Please be patient as the page loads
|
PRKDC_MOUSE
|
||||||
| SwissProt Accessions | P97313, O88187, P97928, Q307W9, Q3V2W8, Q8C2A7, Q9Z341 | Gene names | Prkdc, Xrcc7 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-dependent protein kinase catalytic subunit (EC 2.7.11.1) (DNA-PK catalytic subunit) (DNA-PKcs) (P460). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PRKDC_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.991668 (rank : 2) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P78527, P78528, Q13327, Q13337, Q14175, Q59H99, Q7Z611, Q96SE6, Q9UME3 | Gene names | PRKDC | |||
|
Domain Architecture |

|
|||||
| Description | DNA-dependent protein kinase catalytic subunit (EC 2.7.11.1) (DNA-PK catalytic subunit) (DNA-PKcs) (DNPK1) (p460). | |||||
|
PRKDC_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 92 | |
| SwissProt Accessions | P97313, O88187, P97928, Q307W9, Q3V2W8, Q8C2A7, Q9Z341 | Gene names | Prkdc, Xrcc7 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-dependent protein kinase catalytic subunit (EC 2.7.11.1) (DNA-PK catalytic subunit) (DNA-PKcs) (P460). | |||||
|
FRAP_MOUSE
|
||||||
| θ value | 3.99248e-40 (rank : 3) | NC score | 0.701658 (rank : 3) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9JLN9, Q811J5, Q9CST1 | Gene names | Frap1, Frap | |||
|
Domain Architecture |

|
|||||
| Description | FKBP12-rapamycin complex-associated protein (FK506-binding protein 12- rapamycin complex-associated protein 1) (Rapamycin target protein) (RAPT1) (Mammalian target of rapamycin) (mTOR). | |||||
|
FRAP_HUMAN
|
||||||
| θ value | 5.21438e-40 (rank : 4) | NC score | 0.701475 (rank : 4) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P42345, Q6LE87, Q96QG3, Q9Y4I3 | Gene names | FRAP1, FRAP, FRAP2 | |||
|
Domain Architecture |

|
|||||
| Description | FKBP12-rapamycin complex-associated protein (FK506-binding protein 12- rapamycin complex-associated protein 1) (Rapamycin target protein) (RAPT1) (Mammalian target of rapamycin) (mTOR). | |||||
|
ATR_HUMAN
|
||||||
| θ value | 1.57e-36 (rank : 5) | NC score | 0.696522 (rank : 5) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q13535, Q7KYL3, Q93051, Q9BXK4 | Gene names | ATR, FRP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ATR (EC 2.7.11.1) (Ataxia telangiectasia and Rad3-related protein) (FRAP-related protein 1). | |||||
|
ATR_MOUSE
|
||||||
| θ value | 4.27553e-34 (rank : 6) | NC score | 0.690161 (rank : 6) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9JKK8 | Gene names | Atr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ATR (EC 2.7.11.1) (Ataxia telangiectasia and Rad3-related protein). | |||||
|
ATM_HUMAN
|
||||||
| θ value | 6.17384e-33 (rank : 7) | NC score | 0.666895 (rank : 7) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q13315, O15429, Q12758, Q16551, Q93007, Q9NP02, Q9UCX7 | Gene names | ATM | |||
|
Domain Architecture |

|
|||||
| Description | Serine-protein kinase ATM (EC 2.7.11.1) (Ataxia telangiectasia mutated) (A-T, mutated). | |||||
|
ATM_MOUSE
|
||||||
| θ value | 3.06405e-32 (rank : 8) | NC score | 0.661853 (rank : 8) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q62388 | Gene names | Atm | |||
|
Domain Architecture |

|
|||||
| Description | Serine-protein kinase ATM (EC 2.7.11.1) (Ataxia telangiectasia mutated homolog) (A-T, mutated homolog). | |||||
|
SMG1_HUMAN
|
||||||
| θ value | 1.98606e-31 (rank : 9) | NC score | 0.626354 (rank : 10) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96Q15, O43305, Q13284, Q8NFX2, Q96QV0, Q96RW3 | Gene names | SMG1, ATX, KIAA0421, LIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SMG1 (EC 2.7.11.1) (SMG-1) (hSMG-1) (Lambda/iota protein kinase C-interacting protein) (Lambda-interacting protein) (61E3.4). | |||||
|
SMG1_MOUSE
|
||||||
| θ value | 1.98606e-31 (rank : 10) | NC score | 0.630363 (rank : 9) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8BKX6, Q5DU42, Q6ZQC0, Q8BLU4, Q8BWJ5, Q8BXD3 | Gene names | Smg1, Kiaa0421 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SMG1 (EC 2.7.11.1) (SMG-1). | |||||
|
TRRAP_HUMAN
|
||||||
| θ value | 1.47631e-18 (rank : 11) | NC score | 0.443930 (rank : 14) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Y4A5, O75218, Q9Y631, Q9Y6H4 | Gene names | TRRAP, PAF400 | |||
|
Domain Architecture |

|
|||||
| Description | Transformation/transcription domain-associated protein (350/400 kDa PCAF-associated factor) (PAF350/400) (STAF40) (Tra1 homolog). | |||||
|
TRRAP_MOUSE
|
||||||
| θ value | 3.63628e-17 (rank : 12) | NC score | 0.448781 (rank : 13) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q80YV3, Q8C0Z5, Q8K104 | Gene names | Trrap | |||
|
Domain Architecture |

