Please be patient as the page loads
|
UBP43_MOUSE
|
||||||
| SwissProt Accessions | Q8BUM9, Q8VDP5 | Gene names | Usp43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 43 (EC 3.1.2.15) (Ubiquitin thioesterase 43) (Ubiquitin-specific-processing protease 43) (Deubiquitinating enzyme 43). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
UBP31_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.944435 (rank : 3) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q70CQ4, Q6AW97, Q6ZTC0, Q6ZTN2, Q9ULL7 | Gene names | USP31, KIAA1203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 31 (EC 3.1.2.15) (Ubiquitin thioesterase 31) (Ubiquitin-specific-processing protease 31) (Deubiquitinating enzyme 31). | |||||
|
UBP43_HUMAN
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.988926 (rank : 2) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q70EL4, Q8N2C5, Q96DQ6 | Gene names | USP43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 43 (EC 3.1.2.15) (Ubiquitin thioesterase 43) (Ubiquitin-specific-processing protease 43) (Deubiquitinating enzyme 43). | |||||
|
UBP43_MOUSE
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 130 | |
| SwissProt Accessions | Q8BUM9, Q8VDP5 | Gene names | Usp43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 43 (EC 3.1.2.15) (Ubiquitin thioesterase 43) (Ubiquitin-specific-processing protease 43) (Deubiquitinating enzyme 43). | |||||
|
UBP4_HUMAN
|
||||||
| θ value | 1.16164e-39 (rank : 4) | NC score | 0.856885 (rank : 4) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q13107, O43452, O43453 | Gene names | USP4, UNP, UNPH | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 4 (EC 3.1.2.15) (Ubiquitin thioesterase 4) (Ubiquitin-specific-processing protease 4) (Deubiquitinating enzyme 4) (Ubiquitous nuclear protein homolog). | |||||
|
UBP4_MOUSE
|
||||||
| θ value | 9.83387e-39 (rank : 5) | NC score | 0.855620 (rank : 6) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P35123, O54704 | Gene names | Usp4, Unp | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 4 (EC 3.1.2.15) (Ubiquitin thioesterase 4) (Ubiquitin-specific-processing protease 4) (Deubiquitinating enzyme 4) (Ubiquitous nuclear protein). | |||||
|
UBP15_MOUSE
|
||||||
| θ value | 3.73686e-38 (rank : 6) | NC score | 0.854588 (rank : 7) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q8R5H1, Q3UL25 | Gene names | Usp15 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 15 (EC 3.1.2.15) (Ubiquitin thioesterase 15) (Ubiquitin-specific-processing protease 15) (Deubiquitinating enzyme 15). | |||||
|
UBP15_HUMAN
|
||||||
| θ value | 8.32485e-38 (rank : 7) | NC score | 0.854120 (rank : 8) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9Y4E8, Q9HCA6, Q9UNP0, Q9Y5B5 | Gene names | USP15, KIAA0529 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 15 (EC 3.1.2.15) (Ubiquitin thioesterase 15) (Ubiquitin-specific-processing protease 15) (Deubiquitinating enzyme 15) (Unph-2) (Unph4). | |||||
|
UBP11_HUMAN
|
||||||
| θ value | 2.34606e-32 (rank : 8) | NC score | 0.855853 (rank : 5) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P51784, Q8IUG6, Q9BWE1 | Gene names | USP11, UHX1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 11 (EC 3.1.2.15) (Ubiquitin thioesterase 11) (Ubiquitin-specific-processing protease 11) (Deubiquitinating enzyme 11). | |||||
|
UBP8_MOUSE
|
||||||
| θ value | 2.19584e-30 (rank : 9) | NC score | 0.835821 (rank : 12) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q80U87, Q80YP2, Q8R0D3, Q9EQU1, Q9WVP5 | Gene names | Usp8, Kiaa0055, Ubpy | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 8 (EC 3.1.2.15) (Ubiquitin thioesterase 8) (Ubiquitin-specific-processing protease 8) (Deubiquitinating enzyme 8) (mUBPy). | |||||
|
UBP8_HUMAN
|
||||||
| θ value | 4.89182e-30 (rank : 10) | NC score | 0.796249 (rank : 23) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 751 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | P40818, Q7Z3U2, Q86VA0, Q8IWI7 | Gene names | USP8, KIAA0055, UBPY | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 8 (EC 3.1.2.15) (Ubiquitin thioesterase 8) (Ubiquitin-specific-processing protease 8) (Deubiquitinating enzyme 8) (hUBPy). | |||||
|
UBP2_MOUSE
|
||||||
| θ value | 6.61148e-27 (rank : 11) | NC score | 0.848909 (rank : 9) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | O88623, Q6X4T9, Q6X4U1, Q8VD74, Q9JJ87 | Gene names | Usp2, Ubp41 | |||
|
Domain Architecture |
|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 2 (EC 3.1.2.15) (Ubiquitin thioesterase 2) (Ubiquitin-specific-processing protease 2) (Deubiquitinating enzyme 2) (41 kDa ubiquitin-specific protease). | |||||
|
UBP2_HUMAN
|
||||||
| θ value | 3.28125e-26 (rank : 12) | NC score | 0.847094 (rank : 10) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | O75604, Q96MB9, Q9BQ21 | Gene names | USP2, UBP41 | |||
|
Domain Architecture |
|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 2 (EC 3.1.2.15) (Ubiquitin thioesterase 2) (Ubiquitin-specific-processing protease 2) (Deubiquitinating enzyme 2) (41 kDa ubiquitin-specific protease). | |||||
|
UBP33_HUMAN
|
||||||
| θ value | 7.30988e-26 (rank : 13) | NC score | 0.826362 (rank : 19) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q8TEY7, Q8TEY6, Q96AV6, Q9H9F0, Q9UPQ5 | Gene names | USP33, KIAA1097, VDU1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 33 (EC 3.1.2.15) (Ubiquitin thioesterase 33) (Ubiquitin-specific-processing protease 33) (Deubiquitinating enzyme 33) (VHL-interacting deubiquitinating enzyme 1). | |||||
|
UBP50_MOUSE
|
||||||
| θ value | 2.12685e-25 (rank : 14) | NC score | 0.845316 (rank : 11) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q6P8X6, Q9D558, Q9D9E7 | Gene names | Usp50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ubiquitin carboxyl-terminal hydrolase 50 (EC 3.1.2.15) (Ubiquitin thioesterase 50) (Ubiquitin-specific-processing protease 50) (Deubiquitinating enzyme 50). | |||||
|
UBP20_HUMAN
|
||||||
| θ value | 3.62785e-25 (rank : 15) | NC score | 0.829121 (rank : 16) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9Y2K6, Q96LG5, Q9UQN8, Q9UQP0 | Gene names | USP20, KIAA1003, LSFR3A | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 20 (EC 3.1.2.15) (Ubiquitin thioesterase 20) (Ubiquitin-specific-processing protease 20) (Deubiquitinating enzyme 20). | |||||
|
UBP51_HUMAN
|
||||||
| θ value | 3.07116e-24 (rank : 16) | NC score | 0.814680 (rank : 20) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q70EK9, Q8IWJ8 | Gene names | USP51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 51 (EC 3.1.2.15) (Ubiquitin thioesterase 51) (Ubiquitin-specific-processing protease 51) (Deubiquitinating enzyme 51). | |||||
|
UBP3_MOUSE
|
||||||
| θ value | 8.93572e-24 (rank : 17) | NC score | 0.828976 (rank : 17) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q91W36 | Gene names | Usp3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 3 (EC 3.1.2.15) (Ubiquitin thioesterase 3) (Ubiquitin-specific-processing protease 3) (Deubiquitinating enzyme 3). | |||||
|
UBP3_HUMAN
|
||||||
| θ value | 1.99067e-23 (rank : 18) | NC score | 0.828783 (rank : 18) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9Y6I4 | Gene names | USP3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 3 (EC 3.1.2.15) (Ubiquitin thioesterase 3) (Ubiquitin-specific-processing protease 3) (Deubiquitinating enzyme 3). | |||||
|
UBP6_HUMAN
|
||||||
| θ value | 5.79196e-23 (rank : 19) | NC score | 0.703962 (rank : 37) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P35125, Q15634, Q86WP6, Q8IWT4 | Gene names | USP6, TRE2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 6 (EC 3.1.2.15) (Ubiquitin thioesterase 6) (Ubiquitin-specific-processing protease 6) (Deubiquitinating enzyme 6) (Proto-oncogene TRE-2). | |||||
|
UBP32_HUMAN
|
||||||
| θ value | 7.56453e-23 (rank : 20) | NC score | 0.746949 (rank : 29) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q8NFA0, Q9BX85, Q9Y591 | Gene names | USP32, USP10 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 32 (EC 3.1.2.15) (Ubiquitin thioesterase 32) (Ubiquitin-specific-processing protease 32) (Deubiquitinating enzyme 32) (NY-REN-60 antigen). | |||||
|
UBP22_MOUSE
|
||||||
| θ value | 1.86321e-21 (rank : 21) | NC score | 0.808811 (rank : 21) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q5DU02, Q5SU81, Q66JV8 | Gene names | Usp22, Kiaa1063 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 22 (EC 3.1.2.15) (Ubiquitin thioesterase 22) (Ubiquitin-specific-processing protease 22) (Deubiquitinating enzyme 22). | |||||
|
UBP22_HUMAN
|
||||||
| θ value | 5.42112e-21 (rank : 22) | NC score | 0.807571 (rank : 22) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9UPT9 | Gene names | USP22, KIAA1063, USP3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 22 (EC 3.1.2.15) (Ubiquitin thioesterase 22) (Ubiquitin-specific-processing protease 22) (Deubiquitinating enzyme 22). | |||||
|
UBP21_HUMAN
|
||||||
| θ value | 3.51386e-20 (rank : 23) | NC score | 0.829687 (rank : 15) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q9UK80, Q9BTV1, Q9NYN4 | Gene names | USP21, USP23 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 21 (EC 3.1.2.15) (Ubiquitin thioesterase 21) (Ubiquitin-specific-processing protease 21) (Deubiquitinating enzyme 21) (NEDD8-specific protease). | |||||
|
UBP21_MOUSE
|
||||||
| θ value | 4.58923e-20 (rank : 24) | NC score | 0.830715 (rank : 13) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q9QZL6, Q9D0R1 | Gene names | Usp21, Usp23 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 21 (EC 3.1.2.15) (Ubiquitin thioesterase 21) (Ubiquitin-specific-processing protease 21) (Deubiquitinating enzyme 21). | |||||
|
UBP50_HUMAN
|
||||||
| θ value | 2.35696e-16 (rank : 25) | NC score | 0.830658 (rank : 14) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q70EL3 | Gene names | USP50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inactive ubiquitin carboxyl-terminal hydrolase 50 (Inactive ubiquitin- specific peptidase 50). | |||||
|
UBP44_HUMAN
|
||||||
| θ value | 2.60593e-15 (rank : 26) | NC score | 0.766407 (rank : 25) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9H0E7 | Gene names | USP44 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 44 (EC 3.1.2.15) (Ubiquitin thioesterase 44) (Ubiquitin-specific-processing protease 44) (Deubiquitinating enzyme 44). | |||||
|
UBP49_MOUSE
|
||||||
| θ value | 4.91457e-14 (rank : 27) | NC score | 0.753894 (rank : 26) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q6P9L4 | Gene names | Usp49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 49 (EC 3.1.2.15) (Ubiquitin thioesterase 49) (Ubiquitin-specific-processing protease 49) (Deubiquitinating enzyme 49). | |||||
|
UBP49_HUMAN
|
||||||
| θ value | 8.38298e-14 (rank : 28) | NC score | 0.751469 (rank : 27) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q70CQ1, Q96CK4 | Gene names | USP49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 49 (EC 3.1.2.15) (Ubiquitin thioesterase 49) (Ubiquitin-specific-processing protease 49) (Deubiquitinating enzyme 49). | |||||
|
UBP48_MOUSE
|
||||||
| θ value | 6.00763e-12 (rank : 29) | NC score | 0.690530 (rank : 40) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q3V0C5, Q3TM39, Q571J9, Q8VDJ1 | Gene names | Usp48, Kiaa4202 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 48 (EC 3.1.2.15) (Ubiquitin thioesterase 48) (Ubiquitin-specific-processing protease 48) (Deubiquitinating enzyme 48). | |||||
|
UBP48_HUMAN
|
||||||
