Please be patient as the page loads
|
ZFYV9_HUMAN
|
||||||
| SwissProt Accessions | O95405, Q9UNE1, Q9Y5R7 | Gene names | ZFYVE9, MADHIP, SARA, SMADIP | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger FYVE domain-containing protein 9 (Mothers against decapentaplegic homolog-interacting protein) (Madh-interacting protein) (Smad anchor for receptor activation) (Receptor activation anchor) (hSARA) (Novel serine protease) (NSP). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ZFY16_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.947491 (rank : 2) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q7Z3T8, O15023, Q7LAU7, Q86T69, Q8N5L3, Q8NEK3 | Gene names | ZFYVE16, KIAA0305 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 16 (Endofin) (Endosome- associated FYVE domain protein). | |||||
|
ZFYV9_HUMAN
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 134 | |
| SwissProt Accessions | O95405, Q9UNE1, Q9Y5R7 | Gene names | ZFYVE9, MADHIP, SARA, SMADIP | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger FYVE domain-containing protein 9 (Mothers against decapentaplegic homolog-interacting protein) (Madh-interacting protein) (Smad anchor for receptor activation) (Receptor activation anchor) (hSARA) (Novel serine protease) (NSP). | |||||
|
ZFY16_MOUSE
|
||||||
| θ value | 4.40161e-164 (rank : 3) | NC score | 0.945409 (rank : 3) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q80U44, Q8BRD2, Q8CG97 | Gene names | Zfyve16, Kiaa0305 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 16 (Endofin) (Endosomal- associated FYVE domain protein). | |||||
|
FGD6_HUMAN
|
||||||
| θ value | 4.02038e-16 (rank : 4) | NC score | 0.564634 (rank : 11) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q6ZV73, Q6ZR53, Q7Z2Z7, Q96D44, Q9NUR8, Q9P2I5 | Gene names | FGD6, KIAA1362, ZFYVE24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 6 (Zinc finger FYVE domain-containing protein 24). | |||||
|
FGD6_MOUSE
|
||||||
| θ value | 8.95645e-16 (rank : 5) | NC score | 0.553032 (rank : 14) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q69ZL1, Q8C8W5, Q8K3B0, Q9D3Y7 | Gene names | Fgd6, Kiaa1362 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 6. | |||||
|
ZFY21_MOUSE
|
||||||
| θ value | 7.09661e-13 (rank : 6) | NC score | 0.738308 (rank : 4) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q8VCM3, Q9D1E2 | Gene names | Zfyve21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 21. | |||||
|
FGD2_MOUSE
|
||||||
| θ value | 1.58096e-12 (rank : 7) | NC score | 0.508361 (rank : 21) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8BY35, O88841, Q7TSE3, Q8VDH4 | Gene names | Fgd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 2. | |||||
|
FGD2_HUMAN
|
||||||
| θ value | 2.0648e-12 (rank : 8) | NC score | 0.508113 (rank : 22) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q7Z6J4, Q5T8I1, Q6P6A8, Q6ZNL5, Q8IZ32, Q8N868, Q9H7M2 | Gene names | FGD2, ZFYVE4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 2 (Zinc finger FYVE domain-containing protein 4). | |||||
|
FGD4_HUMAN
|
||||||
| θ value | 4.59992e-12 (rank : 9) | NC score | 0.485848 (rank : 24) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q96M96, Q6ULS2, Q8TCP6 | Gene names | FGD4, FRABP, ZFYVE6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 4 (Actin filament- binding protein frabin) (FGD1-related F-actin-binding protein) (Zinc finger FYVE domain-containing protein 6). | |||||
|
PKHF2_HUMAN
|
||||||
| θ value | 4.59992e-12 (rank : 10) | NC score | 0.685107 (rank : 8) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9H8W4 | Gene names | PLEKHF2, ZFYVE18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family F member 2 (PH domain- containing family F member 2) (PH and FYVE domain-containing protein 2) (Phafin-2) (Zinc finger FYVE domain-containing protein 18). | |||||
|
PKHF2_MOUSE
|
||||||
| θ value | 4.59992e-12 (rank : 11) | NC score | 0.685663 (rank : 7) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q91WB4 | Gene names | Plekhf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family F member 2 (PH domain- containing family F member 2). | |||||
|
ZFY21_HUMAN
|
||||||
| θ value | 6.00763e-12 (rank : 12) | NC score | 0.730044 (rank : 5) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9BQ24, Q86T05, Q96LT1 | Gene names | ZFYVE21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 21. | |||||
|
FGD1_MOUSE
|
||||||
| θ value | 7.84624e-12 (rank : 13) | NC score | 0.480542 (rank : 28) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P52734 | Gene names | Fgd1 | |||
|
Domain Architecture |

|
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 1 (Faciogenital dysplasia 1 protein homolog) (Zinc finger FYVE domain-containing protein 3) (Rho/Rac guanine nucleotide exchange factor FGD1) (Rho/Rac GEF). | |||||
|
FGD4_MOUSE
|
||||||
| θ value | 1.33837e-11 (rank : 14) | NC score | 0.482009 (rank : 27) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q91ZT5, Q3UEB6, Q8BW60, Q8BZI7, Q91ZT3, Q91ZT4 | Gene names | Fgd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 4 (Actin filament- binding protein frabin) (FGD1-related F-actin-binding protein). | |||||
|
FGD1_HUMAN
|
||||||
| θ value | 1.74796e-11 (rank : 15) | NC score | 0.480035 (rank : 29) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 367 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P98174, Q8N4D9 | Gene names | FGD1, ZFYVE3 | |||
|
Domain Architecture |

|
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 1 (Faciogenital dysplasia 1 protein) (Zinc finger FYVE domain-containing protein 3) (Rho/Rac guanine nucleotide exchange factor FGD1) (Rho/Rac GEF). | |||||
|
ZFY28_HUMAN
|
||||||
| θ value | 3.89403e-11 (rank : 16) | NC score | 0.694392 (rank : 6) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9HCC9 | Gene names | ZFYVE28, KIAA1643 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger FYVE domain-containing protein 28. | |||||
|
FYV1_HUMAN
|
||||||
| θ value | 5.08577e-11 (rank : 17) | NC score | 0.516728 (rank : 19) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9Y2I7, Q8NB67 | Gene names | PIP5K3, KIAA0981, PIKFYVE | |||
|
Domain Architecture |

|
|||||
| Description | FYVE finger-containing phosphoinositide kinase (EC 2.7.1.68) (1- phosphatidylinositol-4-phosphate 5-kinase) (Phosphatidylinositol-3- phosphate 5-kinase type III) (PIP5K) (PtdIns(4)P-5-kinase) (PIKfyve) (p235). | |||||
|
FYV1_MOUSE
|
||||||
| θ value | 5.08577e-11 (rank : 18) | NC score | 0.515200 (rank : 20) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9Z1T6 | Gene names | Pip5k3, Pikfyve | |||
|
Domain Architecture |

|
|||||
| Description | FYVE finger-containing phosphoinositide kinase (EC 2.7.1.68) (1- phosphatidylinositol-4-phosphate 5-kinase) (PIP5K) (PtdIns(4)P-5- kinase) (PIKfyve) (p235). | |||||
|
FGD5_HUMAN
|
||||||
| θ value | 1.133e-10 (rank : 19) | NC score | 0.553273 (rank : 13) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q6ZNL6, Q6MZY1, Q7Z303, Q8IYP3, Q8N861, Q8N8G4 | Gene names | FGD5, ZFYVE23 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 5 (Zinc finger FYVE domain-containing protein 23). | |||||
|
MTMR3_HUMAN
|
||||||
| θ value | 1.47974e-10 (rank : 20) | NC score | 0.398026 (rank : 35) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q13615, Q9NYN5, Q9NYN6, Q9UDX6, Q9UEG3 | Gene names | MTMR3, KIAA0371, ZFYVE10 | |||
|
Domain Architecture |

|
|||||
| Description | Myotubularin-related protein 3 (EC 3.1.3.48) (FYVE domain-containing dual specificity protein phosphatase 1) (FYVE-DSP1) (Zinc finger FYVE domain-containing protein 10). | |||||
|
RUFY2_HUMAN
|
||||||
| θ value | 1.47974e-10 (rank : 21) | NC score | 0.308666 (rank : 44) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1208 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q8WXA3, Q96P51, Q9P1Z1 | Gene names | RUFY2, KIAA1537, RABIP4R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 2 (Rab4-interacting protein related). | |||||
|
EEA1_HUMAN
|
||||||
| θ value | 2.52405e-10 (rank : 22) | NC score | 0.216626 (rank : 50) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1796 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q15075, Q14221 | Gene names | EEA1, ZFYVE2 | |||
|
Domain Architecture |

|
|||||
| Description | Early endosome antigen 1 (Endosome-associated protein p162) (Zinc finger FYVE domain-containing protein 2). | |||||
|
EEA1_MOUSE
|
||||||
| θ value | 2.52405e-10 (rank : 23) | NC score | 0.234057 (rank : 49) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1637 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q8BL66, Q6DIC2 | Gene names | Eea1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Early endosome antigen 1. | |||||
|
RUFY1_HUMAN
|
||||||
| θ value | 2.52405e-10 (rank : 24) | NC score | 0.380995 (rank : 38) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 848 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q96T51, Q59FF3, Q71S93, Q9H6I3 | Gene names | RUFY1, RABIP4, ZFYVE12 | |||
|
Domain Architecture |