|
|||||
| Description | Transformation/transcription domain-associated protein (Tra1 homolog). | |||||
|
PK3CG_HUMAN
|
||||||
| θ value | 3.18553e-13 (rank : 13) | NC score | 0.432340 (rank : 15) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P48736, Q8IV23, Q9BZC8 | Gene names | PIK3CG | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit gamma isoform (EC 2.7.1.153) (PI3-kinase p110 subunit gamma) (PtdIns-3- kinase subunit p110) (PI3K) (PI3Kgamma) (p120-PI3K). | |||||
|
PK3CG_MOUSE
|
||||||
| θ value | 9.26847e-13 (rank : 14) | NC score | 0.429758 (rank : 17) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9JHG7 | Gene names | Pik3cg, Pi3kg1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit gamma isoform (EC 2.7.1.153) (PI3-kinase p110 subunit gamma) (PtdIns-3- kinase subunit p110) (PI3K) (PI3Kgamma) (p120-PI3K). | |||||
|
PK3CD_MOUSE
|
||||||
| θ value | 2.28291e-11 (rank : 15) | NC score | 0.430136 (rank : 16) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O35904 | Gene names | Pik3cd | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit delta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit delta) (PtdIns-3- kinase p110) (PI3K) (p110delta). | |||||
|
PK3CD_HUMAN
|
||||||
| θ value | 2.98157e-11 (rank : 16) | NC score | 0.428892 (rank : 19) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O00329, O15445 | Gene names | PIK3CD | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit delta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit delta) (PtdIns-3- kinase p110) (PI3K) (p110delta). | |||||
|
PK3C3_MOUSE
|
||||||
| θ value | 5.08577e-11 (rank : 17) | NC score | 0.473717 (rank : 11) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q6PF93, Q8R3S8 | Gene names | Pik3c3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3). | |||||
|
PK3CB_MOUSE
|
||||||
| θ value | 8.67504e-11 (rank : 18) | NC score | 0.429505 (rank : 18) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BTI9 | Gene names | Pik3cb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit beta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit beta) (PtdIns-3-kinase p110) (PI3K) (PI3Kbeta). | |||||
|
PK3C3_HUMAN
|
||||||
| θ value | 1.133e-10 (rank : 19) | NC score | 0.472772 (rank : 12) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8NEB9, Q15134 | Gene names | PIK3C3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3) (Phosphatidylinositol 3-kinase p100 subunit). | |||||
|
PK3CB_HUMAN
|
||||||
| θ value | 3.29651e-10 (rank : 20) | NC score | 0.427825 (rank : 20) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P42338 | Gene names | PIK3CB | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit beta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit beta) (PtdIns-3-kinase p110) (PI3K) (PI3Kbeta). | |||||
|
PI4KB_HUMAN
|
||||||
| θ value | 1.43324e-05 (rank : 21) | NC score | 0.394248 (rank : 21) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UBF8, O15096, P78405, Q9BWR6 | Gene names | PIK4CB, PI4KB | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4-kinase beta (EC 2.7.1.67) (PtdIns 4-kinase beta) (PI4Kbeta) (PI4K-beta) (NPIK) (PI4K92). | |||||
|
PI4KB_MOUSE
|
||||||
| θ value | 1.43324e-05 (rank : 22) | NC score | 0.392409 (rank : 22) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8BKC8, Q8C146 | Gene names | Pik4cb | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4-kinase beta (EC 2.7.1.67) (PtdIns 4-kinase beta) (PI4Kbeta) (PI4K-beta). | |||||
|
P3C2B_HUMAN
|
||||||
| θ value | 0.000121331 (rank : 23) | NC score | 0.294297 (rank : 30) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O00750, O95666, Q5SW99 | Gene names | PIK3C2B | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing beta polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-beta) (PtdIns-3-kinase C2 beta) (PI3K-C2beta) (C2-PI3K). | |||||
|
P3C2G_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 24) | NC score | 0.344490 (rank : 26) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O75747 | Gene names | PIK3C2G | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing gamma polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-gamma) (PtdIns-3-kinase C2 gamma) (PI3K-C2gamma). | |||||
|
PK3CA_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 25) | NC score | 0.367855 (rank : 23) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P42336, Q99762 | Gene names | PIK3CA | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (EC 2.7.1.153) (PI3-kinase p110 subunit alpha) (PtdIns-3- kinase p110) (PI3K). | |||||
|
PK3CA_MOUSE
|
||||||
| θ value | 0.000158464 (rank : 26) | NC score | 0.367673 (rank : 24) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P42337 | Gene names | Pik3ca | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (EC 2.7.1.153) (PI3-kinase p110 subunit alpha) (PtdIns-3- kinase p110) (PI3K). | |||||
|
P3C2G_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 27) | NC score | 0.349066 (rank : 25) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O70167, O70168 | Gene names | Pik3c2g | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing gamma polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-gamma) (PtdIns-3-kinase C2 gamma) (PI3K-C2gamma). | |||||
|
P3C2A_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 28) | NC score | 0.308168 (rank : 28) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q61194, Q61182 | Gene names | Pik3c2a, Cpk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing alpha polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-alpha) (PtdIns-3-kinase C2 alpha) (PI3K-C2alpha) (Cpk-m) (p170). | |||||
|
P3C2A_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 29) | NC score | 0.304654 (rank : 29) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O00443 | Gene names | PIK3C2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing alpha polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-alpha) (PtdIns-3-kinase C2 alpha) (PI3K-C2alpha). | |||||
|
SIA8A_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 30) | NC score | 0.041627 (rank : 36) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q64687 | Gene names | St8sia1, Siat8, Siat8a | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-N-acetylneuraminide alpha-2,8-sialyltransferase (EC 2.4.99.8) (Ganglioside GD3 synthase) (Ganglioside GT3 synthase) (Alpha-2,8- sialyltransferase 8A) (ST8Sia I). | |||||
|
K0423_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 31) | NC score | 0.055328 (rank : 34) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6A070 | Gene names | Kiaa0423 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0423. | |||||
|
KMHN1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 32) | NC score | 0.027265 (rank : 38) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 950 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8TBZ0, Q86YI9, Q8N7W0 | Gene names | KMHN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cancer/testis antigen KM-HN-1. | |||||
|
CETP_HUMAN
|
||||||
| θ value | 0.125558 (rank : 33) | NC score | 0.069313 (rank : 31) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P11597, Q13987, Q13988 | Gene names | CETP | |||
|
Domain Architecture |
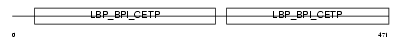
|
|||||
| Description | Cholesteryl ester transfer protein precursor (Lipid transfer protein I). | |||||
|
CCD46_HUMAN
|
||||||
| θ value | 0.21417 (rank : 34) | NC score | 0.019243 (rank : 42) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1430 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8N8E3, Q6PIB5, Q8NCR4, Q8NFR4 | Gene names | CCDC46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 46. | |||||
|
PI4KA_HUMAN
|
||||||
| θ value | 0.47712 (rank : 35) | NC score | 0.319553 (rank : 27) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P42356, Q7Z625, Q9UPG2 | Gene names | PIK4CA, PIK4 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4-kinase alpha (EC 2.7.1.67) (PI4-kinase alpha) (PtdIns-4-kinase alpha) (PI4K-alpha). | |||||
|
SMC4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 36) | NC score | 0.023115 (rank : 39) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NTJ3, O95752, Q8NDL4, Q9UNT9 | Gene names | SMC4L1, CAPC, SMC4 | |||
|
Domain Architecture |
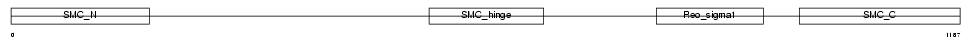
|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (hCAP-C) (XCAP-C homolog). | |||||
|
XPO4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 37) | NC score | 0.056197 (rank : 33) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9C0E2, Q5VUZ5, Q9H934 | Gene names | XPO4, KIAA1721 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Exportin-4 (Exp4). | |||||
|
XPO4_MOUSE
|
||||||
| θ value | 1.06291 (rank : 38) | NC score | 0.056242 (rank : 32) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ESJ0, Q6ZPJ4 | Gene names | Xpo4, Kiaa1721 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Exportin-4 (Exp4). | |||||
|
SIA8A_HUMAN
|
||||||
| θ value | 1.38821 (rank : 39) | NC score | 0.027833 (rank : 37) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92185, Q93064 | Gene names | ST8SIA1, SIAT8, SIAT8A | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-N-acetylneuraminide alpha-2,8-sialyltransferase (EC 2.4.99.8) (Ganglioside GD3 synthase) (Ganglioside GT3 synthase) (Alpha-2,8- sialyltransferase 8A) (ST8Sia I). | |||||
|
SYNE2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 40) | NC score | 0.019075 (rank : 43) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1323 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8WXH0, Q8N1S3, Q8NF49, Q8TER7, Q8WWW3, Q8WWW4, Q8WWW5, Q8WXH1, Q9NU50, Q9UFQ4, Q9Y2L4, Q9Y4R1 | Gene names | SYNE2, KIAA1011, NUA | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-2 (Nuclear envelope spectrin repeat protein 2) (Syne-2) (Synaptic nuclear envelope protein 2) (Nucleus and actin connecting element protein) (Protein NUANCE). | |||||
|
CENPI_HUMAN
|
||||||
| θ value | 1.81305 (rank : 41) | NC score | 0.044465 (rank : 35) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92674, Q5JWZ9, Q96ED0 | Gene names | CENPI, FSHPRH1, ICEN19, LRPR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein I (CENP-I) (Interphase centromere complex protein 19) (Follicle-stimulating hormone primary response protein) (FSH primary response protein 1) (Leucine-rich primary response protein 1). | |||||
|
DHX36_HUMAN
|
||||||
| θ value | 1.81305 (rank : 42) | NC score | 0.009462 (rank : 76) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H2U1, Q70JU3, Q8IYE5, Q9P240 | Gene names | DHX36, DDX36, KIAA1488, MLEL1, RHAU | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DHX36 (EC 3.6.1.-) (DEAH box protein 36) (MLE-like protein 1) (RNA helicase associated with AU-rich element ARE). | |||||
|
GCC2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 43) | NC score | 0.012995 (rank : 56) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1619 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8CHG3, Q8BR44, Q8R2Q5, Q9CT45 | Gene names | Gcc2, Kiaa0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185). | |||||
|
GOGB1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 44) | NC score | 0.012587 (rank : 59) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
ITPR1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 45) | NC score | 0.012160 (rank : 63) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q14643, Q14660, Q99897 | Gene names | ITPR1, INSP3R1 | |||
|
Domain Architecture |

