Please be patient as the page loads
|
PK3CA_MOUSE
|
||||||
| SwissProt Accessions | P42337 | Gene names | Pik3ca | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (EC 2.7.1.153) (PI3-kinase p110 subunit alpha) (PtdIns-3- kinase p110) (PI3K). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PK3CA_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999669 (rank : 2) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P42336, Q99762 | Gene names | PIK3CA | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (EC 2.7.1.153) (PI3-kinase p110 subunit alpha) (PtdIns-3- kinase p110) (PI3K). | |||||
|
PK3CA_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P42337 | Gene names | Pik3ca | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (EC 2.7.1.153) (PI3-kinase p110 subunit alpha) (PtdIns-3- kinase p110) (PI3K). | |||||
|
PK3CB_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.979415 (rank : 5) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P42338 | Gene names | PIK3CB | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit beta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit beta) (PtdIns-3-kinase p110) (PI3K) (PI3Kbeta). | |||||
|
PK3CB_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.979293 (rank : 6) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BTI9 | Gene names | Pik3cb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit beta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit beta) (PtdIns-3-kinase p110) (PI3K) (PI3Kbeta). | |||||
|
PK3CD_HUMAN
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.981681 (rank : 3) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O00329, O15445 | Gene names | PIK3CD | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit delta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit delta) (PtdIns-3- kinase p110) (PI3K) (p110delta). | |||||
|
PK3CD_MOUSE
|
||||||
| θ value | 0 (rank : 6) | NC score | 0.981320 (rank : 4) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O35904 | Gene names | Pik3cd | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit delta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit delta) (PtdIns-3- kinase p110) (PI3K) (p110delta). | |||||
|
PK3CG_HUMAN
|
||||||
| θ value | 3.27187e-151 (rank : 7) | NC score | 0.966367 (rank : 7) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P48736, Q8IV23, Q9BZC8 | Gene names | PIK3CG | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit gamma isoform (EC 2.7.1.153) (PI3-kinase p110 subunit gamma) (PtdIns-3- kinase subunit p110) (PI3K) (PI3Kgamma) (p120-PI3K). | |||||
|
PK3CG_MOUSE
|
||||||
| θ value | 1.62381e-150 (rank : 8) | NC score | 0.965955 (rank : 8) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9JHG7 | Gene names | Pik3cg, Pi3kg1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit gamma isoform (EC 2.7.1.153) (PI3-kinase p110 subunit gamma) (PtdIns-3- kinase subunit p110) (PI3K) (PI3Kgamma) (p120-PI3K). | |||||
|
P3C2A_MOUSE
|
||||||
| θ value | 6.45994e-107 (rank : 9) | NC score | 0.835124 (rank : 13) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q61194, Q61182 | Gene names | Pik3c2a, Cpk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing alpha polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-alpha) (PtdIns-3-kinase C2 alpha) (PI3K-C2alpha) (Cpk-m) (p170). | |||||
|
P3C2B_HUMAN
|
||||||
| θ value | 1.1019e-106 (rank : 10) | NC score | 0.811608 (rank : 16) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O00750, O95666, Q5SW99 | Gene names | PIK3C2B | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing beta polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-beta) (PtdIns-3-kinase C2 beta) (PI3K-C2beta) (C2-PI3K). | |||||
|
P3C2A_HUMAN
|
||||||
| θ value | 3.31772e-103 (rank : 11) | NC score | 0.832559 (rank : 14) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O00443 | Gene names | PIK3C2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing alpha polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-alpha) (PtdIns-3-kinase C2 alpha) (PI3K-C2alpha). | |||||
|
P3C2G_MOUSE
|
||||||
| θ value | 6.07444e-89 (rank : 12) | NC score | 0.907430 (rank : 10) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O70167, O70168 | Gene names | Pik3c2g | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing gamma polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-gamma) (PtdIns-3-kinase C2 gamma) (PI3K-C2gamma). | |||||
|
P3C2G_HUMAN
|
||||||
| θ value | 1.82896e-85 (rank : 13) | NC score | 0.908405 (rank : 9) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O75747 | Gene names | PIK3C2G | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing gamma polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-gamma) (PtdIns-3-kinase C2 gamma) (PI3K-C2gamma). | |||||
|
PK3C3_HUMAN
|
||||||
| θ value | 3.99248e-40 (rank : 14) | NC score | 0.857962 (rank : 11) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8NEB9, Q15134 | Gene names | PIK3C3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3) (Phosphatidylinositol 3-kinase p100 subunit). | |||||
|
PK3C3_MOUSE
|
||||||
| θ value | 1.16164e-39 (rank : 15) | NC score | 0.856610 (rank : 12) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6PF93, Q8R3S8 | Gene names | Pik3c3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3). | |||||
|
PI4KA_HUMAN
|
||||||
| θ value | 5.39604e-37 (rank : 16) | NC score | 0.829407 (rank : 15) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P42356, Q7Z625, Q9UPG2 | Gene names | PIK4CA, PIK4 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4-kinase alpha (EC 2.7.1.67) (PI4-kinase alpha) (PtdIns-4-kinase alpha) (PI4K-alpha). | |||||
|
PI4KB_HUMAN
|
||||||
| θ value | 3.62785e-25 (rank : 17) | NC score | 0.776189 (rank : 18) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UBF8, O15096, P78405, Q9BWR6 | Gene names | PIK4CB, PI4KB | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4-kinase beta (EC 2.7.1.67) (PtdIns 4-kinase beta) (PI4Kbeta) (PI4K-beta) (NPIK) (PI4K92). | |||||
|
PI4KB_MOUSE
|
||||||
| θ value | 3.62785e-25 (rank : 18) | NC score | 0.778480 (rank : 17) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8BKC8, Q8C146 | Gene names | Pik4cb | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4-kinase beta (EC 2.7.1.67) (PtdIns 4-kinase beta) (PI4Kbeta) (PI4K-beta). | |||||
|
FRAP_HUMAN
|
||||||
| θ value | 1.99992e-07 (rank : 19) | NC score | 0.397375 (rank : 20) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P42345, Q6LE87, Q96QG3, Q9Y4I3 | Gene names | FRAP1, FRAP, FRAP2 | |||
|
Domain Architecture |

