protein homolog). | |||||
|
PARP9_HUMAN
|
||||||
| θ value | 7.75577e-52 (rank : 5) | NC score | 0.810188 (rank : 4) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8IXQ6, Q8TCP3, Q9BZL8, Q9BZL9 | Gene names | PARP9, BAL | |||
|
Domain Architecture |
|
|||||
| Description | Poly [ADP-ribose] polymerase 9 (EC 2.4.2.30) (PARP-9) (B aggressive lymphoma protein). | |||||
|
PAR10_HUMAN
|
||||||
| θ value | 4.72714e-33 (rank : 6) | NC score | 0.679239 (rank : 10) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q53GL7, Q8N2I0, Q8WV05, Q96CH7 | Gene names | PARP10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 10 (EC 2.4.2.30) (PARP-10). | |||||
|
PAR12_MOUSE
|
||||||
| θ value | 8.91499e-32 (rank : 7) | NC score | 0.689306 (rank : 8) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BZ20, Q80VL6, Q8K333, Q8R3U2 | Gene names | Parp12, Zc3hdc1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 12 (EC 2.4.2.30) (PARP-12) (Zinc finger CCCH domain-containing protein 1). | |||||
|
PAR12_HUMAN
|
||||||
| θ value | 1.98606e-31 (rank : 8) | NC score | 0.685490 (rank : 9) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9H0J9, Q9H610, Q9NP36, Q9NTI3 | Gene names | PARP12, ZC3HDC1 | |||
|
Domain Architecture |
|
|||||
| Description | Poly [ADP-ribose] polymerase 12 (EC 2.4.2.30) (PARP-12) (Zinc finger CCCH domain-containing protein 1). | |||||
|
PARPT_HUMAN
|
||||||
| θ value | 6.38894e-30 (rank : 9) | NC score | 0.699559 (rank : 7) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q7Z3E1, Q68CY9, Q86VP4, Q9Y4P7 | Gene names | TIPARP, PARP7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TCDD-inducible poly [ADP-ribose] polymerase (EC 2.4.2.30) (Poly [ADP- ribose] polymerase 7) (PARP-7). | |||||
|
PARPT_MOUSE
|
||||||
| θ value | 6.38894e-30 (rank : 10) | NC score | 0.699786 (rank : 6) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8C1B2, Q3UD50, Q8C032 | Gene names | Tiparp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TCDD-inducible poly [ADP-ribose] polymerase (EC 2.4.2.30). | |||||
|
ZCC2_HUMAN
|
||||||
| θ value | 6.62687e-19 (rank : 11) | NC score | 0.641376 (rank : 11) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7Z2W4, Q8IW57, Q8TAJ3, Q96N79, Q9H8R9, Q9P0Y7 | Gene names | ZC3HAV1, ZC3HDC2 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger CCCH type antiviral protein 1 (Zinc finger CCCH domain- containing protein 2). | |||||
|
LRP16_HUMAN
|
||||||
| θ value | 9.56915e-18 (rank : 12) | NC score | 0.546403 (rank : 12) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BQ69, Q9UH96 | Gene names | LRP16 | |||
|
Domain Architecture |
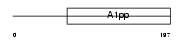
|
|||||
| Description | Protein LRP16. | |||||
|
LRP16_MOUSE
|
||||||
| θ value | 2.60593e-15 (rank : 13) | NC score | 0.534480 (rank : 13) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q922B1 | Gene names | Lrp16 | |||
|
Domain Architecture |
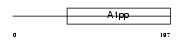
|
|||||
| Description | Protein LRP16. | |||||
|
H2AY_HUMAN
|
||||||
| θ value | 3.76295e-14 (rank : 14) | NC score | 0.296154 (rank : 14) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75367, O75377, Q503A8, Q7Z5E3, Q96D41, Q9H8P3, Q9UP96 | Gene names | H2AFY, MACROH2A1 | |||
|
Domain Architecture |

|
|||||
| Description | Core histone macro-H2A.1 (Histone macroH2A1) (mH2A1) (H2A.y) (H2A/y) (Medulloblastoma antigen MU-MB-50.205). | |||||
|
H2AY_MOUSE
|
||||||
| θ value | 4.91457e-14 (rank : 15) | NC score | 0.295573 (rank : 15) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9QZQ8, Q91VZ2, Q9QZQ7 | Gene names | H2afy | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Core histone macro-H2A.1 (Histone macroH2A1) (mH2A1) (H2A.y) (H2A/y). | |||||
|
H2AW_HUMAN
|
||||||
| θ value | 8.67504e-11 (rank : 16) | NC score | 0.244232 (rank : 16) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9P0M6, Q5SQT2 | Gene names | H2AFY2, MACROH2A2 | |||
|
Domain Architecture |

|
|||||
| Description | Core histone macro-H2A.2 (Histone macroH2A2) (mH2A2). | |||||
|
H2AW_MOUSE
|
||||||
| θ value | 8.67504e-11 (rank : 17) | NC score | 0.241829 (rank : 17) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8CCK0, Q3TNT4, Q925I6 | Gene names | H2afy2, H2afy3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Core histone macro-H2A.2 (Histone macroH2A2) (mH2A2). | |||||
|
TNKS2_HUMAN
|
||||||
| θ value | 4.76016e-09 (rank : 18) | NC score | 0.134615 (rank : 20) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H2K2, Q9H8F2, Q9HAS4 | Gene names | TNKS2, PARP5B, TANK2, TNKL | |||
|
Domain Architecture |

|
|||||
| Description | Tankyrase-2 (EC 2.4.2.30) (TANK2) (Tankyrase II) (TNKS-2) (TRF1- interacting ankyrin-related ADP-ribose polymerase 2) (Tankyrase-like protein) (Tankyrase-related protein). | |||||
|
DTX3L_HUMAN
|
||||||
| θ value | 1.53129e-07 (rank : 19) | NC score | 0.134902 (rank : 19) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8TDB6 | Gene names | DTX3L, BBAP | |||
|
Domain Architecture |

|
|||||
| Description | Protein deltex 3-like protein (B-lymphoma- and BAL-associated protein) (Rhysin-2) (Rhysin2). | |||||
|
TNKS1_HUMAN
|
||||||
| θ value | 1.69304e-06 (rank : 20) | NC score | 0.114106 (rank : 21) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O95271, O95272 | Gene names | TNKS, PARP5A, PARPL, TIN1, TINF1, TNKS1 | |||
|
Domain Architecture |

|
|||||
| Description | Tankyrase-1 (EC 2.4.2.30) (TANK1) (Tankyrase I) (TNKS-1) (TRF1- interacting ankyrin-related ADP-ribose polymerase). | |||||
|
NMI_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 21) | NC score | 0.105963 (rank : 22) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q13287, Q53TI8, Q9BVE5 | Gene names | NMI | |||
|
Domain Architecture |
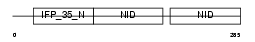
|
|||||
| Description | N-myc-interactor (Nmi) (N-myc and STAT interactor). | |||||
|
PARP3_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 22) | NC score | 0.140620 (rank : 18) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y6F1, Q8NER9, Q96CG2, Q9UG81 | Gene names | PARP3, ADPRT3, ADPRTL3 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 3 (EC 2.4.2.30) (PARP-3) (NAD(+) ADP- ribosyltransferase 3) (Poly[ADP-ribose] synthetase 3) (pADPRT-3) (hPARP-3) (IRT1). | |||||
|
TRM61_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 23) | NC score | 0.064257 (rank : 50) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96FX7, Q8N7Q9 | Gene names | TRM61, C14orf172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA (adenine-N(1)-)-methyltransferase catalytic subunit TRM61 (EC 2.1.1.36) (tRNA(m1A58)-methyltransferase subunit TRM61) (tRNA(m1A58)MTase subunit TRM61). | |||||
|
AKAP9_HUMAN
|
||||||
| θ value | 0.163984 (rank : 24) | NC score | 0.036759 (rank : 57) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1710 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q99996, O14869, O43355, O94895, Q9UQH3, Q9UQQ4, Q9Y6B8, Q9Y6Y2 | Gene names | AKAP9, AKAP350, AKAP450, KIAA0803 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 9 (Protein kinase A-anchoring protein 9) (PRKA9) (A-kinase anchor protein 450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP 350) (hgAKAP 350) (AKAP 120-like protein) (Protein hyperion) (Protein yotiao) (Centrosome- and Golgi-localized PKN-associated protein) (CG-NAP). | |||||
|
VIGLN_HUMAN
|
||||||
| θ value | 0.21417 (rank : 25) | NC score | 0.057964 (rank : 53) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q00341, Q9UCY3 | Gene names | HDLBP, HBP, VGL | |||
|
Domain Architecture |

|
|||||
| Description | Vigilin (High density lipoprotein-binding protein) (HDL-binding protein). | |||||
|
VIGLN_MOUSE
|
||||||
| θ value | 0.21417 (rank : 26) | NC score | 0.058721 (rank : 52) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8VDJ3 | Gene names | Hdlbp | |||
|
Domain Architecture |

|
|||||
| Description | Vigilin (High density lipoprotein-binding protein) (HDL-binding protein). | |||||
|
HUS1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 27) | NC score | 0.035843 (rank : 58) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BQY8, O70543, Q6P8H5 | Gene names | Hus1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Checkpoint protein HUS1 (mHUS1). | |||||
|
TITIN_HUMAN
|
||||||
| θ value | 0.279714 (rank : 28) | NC score | 0.004888 (rank : 134) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
ANR26_MOUSE
|
||||||
| θ value | 0.47712 (rank : 29) | NC score | 0.030456 (rank : 61) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1554 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q811D2, Q69ZS2, Q9CS61 | Gene names | Ankrd26, Kiaa1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
SH3G3_HUMAN
|
||||||
| θ value | 0.47712 (rank : 30) | NC score | 0.020245 (rank : 89) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99963, O43553, O43554 | Gene names | SH3GL3, CNSA3, SH3D2C | |||
|
Domain Architecture |
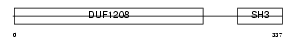
|
|||||
| Description | SH3-containing GRB2-like protein 3 (Endophilin-3) (Endophilin-A3) (SH3 domain protein 2C) (EEN-B2). | |||||
|
ZN287_HUMAN
|
||||||
| θ value | 0.62314 (rank : 31) | NC score | -0.000894 (rank : 140) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9HBT7 | Gene names | ZNF287 | |||
|
Domain Architecture |
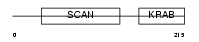
|
|||||
| Description | Zinc finger protein 287. | |||||
|
CP1A2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 32) | NC score | 0.007824 (rank : 128) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P05177, Q16754, Q6NWU5, Q9BXX7, Q9UK49 | Gene names | CYP1A2 | |||
|
Domain Architecture |