|
|||||
| Description | Inositol 1,4,5-trisphosphate receptor type 1 (Type 1 inositol 1,4,5- trisphosphate receptor) (Type 1 InsP3 receptor) (IP3 receptor isoform 1) (InsP3R1) (IP3R). | |||||
|
SPAG5_MOUSE
|
||||||
| θ value | 1.81305 (rank : 46) | NC score | 0.019839 (rank : 41) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 612 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q7TME2, Q8CH61 | Gene names | Spag5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 5 (Mastrin) (Mitotic spindle-associated protein p126) (MAP126). | |||||
|
CE290_HUMAN
|
||||||
| θ value | 2.36792 (rank : 47) | NC score | 0.015252 (rank : 48) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
HMMR_HUMAN
|
||||||
| θ value | 2.36792 (rank : 48) | NC score | 0.014321 (rank : 52) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1082 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O75330, Q92767 | Gene names | HMMR, IHABP, RHAMM | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronan mediated motility receptor (Intracellular hyaluronic acid- binding protein) (Receptor for hyaluronan-mediated motility) (CD168 antigen). | |||||
|
K0776_MOUSE
|
||||||
| θ value | 2.36792 (rank : 49) | NC score | 0.020805 (rank : 40) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CCJ3, Q3V145, Q6ZQ50, Q8BT70, Q8C484, Q9D8I8 | Gene names | Kiaa0776 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0776. | |||||
|
MYH6_HUMAN
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.012475 (rank : 60) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
SPA12_HUMAN
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.005773 (rank : 81) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IW75 | Gene names | SERPINA12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serpin A12 precursor (Visceral adipose-specific serpin) (Visceral adipose tissue-derived serine protease inhibitor) (Vaspin) (OL-64). | |||||
|
BMX_HUMAN
|
||||||
| θ value | 3.0926 (rank : 52) | NC score | 0.001140 (rank : 86) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 942 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P51813, O60564, Q12871 | Gene names | BMX | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic tyrosine-protein kinase BMX (EC 2.7.10.2) (Bone marrow tyrosine kinase gene in chromosome X protein) (Epithelial and endothelial tyrosine kinase) (ETK) (NTK38). | |||||
|
BTK_HUMAN
|
||||||
| θ value | 3.0926 (rank : 53) | NC score | 0.000650 (rank : 88) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1026 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q06187 | Gene names | BTK, AGMX1, ATK, BPK | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase BTK (EC 2.7.10.2) (Bruton tyrosine kinase) (Agammaglobulinaemia tyrosine kinase) (ATK) (B cell progenitor kinase) (BPK). | |||||
|
FMNL_HUMAN
|
||||||
| θ value | 3.0926 (rank : 54) | NC score | 0.011040 (rank : 68) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 529 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O95466, Q6DKG5, Q6IBP3, Q86UH1, Q8N671, Q8TDH1, Q96H10 | Gene names | FMNL1, C17orf1, C17orf1B, FMNL | |||
|
Domain Architecture |
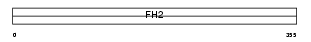
|
|||||
| Description | Formin-like protein 1 (Leukocyte formin) (CLL-associated antigen KW- 13). | |||||
|
MYO1B_MOUSE
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.010234 (rank : 72) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P46735, P70244 | Gene names | Myo1b | |||
|
Domain Architecture |

|
|||||
| Description | Myosin Ib (Myosin I alpha) (MMI-alpha) (MMIa) (MIH-L). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.013045 (rank : 55) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
TPM2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 57) | NC score | 0.016644 (rank : 46) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 677 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P07951, P06468, Q13894, Q53FM4, Q5TCU4, Q5TCU7, Q9UH67 | Gene names | TPM2, TMSB | |||
|
Domain Architecture |
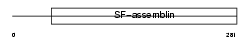
|
|||||
| Description | Tropomyosin beta chain (Tropomyosin 2) (Beta-tropomyosin). | |||||
|
TPM2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.017133 (rank : 45) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P58774, P02560, P46901 | Gene names | Tpm2, Tpm-2 | |||
|
Domain Architecture |
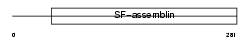
|
|||||
| Description | Tropomyosin beta chain (Tropomyosin 2) (Beta-tropomyosin). | |||||
|
ZN567_HUMAN
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | -0.002469 (rank : 92) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 769 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N184, Q6N044 | Gene names | ZNF567 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 567. | |||||
|
BTK_MOUSE
|
||||||
| θ value | 4.03905 (rank : 60) | NC score | 0.000486 (rank : 89) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1017 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P35991, Q61365 | Gene names | Btk, Bpk | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase BTK (EC 2.7.10.2) (Bruton tyrosine kinase) (Agammaglobulinaemia tyrosine kinase) (ATK) (B cell progenitor kinase) (BPK) (Kinase EMB). | |||||
|
CAB45_HUMAN
|
||||||
| θ value | 4.03905 (rank : 61) | NC score | 0.013516 (rank : 53) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BRK5, Q8NBQ3, Q9NZP7 | Gene names | SDF4, CAB45 | |||
|
Domain Architecture |
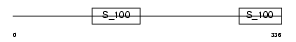
|
|||||
| Description | 45 kDa calcium-binding protein precursor (Cab45) (Stromal cell-derived factor 4) (SDF-4). | |||||
|
CALX_HUMAN
|
||||||
| θ value | 4.03905 (rank : 62) | NC score | 0.010679 (rank : 69) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P27824 | Gene names | CANX | |||
|
Domain Architecture |
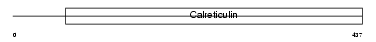
|
|||||
| Description | Calnexin precursor (Major histocompatibility complex class I antigen- binding protein p88) (p90) (IP90). | |||||
|
CD2L7_HUMAN
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | -0.000229 (rank : 90) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1275 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9NYV4, O94978 | Gene names | CRKRS, CRK7, KIAA0904 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle 2-related protein kinase 7 (EC 2.7.11.22) (CDC2- related protein kinase 7) (Cdc2-related kinase, arginine/serine-rich) (CrkRS). | |||||
|
KIF2B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.005110 (rank : 84) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N4N8, Q96MA2, Q9BXG6 | Gene names | KIF2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin-like protein KIF2B. | |||||
|
RPGF2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 65) | NC score | 0.012213 (rank : 61) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y4G8 | Gene names | RAPGEF2, KIAA0313, NRAPGEP, PDZGEF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rap guanine nucleotide exchange factor 2 (Neural RAP guanine nucleotide exchange protein) (nRap GEP) (PDZ domain-containing guanine nucleotide exchange factor 1) (PDZ-GEF1) (RA-GEF). | |||||
|
SORC3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 66) | NC score | 0.011946 (rank : 64) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UPU3, Q5VXF9, Q9NQJ2 | Gene names | SORCS3, KIAA1059 | |||
|
Domain Architecture |

|
|||||
| Description | VPS10 domain-containing receptor SorCS3 precursor. | |||||
|
GDF9_HUMAN
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.007837 (rank : 78) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O60383 | Gene names | GDF9 | |||
|
Domain Architecture |