|
|||||
| Description | FKBP12-rapamycin complex-associated protein (FK506-binding protein 12- rapamycin complex-associated protein 1) (Rapamycin target protein) (RAPT1) (Mammalian target of rapamycin) (mTOR). | |||||
|
FRAP_MOUSE
|
||||||
| θ value | 1.99992e-07 (rank : 20) | NC score | 0.398672 (rank : 19) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9JLN9, Q811J5, Q9CST1 | Gene names | Frap1, Frap | |||
|
Domain Architecture |

|
|||||
| Description | FKBP12-rapamycin complex-associated protein (FK506-binding protein 12- rapamycin complex-associated protein 1) (Rapamycin target protein) (RAPT1) (Mammalian target of rapamycin) (mTOR). | |||||
|
ATM_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 21) | NC score | 0.324958 (rank : 23) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q13315, O15429, Q12758, Q16551, Q93007, Q9NP02, Q9UCX7 | Gene names | ATM | |||
|
Domain Architecture |

|
|||||
| Description | Serine-protein kinase ATM (EC 2.7.11.1) (Ataxia telangiectasia mutated) (A-T, mutated). | |||||
|
PRKDC_MOUSE
|
||||||
| θ value | 0.000158464 (rank : 22) | NC score | 0.367673 (rank : 21) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P97313, O88187, P97928, Q307W9, Q3V2W8, Q8C2A7, Q9Z341 | Gene names | Prkdc, Xrcc7 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-dependent protein kinase catalytic subunit (EC 2.7.11.1) (DNA-PK catalytic subunit) (DNA-PKcs) (P460). | |||||
|
PRKDC_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 23) | NC score | 0.361361 (rank : 22) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P78527, P78528, Q13327, Q13337, Q14175, Q59H99, Q7Z611, Q96SE6, Q9UME3 | Gene names | PRKDC | |||
|
Domain Architecture |

|
|||||
| Description | DNA-dependent protein kinase catalytic subunit (EC 2.7.11.1) (DNA-PK catalytic subunit) (DNA-PKcs) (DNPK1) (p460). | |||||
|
ATM_MOUSE
|
||||||
| θ value | 0.000461057 (rank : 24) | NC score | 0.314750 (rank : 26) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q62388 | Gene names | Atm | |||
|
Domain Architecture |

|
|||||
| Description | Serine-protein kinase ATM (EC 2.7.11.1) (Ataxia telangiectasia mutated homolog) (A-T, mutated homolog). | |||||
|
ATR_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 25) | NC score | 0.321687 (rank : 24) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q13535, Q7KYL3, Q93051, Q9BXK4 | Gene names | ATR, FRP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ATR (EC 2.7.11.1) (Ataxia telangiectasia and Rad3-related protein) (FRAP-related protein 1). | |||||
|
ATR_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 26) | NC score | 0.320237 (rank : 25) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9JKK8 | Gene names | Atr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ATR (EC 2.7.11.1) (Ataxia telangiectasia and Rad3-related protein). | |||||
|
SMG1_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 27) | NC score | 0.265297 (rank : 28) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96Q15, O43305, Q13284, Q8NFX2, Q96QV0, Q96RW3 | Gene names | SMG1, ATX, KIAA0421, LIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SMG1 (EC 2.7.11.1) (SMG-1) (hSMG-1) (Lambda/iota protein kinase C-interacting protein) (Lambda-interacting protein) (61E3.4). | |||||
|
SMG1_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 28) | NC score | 0.267432 (rank : 27) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BKX6, Q5DU42, Q6ZQC0, Q8BLU4, Q8BWJ5, Q8BXD3 | Gene names | Smg1, Kiaa0421 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SMG1 (EC 2.7.11.1) (SMG-1). | |||||
|
LARP4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 29) | NC score | 0.028981 (rank : 49) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q71RC2, Q5CZ97, Q96NF9 | Gene names | LARP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | La-related protein 4 (La ribonucleoprotein domain family member 4). | |||||
|
WASF3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 30) | NC score | 0.029733 (rank : 47) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UPY6, O94974 | Gene names | WASF3, KIAA0900, SCAR3, WAVE3 | |||
|
Domain Architecture |