|
|||||
| Description | Cytochrome P450 1A2 (EC 1.14.14.1) (CYPIA2) (P450-P3) (P(3)450) (P450 4). | |||||
|
MYG_MOUSE
|
||||||
| θ value | 0.813845 (rank : 33) | NC score | 0.030115 (rank : 62) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P04247 | Gene names | Mb | |||
|
Domain Architecture |

|
|||||
| Description | Myoglobin. | |||||
|
NMI_MOUSE
|
||||||
| θ value | 0.813845 (rank : 34) | NC score | 0.079112 (rank : 23) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O35309, Q99M49 | Gene names | Nmi | |||
|
Domain Architecture |
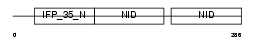
|
|||||
| Description | N-myc-interactor (Nmi) (N-myc and STAT interactor). | |||||
|
PARP1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 35) | NC score | 0.053518 (rank : 54) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P09874, Q8IUZ9 | Gene names | PARP1, ADPRT, PPOL | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 1 (EC 2.4.2.30) (PARP-1) (ADPRT) (NAD(+) ADP-ribosyltransferase 1) (Poly[ADP-ribose] synthetase 1). | |||||
|
SYNE2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 36) | NC score | 0.026604 (rank : 66) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1323 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8WXH0, Q8N1S3, Q8NF49, Q8TER7, Q8WWW3, Q8WWW4, Q8WWW5, Q8WXH1, Q9NU50, Q9UFQ4, Q9Y2L4, Q9Y4R1 | Gene names | SYNE2, KIAA1011, NUA | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-2 (Nuclear envelope spectrin repeat protein 2) (Syne-2) (Synaptic nuclear envelope protein 2) (Nucleus and actin connecting element protein) (Protein NUANCE). | |||||
|
CJ084_MOUSE
|
||||||
| θ value | 1.06291 (rank : 37) | NC score | 0.035212 (rank : 59) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C6C7 | Gene names | D19Ertd737e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf84 homolog. | |||||
|
KIF6_HUMAN
|
||||||
| θ value | 1.06291 (rank : 38) | NC score | 0.017525 (rank : 103) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6ZMV9, Q2MDE3, Q2MDE4, Q5T8J6, Q6ZWE3, Q86T87, Q8WTV4 | Gene names | KIF6, C6orf102 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin-like protein KIF6. | |||||
|
MDR1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 39) | NC score | 0.007989 (rank : 125) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P08183, Q12755, Q14812 | Gene names | ABCB1, MDR1, PGY1 | |||
|
Domain Architecture |

|
|||||
| Description | Multidrug resistance protein 1 (EC 3.6.3.44) (ATP-binding cassette sub-family B member 1) (P-glycoprotein 1) (CD243 antigen). | |||||
|
MYH9_MOUSE
|
||||||
| θ value | 1.06291 (rank : 40) | NC score | 0.020247 (rank : 88) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
RABE1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 41) | NC score | 0.029329 (rank : 63) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1104 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q15276, O95369, Q8IVX3 | Gene names | RABEP1, RABPT5, RABPT5A | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 1 (Rabaptin-5) (Rabaptin-5alpha) (Rabaptin-4) (NY-REN-17 antigen). | |||||
|
RABE1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 42) | NC score | 0.028909 (rank : 64) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O35551, Q99L08, Q9CRP3, Q9EQF9, Q9JI94 | Gene names | Rabep1, Rab5ep, Rabpt5, Rabpt5a | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 1 (Rabaptin-5) (Rabaptin-5alpha). | |||||
|
ARHGC_HUMAN
|
||||||
| θ value | 1.38821 (rank : 43) | NC score | 0.011812 (rank : 114) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NZN5, O15086 | Gene names | ARHGEF12, KIAA0382, LARG | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 12 (Leukemia-associated RhoGEF). | |||||
|
CTRO_HUMAN
|
||||||
| θ value | 1.38821 (rank : 44) | NC score | 0.010484 (rank : 117) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 2092 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O14578, Q6XUH8, Q86UQ9, Q9UPZ7 | Gene names | CIT, KIAA0949, STK21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
DICER_HUMAN
|
||||||
| θ value | 1.38821 (rank : 45) | NC score | 0.018097 (rank : 101) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UPY3, O95943, Q9UQ02 | Gene names | DICER1, DICER, HERNA, KIAA0928 | |||
|
Domain Architecture |

|
|||||
| Description | Endoribonuclease Dicer (EC 3.1.26.-) (Helicase with RNase motif) (Helicase-MOI). | |||||
|
MYH9_HUMAN
|
||||||
| θ value | 1.38821 (rank : 46) | NC score | 0.018393 (rank : 98) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
NUAK2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 47) | NC score | 0.001438 (rank : 136) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9H093 | Gene names | NUAK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NUAK family SNF1-like kinase 2 (EC 2.7.11.1) (SNF1/AMP kinase-related kinase) (SNARK). | |||||
|
RON_MOUSE
|
||||||
| θ value | 1.38821 (rank : 48) | NC score | 0.001337 (rank : 137) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q62190, Q62555 | Gene names | Mst1r, Ron, Stk | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage-stimulating protein receptor precursor (EC 2.7.10.1) (MSP receptor) (p185-Ron) (Stem cell-derived tyrosine kinase) (CDw136 antigen) [Contains: Macrophage-stimulating protein receptor alpha chain; Macrophage-stimulating protein receptor beta chain]. | |||||
|
SYCP1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 49) | NC score | 0.023931 (rank : 73) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1313 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q15431, O14963 | Gene names | SYCP1, SCP1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
TRM61_MOUSE
|
||||||
| θ value | 1.38821 (rank : 50) | NC score | 0.049571 (rank : 55) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80XC2, Q8BMD4, Q8BX33 | Gene names | Trm61 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA (adenine-N(1)-)-methyltransferase catalytic subunit TRM61 (EC 2.1.1.36) (tRNA(m1A58)-methyltransferase subunit TRM61) (tRNA(m1A58)MTase subunit TRM61). | |||||
|
ATRIP_HUMAN
|
||||||
| θ value | 1.81305 (rank : 51) | NC score | 0.026092 (rank : 68) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8WXE1, Q8NHQ2, Q8WUG7, Q96CL3, Q9HA30 | Gene names | ATRIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATR-interacting protein (ATM and Rad3-related-interacting protein). | |||||
|
CENPE_HUMAN
|
||||||
| θ value | 1.81305 (rank : 52) | NC score | 0.023063 (rank : 76) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
FYCO1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 53) | NC score | 0.025249 (rank : 70) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8VDC1, Q3V385, Q7TMT0, Q8BJN2 | Gene names | Fyco1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1. | |||||
|
MBD4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 54) | NC score | 0.023392 (rank : 75) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95243, Q7Z4T3, Q96F09 | Gene names | MBD4, MED1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 4 (EC 3.2.2.-) (Methyl-CpG-binding protein MBD4) (Methyl-CpG-binding endonuclease 1) (Mismatch-specific DNA N-glycosylase). | |||||
|
MRCKA_HUMAN
|
||||||
| θ value | 1.81305 (rank : 55) | NC score | 0.008957 (rank : 120) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1999 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q5VT25, Q59GZ1, Q5H9N9, Q5T797, Q5VT26, Q5VT27, Q86XX2, Q86XX3, Q99646 | Gene names | CDC42BPA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK alpha (EC 2.7.11.1) (CDC42- binding protein kinase alpha) (Myotonic dystrophy kinase-related CDC42-binding kinase alpha) (Myotonic dystrophy protein kinase-like alpha) (MRCK alpha) (DMPK-like alpha). | |||||
|
SNX25_HUMAN
|
||||||
| θ value | 1.81305 (rank : 56) | NC score | 0.025803 (rank : 69) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H3E2 | Gene names | SNX25 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-25. | |||||
|
TOLIP_MOUSE
|
||||||
| θ value | 1.81305 (rank : 57) | NC score | 0.032461 (rank : 60) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QZ06, Q543N1, Q9D5P5 | Gene names | Tollip | |||
|
Domain Architecture |
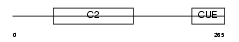
|
|||||
| Description | Toll-interacting protein. | |||||
|
GNAT1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 58) | NC score | 0.008343 (rank : 122) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P11488 | Gene names | GNAT1, GNATR | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-binding protein G(t), alpha-1 subunit (Transducin alpha-1 chain). | |||||
|
KIF4A_HUMAN
|
||||||
| θ value | 2.36792 (rank : 59) | NC score | 0.018987 (rank : 96) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O95239, Q86TN3, Q86XX7, Q9NNY6, Q9NY24, Q9UMW3 | Gene names | KIF4A, KIF4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
MDN1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 60) | NC score | 0.022601 (rank : 80) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9NU22, O15019 | Gene names | MDN1, KIAA0301 | |||
|
Domain Architecture |

|
|||||
| Description | Midasin (MIDAS-containing protein). | |||||
|
NSBP1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 61) | NC score | 0.012497 (rank : 112) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
SH3G3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 62) | NC score | 0.016541 (rank : 105) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q62421 | Gene names | Sh3gl3, Sh3d2c, Sh3d2c2 | |||
|
Domain Architecture |
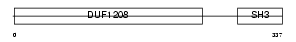
|
|||||
| Description | SH3-containing GRB2-like protein 3 (Endophilin-3) (Endophilin-A3) (SH3 domain protein 2C) (SH3p13). | |||||
|
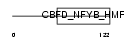
TAF12_MOUSE
|
||||||
| θ value | 2.36792 (rank : 63) | NC score | 0.028457 (rank : 65) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VE65, Q3V0S7, Q9CZ11, Q9D8Q0 | Gene names | Taf12 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription initiation factor TFIID subunit 12 (Transcription initiation factor TFIID 20 kDa subunits) (TAFII-20) (TAFII20). | |||||
|
TXLNB_HUMAN
|
||||||
| θ value | 2.36792 (rank : 64) | NC score | 0.020317 (rank : 87) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 912 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8N3L3, Q76L25, Q86T52, Q8N3S2 | Gene names | TXLNB, C6orf198, MDP77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77) (hMDP77). | |||||
|
CCD46_MOUSE
|
||||||
| θ value | 3.0926 (rank : 65) | NC score | 0.019598 (rank : 94) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q5PR68, Q80ZN6, Q99JS4, Q9D9T1 | Gene names | Ccdc46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled coil domain-containing protein 46. | |||||
|
PTBP2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 66) | NC score | 0.015035 (rank : 109) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UKA9, Q8N0Z1, Q8N160, Q8NFB0, Q8NFB1, Q969N9, Q96Q76 | Gene names | PTBP2, NPTB, PTB, PTBLP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypyrimidine tract-binding protein 2 (Neurally-enriched homolog of PTB) (Neural polypyrimidine tract-binding protein) (PTB-like protein). | |||||
|
PTBP2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 67) | NC score | 0.015056 (rank : 108) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91Z31, Q7TNW7, Q9CUW2, Q9QYC2, Q9R0V9 | Gene names | Ptbp2, Brptb, Nptb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypyrimidine tract-binding protein 2 (Brain-enriched polypyrimidine tract-binding protein) (Brain-enriched PTB) (RRM-type RNA-binding protein brPTB) (Neural polypyrimidine tract-binding protein). | |||||
|
T106C_MOUSE
|
||||||
| θ value | 3.0926 (rank : 68) | NC score | 0.024317 (rank : 71) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80VP8 | Gene names | Tmem106c, D15Ertd405e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 106C. | |||||
|
THNSL_HUMAN
|
||||||
| θ value | 3.0926 (rank : 69) | NC score | 0.018469 (rank : 97) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8IYQ7, Q5VV21 | Gene names | THNSL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Threonine synthase-like 1. | |||||
|
TM9S3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 70) | NC score | 0.016402 (rank : 106) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9HD45, Q6UWE7, Q9NWL8, Q9P0G9, Q9UHW8 | Gene names | TM9SF3, SMBP | |||
|
Domain Architecture |