|
|||||
| Description | Growth/differentiation factor 9 precursor (GDF-9). | |||||
|
M6PBP_MOUSE
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.012162 (rank : 62) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9DBG5, Q3TK05, Q8BKV9, Q9CZK1 | Gene names | M6prbp1, Tip47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mannose-6-phosphate receptor-binding protein 1 (Cargo selection protein TIP47). | |||||
|
PAI2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.004333 (rank : 85) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P12388, O35687, Q9QWP6, Q9QWP7, Q9QWP8, Q9QWP9, Q9QWQ0, Q9QWZ5, Q9QWZ6 | Gene names | Serpinb2, Pai2, Planh2 | |||
|
Domain Architecture |
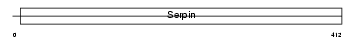
|
|||||
| Description | Plasminogen activator inhibitor 2, macrophage (PAI-2). | |||||
|
PON3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 70) | NC score | 0.010013 (rank : 74) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q62087, Q8C2I7 | Gene names | Pon3 | |||
|
Domain Architecture |

|
|||||
| Description | Serum paraoxonase/lactonase 3 (EC 3.1.1.-). | |||||
|
SLC2B_HUMAN
|
||||||
| θ value | 5.27518 (rank : 71) | NC score | 0.015648 (rank : 47) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NEV8, Q9Y4D6 | Gene names | EXPH5, KIAA0624, SLAC2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Slp homolog lacking C2 domains b (Exophilin-5). | |||||
|
SORC3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 72) | NC score | 0.011322 (rank : 67) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VI51, Q9CTR4 | Gene names | Sorcs3 | |||
|
Domain Architecture |

|
|||||
| Description | VPS10 domain-containing receptor SorCS3 precursor. | |||||
|
ST18_MOUSE
|
||||||
| θ value | 5.27518 (rank : 73) | NC score | 0.013213 (rank : 54) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80TY4, Q3UH00, Q3URH9, Q3UVB9, Q3UZN9, Q811B4, Q8K098 | Gene names | St18, Kiaa0535 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity protein 18. | |||||
|
XPOT_HUMAN
|
||||||
| θ value | 5.27518 (rank : 74) | NC score | 0.018125 (rank : 44) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O43592, O43784, Q8WUG2, Q9BVS7 | Gene names | XPOT | |||
|
Domain Architecture |

|
|||||
| Description | Exportin-T (tRNA exportin) (Exportin(tRNA)). | |||||
|
CP135_MOUSE
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.011699 (rank : 66) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1278 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6P5D4, Q6A030 | Gene names | Cep135, Cep4, Kiaa0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
K1H1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.005277 (rank : 83) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15323 | Gene names | KRTHA1, HHA1, HKA1 | |||
|
Domain Architecture |
|
|||||
| Description | Keratin, type I cuticular Ha1 (Hair keratin, type I Ha1). | |||||
|
K1HA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.005297 (rank : 82) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 502 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O76009 | Gene names | KRTHA3A, HHA3-I, HKA3A | |||
|
Domain Architecture |
|
|||||
| Description | Keratin, type I cuticular Ha3-I (Hair keratin, type I Ha3-I). | |||||
|
NOTC2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | -0.001088 (rank : 91) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1069 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O35516, Q06008, Q60941 | Gene names | Notch2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 2 precursor (Notch 2) (Motch B) [Contains: Notch 2 extracellular truncation; Notch 2 intracellular domain]. | |||||
|
RPGF6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.009949 (rank : 75) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TEU7, Q8NI21, Q8TEU6, Q96PC1 | Gene names | RAPGEF6, PDZGEF2 | |||
|
Domain Architecture |

|
|||||
| Description | Rap guanine nucleotide exchange factor 6 (PDZ domain-containing guanine nucleotide exchange factor 2) (PDZ-GEF2) (RA-GEF-2). | |||||
|
S35A3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.012837 (rank : 58) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y2D2, Q68CR2 | Gene names | SLC35A3 | |||
|
Domain Architecture |
|
|||||
| Description | UDP-N-acetylglucosamine transporter (Golgi UDP-GlcNAc transporter) (Solute carrier family 35 member A3). | |||||
|
S35A3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.012892 (rank : 57) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R1T4, Q8BRW7 | Gene names | Slc35a3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UDP-N-acetylglucosamine transporter (Golgi UDP-GlcNAc transporter) (Solute carrier family 35 member A3). | |||||
|
SMC4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.015207 (rank : 49) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 830 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8CG47, Q8BTS7, Q8BTY9, Q99K21 | Gene names | Smc4l1, Capc, Smc4 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (XCAP-C homolog). | |||||
|
SVIL_HUMAN
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.007613 (rank : 79) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O95425, O60611, O60612, Q5VZK5, Q5VZK6, Q9H1R7 | Gene names | SVIL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Supervillin (Archvillin) (p205/p250). | |||||
|
TPM1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.015207 (rank : 50) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 716 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P09493, P09494, P10469, Q86W64, Q96IK2, Q9UCY9 | Gene names | TPM1, TMSA | |||
|
Domain Architecture |
|
|||||
| Description | Tropomyosin 1 alpha chain (Alpha-tropomyosin). | |||||
|
TPM1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.015111 (rank : 51) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 729 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P58771, P02558, P19354, P46902, P99034 | Gene names | Tpm1, Tpm-1, Tpma | |||
|
Domain Architecture |
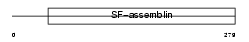
|
|||||
| Description | Tropomyosin 1 alpha chain (Alpha-tropomyosin). | |||||
|
ANPRA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.001126 (rank : 87) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 612 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P16066, Q6P4Q3 | Gene names | NPR1, ANPRA | |||
|
Domain Architecture |
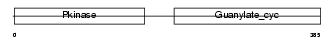
|
|||||
| Description | Atrial natriuretic peptide receptor A precursor (ANP-A) (ANPRA) (GC-A) (Guanylate cyclase) (EC 4.6.1.2) (NPR-A) (Atrial natriuretic peptide A-type receptor). | |||||
|
COG2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.010209 (rank : 73) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q921L5, Q9CSQ1 | Gene names | Cog2, Ldlc | |||
|
Domain Architecture |
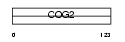
|
|||||
| Description | Conserved oligomeric Golgi complex component 2 (Low density lipoprotein receptor defect C-complementing protein). | |||||
|
HIP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.008504 (rank : 77) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1016 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O00291, O00328, Q8TDL4 | Gene names | HIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-interacting protein 1 (HIP-I). | |||||
|
MAMC2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.010528 (rank : 70) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7Z304, Q5VW47, Q8WX43, Q96BM4 | Gene names | MAMDC2 | |||
|
Domain Architecture |

|
|||||
| Description | MAM domain-containing protein 2 precursor. | |||||
|
MAMC2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.010485 (rank : 71) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CG85, Q8BM24 | Gene names | Mamdc2 | |||
|
Domain Architecture |

|
|||||
| Description | MAM domain-containing protein 2 precursor. | |||||
|
ST18_HUMAN
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.011859 (rank : 65) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O60284 | Gene names | ST18, KIAA0535, ZNF387 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity protein 18 (Zinc finger protein 387). | |||||
|
TRI44_HUMAN
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.006320 (rank : 80) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96DX7, Q96QY2, Q9UGK0 | Gene names | TRIM44, DIPB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 44 (Protein DIPB). | |||||
|
PRKDC_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 92 | |
| SwissProt Accessions | P97313, O88187, P97928, Q307W9, Q3V2W8, Q8C2A7, Q9Z341 | Gene names | Prkdc, Xrcc7 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-dependent protein kinase catalytic subunit (EC 2.7.11.1) (DNA-PK catalytic subunit) (DNA-PKcs) (P460). | |||||
|
PRKDC_HUMAN
|
||||||
| NC score | 0.991668 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P78527, P78528, Q13327, Q13337, Q14175, Q59H99, Q7Z611, Q96SE6, Q9UME3 | Gene names | PRKDC | |||
|
Domain Architecture |