|
|||||
| Description | Wiskott-Aldrich syndrome protein family member 3 (WASP-family protein member 3) (Protein WAVE-3) (Verprolin homology domain-containing protein 3). | |||||
|
WASF3_MOUSE
|
||||||
| θ value | 1.06291 (rank : 31) | NC score | 0.029423 (rank : 48) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8VHI6 | Gene names | Wasf3, Wave3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 3 (WASP-family protein member 3) (Protein WAVE-3). | |||||
|
FBX46_HUMAN
|
||||||
| θ value | 2.36792 (rank : 32) | NC score | 0.024375 (rank : 51) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PJ61 | Gene names | FBXO46, FBX46, FBXO34L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 46 (F-box only protein 34-like). | |||||
|
FBX46_MOUSE
|
||||||
| θ value | 2.36792 (rank : 33) | NC score | 0.025444 (rank : 50) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BG80 | Gene names | Fbxo46, Fbx46, Fbxo34l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 46 (F-box only protein 34-like). | |||||
|
ARMX2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 34) | NC score | 0.010474 (rank : 62) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7L311, O60267, Q5H9D9 | Gene names | ARMCX2, ALEX2, KIAA0512 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 2 (Protein ALEX2) (ARM protein lost in epithelial cancers on chromosome X 2). | |||||
|
WASF1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | 0.020925 (rank : 54) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92558 | Gene names | WASF1, KIAA0269, SCAR1, WAVE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 1 (WASP-family protein member 1) (Protein WAVE-1) (Verprolin homology domain-containing protein 1). | |||||
|
WASF1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.020848 (rank : 55) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R5H6, Q91W51, Q9ERQ9 | Gene names | Wasf1, Wave1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 1 (WASP-family protein member 1) (Protein WAVE-1). | |||||
|
AP4B1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 37) | NC score | 0.021151 (rank : 53) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9WV76 | Gene names | Ap4b1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-4 complex subunit beta-1 (Adapter-related protein complex 4 beta-1 subunit) (Beta subunit of AP-4) (AP-4 adapter complex beta subunit). | |||||
|
ARMX2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 38) | NC score | 0.008514 (rank : 64) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6A058, Q9CXI9 | Gene names | Armcx2, Kiaa0512 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 2. | |||||
|
DYH9_HUMAN
|
||||||
| θ value | 5.27518 (rank : 39) | NC score | 0.005004 (rank : 65) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NYC9, O95494, Q9NQ28 | Gene names | DNAH9, DNAH17L, DNAL1 | |||
|
Domain Architecture |