|
|||||
| Description | RUN and FYVE domain-containing protein 1 (FYVE-finger protein EIP1) (Zinc finger FYVE domain-containing protein 12) (La-binding protein 1) (Rab4-interacting protein). | |||||
|
RUFY2_MOUSE
|
||||||
| θ value | 2.52405e-10 (rank : 25) | NC score | 0.386642 (rank : 37) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 820 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q8R4C2, Q69ZH1 | Gene names | Rufy2, Kiaa1537 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 2 (Leucine zipper FYVE-finger protein) (LZ-FYVE). | |||||
|
FGD5_MOUSE
|
||||||
| θ value | 3.29651e-10 (rank : 26) | NC score | 0.561682 (rank : 12) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q80UZ0, Q8BHM5 | Gene names | Fgd5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 5. | |||||
|
PKHF1_MOUSE
|
||||||
| θ value | 3.29651e-10 (rank : 27) | NC score | 0.670830 (rank : 10) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q3TB82, Q99M16 | Gene names | Plekhf1, Lapf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family F member 1 (PH domain- containing family F member 1) (Lysosome-associated apoptosis-inducing protein containing PH and FYVE domains). | |||||
|
PKHF1_HUMAN
|
||||||
| θ value | 4.30538e-10 (rank : 28) | NC score | 0.673172 (rank : 9) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q96S99, Q96K11, Q9BUB9 | Gene names | PLEKHF1, APPD, LAPF, ZFYVE15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family F member 1 (PH domain- containing family F member 1) (PH and FYVE domain-containing protein 1) (Phafin-1) (Lysosome-associated apoptosis-inducing protein containing PH and FYVE domains) (Apoptosis-inducing protein) (Zinc finger FYVE domain-containing protein 15). | |||||
|
RUFY1_MOUSE
|
||||||
| θ value | 4.30538e-10 (rank : 29) | NC score | 0.388606 (rank : 36) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 765 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q8BIJ7, Q8BKQ4, Q8BL21, Q9EPM6 | Gene names | Rufy1, Rabip4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 1 (Rab4-interacting protein). | |||||
|
HGS_MOUSE
|
||||||
| θ value | 2.79066e-09 (rank : 30) | NC score | 0.523924 (rank : 18) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q99LI8, Q61691, Q8BQW3 | Gene names | Hgs, Hrs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate. | |||||
|
HGS_HUMAN
|
||||||
| θ value | 8.11959e-09 (rank : 31) | NC score | 0.524554 (rank : 17) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | O14964, Q9NR36 | Gene names | HGS, HRS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate (Protein pp110) (Hrs). | |||||
|
WDFY3_MOUSE
|
||||||
| θ value | 1.38499e-08 (rank : 32) | NC score | 0.378903 (rank : 39) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q6VNB8, Q8C8H7 | Gene names | Wdfy3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and FYVE domain-containing protein 3 (Beach domain, WD repeat and FYVE domain-containing protein 1) (BWF1). | |||||
|
FYCO1_HUMAN
|
||||||
| θ value | 1.80886e-08 (rank : 33) | NC score | 0.267908 (rank : 45) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1149 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9BQS8, Q3MJE6, Q86T41, Q86TB1, Q8TEF9, Q96IV5, Q9H8P9 | Gene names | FYCO1, ZFYVE7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1 (Zinc finger FYVE domain-containing protein 7). | |||||
|
WDFY3_HUMAN
|
||||||
| θ value | 1.80886e-08 (rank : 34) | NC score | 0.377244 (rank : 40) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8IZQ1, Q4W5K5, Q6P0Q5, Q8N1T2, Q96BS7, Q96N85, Q9Y2J7 | Gene names | WDFY3, KIAA0993 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and FYVE domain-containing protein 3 (Autophagy-linked FYVE protein) (Alfy). | |||||
|
ZFYV1_HUMAN
|
||||||
| θ value | 1.69304e-06 (rank : 35) | NC score | 0.542479 (rank : 15) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9HBF4, Q8WYX7, Q96K57, Q9BXP9, Q9HCI3 | Gene names | ZFYVE1, DFCP1, KIAA1589, TAFF1, ZNFN2A1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger FYVE domain-containing protein 1 (Double FYVE-containing protein 1) (Tandem FYVE fingers-1) (SR3). | |||||
|
ZFY19_HUMAN
|
||||||
| θ value | 2.21117e-06 (rank : 36) | NC score | 0.488111 (rank : 23) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q96K21, Q86WC2, Q8WU96 | Gene names | ZFYVE19, MPFYVE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 19 (MLL partner containing FYVE domain). | |||||
|
ANFY1_MOUSE
|
||||||
| θ value | 3.77169e-06 (rank : 37) | NC score | 0.252957 (rank : 46) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 480 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q810B6, O54807, Q80TG6, Q80UH8 | Gene names | Ankfy1, Ankhzn, Kiaa1255 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and FYVE domain-containing protein 1 (Ankyrin repeats hooked to a zinc finger motif). | |||||
|
RBNS5_HUMAN
|
||||||
| θ value | 4.92598e-06 (rank : 38) | NC score | 0.482745 (rank : 26) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9H1K0, Q3KP30, Q59EY8, Q8NAQ1 | Gene names | ZFYVE20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rabenosyn-5 (FYVE finger-containing Rab5 effector protein rabenosyn-5) (Zinc finger FYVE domain-containing protein 20) (110 kDa protein). | |||||
|
RBNS5_MOUSE
|
||||||
| θ value | 4.92598e-06 (rank : 39) | NC score | 0.478140 (rank : 30) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q80Y56, Q8K0L6, Q9CTW0 | Gene names | Zfyve20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rabenosyn-5 (FYVE finger-containing Rab5 effector protein rabenosyn-5) (Zinc finger FYVE domain-containing protein 20). | |||||
|
ZFYV1_MOUSE
|
||||||
| θ value | 4.92598e-06 (rank : 40) | NC score | 0.537297 (rank : 16) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q810J8 | Gene names | Zfyve1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger FYVE domain-containing protein 1. | |||||
|
FYCO1_MOUSE
|
||||||
| θ value | 8.40245e-06 (rank : 41) | NC score | 0.246498 (rank : 48) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8VDC1, Q3V385, Q7TMT0, Q8BJN2 | Gene names | Fyco1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1. | |||||
|
ANFY1_HUMAN
|
||||||
| θ value | 1.09739e-05 (rank : 42) | NC score | 0.248644 (rank : 47) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 480 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9P2R3, Q9ULG5 | Gene names | ANKFY1, ANKHZN, KIAA1255 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and FYVE domain-containing protein 1 (Ankyrin repeats hooked to a zinc finger motif). | |||||
|
FGD3_HUMAN
|
||||||
| θ value | 1.09739e-05 (rank : 43) | NC score | 0.439806 (rank : 31) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q5JSP0, Q7Z7D9, Q8N5G1 | Gene names | FGD3, ZFYVE5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 3 (Zinc finger FYVE domain-containing protein 5). | |||||
|
ZFY19_MOUSE
|
||||||
| θ value | 1.87187e-05 (rank : 44) | NC score | 0.483226 (rank : 25) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9DAZ9, Q8VCV7 | Gene names | Zfyve19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 19. | |||||
|
ZFY27_HUMAN
|
||||||
| θ value | 5.44631e-05 (rank : 45) | NC score | 0.414509 (rank : 33) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5T4F4, Q5T4F1, Q5T4F2, Q5T4F3, Q8N1K0, Q8N6D6, Q8NCA0, Q8NDE4, Q96M08 | Gene names | ZFYVE27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 27. | |||||
|
ZFY27_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 46) | NC score | 0.418856 (rank : 32) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q3TXX3, Q8CFP8 | Gene names | Zfyve27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 27. | |||||
|
WDFY2_HUMAN
|
||||||
| θ value | 0.000121331 (rank : 47) | NC score | 0.357331 (rank : 43) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q96P53, Q96CS1 | Gene names | WDFY2, WDF2, ZFYVE22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and FYVE domain-containing protein 2 (WD40- and FYVE domain- containing protein 2) (Zinc finger FYVE domain-containing protein 22). | |||||
|
WDFY2_MOUSE
|
||||||
| θ value | 0.000121331 (rank : 48) | NC score | 0.360060 (rank : 42) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8BUB4 | Gene names | Wdfy2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and FYVE domain-containing protein 2 (WD40- and FYVE domain- containing protein 2). | |||||
|
WDFY1_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 49) | NC score | 0.367154 (rank : 41) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8IWB7, Q9H9D5, Q9P2B3 | Gene names | WDFY1, KIAA1435, WDF1, ZFYVE17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and FYVE domain-containing protein 1 (WD40- and FYVE domain- containing protein 1) (Phosphoinositide-binding protein 1) (FENS-1) (Zinc finger FYVE domain-containing protein 17). | |||||
|
FGD3_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 50) | NC score | 0.401254 (rank : 34) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O88842, Q8BQ72 | Gene names | Fgd3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 3. | |||||
|
DSPP_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 51) | NC score | 0.088359 (rank : 85) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
RFFL_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 52) | NC score | 0.172148 (rank : 52) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8WZ73, Q8NHW0, Q8TBY7, Q96BE6 | Gene names | RFFL, RNF189, RNF34L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rififylin (RING finger and FYVE-like domain-containing protein 1) (FYVE-RING finger protein Sakura) (Fring) (Caspases-8 and -10- associated RING finger protein 2) (CARP-2) (Caspase regulator CARP2) (RING finger protein 189) (RING finger protein 34-like). | |||||
|
RFFL_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 53) | NC score | 0.184984 (rank : 51) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q6ZQM0, Q5SVC2, Q5SVC4, Q9D543, Q9D9B1 | Gene names | Rffl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rififylin (RING finger and FYVE-like domain-containing protein 1) (Fring). | |||||
|
CJ118_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 54) | NC score | 0.051043 (rank : 121) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q7Z3E2, Q2M2V6, Q6NS91, Q7RTP1, Q8N117, Q8N3G3, Q8N6C2, Q9NWA3 | Gene names | C10orf118 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf118 (CTCL tumor antigen HD-CL-01/L14-2). | |||||
|
CJ118_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 55) | NC score | 0.051927 (rank : 120) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1132 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8C9S4, Q6P3D6, Q8CJC0 | Gene names | Otg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf118 homolog (Oocyte-testis gene 1 protein). | |||||
|
MYRIP_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 56) | NC score | 0.072026 (rank : 94) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8K3I4, Q8CFC0, Q8K4H5 | Gene names | Myrip, Slac2c | |||
|
Domain Architecture |
|
|||||
| Description | Rab effector MyRIP (Myosin-VIIa- and Rab-interacting protein) (Exophilin-8) (Slp homolog lacking C2 domains c). | |||||
|
RP1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 57) | NC score | 0.043276 (rank : 124) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P56715 | Gene names | RP1, ORP1 | |||
|
Domain Architecture |
|
|||||
| Description | Oxygen-regulated protein 1 (Retinitis pigmentosa RP1 protein) (Retinitis pigmentosa 1 protein). | |||||
|
ZCHC6_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 58) | NC score | 0.036971 (rank : 129) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5VYS8, Q5H9T0, Q5VYS5, Q5VYS7, Q658Z9, Q659A2, Q6MZJ3, Q8N5F0, Q96N57, Q96NE8, Q9C0F2, Q9H8M6 | Gene names | ZCCHC6, KIAA1711 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 6. | |||||
|
PAF49_HUMAN
|
||||||
| θ value | 0.279714 (rank : 59) | NC score | 0.041050 (rank : 125) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O15446, Q32N11, Q7Z5U2, Q9UPF6 | Gene names | CD3EAP, ASE1, CAST, PAF49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase I-associated factor PAF49 (Anti-sense to ERCC-1 protein) (ASE-1) (CD3-epsilon-associated protein) (CD3E-associated protein) (CAST). | |||||
|
E2F4_HUMAN
|
||||||
| θ value | 0.365318 (rank : 60) | NC score | 0.026716 (rank : 139) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q16254, Q12991, Q15328 | Gene names | E2F4 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor E2F4 (E2F-4). | |||||
|
KS6A5_HUMAN
|
||||||
| θ value | 0.365318 (rank : 61) | NC score | 0.000949 (rank : 188) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 855 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75582, O95316, Q96AF7 | Gene names | RPS6KA5, MSK1 | |||
|
Domain Architecture |
|
|||||
| Description | Ribosomal protein S6 kinase alpha-5 (EC 2.7.11.1) (Nuclear mitogen- and stress-activated protein kinase 1) (90 kDa ribosomal protein S6 kinase 5) (RSK-like protein kinase) (RSKL). | |||||
|
BCAS1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 62) | NC score | 0.040248 (rank : 126) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75363, Q68CZ3 | Gene names | BCAS1, AIBC1, NABC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Breast carcinoma amplified sequence 1 (Novel amplified in breast cancer 1) (Amplified and overexpressed in breast cancer). | |||||
|
RANB9_HUMAN
|
||||||
| θ value | 0.47712 (rank : 63) | NC score | 0.033141 (rank : 131) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96S59, O94764, Q6P3T7, Q7LBR2, Q7Z7F9 | Gene names | RANBP9, RANBPM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 9 (RanBP9) (RanBP7) (Ran-binding protein M) (RanBPM) (BPM90) (BPM-L). | |||||
|
RNF34_HUMAN
|
||||||
| θ value | 0.47712 (rank : 64) | NC score | 0.169860 (rank : 54) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q969K3, Q8NG47, Q9H6W8 | Gene names | RNF34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 34 (RING finger protein RIFF) (FYVE-RING finger protein Momo) (Human RING finger homologous to inhibitor of apoptosis protein) (hRFI) (Caspases-8 and -10-associated RING finger protein 1) (CARP-1) (Caspase regulator CARP1). | |||||
|
DSPP_MOUSE
|
||||||
| θ value | 0.62314 (rank : 65) | NC score | 0.067899 (rank : 102) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
SPRL1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 66) | NC score | 0.028409 (rank : 135) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P70663, P97810, Q99L82 | Gene names | Sparcl1, Ecm2, Sc1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (Matrix glycoprotein Sc1) (Extracellular matrix protein 2). | |||||
|
ZHX2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 67) | NC score | 0.019776 (rank : 148) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8C0C0 | Gene names | Zhx2, Afr1, Raf | |||
|
Domain Architecture |

|
|||||
| Description | Zinc fingers and homeoboxes protein 2 (Zinc finger and homeodomain protein 2) (Alpha-fetoprotein regulator 1) (AFP regulator 1) (Regulator of AFP). | |||||
|
AFF4_HUMAN
|
||||||
| θ value | 0.813845 (rank : 68) | NC score | 0.019473 (rank : 150) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
KS6A5_MOUSE
|
||||||
| θ value | 0.813845 (rank : 69) | NC score | 0.000820 (rank : 189) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 854 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8C050, Q8CI92 | Gene names | Rps6ka5, Msk1 | |||
|
Domain Architecture |
|
|||||
| Description | Ribosomal protein S6 kinase alpha-5 (EC 2.7.11.1) (Nuclear mitogen- and stress-activated protein kinase 1) (90 kDa ribosomal protein S6 kinase 5) (RSK-like protein kinase) (RLSK). | |||||
|
RRMJ3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 70) | NC score | 0.035750 (rank : 130) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IY81, Q8N3A3, Q8WXX1, Q9BWM4, Q9NXT6 | Gene names | FTSJ3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative rRNA methyltransferase 3 (EC 2.1.1.-) (rRNA (uridine-2'-O-)- methyltransferase 3). | |||||
|
TRAIP_MOUSE
|
||||||
| θ value | 0.813845 (rank : 71) | NC score | 0.053060 (rank : 117) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 745 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8VIG6, O08854, Q922M8, Q9CPP4 | Gene names | Traip, Trip | |||
|
Domain Architecture |
|
|||||
| Description | TRAF-interacting protein. | |||||
|
ANR21_HUMAN
|
||||||
| θ value | 1.06291 (rank : 72) | NC score | 0.028431 (rank : 134) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q86YR6 | Gene names | ANKRD21, POTE | |||
|
Domain Architecture |
|
|||||
| Description | Ankyrin repeat domain-containing protein 21 (Protein POTE) (Prostate, ovary, testis-expressed protein). | |||||
|
CYLC1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 73) | NC score | 0.025662 (rank : 140) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
MYST4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 74) | NC score | 0.025257 (rank : 142) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
RNF34_MOUSE
|
||||||
| θ value | 1.06291 (rank : 75) | NC score | 0.170749 (rank : 53) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q99KR6, Q3UV45 | Gene names | Rnf34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 34 (RING finger protein RIFF) (Phafin-1). | |||||
|
SAMN1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 76) | NC score | 0.022419 (rank : 144) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P57725, Q8C6S2 | Gene names | Samsn1 | |||
|
Domain Architecture |

|
|||||
| Description | SAM domain-containing protein SAMSN-1 (SAM domain, SH3 domain and nuclear localization signals protein 1). | |||||
|
DAB2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 77) | NC score | 0.028930 (rank : 133) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P98082, Q13598, Q9BTY0, Q9UK04 | Gene names | DAB2, DOC2 | |||
|
Domain Architecture |

|
|||||
| Description | Disabled homolog 2 (Differentially expressed protein 2) (DOC-2). | |||||
|
K0423_HUMAN
|
||||||
| θ value | 1.81305 (rank : 78) | NC score | 0.027013 (rank : 138) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y4F4, Q68D66, Q6PG27 | Gene names | KIAA0423 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0423. | |||||
|
MBD6_HUMAN
|
||||||
| θ value | 1.81305 (rank : 79) | NC score | 0.025464 (rank : 141) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96DN6, Q8N3M0, Q8NA81, Q96Q00 | Gene names | MBD6, KIAA1887 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 6 (Methyl-CpG-binding protein MBD6). | |||||
|
SOX6_MOUSE
|
||||||
| θ value | 1.81305 (rank : 80) | NC score | 0.015945 (rank : 158) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P40645, Q62250, Q9QWS5 | Gene names | Sox6, Sox-6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-6 (SOX-LZ). | |||||
|
DMP1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 81) | NC score | 0.031379 (rank : 132) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
MAP9_HUMAN
|
||||||
| θ value | 2.36792 (rank : 82) | NC score | 0.039639 (rank : 127) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q49MG5, Q4W5I7, Q68DU1, Q9H781, Q9H7B6 | Gene names | MAP9, ASAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 9 (Aster-associated protein). | |||||
|
POT15_HUMAN
|
||||||
| θ value | 2.36792 (rank : 83) | NC score | 0.027987 (rank : 136) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6S5H4, Q6NXN7, Q6S5H7 | Gene names | POTE15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prostate, ovary, testis-expressed protein on chromosome 15. | |||||
|
AIM1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 84) | NC score | 0.027564 (rank : 137) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y4K1, O00296, Q5VWJ2 | Gene names | AIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Absent in melanoma 1 protein. | |||||
|
CA103_MOUSE
|
||||||
| θ value | 3.0926 (rank : 85) | NC score | 0.022709 (rank : 143) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CDD9, Q8C893 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C1orf103 homolog. | |||||
|
FLRT3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 86) | NC score | 0.003017 (rank : 185) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NZU0, Q96K39, Q96K42, Q96KB1, Q9P259 | Gene names | FLRT3, KIAA1469 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat transmembrane protein FLRT3 precursor (Fibronectin-like domain-containing leucine-rich transmembrane protein 3). | |||||
|
HSF1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 87) | NC score | 0.018641 (rank : 152) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P38532, O70462 | Gene names | Hsf1 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock factor protein 1 (HSF 1) (Heat shock transcription factor 1) (HSTF 1). | |||||
|
NOL4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 88) | NC score | 0.018673 (rank : 151) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P60954 | Gene names | Nol4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar protein 4. | |||||
|
SET1A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 89) | NC score | 0.018147 (rank : 154) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
UBP10_HUMAN
|
||||||
| θ value | 3.0926 (rank : 90) | NC score | 0.013059 (rank : 164) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14694, Q9BWG7, Q9NSL7 | Gene names | USP10, KIAA0190 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 10 (EC 3.1.2.15) (Ubiquitin thioesterase 10) (Ubiquitin-specific-processing protease 10) (Deubiquitinating enzyme 10). | |||||
|
ZBT38_HUMAN
|
||||||
| θ value | 3.0926 (rank : 91) | NC score | -0.001820 (rank : 194) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 885 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8NAP3 | Gene names | ZBTB38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and BTB domain-containing protein 38. | |||||
|
NCOR2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 92) | NC score | 0.011455 (rank : 168) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
S13A4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 93) | NC score | 0.011436 (rank : 169) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UKG4 | Gene names | SLC13A4, SUT1 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 13 member 4 (Na(+)/sulfate cotransporter SUT-1). | |||||
|
ZBT24_MOUSE
|
||||||
| θ value | 4.03905 (rank : 94) | NC score | -0.003897 (rank : 196) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 894 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80X44, Q3UIB7, Q7TPC4, Q8CJC9 | Gene names | Zbtb24, Bif1, Bsg1, Znf450 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and BTB domain-containing protein 24 (Zinc finger protein 450) (Bone morphogenetic protein-induced factor 1) (Brain-specific protein 1). | |||||
|
ADA29_HUMAN
|
||||||
| θ value | 5.27518 (rank : 95) | NC score | 0.002647 (rank : 186) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UKF5, Q9UHP1, Q9UKF3, Q9UKF4 | Gene names | ADAM29 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 29 precursor (A disintegrin and metalloproteinase domain 29). | |||||
|
ATP4A_MOUSE
|
||||||
| θ value | 5.27518 (rank : 96) | NC score | 0.003532 (rank : 184) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q64436, Q9CV46 | Gene names | Atp4a | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 1 (EC 3.6.3.10) (Proton pump) (Gastric H+/K+ ATPase subunit alpha). | |||||
|
BAT2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 97) | NC score | 0.016116 (rank : 157) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
EVI2B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 98) | NC score | 0.016198 (rank : 156) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8VD58, Q3U1A3 | Gene names | Evi2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EVI2B protein precursor (Ecotropic viral integration site 2B protein). | |||||
|
HELC1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 99) | NC score | 0.012024 (rank : 166) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N3C0, O43738, Q9H1I9, Q9NTR0 | Gene names | ASCC3, HELIC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Activating signal cointegrator 1 complex subunit 3 (EC 3.6.1.-) (ASC-1 complex subunit p200) (Trip4 complex subunit p200) (Helicase, ATP binding 1). | |||||
|
MK07_MOUSE
|
||||||
| θ value | 5.27518 (rank : 100) | NC score | -0.001854 (rank : 195) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1054 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9WVS8 | Gene names | Mapk7, Erk5 | |||
|
Domain Architecture |
|
|||||
| Description | Mitogen-activated protein kinase 7 (EC 2.7.11.24) (Extracellular signal-regulated kinase 5) (ERK-5) (BMK1 kinase). | |||||
|
PCDBG_HUMAN
|
||||||
| θ value | 5.27518 (rank : 101) | NC score | 0.000040 (rank : 192) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NRJ7, Q8IYD5, Q96SE9, Q9HCF1 | Gene names | PCDHB16, KIAA1621, PCDH3X | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 16 precursor (PCDH-beta16) (Protocadherin 3X). | |||||
|
TMF1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 102) | NC score | 0.037813 (rank : 128) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 987 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P82094 | Gene names | TMF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA element modulatory factor (TMF). | |||||
|
BTBDC_HUMAN
|
||||||
| θ value | 6.88961 (rank : 103) | NC score | 0.016426 (rank : 155) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8IY92, Q96JP1 | Gene names | BTBD12, KIAA1784 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein 12. | |||||
|
CCD82_HUMAN
|
||||||
| θ value | 6.88961 (rank : 104) | NC score | 0.020274 (rank : 145) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N4S0, Q9H2Q5, Q9H5E3 | Gene names | CCDC82 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 82. | |||||
|
CCDC4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 105) | NC score | 0.018636 (rank : 153) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6ZU67, Q58A26, Q58A27 | Gene names | CCDC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 4. | |||||
|
CSMD3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 106) | NC score | 0.002289 (rank : 187) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z407, Q96PZ3 | Gene names | CSMD3, KIAA1894 | |||
|
Domain Architecture |