|
|||||
| Description | Transmembrane 9 superfamily protein member 3 precursor (SM-11044- binding protein) (EP70-P-iso). | |||||
|
ZN630_HUMAN
|
||||||
| θ value | 3.0926 (rank : 71) | NC score | 0.000140 (rank : 139) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 737 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q2M218 | Gene names | ZNF630 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 630. | |||||
|
C8AP2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.015441 (rank : 107) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UKL3, Q6PH76, Q7LCQ7, Q86YD9, Q9NUQ4, Q9NZV9, Q9P2N1, Q9Y563 | Gene names | CASP8AP2, FLASH, KIAA1315, RIP25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
GCC2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 73) | NC score | 0.020245 (rank : 90) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1619 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8CHG3, Q8BR44, Q8R2Q5, Q9CT45 | Gene names | Gcc2, Kiaa0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185). | |||||
|
GNAT1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 74) | NC score | 0.007426 (rank : 129) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P20612, Q80X34 | Gene names | Gnat1, Gnat-1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-binding protein G(t), alpha-1 subunit (Transducin alpha-1 chain). | |||||
|
KINH_MOUSE
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | 0.018149 (rank : 100) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1082 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q61768, O08711, Q61580 | Gene names | Kif5b, Khcs, Kns1 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin heavy chain (Ubiquitous kinesin heavy chain) (UKHC). | |||||
|
NOL10_HUMAN
|
||||||
| θ value | 4.03905 (rank : 76) | NC score | 0.020040 (rank : 93) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9BSC4, Q53RC9, Q96TA5, Q9H7Y7, Q9H855 | Gene names | NOL10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar protein 10. | |||||
|
PARP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 77) | NC score | 0.045132 (rank : 56) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P11103, Q9JLX4, Q9QVQ3 | Gene names | Parp1, Adprp, Adprt, Adprt1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 1 (EC 2.4.2.30) (PARP-1) (ADPRT) (NAD(+) ADP-ribosyltransferase 1) (Poly[ADP-ribose] synthetase 1) (msPARP). | |||||
|
RT35_HUMAN
|
||||||
| θ value | 4.03905 (rank : 78) | NC score | 0.020165 (rank : 91) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P82673, Q32LZ1, Q6P4C6, Q7L1M6, Q96AI0, Q9H044, Q9HC14, Q9P1R5 | Gene names | MRPS35, MRPS28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 28S ribosomal protein S35, mitochondrial precursor (S35mt) (MRP-S35) (Mitochondrial ribosomal protein S28) (MRP-S28). | |||||
|
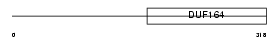
CCDC6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.022012 (rank : 83) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 723 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q16204, Q15250 | Gene names | CCDC6, D10S170 | |||
|
Domain Architecture |
|
|||||
| Description | Coiled-coil domain-containing protein 6 (H4 protein) (Papillary thyroid carcinoma-encoded protein). | |||||
|
CF152_MOUSE
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | 0.020782 (rank : 86) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q80ST9, Q9CYM9, Q9D5J9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf152 homolog. | |||||
|
COHA1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.008900 (rank : 121) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q07563, Q99LK8 | Gene names | Col17a1, Bp180, Bpag2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVII) chain (Bullous pemphigoid antigen 2) (180 kDa bullous pemphigoid antigen 2). | |||||
|
CP250_MOUSE
|
||||||
| θ value | 5.27518 (rank : 82) | NC score | 0.022296 (rank : 81) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1583 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q60952, Q2I8G3, Q3UTR4, Q6PFF6, Q8BLC6 | Gene names | Cep250, Cep2, Inmp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1) (Intranuclear matrix protein). | |||||
|
EP15R_MOUSE
|
||||||
| θ value | 5.27518 (rank : 83) | NC score | 0.016788 (rank : 104) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q60902, Q8CB60, Q8CB70, Q91WH8 | Gene names | Eps15l1, Eps15-rs, Eps15R | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal growth factor receptor substrate 15-like 1 (Eps15-related protein) (Eps15R) (Epidermal growth factor receptor pathway substrate 15-related sequence) (Eps15-rs). | |||||
|
IF3A_MOUSE
|
||||||
| θ value | 5.27518 (rank : 84) | NC score | 0.021090 (rank : 84) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 871 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P23116, Q60697, Q62162 | Gene names | Eif3s10, Csma, Eif3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3 p185) (eIF3a) (p162 protein) (Centrosomin). | |||||
|
KS6A1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 85) | NC score | 0.000491 (rank : 138) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 850 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P18653 | Gene names | Rps6ka1, Rsk1 | |||
|
Domain Architecture |
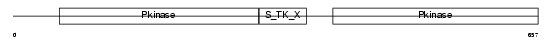
|
|||||
| Description | Ribosomal protein S6 kinase alpha-1 (EC 2.7.11.1) (S6K-alpha 1) (90 kDa ribosomal protein S6 kinase 1) (p90-RSK 1) (Ribosomal S6 kinase 1) (RSK-1) (pp90RSK1) (p90S6K) (MAP kinase-activated protein kinase 1a) (MAPKAPK1A). | |||||
|
LETM1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 86) | NC score | 0.022028 (rank : 82) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Z2I0, Q5PQC5, Q7TMK8, Q80ZX6, Q8CGJ3, Q8K5E5 | Gene names | Letm1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper-EF-hand-containing transmembrane protein 1, mitochondrial precursor. | |||||
|
STA13_HUMAN
|
||||||
| θ value | 5.27518 (rank : 87) | NC score | 0.007414 (rank : 130) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y3M8, Q5HYH1, Q5TAE3, Q6UN61, Q86TP6, Q86WQ3, Q86XT1 | Gene names | STARD13, DLC2, GT650 | |||
|
Domain Architecture |

|
|||||
| Description | StAR-related lipid transfer protein 13 (StARD13) (START domain- containing protein 13) (46H23.2) (Deleted in liver cancer protein 2) (Rho GTPase-activating protein). | |||||
|
SYNE1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 88) | NC score | 0.022771 (rank : 79) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
TMF1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 89) | NC score | 0.022910 (rank : 78) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 987 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P82094 | Gene names | TMF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA element modulatory factor (TMF). | |||||
|
ANR26_HUMAN
|
||||||
| θ value | 6.88961 (rank : 90) | NC score | 0.023594 (rank : 74) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1614 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UPS8, Q2TAZ3, Q6ZR14, Q9H1Q1, Q9NSK9, Q9NTD5, Q9NW69 | Gene names | ANKRD26, KIAA1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
BIRC6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 91) | NC score | 0.009421 (rank : 119) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NR09, Q9ULD1 | Gene names | BIRC6, KIAA1289 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 6 (Ubiquitin-conjugating BIR-domain enzyme apollon). | |||||
|
CENPF_HUMAN
|
||||||
| θ value | 6.88961 (rank : 92) | NC score | 0.026314 (rank : 67) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
CTNA3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | 0.006909 (rank : 131) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q65CL1, Q3UQ61, Q8C0N3 | Gene names | Ctnna3, Catna3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Catenin alpha-3 (Alpha T-catenin) (Cadherin-associated protein). | |||||
|
DUS3L_HUMAN
|
||||||
| θ value | 6.88961 (rank : 94) | NC score | 0.010687 (rank : 116) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96G46, Q96HM5, Q9BSU4, Q9H877, Q9NPR1 | Gene names | DUS3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA-dihydrouridine synthase 3-like (EC 1.-.-.-). | |||||
|
EEA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 95) | NC score | 0.019321 (rank : 95) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1796 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q15075, Q14221 | Gene names | EEA1, ZFYVE2 | |||
|
Domain Architecture |

|
|||||
| Description | Early endosome antigen 1 (Endosome-associated protein p162) (Zinc finger FYVE domain-containing protein 2). | |||||
|
EMIL2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 96) | NC score | 0.014050 (rank : 110) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8K482 | Gene names | Emilin2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Basilin). | |||||
|
ICAL_HUMAN
|
||||||
| θ value | 6.88961 (rank : 97) | NC score | 0.011998 (rank : 113) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P20810, O95360, Q96D08, Q9H1Z5 | Gene names | CAST | |||
|
Domain Architecture |

|
|||||
| Description | Calpastatin (Calpain inhibitor) (Sperm BS-17 component). | |||||
|
MACF1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 98) | NC score | 0.017580 (rank : 102) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1341 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9UPN3, O75053, Q5VW20, Q8WXY2, Q9H540, Q9UKP0, Q9ULG9 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoforms 1/2/3/5 (Actin cross-linking family protein 7) (Macrophin-1) (Trabeculin-alpha) (620 kDa actin-binding protein) (ABP620). | |||||
|
MACF4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 99) | NC score | 0.020814 (rank : 85) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1321 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q96PK2, Q8WXY1 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoform 4. | |||||
|
MMRN1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 100) | NC score | 0.022993 (rank : 77) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q13201, Q6P3T8 | Gene names | MMRN1, ECM, EMILIN4, MMRN | |||
|
Domain Architecture |

|
|||||
| Description | Multimerin-1 precursor (Endothelial cell multimerin 1) (EMILIN-4) (Elastin microfibril interface located protein 4) (Elastin microfibril interfacer 4). | |||||
|
NOS1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 101) | NC score | 0.005222 (rank : 133) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z0J4, Q64208 | Gene names | Nos1 | |||
|
Domain Architecture |