|
|||||
| Description | Ciliary dynein heavy chain 9 (Axonemal beta dynein heavy chain 9). | |||||
|
AP2B1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | 0.014139 (rank : 58) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P63010, P21851, Q96J19 | Gene names | AP2B1, CLAPB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-2 complex subunit beta-1 (Adapter-related protein complex 2 beta-1 subunit) (Beta-adaptin) (Plasma membrane adaptor HA2/AP2 adaptin beta subunit) (Clathrin assembly protein complex 2 beta large chain) (AP105B). | |||||
|
AP2B1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.014141 (rank : 57) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DBG3, Q80XJ4 | Gene names | Ap2b1, Clapb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-2 complex subunit beta-1 (Adapter-related protein complex 2 beta-1 subunit) (Beta-adaptin) (Plasma membrane adaptor HA2/AP2 adaptin beta subunit) (Clathrin assembly protein complex 2 beta large chain) (AP105B). | |||||
|
AP4B1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.020509 (rank : 56) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y6B7 | Gene names | AP4B1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-4 complex subunit beta-1 (Adapter-related protein complex 4 beta 1 subunit) (Beta subunit of AP-4) (AP-4 adapter complex beta subunit). | |||||
|
AP1B1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 43) | NC score | 0.011689 (rank : 60) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q10567, P78436 | Gene names | AP1B1, ADTB1, BAM22 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit beta-1 (Adapter-related protein complex 1 beta-1 subunit) (Beta-adaptin 1) (Adaptor protein complex AP-1 beta-1 subunit) (Golgi adaptor HA1/AP1 adaptin beta subunit) (Clathrin assembly protein complex 1 beta large chain). | |||||
|
AP1B1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 44) | NC score | 0.011657 (rank : 61) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O35643, Q922E2 | Gene names | Ap1b1, Adtb1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit beta-1 (Adapter-related protein complex 1 beta-1 subunit) (Beta-adaptin 1) (Adaptor protein complex AP-1 beta-1 subunit) (Golgi adaptor HA1/AP1 adaptin beta subunit) (Clathrin assembly protein complex 1 beta large chain). | |||||
|
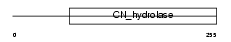
BUP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 45) | NC score | 0.024363 (rank : 52) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UBR1, Q9UIR3 | Gene names | UPB1, BUP1 | |||
|
Domain Architecture |
|
|||||
| Description | Beta-ureidopropionase (EC 3.5.1.6) (Beta-alanine synthase) (N- carbamoyl-beta-alanine amidohydrolase) (BUP-1). | |||||
|
LPPRC_HUMAN
|
||||||
| θ value | 8.99809 (rank : 46) | NC score | 0.008678 (rank : 63) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P42704 | Gene names | LRPPRC, LRP130 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 130 kDa leucine-rich protein (LRP 130) (GP130) (Leucine-rich PPR motif-containing protein). | |||||
|
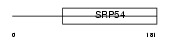
RFC2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 47) | NC score | 0.004777 (rank : 66) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9WUK4 | Gene names | Rfc2 | |||
|
Domain Architecture |
|
|||||
| Description | Replication factor C subunit 2 (Replication factor C 40 kDa subunit) (RF-C 40 kDa subunit) (RFC40) (Activator 1 40 kDa subunit) (A1 40 kDa subunit). | |||||
|
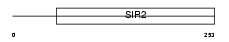
SIRT5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 48) | NC score | 0.013145 (rank : 59) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8K2C6 | Gene names | Sirt5, Sir2l5 | |||
|
Domain Architecture |
|
|||||
| Description | NAD-dependent deacetylase sirtuin-5 (EC 3.5.1.-) (SIR2-like protein 5). | |||||
|
FA62A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 49) | NC score | 0.074720 (rank : 32) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BSJ8, O94848, Q6PJN4, Q9H6J1, Q9H6W2, Q9Y416 | Gene names | FAM62A, KIAA0747, MBC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM62A (Membrane-bound C2 domain-containing protein). | |||||
|
FA62A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 50) | NC score | 0.083750 (rank : 31) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3U7R1, Q8C8R1, Q91X62, Q9CVH0, Q9Z1X5, Q9Z1X6 | Gene names | Fam62a, Mbc2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM62A (Membrane-bound C2 domain-containing protein). | |||||
|
RIMS1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.069961 (rank : 34) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 437 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q86UR5, O15048, Q8TDY9, Q8TDZ5, Q9HBA1, Q9HBA2, Q9HBA3, Q9HBA4, Q9HBA5, Q9HBA6 | Gene names | RIMS1, KIAA0340, RIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 1 (Rab3-interacting molecule 1) (RIM 1). | |||||
|
RIMS2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.059619 (rank : 39) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UQ26, O43413, Q86XL9, Q8IWV9, Q8IWW1 | Gene names | RIMS2, KIAA0751, RIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2). | |||||
|
RIMS2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 53) | NC score | 0.059668 (rank : 38) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9EQZ7, Q8C433, Q8CCK2 | Gene names | Rims2, Rab3ip2, Rim2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2) (Rab3-interacting protein 2). | |||||
|
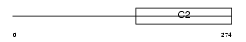
RIMS3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 54) | NC score | 0.053224 (rank : 45) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UJD0, Q92511 | Gene names | RIMS3, KIAA0237 | |||
|
Domain Architecture |
|
|||||
| Description | Regulating synaptic membrane exocytosis protein 3 (Nim3) (Rab-3- interacting molecule 3) (RIM 3) (RIM3 gamma). | |||||
|
RIMS3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.052478 (rank : 46) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80U57 | Gene names | Rims3, Kiaa0237 | |||
|
Domain Architecture |
|
|||||
| Description | Regulating synaptic membrane exocytosis protein 3 (Nim3) (Rab-3- interacting molecule 3) (RIM 3) (RIM3 gamma). | |||||
|
RIMS4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 56) | NC score | 0.084937 (rank : 30) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H426, Q5JWT7 | Gene names | RIMS4 | |||
|
Domain Architecture |
|
|||||
| Description | Regulating synaptic membrane exocytosis protein 4 (Rab3-interacting molecule 4) (RIM 4) (RIM4 gamma). | |||||
|
RIMS4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 57) | NC score | 0.084939 (rank : 29) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P60191 | Gene names | Rims4 | |||
|
Domain Architecture |
|
|||||
| Description | Regulating synaptic membrane exocytosis protein 4 (Rab3-interacting molecule 4) (RIM 4) (RIM4 gamma). | |||||
|
SYTL1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 58) | NC score | 0.063359 (rank : 35) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IYJ3, Q96BB6, Q96GU6, Q96S89, Q96SI0 | Gene names | SYTL1, SLP1 | |||
|
Domain Architecture |
|
|||||
| Description | Synaptotagmin-like protein 1 (Exophilin-7) (Protein JFC1) (SB146). | |||||
|
SYTL1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 59) | NC score | 0.059077 (rank : 40) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99N80, Q99J26 | Gene names | Sytl1, Slp1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 1 (Exophilin-7). | |||||
|
SYTL2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.056710 (rank : 43) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9HCH5, Q2YDA7, Q8ND34, Q96BJ2, Q9H768, Q9NXM1 | Gene names | SYTL2, KIAA1597, SLP2 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 2 (Exophilin-4). | |||||