|
|||||
| Description | CUB and sushi domain-containing protein 3 precursor (CUB and sushi multiple domains protein 3). | |||||
|
DAB2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 107) | NC score | 0.019491 (rank : 149) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P98078, Q3U3K1, Q91W56, Q923E1 | Gene names | Dab2, Doc2 | |||
|
Domain Architecture |
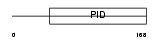
|
|||||
| Description | Disabled homolog 2 (DOC-2) (Mitogen-responsive phosphoprotein). | |||||
|
ICAL_HUMAN
|
||||||
| θ value | 6.88961 (rank : 108) | NC score | 0.011237 (rank : 170) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P20810, O95360, Q96D08, Q9H1Z5 | Gene names | CAST | |||
|
Domain Architecture |

|
|||||
| Description | Calpastatin (Calpain inhibitor) (Sperm BS-17 component). | |||||
|
JHD2C_HUMAN
|
||||||
| θ value | 6.88961 (rank : 109) | NC score | 0.012567 (rank : 165) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15652, Q5SQZ8, Q5SQZ9, Q5SR00, Q7Z3E7, Q8N3U0, Q96KB9, Q9P2G7 | Gene names | JMJD1C, JHDM2C, KIAA1380, TRIP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable JmjC domain-containing histone demethylation protein 2C (EC 1.14.11.-) (Jumonji domain-containing protein 1C) (Thyroid receptor-interacting protein 8) (TRIP-8). | |||||
|
MOCS3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 110) | NC score | 0.014898 (rank : 161) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95396 | Gene names | MOCS3 | |||
|
Domain Architecture |
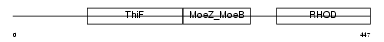
|
|||||
| Description | Molybdenum cofactor synthesis protein 3 (Molybdopterin synthase sulfurylase) (MPT synthase sulfurylase). | |||||
|
RFX3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 111) | NC score | 0.011210 (rank : 172) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P48380, Q5JTL7, Q5JTL8, Q6NW13, Q8WTU4, Q95HL5, Q95HL6 | Gene names | RFX3 | |||
|
Domain Architecture |
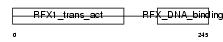
|
|||||
| Description | Transcription factor RFX3. | |||||
|
RFX3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 112) | NC score | 0.011217 (rank : 171) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P48381, Q8BLW2, Q8C0R3, Q8VBY6 | Gene names | Rfx3 | |||
|
Domain Architecture |
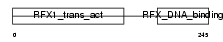
|
|||||
| Description | Transcription factor RFX3. | |||||
|
S28A2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 113) | NC score | 0.010669 (rank : 174) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O88627 | Gene names | Slc28a2, Cnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/nucleoside cotransporter 2 (Na(+)/nucleoside cotransporter 2) (Sodium-coupled nucleoside transporter 2) (Concentrative nucleoside transporter 2) (CNT 2) (Sodium/purine nucleoside cotransporter) (SPNT). | |||||
|
STAP2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 114) | NC score | 0.012000 (rank : 167) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R0L1, Q8BWS2 | Gene names | Stap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-transducing adaptor protein 2 (STAP-2). | |||||
|
THIC_MOUSE
|
||||||
| θ value | 6.88961 (rank : 115) | NC score | 0.007200 (rank : 178) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CAY6, Q3TJY7, Q62292, Q96EB7, Q99K88, Q9CYV2 | Gene names | Acat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acetyl-CoA acetyltransferase, cytosolic (EC 2.3.1.9) (Cytosolic acetoacetyl-CoA thiolase). | |||||
|
UTY_MOUSE
|
||||||
| θ value | 6.88961 (rank : 116) | NC score | 0.009794 (rank : 176) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P79457, O97979 | Gene names | Uty | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitously transcribed Y chromosome tetratricopeptide repeat protein (Ubiquitously transcribed TPR protein ON the Y chromosome) (Male- specific histocompatibility antigen H-YDB). | |||||
|
WDTC1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 117) | NC score | 0.019923 (rank : 146) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 299 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8N5D0, Q9NV87, Q9UPW4 | Gene names | WDTC1, KIAA1037 | |||
|
Domain Architecture |

|
|||||
| Description | WD and tetratricopeptide repeats protein 1. | |||||
|
WDTC1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 118) | NC score | 0.019787 (rank : 147) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80ZK9 | Gene names | Wdtc1 | |||
|
Domain Architecture |

|
|||||
| Description | WD and tetratricopeptide repeats protein 1. | |||||
|
ARID2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 119) | NC score | 0.013421 (rank : 163) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q68CP9, Q5EB51, Q645I3, Q6ZRY5, Q7Z3I5, Q86T28, Q96SJ6, Q9HCL5 | Gene names | ARID2, KIAA1557 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-rich interactive domain-containing protein 2 (ARID domain- containing protein 2) (BRG1-associated factor 200) (BAF200). | |||||
|
CCNB1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 120) | NC score | 0.005071 (rank : 182) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P14635 | Gene names | CCNB1, CCNB | |||
|
Domain Architecture |
|
|||||
| Description | G2/mitotic-specific cyclin-B1. | |||||
|
CLH2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | 0.007175 (rank : 179) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P53675, Q14017, Q15808, Q15809 | Gene names | CLTCL1, CLH22, CLTCL, CLTD | |||
|
Domain Architecture |

|
|||||
| Description | Clathrin heavy chain 2 (CLH-22). | |||||
|
EYA4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 122) | NC score | 0.004938 (rank : 183) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95677, O95464, O95679, Q8IW39, Q9NTR7 | Gene names | EYA4 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 4 (EC 3.1.3.48). | |||||
|
FA53B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | 0.010540 (rank : 175) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q14153, Q8N5S6 | Gene names | FAM53B, KIAA0140 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM53B. | |||||
|
HTF4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.005846 (rank : 181) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99081 | Gene names | TCF12, HEB, HTF4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 12 (Transcription factor HTF-4) (E-box-binding protein) (DNA-binding protein HTF4). | |||||
|
IDUA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 125) | NC score | 0.015036 (rank : 159) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P35475 | Gene names | IDUA | |||
|
Domain Architecture |
|
|||||
| Description | Alpha-L-iduronidase precursor (EC 3.2.1.76). | |||||
|
IDUA_MOUSE
|
||||||
| θ value | 8.99809 (rank : 126) | NC score | 0.014956 (rank : 160) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P48441 | Gene names | Idua | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-L-iduronidase precursor (EC 3.2.1.76). | |||||
|
IRF1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 127) | NC score | 0.006363 (rank : 180) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P15314 | Gene names | Irf1 | |||
|
Domain Architecture |
|
|||||
| Description | Interferon regulatory factor 1 (IRF-1). | |||||
|
NEDD1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 128) | NC score | 0.009089 (rank : 177) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 294 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NHV4, Q8NA30 | Gene names | NEDD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein NEDD1. | |||||
|
PTN13_MOUSE
|
||||||
| θ value | 8.99809 (rank : 129) | NC score | 0.000060 (rank : 191) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q64512, Q61494, Q62135, Q64499 | Gene names | Ptpn13, Ptp14 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 13 (EC 3.1.3.48) (Protein tyrosine phosphatase PTP-BL) (Protein-tyrosine phosphatase RIP) (protein tyrosine phosphatase DPZPTP) (PTP36). | |||||
|
RIMS2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 130) | NC score | 0.013641 (rank : 162) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9EQZ7, Q8C433, Q8CCK2 | Gene names | Rims2, Rab3ip2, Rim2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2) (Rab3-interacting protein 2). | |||||
|
RNF31_MOUSE
|
||||||
| θ value | 8.99809 (rank : 131) | NC score | 0.011030 (rank : 173) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q924T7 | Gene names | Rnf31 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 31. | |||||
|
TITIN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 132) | NC score | -0.001074 (rank : 193) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
TRI10_HUMAN
|
||||||
| θ value | 8.99809 (rank : 133) | NC score | 0.000731 (rank : 190) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UDY6, Q86Z08, Q96QB6, Q9C023, Q9C024 | Gene names | TRIM10, RFB30, RNF9 | |||
|
Domain Architecture |
|
|||||
| Description | Tripartite motif-containing protein 10 (RING finger protein 9) (B30- RING finger protein). | |||||
|
ZNF18_MOUSE
|
||||||
| θ value | 8.99809 (rank : 134) | NC score | -0.004238 (rank : 197) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 743 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q810A1, Q9D972 | Gene names | Znf18, Zfp535 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 18 (Zinc finger protein 535). | |||||
|
ABR_HUMAN
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.088868 (rank : 83) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q12979, Q13693, Q13694 | Gene names | ABR | |||
|
Domain Architecture |

|
|||||
| Description | Active breakpoint cluster region-related protein. | |||||
|
ALS2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.058957 (rank : 108) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q920R0, Q8JZR1, Q9CXJ3 | Gene names | Als2 | |||
|
Domain Architecture |

|
|||||
| Description | Alsin (Amyotrophic lateral sclerosis protein 2 homolog). | |||||
|
ARHG1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.088610 (rank : 84) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q92888, O00513, Q8N4J4, Q96BF4, Q96F17, Q9BSB1 | Gene names | ARHGEF1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 1 (p115-RhoGEF) (p115RhoGEF) (115 kDa guanine nucleotide exchange factor) (Sub1.5). | |||||
|
ARHG1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.087898 (rank : 86) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61210, O89074, Q80YE8, Q80YE9, Q91VL3 | Gene names | Arhgef1, Lbcl2, Lsc | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 1 (Lymphoid blast crisis-like 2) (Lbc's second cousin). | |||||
|
ARHG2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.073184 (rank : 90) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92974, O75142, Q15079 | Gene names | ARHGEF2, KIAA0651, LFP40 | |||
|
Domain Architecture |
|
|||||
| Description | Rho/Rac guanine nucleotide exchange factor 2 (GEF-H1 protein) (Proliferating cell nucleolar antigen p40). | |||||
|
ARHG2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.100305 (rank : 72) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q60875, O09115 | Gene names | Arhgef2, Lbcl1, Lfc | |||
|
Domain Architecture |
|
|||||
| Description | Rho/Rac guanine nucleotide exchange factor 2 (Lymphoid blast crisis- like 1) (LBC'S first cousin) (Oncogene LFC) (RHOBIN). | |||||
|
ARHG3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 141) | NC score | 0.057863 (rank : 110) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NR81, Q6NUN3, Q7Z4U2, Q7Z5T2, Q9H7T4 | Gene names | ARHGEF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 3 (Exchange factor found in platelets and leukemic and neuronal tissues) (XPLN). | |||||
|
ARHG3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.059371 (rank : 106) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91X46, Q8CDM0, Q91VY4, Q99K14, Q9DC31 | Gene names | Arhgef3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 3. | |||||
|
ARHG4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.092416 (rank : 78) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NR80, Q9HDC6, Q9UPP0 | Gene names | ARHGEF4, KIAA1112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 4 (APC-stimulated guanine nucleotide exchange factor) (Asef). | |||||
|
ARHG4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.091699 (rank : 81) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TNR9, Q80TJ6 | Gene names | Arhgef4, Kiaa1112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 4 (APC-stimulated guanine nucleotide exchange factor) (Asef). | |||||
|
ARHG5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.123508 (rank : 61) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q12774 | Gene names | ARHGEF5, TIM | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 5 (Guanine nucleotide regulatory protein TIM) (Oncogene TIM) (p60 TIM) (Transforming immortalized mammary oncogene). | |||||
|
ARHG7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 146) | NC score | 0.069798 (rank : 98) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q14155, Q6P9G3, Q6PII2, Q86W63, Q8N3M1 | Gene names | ARHGEF7, COOL1, KIAA0142, P85SPR, PAK3BP, PIXB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 7 (PAK-interacting exchange factor beta) (Beta-Pix) (COOL-1) (p85). | |||||
|
ARHG7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.058494 (rank : 109) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9ES28, O08757, Q9ES27 | Gene names | Arhgef7, Pak3bp | |||
|
Domain Architecture |
|
|||||
| Description | Rho guanine nucleotide exchange factor 7 (PAK-interacting exchange factor beta) (Beta-Pix) (p85SPR). | |||||
|
ARHG8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.057837 (rank : 111) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7Z628, Q12773, Q96D82, Q99903, Q9UEN6 | Gene names | NET1, ARHGEF8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuroepithelial cell-transforming gene 1 protein (p65 Net1 proto- oncogene) (Rho guanine nucleotide exchange factor 8). | |||||
|
ARHG9_HUMAN
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.085659 (rank : 87) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43307, Q5JSL6 | Gene names | ARHGEF9, KIAA0424 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 9 (Rac/Cdc42 guanine nucleotide exchange factor 9) (Collybistin) (PEM-2 homolog). | |||||
|
ARHG9_MOUSE
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.084417 (rank : 88) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q3UTH8, Q3TQ60, Q80U06, Q8CAF9 | Gene names | Arhgef9, Kiaa0424 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 9 (Rac/Cdc42 guanine nucleotide exchange factor 9) (Collybistin). | |||||
|
ARHGA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.116253 (rank : 66) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O15013, O14665, Q68D55, Q8IWD9, Q8IY77 | Gene names | ARHGEF10, KIAA0294 | |||
|
Domain Architecture |
|
|||||
| Description | Rho guanine nucleotide exchange factor 10. | |||||
|
ARHGA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.116831 (rank : 65) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8C033, Q5DU38, Q80VH8, Q8BW76, Q922S7 | Gene names | Arhgef10, Kiaa0294 | |||
|
Domain Architecture |
|
|||||
| Description | Rho guanine nucleotide exchange factor 10. | |||||
|
ARHGB_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.070412 (rank : 97) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 375 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O15085 | Gene names | ARHGEF11, KIAA0380 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 11 (PDZ-RhoGEF). | |||||
|
ARHGC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.069573 (rank : 99) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NZN5, O15086 | Gene names | ARHGEF12, KIAA0382, LARG | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 12 (Leukemia-associated RhoGEF). | |||||
|
ARHGC_MOUSE
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.068839 (rank : 101) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8R4H2, Q80U18 | Gene names | Arhgef12, Kiaa0382, Larg | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 12 (Leukemia-associated RhoGEF). | |||||
|
ARHGF_HUMAN
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.110102 (rank : 69) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O94989, Q8N449, Q9H8B4 | Gene names | ARHGEF15, KIAA0915 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 15 (Vsm-RhoGEF). | |||||
|
ARHGG_HUMAN
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.106960 (rank : 71) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5VV41, Q86TF0, Q99434 | Gene names | ARHGEF16, NBR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 16. | |||||
|
ARHGG_MOUSE
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.099051 (rank : 73) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q3U5C8, Q501M8, Q8VCE8 | Gene names | Arhgef16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 16. | |||||
|
BCR_HUMAN
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.090603 (rank : 82) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P11274, P78501, Q12842 | Gene names | BCR, BCR1 | |||
|
Domain Architecture |