| θ value | 1.33837e-11 (rank : 30) | NC score | 0.691561 (rank : 39) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q86UV5, Q2M3I4, Q5SZI4, Q5T3T5, Q6NX53, Q8N3F6, Q96F64, Q96IQ3, Q9H5N3, Q9H5T7, Q9NUJ6, Q9NXR0 | Gene names | USP48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 48 (EC 3.1.2.15) (Ubiquitin thioesterase 48) (Ubiquitin-specific-processing protease 48) (Deubiquitinating enzyme 48). | |||||
|
UBP16_HUMAN
|
||||||
| θ value | 1.74796e-11 (rank : 31) | NC score | 0.778304 (rank : 24) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9Y5T5, Q8NEL3 | Gene names | USP16 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 16 (EC 3.1.2.15) (Ubiquitin thioesterase 16) (Ubiquitin-specific-processing protease 16) (Deubiquitinating enzyme 16) (Ubiquitin-processing protease UBP-M). | |||||
|
UBP18_HUMAN
|
||||||
| θ value | 1.47974e-10 (rank : 32) | NC score | 0.740164 (rank : 30) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9UMW8, Q9NY71 | Gene names | USP18, ISG43 | |||
|
Domain Architecture |
|
|||||
| Description | Ubl carboxyl-terminal hydrolase 18 (EC 3.1.2.-) (Ubl thioesterase 18) (ISG15-specific-processing protease) (43 kDa ISG15-specific protease) (hUBP43). | |||||
|
UB17L_HUMAN
|
||||||
| θ value | 2.52405e-10 (rank : 33) | NC score | 0.731769 (rank : 34) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q7RTZ2 | Gene names | USP17L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 17-like protein (EC 3.1.2.15) (Ubiquitin thioesterase 17-like) (Ubiquitin-specific-processing protease 17-like) (Deubiquitinating enzyme 17-like). | |||||
|
UBPW_MOUSE
|
||||||
| θ value | 3.29651e-10 (rank : 34) | NC score | 0.751262 (rank : 28) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q61068 | Gene names | Dub1, Dub-1 | |||
|
Domain Architecture |
|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase DUB-1 (EC 3.1.2.15) (Ubiquitin thioesterase DUB-1) (Ubiquitin-specific-processing protease DUB-1) (Deubiquitinating enzyme 1). | |||||
|
UBP18_MOUSE
|
||||||
| θ value | 5.62301e-10 (rank : 35) | NC score | 0.726491 (rank : 35) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9WTV6 | Gene names | Usp18, Ubp43 | |||
|
Domain Architecture |
|
|||||
| Description | Ubl carboxyl-terminal hydrolase 18 (EC 3.1.2.-) (Ubl thioesterase 18) (ISG15-specific-processing protease) (43 kDa ISG15-specific protease). | |||||
|
UBP46_HUMAN
|
||||||
| θ value | 1.25267e-09 (rank : 36) | NC score | 0.732955 (rank : 32) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P62068, Q80V95, Q9H7U4, Q9H9T8 | Gene names | USP46 | |||
|
Domain Architecture |
|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 46 (EC 3.1.2.15) (Ubiquitin thioesterase 46) (Ubiquitin-specific-processing protease 46) (Deubiquitinating enzyme 46). | |||||
|
UBP46_MOUSE
|
||||||
| θ value | 1.25267e-09 (rank : 37) | NC score | 0.732955 (rank : 33) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P62069, Q80V95, Q9H7U4, Q9H9T8 | Gene names | Usp46 | |||
|
Domain Architecture |
|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 46 (EC 3.1.2.15) (Ubiquitin thioesterase 46) (Ubiquitin-specific-processing protease 46) (Deubiquitinating enzyme 46). | |||||
|
UBP12_HUMAN
|
||||||
| θ value | 2.79066e-09 (rank : 38) | NC score | 0.736269 (rank : 31) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | O75317 | Gene names | USP12, UBH1, USP12L1 | |||
|
Domain Architecture |
|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 12 (EC 3.1.2.15) (Ubiquitin thioesterase 12) (Ubiquitin-specific-processing protease 12) (Deubiquitinating enzyme 12) (Ubiquitin-hydrolyzing enzyme 1). | |||||
|
UBP37_HUMAN
|
||||||
| θ value | 2.79066e-09 (rank : 39) | NC score | 0.613506 (rank : 47) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q86T82, Q9HCH8 | Gene names | USP37, KIAA1594 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 37 (EC 3.1.2.15) (Ubiquitin thioesterase 37) (Ubiquitin-specific-processing protease 37) (Deubiquitinating enzyme 37). | |||||
|
UBP42_HUMAN
|
||||||
| θ value | 8.11959e-09 (rank : 40) | NC score | 0.698324 (rank : 38) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q9H9J4, Q3C166, Q6P9B4 | Gene names | USP42 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 42 (EC 3.1.2.15) (Ubiquitin thioesterase 42) (Ubiquitin-specific-processing protease 42) (Deubiquitinating enzyme 42). | |||||
|
UBP24_HUMAN
|
||||||
| θ value | 1.80886e-08 (rank : 41) | NC score | 0.618603 (rank : 44) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9UPU5, Q6ZSY2, Q8N2Y4, Q9NXD1 | Gene names | USP24, KIAA1057 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 24 (EC 3.1.2.15) (Ubiquitin thioesterase 24) (Ubiquitin-specific-processing protease 24) (Deubiquitinating enzyme 24). | |||||
|
UBP36_HUMAN
|
||||||
| θ value | 5.26297e-08 (rank : 42) | NC score | 0.715724 (rank : 36) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q9P275, Q8NDM8, Q9NVC8 | Gene names | USP36, KIAA1453 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 36 (EC 3.1.2.15) (Ubiquitin thioesterase 36) (Ubiquitin-specific-processing protease 36) (Deubiquitinating enzyme 36). | |||||
|
UBP38_HUMAN
|
||||||
| θ value | 8.97725e-08 (rank : 43) | NC score | 0.605385 (rank : 48) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q8NB14, Q8NDF5, Q96DK6, Q96PZ6, Q9BY55 | Gene names | USP38, KIAA1891 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 38 (EC 3.1.2.15) (Ubiquitin thioesterase 38) (Ubiquitin-specific-processing protease 38) (Deubiquitinating enzyme 38) (HP43.8KD). | |||||
|
UBP29_HUMAN
|
||||||
| θ value | 3.41135e-07 (rank : 44) | NC score | 0.549655 (rank : 57) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9HBJ7 | Gene names | USP29 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 29 (EC 3.1.2.15) (Ubiquitin thioesterase 29) (Ubiquitin-specific-processing protease 29) (Deubiquitinating enzyme 29). | |||||
|
UBP26_HUMAN
|
||||||
| θ value | 5.81887e-07 (rank : 45) | NC score | 0.540954 (rank : 63) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9BXU7 | Gene names | USP26 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 26 (EC 3.1.2.15) (Ubiquitin thioesterase 26) (Ubiquitin-specific-processing protease 26) (Deubiquitinating enzyme 26). | |||||
|
UBP38_MOUSE
|
||||||
| θ value | 5.81887e-07 (rank : 46) | NC score | 0.604101 (rank : 49) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q8BW70, Q8BWL1, Q8BX03 | Gene names | Usp38 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 38 (EC 3.1.2.15) (Ubiquitin thioesterase 38) (Ubiquitin-specific-processing protease 38) (Deubiquitinating enzyme 38). | |||||
|
USP9X_HUMAN
|
||||||
| θ value | 9.92553e-07 (rank : 47) | NC score | 0.542309 (rank : 60) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q93008, O75550, Q8WWT3, Q8WX12 | Gene names | USP9X, DFFRX, USP9 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome). | |||||
|
USP9X_MOUSE
|
||||||
| θ value | 1.69304e-06 (rank : 48) | NC score | 0.541326 (rank : 61) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P70398, Q62497 | Gene names | Usp9x, Fafl, Fam | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome) (Ubiquitin carboxyl-terminal hydrolase FAM) (Fat facets homolog). | |||||
|
USP9Y_HUMAN
|
||||||
| θ value | 2.21117e-06 (rank : 49) | NC score | 0.538758 (rank : 64) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | O00507, O14601 | Gene names | USP9Y, DFFRY, USP10 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-Y (EC 3.1.2.15) (Ubiquitin thioesterase FAF-Y) (Ubiquitin-specific-processing protease FAF-Y) (Deubiquitinating enzyme FAF-Y) (Fat facets protein-related, Y- linked) (Ubiquitin-specific protease 9, Y chromosome). | |||||
|
UBP47_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 50) | NC score | 0.590076 (rank : 54) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q96K76, Q658U0, Q86Y73, Q8TEP6, Q9BWI0, Q9NWN1 | Gene names | USP47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 47 (EC 3.1.2.15) (Ubiquitin thioesterase 47) (Ubiquitin-specific-processing protease 47) (Deubiquitinating enzyme 47). | |||||
|
UBP47_MOUSE
|
||||||
| θ value | 6.43352e-06 (rank : 51) | NC score | 0.592883 (rank : 53) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q8BY87, Q5EBP2, Q6KAR9, Q80V06, Q8BHU1, Q8BI15, Q8BI16, Q8BUW4, Q91X25 | Gene names | Usp47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 47 (EC 3.1.2.15) (Ubiquitin thioesterase 47) (Ubiquitin-specific-processing protease 47) (Deubiquitinating enzyme 47). | |||||
|
UBP10_HUMAN
|
||||||
| θ value | 1.09739e-05 (rank : 52) | NC score | 0.614975 (rank : 46) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q14694, Q9BWG7, Q9NSL7 | Gene names | USP10, KIAA0190 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 10 (EC 3.1.2.15) (Ubiquitin thioesterase 10) (Ubiquitin-specific-processing protease 10) (Deubiquitinating enzyme 10). | |||||
|
UBP10_MOUSE
|
||||||
| θ value | 1.09739e-05 (rank : 53) | NC score | 0.618843 (rank : 43) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | P52479, Q91VY7 | Gene names | Usp10, Ode-1, Uchrp | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 10 (EC 3.1.2.15) (Ubiquitin thioesterase 10) (Ubiquitin-specific-processing protease 10) (Deubiquitinating enzyme 10). | |||||
|
UBP35_HUMAN
|
||||||
| θ value | 1.43324e-05 (rank : 54) | NC score | 0.619730 (rank : 42) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9P2H5 | Gene names | USP35, KIAA1372, USP34 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 35 (EC 3.1.2.15) (Ubiquitin thioesterase 35) (Ubiquitin-specific-processing protease 35) (Deubiquitinating enzyme 35). | |||||
|
UBP13_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 55) | NC score | 0.495528 (rank : 66) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q92995 | Gene names | USP13, ISOT3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 13 (EC 3.1.2.15) (Ubiquitin thioesterase 13) (Ubiquitin-specific-processing protease 13) (Deubiquitinating enzyme 13) (Isopeptidase T-3) (ISOT-3). | |||||
|
UBP30_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 56) | NC score | 0.683937 (rank : 41) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q70CQ3, Q8WTU7, Q96JX4, Q9BSS3 | Gene names | USP30 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 30 (EC 3.1.2.15) (Ubiquitin thioesterase 30) (Ubiquitin-specific-processing protease 30) (Deubiquitinating enzyme 30). | |||||
|
UBP34_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 57) | NC score | 0.542481 (rank : 59) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q70CQ2, O60316, O94834, Q3B777, Q6P6C9, Q7L8P6, Q8N3T9, Q8TBW2, Q9UGA1 | Gene names | USP34, KIAA0570, KIAA0729 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 34 (EC 3.1.2.15) (Ubiquitin thioesterase 34) (Ubiquitin-specific-processing protease 34) (Deubiquitinating enzyme 34). | |||||
|
UBP34_MOUSE
|
||||||
| θ value | 0.000121331 (rank : 58) | NC score | 0.541165 (rank : 62) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q6ZQ93, Q3UPN0, Q6P563, Q7TMJ6, Q8CCH0 | Gene names | Usp34, Kiaa0570 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 34 (EC 3.1.2.15) (Ubiquitin thioesterase 34) (Ubiquitin-specific-processing protease 34) (Deubiquitinating enzyme 34). | |||||
|
UBP40_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 59) | NC score | 0.603563 (rank : 50) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9NVE5, Q6NX38, Q70EL0 | Gene names | USP40 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 40 (EC 3.1.2.15) (Ubiquitin thioesterase 40) (Ubiquitin-specific-processing protease 40) (Deubiquitinating enzyme 40). | |||||