|
SYTL2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.055302 (rank : 44) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 253 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99N50, Q8BT37, Q99J89, Q99J90, Q99N51, Q99N52, Q99N55, Q99N56 | Gene names | Sytl2, Slp2 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 2 (Exophilin-4). | |||||
|
SYTL3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.074198 (rank : 33) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99N48, Q8C506, Q99N47, Q99N49, Q99N54, Q99N79 | Gene names | Sytl3, Slp3 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 3 (Exophilin-6). | |||||
|
SYTL4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.057468 (rank : 41) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96C24, Q5H9J3, Q5JPG8, Q8N9P4, Q9H4R0, Q9H4R1 | Gene names | SYTL4 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 4 (Exophilin-2) (Granuphilin). | |||||
|
SYTL4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.057126 (rank : 42) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9R0Q1, Q8R321, Q9R0Q0 | Gene names | Sytl4, Slp4 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 4 (Exophilin-2) (Granuphilin). | |||||
|
SYTL5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.061981 (rank : 36) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TDW5 | Gene names | SYTL5, SLP5 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 5. | |||||
|
SYTL5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.061346 (rank : 37) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80T23, Q812E5 | Gene names | Sytl5, Slp5 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 5. | |||||
|
PK3CA_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P42337 | Gene names | Pik3ca | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (EC 2.7.1.153) (PI3-kinase p110 subunit alpha) (PtdIns-3- kinase p110) (PI3K). | |||||
|
PK3CA_HUMAN
|
||||||
| NC score | 0.999669 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P42336, Q99762 | Gene names | PIK3CA | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (EC 2.7.1.153) (PI3-kinase p110 subunit alpha) (PtdIns-3- kinase p110) (PI3K). | |||||
|
PK3CD_HUMAN
|
||||||
| NC score | 0.981681 (rank : 3) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O00329, O15445 | Gene names | PIK3CD | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit delta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit delta) (PtdIns-3- kinase p110) (PI3K) (p110delta). | |||||
|
PK3CD_MOUSE
|
||||||
| NC score | 0.981320 (rank : 4) | θ value | 0 (rank : 6) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O35904 | Gene names | Pik3cd | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit delta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit delta) (PtdIns-3- kinase p110) (PI3K) (p110delta). | |||||
|
PK3CB_HUMAN
|
||||||
| NC score | 0.979415 (rank : 5) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P42338 | Gene names | PIK3CB | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit beta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit beta) (PtdIns-3-kinase p110) (PI3K) (PI3Kbeta). | |||||
|
PK3CB_MOUSE
|
||||||
| NC score | 0.979293 (rank : 6) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BTI9 | Gene names | Pik3cb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit beta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit beta) (PtdIns-3-kinase p110) (PI3K) (PI3Kbeta). | |||||
|
PK3CG_HUMAN
|
||||||
| NC score | 0.966367 (rank : 7) | θ value | 3.27187e-151 (rank : 7) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P48736, Q8IV23, Q9BZC8 | Gene names | PIK3CG | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit gamma isoform (EC 2.7.1.153) (PI3-kinase p110 subunit gamma) (PtdIns-3- kinase subunit p110) (PI3K) (PI3Kgamma) (p120-PI3K). | |||||
|
PK3CG_MOUSE
|
||||||
| NC score | 0.965955 (rank : 8) | θ value | 1.62381e-150 (rank : 8) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9JHG7 | Gene names | Pik3cg, Pi3kg1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit gamma isoform (EC 2.7.1.153) (PI3-kinase p110 subunit gamma) (PtdIns-3- kinase subunit p110) (PI3K) (PI3Kgamma) (p120-PI3K). | |||||
|
P3C2G_HUMAN
|
||||||
| NC score | 0.908405 (rank : 9) | θ value | 1.82896e-85 (rank : 13) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O75747 | Gene names | PIK3C2G | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing gamma polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-gamma) (PtdIns-3-kinase C2 gamma) (PI3K-C2gamma). | |||||
|
P3C2G_MOUSE
|
||||||
| NC score | 0.907430 (rank : 10) | θ value | 6.07444e-89 (rank : 12) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O70167, O70168 | Gene names | Pik3c2g | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing gamma polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-gamma) (PtdIns-3-kinase C2 gamma) (PI3K-C2gamma). | |||||
|
PK3C3_HUMAN
|
||||||
| NC score | 0.857962 (rank : 11) | θ value | 3.99248e-40 (rank : 14) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8NEB9, Q15134 | Gene names | PIK3C3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3) (Phosphatidylinositol 3-kinase p100 subunit). | |||||
|
PK3C3_MOUSE
|
||||||
| NC score | 0.856610 (rank : 12) | θ value | 1.16164e-39 (rank : 15) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6PF93, Q8R3S8 | Gene names | Pik3c3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3). | |||||
|
P3C2A_MOUSE
|
||||||
| NC score | 0.835124 (rank : 13) | θ value | 6.45994e-107 (rank : 9) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q61194, Q61182 | Gene names | Pik3c2a, Cpk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing alpha polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-alpha) (PtdIns-3-kinase C2 alpha) (PI3K-C2alpha) (Cpk-m) (p170). | |||||
|
P3C2A_HUMAN
|
||||||
| NC score | 0.832559 (rank : 14) | θ value | 3.31772e-103 (rank : 11) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O00443 | Gene names | PIK3C2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing alpha polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-alpha) (PtdIns-3-kinase C2 alpha) (PI3K-C2alpha). | |||||
|
PI4KA_HUMAN
|
||||||
| NC score | 0.829407 (rank : 15) | θ value | 5.39604e-37 (rank : 16) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P42356, Q7Z625, Q9UPG2 | Gene names | PIK4CA, PIK4 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4-kinase alpha (EC 2.7.1.67) (PI4-kinase alpha) (PtdIns-4-kinase alpha) (PI4K-alpha). | |||||
|
P3C2B_HUMAN
|
||||||
| NC score | 0.811608 (rank : 16) | θ value | 1.1019e-106 (rank : 10) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O00750, O95666, Q5SW99 | Gene names | PIK3C2B | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing beta polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-beta) (PtdIns-3-kinase C2 beta) (PI3K-C2beta) (C2-PI3K). | |||||
|
PI4KB_MOUSE
|
||||||
| NC score | 0.778480 (rank : 17) | θ value | 3.62785e-25 (rank : 18) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8BKC8, Q8C146 | Gene names | Pik4cb | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4-kinase beta (EC 2.7.1.67) (PtdIns 4-kinase beta) (PI4Kbeta) (PI4K-beta). | |||||
|
PI4KB_HUMAN
|
||||||
| NC score | 0.776189 (rank : 18) | θ value | 3.62785e-25 (rank : 17) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UBF8, O15096, P78405, Q9BWR6 | Gene names | PIK4CB, PI4KB | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4-kinase beta (EC 2.7.1.67) (PtdIns 4-kinase beta) (PI4Kbeta) (PI4K-beta) (NPIK) (PI4K92). | |||||
|
FRAP_MOUSE
|
||||||
| NC score | 0.398672 (rank : 19) | θ value | 1.99992e-07 (rank : 20) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9JLN9, Q811J5, Q9CST1 | Gene names | Frap1, Frap | |||
|
Domain Architecture |