|
|||||
| Description | Breakpoint cluster region protein (EC 2.7.11.1) (NY-REN-26 antigen). | |||||
|
CJ064_HUMAN
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.080233 (rank : 89) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6ZS81, Q8N4A3 | Gene names | C10orf64 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C10orf64. | |||||
|
ECT2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.138210 (rank : 60) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9H8V3, Q9NSV8, Q9NVW9 | Gene names | ECT2 | |||
|
Domain Architecture |

|
|||||
| Description | ECT2 protein (Epithelial cell-transforming sequence 2 oncogene). | |||||
|
ECT2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.141727 (rank : 59) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q07139 | Gene names | Ect2 | |||
|
Domain Architecture |

|
|||||
| Description | ECT2 protein (Epithelial cell-transforming sequence 2 oncogene). | |||||
|
FARP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 163) | NC score | 0.147561 (rank : 58) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y4F1, Q6IQ29 | Gene names | FARP1, CDEP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 1 (Chondrocyte- derived ezrin-like protein). | |||||
|
FARP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 164) | NC score | 0.152465 (rank : 55) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O94887, Q53QM5, Q8WU27, Q9UFE7 | Gene names | FARP2, KIAA0793 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 2 (FERM domain including RhoGEF) (FIR). | |||||
|
FARP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 165) | NC score | 0.151102 (rank : 56) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q91VS8, Q3TAP2, Q69ZZ0 | Gene names | Farp2, Kiaa0793 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 2 (FERM domain including RhoGEF) (FIR). | |||||
|
GNRP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 166) | NC score | 0.091863 (rank : 80) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q13972, Q16027 | Gene names | RASGRF1, CDC25 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-releasing protein (GNRP) (Ras-specific nucleotide exchange factor CDC25) (Ras-specific guanine nucleotide-releasing factor). | |||||
|
GNRP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 167) | NC score | 0.092161 (rank : 79) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P27671 | Gene names | Rasgrf1, Cdc25, Grf1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-releasing protein (GNRP) (Ras-specific nucleotide exchange factor CDC25) (CDC25Mm). | |||||
|
HAPIP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 168) | NC score | 0.112082 (rank : 67) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O60229 | Gene names | HAPIP, DUO | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-associated protein-interacting protein (Protein Duo). | |||||
|
ITSN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 169) | NC score | 0.072704 (rank : 92) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1757 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q15811, O95216, Q9UET5, Q9UK60, Q9UNK1, Q9UNK2, Q9UQ92 | Gene names | ITSN1, ITSN, SH3D1A | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (SH3 domain-containing protein 1A) (SH3P17). | |||||
|
ITSN1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 170) | NC score | 0.071598 (rank : 95) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1669 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9Z0R4, Q9R143 | Gene names | Itsn1, Ese1, Itsn | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (EH and SH3 domains protein 1). | |||||
|
ITSN2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 171) | NC score | 0.072517 (rank : 93) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1173 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9NZM3, O95062, Q15812, Q9HAK4, Q9NXE6, Q9NYG0, Q9NZM2, Q9ULG4 | Gene names | ITSN2, KIAA1256, SH3D1B, SWAP | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-2 (SH3 domain-containing protein 1B) (SH3P18) (SH3P18-like WASP-associated protein). | |||||
|
ITSN2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 172) | NC score | 0.072931 (rank : 91) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1371 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Z0R6, Q9Z0R5 | Gene names | Itsn2, Ese2, Sh3d1B | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-2 (SH3 domain-containing protein 1B) (EH and SH3 domains protein 2) (EH domain and SH3 domain regulator of endocytosis 2). | |||||
|
LBC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 173) | NC score | 0.093197 (rank : 77) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q12802 | Gene names | LBC | |||
|
Domain Architecture |
|
|||||
| Description | LBC oncogene (P47) (Lymphoid blast crisis oncogene). | |||||
|
MCF2L_MOUSE
|
||||||
| θ value | θ > 10 (rank : 174) | NC score | 0.098156 (rank : 74) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q64096 | Gene names | Mcf2l, Dbs | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide exchange factor DBS (DBL's big sister) (MCF2- transforming sequence-like protein). | |||||
|
MCF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 175) | NC score | 0.096812 (rank : 75) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P10911, P14919, Q5JYJ4, Q8IUF3, Q8IUF4, Q9UJB3 | Gene names | MCF2, DBL | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene DBL (Proto-oncogene MCF-2) [Contains: MCF2-transforming protein; DBL-transforming protein]. | |||||
|
NBEAL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 176) | NC score | 0.051030 (rank : 122) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6ZS30, Q6Y876, Q6ZQY5, Q96Q30 | Gene names | NBEAL1, ALS2CR17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurobeachin-like 1 (Amyotrophic lateral sclerosis 2 chromosomal region candidate gene 17 protein). | |||||
|
NGEF_HUMAN
|
||||||
| θ value | θ > 10 (rank : 177) | NC score | 0.110213 (rank : 68) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8N5V2, Q53QQ4, Q53ST7, Q6GMQ5, Q9NQD6 | Gene names | NGEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ephexin-1 (Eph-interacting exchange protein) (Neuronal guanine nucleotide exchange factor). | |||||
|
NGEF_MOUSE
|
||||||
| θ value | θ > 10 (rank : 178) | NC score | 0.108068 (rank : 70) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8CHT1, Q8R204, Q923H2, Q9JHT9 | Gene names | Ngef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ephexin-1 (Eph-interacting exchange protein) (Neuronal guanine nucleotide exchange factor). | |||||
|
PKHA6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 179) | NC score | 0.065711 (rank : 103) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Y2H5 | Gene names | PLEKHA6, KIAA0969, PEPP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
PKHA6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 180) | NC score | 0.063182 (rank : 104) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q7TQG1, Q8K0J5 | Gene names | Plekha6, Pepp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
PKHG1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 181) | NC score | 0.121154 (rank : 64) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9ULL1, Q5T1F2 | Gene names | PLEKHG1, KIAA1209 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family G member 1. | |||||
|
PKHG4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 182) | NC score | 0.095072 (rank : 76) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q58EX7, Q4G0J8, Q4H485, Q56A69, Q9H7K4, Q9UFW0 | Gene names | PLEKHG4, PRTPHN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Puratrophin-1 (Pleckstrin homology domain-containing family G member 4) (Purkinje cell atrophy-associated protein 1). | |||||
|
PREX1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 183) | NC score | 0.148164 (rank : 57) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8TCU6, Q5JS95, Q5JS96, Q69YL2, Q7Z2L9, Q9BQH0, Q9BX55, Q9H4Q6, Q9P2D2, Q9UGQ4 | Gene names | PREX1, KIAA1415 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3,4,5-trisphosphate-dependent Rac exchanger 1 protein (P-Rex1 protein). | |||||
|
RGNEF_MOUSE
|
||||||
| θ value | θ > 10 (rank : 184) | NC score | 0.050088 (rank : 123) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P97433 | Gene names | Rgnef, Rhoip2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-guanine nucleotide exchange factor (Rho-interacting protein 2) (RhoGEF) (RIP2). | |||||
|
RUFY3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 185) | NC score | 0.122507 (rank : 63) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 594 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q7L099, O94948, Q9UI00 | Gene names | RUFY3, KIAA0871 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein RUFY3 (Rap2-interacting protein x) (RIPx). | |||||
|
RUFY3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 186) | NC score | 0.123456 (rank : 62) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 579 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9D394, Q3U348, Q3UF96, Q6PE64, Q8VD10 | Gene names | Rufy3, D5Bwg0860e, Ripx | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein RUFY3 (Rap2-interacting protein x) (RIPx). | |||||
|
SOS2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 187) | NC score | 0.052636 (rank : 118) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q07890, Q15503 | Gene names | SOS2 | |||
|
Domain Architecture |

|
|||||
| Description | Son of sevenless homolog 2 (SOS-2). | |||||
|
SWP70_HUMAN
|
||||||
| θ value | θ > 10 (rank : 188) | NC score | 0.057312 (rank : 112) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 973 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UH65, O75135, Q7LCY6, Q9P061, Q9P0Z8 | Gene names | SWAP70, KIAA0640 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Switch-associated protein 70 (SWAP-70). | |||||
|
SWP70_MOUSE
|
||||||
| θ value | θ > 10 (rank : 189) | NC score | 0.054337 (rank : 116) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1097 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q6A028, O88443, Q3TQR6, Q3UCA3, Q6P1D0 | Gene names | Swap70, Kiaa0640 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Switch-associated protein 70 (SWAP-70). | |||||
|
TIAM1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 190) | NC score | 0.071264 (rank : 96) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13009 | Gene names | TIAM1 | |||
|
Domain Architecture |

|
|||||
| Description | T-lymphoma invasion and metastasis-inducing protein 1 (TIAM-1 protein). | |||||
|
TIAM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 191) | NC score | 0.069312 (rank : 100) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q60610 | Gene names | Tiam1, Tiam-1 | |||
|
Domain Architecture |