|
UBP29_MOUSE
|
||||||
| θ value | 0.000270298 (rank : 60) | NC score | 0.512339 (rank : 65) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9ES63 | Gene names | Usp29 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 29 (EC 3.1.2.15) (Ubiquitin thioesterase 29) (Ubiquitin-specific-processing protease 29) (Deubiquitinating enzyme 29). | |||||
|
UBP7_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 61) | NC score | 0.595209 (rank : 51) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q93009 | Gene names | USP7, HAUSP | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 7 (EC 3.1.2.15) (Ubiquitin thioesterase 7) (Ubiquitin-specific-processing protease 7) (Deubiquitinating enzyme 7) (Herpesvirus-associated ubiquitin-specific protease). | |||||
|
UBP14_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 62) | NC score | 0.559097 (rank : 56) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P54578 | Gene names | USP14, TGT | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 14 (EC 3.1.2.15) (Ubiquitin thioesterase 14) (Ubiquitin-specific-processing protease 14) (Deubiquitinating enzyme 14). | |||||
|
UBP26_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 63) | NC score | 0.431573 (rank : 69) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q99MX1 | Gene names | Usp26 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 26 (EC 3.1.2.15) (Ubiquitin thioesterase 26) (Ubiquitin-specific-processing protease 26) (Deubiquitinating enzyme 26). | |||||
|
SNUT2_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 64) | NC score | 0.587850 (rank : 55) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q53GS9, Q6NX47, Q96RK9, Q9BV89, Q9H381, Q9P050, Q9Y310 | Gene names | USP39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 2 (U4/U6.U5 tri-snRNP-associated 65 kDa protein) (65K) (Inactive ubiquitin-specific peptidase 39) (SAD1 homolog). | |||||
|
SNUT2_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 65) | NC score | 0.594749 (rank : 52) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q3TIX9, Q3TI55, Q8BLI5, Q8BZZ5, Q9CQR1 | Gene names | Usp39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 2 (Inactive ubiquitin-specific peptidase 39). | |||||
|
UBP5_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 66) | NC score | 0.475352 (rank : 68) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P56399 | Gene names | Usp5, Isot | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
UBP5_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 67) | NC score | 0.479187 (rank : 67) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P45974, Q96J22 | Gene names | USP5, ISOT | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
UBP28_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 68) | NC score | 0.357197 (rank : 70) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q5I043, Q6NZP3, Q6ZPP1, Q8BWI1 | Gene names | Usp28, Kiaa1515 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 28 (EC 3.1.2.15) (Ubiquitin thioesterase 28) (Ubiquitin-specific-processing protease 28) (Deubiquitinating enzyme 28). | |||||
|
UBP1_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 69) | NC score | 0.617728 (rank : 45) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | O94782, Q9UFR0, Q9UNJ3 | Gene names | USP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 1 (EC 3.1.2.15) (Ubiquitin thioesterase 1) (Ubiquitin-specific-processing protease 1) (Deubiquitinating enzyme 1) (hUBP). | |||||
|
UBP14_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 70) | NC score | 0.545808 (rank : 58) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9JMA1, Q543U5, Q923F2, Q9D0L0 | Gene names | Usp14 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 14 (EC 3.1.2.15) (Ubiquitin thioesterase 14) (Ubiquitin-specific-processing protease 14) (Deubiquitinating enzyme 14). | |||||
|
UBP28_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 71) | NC score | 0.352691 (rank : 71) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q96RU2, Q9P213 | Gene names | USP28, KIAA1515 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 28 (EC 3.1.2.15) (Ubiquitin thioesterase 28) (Ubiquitin-specific-processing protease 28) (Deubiquitinating enzyme 28). | |||||
|
UBP25_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 72) | NC score | 0.327910 (rank : 73) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9UHP3, Q9H9W1 | Gene names | USP25, USP21 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 25 (EC 3.1.2.15) (Ubiquitin thioesterase 25) (Ubiquitin-specific-processing protease 25) (Deubiquitinating enzyme 25) (USP on chromosome 21). | |||||
|
UBP25_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 73) | NC score | 0.329089 (rank : 72) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P57080, Q80ZT9 | Gene names | Usp25 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 25 (EC 3.1.2.15) (Ubiquitin thioesterase 25) (Ubiquitin-specific-processing protease 25) (Deubiquitinating enzyme 25) (mUSP25). | |||||
|
ENL_HUMAN
|
||||||
| θ value | 0.125558 (rank : 74) | NC score | 0.037906 (rank : 81) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
MUC1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 75) | NC score | 0.016493 (rank : 94) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
BSN_HUMAN
|
||||||
| θ value | 0.163984 (rank : 76) | NC score | 0.026847 (rank : 85) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
CO5A2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 77) | NC score | 0.016648 (rank : 93) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P05997 | Gene names | COL5A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(V) chain precursor. | |||||
|
ZN385_HUMAN
|
||||||
| θ value | 0.163984 (rank : 78) | NC score | 0.027773 (rank : 83) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q96PM9, Q5VH53, Q9H7R6, Q9UFU3 | Gene names | ZNF385, HZF, RZF | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 385 (Hematopoietic zinc finger protein) (Retinal zinc finger protein). | |||||
|
I12R1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 79) | NC score | 0.031083 (rank : 82) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q60837 | Gene names | Il12rb1, Il12rb | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-12 receptor beta-1 chain precursor (IL-12R-beta1) (Interleukin-12 receptor beta) (IL-12 receptor beta component) (CD212 antigen). | |||||
|
TACC2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 80) | NC score | 0.014749 (rank : 99) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O95359, Q9NZ41, Q9NZR5 | Gene names | TACC2 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 2 (Anti Zuai-1) (AZU-1). | |||||
|
CSF1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 81) | NC score | 0.024742 (rank : 86) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P09603, Q13130, Q14086, Q14806, Q9UQR8 | Gene names | CSF1 | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage colony-stimulating factor 1 precursor (CSF-1) (MCSF) (M- CSF) (Lanimostim). | |||||
|
DEMA_HUMAN
|
||||||
| θ value | 0.62314 (rank : 82) | NC score | 0.019417 (rank : 89) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q08495, Q13215, Q9BRE3 | Gene names | EPB49, DMT | |||
|
Domain Architecture |

|
|||||
| Description | Dematin (Erythrocyte membrane protein band 4.9). | |||||
|
SALL3_HUMAN
|
||||||
| θ value | 0.62314 (rank : 83) | NC score | 0.000915 (rank : 128) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 1066 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9BXA9, Q9UGH1 | Gene names | SALL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 3 (Zinc finger protein SALL3) (hSALL3). | |||||
|
CIC_HUMAN
|
||||||
| θ value | 0.813845 (rank : 84) | NC score | 0.017152 (rank : 92) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
ISL1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 85) | NC score | 0.009143 (rank : 108) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 421 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P61371, P20663, P47894 | Gene names | ISL1 | |||
|
Domain Architecture |
|
|||||
| Description | Insulin gene enhancer protein ISL-1 (Islet-1). | |||||
|
ISL1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 86) | NC score | 0.009143 (rank : 109) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 421 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P61372, P20663, P47894, Q812D8 | Gene names | Isl1 | |||
|
Domain Architecture |
|
|||||
| Description | Insulin gene enhancer protein ISL-1 (Islet-1). | |||||
|
BPA1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 87) | NC score | 0.000578 (rank : 131) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
INP5E_HUMAN
|
||||||
| θ value | 1.38821 (rank : 88) | NC score | 0.015850 (rank : 95) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NRR6, Q15734 | Gene names | INPP5E | |||
|
Domain Architecture |
|
|||||
| Description | 72 kDa inositol polyphosphate 5-phosphatase (EC 3.1.3.36) (Phosphatidylinositol-4,5-bisphosphate 5-phosphatase) (Phosphatidylinositol polyphosphate 5-phosphatase type IV). | |||||
|
LIRB3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 89) | NC score | 0.010391 (rank : 103) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75022, O15471, Q86U49 | Gene names | LILRB3, ILT5, LIR3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leukocyte immunoglobulin-like receptor subfamily B member 3 precursor (Leukocyte immunoglobulin-like receptor 3) (LIR-3) (Immunoglobulin- like transcript 5) (ILT-5) (Monocyte inhibitory receptor HL9) (CD85a antigen). | |||||
|
ABCA2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 90) | NC score | 0.005234 (rank : 121) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P41234 | Gene names | Abca2, Abc2 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-binding cassette sub-family A member 2 (ATP-binding cassette transporter 2) (ATP-binding cassette 2). | |||||
|
MYO15_HUMAN
|
||||||
| θ value | 1.81305 (rank : 91) | NC score | 0.004241 (rank : 124) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
SYN3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 92) | NC score | 0.015212 (rank : 97) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14994 | Gene names | SYN3 | |||
|
Domain Architecture |
|
|||||
| Description | Synapsin-3 (Synapsin III). | |||||
|
IL6RA_MOUSE
|
||||||
| θ value | 2.36792 (rank : 93) | NC score | 0.012113 (rank : 100) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P22272 | Gene names | Il6ra, Il6r | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-6 receptor alpha chain precursor (IL-6R-alpha) (IL-6R 1) (CD126 antigen). | |||||
|
LIRB1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 94) | NC score | 0.009805 (rank : 106) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NHL6, O75024, O75025, Q8NHJ9, Q8NHK0 | Gene names | LILRB1, ILT2, LIR1, MIR7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leukocyte immunoglobulin-like receptor subfamily B member 1 precursor (Leukocyte immunoglobulin-like receptor 1) (LIR-1) (Immunoglobulin- like transcript 2) (ILT-2) (Monocyte/macrophage immunoglobulin-like receptor 7) (MIR-7) (CD85j antigen). | |||||
|
PKD1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 95) | NC score | 0.008398 (rank : 113) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P98161, Q15140, Q15141 | Gene names | PKD1 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystin-1 precursor (Autosomal dominant polycystic kidney disease protein 1). | |||||
