Please be patient as the page loads
|
MCM4_MOUSE
|
||||||
| SwissProt Accessions | P49717, O89056 | Gene names | Mcm4, Cdc21, Mcmd4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM4 (CDC21 homolog) (P1-CDC21). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
MCM4_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.995271 (rank : 2) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P33991, Q8NEH1, Q99658 | Gene names | MCM4, CDC21 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM4 (CDC21 homolog) (P1-CDC21). | |||||
|
MCM4_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P49717, O89056 | Gene names | Mcm4, Cdc21, Mcmd4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM4 (CDC21 homolog) (P1-CDC21). | |||||
|
MCM7_MOUSE
|
||||||
| θ value | 6.93392e-93 (rank : 3) | NC score | 0.952178 (rank : 3) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q61881 | Gene names | Mcm7, Cdc47, Mcmd7 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM7 (CDC47 homolog). | |||||
|
MCM2_HUMAN
|
||||||
| θ value | 4.49445e-92 (rank : 4) | NC score | 0.948108 (rank : 5) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P49736, Q14577, Q15023, Q8N2V1, Q969W7, Q96AE1, Q9BRM7 | Gene names | MCM2, BM28, CDCL1, KIAA0030 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM2 (Minichromosome maintenance protein 2 homolog) (Nuclear protein BM28). | |||||
|
MCM7_HUMAN
|
||||||
| θ value | 2.91321e-91 (rank : 5) | NC score | 0.951683 (rank : 4) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P33993, Q15076, Q96D34, Q96GL1 | Gene names | MCM7, CDC47, MCM2 | |||
|
Domain Architecture |
|
|||||
| Description | DNA replication licensing factor MCM7 (CDC47 homolog) (P1.1-MCM3). | |||||
|
MCM2_MOUSE
|
||||||
| θ value | 8.47616e-91 (rank : 6) | NC score | 0.947515 (rank : 6) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P97310, O08971, O89057, Q8C2R0 | Gene names | Mcm2, Bm28, Cdcl1, Kiaa0030, Mcmd2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM2 (Minichromosome maintenance protein 2 homolog) (Nuclear protein BM28). | |||||
|
MCM5_MOUSE
|
||||||
| θ value | 3.33315e-87 (rank : 7) | NC score | 0.946010 (rank : 8) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P49718 | Gene names | Mcm5, Cdc46, Mcmd5 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM5 (CDC46 homolog) (P1-CDC46). | |||||
|
MCM5_HUMAN
|
||||||
| θ value | 2.82168e-86 (rank : 8) | NC score | 0.946426 (rank : 7) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P33992, O60785, Q14578, Q9BTJ4, Q9BWL8 | Gene names | MCM5, CDC46 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM5 (CDC46 homolog) (P1-CDC46). | |||||
|
MCM6_HUMAN
|
||||||
| θ value | 5.50685e-82 (rank : 9) | NC score | 0.941052 (rank : 9) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q14566, Q13504, Q99859 | Gene names | MCM6 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM6 (p105MCM). | |||||
|
MCM3_MOUSE
|
||||||
| θ value | 7.19216e-82 (rank : 10) | NC score | 0.932779 (rank : 14) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P25206, Q61492 | Gene names | Mcm3, Mcmd, Mcmd3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM3 (DNA polymerase alpha holoenzyme-associated protein P1) (P1-MCM3). | |||||
|
MCM3_HUMAN
|
||||||
| θ value | 1.2268e-81 (rank : 11) | NC score | 0.935933 (rank : 11) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P25205, Q92660, Q9BTR3, Q9NUE7 | Gene names | MCM3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM3 (DNA polymerase alpha holoenzyme-associated protein P1) (RLF subunit beta) (P102 protein) (P1-MCM3). | |||||
|
MCM6_MOUSE
|
||||||
| θ value | 7.95186e-81 (rank : 12) | NC score | 0.940814 (rank : 10) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P97311, Q80YQ7 | Gene names | Mcm6, Mcmd6, Mis5 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM6 (Mis5 homolog). | |||||
|
MCM8_MOUSE
|
||||||
| θ value | 1.26953e-78 (rank : 13) | NC score | 0.935090 (rank : 12) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9CWV1, Q80US2, Q80VI0 | Gene names | Mcm8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA replication licensing factor MCM8 (Minichromosome maintenance 8). | |||||
|
MCM8_HUMAN
|
||||||
| θ value | 1.8332e-77 (rank : 14) | NC score | 0.935045 (rank : 13) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UJA3, Q495R4, Q495R7, Q86US4, Q969I5 | Gene names | MCM8, C20orf154 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM8 (Minichromosome maintenance 8). | |||||
|
AATK_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 15) | NC score | 0.008082 (rank : 36) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1243 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6ZMQ8, O75136, Q6ZN31, Q86X28 | Gene names | AATK, KIAA0641 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Apoptosis-associated tyrosine-protein kinase (EC 2.7.10.2) (AATYK) (Brain apoptosis-associated tyrosine kinase) (CDK5-binding protein) (p35-binding protein) (p35BP). | |||||
|
GON4L_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 16) | NC score | 0.029128 (rank : 19) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 555 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q3T8J9, Q3T8J8, Q5VYZ5, Q5W0D5, Q6AWA6, Q6P1Q6, Q7Z3L3, Q8IY79, Q9BQI1, Q9HCG6 | Gene names | GON4L, GON4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
DYH11_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 17) | NC score | 0.038273 (rank : 17) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96DT5, Q9UJ82 | Gene names | DNAH11 | |||
|
Domain Architecture |