|
|||||
| Description | T-lymphoma invasion and metastasis-inducing protein 1 (TIAM-1 protein). | |||||
|
VAV2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 192) | NC score | 0.052069 (rank : 119) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P52735 | Gene names | VAV2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
VAV2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 193) | NC score | 0.056409 (rank : 113) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 456 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q60992 | Gene names | Vav2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
VAV3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 194) | NC score | 0.056399 (rank : 114) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 398 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UKW4, O95230, Q9Y5X8 | Gene names | VAV3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-3. | |||||
|
VAV3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 195) | NC score | 0.055649 (rank : 115) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 404 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R0C8, Q7TS85 | Gene names | Vav3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-3. | |||||
|
VAV_HUMAN
|
||||||
| θ value | θ > 10 (rank : 196) | NC score | 0.061431 (rank : 105) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P15498, Q15860 | Gene names | VAV1, VAV | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene vav. | |||||
|
VAV_MOUSE
|
||||||
| θ value | θ > 10 (rank : 197) | NC score | 0.058961 (rank : 107) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 430 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P27870, Q8BTV7 | Gene names | Vav1, Vav | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene vav (p95vav). | |||||
|
ZFYV9_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 134 | |
| SwissProt Accessions | O95405, Q9UNE1, Q9Y5R7 | Gene names | ZFYVE9, MADHIP, SARA, SMADIP | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger FYVE domain-containing protein 9 (Mothers against decapentaplegic homolog-interacting protein) (Madh-interacting protein) (Smad anchor for receptor activation) (Receptor activation anchor) (hSARA) (Novel serine protease) (NSP). | |||||
|
ZFY16_HUMAN
|
||||||
| NC score | 0.947491 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q7Z3T8, O15023, Q7LAU7, Q86T69, Q8N5L3, Q8NEK3 | Gene names | ZFYVE16, KIAA0305 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 16 (Endofin) (Endosome- associated FYVE domain protein). | |||||
|
ZFY16_MOUSE
|
||||||
| NC score | 0.945409 (rank : 3) | θ value | 4.40161e-164 (rank : 3) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q80U44, Q8BRD2, Q8CG97 | Gene names | Zfyve16, Kiaa0305 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 16 (Endofin) (Endosomal- associated FYVE domain protein). | |||||
|
ZFY21_MOUSE
|
||||||
| NC score | 0.738308 (rank : 4) | θ value | 7.09661e-13 (rank : 6) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q8VCM3, Q9D1E2 | Gene names | Zfyve21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 21. | |||||
|
ZFY21_HUMAN
|
||||||
| NC score | 0.730044 (rank : 5) | θ value | 6.00763e-12 (rank : 12) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9BQ24, Q86T05, Q96LT1 | Gene names | ZFYVE21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 21. | |||||
|
ZFY28_HUMAN
|
||||||
| NC score | 0.694392 (rank : 6) | θ value | 3.89403e-11 (rank : 16) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9HCC9 | Gene names | ZFYVE28, KIAA1643 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger FYVE domain-containing protein 28. | |||||
|
PKHF2_MOUSE
|
||||||
| NC score | 0.685663 (rank : 7) | θ value | 4.59992e-12 (rank : 11) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q91WB4 | Gene names | Plekhf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family F member 2 (PH domain- containing family F member 2). | |||||
|
PKHF2_HUMAN
|
||||||
| NC score | 0.685107 (rank : 8) | θ value | 4.59992e-12 (rank : 10) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9H8W4 | Gene names | PLEKHF2, ZFYVE18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family F member 2 (PH domain- containing family F member 2) (PH and FYVE domain-containing protein 2) (Phafin-2) (Zinc finger FYVE domain-containing protein 18). | |||||
|
PKHF1_HUMAN
|
||||||
| NC score | 0.673172 (rank : 9) | θ value | 4.30538e-10 (rank : 28) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q96S99, Q96K11, Q9BUB9 | Gene names | PLEKHF1, APPD, LAPF, ZFYVE15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family F member 1 (PH domain- containing family F member 1) (PH and FYVE domain-containing protein 1) (Phafin-1) (Lysosome-associated apoptosis-inducing protein containing PH and FYVE domains) (Apoptosis-inducing protein) (Zinc finger FYVE domain-containing protein 15). | |||||
|
PKHF1_MOUSE
|
||||||
| NC score | 0.670830 (rank : 10) | θ value | 3.29651e-10 (rank : 27) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q3TB82, Q99M16 | Gene names | Plekhf1, Lapf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family F member 1 (PH domain- containing family F member 1) (Lysosome-associated apoptosis-inducing protein containing PH and FYVE domains). | |||||
|
FGD6_HUMAN
|
||||||
| NC score | 0.564634 (rank : 11) | θ value | 4.02038e-16 (rank : 4) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q6ZV73, Q6ZR53, Q7Z2Z7, Q96D44, Q9NUR8, Q9P2I5 | Gene names | FGD6, KIAA1362, ZFYVE24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 6 (Zinc finger FYVE domain-containing protein 24). | |||||
|
FGD5_MOUSE
|
||||||
| NC score | 0.561682 (rank : 12) | θ value | 3.29651e-10 (rank : 26) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q80UZ0, Q8BHM5 | Gene names | Fgd5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 5. | |||||
|
FGD5_HUMAN
|
||||||
| NC score | 0.553273 (rank : 13) | θ value | 1.133e-10 (rank : 19) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q6ZNL6, Q6MZY1, Q7Z303, Q8IYP3, Q8N861, Q8N8G4 | Gene names | FGD5, ZFYVE23 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 5 (Zinc finger FYVE domain-containing protein 23). | |||||
|
FGD6_MOUSE
|
||||||
| NC score | 0.553032 (rank : 14) | θ value | 8.95645e-16 (rank : 5) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q69ZL1, Q8C8W5, Q8K3B0, Q9D3Y7 | Gene names | Fgd6, Kiaa1362 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 6. | |||||
|
ZFYV1_HUMAN
|
||||||
| NC score | 0.542479 (rank : 15) | θ value | 1.69304e-06 (rank : 35) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9HBF4, Q8WYX7, Q96K57, Q9BXP9, Q9HCI3 | Gene names | ZFYVE1, DFCP1, KIAA1589, TAFF1, ZNFN2A1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger FYVE domain-containing protein 1 (Double FYVE-containing protein 1) (Tandem FYVE fingers-1) (SR3). | |||||
|
ZFYV1_MOUSE
|
||||||
| NC score | 0.537297 (rank : 16) | θ value | 4.92598e-06 (rank : 40) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q810J8 | Gene names | Zfyve1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger FYVE domain-containing protein 1. | |||||
|
HGS_HUMAN
|
||||||
| NC score | 0.524554 (rank : 17) | θ value | 8.11959e-09 (rank : 31) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | O14964, Q9NR36 | Gene names | HGS, HRS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate (Protein pp110) (Hrs). | |||||
|
HGS_MOUSE
|
||||||
| NC score | 0.523924 (rank : 18) | θ value | 2.79066e-09 (rank : 30) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q99LI8, Q61691, Q8BQW3 | Gene names | Hgs, Hrs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate. | |||||
|
FYV1_HUMAN
|
||||||
| NC score | 0.516728 (rank : 19) | θ value | 5.08577e-11 (rank : 17) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9Y2I7, Q8NB67 | Gene names | PIP5K3, KIAA0981, PIKFYVE | |||
|
Domain Architecture |

|
|||||
| Description | FYVE finger-containing phosphoinositide kinase (EC 2.7.1.68) (1- phosphatidylinositol-4-phosphate 5-kinase) (Phosphatidylinositol-3- phosphate 5-kinase type III) (PIP5K) (PtdIns(4)P-5-kinase) (PIKfyve) (p235). | |||||
|
FYV1_MOUSE
|
||||||
| NC score | 0.515200 (rank : 20) | θ value | 5.08577e-11 (rank : 18) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9Z1T6 | Gene names | Pip5k3, Pikfyve | |||
|
Domain Architecture |

|
|||||
| Description | FYVE finger-containing phosphoinositide kinase (EC 2.7.1.68) (1- phosphatidylinositol-4-phosphate 5-kinase) (PIP5K) (PtdIns(4)P-5- kinase) (PIKfyve) (p235). | |||||
|
FGD2_MOUSE
|
||||||
| NC score | 0.508361 (rank : 21) | θ value | 1.58096e-12 (rank : 7) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8BY35, O88841, Q7TSE3, Q8VDH4 | Gene names | Fgd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 2. | |||||
|
FGD2_HUMAN
|
||||||
| NC score | 0.508113 (rank : 22) | θ value | 2.0648e-12 (rank : 8) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q7Z6J4, Q5T8I1, Q6P6A8, Q6ZNL5, Q8IZ32, Q8N868, Q9H7M2 | Gene names | FGD2, ZFYVE4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 2 (Zinc finger FYVE domain-containing protein 4). | |||||
|
ZFY19_HUMAN
|
||||||
| NC score | 0.488111 (rank : 23) | θ value | 2.21117e-06 (rank : 36) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q96K21, Q86WC2, Q8WU96 | Gene names | ZFYVE19, MPFYVE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 19 (MLL partner containing FYVE domain). | |||||
|
FGD4_HUMAN
|
||||||
| NC score | 0.485848 (rank : 24) | θ value | 4.59992e-12 (rank : 9) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q96M96, Q6ULS2, Q8TCP6 | Gene names | FGD4, FRABP, ZFYVE6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 4 (Actin filament- binding protein frabin) (FGD1-related F-actin-binding protein) (Zinc finger FYVE domain-containing protein 6). | |||||
|
ZFY19_MOUSE
|
||||||
| NC score | 0.483226 (rank : 25) | θ value | 1.87187e-05 (rank : 44) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9DAZ9, Q8VCV7 | Gene names | Zfyve19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 19. | |||||
|
RBNS5_HUMAN
|
||||||
| NC score | 0.482745 (rank : 26) | θ value | 4.92598e-06 (rank : 38) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9H1K0, Q3KP30, Q59EY8, Q8NAQ1 | Gene names | ZFYVE20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rabenosyn-5 (FYVE finger-containing Rab5 effector protein rabenosyn-5) (Zinc finger FYVE domain-containing protein 20) (110 kDa protein). | |||||
|
FGD4_MOUSE
|
||||||
| NC score | 0.482009 (rank : 27) | θ value | 1.33837e-11 (rank : 14) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q91ZT5, Q3UEB6, Q8BW60, Q8BZI7, Q91ZT3, Q91ZT4 | Gene names | Fgd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 4 (Actin filament- binding protein frabin) (FGD1-related F-actin-binding protein). | |||||
|
FGD1_MOUSE
|
||||||
| NC score | 0.480542 (rank : 28) | θ value | 7.84624e-12 (rank : 13) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P52734 | Gene names | Fgd1 | |||
|
Domain Architecture |

|
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 1 (Faciogenital dysplasia 1 protein homolog) (Zinc finger FYVE domain-containing protein 3) (Rho/Rac guanine nucleotide exchange factor FGD1) (Rho/Rac GEF). | |||||
|
FGD1_HUMAN
|
||||||
| NC score | 0.480035 (rank : 29) | θ value | 1.74796e-11 (rank : 15) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 367 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P98174, Q8N4D9 | Gene names | FGD1, ZFYVE3 | |||
|
Domain Architecture |

|
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 1 (Faciogenital dysplasia 1 protein) (Zinc finger FYVE domain-containing protein 3) (Rho/Rac guanine nucleotide exchange factor FGD1) (Rho/Rac GEF). | |||||
|
RBNS5_MOUSE
|
||||||
| NC score | 0.478140 (rank : 30) | θ value | 4.92598e-06 (rank : 39) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q80Y56, Q8K0L6, Q9CTW0 | Gene names | Zfyve20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rabenosyn-5 (FYVE finger-containing Rab5 effector protein rabenosyn-5) (Zinc finger FYVE domain-containing protein 20). | |||||
|
FGD3_HUMAN
|
||||||
| NC score | 0.439806 (rank : 31) | θ value | 1.09739e-05 (rank : 43) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q5JSP0, Q7Z7D9, Q8N5G1 | Gene names | FGD3, ZFYVE5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 3 (Zinc finger FYVE domain-containing protein 5). | |||||
|
ZFY27_MOUSE
|
||||||
| NC score | 0.418856 (rank : 32) | θ value | 7.1131e-05 (rank : 46) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q3TXX3, Q8CFP8 | Gene names | Zfyve27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 27. | |||||
|
ZFY27_HUMAN
|
||||||
| NC score | 0.414509 (rank : 33) | θ value | 5.44631e-05 (rank : 45) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5T4F4, Q5T4F1, Q5T4F2, Q5T4F3, Q8N1K0, Q8N6D6, Q8NCA0, Q8NDE4, Q96M08 | Gene names | ZFYVE27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 27. | |||||
|
FGD3_MOUSE
|
||||||
| NC score | 0.401254 (rank : 34) | θ value | 0.00035302 (rank : 50) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O88842, Q8BQ72 | Gene names | Fgd3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 3. | |||||
|
MTMR3_HUMAN
|
||||||
| NC score | 0.398026 (rank : 35) | θ value | 1.47974e-10 (rank : 20) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q13615, Q9NYN5, Q9NYN6, Q9UDX6, Q9UEG3 | Gene names | MTMR3, KIAA0371, ZFYVE10 | |||
|
Domain Architecture |

|
|||||
| Description | Myotubularin-related protein 3 (EC 3.1.3.48) (FYVE domain-containing dual specificity protein phosphatase 1) (FYVE-DSP1) (Zinc finger FYVE domain-containing protein 10). | |||||
|
RUFY1_MOUSE
|
||||||
| NC score | 0.388606 (rank : 36) | θ value | 4.30538e-10 (rank : 29) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 765 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q8BIJ7, Q8BKQ4, Q8BL21, Q9EPM6 | Gene names | Rufy1, Rabip4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 1 (Rab4-interacting protein). | |||||
|
RUFY2_MOUSE
|
||||||
| NC score | 0.386642 (rank : 37) | θ value | 2.52405e-10 (rank : 25) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 820 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q8R4C2, Q69ZH1 | Gene names | Rufy2, Kiaa1537 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 2 (Leucine zipper FYVE-finger protein) (LZ-FYVE). | |||||
|
RUFY1_HUMAN
|
||||||
| NC score | 0.380995 (rank : 38) | θ value | 2.52405e-10 (rank : 24) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 848 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q96T51, Q59FF3, Q71S93, Q9H6I3 | Gene names | RUFY1, RABIP4, ZFYVE12 | |||
|
Domain Architecture |

|
|||||
| Description | RUN and FYVE domain-containing protein 1 (FYVE-finger protein EIP1) (Zinc finger FYVE domain-containing protein 12) (La-binding protein 1) (Rab4-interacting protein). | |||||
|
WDFY3_MOUSE
|
||||||
| NC score | 0.378903 (rank : 39) | θ value | 1.38499e-08 (rank : 32) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q6VNB8, Q8C8H7 | Gene names | Wdfy3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and FYVE domain-containing protein 3 (Beach domain, WD repeat and FYVE domain-containing protein 1) (BWF1). | |||||
|
WDFY3_HUMAN
|
||||||
| NC score | 0.377244 (rank : 40) | θ value | 1.80886e-08 (rank : 34) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8IZQ1, Q4W5K5, Q6P0Q5, Q8N1T2, Q96BS7, Q96N85, Q9Y2J7 | Gene names | WDFY3, KIAA0993 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and FYVE domain-containing protein 3 (Autophagy-linked FYVE protein) (Alfy). | |||||
|
WDFY1_HUMAN
|
||||||
| NC score | 0.367154 (rank : 41) | θ value | 0.000270298 (rank : 49) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8IWB7, Q9H9D5, Q9P2B3 | Gene names | WDFY1, KIAA1435, WDF1, ZFYVE17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and FYVE domain-containing protein 1 (WD40- and FYVE domain- containing protein 1) (Phosphoinositide-binding protein 1) (FENS-1) (Zinc finger FYVE domain-containing protein 17). | |||||
|
WDFY2_MOUSE
|
||||||
| NC score | 0.360060 (rank : 42) | θ value | 0.000121331 (rank : 48) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8BUB4 | Gene names | Wdfy2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and FYVE domain-containing protein 2 (WD40- and FYVE domain- containing protein 2). | |||||
|
WDFY2_HUMAN
|
||||||
| NC score | 0.357331 (rank : 43) | θ value | 0.000121331 (rank : 47) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q96P53, Q96CS1 | Gene names | WDFY2, WDF2, ZFYVE22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and FYVE domain-containing protein 2 (WD40- and FYVE domain- containing protein 2) (Zinc finger FYVE domain-containing protein 22). | |||||
|
RUFY2_HUMAN
|
||||||
| NC score | 0.308666 (rank : 44) | θ value | 1.47974e-10 (rank : 21) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1208 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q8WXA3, Q96P51, Q9P1Z1 | Gene names | RUFY2, KIAA1537, RABIP4R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 2 (Rab4-interacting protein related). | |||||
|
FYCO1_HUMAN
|
||||||
| NC score | 0.267908 (rank : 45) | θ value | 1.80886e-08 (rank : 33) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1149 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9BQS8, Q3MJE6, Q86T41, Q86TB1, Q8TEF9, Q96IV5, Q9H8P9 | Gene names | FYCO1, ZFYVE7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1 (Zinc finger FYVE domain-containing protein 7). | |||||
|
ANFY1_MOUSE
|
||||||
| NC score | 0.252957 (rank : 46) | θ value | 3.77169e-06 (rank : 37) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 480 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q810B6, O54807, Q80TG6, Q80UH8 | Gene names | Ankfy1, Ankhzn, Kiaa1255 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and FYVE domain-containing protein 1 (Ankyrin repeats hooked to a zinc finger motif). | |||||
|
ANFY1_HUMAN
|
||||||
| NC score | 0.248644 (rank : 47) | θ value | 1.09739e-05 (rank : 42) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 480 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9P2R3, Q9ULG5 | Gene names | ANKFY1, ANKHZN, KIAA1255 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and FYVE domain-containing protein 1 (Ankyrin repeats hooked to a zinc finger motif). | |||||
|
FYCO1_MOUSE
|
||||||
| NC score | 0.246498 (rank : 48) | θ value | 8.40245e-06 (rank : 41) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8VDC1, Q3V385, Q7TMT0, Q8BJN2 | Gene names | Fyco1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1. | |||||
|
EEA1_MOUSE
|
||||||
| NC score | 0.234057 (rank : 49) | θ value | 2.52405e-10 (rank : 23) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1637 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q8BL66, Q6DIC2 | Gene names | Eea1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Early endosome antigen 1. | |||||
|
EEA1_HUMAN
|
||||||
| NC score | 0.216626 (rank : 50) | θ value | 2.52405e-10 (rank : 22) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1796 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q15075, Q14221 | Gene names | EEA1, ZFYVE2 | |||
|
Domain Architecture |