|
41_HUMAN
|
||||||
| θ value | 3.0926 (rank : 96) | NC score | 0.004057 (rank : 125) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P11171, P11176, Q14245, Q5TB35, Q5VXN8, Q8IXV9, Q9Y578, Q9Y579 | Gene names | EPB41, E41P | |||
|
Domain Architecture |

|
|||||
| Description | Protein 4.1 (Band 4.1) (P4.1) (EPB4.1) (4.1R). | |||||
|
FBS1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 97) | NC score | 0.027403 (rank : 84) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P62706 | Gene names | FBS1, FBS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibrosin-1. | |||||
|
NETR_MOUSE
|
||||||
| θ value | 3.0926 (rank : 98) | NC score | -0.000193 (rank : 133) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O08762 | Gene names | Prss12, Bssp3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurotrypsin precursor (EC 3.4.21.-) (Motopsin) (Brain-specific serine protease 3) (BSSP-3). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 4.03905 (rank : 99) | NC score | 0.015035 (rank : 98) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
PDZD2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 100) | NC score | 0.019279 (rank : 90) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O15018, Q9BXD4 | Gene names | PDZD2, AIPC, KIAA0300, PDZK3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 2 (PDZ domain-containing protein 3) (Activated in prostate cancer protein). | |||||
|
TNR9_HUMAN
|
||||||
| θ value | 4.03905 (rank : 101) | NC score | 0.008233 (rank : 114) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q07011 | Gene names | TNFRSF9, CD137, ILA | |||
|
Domain Architecture |
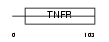
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 9 precursor (4-1BB ligand receptor) (T-cell antigen 4-1BB homolog) (T-cell antigen ILA) (CD137 antigen). | |||||
|
TRIM8_HUMAN
|
||||||
| θ value | 4.03905 (rank : 102) | NC score | 0.001660 (rank : 127) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BZR9, Q9C028 | Gene names | TRIM8, GERP, RNF27 | |||
|
Domain Architecture |
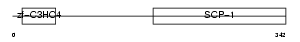
|
|||||
| Description | Tripartite motif-containing protein 8 (RING finger protein 27) (Glioblastoma-expressed RING finger protein). | |||||
|
CAC1F_HUMAN
|
||||||
| θ value | 5.27518 (rank : 103) | NC score | 0.002511 (rank : 126) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60840, O43901 | Gene names | CACNA1F, CACNAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1F (Voltage- gated calcium channel subunit alpha Cav1.4). | |||||
|
CEL_HUMAN
|
||||||
| θ value | 5.27518 (rank : 104) | NC score | 0.007676 (rank : 116) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
CIC_MOUSE
|
||||||
| θ value | 5.27518 (rank : 105) | NC score | 0.017298 (rank : 91) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 742 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q924A2, Q6PDJ8, Q8CGE4, Q8CHH0, Q9CW61 | Gene names | Cic, Kiaa0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
DOCK6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 106) | NC score | 0.006460 (rank : 120) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96HP0, Q9P2F2 | Gene names | DOCK6, KIAA1395 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 6. | |||||
|
GPTC8_HUMAN
|
||||||
| θ value | 5.27518 (rank : 107) | NC score | 0.011457 (rank : 101) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UKJ3, O60300 | Gene names | GPATC8, KIAA0553 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 8. | |||||
|
ODPX_MOUSE
|
||||||
| θ value | 5.27518 (rank : 108) | NC score | 0.008937 (rank : 111) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BKZ9 | Gene names | Pdhx | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pyruvate dehydrogenase protein X component, mitochondrial precursor (Dihydrolipoamide dehydrogenase-binding protein of pyruvate dehydrogenase complex) (Lipoyl-containing pyruvate dehydrogenase complex component X). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 109) | NC score | 0.008942 (rank : 110) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
DPOLZ_HUMAN
|
||||||
| θ value | 6.88961 (rank : 110) | NC score | 0.006872 (rank : 118) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60673, O43214 | Gene names | REV3L, POLZ, REV3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (hREV3). | |||||
|
EVX1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 111) | NC score | -0.000100 (rank : 132) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P23683 | Gene names | Evx1 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox even-skipped homolog protein 1 (EVX-1). | |||||
|
FSTL4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 112) | NC score | 0.000634 (rank : 129) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6MZW2, Q8TBU0, Q9UPU1 | Gene names | FSTL4, KIAA1061 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Follistatin-related protein 4 precursor (Follistatin-like 4). | |||||
|
IKKE_MOUSE
|
||||||
| θ value | 6.88961 (rank : 113) | NC score | -0.005146 (rank : 136) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 848 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9R0T8 | Gene names | Ikbke, Ikke, Ikki | |||
|
Domain Architecture |

|
|||||
| Description | Inhibitor of nuclear factor kappa-B kinase epsilon subunit (EC 2.7.11.10) (I kappa-B kinase epsilon) (IkBKE) (IKK-epsilon) (IKK- E) (Inducible I kappa-B kinase) (IKK-i). | |||||
|
K1210_HUMAN
|
||||||
| θ value | 6.88961 (rank : 114) | NC score | 0.021995 (rank : 87) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9ULL0, Q5JPN4 | Gene names | KIAA1210 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1210. | |||||
|
LIPA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 115) | NC score | 0.000626 (rank : 130) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 1294 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13136, Q13135, Q14567, Q8N4I2 | Gene names | PPFIA1, LIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-1) (PTPRF-interacting protein alpha-1) (LAR-interacting protein 1) (LIP.1). | |||||
|
MEF2B_MOUSE
|
||||||
| θ value | 6.88961 (rank : 116) | NC score | 0.006623 (rank : 119) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O55087, O55088, O55089, O55090, O55231, Q61843 | Gene names | Mef2b | |||
|
Domain Architecture |
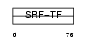
|
|||||
| Description | Myocyte-specific enhancer factor 2B. | |||||
|
TROAP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 117) | NC score | 0.008809 (rank : 112) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q12815 | Gene names | TROAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trophinin-associated protein (Tastin) (Trophinin-assisting protein). | |||||
|
CGRE1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 118) | NC score | 0.010568 (rank : 102) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q99674 | Gene names | CGREF1, CGR11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with EF hand domain 1 (Cell growth regulatory gene 11 protein). | |||||
|
CO4A2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 119) | NC score | 0.007480 (rank : 117) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 454 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P08572 | Gene names | COL4A2 | |||
|
Domain Architecture |
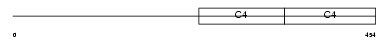
|
|||||
| Description | Collagen alpha-2(IV) chain precursor. | |||||
|
HMX2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 120) | NC score | -0.001294 (rank : 134) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P43687, Q8R1Q2 | Gene names | Hmx2, Nkx-5.2, Nkx5-2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein HMX2 (Homeobox protein Nkx-5.2). | |||||
|
LIRB4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | 0.008179 (rank : 115) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NHJ6, O15468, O75021, Q8N1C7, Q8NHL5 | Gene names | LILRB4, ILT3, LIR5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leukocyte immunoglobulin-like receptor subfamily B member 4 precursor (Leukocyte immunoglobulin-like receptor 5) (LIR-5) (Immunoglobulin- like transcript 3) (ILT-3) (Monocyte inhibitory receptor HM18) (CD85k antigen). | |||||
|
MARK2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 122) | NC score | -0.004114 (rank : 135) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q05512, Q6PDR4, Q8BR95 | Gene names | Mark2, Emk | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL Motif Kinase) (EMK1). | |||||
|
NU153_HUMAN
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | 0.015573 (rank : 96) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P49790 | Gene names | NUP153 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup153 (Nucleoporin Nup153) (153 kDa nucleoporin). | |||||
|
PTC2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.004452 (rank : 123) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y6C5, O95341, O95856, Q5QP87, Q6UX14 | Gene names | PTCH2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein patched homolog 2 (PTC2). | |||||
|
SPT5H_HUMAN
|
||||||
| θ value | 8.99809 (rank : 125) | NC score | 0.010170 (rank : 104) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O00267, O43279, Q59G52, Q99639 | Gene names | SUPT5H, SPT5, SPT5H | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT5 (hSPT5) (DRB sensitivity-inducing factor large subunit) (DSIF large subunit) (DSIF p160) (Tat- cotransactivator 1 protein) (Tat-CT1 protein). | |||||
|
SPT5H_MOUSE
|
||||||
| θ value | 8.99809 (rank : 126) | NC score | 0.009627 (rank : 107) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 437 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O55201, Q3UJ77, Q3UJH1, Q3UKD7, Q3UM54, Q6PB73, Q6PDP0, Q6PFR4 | Gene names | Supt5h | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT5 (DRB sensitivity-inducing factor large subunit) (DSIF large subunit). | |||||
|
TACC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 127) | NC score | 0.004758 (rank : 122) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75410, Q6Y687, Q86YG7, Q8IUJ2, Q8IUJ3, Q8IUJ4, Q8IZG2, Q8NEY7, Q9UPP9 | Gene names | TACC1, KIAA1103 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 1 (Taxin 1) (Gastric cancer antigen Ga55). | |||||
|
TM119_MOUSE
|
||||||
| θ value | 8.99809 (rank : 128) | NC score | 0.009882 (rank : 105) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R138, Q8BP14 | Gene names | Tmem119 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 119 precursor. | |||||
|
TXND2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 129) | NC score | 0.021780 (rank : 88) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
TYK2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 130) | NC score | -0.005721 (rank : 137) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 841 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P29597, Q6QB10, Q96CH0 | Gene names | TYK2 | |||
|
Domain Architecture |

|
|||||
| Description | Non-receptor tyrosine-protein kinase TYK2 (EC 2.7.10.2). | |||||
|
BRAP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.087667 (rank : 77) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q7Z569, O43238, O75341 | Gene names | BRAP, RNF52 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BRCA1-associated protein (EC 6.3.2.-) (BRAP2) (Impedes mitogenic signal propagation) (IMP) (RING finger protein 52) (NY-REN-63 antigen). | |||||
|