|
|||||
| Description | Ciliary dynein heavy chain 11 (Axonemal beta dynein heavy chain 11). | |||||
|
CJ068_HUMAN
|
||||||
| θ value | 0.125558 (rank : 18) | NC score | 0.048071 (rank : 15) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 316 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H943, Q8N7T7 | Gene names | C10orf68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf68. | |||||
|
GIMA7_HUMAN
|
||||||
| θ value | 0.279714 (rank : 19) | NC score | 0.022772 (rank : 23) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NHV1 | Gene names | GIMAP7, IAN7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GTPase, IMAP family member 7 (Immunity-associated nucleotide 7 protein). | |||||
|
ABCD1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 20) | NC score | 0.026321 (rank : 20) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P48410, Q9QY41, Q9QZ32 | Gene names | Abcd1, Ald, Aldgh | |||
|
Domain Architecture |
|
|||||
| Description | ATP-binding cassette sub-family D member 1 (Adrenoleukodystrophy protein) (ALDP). | |||||
|
ABCD1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 21) | NC score | 0.025073 (rank : 21) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P33897, Q6GTZ2 | Gene names | ABCD1, ALD | |||
|
Domain Architecture |

|
|||||
| Description | ATP-binding cassette sub-family D member 1 (Adrenoleukodystrophy protein) (ALDP). | |||||
|
MYH11_HUMAN
|
||||||
| θ value | 0.47712 (rank : 22) | NC score | 0.005076 (rank : 43) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
FBX11_HUMAN
|
||||||
| θ value | 1.06291 (rank : 23) | NC score | 0.023725 (rank : 22) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q86XK2, Q8IXG3, Q96E90, Q9H6V8, Q9H9L1, Q9NR14, Q9UFK1, Q9UHI1, Q9UKC2 | Gene names | FBXO11, FBX11, VIT1 | |||
|
Domain Architecture |