|
|||||
| Description | DNA-dependent protein kinase catalytic subunit (EC 2.7.11.1) (DNA-PK catalytic subunit) (DNA-PKcs) (DNPK1) (p460). | |||||
|
FRAP_MOUSE
|
||||||
| NC score | 0.701658 (rank : 3) | θ value | 3.99248e-40 (rank : 3) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9JLN9, Q811J5, Q9CST1 | Gene names | Frap1, Frap | |||
|
Domain Architecture |

|
|||||
| Description | FKBP12-rapamycin complex-associated protein (FK506-binding protein 12- rapamycin complex-associated protein 1) (Rapamycin target protein) (RAPT1) (Mammalian target of rapamycin) (mTOR). | |||||
|
FRAP_HUMAN
|
||||||
| NC score | 0.701475 (rank : 4) | θ value | 5.21438e-40 (rank : 4) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P42345, Q6LE87, Q96QG3, Q9Y4I3 | Gene names | FRAP1, FRAP, FRAP2 | |||
|
Domain Architecture |

|
|||||
| Description | FKBP12-rapamycin complex-associated protein (FK506-binding protein 12- rapamycin complex-associated protein 1) (Rapamycin target protein) (RAPT1) (Mammalian target of rapamycin) (mTOR). | |||||
|
ATR_HUMAN
|
||||||
| NC score | 0.696522 (rank : 5) | θ value | 1.57e-36 (rank : 5) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q13535, Q7KYL3, Q93051, Q9BXK4 | Gene names | ATR, FRP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ATR (EC 2.7.11.1) (Ataxia telangiectasia and Rad3-related protein) (FRAP-related protein 1). | |||||
|
ATR_MOUSE
|
||||||
| NC score | 0.690161 (rank : 6) | θ value | 4.27553e-34 (rank : 6) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9JKK8 | Gene names | Atr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ATR (EC 2.7.11.1) (Ataxia telangiectasia and Rad3-related protein). | |||||
|
ATM_HUMAN
|
||||||
| NC score | 0.666895 (rank : 7) | θ value | 6.17384e-33 (rank : 7) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q13315, O15429, Q12758, Q16551, Q93007, Q9NP02, Q9UCX7 | Gene names | ATM | |||
|
Domain Architecture |

|
|||||
| Description | Serine-protein kinase ATM (EC 2.7.11.1) (Ataxia telangiectasia mutated) (A-T, mutated). | |||||
|
ATM_MOUSE
|
||||||
| NC score | 0.661853 (rank : 8) | θ value | 3.06405e-32 (rank : 8) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q62388 | Gene names | Atm | |||
|
Domain Architecture |

|
|||||
| Description | Serine-protein kinase ATM (EC 2.7.11.1) (Ataxia telangiectasia mutated homolog) (A-T, mutated homolog). | |||||
|
SMG1_MOUSE
|
||||||
| NC score | 0.630363 (rank : 9) | θ value | 1.98606e-31 (rank : 10) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8BKX6, Q5DU42, Q6ZQC0, Q8BLU4, Q8BWJ5, Q8BXD3 | Gene names | Smg1, Kiaa0421 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SMG1 (EC 2.7.11.1) (SMG-1). | |||||
|
SMG1_HUMAN
|
||||||
| NC score | 0.626354 (rank : 10) | θ value | 1.98606e-31 (rank : 9) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96Q15, O43305, Q13284, Q8NFX2, Q96QV0, Q96RW3 | Gene names | SMG1, ATX, KIAA0421, LIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SMG1 (EC 2.7.11.1) (SMG-1) (hSMG-1) (Lambda/iota protein kinase C-interacting protein) (Lambda-interacting protein) (61E3.4). | |||||
|
PK3C3_MOUSE
|
||||||
| NC score | 0.473717 (rank : 11) | θ value | 5.08577e-11 (rank : 17) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q6PF93, Q8R3S8 | Gene names | Pik3c3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3). | |||||
|
PK3C3_HUMAN
|
||||||
| NC score | 0.472772 (rank : 12) | θ value | 1.133e-10 (rank : 19) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8NEB9, Q15134 | Gene names | PIK3C3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3) (Phosphatidylinositol 3-kinase p100 subunit). | |||||
|
TRRAP_MOUSE
|
||||||
| NC score | 0.448781 (rank : 13) | θ value | 3.63628e-17 (rank : 12) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q80YV3, Q8C0Z5, Q8K104 | Gene names | Trrap | |||
|
Domain Architecture |

|
|||||
| Description | Transformation/transcription domain-associated protein (Tra1 homolog). | |||||
|
TRRAP_HUMAN
|
||||||
| NC score | 0.443930 (rank : 14) | θ value | 1.47631e-18 (rank : 11) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Y4A5, O75218, Q9Y631, Q9Y6H4 | Gene names | TRRAP, PAF400 | |||
|
Domain Architecture |

|
|||||
| Description | Transformation/transcription domain-associated protein (350/400 kDa PCAF-associated factor) (PAF350/400) (STAF40) (Tra1 homolog). | |||||
|
PK3CG_HUMAN
|
||||||
| NC score | 0.432340 (rank : 15) | θ value | 3.18553e-13 (rank : 13) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P48736, Q8IV23, Q9BZC8 | Gene names | PIK3CG | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit gamma isoform (EC 2.7.1.153) (PI3-kinase p110 subunit gamma) (PtdIns-3- kinase subunit p110) (PI3K) (PI3Kgamma) (p120-PI3K). | |||||
|
PK3CD_MOUSE
|
||||||
| NC score | 0.430136 (rank : 16) | θ value | 2.28291e-11 (rank : 15) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O35904 | Gene names | Pik3cd | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit delta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit delta) (PtdIns-3- kinase p110) (PI3K) (p110delta). | |||||
|
PK3CG_MOUSE
|
||||||
| NC score | 0.429758 (rank : 17) | θ value | 9.26847e-13 (rank : 14) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9JHG7 | Gene names | Pik3cg, Pi3kg1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit gamma isoform (EC 2.7.1.153) (PI3-kinase p110 subunit gamma) (PtdIns-3- kinase subunit p110) (PI3K) (PI3Kgamma) (p120-PI3K). | |||||
|
PK3CB_MOUSE
|
||||||
| NC score | 0.429505 (rank : 18) | θ value | 8.67504e-11 (rank : 18) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BTI9 | Gene names | Pik3cb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit beta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit beta) (PtdIns-3-kinase p110) (PI3K) (PI3Kbeta). | |||||
|
PK3CD_HUMAN
|
||||||
| NC score | 0.428892 (rank : 19) | θ value | 2.98157e-11 (rank : 16) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O00329, O15445 | Gene names | PIK3CD | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit delta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit delta) (PtdIns-3- kinase p110) (PI3K) (p110delta). | |||||
|
PK3CB_HUMAN
|
||||||
| NC score | 0.427825 (rank : 20) | θ value | 3.29651e-10 (rank : 20) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P42338 | Gene names | PIK3CB | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit beta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit beta) (PtdIns-3-kinase p110) (PI3K) (PI3Kbeta). | |||||
|
PI4KB_HUMAN
|
||||||
| NC score | 0.394248 (rank : 21) | θ value | 1.43324e-05 (rank : 21) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UBF8, O15096, P78405, Q9BWR6 | Gene names | PIK4CB, PI4KB | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4-kinase beta (EC 2.7.1.67) (PtdIns 4-kinase beta) (PI4Kbeta) (PI4K-beta) (NPIK) (PI4K92). | |||||
|
PI4KB_MOUSE
|
||||||
| NC score | 0.392409 (rank : 22) | θ value | 1.43324e-05 (rank : 22) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8BKC8, Q8C146 | Gene names | Pik4cb | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4-kinase beta (EC 2.7.1.67) (PtdIns 4-kinase beta) (PI4Kbeta) (PI4K-beta). | |||||
|
PK3CA_HUMAN
|
||||||
| NC score | 0.367855 (rank : 23) | θ value | 0.000158464 (rank : 25) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P42336, Q99762 | Gene names | PIK3CA | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (EC 2.7.1.153) (PI3-kinase p110 subunit alpha) (PtdIns-3- kinase p110) (PI3K). | |||||
|
PK3CA_MOUSE
|
||||||
| NC score | 0.367673 (rank : 24) | θ value | 0.000158464 (rank : 26) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P42337 | Gene names | Pik3ca | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (EC 2.7.1.153) (PI3-kinase p110 subunit alpha) (PtdIns-3- kinase p110) (PI3K). | |||||
|
P3C2G_MOUSE
|
||||||
| NC score | 0.349066 (rank : 25) | θ value | 0.00035302 (rank : 27) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O70167, O70168 | Gene names | Pik3c2g | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing gamma polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-gamma) (PtdIns-3-kinase C2 gamma) (PI3K-C2gamma). | |||||
|
P3C2G_HUMAN
|
||||||
| NC score | 0.344490 (rank : 26) | θ value | 0.000158464 (rank : 24) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O75747 | Gene names | PIK3C2G | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing gamma polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-gamma) (PtdIns-3-kinase C2 gamma) (PI3K-C2gamma). | |||||
|
PI4KA_HUMAN
|
||||||
| NC score | 0.319553 (rank : 27) | θ value | 0.47712 (rank : 35) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P42356, Q7Z625, Q9UPG2 | Gene names | PIK4CA, PIK4 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4-kinase alpha (EC 2.7.1.67) (PI4-kinase alpha) (PtdIns-4-kinase alpha) (PI4K-alpha). | |||||
|
P3C2A_MOUSE
|
||||||
| NC score | 0.308168 (rank : 28) | θ value | 0.00134147 (rank : 28) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q61194, Q61182 | Gene names | Pik3c2a, Cpk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing alpha polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-alpha) (PtdIns-3-kinase C2 alpha) (PI3K-C2alpha) (Cpk-m) (p170). | |||||
|
P3C2A_HUMAN
|
||||||
| NC score | 0.304654 (rank : 29) | θ value | 0.00298849 (rank : 29) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O00443 | Gene names | PIK3C2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing alpha polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-alpha) (PtdIns-3-kinase C2 alpha) (PI3K-C2alpha). | |||||
|
P3C2B_HUMAN
|
||||||
| NC score | 0.294297 (rank : 30) | θ value | 0.000121331 (rank : 23) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O00750, O95666, Q5SW99 | Gene names | PIK3C2B | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing beta polypeptide (EC 2.7.1.154) (Phosphoinositide 3-Kinase-C2-beta) (PtdIns-3-kinase C2 beta) (PI3K-C2beta) (C2-PI3K). | |||||
|
CETP_HUMAN
|
||||||
| NC score | 0.069313 (rank : 31) | θ value | 0.125558 (rank : 33) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P11597, Q13987, Q13988 | Gene names | CETP | |||
|
Domain Architecture |