BRAP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.101098 (rank : 75) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99MP8, Q8CC00, Q8CHX1, Q99MP7, Q9CXX8 | Gene names | Brap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BRCA1-associated protein (EC 6.3.2.-) (BRAP2) (Impedes mitogenic signal propagation) (IMP). | |||||
|
GGNB1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.095835 (rank : 76) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5YKI7, Q5YKI8 | Gene names | GGNBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin-binding protein 1. | |||||
|
GGNB1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.125941 (rank : 74) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6K1E7, Q8C5T9 | Gene names | Ggnbp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin-binding protein 1. | |||||
|
HDAC6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.064282 (rank : 79) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UBN7 | Gene names | HDAC6 | |||
|
Domain Architecture |

|
|||||
| Description | Histone deacetylase 6 (HD6). | |||||
|
HDAC6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.069252 (rank : 78) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Z2V5 | Gene names | Hdac6 | |||
|
Domain Architecture |

|
|||||
| Description | Histone deacetylase 6 (HD6) (Histone deacetylase mHDA2). | |||||
|
TBCD3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.061103 (rank : 80) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8IZP1, Q9H0B9, Q9UDD4 | Gene names | TBC1D3, PRC17 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 3 (Rab GTPase-activating protein PRC17) (Prostate cancer gene 17 protein) (TRE17 alpha protein). | |||||
|
UBP43_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 130 | |
| SwissProt Accessions | Q8BUM9, Q8VDP5 | Gene names | Usp43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 43 (EC 3.1.2.15) (Ubiquitin thioesterase 43) (Ubiquitin-specific-processing protease 43) (Deubiquitinating enzyme 43). | |||||
|
UBP43_HUMAN
|
||||||
| NC score | 0.988926 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q70EL4, Q8N2C5, Q96DQ6 | Gene names | USP43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 43 (EC 3.1.2.15) (Ubiquitin thioesterase 43) (Ubiquitin-specific-processing protease 43) (Deubiquitinating enzyme 43). | |||||
|
UBP31_HUMAN
|
||||||
| NC score | 0.944435 (rank : 3) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q70CQ4, Q6AW97, Q6ZTC0, Q6ZTN2, Q9ULL7 | Gene names | USP31, KIAA1203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 31 (EC 3.1.2.15) (Ubiquitin thioesterase 31) (Ubiquitin-specific-processing protease 31) (Deubiquitinating enzyme 31). | |||||
|
UBP4_HUMAN
|
||||||
| NC score | 0.856885 (rank : 4) | θ value | 1.16164e-39 (rank : 4) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q13107, O43452, O43453 | Gene names | USP4, UNP, UNPH | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 4 (EC 3.1.2.15) (Ubiquitin thioesterase 4) (Ubiquitin-specific-processing protease 4) (Deubiquitinating enzyme 4) (Ubiquitous nuclear protein homolog). | |||||
|
UBP11_HUMAN
|
||||||
| NC score | 0.855853 (rank : 5) | θ value | 2.34606e-32 (rank : 8) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P51784, Q8IUG6, Q9BWE1 | Gene names | USP11, UHX1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 11 (EC 3.1.2.15) (Ubiquitin thioesterase 11) (Ubiquitin-specific-processing protease 11) (Deubiquitinating enzyme 11). | |||||
|
UBP4_MOUSE
|
||||||
| NC score | 0.855620 (rank : 6) | θ value | 9.83387e-39 (rank : 5) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P35123, O54704 | Gene names | Usp4, Unp | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 4 (EC 3.1.2.15) (Ubiquitin thioesterase 4) (Ubiquitin-specific-processing protease 4) (Deubiquitinating enzyme 4) (Ubiquitous nuclear protein). | |||||
|
UBP15_MOUSE
|
||||||
| NC score | 0.854588 (rank : 7) | θ value | 3.73686e-38 (rank : 6) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q8R5H1, Q3UL25 | Gene names | Usp15 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 15 (EC 3.1.2.15) (Ubiquitin thioesterase 15) (Ubiquitin-specific-processing protease 15) (Deubiquitinating enzyme 15). | |||||
|
UBP15_HUMAN
|
||||||
| NC score | 0.854120 (rank : 8) | θ value | 8.32485e-38 (rank : 7) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9Y4E8, Q9HCA6, Q9UNP0, Q9Y5B5 | Gene names | USP15, KIAA0529 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 15 (EC 3.1.2.15) (Ubiquitin thioesterase 15) (Ubiquitin-specific-processing protease 15) (Deubiquitinating enzyme 15) (Unph-2) (Unph4). | |||||
|
UBP2_MOUSE
|
||||||
| NC score | 0.848909 (rank : 9) | θ value | 6.61148e-27 (rank : 11) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | O88623, Q6X4T9, Q6X4U1, Q8VD74, Q9JJ87 | Gene names | Usp2, Ubp41 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 2 (EC 3.1.2.15) (Ubiquitin thioesterase 2) (Ubiquitin-specific-processing protease 2) (Deubiquitinating enzyme 2) (41 kDa ubiquitin-specific protease). | |||||
|
UBP2_HUMAN
|
||||||
| NC score | 0.847094 (rank : 10) | θ value | 3.28125e-26 (rank : 12) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | O75604, Q96MB9, Q9BQ21 | Gene names | USP2, UBP41 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 2 (EC 3.1.2.15) (Ubiquitin thioesterase 2) (Ubiquitin-specific-processing protease 2) (Deubiquitinating enzyme 2) (41 kDa ubiquitin-specific protease). | |||||
|
UBP50_MOUSE
|
||||||
| NC score | 0.845316 (rank : 11) | θ value | 2.12685e-25 (rank : 14) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q6P8X6, Q9D558, Q9D9E7 | Gene names | Usp50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ubiquitin carboxyl-terminal hydrolase 50 (EC 3.1.2.15) (Ubiquitin thioesterase 50) (Ubiquitin-specific-processing protease 50) (Deubiquitinating enzyme 50). | |||||
|
UBP8_MOUSE
|
||||||
| NC score | 0.835821 (rank : 12) | θ value | 2.19584e-30 (rank : 9) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q80U87, Q80YP2, Q8R0D3, Q9EQU1, Q9WVP5 | Gene names | Usp8, Kiaa0055, Ubpy | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 8 (EC 3.1.2.15) (Ubiquitin thioesterase 8) (Ubiquitin-specific-processing protease 8) (Deubiquitinating enzyme 8) (mUBPy). | |||||
|
UBP21_MOUSE
|
||||||
| NC score | 0.830715 (rank : 13) | θ value | 4.58923e-20 (rank : 24) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q9QZL6, Q9D0R1 | Gene names | Usp21, Usp23 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 21 (EC 3.1.2.15) (Ubiquitin thioesterase 21) (Ubiquitin-specific-processing protease 21) (Deubiquitinating enzyme 21). | |||||
|
UBP50_HUMAN
|
||||||
| NC score | 0.830658 (rank : 14) | θ value | 2.35696e-16 (rank : 25) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q70EL3 | Gene names | USP50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inactive ubiquitin carboxyl-terminal hydrolase 50 (Inactive ubiquitin- specific peptidase 50). | |||||
|
UBP21_HUMAN
|
||||||
| NC score | 0.829687 (rank : 15) | θ value | 3.51386e-20 (rank : 23) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q9UK80, Q9BTV1, Q9NYN4 | Gene names | USP21, USP23 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 21 (EC 3.1.2.15) (Ubiquitin thioesterase 21) (Ubiquitin-specific-processing protease 21) (Deubiquitinating enzyme 21) (NEDD8-specific protease). | |||||
|
UBP20_HUMAN
|
||||||
| NC score | 0.829121 (rank : 16) | θ value | 3.62785e-25 (rank : 15) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9Y2K6, Q96LG5, Q9UQN8, Q9UQP0 | Gene names | USP20, KIAA1003, LSFR3A | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 20 (EC 3.1.2.15) (Ubiquitin thioesterase 20) (Ubiquitin-specific-processing protease 20) (Deubiquitinating enzyme 20). | |||||
|
UBP3_MOUSE
|
||||||
| NC score | 0.828976 (rank : 17) | θ value | 8.93572e-24 (rank : 17) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q91W36 | Gene names | Usp3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 3 (EC 3.1.2.15) (Ubiquitin thioesterase 3) (Ubiquitin-specific-processing protease 3) (Deubiquitinating enzyme 3). | |||||
|
UBP3_HUMAN
|
||||||
| NC score | 0.828783 (rank : 18) | θ value | 1.99067e-23 (rank : 18) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9Y6I4 | Gene names | USP3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 3 (EC 3.1.2.15) (Ubiquitin thioesterase 3) (Ubiquitin-specific-processing protease 3) (Deubiquitinating enzyme 3). | |||||
|
UBP33_HUMAN
|
||||||
| NC score | 0.826362 (rank : 19) | θ value | 7.30988e-26 (rank : 13) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q8TEY7, Q8TEY6, Q96AV6, Q9H9F0, Q9UPQ5 | Gene names | USP33, KIAA1097, VDU1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 33 (EC 3.1.2.15) (Ubiquitin thioesterase 33) (Ubiquitin-specific-processing protease 33) (Deubiquitinating enzyme 33) (VHL-interacting deubiquitinating enzyme 1). | |||||
|
UBP51_HUMAN
|
||||||
| NC score | 0.814680 (rank : 20) | θ value | 3.07116e-24 (rank : 16) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q70EK9, Q8IWJ8 | Gene names | USP51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 51 (EC 3.1.2.15) (Ubiquitin thioesterase 51) (Ubiquitin-specific-processing protease 51) (Deubiquitinating enzyme 51). | |||||
|
UBP22_MOUSE
|
||||||
| NC score | 0.808811 (rank : 21) | θ value | 1.86321e-21 (rank : 21) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q5DU02, Q5SU81, Q66JV8 | Gene names | Usp22, Kiaa1063 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 22 (EC 3.1.2.15) (Ubiquitin thioesterase 22) (Ubiquitin-specific-processing protease 22) (Deubiquitinating enzyme 22). | |||||
|
UBP22_HUMAN
|
||||||
| NC score | 0.807571 (rank : 22) | θ value | 5.42112e-21 (rank : 22) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9UPT9 | Gene names | USP22, KIAA1063, USP3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 22 (EC 3.1.2.15) (Ubiquitin thioesterase 22) (Ubiquitin-specific-processing protease 22) (Deubiquitinating enzyme 22). | |||||
|
UBP8_HUMAN
|
||||||
| NC score | 0.796249 (rank : 23) | θ value | 4.89182e-30 (rank : 10) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 751 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | P40818, Q7Z3U2, Q86VA0, Q8IWI7 | Gene names | USP8, KIAA0055, UBPY | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 8 (EC 3.1.2.15) (Ubiquitin thioesterase 8) (Ubiquitin-specific-processing protease 8) (Deubiquitinating enzyme 8) (hUBPy). | |||||
|
UBP16_HUMAN
|
||||||
| NC score | 0.778304 (rank : 24) | θ value | 1.74796e-11 (rank : 31) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9Y5T5, Q8NEL3 | Gene names | USP16 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 16 (EC 3.1.2.15) (Ubiquitin thioesterase 16) (Ubiquitin-specific-processing protease 16) (Deubiquitinating enzyme 16) (Ubiquitin-processing protease UBP-M). | |||||
|
UBP44_HUMAN
|
||||||
| NC score | 0.766407 (rank : 25) | θ value | 2.60593e-15 (rank : 26) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9H0E7 | Gene names | USP44 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 44 (EC 3.1.2.15) (Ubiquitin thioesterase 44) (Ubiquitin-specific-processing protease 44) (Deubiquitinating enzyme 44). | |||||
|
UBP49_MOUSE
|
||||||
| NC score | 0.753894 (rank : 26) | θ value | 4.91457e-14 (rank : 27) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q6P9L4 | Gene names | Usp49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 49 (EC 3.1.2.15) (Ubiquitin thioesterase 49) (Ubiquitin-specific-processing protease 49) (Deubiquitinating enzyme 49). | |||||
|
UBP49_HUMAN
|
||||||