|
|||||
| Description | F-box only protein 11 (Vitiligo-associated protein VIT-1). | |||||
|
NENF_HUMAN
|
||||||
| θ value | 1.06291 (rank : 24) | NC score | 0.041908 (rank : 16) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UMX5, Q53FZ6, Q5TM90 | Gene names | NENF, SPUF | |||
|
Domain Architecture |
|
|||||
| Description | Neudesin precursor (Neuron-derived neurotrophic factor) (Secreted protein of unknown function) (SPUF protein). | |||||
|
PAPOG_MOUSE
|
||||||
| θ value | 1.38821 (rank : 25) | NC score | 0.012403 (rank : 31) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PCL9, Q8BZC9 | Gene names | Papolg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(A) polymerase gamma (EC 2.7.7.19) (PAP gamma) (Polynucleotide adenylyltransferase gamma) (SRP RNA 3' adenylating enzyme). | |||||
|
CAP2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 26) | NC score | 0.018507 (rank : 25) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CYT6, Q80YU7, Q9D6L0 | Gene names | Cap2 | |||
|
Domain Architecture |
|
|||||
| Description | Adenylyl cyclase-associated protein 2 (CAP 2). | |||||
|
PK1IP_HUMAN
|
||||||
| θ value | 1.81305 (rank : 27) | NC score | 0.008241 (rank : 35) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NWT1, Q5T4J2, Q96QJ8, Q96T87 | Gene names | PAK1IP1, PIP1, WDR84 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | p21-activated protein kinase-interacting protein 1 (PAK1-interacting protein 1) (PAK/PLC-interacting protein 1) (hPIP1) (WD repeat protein 84). | |||||
|
RHG18_HUMAN
|
||||||
| θ value | 1.81305 (rank : 28) | NC score | 0.018984 (rank : 24) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N392, Q58EZ3, Q6P679, Q6PJD7, Q96S64 | Gene names | ARHGAP18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho-GTPase-activating protein 18 (MacGAP). | |||||
|
DYH9_HUMAN
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.031686 (rank : 18) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NYC9, O95494, Q9NQ28 | Gene names | DNAH9, DNAH17L, DNAL1 | |||
|
Domain Architecture |

|
|||||
| Description | Ciliary dynein heavy chain 9 (Axonemal beta dynein heavy chain 9). | |||||
|
K1802_MOUSE
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.005908 (rank : 40) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
MYH11_MOUSE
|
||||||
| θ value | 3.0926 (rank : 31) | NC score | 0.004304 (rank : 47) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
NU133_MOUSE
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.014316 (rank : 27) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R0G9 | Gene names | Nup133 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup133 (Nucleoporin Nup133) (133 kDa nucleoporin). | |||||
|
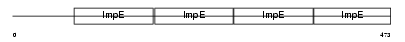
OGT1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.017375 (rank : 26) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15294, Q7Z3K0, Q8WWM8, Q96CC1, Q9UG57 | Gene names | OGT | |||
|
Domain Architecture |
|
|||||
| Description | UDP-N-acetylglucosamine--peptide N-acetylglucosaminyltransferase 110 kDa subunit (EC 2.4.1.-) (O-GlcNAc transferase p110 subunit). | |||||
|
GLIS3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 34) | NC score | -0.001417 (rank : 52) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 758 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NEA6 | Gene names | GLIS3, ZNF515 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLIS3 (GLI-similar 3) (Zinc finger protein 515). | |||||
|
MARK2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | -0.001610 (rank : 53) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1056 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7KZI7, Q15449, Q15524, Q5XGA3, Q68A18, Q96HB3, Q96RG0 | Gene names | MARK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL motif kinase) (EMK1) (PAR1 homolog). | |||||
|
MYH9_HUMAN
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.004110 (rank : 48) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
PHLB2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.006976 (rank : 37) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1016 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8K1N2, Q3TNI3, Q80Y16, Q8BKV3 | Gene names | Phldb2, Ll5b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 2 (Protein LL5-beta). | |||||
|
RHG18_MOUSE
|
||||||
| θ value | 4.03905 (rank : 38) | NC score | 0.012613 (rank : 30) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K0Q5, Q8BP03, Q8R196 | Gene names | Arhgap18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho-GTPase-activating protein 18. | |||||
|
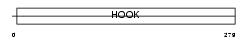
HOOK3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 39) | NC score | 0.008419 (rank : 34) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8BUK6, Q8BV48, Q8BW57, Q8BY41 | Gene names | Hook3 | |||
|
Domain Architecture |
|
|||||
| Description | Hook homolog 3. | |||||
|
SC24D_HUMAN
|
||||||
| θ value | 5.27518 (rank : 40) | NC score | 0.008534 (rank : 33) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O94855 | Gene names | SEC24D, KIAA0755 | |||
|
Domain Architecture |