|
|||||
| Description | Early endosome antigen 1 (Endosome-associated protein p162) (Zinc finger FYVE domain-containing protein 2). | |||||
|
RFFL_MOUSE
|
||||||
| NC score | 0.184984 (rank : 51) | θ value | 0.0193708 (rank : 53) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q6ZQM0, Q5SVC2, Q5SVC4, Q9D543, Q9D9B1 | Gene names | Rffl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rififylin (RING finger and FYVE-like domain-containing protein 1) (Fring). | |||||
|
RFFL_HUMAN
|
||||||
| NC score | 0.172148 (rank : 52) | θ value | 0.0193708 (rank : 52) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8WZ73, Q8NHW0, Q8TBY7, Q96BE6 | Gene names | RFFL, RNF189, RNF34L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rififylin (RING finger and FYVE-like domain-containing protein 1) (FYVE-RING finger protein Sakura) (Fring) (Caspases-8 and -10- associated RING finger protein 2) (CARP-2) (Caspase regulator CARP2) (RING finger protein 189) (RING finger protein 34-like). | |||||
|
RNF34_MOUSE
|
||||||
| NC score | 0.170749 (rank : 53) | θ value | 1.06291 (rank : 75) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q99KR6, Q3UV45 | Gene names | Rnf34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 34 (RING finger protein RIFF) (Phafin-1). | |||||
|
RNF34_HUMAN
|
||||||
| NC score | 0.169860 (rank : 54) | θ value | 0.47712 (rank : 64) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q969K3, Q8NG47, Q9H6W8 | Gene names | RNF34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 34 (RING finger protein RIFF) (FYVE-RING finger protein Momo) (Human RING finger homologous to inhibitor of apoptosis protein) (hRFI) (Caspases-8 and -10-associated RING finger protein 1) (CARP-1) (Caspase regulator CARP1). | |||||
|
FARP2_HUMAN
|
||||||
| NC score | 0.152465 (rank : 55) | θ value | θ > 10 (rank : 164) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O94887, Q53QM5, Q8WU27, Q9UFE7 | Gene names | FARP2, KIAA0793 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 2 (FERM domain including RhoGEF) (FIR). | |||||
|
FARP2_MOUSE
|
||||||
| NC score | 0.151102 (rank : 56) | θ value | θ > 10 (rank : 165) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q91VS8, Q3TAP2, Q69ZZ0 | Gene names | Farp2, Kiaa0793 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 2 (FERM domain including RhoGEF) (FIR). | |||||
|
PREX1_HUMAN
|
||||||
| NC score | 0.148164 (rank : 57) | θ value | θ > 10 (rank : 183) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8TCU6, Q5JS95, Q5JS96, Q69YL2, Q7Z2L9, Q9BQH0, Q9BX55, Q9H4Q6, Q9P2D2, Q9UGQ4 | Gene names | PREX1, KIAA1415 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3,4,5-trisphosphate-dependent Rac exchanger 1 protein (P-Rex1 protein). | |||||
|
FARP1_HUMAN
|
||||||
| NC score | 0.147561 (rank : 58) | θ value | θ > 10 (rank : 163) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y4F1, Q6IQ29 | Gene names | FARP1, CDEP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 1 (Chondrocyte- derived ezrin-like protein). | |||||
|
ECT2_MOUSE
|
||||||
| NC score | 0.141727 (rank : 59) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q07139 | Gene names | Ect2 | |||
|
Domain Architecture |

|
|||||
| Description | ECT2 protein (Epithelial cell-transforming sequence 2 oncogene). | |||||
|
ECT2_HUMAN
|
||||||
| NC score | 0.138210 (rank : 60) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9H8V3, Q9NSV8, Q9NVW9 | Gene names | ECT2 | |||
|
Domain Architecture |

|
|||||
| Description | ECT2 protein (Epithelial cell-transforming sequence 2 oncogene). | |||||
|
ARHG5_HUMAN
|
||||||
| NC score | 0.123508 (rank : 61) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q12774 | Gene names | ARHGEF5, TIM | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 5 (Guanine nucleotide regulatory protein TIM) (Oncogene TIM) (p60 TIM) (Transforming immortalized mammary oncogene). | |||||
|
RUFY3_MOUSE
|
||||||
| NC score | 0.123456 (rank : 62) | θ value | θ > 10 (rank : 186) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 579 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9D394, Q3U348, Q3UF96, Q6PE64, Q8VD10 | Gene names | Rufy3, D5Bwg0860e, Ripx | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein RUFY3 (Rap2-interacting protein x) (RIPx). | |||||
|
RUFY3_HUMAN
|
||||||
| NC score | 0.122507 (rank : 63) | θ value | θ > 10 (rank : 185) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 594 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q7L099, O94948, Q9UI00 | Gene names | RUFY3, KIAA0871 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein RUFY3 (Rap2-interacting protein x) (RIPx). | |||||
|
PKHG1_HUMAN
|
||||||
| NC score | 0.121154 (rank : 64) | θ value | θ > 10 (rank : 181) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9ULL1, Q5T1F2 | Gene names | PLEKHG1, KIAA1209 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family G member 1. | |||||
|
ARHGA_MOUSE
|
||||||
| NC score | 0.116831 (rank : 65) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8C033, Q5DU38, Q80VH8, Q8BW76, Q922S7 | Gene names | Arhgef10, Kiaa0294 | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 10. | |||||
|
ARHGA_HUMAN
|
||||||
| NC score | 0.116253 (rank : 66) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O15013, O14665, Q68D55, Q8IWD9, Q8IY77 | Gene names | ARHGEF10, KIAA0294 | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 10. | |||||
|
HAPIP_HUMAN
|
||||||
| NC score | 0.112082 (rank : 67) | θ value | θ > 10 (rank : 168) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O60229 | Gene names | HAPIP, DUO | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-associated protein-interacting protein (Protein Duo). | |||||
|
NGEF_HUMAN
|
||||||
| NC score | 0.110213 (rank : 68) | θ value | θ > 10 (rank : 177) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8N5V2, Q53QQ4, Q53ST7, Q6GMQ5, Q9NQD6 | Gene names | NGEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ephexin-1 (Eph-interacting exchange protein) (Neuronal guanine nucleotide exchange factor). | |||||
|
ARHGF_HUMAN
|
||||||
| NC score | 0.110102 (rank : 69) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O94989, Q8N449, Q9H8B4 | Gene names | ARHGEF15, KIAA0915 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 15 (Vsm-RhoGEF). | |||||
|
NGEF_MOUSE
|
||||||
| NC score | 0.108068 (rank : 70) | θ value | θ > 10 (rank : 178) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8CHT1, Q8R204, Q923H2, Q9JHT9 | Gene names | Ngef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ephexin-1 (Eph-interacting exchange protein) (Neuronal guanine nucleotide exchange factor). | |||||
|
ARHGG_HUMAN
|
||||||
| NC score | 0.106960 (rank : 71) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5VV41, Q86TF0, Q99434 | Gene names | ARHGEF16, NBR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 16. | |||||
|
ARHG2_MOUSE
|
||||||
| NC score | 0.100305 (rank : 72) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q60875, O09115 | Gene names | Arhgef2, Lbcl1, Lfc | |||
|
Domain Architecture |

|
|||||
| Description | Rho/Rac guanine nucleotide exchange factor 2 (Lymphoid blast crisis- like 1) (LBC'S first cousin) (Oncogene LFC) (RHOBIN). | |||||
|
ARHGG_MOUSE
|
||||||
| NC score | 0.099051 (rank : 73) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q3U5C8, Q501M8, Q8VCE8 | Gene names | Arhgef16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 16. | |||||
|
MCF2L_MOUSE
|
||||||
| NC score | 0.098156 (rank : 74) | θ value | θ > 10 (rank : 174) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q64096 | Gene names | Mcf2l, Dbs | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide exchange factor DBS (DBL's big sister) (MCF2- transforming sequence-like protein). | |||||
|
MCF2_HUMAN
|
||||||
| NC score | 0.096812 (rank : 75) | θ value | θ > 10 (rank : 175) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P10911, P14919, Q5JYJ4, Q8IUF3, Q8IUF4, Q9UJB3 | Gene names | MCF2, DBL | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene DBL (Proto-oncogene MCF-2) [Contains: MCF2-transforming protein; DBL-transforming protein]. | |||||
|
PKHG4_HUMAN
|
||||||
| NC score | 0.095072 (rank : 76) | θ value | θ > 10 (rank : 182) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q58EX7, Q4G0J8, Q4H485, Q56A69, Q9H7K4, Q9UFW0 | Gene names | PLEKHG4, PRTPHN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Puratrophin-1 (Pleckstrin homology domain-containing family G member 4) (Purkinje cell atrophy-associated protein 1). | |||||
|
LBC_HUMAN
|
||||||
| NC score | 0.093197 (rank : 77) | θ value | θ > 10 (rank : 173) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q12802 | Gene names | LBC | |||
|
Domain Architecture |

|
|||||
| Description | LBC oncogene (P47) (Lymphoid blast crisis oncogene). | |||||
|
ARHG4_HUMAN
|
||||||
| NC score | 0.092416 (rank : 78) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NR80, Q9HDC6, Q9UPP0 | Gene names | ARHGEF4, KIAA1112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 4 (APC-stimulated guanine nucleotide exchange factor) (Asef). | |||||
|
GNRP_MOUSE
|
||||||
| NC score | 0.092161 (rank : 79) | θ value | θ > 10 (rank : 167) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P27671 | Gene names | Rasgrf1, Cdc25, Grf1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-releasing protein (GNRP) (Ras-specific nucleotide exchange factor CDC25) (CDC25Mm). | |||||
|
GNRP_HUMAN
|
||||||
| NC score | 0.091863 (rank : 80) | θ value | θ > 10 (rank : 166) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q13972, Q16027 | Gene names | RASGRF1, CDC25 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-releasing protein (GNRP) (Ras-specific nucleotide exchange factor CDC25) (Ras-specific guanine nucleotide-releasing factor). | |||||
|
ARHG4_MOUSE
|
||||||
| NC score | 0.091699 (rank : 81) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TNR9, Q80TJ6 | Gene names | Arhgef4, Kiaa1112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 4 (APC-stimulated guanine nucleotide exchange factor) (Asef). | |||||
|
BCR_HUMAN
|
||||||
| NC score | 0.090603 (rank : 82) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P11274, P78501, Q12842 | Gene names | BCR, BCR1 | |||
|
Domain Architecture |

|
|||||
| Description | Breakpoint cluster region protein (EC 2.7.11.1) (NY-REN-26 antigen). | |||||
|
ABR_HUMAN
|
||||||
| NC score | 0.088868 (rank : 83) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q12979, Q13693, Q13694 | Gene names | ABR | |||
|
Domain Architecture |

|
|||||
| Description | Active breakpoint cluster region-related protein. | |||||
|
ARHG1_HUMAN
|
||||||
| NC score | 0.088610 (rank : 84) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q92888, O00513, Q8N4J4, Q96BF4, Q96F17, Q9BSB1 | Gene names | ARHGEF1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 1 (p115-RhoGEF) (p115RhoGEF) (115 kDa guanine nucleotide exchange factor) (Sub1.5). | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.088359 (rank : 85) | θ value | 0.00298849 (rank : 51) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
ARHG1_MOUSE
|
||||||
| NC score | 0.087898 (rank : 86) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61210, O89074, Q80YE8, Q80YE9, Q91VL3 | Gene names | Arhgef1, Lbcl2, Lsc | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 1 (Lymphoid blast crisis-like 2) (Lbc's second cousin). | |||||
|
ARHG9_HUMAN
|
||||||
| NC score | 0.085659 (rank : 87) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43307, Q5JSL6 | Gene names | ARHGEF9, KIAA0424 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 9 (Rac/Cdc42 guanine nucleotide exchange factor 9) (Collybistin) (PEM-2 homolog). | |||||
|
ARHG9_MOUSE
|
||||||
| NC score | 0.084417 (rank : 88) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q3UTH8, Q3TQ60, Q80U06, Q8CAF9 | Gene names | Arhgef9, Kiaa0424 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 9 (Rac/Cdc42 guanine nucleotide exchange factor 9) (Collybistin). | |||||
|
CJ064_HUMAN
|
||||||
| NC score | 0.080233 (rank : 89) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6ZS81, Q8N4A3 | Gene names | C10orf64 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C10orf64. | |||||
|
ARHG2_HUMAN
|
||||||
| NC score | 0.073184 (rank : 90) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92974, O75142, Q15079 | Gene names | ARHGEF2, KIAA0651, LFP40 | |||
|
Domain Architecture |

|
|||||
| Description | Rho/Rac guanine nucleotide exchange factor 2 (GEF-H1 protein) (Proliferating cell nucleolar antigen p40). | |||||
|
ITSN2_MOUSE
|
||||||
| NC score | 0.072931 (rank : 91) | θ value | θ > 10 (rank : 172) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1371 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Z0R6, Q9Z0R5 | Gene names | Itsn2, Ese2, Sh3d1B | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-2 (SH3 domain-containing protein 1B) (EH and SH3 domains protein 2) (EH domain and SH3 domain regulator of endocytosis 2). | |||||
|
ITSN1_HUMAN
|
||||||
| NC score | 0.072704 (rank : 92) | θ value | θ > 10 (rank : 169) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1757 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q15811, O95216, Q9UET5, Q9UK60, Q9UNK1, Q9UNK2, Q9UQ92 | Gene names | ITSN1, ITSN, SH3D1A | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (SH3 domain-containing protein 1A) (SH3P17). | |||||
|
ITSN2_HUMAN
|
||||||
| NC score | 0.072517 (rank : 93) | θ value | θ > 10 (rank : 171) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1173 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9NZM3, O95062, Q15812, Q9HAK4, Q9NXE6, Q9NYG0, Q9NZM2, Q9ULG4 | Gene names | ITSN2, KIAA1256, SH3D1B, SWAP | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-2 (SH3 domain-containing protein 1B) (SH3P18) (SH3P18-like WASP-associated protein). | |||||
|
MYRIP_MOUSE
|
||||||
| NC score | 0.072026 (rank : 94) | θ value | 0.0563607 (rank : 56) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8K3I4, Q8CFC0, Q8K4H5 | Gene names | Myrip, Slac2c | |||
|
Domain Architecture |

|
|||||
| Description | Rab effector MyRIP (Myosin-VIIa- and Rab-interacting protein) (Exophilin-8) (Slp homolog lacking C2 domains c). | |||||
|
ITSN1_MOUSE
|
||||||
| NC score | 0.071598 (rank : 95) | θ value | θ > 10 (rank : 170) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1669 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9Z0R4, Q9R143 | Gene names | Itsn1, Ese1, Itsn | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (EH and SH3 domains protein 1). | |||||
|
TIAM1_HUMAN
|
||||||
| NC score | 0.071264 (rank : 96) | θ value | θ > 10 (rank : 190) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13009 | Gene names | TIAM1 | |||
|
Domain Architecture |