| NC score | 0.751469 (rank : 27) | θ value | 8.38298e-14 (rank : 28) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q70CQ1, Q96CK4 | Gene names | USP49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 49 (EC 3.1.2.15) (Ubiquitin thioesterase 49) (Ubiquitin-specific-processing protease 49) (Deubiquitinating enzyme 49). | |||||
|
UBPW_MOUSE
|
||||||
| NC score | 0.751262 (rank : 28) | θ value | 3.29651e-10 (rank : 34) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q61068 | Gene names | Dub1, Dub-1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase DUB-1 (EC 3.1.2.15) (Ubiquitin thioesterase DUB-1) (Ubiquitin-specific-processing protease DUB-1) (Deubiquitinating enzyme 1). | |||||
|
UBP32_HUMAN
|
||||||
| NC score | 0.746949 (rank : 29) | θ value | 7.56453e-23 (rank : 20) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q8NFA0, Q9BX85, Q9Y591 | Gene names | USP32, USP10 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 32 (EC 3.1.2.15) (Ubiquitin thioesterase 32) (Ubiquitin-specific-processing protease 32) (Deubiquitinating enzyme 32) (NY-REN-60 antigen). | |||||
|
UBP18_HUMAN
|
||||||
| NC score | 0.740164 (rank : 30) | θ value | 1.47974e-10 (rank : 32) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9UMW8, Q9NY71 | Gene names | USP18, ISG43 | |||
|
Domain Architecture |

|
|||||
| Description | Ubl carboxyl-terminal hydrolase 18 (EC 3.1.2.-) (Ubl thioesterase 18) (ISG15-specific-processing protease) (43 kDa ISG15-specific protease) (hUBP43). | |||||
|
UBP12_HUMAN
|
||||||
| NC score | 0.736269 (rank : 31) | θ value | 2.79066e-09 (rank : 38) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | O75317 | Gene names | USP12, UBH1, USP12L1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 12 (EC 3.1.2.15) (Ubiquitin thioesterase 12) (Ubiquitin-specific-processing protease 12) (Deubiquitinating enzyme 12) (Ubiquitin-hydrolyzing enzyme 1). | |||||
|
UBP46_HUMAN
|
||||||
| NC score | 0.732955 (rank : 32) | θ value | 1.25267e-09 (rank : 36) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P62068, Q80V95, Q9H7U4, Q9H9T8 | Gene names | USP46 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 46 (EC 3.1.2.15) (Ubiquitin thioesterase 46) (Ubiquitin-specific-processing protease 46) (Deubiquitinating enzyme 46). | |||||
|
UBP46_MOUSE
|
||||||
| NC score | 0.732955 (rank : 33) | θ value | 1.25267e-09 (rank : 37) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P62069, Q80V95, Q9H7U4, Q9H9T8 | Gene names | Usp46 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 46 (EC 3.1.2.15) (Ubiquitin thioesterase 46) (Ubiquitin-specific-processing protease 46) (Deubiquitinating enzyme 46). | |||||
|
UB17L_HUMAN
|
||||||
| NC score | 0.731769 (rank : 34) | θ value | 2.52405e-10 (rank : 33) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q7RTZ2 | Gene names | USP17L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 17-like protein (EC 3.1.2.15) (Ubiquitin thioesterase 17-like) (Ubiquitin-specific-processing protease 17-like) (Deubiquitinating enzyme 17-like). | |||||
|
UBP18_MOUSE
|
||||||
| NC score | 0.726491 (rank : 35) | θ value | 5.62301e-10 (rank : 35) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9WTV6 | Gene names | Usp18, Ubp43 | |||
|
Domain Architecture |

|
|||||
| Description | Ubl carboxyl-terminal hydrolase 18 (EC 3.1.2.-) (Ubl thioesterase 18) (ISG15-specific-processing protease) (43 kDa ISG15-specific protease). | |||||
|
UBP36_HUMAN
|
||||||
| NC score | 0.715724 (rank : 36) | θ value | 5.26297e-08 (rank : 42) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q9P275, Q8NDM8, Q9NVC8 | Gene names | USP36, KIAA1453 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 36 (EC 3.1.2.15) (Ubiquitin thioesterase 36) (Ubiquitin-specific-processing protease 36) (Deubiquitinating enzyme 36). | |||||
|
UBP6_HUMAN
|
||||||
| NC score | 0.703962 (rank : 37) | θ value | 5.79196e-23 (rank : 19) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P35125, Q15634, Q86WP6, Q8IWT4 | Gene names | USP6, TRE2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 6 (EC 3.1.2.15) (Ubiquitin thioesterase 6) (Ubiquitin-specific-processing protease 6) (Deubiquitinating enzyme 6) (Proto-oncogene TRE-2). | |||||
|
UBP42_HUMAN
|
||||||
| NC score | 0.698324 (rank : 38) | θ value | 8.11959e-09 (rank : 40) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q9H9J4, Q3C166, Q6P9B4 | Gene names | USP42 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 42 (EC 3.1.2.15) (Ubiquitin thioesterase 42) (Ubiquitin-specific-processing protease 42) (Deubiquitinating enzyme 42). | |||||
|
UBP48_HUMAN
|
||||||
| NC score | 0.691561 (rank : 39) | θ value | 1.33837e-11 (rank : 30) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q86UV5, Q2M3I4, Q5SZI4, Q5T3T5, Q6NX53, Q8N3F6, Q96F64, Q96IQ3, Q9H5N3, Q9H5T7, Q9NUJ6, Q9NXR0 | Gene names | USP48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 48 (EC 3.1.2.15) (Ubiquitin thioesterase 48) (Ubiquitin-specific-processing protease 48) (Deubiquitinating enzyme 48). | |||||
|
UBP48_MOUSE
|
||||||
| NC score | 0.690530 (rank : 40) | θ value | 6.00763e-12 (rank : 29) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q3V0C5, Q3TM39, Q571J9, Q8VDJ1 | Gene names | Usp48, Kiaa4202 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 48 (EC 3.1.2.15) (Ubiquitin thioesterase 48) (Ubiquitin-specific-processing protease 48) (Deubiquitinating enzyme 48). | |||||
|
UBP30_HUMAN
|
||||||
| NC score | 0.683937 (rank : 41) | θ value | 4.1701e-05 (rank : 56) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q70CQ3, Q8WTU7, Q96JX4, Q9BSS3 | Gene names | USP30 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 30 (EC 3.1.2.15) (Ubiquitin thioesterase 30) (Ubiquitin-specific-processing protease 30) (Deubiquitinating enzyme 30). | |||||
|
UBP35_HUMAN
|
||||||
| NC score | 0.619730 (rank : 42) | θ value | 1.43324e-05 (rank : 54) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9P2H5 | Gene names | USP35, KIAA1372, USP34 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 35 (EC 3.1.2.15) (Ubiquitin thioesterase 35) (Ubiquitin-specific-processing protease 35) (Deubiquitinating enzyme 35). | |||||
|
UBP10_MOUSE
|
||||||
| NC score | 0.618843 (rank : 43) | θ value | 1.09739e-05 (rank : 53) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | P52479, Q91VY7 | Gene names | Usp10, Ode-1, Uchrp | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 10 (EC 3.1.2.15) (Ubiquitin thioesterase 10) (Ubiquitin-specific-processing protease 10) (Deubiquitinating enzyme 10). | |||||
|
UBP24_HUMAN
|
||||||
| NC score | 0.618603 (rank : 44) | θ value | 1.80886e-08 (rank : 41) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9UPU5, Q6ZSY2, Q8N2Y4, Q9NXD1 | Gene names | USP24, KIAA1057 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 24 (EC 3.1.2.15) (Ubiquitin thioesterase 24) (Ubiquitin-specific-processing protease 24) (Deubiquitinating enzyme 24). | |||||
|
UBP1_HUMAN
|
||||||
| NC score | 0.617728 (rank : 45) | θ value | 0.00665767 (rank : 69) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | O94782, Q9UFR0, Q9UNJ3 | Gene names | USP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 1 (EC 3.1.2.15) (Ubiquitin thioesterase 1) (Ubiquitin-specific-processing protease 1) (Deubiquitinating enzyme 1) (hUBP). | |||||
|
UBP10_HUMAN
|
||||||
| NC score | 0.614975 (rank : 46) | θ value | 1.09739e-05 (rank : 52) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q14694, Q9BWG7, Q9NSL7 | Gene names | USP10, KIAA0190 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 10 (EC 3.1.2.15) (Ubiquitin thioesterase 10) (Ubiquitin-specific-processing protease 10) (Deubiquitinating enzyme 10). | |||||
|
UBP37_HUMAN
|
||||||
| NC score | 0.613506 (rank : 47) | θ value | 2.79066e-09 (rank : 39) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q86T82, Q9HCH8 | Gene names | USP37, KIAA1594 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 37 (EC 3.1.2.15) (Ubiquitin thioesterase 37) (Ubiquitin-specific-processing protease 37) (Deubiquitinating enzyme 37). | |||||
|
UBP38_HUMAN
|
||||||
| NC score | 0.605385 (rank : 48) | θ value | 8.97725e-08 (rank : 43) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q8NB14, Q8NDF5, Q96DK6, Q96PZ6, Q9BY55 | Gene names | USP38, KIAA1891 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 38 (EC 3.1.2.15) (Ubiquitin thioesterase 38) (Ubiquitin-specific-processing protease 38) (Deubiquitinating enzyme 38) (HP43.8KD). | |||||
|
UBP38_MOUSE
|
||||||
| NC score | 0.604101 (rank : 49) | θ value | 5.81887e-07 (rank : 46) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q8BW70, Q8BWL1, Q8BX03 | Gene names | Usp38 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 38 (EC 3.1.2.15) (Ubiquitin thioesterase 38) (Ubiquitin-specific-processing protease 38) (Deubiquitinating enzyme 38). | |||||
|
UBP40_HUMAN
|
||||||
| NC score | 0.603563 (rank : 50) | θ value | 0.000158464 (rank : 59) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9NVE5, Q6NX38, Q70EL0 | Gene names | USP40 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 40 (EC 3.1.2.15) (Ubiquitin thioesterase 40) (Ubiquitin-specific-processing protease 40) (Deubiquitinating enzyme 40). | |||||
|
UBP7_HUMAN
|
||||||
| NC score | 0.595209 (rank : 51) | θ value | 0.00035302 (rank : 61) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q93009 | Gene names | USP7, HAUSP | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 7 (EC 3.1.2.15) (Ubiquitin thioesterase 7) (Ubiquitin-specific-processing protease 7) (Deubiquitinating enzyme 7) (Herpesvirus-associated ubiquitin-specific protease). | |||||
|
SNUT2_MOUSE
|
||||||
| NC score | 0.594749 (rank : 52) | θ value | 0.00175202 (rank : 65) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q3TIX9, Q3TI55, Q8BLI5, Q8BZZ5, Q9CQR1 | Gene names | Usp39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 2 (Inactive ubiquitin-specific peptidase 39). | |||||
|
UBP47_MOUSE
|
||||||
| NC score | 0.592883 (rank : 53) | θ value | 6.43352e-06 (rank : 51) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q8BY87, Q5EBP2, Q6KAR9, Q80V06, Q8BHU1, Q8BI15, Q8BI16, Q8BUW4, Q91X25 | Gene names | Usp47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 47 (EC 3.1.2.15) (Ubiquitin thioesterase 47) (Ubiquitin-specific-processing protease 47) (Deubiquitinating enzyme 47). | |||||
|
UBP47_HUMAN
|
||||||
| NC score | 0.590076 (rank : 54) | θ value | 6.43352e-06 (rank : 50) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q96K76, Q658U0, Q86Y73, Q8TEP6, Q9BWI0, Q9NWN1 | Gene names | USP47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 47 (EC 3.1.2.15) (Ubiquitin thioesterase 47) (Ubiquitin-specific-processing protease 47) (Deubiquitinating enzyme 47). | |||||
|
SNUT2_HUMAN
|
||||||
| NC score | 0.587850 (rank : 55) | θ value | 0.00175202 (rank : 64) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q53GS9, Q6NX47, Q96RK9, Q9BV89, Q9H381, Q9P050, Q9Y310 | Gene names | USP39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 2 (U4/U6.U5 tri-snRNP-associated 65 kDa protein) (65K) (Inactive ubiquitin-specific peptidase 39) (SAD1 homolog). | |||||
|
UBP14_HUMAN
|
||||||
| NC score | 0.559097 (rank : 56) | θ value | 0.00102713 (rank : 62) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P54578 | Gene names | USP14, TGT | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 14 (EC 3.1.2.15) (Ubiquitin thioesterase 14) (Ubiquitin-specific-processing protease 14) (Deubiquitinating enzyme 14). | |||||
|
UBP29_HUMAN
|
||||||