|
|||||
| Description | Protein transport protein Sec24D (SEC24-related protein D). | |||||
|
SNX13_HUMAN
|
||||||
| θ value | 5.27518 (rank : 41) | NC score | 0.013379 (rank : 28) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y5W8, O94821, Q8WVZ2, Q8WXH8 | Gene names | SNX13, KIAA0713 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-13 (RGS domain- and PHOX domain-containing protein) (RGS-PX1). | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 42) | NC score | 0.006894 (rank : 38) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
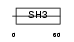
BCAR1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.004551 (rank : 46) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61140, Q60869 | Gene names | Bcar1, Cas, Crkas | |||
|
Domain Architecture |
|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
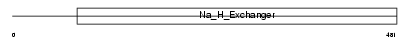
SL9A2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.005717 (rank : 41) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UBY0 | Gene names | SLC9A2, NHE2 | |||
|
Domain Architecture |
|
|||||
| Description | Sodium/hydrogen exchanger 2 (Na(+)/H(+) exchanger 2) (NHE-2) (Solute carrier family 9 member 2). | |||||
|
SMRC2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.005464 (rank : 42) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PDG5, Q6P6P3 | Gene names | Smarcc2, Baf170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
TGFB3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.006634 (rank : 39) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P17125 | Gene names | Tgfb3 | |||
|
Domain Architecture |
|
|||||
| Description | Transforming growth factor beta-3 precursor (TGF-beta-3). | |||||
|
CAP2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 47) | NC score | 0.013357 (rank : 29) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P40123, Q6IAY2 | Gene names | CAP2 | |||
|
Domain Architecture |
|
|||||
| Description | Adenylyl cyclase-associated protein 2 (CAP 2). | |||||
|
GPR64_HUMAN
|
||||||
| θ value | 8.99809 (rank : 48) | NC score | 0.002404 (rank : 51) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IZP9, O00406, Q8IWT2, Q8IZE4, Q8IZE5, Q8IZE6, Q8IZE7, Q8IZP3, Q8IZP4 | Gene names | GPR64, HE6, TM7LN2 | |||
|
Domain Architecture |

|
|||||
| Description | G-protein coupled receptor 64 precursor (Epididymis-specific protein 6) (He6 receptor). | |||||
|
GRIP2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.004844 (rank : 45) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9C0E4, Q8TEH9, Q9H7H3 | Gene names | GRIP2, KIAA1719 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor-interacting protein 2 (GRIP2 protein). | |||||
|
PEX1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 50) | NC score | 0.005050 (rank : 44) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43933, Q96S71, Q96S72, Q96S73, Q99994 | Gene names | PEX1 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisome biogenesis factor 1 (Peroxin-1) (Peroxisome biogenesis disorder protein 1). | |||||
|
PLCB4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.003417 (rank : 50) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q15147, Q5JYS8, Q5JYT0, Q5JYT4, Q9BQW5, Q9BQW6, Q9BQW8, Q9UJQ2 | Gene names | PLCB4 | |||
|
Domain Architecture |

|
|||||
| Description | 1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase beta 4 (EC 3.1.4.11) (Phosphoinositide phospholipase C) (Phospholipase C- beta-4) (PLC-beta-4). | |||||
|
RYR3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.008864 (rank : 32) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15413, O15175, Q15412 | Gene names | RYR3, HBRR | |||
|
Domain Architecture |