|
|||||
| Description | Cholesteryl ester transfer protein precursor (Lipid transfer protein I). | |||||
|
XPO4_MOUSE
|
||||||
| NC score | 0.056242 (rank : 32) | θ value | 1.06291 (rank : 38) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ESJ0, Q6ZPJ4 | Gene names | Xpo4, Kiaa1721 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Exportin-4 (Exp4). | |||||
|
XPO4_HUMAN
|
||||||
| NC score | 0.056197 (rank : 33) | θ value | 1.06291 (rank : 37) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9C0E2, Q5VUZ5, Q9H934 | Gene names | XPO4, KIAA1721 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Exportin-4 (Exp4). | |||||
|
K0423_MOUSE
|
||||||
| NC score | 0.055328 (rank : 34) | θ value | 0.0563607 (rank : 31) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6A070 | Gene names | Kiaa0423 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0423. | |||||
|
CENPI_HUMAN
|
||||||
| NC score | 0.044465 (rank : 35) | θ value | 1.81305 (rank : 41) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92674, Q5JWZ9, Q96ED0 | Gene names | CENPI, FSHPRH1, ICEN19, LRPR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein I (CENP-I) (Interphase centromere complex protein 19) (Follicle-stimulating hormone primary response protein) (FSH primary response protein 1) (Leucine-rich primary response protein 1). | |||||
|
SIA8A_MOUSE
|
||||||
| NC score | 0.041627 (rank : 36) | θ value | 0.0113563 (rank : 30) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q64687 | Gene names | St8sia1, Siat8, Siat8a | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-N-acetylneuraminide alpha-2,8-sialyltransferase (EC 2.4.99.8) (Ganglioside GD3 synthase) (Ganglioside GT3 synthase) (Alpha-2,8- sialyltransferase 8A) (ST8Sia I). | |||||
|
SIA8A_HUMAN
|
||||||
| NC score | 0.027833 (rank : 37) | θ value | 1.38821 (rank : 39) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92185, Q93064 | Gene names | ST8SIA1, SIAT8, SIAT8A | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-N-acetylneuraminide alpha-2,8-sialyltransferase (EC 2.4.99.8) (Ganglioside GD3 synthase) (Ganglioside GT3 synthase) (Alpha-2,8- sialyltransferase 8A) (ST8Sia I). | |||||
|
KMHN1_HUMAN
|
||||||
| NC score | 0.027265 (rank : 38) | θ value | 0.0961366 (rank : 32) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 950 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8TBZ0, Q86YI9, Q8N7W0 | Gene names | KMHN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cancer/testis antigen KM-HN-1. | |||||
|
SMC4_HUMAN
|
||||||
| NC score | 0.023115 (rank : 39) | θ value | 1.06291 (rank : 36) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NTJ3, O95752, Q8NDL4, Q9UNT9 | Gene names | SMC4L1, CAPC, SMC4 | |||
|
Domain Architecture |
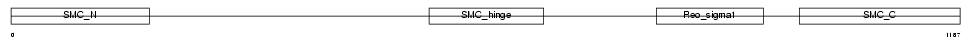
|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (hCAP-C) (XCAP-C homolog). | |||||
|
K0776_MOUSE
|
||||||
| NC score | 0.020805 (rank : 40) | θ value | 2.36792 (rank : 49) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CCJ3, Q3V145, Q6ZQ50, Q8BT70, Q8C484, Q9D8I8 | Gene names | Kiaa0776 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0776. | |||||
|
SPAG5_MOUSE
|
||||||
| NC score | 0.019839 (rank : 41) | θ value | 1.81305 (rank : 46) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 612 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q7TME2, Q8CH61 | Gene names | Spag5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 5 (Mastrin) (Mitotic spindle-associated protein p126) (MAP126). | |||||
|
CCD46_HUMAN
|
||||||
| NC score | 0.019243 (rank : 42) | θ value | 0.21417 (rank : 34) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1430 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8N8E3, Q6PIB5, Q8NCR4, Q8NFR4 | Gene names | CCDC46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 46. | |||||
|
SYNE2_HUMAN
|
||||||
| NC score | 0.019075 (rank : 43) | θ value | 1.38821 (rank : 40) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1323 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8WXH0, Q8N1S3, Q8NF49, Q8TER7, Q8WWW3, Q8WWW4, Q8WWW5, Q8WXH1, Q9NU50, Q9UFQ4, Q9Y2L4, Q9Y4R1 | Gene names | SYNE2, KIAA1011, NUA | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-2 (Nuclear envelope spectrin repeat protein 2) (Syne-2) (Synaptic nuclear envelope protein 2) (Nucleus and actin connecting element protein) (Protein NUANCE). | |||||
|
XPOT_HUMAN
|
||||||
| NC score | 0.018125 (rank : 44) | θ value | 5.27518 (rank : 74) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O43592, O43784, Q8WUG2, Q9BVS7 | Gene names | XPOT | |||
|
Domain Architecture |