| NC score | 0.549655 (rank : 57) | θ value | 3.41135e-07 (rank : 44) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9HBJ7 | Gene names | USP29 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 29 (EC 3.1.2.15) (Ubiquitin thioesterase 29) (Ubiquitin-specific-processing protease 29) (Deubiquitinating enzyme 29). | |||||
|
UBP14_MOUSE
|
||||||
| NC score | 0.545808 (rank : 58) | θ value | 0.0113563 (rank : 70) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9JMA1, Q543U5, Q923F2, Q9D0L0 | Gene names | Usp14 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 14 (EC 3.1.2.15) (Ubiquitin thioesterase 14) (Ubiquitin-specific-processing protease 14) (Deubiquitinating enzyme 14). | |||||
|
UBP34_HUMAN
|
||||||
| NC score | 0.542481 (rank : 59) | θ value | 7.1131e-05 (rank : 57) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q70CQ2, O60316, O94834, Q3B777, Q6P6C9, Q7L8P6, Q8N3T9, Q8TBW2, Q9UGA1 | Gene names | USP34, KIAA0570, KIAA0729 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 34 (EC 3.1.2.15) (Ubiquitin thioesterase 34) (Ubiquitin-specific-processing protease 34) (Deubiquitinating enzyme 34). | |||||
|
USP9X_HUMAN
|
||||||
| NC score | 0.542309 (rank : 60) | θ value | 9.92553e-07 (rank : 47) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q93008, O75550, Q8WWT3, Q8WX12 | Gene names | USP9X, DFFRX, USP9 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome). | |||||
|
USP9X_MOUSE
|
||||||
| NC score | 0.541326 (rank : 61) | θ value | 1.69304e-06 (rank : 48) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P70398, Q62497 | Gene names | Usp9x, Fafl, Fam | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome) (Ubiquitin carboxyl-terminal hydrolase FAM) (Fat facets homolog). | |||||
|
UBP34_MOUSE
|
||||||
| NC score | 0.541165 (rank : 62) | θ value | 0.000121331 (rank : 58) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q6ZQ93, Q3UPN0, Q6P563, Q7TMJ6, Q8CCH0 | Gene names | Usp34, Kiaa0570 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 34 (EC 3.1.2.15) (Ubiquitin thioesterase 34) (Ubiquitin-specific-processing protease 34) (Deubiquitinating enzyme 34). | |||||
|
UBP26_HUMAN
|
||||||
| NC score | 0.540954 (rank : 63) | θ value | 5.81887e-07 (rank : 45) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9BXU7 | Gene names | USP26 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 26 (EC 3.1.2.15) (Ubiquitin thioesterase 26) (Ubiquitin-specific-processing protease 26) (Deubiquitinating enzyme 26). | |||||
|
USP9Y_HUMAN
|
||||||
| NC score | 0.538758 (rank : 64) | θ value | 2.21117e-06 (rank : 49) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | O00507, O14601 | Gene names | USP9Y, DFFRY, USP10 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-Y (EC 3.1.2.15) (Ubiquitin thioesterase FAF-Y) (Ubiquitin-specific-processing protease FAF-Y) (Deubiquitinating enzyme FAF-Y) (Fat facets protein-related, Y- linked) (Ubiquitin-specific protease 9, Y chromosome). | |||||
|
UBP29_MOUSE
|
||||||
| NC score | 0.512339 (rank : 65) | θ value | 0.000270298 (rank : 60) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9ES63 | Gene names | Usp29 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 29 (EC 3.1.2.15) (Ubiquitin thioesterase 29) (Ubiquitin-specific-processing protease 29) (Deubiquitinating enzyme 29). | |||||
|
UBP13_HUMAN
|
||||||
| NC score | 0.495528 (rank : 66) | θ value | 2.44474e-05 (rank : 55) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q92995 | Gene names | USP13, ISOT3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 13 (EC 3.1.2.15) (Ubiquitin thioesterase 13) (Ubiquitin-specific-processing protease 13) (Deubiquitinating enzyme 13) (Isopeptidase T-3) (ISOT-3). | |||||
|
UBP5_HUMAN
|
||||||
| NC score | 0.479187 (rank : 67) | θ value | 0.00228821 (rank : 67) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P45974, Q96J22 | Gene names | USP5, ISOT | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
UBP5_MOUSE
|
||||||
| NC score | 0.475352 (rank : 68) | θ value | 0.00175202 (rank : 66) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P56399 | Gene names | Usp5, Isot | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
UBP26_MOUSE
|
||||||
| NC score | 0.431573 (rank : 69) | θ value | 0.00134147 (rank : 63) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q99MX1 | Gene names | Usp26 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 26 (EC 3.1.2.15) (Ubiquitin thioesterase 26) (Ubiquitin-specific-processing protease 26) (Deubiquitinating enzyme 26). | |||||
|
UBP28_MOUSE
|
||||||
| NC score | 0.357197 (rank : 70) | θ value | 0.00298849 (rank : 68) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q5I043, Q6NZP3, Q6ZPP1, Q8BWI1 | Gene names | Usp28, Kiaa1515 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 28 (EC 3.1.2.15) (Ubiquitin thioesterase 28) (Ubiquitin-specific-processing protease 28) (Deubiquitinating enzyme 28). | |||||
|
UBP28_HUMAN
|
||||||
| NC score | 0.352691 (rank : 71) | θ value | 0.0563607 (rank : 71) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q96RU2, Q9P213 | Gene names | USP28, KIAA1515 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 28 (EC 3.1.2.15) (Ubiquitin thioesterase 28) (Ubiquitin-specific-processing protease 28) (Deubiquitinating enzyme 28). | |||||
|
UBP25_MOUSE
|
||||||
| NC score | 0.329089 (rank : 72) | θ value | 0.0961366 (rank : 73) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P57080, Q80ZT9 | Gene names | Usp25 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 25 (EC 3.1.2.15) (Ubiquitin thioesterase 25) (Ubiquitin-specific-processing protease 25) (Deubiquitinating enzyme 25) (mUSP25). | |||||
|
UBP25_HUMAN
|
||||||
| NC score | 0.327910 (rank : 73) | θ value | 0.0961366 (rank : 72) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9UHP3, Q9H9W1 | Gene names | USP25, USP21 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 25 (EC 3.1.2.15) (Ubiquitin thioesterase 25) (Ubiquitin-specific-processing protease 25) (Deubiquitinating enzyme 25) (USP on chromosome 21). | |||||
|
GGNB1_MOUSE
|
||||||
| NC score | 0.125941 (rank : 74) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6K1E7, Q8C5T9 | Gene names | Ggnbp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin-binding protein 1. | |||||
|
BRAP_MOUSE
|
||||||
| NC score | 0.101098 (rank : 75) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99MP8, Q8CC00, Q8CHX1, Q99MP7, Q9CXX8 | Gene names | Brap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BRCA1-associated protein (EC 6.3.2.-) (BRAP2) (Impedes mitogenic signal propagation) (IMP). | |||||
|
GGNB1_HUMAN
|
||||||
| NC score | 0.095835 (rank : 76) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5YKI7, Q5YKI8 | Gene names | GGNBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin-binding protein 1. | |||||
|
BRAP_HUMAN
|
||||||
| NC score | 0.087667 (rank : 77) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q7Z569, O43238, O75341 | Gene names | BRAP, RNF52 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BRCA1-associated protein (EC 6.3.2.-) (BRAP2) (Impedes mitogenic signal propagation) (IMP) (RING finger protein 52) (NY-REN-63 antigen). | |||||
|
HDAC6_MOUSE
|
||||||
| NC score | 0.069252 (rank : 78) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Z2V5 | Gene names | Hdac6 | |||
|
Domain Architecture |

|
|||||
| Description | Histone deacetylase 6 (HD6) (Histone deacetylase mHDA2). | |||||
|
HDAC6_HUMAN
|
||||||
| NC score | 0.064282 (rank : 79) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UBN7 | Gene names | HDAC6 | |||
|
Domain Architecture |

|
|||||
| Description | Histone deacetylase 6 (HD6). | |||||
|
TBCD3_HUMAN
|
||||||
| NC score | 0.061103 (rank : 80) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8IZP1, Q9H0B9, Q9UDD4 | Gene names | TBC1D3, PRC17 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 3 (Rab GTPase-activating protein PRC17) (Prostate cancer gene 17 protein) (TRE17 alpha protein). | |||||
|
ENL_HUMAN
|
||||||
| NC score | 0.037906 (rank : 81) | θ value | 0.125558 (rank : 74) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
I12R1_MOUSE
|
||||||
| NC score | 0.031083 (rank : 82) | θ value | 0.21417 (rank : 79) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q60837 | Gene names | Il12rb1, Il12rb | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-12 receptor beta-1 chain precursor (IL-12R-beta1) (Interleukin-12 receptor beta) (IL-12 receptor beta component) (CD212 antigen). | |||||
|
ZN385_HUMAN
|
||||||
| NC score | 0.027773 (rank : 83) | θ value | 0.163984 (rank : 78) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q96PM9, Q5VH53, Q9H7R6, Q9UFU3 | Gene names | ZNF385, HZF, RZF | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 385 (Hematopoietic zinc finger protein) (Retinal zinc finger protein). | |||||
|
FBS1_HUMAN
|
||||||
| NC score | 0.027403 (rank : 84) | θ value | 3.0926 (rank : 97) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P62706 | Gene names | FBS1, FBS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibrosin-1. | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.026847 (rank : 85) | θ value | 0.163984 (rank : 76) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
CSF1_HUMAN
|
||||||
| NC score | 0.024742 (rank : 86) | θ value | 0.47712 (rank : 81) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P09603, Q13130, Q14086, Q14806, Q9UQR8 | Gene names | CSF1 | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage colony-stimulating factor 1 precursor (CSF-1) (MCSF) (M- CSF) (Lanimostim). | |||||
|
K1210_HUMAN
|
||||||
| NC score | 0.021995 (rank : 87) | θ value | 6.88961 (rank : 114) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9ULL0, Q5JPN4 | Gene names | KIAA1210 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1210. | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.021780 (rank : 88) | θ value | 8.99809 (rank : 129) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
DEMA_HUMAN
|
||||||
| NC score | 0.019417 (rank : 89) | θ value | 0.62314 (rank : 82) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q08495, Q13215, Q9BRE3 | Gene names | EPB49, DMT | |||
|
Domain Architecture |

|
|||||
| Description | Dematin (Erythrocyte membrane protein band 4.9). | |||||
|
PDZD2_HUMAN
|
||||||
| NC score | 0.019279 (rank : 90) | θ value | 4.03905 (rank : 100) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O15018, Q9BXD4 | Gene names | PDZD2, AIPC, KIAA0300, PDZK3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 2 (PDZ domain-containing protein 3) (Activated in prostate cancer protein). | |||||
|
CIC_MOUSE
|
||||||
| NC score | 0.017298 (rank : 91) | θ value | 5.27518 (rank : 105) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 742 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q924A2, Q6PDJ8, Q8CGE4, Q8CHH0, Q9CW61 | Gene names | Cic, Kiaa0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
CIC_HUMAN
|
||||||
| NC score | 0.017152 (rank : 92) | θ value | 0.813845 (rank : 84) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
CO5A2_HUMAN
|
||||||
| NC score | 0.016648 (rank : 93) | θ value | 0.163984 (rank : 77) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P05997 | Gene names | COL5A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(V) chain precursor. | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.016493 (rank : 94) | θ value | 0.125558 (rank : 75) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
INP5E_HUMAN
|
||||||
| NC score | 0.015850 (rank : 95) | θ value | 1.38821 (rank : 88) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NRR6, Q15734 | Gene names | INPP5E | |||
|
Domain Architecture |
|
|||||