|
|||||
| Description | FKBP12-rapamycin complex-associated protein (FK506-binding protein 12- rapamycin complex-associated protein 1) (Rapamycin target protein) (RAPT1) (Mammalian target of rapamycin) (mTOR). | |||||
|
FRAP_HUMAN
|
||||||
| NC score | 0.397375 (rank : 20) | θ value | 1.99992e-07 (rank : 19) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P42345, Q6LE87, Q96QG3, Q9Y4I3 | Gene names | FRAP1, FRAP, FRAP2 | |||
|
Domain Architecture |

|
|||||
| Description | FKBP12-rapamycin complex-associated protein (FK506-binding protein 12- rapamycin complex-associated protein 1) (Rapamycin target protein) (RAPT1) (Mammalian target of rapamycin) (mTOR). | |||||
|
PRKDC_MOUSE
|
||||||
| NC score | 0.367673 (rank : 21) | θ value | 0.000158464 (rank : 22) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P97313, O88187, P97928, Q307W9, Q3V2W8, Q8C2A7, Q9Z341 | Gene names | Prkdc, Xrcc7 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-dependent protein kinase catalytic subunit (EC 2.7.11.1) (DNA-PK catalytic subunit) (DNA-PKcs) (P460). | |||||
|
PRKDC_HUMAN
|
||||||
| NC score | 0.361361 (rank : 22) | θ value | 0.00020696 (rank : 23) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P78527, P78528, Q13327, Q13337, Q14175, Q59H99, Q7Z611, Q96SE6, Q9UME3 | Gene names | PRKDC | |||
|
Domain Architecture |

|
|||||
| Description | DNA-dependent protein kinase catalytic subunit (EC 2.7.11.1) (DNA-PK catalytic subunit) (DNA-PKcs) (DNPK1) (p460). | |||||
|
ATM_HUMAN
|
||||||
| NC score | 0.324958 (rank : 23) | θ value | 0.000158464 (rank : 21) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q13315, O15429, Q12758, Q16551, Q93007, Q9NP02, Q9UCX7 | Gene names | ATM | |||
|
Domain Architecture |

|
|||||
| Description | Serine-protein kinase ATM (EC 2.7.11.1) (Ataxia telangiectasia mutated) (A-T, mutated). | |||||
|
ATR_HUMAN
|
||||||
| NC score | 0.321687 (rank : 24) | θ value | 0.00175202 (rank : 25) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q13535, Q7KYL3, Q93051, Q9BXK4 | Gene names | ATR, FRP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ATR (EC 2.7.11.1) (Ataxia telangiectasia and Rad3-related protein) (FRAP-related protein 1). | |||||
|
ATR_MOUSE
|
||||||
| NC score | 0.320237 (rank : 25) | θ value | 0.00175202 (rank : 26) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9JKK8 | Gene names | Atr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ATR (EC 2.7.11.1) (Ataxia telangiectasia and Rad3-related protein). | |||||
|
ATM_MOUSE
|
||||||
| NC score | 0.314750 (rank : 26) | θ value | 0.000461057 (rank : 24) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q62388 | Gene names | Atm | |||
|
Domain Architecture |