|
|||||
| Description | Ryanodine receptor 3 (Brain-type ryanodine receptor) (RyR3) (RYR-3) (Brain ryanodine receptor-calcium release channel). | |||||
|
TARA_MOUSE
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.003491 (rank : 49) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
MCM4_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P49717, O89056 | Gene names | Mcm4, Cdc21, Mcmd4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM4 (CDC21 homolog) (P1-CDC21). | |||||
|
MCM4_HUMAN
|
||||||
| NC score | 0.995271 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P33991, Q8NEH1, Q99658 | Gene names | MCM4, CDC21 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM4 (CDC21 homolog) (P1-CDC21). | |||||
|
MCM7_MOUSE
|
||||||
| NC score | 0.952178 (rank : 3) | θ value | 6.93392e-93 (rank : 3) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q61881 | Gene names | Mcm7, Cdc47, Mcmd7 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM7 (CDC47 homolog). | |||||
|
MCM7_HUMAN
|
||||||
| NC score | 0.951683 (rank : 4) | θ value | 2.91321e-91 (rank : 5) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P33993, Q15076, Q96D34, Q96GL1 | Gene names | MCM7, CDC47, MCM2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM7 (CDC47 homolog) (P1.1-MCM3). | |||||
|
MCM2_HUMAN
|
||||||
| NC score | 0.948108 (rank : 5) | θ value | 4.49445e-92 (rank : 4) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P49736, Q14577, Q15023, Q8N2V1, Q969W7, Q96AE1, Q9BRM7 | Gene names | MCM2, BM28, CDCL1, KIAA0030 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM2 (Minichromosome maintenance protein 2 homolog) (Nuclear protein BM28). | |||||
|
MCM2_MOUSE
|
||||||
| NC score | 0.947515 (rank : 6) | θ value | 8.47616e-91 (rank : 6) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P97310, O08971, O89057, Q8C2R0 | Gene names | Mcm2, Bm28, Cdcl1, Kiaa0030, Mcmd2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM2 (Minichromosome maintenance protein 2 homolog) (Nuclear protein BM28). | |||||
|
MCM5_HUMAN
|
||||||
| NC score | 0.946426 (rank : 7) | θ value | 2.82168e-86 (rank : 8) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P33992, O60785, Q14578, Q9BTJ4, Q9BWL8 | Gene names | MCM5, CDC46 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM5 (CDC46 homolog) (P1-CDC46). | |||||
|
MCM5_MOUSE
|
||||||
| NC score | 0.946010 (rank : 8) | θ value | 3.33315e-87 (rank : 7) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P49718 | Gene names | Mcm5, Cdc46, Mcmd5 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM5 (CDC46 homolog) (P1-CDC46). | |||||
|
MCM6_HUMAN
|
||||||
| NC score | 0.941052 (rank : 9) | θ value | 5.50685e-82 (rank : 9) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q14566, Q13504, Q99859 | Gene names | MCM6 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM6 (p105MCM). | |||||
|
MCM6_MOUSE
|
||||||
| NC score | 0.940814 (rank : 10) | θ value | 7.95186e-81 (rank : 12) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P97311, Q80YQ7 | Gene names | Mcm6, Mcmd6, Mis5 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM6 (Mis5 homolog). | |||||
|
MCM3_HUMAN
|
||||||
| NC score | 0.935933 (rank : 11) | θ value | 1.2268e-81 (rank : 11) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P25205, Q92660, Q9BTR3, Q9NUE7 | Gene names | MCM3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM3 (DNA polymerase alpha holoenzyme-associated protein P1) (RLF subunit beta) (P102 protein) (P1-MCM3). | |||||
|
MCM8_MOUSE
|
||||||
| NC score | 0.935090 (rank : 12) | θ value | 1.26953e-78 (rank : 13) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9CWV1, Q80US2, Q80VI0 | Gene names | Mcm8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA replication licensing factor MCM8 (Minichromosome maintenance 8). | |||||
|
MCM8_HUMAN
|
||||||
| NC score | 0.935045 (rank : 13) | θ value | 1.8332e-77 (rank : 14) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UJA3, Q495R4, Q495R7, Q86US4, Q969I5 | Gene names | MCM8, C20orf154 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM8 (Minichromosome maintenance 8). | |||||
|
MCM3_MOUSE
|
||||||
| NC score | 0.932779 (rank : 14) | θ value | 7.19216e-82 (rank : 10) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P25206, Q61492 | Gene names | Mcm3, Mcmd, Mcmd3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM3 (DNA polymerase alpha holoenzyme-associated protein P1) (P1-MCM3). | |||||
|
CJ068_HUMAN
|
||||||
| NC score | 0.048071 (rank : 15) | θ value | 0.125558 (rank : 18) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 316 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H943, Q8N7T7 | Gene names | C10orf68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf68. | |||||
|
NENF_HUMAN
|
||||||
| NC score | 0.041908 (rank : 16) | θ value | 1.06291 (rank : 24) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UMX5, Q53FZ6, Q5TM90 | Gene names | NENF, SPUF | |||
|
Domain Architecture |