|
|||||
| Description | Nitric-oxide synthase, brain (EC 1.14.13.39) (NOS type I) (Neuronal NOS) (N-NOS) (nNOS) (Constitutive NOS) (NC-NOS) (bNOS). | |||||
|
REST_HUMAN
|
||||||
| θ value | 6.88961 (rank : 102) | NC score | 0.018237 (rank : 99) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1605 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P30622 | Gene names | RSN, CYLN1 | |||
|
Domain Architecture |

|
|||||
| Description | Restin (Cytoplasmic linker protein 170 alpha-2) (CLIP-170) (Reed- Sternberg intermediate filament-associated protein) (Cytoplasmic linker protein 1). | |||||
|
RHG18_HUMAN
|
||||||
| θ value | 6.88961 (rank : 103) | NC score | 0.008333 (rank : 123) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8N392, Q58EZ3, Q6P679, Q6PJD7, Q96S64 | Gene names | ARHGAP18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho-GTPase-activating protein 18 (MacGAP). | |||||
|
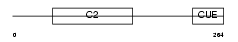
TOLIP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 104) | NC score | 0.024278 (rank : 72) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H0E2, Q9H9E6, Q9UJ69 | Gene names | TOLLIP | |||
|
Domain Architecture |
|
|||||
| Description | Toll-interacting protein. | |||||
|
IF3A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 105) | NC score | 0.020158 (rank : 92) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 875 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q14152, O00653 | Gene names | EIF3S10, KIAA0139 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3 p185) (eIF3a). | |||||
|
IPO8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 106) | NC score | 0.006400 (rank : 132) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15397 | Gene names | IPO8, RANBP8 | |||
|
Domain Architecture |
|
|||||
| Description | Importin-8 (Imp8) (Ran-binding protein 8) (RanBP8). | |||||
|
KI13A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 107) | NC score | 0.011366 (rank : 115) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9H1H9, Q9H1H8 | Gene names | KIF13A, RBKIN | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13A (Kinesin-like protein RBKIN). | |||||
|
RNF34_MOUSE
|
||||||
| θ value | 8.99809 (rank : 108) | NC score | 0.007847 (rank : 127) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99KR6, Q3UV45 | Gene names | Rnf34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 34 (RING finger protein RIFF) (Phafin-1). | |||||
|
SCFD2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 109) | NC score | 0.009985 (rank : 118) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8WU76, Q8N5F3, Q8N8H0, Q96ED3 | Gene names | SCFD2, STXBP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Sec1 family domain-containing protein 2 (Syntaxin-binding protein 1- like 1). | |||||
|
SMCA5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 110) | NC score | 0.002693 (rank : 135) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60264 | Gene names | SMARCA5, SNF2H, WCRF135 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 5 (EC 3.6.1.-) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin A5) (Sucrose nonfermenting protein 2 homolog) (hSNF2H). | |||||
|
STA5A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 111) | NC score | 0.007939 (rank : 126) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P42230 | Gene names | Stat5a, Mgf, Mpf | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 5A (Mammary gland factor). | |||||
|
STA5B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 112) | NC score | 0.008090 (rank : 124) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P42232, Q541Q5, Q60804, Q8K3Q1, Q9JKM1 | Gene names | Stat5b | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 5B. | |||||
|
TM9S3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 113) | NC score | 0.012694 (rank : 111) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ET30 | Gene names | Tm9sf3, Smbp | |||
|
Domain Architecture |

|
|||||
| Description | Transmembrane 9 superfamily protein member 3 precursor. | |||||
|
H2A1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.065953 (rank : 31) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96QV6 | Gene names | HIST1H2AA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 1-A. | |||||
|
H2A1B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.064586 (rank : 41) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P04908 | Gene names | HIST1H2AB, H2AFM | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A type 1-B. | |||||
|
H2A1C_HUMAN
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.064280 (rank : 49) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q93077, O00775, O00776, O00777, O00778, Q540R1 | Gene names | HIST1H2AC, H2AFL | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A type 1-C. | |||||
|
H2A1D_HUMAN
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.064433 (rank : 45) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P20671, P57754, Q6FGY6 | Gene names | HIST1H2AD, H2AFG | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A type 1-D (H2A.3). | |||||
|
H2A1E_HUMAN
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.064462 (rank : 43) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P28001, Q76P63 | Gene names | HIST1H2AE, H2AFA | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A type 1-E (H2A.2) (H2A/m). | |||||
|
H2A1F_MOUSE
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.064520 (rank : 42) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CGP5 | Gene names | Hist1h2af | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 1-F. | |||||
|
H2A1H_HUMAN
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.064678 (rank : 37) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96KK5 | Gene names | HIST1H2AH, HIST1H2AI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 1-H. | |||||
|
H2A1H_MOUSE
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.064812 (rank : 35) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CGP6 | Gene names | Hist1h2ah | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 1-H. | |||||
|
H2A1J_HUMAN
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.064804 (rank : 36) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99878, Q5JXQ5 | Gene names | HIST1H2AJ, H2AFE | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A type 1-J. | |||||
|
H2A1K_MOUSE
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.064593 (rank : 40) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CGP7 | Gene names | Hist1h2ak | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 1-K. | |||||
|
H2A1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.064328 (rank : 48) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P0C0S8, P02261, Q2M1R2, Q76PA6 | Gene names | HIST1H2AG, H2AFP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 1 (H2A.1). | |||||
|
H2A1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.064462 (rank : 44) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P22752, P10812, Q5SZZ2 | Gene names | Hist1h2ab | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A type 1. | |||||
|
H2A2A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.064600 (rank : 38) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6FI13, P20670 | Gene names | HIST2H2AA3, H2AFO, HIST2H2AA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 2-A (H2A.2). | |||||
|
H2A2A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.064600 (rank : 39) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6GSS7, P20670, Q811M3 | Gene names | Hist2h2aa1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 2-A (H2A.2) (H2a-614) (H2a-615). | |||||
|
H2A2B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.065920 (rank : 32) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IUE6 | Gene names | HIST2H2AB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 2-B. | |||||
|
H2A2B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.066270 (rank : 30) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64522 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 2-B (H2a-613A). | |||||
|
H2A2C_HUMAN
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.065228 (rank : 33) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q16777, Q6DRA7, Q8IUE5 | Gene names | HIST2H2AC, H2AFQ | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A type 2-C (H2A-GL101) (H2A/r). | |||||
|
H2A2C_MOUSE
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.065228 (rank : 34) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64523, Q8CGP3 | Gene names | Hist2h2ac, Hist2h2ab | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 2-C (H2a-613B). | |||||
|
H2A3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.064384 (rank : 46) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7L7L0 | Gene names | HIST3H2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 3. | |||||
|
H2A3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.064384 (rank : 47) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BFU2 | Gene names | Hist3h2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 3. | |||||
|
H2AFB_HUMAN
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.063822 (rank : 51) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P98176, Q5TZB2, Q6FG78, Q96PR7 | Gene names | H2AFB3, H2ABBD, H2AFB | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A-Bbd (H2A Barr body-deficient) (H2A.Bbd). | |||||
|
H2AV_HUMAN
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.069199 (rank : 26) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q71UI9, Q59GV8, Q6PK98 | Gene names | H2AFV, H2AV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2AV (H2A.F/Z). | |||||
|
H2AV_MOUSE
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.069199 (rank : 27) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3THW5, Q5NC92, Q6ZWU0 | Gene names | H2afv, H2av | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2AV (H2A.F/Z). | |||||
|
H2AX_HUMAN
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.067347 (rank : 29) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P16104, Q4ZGJ7, Q6IAS5 | Gene names | H2AFX, H2AX | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A.x (H2a/x). | |||||
|
H2AX_MOUSE
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.067617 (rank : 28) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P27661 | Gene names | H2afx, H2a.x, H2ax, Hist5-2ax | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A.x (H2a/x). | |||||
|
H2AZ_HUMAN
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.069319 (rank : 24) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P0C0S5, P17317, Q6I9U0 | Gene names | H2AFZ, H2AZ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A.Z (H2A/z). | |||||
|
H2AZ_MOUSE
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.069319 (rank : 25) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P0C0S6, P17317, Q3UA55, Q3UK43, Q68FD2 | Gene names | H2afz, H2az | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A.Z (H2A/z). | |||||
|
PAR14_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 113 | |
| SwissProt Accessions | Q2EMV9, Q571C6, Q8BN35, Q8VDT6 | Gene names | Parp14, Kiaa1268 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 14 (EC 2.4.2.30) (PARP-14) (Collaborator of STAT6) (CoaSt6). | |||||
|
PAR14_HUMAN
|
||||||
| NC score | 0.983019 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q460N5, Q460N4, Q8J027, Q9H9X9, Q9NV60, Q9ULF2 | Gene names | PARP14, BAL2, KIAA1268 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 14 (EC 2.4.2.30) (PARP-14) (B aggressive lymphoma protein 2). | |||||
|
PAR15_HUMAN
|
||||||
| NC score | 0.925679 (rank : 3) | θ value | 8.73085e-104 (rank : 3) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q460N3, Q8N1K3 | Gene names | PARP15, BAL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 15 (EC 2.4.2.30) (PARP-15) (B-aggressive lymphoma protein 3). | |||||
|
PARP9_HUMAN
|
||||||
| NC score | 0.810188 (rank : 4) | θ value | 7.75577e-52 (rank : 5) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8IXQ6, Q8TCP3, Q9BZL8, Q9BZL9 | Gene names | PARP9, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 9 (EC 2.4.2.30) (PARP-9) (B aggressive lymphoma protein). | |||||
|
PARP9_MOUSE
|
||||||
| NC score | 0.807987 (rank : 5) | θ value | 1.36585e-56 (rank : 4) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8CAS9, Q99LF9 | Gene names | Parp9, Bal | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 9 (EC 2.4.2.30) (PARP-9) (B aggressive lymphoma protein homolog). | |||||
|
PARPT_MOUSE
|
||||||
| NC score | 0.699786 (rank : 6) | θ value | 6.38894e-30 (rank : 10) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8C1B2, Q3UD50, Q8C032 | Gene names | Tiparp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TCDD-inducible poly [ADP-ribose] polymerase (EC 2.4.2.30). | |||||
|
PARPT_HUMAN
|
||||||
| NC score | 0.699559 (rank : 7) | θ value | 6.38894e-30 (rank : 9) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q7Z3E1, Q68CY9, Q86VP4, Q9Y4P7 | Gene names | TIPARP, PARP7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TCDD-inducible poly [ADP-ribose] polymerase (EC 2.4.2.30) (Poly [ADP- ribose] polymerase 7) (PARP-7). | |||||
|
PAR12_MOUSE
|
||||||
| NC score | 0.689306 (rank : 8) | θ value | 8.91499e-32 (rank : 7) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BZ20, Q80VL6, Q8K333, Q8R3U2 | Gene names | Parp12, Zc3hdc1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 12 (EC 2.4.2.30) (PARP-12) (Zinc finger CCCH domain-containing protein 1). | |||||
|
PAR12_HUMAN
|
||||||
| NC score | 0.685490 (rank : 9) | θ value | 1.98606e-31 (rank : 8) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9H0J9, Q9H610, Q9NP36, Q9NTI3 | Gene names | PARP12, ZC3HDC1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 12 (EC 2.4.2.30) (PARP-12) (Zinc finger CCCH domain-containing protein 1). | |||||
|
PAR10_HUMAN
|
||||||
| NC score | 0.679239 (rank : 10) | θ value | 4.72714e-33 (rank : 6) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q53GL7, Q8N2I0, Q8WV05, Q96CH7 | Gene names | PARP10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 10 (EC 2.4.2.30) (PARP-10). | |||||
|
ZCC2_HUMAN
|
||||||
| NC score | 0.641376 (rank : 11) | θ value | 6.62687e-19 (rank : 11) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7Z2W4, Q8IW57, Q8TAJ3, Q96N79, Q9H8R9, Q9P0Y7 | Gene names | ZC3HAV1, ZC3HDC2 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger CCCH type antiviral protein 1 (Zinc finger CCCH domain- containing protein 2). | |||||
|
LRP16_HUMAN
|
||||||
| NC score | 0.546403 (rank : 12) | θ value | 9.56915e-18 (rank : 12) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BQ69, Q9UH96 | Gene names | LRP16 | |||
|
Domain Architecture |