|
|||||
| Description | T-lymphoma invasion and metastasis-inducing protein 1 (TIAM-1 protein). | |||||
|
ARHGB_HUMAN
|
||||||
| NC score | 0.070412 (rank : 97) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 375 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O15085 | Gene names | ARHGEF11, KIAA0380 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 11 (PDZ-RhoGEF). | |||||
|
ARHG7_HUMAN
|
||||||
| NC score | 0.069798 (rank : 98) | θ value | θ > 10 (rank : 146) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q14155, Q6P9G3, Q6PII2, Q86W63, Q8N3M1 | Gene names | ARHGEF7, COOL1, KIAA0142, P85SPR, PAK3BP, PIXB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 7 (PAK-interacting exchange factor beta) (Beta-Pix) (COOL-1) (p85). | |||||
|
ARHGC_HUMAN
|
||||||
| NC score | 0.069573 (rank : 99) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NZN5, O15086 | Gene names | ARHGEF12, KIAA0382, LARG | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 12 (Leukemia-associated RhoGEF). | |||||
|
TIAM1_MOUSE
|
||||||
| NC score | 0.069312 (rank : 100) | θ value | θ > 10 (rank : 191) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q60610 | Gene names | Tiam1, Tiam-1 | |||
|
Domain Architecture |

|
|||||
| Description | T-lymphoma invasion and metastasis-inducing protein 1 (TIAM-1 protein). | |||||
|
ARHGC_MOUSE
|
||||||
| NC score | 0.068839 (rank : 101) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8R4H2, Q80U18 | Gene names | Arhgef12, Kiaa0382, Larg | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 12 (Leukemia-associated RhoGEF). | |||||
|
DSPP_MOUSE
|
||||||
| NC score | 0.067899 (rank : 102) | θ value | 0.62314 (rank : 65) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
PKHA6_HUMAN
|
||||||
| NC score | 0.065711 (rank : 103) | θ value | θ > 10 (rank : 179) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Y2H5 | Gene names | PLEKHA6, KIAA0969, PEPP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
PKHA6_MOUSE
|
||||||
| NC score | 0.063182 (rank : 104) | θ value | θ > 10 (rank : 180) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q7TQG1, Q8K0J5 | Gene names | Plekha6, Pepp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
VAV_HUMAN
|
||||||
| NC score | 0.061431 (rank : 105) | θ value | θ > 10 (rank : 196) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P15498, Q15860 | Gene names | VAV1, VAV | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene vav. | |||||
|
ARHG3_MOUSE
|
||||||
| NC score | 0.059371 (rank : 106) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91X46, Q8CDM0, Q91VY4, Q99K14, Q9DC31 | Gene names | Arhgef3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 3. | |||||
|
VAV_MOUSE
|
||||||
| NC score | 0.058961 (rank : 107) | θ value | θ > 10 (rank : 197) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 430 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P27870, Q8BTV7 | Gene names | Vav1, Vav | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene vav (p95vav). | |||||
|
ALS2_MOUSE
|
||||||
| NC score | 0.058957 (rank : 108) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q920R0, Q8JZR1, Q9CXJ3 | Gene names | Als2 | |||
|
Domain Architecture |

|
|||||
| Description | Alsin (Amyotrophic lateral sclerosis protein 2 homolog). | |||||
|
ARHG7_MOUSE
|
||||||
| NC score | 0.058494 (rank : 109) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9ES28, O08757, Q9ES27 | Gene names | Arhgef7, Pak3bp | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 7 (PAK-interacting exchange factor beta) (Beta-Pix) (p85SPR). | |||||
|
ARHG3_HUMAN
|
||||||
| NC score | 0.057863 (rank : 110) | θ value | θ > 10 (rank : 141) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NR81, Q6NUN3, Q7Z4U2, Q7Z5T2, Q9H7T4 | Gene names | ARHGEF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 3 (Exchange factor found in platelets and leukemic and neuronal tissues) (XPLN). | |||||
|
ARHG8_HUMAN
|
||||||
| NC score | 0.057837 (rank : 111) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7Z628, Q12773, Q96D82, Q99903, Q9UEN6 | Gene names | NET1, ARHGEF8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuroepithelial cell-transforming gene 1 protein (p65 Net1 proto- oncogene) (Rho guanine nucleotide exchange factor 8). | |||||
|
SWP70_HUMAN
|
||||||
| NC score | 0.057312 (rank : 112) | θ value | θ > 10 (rank : 188) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 973 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UH65, O75135, Q7LCY6, Q9P061, Q9P0Z8 | Gene names | SWAP70, KIAA0640 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Switch-associated protein 70 (SWAP-70). | |||||
|
VAV2_MOUSE
|
||||||
| NC score | 0.056409 (rank : 113) | θ value | θ > 10 (rank : 193) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 456 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q60992 | Gene names | Vav2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
VAV3_HUMAN
|
||||||
| NC score | 0.056399 (rank : 114) | θ value | θ > 10 (rank : 194) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 398 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UKW4, O95230, Q9Y5X8 | Gene names | VAV3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-3. | |||||
|
VAV3_MOUSE
|
||||||
| NC score | 0.055649 (rank : 115) | θ value | θ > 10 (rank : 195) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 404 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R0C8, Q7TS85 | Gene names | Vav3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-3. | |||||
|
SWP70_MOUSE
|
||||||
| NC score | 0.054337 (rank : 116) | θ value | θ > 10 (rank : 189) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1097 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q6A028, O88443, Q3TQR6, Q3UCA3, Q6P1D0 | Gene names | Swap70, Kiaa0640 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Switch-associated protein 70 (SWAP-70). | |||||
|
TRAIP_MOUSE
|
||||||
| NC score | 0.053060 (rank : 117) | θ value | 0.813845 (rank : 71) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 745 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8VIG6, O08854, Q922M8, Q9CPP4 | Gene names | Traip, Trip | |||
|
Domain Architecture |

|
|||||
| Description | TRAF-interacting protein. | |||||
|
SOS2_HUMAN
|
||||||
| NC score | 0.052636 (rank : 118) | θ value | θ > 10 (rank : 187) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q07890, Q15503 | Gene names | SOS2 | |||
|
Domain Architecture |

|
|||||
| Description | Son of sevenless homolog 2 (SOS-2). | |||||
|
VAV2_HUMAN
|
||||||
| NC score | 0.052069 (rank : 119) | θ value | θ > 10 (rank : 192) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P52735 | Gene names | VAV2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
CJ118_MOUSE
|
||||||
| NC score | 0.051927 (rank : 120) | θ value | 0.0563607 (rank : 55) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1132 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8C9S4, Q6P3D6, Q8CJC0 | Gene names | Otg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf118 homolog (Oocyte-testis gene 1 protein). | |||||
|
CJ118_HUMAN
|
||||||
| NC score | 0.051043 (rank : 121) | θ value | 0.0431538 (rank : 54) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q7Z3E2, Q2M2V6, Q6NS91, Q7RTP1, Q8N117, Q8N3G3, Q8N6C2, Q9NWA3 | Gene names | C10orf118 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf118 (CTCL tumor antigen HD-CL-01/L14-2). | |||||
|
NBEAL_HUMAN
|
||||||
| NC score | 0.051030 (rank : 122) | θ value | θ > 10 (rank : 176) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6ZS30, Q6Y876, Q6ZQY5, Q96Q30 | Gene names | NBEAL1, ALS2CR17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurobeachin-like 1 (Amyotrophic lateral sclerosis 2 chromosomal region candidate gene 17 protein). | |||||
|
RGNEF_MOUSE
|
||||||
| NC score | 0.050088 (rank : 123) | θ value | θ > 10 (rank : 184) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P97433 | Gene names | Rgnef, Rhoip2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-guanine nucleotide exchange factor (Rho-interacting protein 2) (RhoGEF) (RIP2). | |||||
|
RP1_HUMAN
|
||||||
| NC score | 0.043276 (rank : 124) | θ value | 0.0736092 (rank : 57) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P56715 | Gene names | RP1, ORP1 | |||
|
Domain Architecture |

|
|||||
| Description | Oxygen-regulated protein 1 (Retinitis pigmentosa RP1 protein) (Retinitis pigmentosa 1 protein). | |||||
|
PAF49_HUMAN
|
||||||
| NC score | 0.041050 (rank : 125) | θ value | 0.279714 (rank : 59) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O15446, Q32N11, Q7Z5U2, Q9UPF6 | Gene names | CD3EAP, ASE1, CAST, PAF49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase I-associated factor PAF49 (Anti-sense to ERCC-1 protein) (ASE-1) (CD3-epsilon-associated protein) (CD3E-associated protein) (CAST). | |||||
|
BCAS1_HUMAN
|
||||||
| NC score | 0.040248 (rank : 126) | θ value | 0.47712 (rank : 62) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75363, Q68CZ3 | Gene names | BCAS1, AIBC1, NABC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Breast carcinoma amplified sequence 1 (Novel amplified in breast cancer 1) (Amplified and overexpressed in breast cancer). | |||||
|
MAP9_HUMAN
|
||||||
| NC score | 0.039639 (rank : 127) | θ value | 2.36792 (rank : 82) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q49MG5, Q4W5I7, Q68DU1, Q9H781, Q9H7B6 | Gene names | MAP9, ASAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 9 (Aster-associated protein). | |||||
|
TMF1_HUMAN
|
||||||
| NC score | 0.037813 (rank : 128) | θ value | 5.27518 (rank : 102) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 987 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P82094 | Gene names | TMF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA element modulatory factor (TMF). | |||||
|
ZCHC6_HUMAN
|
||||||
| NC score | 0.036971 (rank : 129) | θ value | 0.0961366 (rank : 58) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5VYS8, Q5H9T0, Q5VYS5, Q5VYS7, Q658Z9, Q659A2, Q6MZJ3, Q8N5F0, Q96N57, Q96NE8, Q9C0F2, Q9H8M6 | Gene names | ZCCHC6, KIAA1711 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 6. | |||||
|
RRMJ3_HUMAN
|
||||||
| NC score | 0.035750 (rank : 130) | θ value | 0.813845 (rank : 70) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IY81, Q8N3A3, Q8WXX1, Q9BWM4, Q9NXT6 | Gene names | FTSJ3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative rRNA methyltransferase 3 (EC 2.1.1.-) (rRNA (uridine-2'-O-)- methyltransferase 3). | |||||
|
RANB9_HUMAN
|
||||||
| NC score | 0.033141 (rank : 131) | θ value | 0.47712 (rank : 63) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96S59, O94764, Q6P3T7, Q7LBR2, Q7Z7F9 | Gene names | RANBP9, RANBPM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 9 (RanBP9) (RanBP7) (Ran-binding protein M) (RanBPM) (BPM90) (BPM-L). | |||||
|
DMP1_HUMAN
|
||||||
| NC score | 0.031379 (rank : 132) | θ value | 2.36792 (rank : 81) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
DAB2_HUMAN
|
||||||
| NC score | 0.028930 (rank : 133) | θ value | 1.38821 (rank : 77) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P98082, Q13598, Q9BTY0, Q9UK04 | Gene names | DAB2, DOC2 | |||
|
Domain Architecture |

|
|||||
| Description | Disabled homolog 2 (Differentially expressed protein 2) (DOC-2). | |||||
|
ANR21_HUMAN
|
||||||
| NC score | 0.028431 (rank : 134) | θ value | 1.06291 (rank : 72) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q86YR6 | Gene names | ANKRD21, POTE | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat domain-containing protein 21 (Protein POTE) (Prostate, ovary, testis-expressed protein). | |||||
|
SPRL1_MOUSE
|
||||||
| NC score | 0.028409 (rank : 135) | θ value | 0.62314 (rank : 66) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P70663, P97810, Q99L82 | Gene names | Sparcl1, Ecm2, Sc1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (Matrix glycoprotein Sc1) (Extracellular matrix protein 2). | |||||
|
POT15_HUMAN
|
||||||
| NC score | 0.027987 (rank : 136) | θ value | 2.36792 (rank : 83) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6S5H4, Q6NXN7, Q6S5H7 | Gene names | POTE15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prostate, ovary, testis-expressed protein on chromosome 15. | |||||
|
AIM1_HUMAN
|
||||||
| NC score | 0.027564 (rank : 137) | θ value | 3.0926 (rank : 84) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y4K1, O00296, Q5VWJ2 | Gene names | AIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Absent in melanoma 1 protein. | |||||
|
K0423_HUMAN
|
||||||
| NC score | 0.027013 (rank : 138) | θ value | 1.81305 (rank : 78) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y4F4, Q68D66, Q6PG27 | Gene names | KIAA0423 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0423. | |||||
|
E2F4_HUMAN
|
||||||
| NC score | 0.026716 (rank : 139) | θ value | 0.365318 (rank : 60) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q16254, Q12991, Q15328 | Gene names | E2F4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E2F4 (E2F-4). | |||||
|
CYLC1_HUMAN
|
||||||
| NC score | 0.025662 (rank : 140) | θ value | 1.06291 (rank : 73) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
MBD6_HUMAN
|
||||||
| NC score | 0.025464 (rank : 141) | θ value | 1.81305 (rank : 79) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96DN6, Q8N3M0, Q8NA81, Q96Q00 | Gene names | MBD6, KIAA1887 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 6 (Methyl-CpG-binding protein MBD6). | |||||
|
MYST4_HUMAN
|
||||||
| NC score | 0.025257 (rank : 142) | θ value | 1.06291 (rank : 74) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
CA103_MOUSE
|
||||||
| NC score | 0.022709 (rank : 143) | θ value | 3.0926 (rank : 85) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CDD9, Q8C893 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C1orf103 homolog. | |||||
|
SAMN1_MOUSE
|
||||||
| NC score | 0.022419 (rank : 144) | θ value | 1.06291 (rank : 76) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P57725, Q8C6S2 | Gene names | Samsn1 | |||
|
Domain Architecture |

|
|||||
| Description | SAM domain-containing protein SAMSN-1 (SAM domain, SH3 domain and nuclear localization signals protein 1). | |||||
|
CCD82_HUMAN
|
||||||
| NC score | 0.020274 (rank : 145) | θ value | 6.88961 (rank : 104) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N4S0, Q9H2Q5, Q9H5E3 | Gene names | CCDC82 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 82. | |||||
|
WDTC1_HUMAN
|
||||||
| NC score | 0.019923 (rank : 146) | θ value | 6.88961 (rank : 117) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 299 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8N5D0, Q9NV87, Q9UPW4 | Gene names | WDTC1, KIAA1037 | |||
|
Domain Architecture |

|
|||||
| Description | WD and tetratricopeptide repeats protein 1. | |||||
|
WDTC1_MOUSE
|
||||||
| NC score | 0.019787 (rank : 147) | θ value | 6.88961 (rank : 118) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80ZK9 | Gene names | Wdtc1 | |||
|
Domain Architecture |

|
|||||
| Description | WD and tetratricopeptide repeats protein 1. | |||||
|
ZHX2_MOUSE
|
||||||
| NC score | 0.019776 (rank : 148) | θ value | 0.62314 (rank : 67) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8C0C0 | Gene names | Zhx2, Afr1, Raf | |||
|
Domain Architecture |

|
|||||
| Description | Zinc fingers and homeoboxes protein 2 (Zinc finger and homeodomain protein 2) (Alpha-fetoprotein regulator 1) (AFP regulator 1) (Regulator of AFP). | |||||
|
DAB2_MOUSE
|
||||||
| NC score | 0.019491 (rank : 149) | θ value | 6.88961 (rank : 107) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P98078, Q3U3K1, Q91W56, Q923E1 | Gene names | Dab2, Doc2 | |||
|
Domain Architecture |