|
|||||
| Description | Serine-protein kinase ATM (EC 2.7.11.1) (Ataxia telangiectasia mutated homolog) (A-T, mutated homolog). | |||||
|
SMG1_MOUSE
|
||||||
| NC score | 0.267432 (rank : 27) | θ value | 0.00869519 (rank : 28) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BKX6, Q5DU42, Q6ZQC0, Q8BLU4, Q8BWJ5, Q8BXD3 | Gene names | Smg1, Kiaa0421 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SMG1 (EC 2.7.11.1) (SMG-1). | |||||
|
SMG1_HUMAN
|
||||||
| NC score | 0.265297 (rank : 28) | θ value | 0.00869519 (rank : 27) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96Q15, O43305, Q13284, Q8NFX2, Q96QV0, Q96RW3 | Gene names | SMG1, ATX, KIAA0421, LIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SMG1 (EC 2.7.11.1) (SMG-1) (hSMG-1) (Lambda/iota protein kinase C-interacting protein) (Lambda-interacting protein) (61E3.4). | |||||
|
RIMS4_MOUSE
|
||||||
| NC score | 0.084939 (rank : 29) | θ value | θ > 10 (rank : 57) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P60191 | Gene names | Rims4 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 4 (Rab3-interacting molecule 4) (RIM 4) (RIM4 gamma). | |||||
|
RIMS4_HUMAN
|
||||||
| NC score | 0.084937 (rank : 30) | θ value | θ > 10 (rank : 56) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H426, Q5JWT7 | Gene names | RIMS4 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 4 (Rab3-interacting molecule 4) (RIM 4) (RIM4 gamma). | |||||
|
FA62A_MOUSE
|
||||||
| NC score | 0.083750 (rank : 31) | θ value | θ > 10 (rank : 50) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3U7R1, Q8C8R1, Q91X62, Q9CVH0, Q9Z1X5, Q9Z1X6 | Gene names | Fam62a, Mbc2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM62A (Membrane-bound C2 domain-containing protein). | |||||
|
FA62A_HUMAN
|
||||||
| NC score | 0.074720 (rank : 32) | θ value | θ > 10 (rank : 49) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BSJ8, O94848, Q6PJN4, Q9H6J1, Q9H6W2, Q9Y416 | Gene names | FAM62A, KIAA0747, MBC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM62A (Membrane-bound C2 domain-containing protein). | |||||
|
SYTL3_MOUSE
|
||||||
| NC score | 0.074198 (rank : 33) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99N48, Q8C506, Q99N47, Q99N49, Q99N54, Q99N79 | Gene names | Sytl3, Slp3 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 3 (Exophilin-6). | |||||
|
RIMS1_HUMAN
|
||||||
| NC score | 0.069961 (rank : 34) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 437 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q86UR5, O15048, Q8TDY9, Q8TDZ5, Q9HBA1, Q9HBA2, Q9HBA3, Q9HBA4, Q9HBA5, Q9HBA6 | Gene names | RIMS1, KIAA0340, RIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 1 (Rab3-interacting molecule 1) (RIM 1). | |||||
|
SYTL1_HUMAN
|
||||||
| NC score | 0.063359 (rank : 35) | θ value | θ > 10 (rank : 58) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IYJ3, Q96BB6, Q96GU6, Q96S89, Q96SI0 | Gene names | SYTL1, SLP1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 1 (Exophilin-7) (Protein JFC1) (SB146). | |||||
|
SYTL5_HUMAN
|
||||||
| NC score | 0.061981 (rank : 36) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TDW5 | Gene names | SYTL5, SLP5 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 5. | |||||
|
SYTL5_MOUSE
|
||||||
| NC score | 0.061346 (rank : 37) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80T23, Q812E5 | Gene names | Sytl5, Slp5 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 5. | |||||
|
RIMS2_MOUSE
|
||||||
| NC score | 0.059668 (rank : 38) | θ value | θ > 10 (rank : 53) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9EQZ7, Q8C433, Q8CCK2 | Gene names | Rims2, Rab3ip2, Rim2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2) (Rab3-interacting protein 2). | |||||
|
RIMS2_HUMAN
|
||||||
| NC score | 0.059619 (rank : 39) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UQ26, O43413, Q86XL9, Q8IWV9, Q8IWW1 | Gene names | RIMS2, KIAA0751, RIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2). | |||||
|
SYTL1_MOUSE
|
||||||
| NC score | 0.059077 (rank : 40) | θ value | θ > 10 (rank : 59) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99N80, Q99J26 | Gene names | Sytl1, Slp1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 1 (Exophilin-7). | |||||
|
SYTL4_HUMAN
|
||||||
| NC score | 0.057468 (rank : 41) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96C24, Q5H9J3, Q5JPG8, Q8N9P4, Q9H4R0, Q9H4R1 | Gene names | SYTL4 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 4 (Exophilin-2) (Granuphilin). | |||||
|
SYTL4_MOUSE
|
||||||
| NC score | 0.057126 (rank : 42) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9R0Q1, Q8R321, Q9R0Q0 | Gene names | Sytl4, Slp4 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 4 (Exophilin-2) (Granuphilin). | |||||
|
SYTL2_HUMAN
|
||||||
| NC score | 0.056710 (rank : 43) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9HCH5, Q2YDA7, Q8ND34, Q96BJ2, Q9H768, Q9NXM1 | Gene names | SYTL2, KIAA1597, SLP2 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 2 (Exophilin-4). | |||||
|
SYTL2_MOUSE
|
||||||
| NC score | 0.055302 (rank : 44) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 253 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99N50, Q8BT37, Q99J89, Q99J90, Q99N51, Q99N52, Q99N55, Q99N56 | Gene names | Sytl2, Slp2 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 2 (Exophilin-4). | |||||
|
RIMS3_HUMAN
|
||||||
| NC score | 0.053224 (rank : 45) | θ value | θ > 10 (rank : 54) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UJD0, Q92511 | Gene names | RIMS3, KIAA0237 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 3 (Nim3) (Rab-3- interacting molecule 3) (RIM 3) (RIM3 gamma). | |||||
|
RIMS3_MOUSE
|
||||||
| NC score | 0.052478 (rank : 46) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80U57 | Gene names | Rims3, Kiaa0237 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 3 (Nim3) (Rab-3- interacting molecule 3) (RIM 3) (RIM3 gamma). | |||||
|
WASF3_HUMAN
|
||||||
| NC score | 0.029733 (rank : 47) | θ value | 1.06291 (rank : 30) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UPY6, O94974 | Gene names | WASF3, KIAA0900, SCAR3, WAVE3 | |||
|
Domain Architecture |