|
|||||
| Description | Exportin-T (tRNA exportin) (Exportin(tRNA)). | |||||
|
TPM2_MOUSE
|
||||||
| NC score | 0.017133 (rank : 45) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P58774, P02560, P46901 | Gene names | Tpm2, Tpm-2 | |||
|
Domain Architecture |

|
|||||
| Description | Tropomyosin beta chain (Tropomyosin 2) (Beta-tropomyosin). | |||||
|
TPM2_HUMAN
|
||||||
| NC score | 0.016644 (rank : 46) | θ value | 3.0926 (rank : 57) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 677 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P07951, P06468, Q13894, Q53FM4, Q5TCU4, Q5TCU7, Q9UH67 | Gene names | TPM2, TMSB | |||
|
Domain Architecture |

|
|||||
| Description | Tropomyosin beta chain (Tropomyosin 2) (Beta-tropomyosin). | |||||
|
SLC2B_HUMAN
|
||||||
| NC score | 0.015648 (rank : 47) | θ value | 5.27518 (rank : 71) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NEV8, Q9Y4D6 | Gene names | EXPH5, KIAA0624, SLAC2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Slp homolog lacking C2 domains b (Exophilin-5). | |||||
|
CE290_HUMAN
|
||||||
| NC score | 0.015252 (rank : 48) | θ value | 2.36792 (rank : 47) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
SMC4_MOUSE
|
||||||
| NC score | 0.015207 (rank : 49) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 830 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8CG47, Q8BTS7, Q8BTY9, Q99K21 | Gene names | Smc4l1, Capc, Smc4 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (XCAP-C homolog). | |||||
|
TPM1_HUMAN
|
||||||
| NC score | 0.015207 (rank : 50) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 716 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P09493, P09494, P10469, Q86W64, Q96IK2, Q9UCY9 | Gene names | TPM1, TMSA | |||
|
Domain Architecture |

|
|||||
| Description | Tropomyosin 1 alpha chain (Alpha-tropomyosin). | |||||
|
TPM1_MOUSE
|
||||||
| NC score | 0.015111 (rank : 51) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 729 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P58771, P02558, P19354, P46902, P99034 | Gene names | Tpm1, Tpm-1, Tpma | |||
|
Domain Architecture |

|
|||||
| Description | Tropomyosin 1 alpha chain (Alpha-tropomyosin). | |||||
|
HMMR_HUMAN
|
||||||
| NC score | 0.014321 (rank : 52) | θ value | 2.36792 (rank : 48) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1082 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O75330, Q92767 | Gene names | HMMR, IHABP, RHAMM | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronan mediated motility receptor (Intracellular hyaluronic acid- binding protein) (Receptor for hyaluronan-mediated motility) (CD168 antigen). | |||||
|
CAB45_HUMAN
|
||||||
| NC score | 0.013516 (rank : 53) | θ value | 4.03905 (rank : 61) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BRK5, Q8NBQ3, Q9NZP7 | Gene names | SDF4, CAB45 | |||
|
Domain Architecture |

|
|||||
| Description | 45 kDa calcium-binding protein precursor (Cab45) (Stromal cell-derived factor 4) (SDF-4). | |||||
|
ST18_MOUSE
|
||||||
| NC score | 0.013213 (rank : 54) | θ value | 5.27518 (rank : 73) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80TY4, Q3UH00, Q3URH9, Q3UVB9, Q3UZN9, Q811B4, Q8K098 | Gene names | St18, Kiaa0535 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity protein 18. | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.013045 (rank : 55) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
GCC2_MOUSE
|
||||||
| NC score | 0.012995 (rank : 56) | θ value | 1.81305 (rank : 43) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1619 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8CHG3, Q8BR44, Q8R2Q5, Q9CT45 | Gene names | Gcc2, Kiaa0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185). | |||||
|
S35A3_MOUSE
|
||||||
| NC score | 0.012892 (rank : 57) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R1T4, Q8BRW7 | Gene names | Slc35a3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UDP-N-acetylglucosamine transporter (Golgi UDP-GlcNAc transporter) (Solute carrier family 35 member A3). | |||||
|
S35A3_HUMAN
|
||||||
| NC score | 0.012837 (rank : 58) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y2D2, Q68CR2 | Gene names | SLC35A3 | |||
|
Domain Architecture |
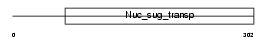
|
|||||
| Description | UDP-N-acetylglucosamine transporter (Golgi UDP-GlcNAc transporter) (Solute carrier family 35 member A3). | |||||
|
GOGB1_HUMAN
|
||||||
| NC score | 0.012587 (rank : 59) | θ value | 1.81305 (rank : 44) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
MYH6_HUMAN
|
||||||
| NC score | 0.012475 (rank : 60) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
RPGF2_HUMAN
|
||||||
| NC score | 0.012213 (rank : 61) | θ value | 4.03905 (rank : 65) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y4G8 | Gene names | RAPGEF2, KIAA0313, NRAPGEP, PDZGEF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rap guanine nucleotide exchange factor 2 (Neural RAP guanine nucleotide exchange protein) (nRap GEP) (PDZ domain-containing guanine nucleotide exchange factor 1) (PDZ-GEF1) (RA-GEF). | |||||
|
M6PBP_MOUSE
|
||||||
| NC score | 0.012162 (rank : 62) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9DBG5, Q3TK05, Q8BKV9, Q9CZK1 | Gene names | M6prbp1, Tip47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mannose-6-phosphate receptor-binding protein 1 (Cargo selection protein TIP47). | |||||
|
ITPR1_HUMAN
|
||||||
| NC score | 0.012160 (rank : 63) | θ value | 1.81305 (rank : 45) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q14643, Q14660, Q99897 | Gene names | ITPR1, INSP3R1 | |||
|
Domain Architecture |

|
|||||
| Description | Inositol 1,4,5-trisphosphate receptor type 1 (Type 1 inositol 1,4,5- trisphosphate receptor) (Type 1 InsP3 receptor) (IP3 receptor isoform 1) (InsP3R1) (IP3R). | |||||
|
SORC3_HUMAN
|
||||||
| NC score | 0.011946 (rank : 64) | θ value | 4.03905 (rank : 66) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UPU3, Q5VXF9, Q9NQJ2 | Gene names | SORCS3, KIAA1059 | |||
|
Domain Architecture |

|
|||||
| Description | VPS10 domain-containing receptor SorCS3 precursor. | |||||
|
ST18_HUMAN
|
||||||
| NC score | 0.011859 (rank : 65) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O60284 | Gene names | ST18, KIAA0535, ZNF387 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity protein 18 (Zinc finger protein 387). | |||||
|
CP135_MOUSE
|
||||||
| NC score | 0.011699 (rank : 66) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1278 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6P5D4, Q6A030 | Gene names | Cep135, Cep4, Kiaa0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
SORC3_MOUSE
|
||||||
| NC score | 0.011322 (rank : 67) | θ value | 5.27518 (rank : 72) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VI51, Q9CTR4 | Gene names | Sorcs3 | |||
|
Domain Architecture |

|
|||||
| Description | VPS10 domain-containing receptor SorCS3 precursor. | |||||
|
FMNL_HUMAN
|
||||||
| NC score | 0.011040 (rank : 68) | θ value | 3.0926 (rank : 54) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 529 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O95466, Q6DKG5, Q6IBP3, Q86UH1, Q8N671, Q8TDH1, Q96H10 | Gene names | FMNL1, C17orf1, C17orf1B, FMNL | |||
|
Domain Architecture |

|
|||||
| Description | Formin-like protein 1 (Leukocyte formin) (CLL-associated antigen KW- 13). | |||||
|
CALX_HUMAN
|
||||||
| NC score | 0.010679 (rank : 69) | θ value | 4.03905 (rank : 62) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P27824 | Gene names | CANX | |||
|
Domain Architecture |