|
|||||
| Description | Disabled homolog 2 (DOC-2) (Mitogen-responsive phosphoprotein). | |||||
|
AFF4_HUMAN
|
||||||
| NC score | 0.019473 (rank : 150) | θ value | 0.813845 (rank : 68) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
NOL4_MOUSE
|
||||||
| NC score | 0.018673 (rank : 151) | θ value | 3.0926 (rank : 88) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P60954 | Gene names | Nol4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar protein 4. | |||||
|
HSF1_MOUSE
|
||||||
| NC score | 0.018641 (rank : 152) | θ value | 3.0926 (rank : 87) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P38532, O70462 | Gene names | Hsf1 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock factor protein 1 (HSF 1) (Heat shock transcription factor 1) (HSTF 1). | |||||
|
CCDC4_HUMAN
|
||||||
| NC score | 0.018636 (rank : 153) | θ value | 6.88961 (rank : 105) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6ZU67, Q58A26, Q58A27 | Gene names | CCDC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 4. | |||||
|
SET1A_HUMAN
|
||||||
| NC score | 0.018147 (rank : 154) | θ value | 3.0926 (rank : 89) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
BTBDC_HUMAN
|
||||||
| NC score | 0.016426 (rank : 155) | θ value | 6.88961 (rank : 103) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8IY92, Q96JP1 | Gene names | BTBD12, KIAA1784 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein 12. | |||||
|
EVI2B_MOUSE
|
||||||
| NC score | 0.016198 (rank : 156) | θ value | 5.27518 (rank : 98) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8VD58, Q3U1A3 | Gene names | Evi2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EVI2B protein precursor (Ecotropic viral integration site 2B protein). | |||||
|
BAT2_MOUSE
|
||||||
| NC score | 0.016116 (rank : 157) | θ value | 5.27518 (rank : 97) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
SOX6_MOUSE
|
||||||
| NC score | 0.015945 (rank : 158) | θ value | 1.81305 (rank : 80) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P40645, Q62250, Q9QWS5 | Gene names | Sox6, Sox-6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-6 (SOX-LZ). | |||||
|
IDUA_HUMAN
|
||||||
| NC score | 0.015036 (rank : 159) | θ value | 8.99809 (rank : 125) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P35475 | Gene names | IDUA | |||
|
Domain Architecture |
|
|||||
| Description | Alpha-L-iduronidase precursor (EC 3.2.1.76). | |||||
|
IDUA_MOUSE
|
||||||
| NC score | 0.014956 (rank : 160) | θ value | 8.99809 (rank : 126) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P48441 | Gene names | Idua | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-L-iduronidase precursor (EC 3.2.1.76). | |||||
|
MOCS3_HUMAN
|
||||||
| NC score | 0.014898 (rank : 161) | θ value | 6.88961 (rank : 110) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95396 | Gene names | MOCS3 | |||
|
Domain Architecture |

|
|||||
| Description | Molybdenum cofactor synthesis protein 3 (Molybdopterin synthase sulfurylase) (MPT synthase sulfurylase). | |||||
|
RIMS2_MOUSE
|
||||||
| NC score | 0.013641 (rank : 162) | θ value | 8.99809 (rank : 130) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9EQZ7, Q8C433, Q8CCK2 | Gene names | Rims2, Rab3ip2, Rim2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2) (Rab3-interacting protein 2). | |||||
|
ARID2_HUMAN
|
||||||
| NC score | 0.013421 (rank : 163) | θ value | 8.99809 (rank : 119) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q68CP9, Q5EB51, Q645I3, Q6ZRY5, Q7Z3I5, Q86T28, Q96SJ6, Q9HCL5 | Gene names | ARID2, KIAA1557 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-rich interactive domain-containing protein 2 (ARID domain- containing protein 2) (BRG1-associated factor 200) (BAF200). | |||||
|
UBP10_HUMAN
|
||||||
| NC score | 0.013059 (rank : 164) | θ value | 3.0926 (rank : 90) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14694, Q9BWG7, Q9NSL7 | Gene names | USP10, KIAA0190 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 10 (EC 3.1.2.15) (Ubiquitin thioesterase 10) (Ubiquitin-specific-processing protease 10) (Deubiquitinating enzyme 10). | |||||
|
JHD2C_HUMAN
|
||||||
| NC score | 0.012567 (rank : 165) | θ value | 6.88961 (rank : 109) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15652, Q5SQZ8, Q5SQZ9, Q5SR00, Q7Z3E7, Q8N3U0, Q96KB9, Q9P2G7 | Gene names | JMJD1C, JHDM2C, KIAA1380, TRIP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable JmjC domain-containing histone demethylation protein 2C (EC 1.14.11.-) (Jumonji domain-containing protein 1C) (Thyroid receptor-interacting protein 8) (TRIP-8). | |||||
|
HELC1_HUMAN
|
||||||
| NC score | 0.012024 (rank : 166) | θ value | 5.27518 (rank : 99) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N3C0, O43738, Q9H1I9, Q9NTR0 | Gene names | ASCC3, HELIC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Activating signal cointegrator 1 complex subunit 3 (EC 3.6.1.-) (ASC-1 complex subunit p200) (Trip4 complex subunit p200) (Helicase, ATP binding 1). | |||||
|
STAP2_MOUSE
|
||||||
| NC score | 0.012000 (rank : 167) | θ value | 6.88961 (rank : 114) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R0L1, Q8BWS2 | Gene names | Stap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-transducing adaptor protein 2 (STAP-2). | |||||
|
NCOR2_MOUSE
|
||||||
| NC score | 0.011455 (rank : 168) | θ value | 4.03905 (rank : 92) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
S13A4_HUMAN
|
||||||
| NC score | 0.011436 (rank : 169) | θ value | 4.03905 (rank : 93) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UKG4 | Gene names | SLC13A4, SUT1 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 13 member 4 (Na(+)/sulfate cotransporter SUT-1). | |||||
|
ICAL_HUMAN
|
||||||
| NC score | 0.011237 (rank : 170) | θ value | 6.88961 (rank : 108) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P20810, O95360, Q96D08, Q9H1Z5 | Gene names | CAST | |||
|
Domain Architecture |

|
|||||
| Description | Calpastatin (Calpain inhibitor) (Sperm BS-17 component). | |||||
|
RFX3_MOUSE
|
||||||
| NC score | 0.011217 (rank : 171) | θ value | 6.88961 (rank : 112) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P48381, Q8BLW2, Q8C0R3, Q8VBY6 | Gene names | Rfx3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor RFX3. | |||||
|
RFX3_HUMAN
|
||||||
| NC score | 0.011210 (rank : 172) | θ value | 6.88961 (rank : 111) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P48380, Q5JTL7, Q5JTL8, Q6NW13, Q8WTU4, Q95HL5, Q95HL6 | Gene names | RFX3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor RFX3. | |||||
|
RNF31_MOUSE
|
||||||
| NC score | 0.011030 (rank : 173) | θ value | 8.99809 (rank : 131) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q924T7 | Gene names | Rnf31 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 31. | |||||
|
S28A2_MOUSE
|
||||||
| NC score | 0.010669 (rank : 174) | θ value | 6.88961 (rank : 113) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O88627 | Gene names | Slc28a2, Cnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/nucleoside cotransporter 2 (Na(+)/nucleoside cotransporter 2) (Sodium-coupled nucleoside transporter 2) (Concentrative nucleoside transporter 2) (CNT 2) (Sodium/purine nucleoside cotransporter) (SPNT). | |||||
|
FA53B_HUMAN
|
||||||
| NC score | 0.010540 (rank : 175) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q14153, Q8N5S6 | Gene names | FAM53B, KIAA0140 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM53B. | |||||
|
UTY_MOUSE
|
||||||
| NC score | 0.009794 (rank : 176) | θ value | 6.88961 (rank : 116) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P79457, O97979 | Gene names | Uty | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitously transcribed Y chromosome tetratricopeptide repeat protein (Ubiquitously transcribed TPR protein ON the Y chromosome) (Male- specific histocompatibility antigen H-YDB). | |||||
|
NEDD1_HUMAN
|
||||||
| NC score | 0.009089 (rank : 177) | θ value | 8.99809 (rank : 128) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 294 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NHV4, Q8NA30 | Gene names | NEDD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein NEDD1. | |||||
|
THIC_MOUSE
|
||||||
| NC score | 0.007200 (rank : 178) | θ value | 6.88961 (rank : 115) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CAY6, Q3TJY7, Q62292, Q96EB7, Q99K88, Q9CYV2 | Gene names | Acat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acetyl-CoA acetyltransferase, cytosolic (EC 2.3.1.9) (Cytosolic acetoacetyl-CoA thiolase). | |||||
|
CLH2_HUMAN
|
||||||
| NC score | 0.007175 (rank : 179) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P53675, Q14017, Q15808, Q15809 | Gene names | CLTCL1, CLH22, CLTCL, CLTD | |||
|
Domain Architecture |

|
|||||
| Description | Clathrin heavy chain 2 (CLH-22). | |||||
|
IRF1_MOUSE
|
||||||
| NC score | 0.006363 (rank : 180) | θ value | 8.99809 (rank : 127) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P15314 | Gene names | Irf1 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon regulatory factor 1 (IRF-1). | |||||
|
HTF4_HUMAN
|
||||||
| NC score | 0.005846 (rank : 181) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99081 | Gene names | TCF12, HEB, HTF4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 12 (Transcription factor HTF-4) (E-box-binding protein) (DNA-binding protein HTF4). | |||||
|
CCNB1_HUMAN
|
||||||
| NC score | 0.005071 (rank : 182) | θ value | 8.99809 (rank : 120) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P14635 | Gene names | CCNB1, CCNB | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B1. | |||||
|
EYA4_HUMAN
|
||||||
| NC score | 0.004938 (rank : 183) | θ value | 8.99809 (rank : 122) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95677, O95464, O95679, Q8IW39, Q9NTR7 | Gene names | EYA4 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 4 (EC 3.1.3.48). | |||||
|
ATP4A_MOUSE
|
||||||
| NC score | 0.003532 (rank : 184) | θ value | 5.27518 (rank : 96) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q64436, Q9CV46 | Gene names | Atp4a | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 1 (EC 3.6.3.10) (Proton pump) (Gastric H+/K+ ATPase subunit alpha). | |||||
|
FLRT3_HUMAN
|
||||||
| NC score | 0.003017 (rank : 185) | θ value | 3.0926 (rank : 86) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NZU0, Q96K39, Q96K42, Q96KB1, Q9P259 | Gene names | FLRT3, KIAA1469 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat transmembrane protein FLRT3 precursor (Fibronectin-like domain-containing leucine-rich transmembrane protein 3). | |||||
|
ADA29_HUMAN
|
||||||
| NC score | 0.002647 (rank : 186) | θ value | 5.27518 (rank : 95) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UKF5, Q9UHP1, Q9UKF3, Q9UKF4 | Gene names | ADAM29 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 29 precursor (A disintegrin and metalloproteinase domain 29). | |||||
|
CSMD3_HUMAN
|
||||||
| NC score | 0.002289 (rank : 187) | θ value | 6.88961 (rank : 106) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z407, Q96PZ3 | Gene names | CSMD3, KIAA1894 | |||
|
Domain Architecture |

|
|||||
| Description | CUB and sushi domain-containing protein 3 precursor (CUB and sushi multiple domains protein 3). | |||||
|
KS6A5_HUMAN
|
||||||
| NC score | 0.000949 (rank : 188) | θ value | 0.365318 (rank : 61) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 855 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75582, O95316, Q96AF7 | Gene names | RPS6KA5, MSK1 | |||
|
Domain Architecture |
|
|||||
| Description | Ribosomal protein S6 kinase alpha-5 (EC 2.7.11.1) (Nuclear mitogen- and stress-activated protein kinase 1) (90 kDa ribosomal protein S6 kinase 5) (RSK-like protein kinase) (RSKL). | |||||
|
KS6A5_MOUSE
|
||||||
| NC score | 0.000820 (rank : 189) | θ value | 0.813845 (rank : 69) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 854 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8C050, Q8CI92 | Gene names | Rps6ka5, Msk1 | |||
|
Domain Architecture |
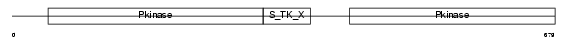
|
|||||
| Description | Ribosomal protein S6 kinase alpha-5 (EC 2.7.11.1) (Nuclear mitogen- and stress-activated protein kinase 1) (90 kDa ribosomal protein S6 kinase 5) (RSK-like protein kinase) (RLSK). | |||||
|
TRI10_HUMAN
|
||||||
| NC score | 0.000731 (rank : 190) | θ value | 8.99809 (rank : 133) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UDY6, Q86Z08, Q96QB6, Q9C023, Q9C024 | Gene names | TRIM10, RFB30, RNF9 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 10 (RING finger protein 9) (B30- RING finger protein). | |||||
|
PTN13_MOUSE
|
||||||
| NC score | 0.000060 (rank : 191) | θ value | 8.99809 (rank : 129) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q64512, Q61494, Q62135, Q64499 | Gene names | Ptpn13, Ptp14 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 13 (EC 3.1.3.48) (Protein tyrosine phosphatase PTP-BL) (Protein-tyrosine phosphatase RIP) (protein tyrosine phosphatase DPZPTP) (PTP36). | |||||
|
PCDBG_HUMAN
|
||||||
| NC score | 0.000040 (rank : 192) | θ value | 5.27518 (rank : 101) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NRJ7, Q8IYD5, Q96SE9, Q9HCF1 | Gene names | PCDHB16, KIAA1621, PCDH3X | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 16 precursor (PCDH-beta16) (Protocadherin 3X). | |||||
|
TITIN_HUMAN
|
||||||
| NC score | -0.001074 (rank : 193) | θ value | 8.99809 (rank : 132) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
ZBT38_HUMAN
|
||||||
| NC score | -0.001820 (rank : 194) | θ value | 3.0926 (rank : 91) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 885 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8NAP3 | Gene names | ZBTB38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and BTB domain-containing protein 38. | |||||
|
MK07_MOUSE
|
||||||
| NC score | -0.001854 (rank : 195) | θ value | 5.27518 (rank : 100) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 1054 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9WVS8 | Gene names | Mapk7, Erk5 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase 7 (EC 2.7.11.24) (Extracellular signal-regulated kinase 5) (ERK-5) (BMK1 kinase). | |||||
|
ZBT24_MOUSE
|
||||||
| NC score | -0.003897 (rank : 196) | θ value | 4.03905 (rank : 94) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 894 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80X44, Q3UIB7, Q7TPC4, Q8CJC9 | Gene names | Zbtb24, Bif1, Bsg1, Znf450 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and BTB domain-containing protein 24 (Zinc finger protein 450) (Bone morphogenetic protein-induced factor 1) (Brain-specific protein 1). | |||||
|
ZNF18_MOUSE
|
||||||
| NC score | -0.004238 (rank : 197) | θ value | 8.99809 (rank : 134) | |||
| Query Neighborhood Hits | 134 | Target Neighborhood Hits | 743 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q810A1, Q9D972 | Gene names | Znf18, Zfp535 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 18 (Zinc finger protein 535). | |||||