|
|||||
| Description | Neudesin precursor (Neuron-derived neurotrophic factor) (Secreted protein of unknown function) (SPUF protein). | |||||
|
DYH11_HUMAN
|
||||||
| NC score | 0.038273 (rank : 17) | θ value | 0.0736092 (rank : 17) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96DT5, Q9UJ82 | Gene names | DNAH11 | |||
|
Domain Architecture |

|
|||||
| Description | Ciliary dynein heavy chain 11 (Axonemal beta dynein heavy chain 11). | |||||
|
DYH9_HUMAN
|
||||||
| NC score | 0.031686 (rank : 18) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NYC9, O95494, Q9NQ28 | Gene names | DNAH9, DNAH17L, DNAL1 | |||
|
Domain Architecture |

|
|||||
| Description | Ciliary dynein heavy chain 9 (Axonemal beta dynein heavy chain 9). | |||||
|
GON4L_HUMAN
|
||||||
| NC score | 0.029128 (rank : 19) | θ value | 0.0330416 (rank : 16) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 555 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q3T8J9, Q3T8J8, Q5VYZ5, Q5W0D5, Q6AWA6, Q6P1Q6, Q7Z3L3, Q8IY79, Q9BQI1, Q9HCG6 | Gene names | GON4L, GON4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
ABCD1_MOUSE
|
||||||
| NC score | 0.026321 (rank : 20) | θ value | 0.365318 (rank : 20) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P48410, Q9QY41, Q9QZ32 | Gene names | Abcd1, Ald, Aldgh | |||
|
Domain Architecture |

|
|||||
| Description | ATP-binding cassette sub-family D member 1 (Adrenoleukodystrophy protein) (ALDP). | |||||
|
ABCD1_HUMAN
|
||||||
| NC score | 0.025073 (rank : 21) | θ value | 0.47712 (rank : 21) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P33897, Q6GTZ2 | Gene names | ABCD1, ALD | |||
|
Domain Architecture |

|
|||||
| Description | ATP-binding cassette sub-family D member 1 (Adrenoleukodystrophy protein) (ALDP). | |||||
|
FBX11_HUMAN
|
||||||
| NC score | 0.023725 (rank : 22) | θ value | 1.06291 (rank : 23) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q86XK2, Q8IXG3, Q96E90, Q9H6V8, Q9H9L1, Q9NR14, Q9UFK1, Q9UHI1, Q9UKC2 | Gene names | FBXO11, FBX11, VIT1 | |||
|
Domain Architecture |

|
|||||
| Description | F-box only protein 11 (Vitiligo-associated protein VIT-1). | |||||
|
GIMA7_HUMAN
|
||||||
| NC score | 0.022772 (rank : 23) | θ value | 0.279714 (rank : 19) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NHV1 | Gene names | GIMAP7, IAN7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GTPase, IMAP family member 7 (Immunity-associated nucleotide 7 protein). | |||||
|
RHG18_HUMAN
|
||||||
| NC score | 0.018984 (rank : 24) | θ value | 1.81305 (rank : 28) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N392, Q58EZ3, Q6P679, Q6PJD7, Q96S64 | Gene names | ARHGAP18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho-GTPase-activating protein 18 (MacGAP). | |||||
|
CAP2_MOUSE
|
||||||
| NC score | 0.018507 (rank : 25) | θ value | 1.81305 (rank : 26) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CYT6, Q80YU7, Q9D6L0 | Gene names | Cap2 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylyl cyclase-associated protein 2 (CAP 2). | |||||
|
OGT1_HUMAN
|
||||||
| NC score | 0.017375 (rank : 26) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15294, Q7Z3K0, Q8WWM8, Q96CC1, Q9UG57 | Gene names | OGT | |||
|
Domain Architecture |