|
|||||
| Description | Wiskott-Aldrich syndrome protein family member 3 (WASP-family protein member 3) (Protein WAVE-3) (Verprolin homology domain-containing protein 3). | |||||
|
WASF3_MOUSE
|
||||||
| NC score | 0.029423 (rank : 48) | θ value | 1.06291 (rank : 31) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8VHI6 | Gene names | Wasf3, Wave3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 3 (WASP-family protein member 3) (Protein WAVE-3). | |||||
|
LARP4_HUMAN
|
||||||
| NC score | 0.028981 (rank : 49) | θ value | 1.06291 (rank : 29) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q71RC2, Q5CZ97, Q96NF9 | Gene names | LARP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | La-related protein 4 (La ribonucleoprotein domain family member 4). | |||||
|
FBX46_MOUSE
|
||||||
| NC score | 0.025444 (rank : 50) | θ value | 2.36792 (rank : 33) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BG80 | Gene names | Fbxo46, Fbx46, Fbxo34l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 46 (F-box only protein 34-like). | |||||
|
FBX46_HUMAN
|
||||||
| NC score | 0.024375 (rank : 51) | θ value | 2.36792 (rank : 32) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PJ61 | Gene names | FBXO46, FBX46, FBXO34L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 46 (F-box only protein 34-like). | |||||
|
BUP1_HUMAN
|
||||||
| NC score | 0.024363 (rank : 52) | θ value | 8.99809 (rank : 45) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UBR1, Q9UIR3 | Gene names | UPB1, BUP1 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-ureidopropionase (EC 3.5.1.6) (Beta-alanine synthase) (N- carbamoyl-beta-alanine amidohydrolase) (BUP-1). | |||||
|
AP4B1_MOUSE
|
||||||
| NC score | 0.021151 (rank : 53) | θ value | 5.27518 (rank : 37) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9WV76 | Gene names | Ap4b1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-4 complex subunit beta-1 (Adapter-related protein complex 4 beta-1 subunit) (Beta subunit of AP-4) (AP-4 adapter complex beta subunit). | |||||
|
WASF1_HUMAN
|
||||||
| NC score | 0.020925 (rank : 54) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92558 | Gene names | WASF1, KIAA0269, SCAR1, WAVE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 1 (WASP-family protein member 1) (Protein WAVE-1) (Verprolin homology domain-containing protein 1). | |||||
|
WASF1_MOUSE
|
||||||
| NC score | 0.020848 (rank : 55) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R5H6, Q91W51, Q9ERQ9 | Gene names | Wasf1, Wave1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 1 (WASP-family protein member 1) (Protein WAVE-1). | |||||
|
AP4B1_HUMAN
|
||||||
| NC score | 0.020509 (rank : 56) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y6B7 | Gene names | AP4B1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-4 complex subunit beta-1 (Adapter-related protein complex 4 beta 1 subunit) (Beta subunit of AP-4) (AP-4 adapter complex beta subunit). | |||||
|
AP2B1_MOUSE
|
||||||
| NC score | 0.014141 (rank : 57) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DBG3, Q80XJ4 | Gene names | Ap2b1, Clapb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-2 complex subunit beta-1 (Adapter-related protein complex 2 beta-1 subunit) (Beta-adaptin) (Plasma membrane adaptor HA2/AP2 adaptin beta subunit) (Clathrin assembly protein complex 2 beta large chain) (AP105B). | |||||
|
AP2B1_HUMAN
|
||||||
| NC score | 0.014139 (rank : 58) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P63010, P21851, Q96J19 | Gene names | AP2B1, CLAPB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-2 complex subunit beta-1 (Adapter-related protein complex 2 beta-1 subunit) (Beta-adaptin) (Plasma membrane adaptor HA2/AP2 adaptin beta subunit) (Clathrin assembly protein complex 2 beta large chain) (AP105B). | |||||
|
SIRT5_MOUSE
|
||||||
| NC score | 0.013145 (rank : 59) | θ value | 8.99809 (rank : 48) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8K2C6 | Gene names | Sirt5, Sir2l5 | |||
|
Domain Architecture |

|
|||||
| Description | NAD-dependent deacetylase sirtuin-5 (EC 3.5.1.-) (SIR2-like protein 5). | |||||
|
AP1B1_HUMAN
|
||||||
| NC score | 0.011689 (rank : 60) | θ value | 8.99809 (rank : 43) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q10567, P78436 | Gene names | AP1B1, ADTB1, BAM22 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit beta-1 (Adapter-related protein complex 1 beta-1 subunit) (Beta-adaptin 1) (Adaptor protein complex AP-1 beta-1 subunit) (Golgi adaptor HA1/AP1 adaptin beta subunit) (Clathrin assembly protein complex 1 beta large chain). | |||||
|
AP1B1_MOUSE
|
||||||
| NC score | 0.011657 (rank : 61) | θ value | 8.99809 (rank : 44) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O35643, Q922E2 | Gene names | Ap1b1, Adtb1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit beta-1 (Adapter-related protein complex 1 beta-1 subunit) (Beta-adaptin 1) (Adaptor protein complex AP-1 beta-1 subunit) (Golgi adaptor HA1/AP1 adaptin beta subunit) (Clathrin assembly protein complex 1 beta large chain). | |||||
|
ARMX2_HUMAN
|
||||||
| NC score | 0.010474 (rank : 62) | θ value | 3.0926 (rank : 34) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7L311, O60267, Q5H9D9 | Gene names | ARMCX2, ALEX2, KIAA0512 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 2 (Protein ALEX2) (ARM protein lost in epithelial cancers on chromosome X 2). | |||||
|
LPPRC_HUMAN
|
||||||
| NC score | 0.008678 (rank : 63) | θ value | 8.99809 (rank : 46) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P42704 | Gene names | LRPPRC, LRP130 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 130 kDa leucine-rich protein (LRP 130) (GP130) (Leucine-rich PPR motif-containing protein). | |||||
|
ARMX2_MOUSE
|
||||||
| NC score | 0.008514 (rank : 64) | θ value | 5.27518 (rank : 38) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6A058, Q9CXI9 | Gene names | Armcx2, Kiaa0512 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 2. | |||||
|
DYH9_HUMAN
|
||||||
| NC score | 0.005004 (rank : 65) | θ value | 5.27518 (rank : 39) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NYC9, O95494, Q9NQ28 | Gene names | DNAH9, DNAH17L, DNAL1 | |||
|
Domain Architecture |

|
|||||
| Description | Ciliary dynein heavy chain 9 (Axonemal beta dynein heavy chain 9). | |||||
|
RFC2_MOUSE
|
||||||
| NC score | 0.004777 (rank : 66) | θ value | 8.99809 (rank : 47) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9WUK4 | Gene names | Rfc2 | |||
|
Domain Architecture |

|
|||||
| Description | Replication factor C subunit 2 (Replication factor C 40 kDa subunit) (RF-C 40 kDa subunit) (RFC40) (Activator 1 40 kDa subunit) (A1 40 kDa subunit). | |||||