|
|||||
| Description | Calnexin precursor (Major histocompatibility complex class I antigen- binding protein p88) (p90) (IP90). | |||||
|
MAMC2_HUMAN
|
||||||
| NC score | 0.010528 (rank : 70) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7Z304, Q5VW47, Q8WX43, Q96BM4 | Gene names | MAMDC2 | |||
|
Domain Architecture |

|
|||||
| Description | MAM domain-containing protein 2 precursor. | |||||
|
MAMC2_MOUSE
|
||||||
| NC score | 0.010485 (rank : 71) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CG85, Q8BM24 | Gene names | Mamdc2 | |||
|
Domain Architecture |

|
|||||
| Description | MAM domain-containing protein 2 precursor. | |||||
|
MYO1B_MOUSE
|
||||||
| NC score | 0.010234 (rank : 72) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P46735, P70244 | Gene names | Myo1b | |||
|
Domain Architecture |

|
|||||
| Description | Myosin Ib (Myosin I alpha) (MMI-alpha) (MMIa) (MIH-L). | |||||
|
COG2_MOUSE
|
||||||
| NC score | 0.010209 (rank : 73) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q921L5, Q9CSQ1 | Gene names | Cog2, Ldlc | |||
|
Domain Architecture |

|
|||||
| Description | Conserved oligomeric Golgi complex component 2 (Low density lipoprotein receptor defect C-complementing protein). | |||||
|
PON3_MOUSE
|
||||||
| NC score | 0.010013 (rank : 74) | θ value | 5.27518 (rank : 70) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q62087, Q8C2I7 | Gene names | Pon3 | |||
|
Domain Architecture |

|
|||||
| Description | Serum paraoxonase/lactonase 3 (EC 3.1.1.-). | |||||
|
RPGF6_HUMAN
|
||||||
| NC score | 0.009949 (rank : 75) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TEU7, Q8NI21, Q8TEU6, Q96PC1 | Gene names | RAPGEF6, PDZGEF2 | |||
|
Domain Architecture |

|
|||||
| Description | Rap guanine nucleotide exchange factor 6 (PDZ domain-containing guanine nucleotide exchange factor 2) (PDZ-GEF2) (RA-GEF-2). | |||||
|
DHX36_HUMAN
|
||||||
| NC score | 0.009462 (rank : 76) | θ value | 1.81305 (rank : 42) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H2U1, Q70JU3, Q8IYE5, Q9P240 | Gene names | DHX36, DDX36, KIAA1488, MLEL1, RHAU | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DHX36 (EC 3.6.1.-) (DEAH box protein 36) (MLE-like protein 1) (RNA helicase associated with AU-rich element ARE). | |||||
|
HIP1_HUMAN
|
||||||
| NC score | 0.008504 (rank : 77) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1016 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O00291, O00328, Q8TDL4 | Gene names | HIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-interacting protein 1 (HIP-I). | |||||
|
GDF9_HUMAN
|
||||||
| NC score | 0.007837 (rank : 78) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O60383 | Gene names | GDF9 | |||
|
Domain Architecture |

|
|||||
| Description | Growth/differentiation factor 9 precursor (GDF-9). | |||||
|
SVIL_HUMAN
|
||||||
| NC score | 0.007613 (rank : 79) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O95425, O60611, O60612, Q5VZK5, Q5VZK6, Q9H1R7 | Gene names | SVIL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Supervillin (Archvillin) (p205/p250). | |||||
|
TRI44_HUMAN
|
||||||
| NC score | 0.006320 (rank : 80) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96DX7, Q96QY2, Q9UGK0 | Gene names | TRIM44, DIPB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 44 (Protein DIPB). | |||||
|
SPA12_HUMAN
|
||||||
| NC score | 0.005773 (rank : 81) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IW75 | Gene names | SERPINA12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serpin A12 precursor (Visceral adipose-specific serpin) (Visceral adipose tissue-derived serine protease inhibitor) (Vaspin) (OL-64). | |||||
|
K1HA_HUMAN
|
||||||
| NC score | 0.005297 (rank : 82) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 502 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O76009 | Gene names | KRTHA3A, HHA3-I, HKA3A | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cuticular Ha3-I (Hair keratin, type I Ha3-I). | |||||
|
K1H1_HUMAN
|
||||||
| NC score | 0.005277 (rank : 83) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15323 | Gene names | KRTHA1, HHA1, HKA1 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cuticular Ha1 (Hair keratin, type I Ha1). | |||||
|
KIF2B_HUMAN
|
||||||
| NC score | 0.005110 (rank : 84) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N4N8, Q96MA2, Q9BXG6 | Gene names | KIF2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin-like protein KIF2B. | |||||
|
PAI2_MOUSE
|
||||||
| NC score | 0.004333 (rank : 85) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P12388, O35687, Q9QWP6, Q9QWP7, Q9QWP8, Q9QWP9, Q9QWQ0, Q9QWZ5, Q9QWZ6 | Gene names | Serpinb2, Pai2, Planh2 | |||
|
Domain Architecture |

|
|||||
| Description | Plasminogen activator inhibitor 2, macrophage (PAI-2). | |||||
|
BMX_HUMAN
|
||||||
| NC score | 0.001140 (rank : 86) | θ value | 3.0926 (rank : 52) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 942 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P51813, O60564, Q12871 | Gene names | BMX | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic tyrosine-protein kinase BMX (EC 2.7.10.2) (Bone marrow tyrosine kinase gene in chromosome X protein) (Epithelial and endothelial tyrosine kinase) (ETK) (NTK38). | |||||
|
ANPRA_HUMAN
|
||||||
| NC score | 0.001126 (rank : 87) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 612 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P16066, Q6P4Q3 | Gene names | NPR1, ANPRA | |||
|
Domain Architecture |

|
|||||
| Description | Atrial natriuretic peptide receptor A precursor (ANP-A) (ANPRA) (GC-A) (Guanylate cyclase) (EC 4.6.1.2) (NPR-A) (Atrial natriuretic peptide A-type receptor). | |||||
|
BTK_HUMAN
|
||||||
| NC score | 0.000650 (rank : 88) | θ value | 3.0926 (rank : 53) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1026 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q06187 | Gene names | BTK, AGMX1, ATK, BPK | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase BTK (EC 2.7.10.2) (Bruton tyrosine kinase) (Agammaglobulinaemia tyrosine kinase) (ATK) (B cell progenitor kinase) (BPK). | |||||
|
BTK_MOUSE
|
||||||
| NC score | 0.000486 (rank : 89) | θ value | 4.03905 (rank : 60) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1017 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P35991, Q61365 | Gene names | Btk, Bpk | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase BTK (EC 2.7.10.2) (Bruton tyrosine kinase) (Agammaglobulinaemia tyrosine kinase) (ATK) (B cell progenitor kinase) (BPK) (Kinase EMB). | |||||
|
CD2L7_HUMAN
|
||||||
| NC score | -0.000229 (rank : 90) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1275 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9NYV4, O94978 | Gene names | CRKRS, CRK7, KIAA0904 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle 2-related protein kinase 7 (EC 2.7.11.22) (CDC2- related protein kinase 7) (Cdc2-related kinase, arginine/serine-rich) (CrkRS). | |||||
|
NOTC2_MOUSE
|
||||||
| NC score | -0.001088 (rank : 91) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1069 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O35516, Q06008, Q60941 | Gene names | Notch2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 2 precursor (Notch 2) (Motch B) [Contains: Notch 2 extracellular truncation; Notch 2 intracellular domain]. | |||||
|
ZN567_HUMAN
|
||||||
| NC score | -0.002469 (rank : 92) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 769 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N184, Q6N044 | Gene names | ZNF567 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 567. | |||||