|
|||||
| Description | UDP-N-acetylglucosamine--peptide N-acetylglucosaminyltransferase 110 kDa subunit (EC 2.4.1.-) (O-GlcNAc transferase p110 subunit). | |||||
|
NU133_MOUSE
|
||||||
| NC score | 0.014316 (rank : 27) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R0G9 | Gene names | Nup133 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup133 (Nucleoporin Nup133) (133 kDa nucleoporin). | |||||
|
SNX13_HUMAN
|
||||||
| NC score | 0.013379 (rank : 28) | θ value | 5.27518 (rank : 41) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y5W8, O94821, Q8WVZ2, Q8WXH8 | Gene names | SNX13, KIAA0713 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-13 (RGS domain- and PHOX domain-containing protein) (RGS-PX1). | |||||
|
CAP2_HUMAN
|
||||||
| NC score | 0.013357 (rank : 29) | θ value | 8.99809 (rank : 47) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P40123, Q6IAY2 | Gene names | CAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylyl cyclase-associated protein 2 (CAP 2). | |||||
|
RHG18_MOUSE
|
||||||
| NC score | 0.012613 (rank : 30) | θ value | 4.03905 (rank : 38) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K0Q5, Q8BP03, Q8R196 | Gene names | Arhgap18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho-GTPase-activating protein 18. | |||||
|
PAPOG_MOUSE
|
||||||
| NC score | 0.012403 (rank : 31) | θ value | 1.38821 (rank : 25) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PCL9, Q8BZC9 | Gene names | Papolg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(A) polymerase gamma (EC 2.7.7.19) (PAP gamma) (Polynucleotide adenylyltransferase gamma) (SRP RNA 3' adenylating enzyme). | |||||
|
RYR3_HUMAN
|
||||||
| NC score | 0.008864 (rank : 32) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15413, O15175, Q15412 | Gene names | RYR3, HBRR | |||
|
Domain Architecture |

|
|||||
| Description | Ryanodine receptor 3 (Brain-type ryanodine receptor) (RyR3) (RYR-3) (Brain ryanodine receptor-calcium release channel). | |||||
|
SC24D_HUMAN
|
||||||
| NC score | 0.008534 (rank : 33) | θ value | 5.27518 (rank : 40) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O94855 | Gene names | SEC24D, KIAA0755 | |||
|
Domain Architecture |

|
|||||
| Description | Protein transport protein Sec24D (SEC24-related protein D). | |||||
|
HOOK3_MOUSE
|
||||||
| NC score | 0.008419 (rank : 34) | θ value | 5.27518 (rank : 39) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8BUK6, Q8BV48, Q8BW57, Q8BY41 | Gene names | Hook3 | |||
|
Domain Architecture |