|
|||||
| Description | Protein LRP16. | |||||
|
LRP16_MOUSE
|
||||||
| NC score | 0.534480 (rank : 13) | θ value | 2.60593e-15 (rank : 13) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q922B1 | Gene names | Lrp16 | |||
|
Domain Architecture |

|
|||||
| Description | Protein LRP16. | |||||
|
H2AY_HUMAN
|
||||||
| NC score | 0.296154 (rank : 14) | θ value | 3.76295e-14 (rank : 14) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75367, O75377, Q503A8, Q7Z5E3, Q96D41, Q9H8P3, Q9UP96 | Gene names | H2AFY, MACROH2A1 | |||
|
Domain Architecture |

|
|||||
| Description | Core histone macro-H2A.1 (Histone macroH2A1) (mH2A1) (H2A.y) (H2A/y) (Medulloblastoma antigen MU-MB-50.205). | |||||
|
H2AY_MOUSE
|
||||||
| NC score | 0.295573 (rank : 15) | θ value | 4.91457e-14 (rank : 15) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9QZQ8, Q91VZ2, Q9QZQ7 | Gene names | H2afy | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Core histone macro-H2A.1 (Histone macroH2A1) (mH2A1) (H2A.y) (H2A/y). | |||||
|
H2AW_HUMAN
|
||||||
| NC score | 0.244232 (rank : 16) | θ value | 8.67504e-11 (rank : 16) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9P0M6, Q5SQT2 | Gene names | H2AFY2, MACROH2A2 | |||
|
Domain Architecture |

|
|||||
| Description | Core histone macro-H2A.2 (Histone macroH2A2) (mH2A2). | |||||
|
H2AW_MOUSE
|
||||||
| NC score | 0.241829 (rank : 17) | θ value | 8.67504e-11 (rank : 17) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8CCK0, Q3TNT4, Q925I6 | Gene names | H2afy2, H2afy3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Core histone macro-H2A.2 (Histone macroH2A2) (mH2A2). | |||||
|
PARP3_HUMAN
|
||||||
| NC score | 0.140620 (rank : 18) | θ value | 0.0113563 (rank : 22) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y6F1, Q8NER9, Q96CG2, Q9UG81 | Gene names | PARP3, ADPRT3, ADPRTL3 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 3 (EC 2.4.2.30) (PARP-3) (NAD(+) ADP- ribosyltransferase 3) (Poly[ADP-ribose] synthetase 3) (pADPRT-3) (hPARP-3) (IRT1). | |||||
|
DTX3L_HUMAN
|
||||||
| NC score | 0.134902 (rank : 19) | θ value | 1.53129e-07 (rank : 19) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8TDB6 | Gene names | DTX3L, BBAP | |||
|
Domain Architecture |

|
|||||
| Description | Protein deltex 3-like protein (B-lymphoma- and BAL-associated protein) (Rhysin-2) (Rhysin2). | |||||
|
TNKS2_HUMAN
|
||||||
| NC score | 0.134615 (rank : 20) | θ value | 4.76016e-09 (rank : 18) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H2K2, Q9H8F2, Q9HAS4 | Gene names | TNKS2, PARP5B, TANK2, TNKL | |||
|
Domain Architecture |

|
|||||
| Description | Tankyrase-2 (EC 2.4.2.30) (TANK2) (Tankyrase II) (TNKS-2) (TRF1- interacting ankyrin-related ADP-ribose polymerase 2) (Tankyrase-like protein) (Tankyrase-related protein). | |||||
|
TNKS1_HUMAN
|
||||||
| NC score | 0.114106 (rank : 21) | θ value | 1.69304e-06 (rank : 20) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O95271, O95272 | Gene names | TNKS, PARP5A, PARPL, TIN1, TINF1, TNKS1 | |||
|
Domain Architecture |

|
|||||
| Description | Tankyrase-1 (EC 2.4.2.30) (TANK1) (Tankyrase I) (TNKS-1) (TRF1- interacting ankyrin-related ADP-ribose polymerase). | |||||
|
NMI_HUMAN
|
||||||
| NC score | 0.105963 (rank : 22) | θ value | 0.00665767 (rank : 21) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q13287, Q53TI8, Q9BVE5 | Gene names | NMI | |||
|
Domain Architecture |

|
|||||
| Description | N-myc-interactor (Nmi) (N-myc and STAT interactor). | |||||
|
NMI_MOUSE
|
||||||
| NC score | 0.079112 (rank : 23) | θ value | 0.813845 (rank : 34) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O35309, Q99M49 | Gene names | Nmi | |||
|
Domain Architecture |

|
|||||
| Description | N-myc-interactor (Nmi) (N-myc and STAT interactor). | |||||
|
H2AZ_HUMAN
|
||||||
| NC score | 0.069319 (rank : 24) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P0C0S5, P17317, Q6I9U0 | Gene names | H2AFZ, H2AZ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A.Z (H2A/z). | |||||
|
H2AZ_MOUSE
|
||||||
| NC score | 0.069319 (rank : 25) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P0C0S6, P17317, Q3UA55, Q3UK43, Q68FD2 | Gene names | H2afz, H2az | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A.Z (H2A/z). | |||||
|
H2AV_HUMAN
|
||||||
| NC score | 0.069199 (rank : 26) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q71UI9, Q59GV8, Q6PK98 | Gene names | H2AFV, H2AV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2AV (H2A.F/Z). | |||||
|
H2AV_MOUSE
|
||||||
| NC score | 0.069199 (rank : 27) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3THW5, Q5NC92, Q6ZWU0 | Gene names | H2afv, H2av | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2AV (H2A.F/Z). | |||||
|
H2AX_MOUSE
|
||||||
| NC score | 0.067617 (rank : 28) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P27661 | Gene names | H2afx, H2a.x, H2ax, Hist5-2ax | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A.x (H2a/x). | |||||
|
H2AX_HUMAN
|
||||||
| NC score | 0.067347 (rank : 29) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P16104, Q4ZGJ7, Q6IAS5 | Gene names | H2AFX, H2AX | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A.x (H2a/x). | |||||
|
H2A2B_MOUSE
|
||||||
| NC score | 0.066270 (rank : 30) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64522 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 2-B (H2a-613A). | |||||
|
H2A1A_HUMAN
|
||||||
| NC score | 0.065953 (rank : 31) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96QV6 | Gene names | HIST1H2AA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 1-A. | |||||
|
H2A2B_HUMAN
|
||||||
| NC score | 0.065920 (rank : 32) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IUE6 | Gene names | HIST2H2AB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 2-B. | |||||
|
H2A2C_HUMAN
|
||||||
| NC score | 0.065228 (rank : 33) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q16777, Q6DRA7, Q8IUE5 | Gene names | HIST2H2AC, H2AFQ | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A type 2-C (H2A-GL101) (H2A/r). | |||||
|
H2A2C_MOUSE
|
||||||
| NC score | 0.065228 (rank : 34) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64523, Q8CGP3 | Gene names | Hist2h2ac, Hist2h2ab | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 2-C (H2a-613B). | |||||
|
H2A1H_MOUSE
|
||||||
| NC score | 0.064812 (rank : 35) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CGP6 | Gene names | Hist1h2ah | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 1-H. | |||||
|
H2A1J_HUMAN
|
||||||
| NC score | 0.064804 (rank : 36) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99878, Q5JXQ5 | Gene names | HIST1H2AJ, H2AFE | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A type 1-J. | |||||
|
H2A1H_HUMAN
|
||||||
| NC score | 0.064678 (rank : 37) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96KK5 | Gene names | HIST1H2AH, HIST1H2AI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 1-H. | |||||
|
H2A2A_HUMAN
|
||||||
| NC score | 0.064600 (rank : 38) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6FI13, P20670 | Gene names | HIST2H2AA3, H2AFO, HIST2H2AA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 2-A (H2A.2). | |||||
|
H2A2A_MOUSE
|
||||||
| NC score | 0.064600 (rank : 39) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6GSS7, P20670, Q811M3 | Gene names | Hist2h2aa1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 2-A (H2A.2) (H2a-614) (H2a-615). | |||||
|
H2A1K_MOUSE
|
||||||
| NC score | 0.064593 (rank : 40) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CGP7 | Gene names | Hist1h2ak | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 1-K. | |||||
|
H2A1B_HUMAN
|
||||||
| NC score | 0.064586 (rank : 41) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P04908 | Gene names | HIST1H2AB, H2AFM | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A type 1-B. | |||||
|
H2A1F_MOUSE
|
||||||
| NC score | 0.064520 (rank : 42) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CGP5 | Gene names | Hist1h2af | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 1-F. | |||||
|
H2A1E_HUMAN
|
||||||
| NC score | 0.064462 (rank : 43) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P28001, Q76P63 | Gene names | HIST1H2AE, H2AFA | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A type 1-E (H2A.2) (H2A/m). | |||||
|
H2A1_MOUSE
|
||||||
| NC score | 0.064462 (rank : 44) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P22752, P10812, Q5SZZ2 | Gene names | Hist1h2ab | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A type 1. | |||||
|
H2A1D_HUMAN
|
||||||
| NC score | 0.064433 (rank : 45) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P20671, P57754, Q6FGY6 | Gene names | HIST1H2AD, H2AFG | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A type 1-D (H2A.3). | |||||
|
H2A3_HUMAN
|
||||||
| NC score | 0.064384 (rank : 46) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7L7L0 | Gene names | HIST3H2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 3. | |||||
|
H2A3_MOUSE
|
||||||
| NC score | 0.064384 (rank : 47) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BFU2 | Gene names | Hist3h2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 3. | |||||
|
H2A1_HUMAN
|
||||||
| NC score | 0.064328 (rank : 48) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P0C0S8, P02261, Q2M1R2, Q76PA6 | Gene names | HIST1H2AG, H2AFP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone H2A type 1 (H2A.1). | |||||
|
H2A1C_HUMAN
|
||||||
| NC score | 0.064280 (rank : 49) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q93077, O00775, O00776, O00777, O00778, Q540R1 | Gene names | HIST1H2AC, H2AFL | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A type 1-C. | |||||
|
TRM61_HUMAN
|
||||||
| NC score | 0.064257 (rank : 50) | θ value | 0.0961366 (rank : 23) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96FX7, Q8N7Q9 | Gene names | TRM61, C14orf172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA (adenine-N(1)-)-methyltransferase catalytic subunit TRM61 (EC 2.1.1.36) (tRNA(m1A58)-methyltransferase subunit TRM61) (tRNA(m1A58)MTase subunit TRM61). | |||||
|
H2AFB_HUMAN
|
||||||
| NC score | 0.063822 (rank : 51) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P98176, Q5TZB2, Q6FG78, Q96PR7 | Gene names | H2AFB3, H2ABBD, H2AFB | |||
|
Domain Architecture |

|
|||||
| Description | Histone H2A-Bbd (H2A Barr body-deficient) (H2A.Bbd). | |||||
|
VIGLN_MOUSE
|
||||||
| NC score | 0.058721 (rank : 52) | θ value | 0.21417 (rank : 26) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8VDJ3 | Gene names | Hdlbp | |||
|
Domain Architecture |

|
|||||
| Description | Vigilin (High density lipoprotein-binding protein) (HDL-binding protein). | |||||
|
VIGLN_HUMAN
|
||||||
| NC score | 0.057964 (rank : 53) | θ value | 0.21417 (rank : 25) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q00341, Q9UCY3 | Gene names | HDLBP, HBP, VGL | |||
|
Domain Architecture |

|
|||||
| Description | Vigilin (High density lipoprotein-binding protein) (HDL-binding protein). | |||||
|
PARP1_HUMAN
|
||||||
| NC score | 0.053518 (rank : 54) | θ value | 0.813845 (rank : 35) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P09874, Q8IUZ9 | Gene names | PARP1, ADPRT, PPOL | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 1 (EC 2.4.2.30) (PARP-1) (ADPRT) (NAD(+) ADP-ribosyltransferase 1) (Poly[ADP-ribose] synthetase 1). | |||||
|
TRM61_MOUSE
|
||||||
| NC score | 0.049571 (rank : 55) | θ value | 1.38821 (rank : 50) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80XC2, Q8BMD4, Q8BX33 | Gene names | Trm61 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA (adenine-N(1)-)-methyltransferase catalytic subunit TRM61 (EC 2.1.1.36) (tRNA(m1A58)-methyltransferase subunit TRM61) (tRNA(m1A58)MTase subunit TRM61). | |||||
|
PARP1_MOUSE
|
||||||
| NC score | 0.045132 (rank : 56) | θ value | 4.03905 (rank : 77) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P11103, Q9JLX4, Q9QVQ3 | Gene names | Parp1, Adprp, Adprt, Adprt1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 1 (EC 2.4.2.30) (PARP-1) (ADPRT) (NAD(+) ADP-ribosyltransferase 1) (Poly[ADP-ribose] synthetase 1) (msPARP). | |||||
|
AKAP9_HUMAN
|
||||||
| NC score | 0.036759 (rank : 57) | θ value | 0.163984 (rank : 24) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1710 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q99996, O14869, O43355, O94895, Q9UQH3, Q9UQQ4, Q9Y6B8, Q9Y6Y2 | Gene names | AKAP9, AKAP350, AKAP450, KIAA0803 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 9 (Protein kinase A-anchoring protein 9) (PRKA9) (A-kinase anchor protein 450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP 350) (hgAKAP 350) (AKAP 120-like protein) (Protein hyperion) (Protein yotiao) (Centrosome- and Golgi-localized PKN-associated protein) (CG-NAP). | |||||
|
HUS1_MOUSE
|
||||||
| NC score | 0.035843 (rank : 58) | θ value | 0.279714 (rank : 27) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BQY8, O70543, Q6P8H5 | Gene names | Hus1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Checkpoint protein HUS1 (mHUS1). | |||||
|
CJ084_MOUSE
|
||||||
| NC score | 0.035212 (rank : 59) | θ value | 1.06291 (rank : 37) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C6C7 | Gene names | D19Ertd737e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf84 homolog. | |||||
|
TOLIP_MOUSE
|
||||||
| NC score | 0.032461 (rank : 60) | θ value | 1.81305 (rank : 57) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QZ06, Q543N1, Q9D5P5 | Gene names | Tollip | |||
|
Domain Architecture |

|
|||||
| Description | Toll-interacting protein. | |||||
|
ANR26_MOUSE
|
||||||
| NC score | 0.030456 (rank : 61) | θ value | 0.47712 (rank : 29) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1554 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q811D2, Q69ZS2, Q9CS61 | Gene names | Ankrd26, Kiaa1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
MYG_MOUSE
|
||||||
| NC score | 0.030115 (rank : 62) | θ value | 0.813845 (rank : 33) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P04247 | Gene names | Mb | |||
|
Domain Architecture |

|
|||||
| Description | Myoglobin. | |||||
|
RABE1_HUMAN
|
||||||
| NC score | 0.029329 (rank : 63) | θ value | 1.06291 (rank : 41) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1104 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q15276, O95369, Q8IVX3 | Gene names | RABEP1, RABPT5, RABPT5A | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 1 (Rabaptin-5) (Rabaptin-5alpha) (Rabaptin-4) (NY-REN-17 antigen). | |||||
|
RABE1_MOUSE
|
||||||
| NC score | 0.028909 (rank : 64) | θ value | 1.06291 (rank : 42) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O35551, Q99L08, Q9CRP3, Q9EQF9, Q9JI94 | Gene names | Rabep1, Rab5ep, Rabpt5, Rabpt5a | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 1 (Rabaptin-5) (Rabaptin-5alpha). | |||||
|
TAF12_MOUSE
|
||||||
| NC score | 0.028457 (rank : 65) | θ value | 2.36792 (rank : 63) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VE65, Q3V0S7, Q9CZ11, Q9D8Q0 | Gene names | Taf12 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 12 (Transcription initiation factor TFIID 20 kDa subunits) (TAFII-20) (TAFII20). | |||||
|
SYNE2_HUMAN
|
||||||
| NC score | 0.026604 (rank : 66) | θ value | 0.813845 (rank : 36) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1323 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8WXH0, Q8N1S3, Q8NF49, Q8TER7, Q8WWW3, Q8WWW4, Q8WWW5, Q8WXH1, Q9NU50, Q9UFQ4, Q9Y2L4, Q9Y4R1 | Gene names | SYNE2, KIAA1011, NUA | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-2 (Nuclear envelope spectrin repeat protein 2) (Syne-2) (Synaptic nuclear envelope protein 2) (Nucleus and actin connecting element protein) (Protein NUANCE). | |||||
|
CENPF_HUMAN
|
||||||
| NC score | 0.026314 (rank : 67) | θ value | 6.88961 (rank : 92) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
ATRIP_HUMAN
|
||||||
| NC score | 0.026092 (rank : 68) | θ value | 1.81305 (rank : 51) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8WXE1, Q8NHQ2, Q8WUG7, Q96CL3, Q9HA30 | Gene names | ATRIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATR-interacting protein (ATM and Rad3-related-interacting protein). | |||||
|
SNX25_HUMAN
|
||||||
| NC score | 0.025803 (rank : 69) | θ value | 1.81305 (rank : 56) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H3E2 | Gene names | SNX25 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-25. | |||||
|
FYCO1_MOUSE
|
||||||
| NC score | 0.025249 (rank : 70) | θ value | 1.81305 (rank : 53) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8VDC1, Q3V385, Q7TMT0, Q8BJN2 | Gene names | Fyco1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1. | |||||
|
T106C_MOUSE
|
||||||
| NC score | 0.024317 (rank : 71) | θ value | 3.0926 (rank : 68) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80VP8 | Gene names | Tmem106c, D15Ertd405e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 106C. | |||||
|
TOLIP_HUMAN
|
||||||
| NC score | 0.024278 (rank : 72) | θ value | 6.88961 (rank : 104) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H0E2, Q9H9E6, Q9UJ69 | Gene names | TOLLIP | |||
|
Domain Architecture |

|
|||||
| Description | Toll-interacting protein. | |||||
|
SYCP1_HUMAN
|
||||||
| NC score | 0.023931 (rank : 73) | θ value | 1.38821 (rank : 49) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1313 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q15431, O14963 | Gene names | SYCP1, SCP1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
ANR26_HUMAN
|
||||||
| NC score | 0.023594 (rank : 74) | θ value | 6.88961 (rank : 90) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1614 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UPS8, Q2TAZ3, Q6ZR14, Q9H1Q1, Q9NSK9, Q9NTD5, Q9NW69 | Gene names | ANKRD26, KIAA1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
MBD4_HUMAN
|
||||||
| NC score | 0.023392 (rank : 75) | θ value | 1.81305 (rank : 54) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95243, Q7Z4T3, Q96F09 | Gene names | MBD4, MED1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 4 (EC 3.2.2.-) (Methyl-CpG-binding protein MBD4) (Methyl-CpG-binding endonuclease 1) (Mismatch-specific DNA N-glycosylase). | |||||
|
CENPE_HUMAN
|
||||||
| NC score | 0.023063 (rank : 76) | θ value | 1.81305 (rank : 52) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
MMRN1_HUMAN
|
||||||
| NC score | 0.022993 (rank : 77) | θ value | 6.88961 (rank : 100) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q13201, Q6P3T8 | Gene names | MMRN1, ECM, EMILIN4, MMRN | |||
|
Domain Architecture |