|
|||||
| Description | Hook homolog 3. | |||||
|
PK1IP_HUMAN
|
||||||
| NC score | 0.008241 (rank : 35) | θ value | 1.81305 (rank : 27) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NWT1, Q5T4J2, Q96QJ8, Q96T87 | Gene names | PAK1IP1, PIP1, WDR84 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | p21-activated protein kinase-interacting protein 1 (PAK1-interacting protein 1) (PAK/PLC-interacting protein 1) (hPIP1) (WD repeat protein 84). | |||||
|
AATK_HUMAN
|
||||||
| NC score | 0.008082 (rank : 36) | θ value | 0.00869519 (rank : 15) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1243 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6ZMQ8, O75136, Q6ZN31, Q86X28 | Gene names | AATK, KIAA0641 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Apoptosis-associated tyrosine-protein kinase (EC 2.7.10.2) (AATYK) (Brain apoptosis-associated tyrosine kinase) (CDK5-binding protein) (p35-binding protein) (p35BP). | |||||
|
PHLB2_MOUSE
|
||||||
| NC score | 0.006976 (rank : 37) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1016 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8K1N2, Q3TNI3, Q80Y16, Q8BKV3 | Gene names | Phldb2, Ll5b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 2 (Protein LL5-beta). | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.006894 (rank : 38) | θ value | 5.27518 (rank : 42) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
TGFB3_MOUSE
|
||||||
| NC score | 0.006634 (rank : 39) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P17125 | Gene names | Tgfb3 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming growth factor beta-3 precursor (TGF-beta-3). | |||||
|
K1802_MOUSE
|
||||||
| NC score | 0.005908 (rank : 40) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
SL9A2_HUMAN
|
||||||
| NC score | 0.005717 (rank : 41) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UBY0 | Gene names | SLC9A2, NHE2 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/hydrogen exchanger 2 (Na(+)/H(+) exchanger 2) (NHE-2) (Solute carrier family 9 member 2). | |||||
|
SMRC2_MOUSE
|
||||||
| NC score | 0.005464 (rank : 42) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PDG5, Q6P6P3 | Gene names | Smarcc2, Baf170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
MYH11_HUMAN
|
||||||
| NC score | 0.005076 (rank : 43) | θ value | 0.47712 (rank : 22) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
PEX1_HUMAN
|
||||||
| NC score | 0.005050 (rank : 44) | θ value | 8.99809 (rank : 50) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43933, Q96S71, Q96S72, Q96S73, Q99994 | Gene names | PEX1 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisome biogenesis factor 1 (Peroxin-1) (Peroxisome biogenesis disorder protein 1). | |||||
|
GRIP2_HUMAN
|
||||||
| NC score | 0.004844 (rank : 45) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9C0E4, Q8TEH9, Q9H7H3 | Gene names | GRIP2, KIAA1719 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor-interacting protein 2 (GRIP2 protein). | |||||
|
BCAR1_MOUSE
|
||||||
| NC score | 0.004551 (rank : 46) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61140, Q60869 | Gene names | Bcar1, Cas, Crkas | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
MYH11_MOUSE
|
||||||
| NC score | 0.004304 (rank : 47) | θ value | 3.0926 (rank : 31) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
MYH9_HUMAN
|
||||||
| NC score | 0.004110 (rank : 48) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
TARA_MOUSE
|
||||||
| NC score | 0.003491 (rank : 49) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
PLCB4_HUMAN
|
||||||
| NC score | 0.003417 (rank : 50) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q15147, Q5JYS8, Q5JYT0, Q5JYT4, Q9BQW5, Q9BQW6, Q9BQW8, Q9UJQ2 | Gene names | PLCB4 | |||
|
Domain Architecture |

|
|||||
| Description | 1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase beta 4 (EC 3.1.4.11) (Phosphoinositide phospholipase C) (Phospholipase C- beta-4) (PLC-beta-4). | |||||
|
GPR64_HUMAN
|
||||||
| NC score | 0.002404 (rank : 51) | θ value | 8.99809 (rank : 48) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IZP9, O00406, Q8IWT2, Q8IZE4, Q8IZE5, Q8IZE6, Q8IZE7, Q8IZP3, Q8IZP4 | Gene names | GPR64, HE6, TM7LN2 | |||
|
Domain Architecture |

|
|||||
| Description | G-protein coupled receptor 64 precursor (Epididymis-specific protein 6) (He6 receptor). | |||||
|
GLIS3_HUMAN
|
||||||
| NC score | -0.001417 (rank : 52) | θ value | 4.03905 (rank : 34) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 758 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NEA6 | Gene names | GLIS3, ZNF515 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLIS3 (GLI-similar 3) (Zinc finger protein 515). | |||||
|
MARK2_HUMAN
|
||||||
| NC score | -0.001610 (rank : 53) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1056 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7KZI7, Q15449, Q15524, Q5XGA3, Q68A18, Q96HB3, Q96RG0 | Gene names | MARK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL motif kinase) (EMK1) (PAR1 homolog). | |||||