|
|||||
| Description | Multimerin-1 precursor (Endothelial cell multimerin 1) (EMILIN-4) (Elastin microfibril interface located protein 4) (Elastin microfibril interfacer 4). | |||||
|
TMF1_HUMAN
|
||||||
| NC score | 0.022910 (rank : 78) | θ value | 5.27518 (rank : 89) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 987 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P82094 | Gene names | TMF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA element modulatory factor (TMF). | |||||
|
SYNE1_HUMAN
|
||||||
| NC score | 0.022771 (rank : 79) | θ value | 5.27518 (rank : 88) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
MDN1_HUMAN
|
||||||
| NC score | 0.022601 (rank : 80) | θ value | 2.36792 (rank : 60) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9NU22, O15019 | Gene names | MDN1, KIAA0301 | |||
|
Domain Architecture |

|
|||||
| Description | Midasin (MIDAS-containing protein). | |||||
|
CP250_MOUSE
|
||||||
| NC score | 0.022296 (rank : 81) | θ value | 5.27518 (rank : 82) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1583 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q60952, Q2I8G3, Q3UTR4, Q6PFF6, Q8BLC6 | Gene names | Cep250, Cep2, Inmp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1) (Intranuclear matrix protein). | |||||
|
LETM1_MOUSE
|
||||||
| NC score | 0.022028 (rank : 82) | θ value | 5.27518 (rank : 86) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Z2I0, Q5PQC5, Q7TMK8, Q80ZX6, Q8CGJ3, Q8K5E5 | Gene names | Letm1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper-EF-hand-containing transmembrane protein 1, mitochondrial precursor. | |||||
|
CCDC6_HUMAN
|
||||||
| NC score | 0.022012 (rank : 83) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 723 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q16204, Q15250 | Gene names | CCDC6, D10S170 | |||
|
Domain Architecture |

|
|||||
| Description | Coiled-coil domain-containing protein 6 (H4 protein) (Papillary thyroid carcinoma-encoded protein). | |||||
|
IF3A_MOUSE
|
||||||
| NC score | 0.021090 (rank : 84) | θ value | 5.27518 (rank : 84) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 871 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P23116, Q60697, Q62162 | Gene names | Eif3s10, Csma, Eif3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3 p185) (eIF3a) (p162 protein) (Centrosomin). | |||||
|
MACF4_HUMAN
|
||||||
| NC score | 0.020814 (rank : 85) | θ value | 6.88961 (rank : 99) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1321 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q96PK2, Q8WXY1 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoform 4. | |||||
|
CF152_MOUSE
|
||||||
| NC score | 0.020782 (rank : 86) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q80ST9, Q9CYM9, Q9D5J9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf152 homolog. | |||||
|
TXLNB_HUMAN
|
||||||
| NC score | 0.020317 (rank : 87) | θ value | 2.36792 (rank : 64) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 912 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8N3L3, Q76L25, Q86T52, Q8N3S2 | Gene names | TXLNB, C6orf198, MDP77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77) (hMDP77). | |||||
|
MYH9_MOUSE
|
||||||
| NC score | 0.020247 (rank : 88) | θ value | 1.06291 (rank : 40) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
SH3G3_HUMAN
|
||||||
| NC score | 0.020245 (rank : 89) | θ value | 0.47712 (rank : 30) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99963, O43553, O43554 | Gene names | SH3GL3, CNSA3, SH3D2C | |||
|
Domain Architecture |

|
|||||
| Description | SH3-containing GRB2-like protein 3 (Endophilin-3) (Endophilin-A3) (SH3 domain protein 2C) (EEN-B2). | |||||
|
GCC2_MOUSE
|
||||||
| NC score | 0.020245 (rank : 90) | θ value | 4.03905 (rank : 73) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1619 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8CHG3, Q8BR44, Q8R2Q5, Q9CT45 | Gene names | Gcc2, Kiaa0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185). | |||||
|
RT35_HUMAN
|
||||||
| NC score | 0.020165 (rank : 91) | θ value | 4.03905 (rank : 78) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P82673, Q32LZ1, Q6P4C6, Q7L1M6, Q96AI0, Q9H044, Q9HC14, Q9P1R5 | Gene names | MRPS35, MRPS28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 28S ribosomal protein S35, mitochondrial precursor (S35mt) (MRP-S35) (Mitochondrial ribosomal protein S28) (MRP-S28). | |||||
|
IF3A_HUMAN
|
||||||
| NC score | 0.020158 (rank : 92) | θ value | 8.99809 (rank : 105) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 875 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q14152, O00653 | Gene names | EIF3S10, KIAA0139 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3 p185) (eIF3a). | |||||
|
NOL10_HUMAN
|
||||||
| NC score | 0.020040 (rank : 93) | θ value | 4.03905 (rank : 76) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9BSC4, Q53RC9, Q96TA5, Q9H7Y7, Q9H855 | Gene names | NOL10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar protein 10. | |||||
|
CCD46_MOUSE
|
||||||
| NC score | 0.019598 (rank : 94) | θ value | 3.0926 (rank : 65) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q5PR68, Q80ZN6, Q99JS4, Q9D9T1 | Gene names | Ccdc46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled coil domain-containing protein 46. | |||||
|
EEA1_HUMAN
|
||||||
| NC score | 0.019321 (rank : 95) | θ value | 6.88961 (rank : 95) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1796 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q15075, Q14221 | Gene names | EEA1, ZFYVE2 | |||
|
Domain Architecture |

|
|||||
| Description | Early endosome antigen 1 (Endosome-associated protein p162) (Zinc finger FYVE domain-containing protein 2). | |||||
|
KIF4A_HUMAN
|
||||||
| NC score | 0.018987 (rank : 96) | θ value | 2.36792 (rank : 59) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O95239, Q86TN3, Q86XX7, Q9NNY6, Q9NY24, Q9UMW3 | Gene names | KIF4A, KIF4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
THNSL_HUMAN
|
||||||
| NC score | 0.018469 (rank : 97) | θ value | 3.0926 (rank : 69) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8IYQ7, Q5VV21 | Gene names | THNSL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Threonine synthase-like 1. | |||||
|
MYH9_HUMAN
|
||||||
| NC score | 0.018393 (rank : 98) | θ value | 1.38821 (rank : 46) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
REST_HUMAN
|
||||||
| NC score | 0.018237 (rank : 99) | θ value | 6.88961 (rank : 102) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1605 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P30622 | Gene names | RSN, CYLN1 | |||
|
Domain Architecture |

|
|||||
| Description | Restin (Cytoplasmic linker protein 170 alpha-2) (CLIP-170) (Reed- Sternberg intermediate filament-associated protein) (Cytoplasmic linker protein 1). | |||||
|
KINH_MOUSE
|
||||||
| NC score | 0.018149 (rank : 100) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1082 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q61768, O08711, Q61580 | Gene names | Kif5b, Khcs, Kns1 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin heavy chain (Ubiquitous kinesin heavy chain) (UKHC). | |||||
|
DICER_HUMAN
|
||||||
| NC score | 0.018097 (rank : 101) | θ value | 1.38821 (rank : 45) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UPY3, O95943, Q9UQ02 | Gene names | DICER1, DICER, HERNA, KIAA0928 | |||
|
Domain Architecture |

|
|||||
| Description | Endoribonuclease Dicer (EC 3.1.26.-) (Helicase with RNase motif) (Helicase-MOI). | |||||
|
MACF1_HUMAN
|
||||||
| NC score | 0.017580 (rank : 102) | θ value | 6.88961 (rank : 98) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1341 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9UPN3, O75053, Q5VW20, Q8WXY2, Q9H540, Q9UKP0, Q9ULG9 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoforms 1/2/3/5 (Actin cross-linking family protein 7) (Macrophin-1) (Trabeculin-alpha) (620 kDa actin-binding protein) (ABP620). | |||||
|
KIF6_HUMAN
|
||||||
| NC score | 0.017525 (rank : 103) | θ value | 1.06291 (rank : 38) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6ZMV9, Q2MDE3, Q2MDE4, Q5T8J6, Q6ZWE3, Q86T87, Q8WTV4 | Gene names | KIF6, C6orf102 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin-like protein KIF6. | |||||
|
EP15R_MOUSE
|
||||||
| NC score | 0.016788 (rank : 104) | θ value | 5.27518 (rank : 83) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q60902, Q8CB60, Q8CB70, Q91WH8 | Gene names | Eps15l1, Eps15-rs, Eps15R | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal growth factor receptor substrate 15-like 1 (Eps15-related protein) (Eps15R) (Epidermal growth factor receptor pathway substrate 15-related sequence) (Eps15-rs). | |||||
|
SH3G3_MOUSE
|
||||||
| NC score | 0.016541 (rank : 105) | θ value | 2.36792 (rank : 62) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q62421 | Gene names | Sh3gl3, Sh3d2c, Sh3d2c2 | |||
|
Domain Architecture |

|
|||||
| Description | SH3-containing GRB2-like protein 3 (Endophilin-3) (Endophilin-A3) (SH3 domain protein 2C) (SH3p13). | |||||
|
TM9S3_HUMAN
|
||||||
| NC score | 0.016402 (rank : 106) | θ value | 3.0926 (rank : 70) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9HD45, Q6UWE7, Q9NWL8, Q9P0G9, Q9UHW8 | Gene names | TM9SF3, SMBP | |||
|
Domain Architecture |

|
|||||
| Description | Transmembrane 9 superfamily protein member 3 precursor (SM-11044- binding protein) (EP70-P-iso). | |||||
|
C8AP2_HUMAN
|
||||||
| NC score | 0.015441 (rank : 107) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UKL3, Q6PH76, Q7LCQ7, Q86YD9, Q9NUQ4, Q9NZV9, Q9P2N1, Q9Y563 | Gene names | CASP8AP2, FLASH, KIAA1315, RIP25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
PTBP2_MOUSE
|
||||||
| NC score | 0.015056 (rank : 108) | θ value | 3.0926 (rank : 67) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91Z31, Q7TNW7, Q9CUW2, Q9QYC2, Q9R0V9 | Gene names | Ptbp2, Brptb, Nptb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypyrimidine tract-binding protein 2 (Brain-enriched polypyrimidine tract-binding protein) (Brain-enriched PTB) (RRM-type RNA-binding protein brPTB) (Neural polypyrimidine tract-binding protein). | |||||
|
PTBP2_HUMAN
|
||||||
| NC score | 0.015035 (rank : 109) | θ value | 3.0926 (rank : 66) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UKA9, Q8N0Z1, Q8N160, Q8NFB0, Q8NFB1, Q969N9, Q96Q76 | Gene names | PTBP2, NPTB, PTB, PTBLP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypyrimidine tract-binding protein 2 (Neurally-enriched homolog of PTB) (Neural polypyrimidine tract-binding protein) (PTB-like protein). | |||||
|
EMIL2_MOUSE
|
||||||
| NC score | 0.014050 (rank : 110) | θ value | 6.88961 (rank : 96) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8K482 | Gene names | Emilin2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Basilin). | |||||
|
TM9S3_MOUSE
|
||||||
| NC score | 0.012694 (rank : 111) | θ value | 8.99809 (rank : 113) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ET30 | Gene names | Tm9sf3, Smbp | |||
|
Domain Architecture |

|
|||||
| Description | Transmembrane 9 superfamily protein member 3 precursor. | |||||
|
NSBP1_MOUSE
|
||||||
| NC score | 0.012497 (rank : 112) | θ value | 2.36792 (rank : 61) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
ICAL_HUMAN
|
||||||
| NC score | 0.011998 (rank : 113) | θ value | 6.88961 (rank : 97) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P20810, O95360, Q96D08, Q9H1Z5 | Gene names | CAST | |||
|
Domain Architecture |

|
|||||
| Description | Calpastatin (Calpain inhibitor) (Sperm BS-17 component). | |||||
|
ARHGC_HUMAN
|
||||||
| NC score | 0.011812 (rank : 114) | θ value | 1.38821 (rank : 43) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NZN5, O15086 | Gene names | ARHGEF12, KIAA0382, LARG | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 12 (Leukemia-associated RhoGEF). | |||||
|
KI13A_HUMAN
|
||||||
| NC score | 0.011366 (rank : 115) | θ value | 8.99809 (rank : 107) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9H1H9, Q9H1H8 | Gene names | KIF13A, RBKIN | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13A (Kinesin-like protein RBKIN). | |||||
|
DUS3L_HUMAN
|
||||||
| NC score | 0.010687 (rank : 116) | θ value | 6.88961 (rank : 94) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96G46, Q96HM5, Q9BSU4, Q9H877, Q9NPR1 | Gene names | DUS3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA-dihydrouridine synthase 3-like (EC 1.-.-.-). | |||||
|
CTRO_HUMAN
|
||||||
| NC score | 0.010484 (rank : 117) | θ value | 1.38821 (rank : 44) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 2092 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O14578, Q6XUH8, Q86UQ9, Q9UPZ7 | Gene names | CIT, KIAA0949, STK21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
SCFD2_HUMAN
|
||||||
| NC score | 0.009985 (rank : 118) | θ value | 8.99809 (rank : 109) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8WU76, Q8N5F3, Q8N8H0, Q96ED3 | Gene names | SCFD2, STXBP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Sec1 family domain-containing protein 2 (Syntaxin-binding protein 1- like 1). | |||||
|
BIRC6_HUMAN
|
||||||
| NC score | 0.009421 (rank : 119) | θ value | 6.88961 (rank : 91) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NR09, Q9ULD1 | Gene names | BIRC6, KIAA1289 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 6 (Ubiquitin-conjugating BIR-domain enzyme apollon). | |||||
|
MRCKA_HUMAN
|
||||||
| NC score | 0.008957 (rank : 120) | θ value | 1.81305 (rank : 55) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 1999 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q5VT25, Q59GZ1, Q5H9N9, Q5T797, Q5VT26, Q5VT27, Q86XX2, Q86XX3, Q99646 | Gene names | CDC42BPA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK alpha (EC 2.7.11.1) (CDC42- binding protein kinase alpha) (Myotonic dystrophy kinase-related CDC42-binding kinase alpha) (Myotonic dystrophy protein kinase-like alpha) (MRCK alpha) (DMPK-like alpha). | |||||
|
COHA1_MOUSE
|
||||||
| NC score | 0.008900 (rank : 121) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q07563, Q99LK8 | Gene names | Col17a1, Bp180, Bpag2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVII) chain (Bullous pemphigoid antigen 2) (180 kDa bullous pemphigoid antigen 2). | |||||
|
GNAT1_HUMAN
|
||||||
| NC score | 0.008343 (rank : 122) | θ value | 2.36792 (rank : 58) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P11488 | Gene names | GNAT1, GNATR | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-binding protein G(t), alpha-1 subunit (Transducin alpha-1 chain). | |||||
|
RHG18_HUMAN
|
||||||
| NC score | 0.008333 (rank : 123) | θ value | 6.88961 (rank : 103) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8N392, Q58EZ3, Q6P679, Q6PJD7, Q96S64 | Gene names | ARHGAP18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho-GTPase-activating protein 18 (MacGAP). | |||||
|
STA5B_MOUSE
|
||||||
| NC score | 0.008090 (rank : 124) | θ value | 8.99809 (rank : 112) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P42232, Q541Q5, Q60804, Q8K3Q1, Q9JKM1 | Gene names | Stat5b | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 5B. | |||||
|
MDR1_HUMAN
|
||||||
| NC score | 0.007989 (rank : 125) | θ value | 1.06291 (rank : 39) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P08183, Q12755, Q14812 | Gene names | ABCB1, MDR1, PGY1 | |||
|
Domain Architecture |

|
|||||
| Description | Multidrug resistance protein 1 (EC 3.6.3.44) (ATP-binding cassette sub-family B member 1) (P-glycoprotein 1) (CD243 antigen). | |||||
|
STA5A_MOUSE
|
||||||
| NC score | 0.007939 (rank : 126) | θ value | 8.99809 (rank : 111) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P42230 | Gene names | Stat5a, Mgf, Mpf | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 5A (Mammary gland factor). | |||||
|
RNF34_MOUSE
|
||||||
| NC score | 0.007847 (rank : 127) | θ value | 8.99809 (rank : 108) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99KR6, Q3UV45 | Gene names | Rnf34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 34 (RING finger protein RIFF) (Phafin-1). | |||||
|
CP1A2_HUMAN
|
||||||
| NC score | 0.007824 (rank : 128) | θ value | 0.813845 (rank : 32) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P05177, Q16754, Q6NWU5, Q9BXX7, Q9UK49 | Gene names | CYP1A2 | |||
|
Domain Architecture |

|
|||||
| Description | Cytochrome P450 1A2 (EC 1.14.14.1) (CYPIA2) (P450-P3) (P(3)450) (P450 4). | |||||
|
GNAT1_MOUSE
|
||||||
| NC score | 0.007426 (rank : 129) | θ value | 4.03905 (rank : 74) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P20612, Q80X34 | Gene names | Gnat1, Gnat-1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-binding protein G(t), alpha-1 subunit (Transducin alpha-1 chain). | |||||
|
STA13_HUMAN
|
||||||
| NC score | 0.007414 (rank : 130) | θ value | 5.27518 (rank : 87) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y3M8, Q5HYH1, Q5TAE3, Q6UN61, Q86TP6, Q86WQ3, Q86XT1 | Gene names | STARD13, DLC2, GT650 | |||
|
Domain Architecture |

|
|||||
| Description | StAR-related lipid transfer protein 13 (StARD13) (START domain- containing protein 13) (46H23.2) (Deleted in liver cancer protein 2) (Rho GTPase-activating protein). | |||||
|
CTNA3_MOUSE
|
||||||
| NC score | 0.006909 (rank : 131) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q65CL1, Q3UQ61, Q8C0N3 | Gene names | Ctnna3, Catna3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Catenin alpha-3 (Alpha T-catenin) (Cadherin-associated protein). | |||||
|
IPO8_HUMAN
|
||||||
| NC score | 0.006400 (rank : 132) | θ value | 8.99809 (rank : 106) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15397 | Gene names | IPO8, RANBP8 | |||
|
Domain Architecture |

|
|||||
| Description | Importin-8 (Imp8) (Ran-binding protein 8) (RanBP8). | |||||
|
NOS1_MOUSE
|
||||||
| NC score | 0.005222 (rank : 133) | θ value | 6.88961 (rank : 101) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z0J4, Q64208 | Gene names | Nos1 | |||
|
Domain Architecture |

|
|||||
| Description | Nitric-oxide synthase, brain (EC 1.14.13.39) (NOS type I) (Neuronal NOS) (N-NOS) (nNOS) (Constitutive NOS) (NC-NOS) (bNOS). | |||||
|
TITIN_HUMAN
|
||||||
| NC score | 0.004888 (rank : 134) | θ value | 0.279714 (rank : 28) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
SMCA5_HUMAN
|
||||||
| NC score | 0.002693 (rank : 135) | θ value | 8.99809 (rank : 110) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60264 | Gene names | SMARCA5, SNF2H, WCRF135 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 5 (EC 3.6.1.-) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin A5) (Sucrose nonfermenting protein 2 homolog) (hSNF2H). | |||||
|
NUAK2_HUMAN
|
||||||
| NC score | 0.001438 (rank : 136) | θ value | 1.38821 (rank : 47) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9H093 | Gene names | NUAK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NUAK family SNF1-like kinase 2 (EC 2.7.11.1) (SNF1/AMP kinase-related kinase) (SNARK). | |||||
|
RON_MOUSE
|
||||||
| NC score | 0.001337 (rank : 137) | θ value | 1.38821 (rank : 48) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q62190, Q62555 | Gene names | Mst1r, Ron, Stk | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage-stimulating protein receptor precursor (EC 2.7.10.1) (MSP receptor) (p185-Ron) (Stem cell-derived tyrosine kinase) (CDw136 antigen) [Contains: Macrophage-stimulating protein receptor alpha chain; Macrophage-stimulating protein receptor beta chain]. | |||||
|
KS6A1_MOUSE
|
||||||
| NC score | 0.000491 (rank : 138) | θ value | 5.27518 (rank : 85) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 850 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P18653 | Gene names | Rps6ka1, Rsk1 | |||
|
Domain Architecture |
|
|||||
| Description | Ribosomal protein S6 kinase alpha-1 (EC 2.7.11.1) (S6K-alpha 1) (90 kDa ribosomal protein S6 kinase 1) (p90-RSK 1) (Ribosomal S6 kinase 1) (RSK-1) (pp90RSK1) (p90S6K) (MAP kinase-activated protein kinase 1a) (MAPKAPK1A). | |||||
|
ZN630_HUMAN
|
||||||
| NC score | 0.000140 (rank : 139) | θ value | 3.0926 (rank : 71) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 737 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q2M218 | Gene names | ZNF630 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 630. | |||||
|
ZN287_HUMAN
|
||||||
| NC score | -0.000894 (rank : 140) | θ value | 0.62314 (rank : 31) | |||
| Query Neighborhood Hits | 113 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9HBT7 | Gene names | ZNF287 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 287. | |||||