| Description | 72 kDa inositol polyphosphate 5-phosphatase (EC 3.1.3.36) (Phosphatidylinositol-4,5-bisphosphate 5-phosphatase) (Phosphatidylinositol polyphosphate 5-phosphatase type IV). | |||||
|
NU153_HUMAN
|
||||||
| NC score | 0.015573 (rank : 96) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P49790 | Gene names | NUP153 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup153 (Nucleoporin Nup153) (153 kDa nucleoporin). | |||||
|
SYN3_HUMAN
|
||||||
| NC score | 0.015212 (rank : 97) | θ value | 1.81305 (rank : 92) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14994 | Gene names | SYN3 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-3 (Synapsin III). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.015035 (rank : 98) | θ value | 4.03905 (rank : 99) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
TACC2_HUMAN
|
||||||
| NC score | 0.014749 (rank : 99) | θ value | 0.279714 (rank : 80) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O95359, Q9NZ41, Q9NZR5 | Gene names | TACC2 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 2 (Anti Zuai-1) (AZU-1). | |||||
|
IL6RA_MOUSE
|
||||||
| NC score | 0.012113 (rank : 100) | θ value | 2.36792 (rank : 93) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P22272 | Gene names | Il6ra, Il6r | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-6 receptor alpha chain precursor (IL-6R-alpha) (IL-6R 1) (CD126 antigen). | |||||
|
GPTC8_HUMAN
|
||||||
| NC score | 0.011457 (rank : 101) | θ value | 5.27518 (rank : 107) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UKJ3, O60300 | Gene names | GPATC8, KIAA0553 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 8. | |||||
|
CGRE1_HUMAN
|
||||||
| NC score | 0.010568 (rank : 102) | θ value | 8.99809 (rank : 118) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q99674 | Gene names | CGREF1, CGR11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with EF hand domain 1 (Cell growth regulatory gene 11 protein). | |||||
|
LIRB3_HUMAN
|
||||||
| NC score | 0.010391 (rank : 103) | θ value | 1.38821 (rank : 89) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75022, O15471, Q86U49 | Gene names | LILRB3, ILT5, LIR3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leukocyte immunoglobulin-like receptor subfamily B member 3 precursor (Leukocyte immunoglobulin-like receptor 3) (LIR-3) (Immunoglobulin- like transcript 5) (ILT-5) (Monocyte inhibitory receptor HL9) (CD85a antigen). | |||||
|
SPT5H_HUMAN
|
||||||
| NC score | 0.010170 (rank : 104) | θ value | 8.99809 (rank : 125) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O00267, O43279, Q59G52, Q99639 | Gene names | SUPT5H, SPT5, SPT5H | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT5 (hSPT5) (DRB sensitivity-inducing factor large subunit) (DSIF large subunit) (DSIF p160) (Tat- cotransactivator 1 protein) (Tat-CT1 protein). | |||||
|
TM119_MOUSE
|
||||||
| NC score | 0.009882 (rank : 105) | θ value | 8.99809 (rank : 128) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R138, Q8BP14 | Gene names | Tmem119 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 119 precursor. | |||||
|
LIRB1_HUMAN
|
||||||
| NC score | 0.009805 (rank : 106) | θ value | 2.36792 (rank : 94) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NHL6, O75024, O75025, Q8NHJ9, Q8NHK0 | Gene names | LILRB1, ILT2, LIR1, MIR7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leukocyte immunoglobulin-like receptor subfamily B member 1 precursor (Leukocyte immunoglobulin-like receptor 1) (LIR-1) (Immunoglobulin- like transcript 2) (ILT-2) (Monocyte/macrophage immunoglobulin-like receptor 7) (MIR-7) (CD85j antigen). | |||||
|
SPT5H_MOUSE
|
||||||
| NC score | 0.009627 (rank : 107) | θ value | 8.99809 (rank : 126) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 437 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O55201, Q3UJ77, Q3UJH1, Q3UKD7, Q3UM54, Q6PB73, Q6PDP0, Q6PFR4 | Gene names | Supt5h | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT5 (DRB sensitivity-inducing factor large subunit) (DSIF large subunit). | |||||
|
ISL1_HUMAN
|
||||||
| NC score | 0.009143 (rank : 108) | θ value | 0.813845 (rank : 85) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 421 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P61371, P20663, P47894 | Gene names | ISL1 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin gene enhancer protein ISL-1 (Islet-1). | |||||
|
ISL1_MOUSE
|
||||||
| NC score | 0.009143 (rank : 109) | θ value | 0.813845 (rank : 86) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 421 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P61372, P20663, P47894, Q812D8 | Gene names | Isl1 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin gene enhancer protein ISL-1 (Islet-1). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.008942 (rank : 110) | θ value | 5.27518 (rank : 109) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
ODPX_MOUSE
|
||||||
| NC score | 0.008937 (rank : 111) | θ value | 5.27518 (rank : 108) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BKZ9 | Gene names | Pdhx | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pyruvate dehydrogenase protein X component, mitochondrial precursor (Dihydrolipoamide dehydrogenase-binding protein of pyruvate dehydrogenase complex) (Lipoyl-containing pyruvate dehydrogenase complex component X). | |||||
|
TROAP_HUMAN
|
||||||
| NC score | 0.008809 (rank : 112) | θ value | 6.88961 (rank : 117) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q12815 | Gene names | TROAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trophinin-associated protein (Tastin) (Trophinin-assisting protein). | |||||
|
PKD1_HUMAN
|
||||||
| NC score | 0.008398 (rank : 113) | θ value | 2.36792 (rank : 95) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P98161, Q15140, Q15141 | Gene names | PKD1 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystin-1 precursor (Autosomal dominant polycystic kidney disease protein 1). | |||||
|
TNR9_HUMAN
|
||||||
| NC score | 0.008233 (rank : 114) | θ value | 4.03905 (rank : 101) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q07011 | Gene names | TNFRSF9, CD137, ILA | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 9 precursor (4-1BB ligand receptor) (T-cell antigen 4-1BB homolog) (T-cell antigen ILA) (CD137 antigen). | |||||
|
LIRB4_HUMAN
|
||||||
| NC score | 0.008179 (rank : 115) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NHJ6, O15468, O75021, Q8N1C7, Q8NHL5 | Gene names | LILRB4, ILT3, LIR5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leukocyte immunoglobulin-like receptor subfamily B member 4 precursor (Leukocyte immunoglobulin-like receptor 5) (LIR-5) (Immunoglobulin- like transcript 3) (ILT-3) (Monocyte inhibitory receptor HM18) (CD85k antigen). | |||||
|
CEL_HUMAN
|
||||||
| NC score | 0.007676 (rank : 116) | θ value | 5.27518 (rank : 104) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
CO4A2_HUMAN
|
||||||
| NC score | 0.007480 (rank : 117) | θ value | 8.99809 (rank : 119) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 454 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P08572 | Gene names | COL4A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(IV) chain precursor. | |||||
|
DPOLZ_HUMAN
|
||||||
| NC score | 0.006872 (rank : 118) | θ value | 6.88961 (rank : 110) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60673, O43214 | Gene names | REV3L, POLZ, REV3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (hREV3). | |||||
|
MEF2B_MOUSE
|
||||||
| NC score | 0.006623 (rank : 119) | θ value | 6.88961 (rank : 116) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O55087, O55088, O55089, O55090, O55231, Q61843 | Gene names | Mef2b | |||
|
Domain Architecture |

|
|||||
| Description | Myocyte-specific enhancer factor 2B. | |||||
|
DOCK6_HUMAN
|
||||||
| NC score | 0.006460 (rank : 120) | θ value | 5.27518 (rank : 106) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96HP0, Q9P2F2 | Gene names | DOCK6, KIAA1395 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 6. | |||||
|
ABCA2_MOUSE
|
||||||
| NC score | 0.005234 (rank : 121) | θ value | 1.81305 (rank : 90) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P41234 | Gene names | Abca2, Abc2 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-binding cassette sub-family A member 2 (ATP-binding cassette transporter 2) (ATP-binding cassette 2). | |||||
|
TACC1_HUMAN
|
||||||
| NC score | 0.004758 (rank : 122) | θ value | 8.99809 (rank : 127) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75410, Q6Y687, Q86YG7, Q8IUJ2, Q8IUJ3, Q8IUJ4, Q8IZG2, Q8NEY7, Q9UPP9 | Gene names | TACC1, KIAA1103 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 1 (Taxin 1) (Gastric cancer antigen Ga55). | |||||
|
PTC2_HUMAN
|
||||||
| NC score | 0.004452 (rank : 123) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y6C5, O95341, O95856, Q5QP87, Q6UX14 | Gene names | PTCH2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein patched homolog 2 (PTC2). | |||||
|
MYO15_HUMAN
|
||||||
| NC score | 0.004241 (rank : 124) | θ value | 1.81305 (rank : 91) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
41_HUMAN
|
||||||
| NC score | 0.004057 (rank : 125) | θ value | 3.0926 (rank : 96) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P11171, P11176, Q14245, Q5TB35, Q5VXN8, Q8IXV9, Q9Y578, Q9Y579 | Gene names | EPB41, E41P | |||
|
Domain Architecture |

|
|||||
| Description | Protein 4.1 (Band 4.1) (P4.1) (EPB4.1) (4.1R). | |||||
|
CAC1F_HUMAN
|
||||||
| NC score | 0.002511 (rank : 126) | θ value | 5.27518 (rank : 103) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60840, O43901 | Gene names | CACNA1F, CACNAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1F (Voltage- gated calcium channel subunit alpha Cav1.4). | |||||
|
TRIM8_HUMAN
|
||||||
| NC score | 0.001660 (rank : 127) | θ value | 4.03905 (rank : 102) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BZR9, Q9C028 | Gene names | TRIM8, GERP, RNF27 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 8 (RING finger protein 27) (Glioblastoma-expressed RING finger protein). | |||||
|
SALL3_HUMAN
|
||||||
| NC score | 0.000915 (rank : 128) | θ value | 0.62314 (rank : 83) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 1066 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9BXA9, Q9UGH1 | Gene names | SALL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 3 (Zinc finger protein SALL3) (hSALL3). | |||||
|
FSTL4_HUMAN
|
||||||
| NC score | 0.000634 (rank : 129) | θ value | 6.88961 (rank : 112) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6MZW2, Q8TBU0, Q9UPU1 | Gene names | FSTL4, KIAA1061 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Follistatin-related protein 4 precursor (Follistatin-like 4). | |||||
|
LIPA1_HUMAN
|
||||||
| NC score | 0.000626 (rank : 130) | θ value | 6.88961 (rank : 115) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 1294 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13136, Q13135, Q14567, Q8N4I2 | Gene names | PPFIA1, LIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-1) (PTPRF-interacting protein alpha-1) (LAR-interacting protein 1) (LIP.1). | |||||
|
BPA1_MOUSE
|
||||||
| NC score | 0.000578 (rank : 131) | θ value | 1.38821 (rank : 87) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
EVX1_MOUSE
|
||||||
| NC score | -0.000100 (rank : 132) | θ value | 6.88961 (rank : 111) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P23683 | Gene names | Evx1 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox even-skipped homolog protein 1 (EVX-1). | |||||
|
NETR_MOUSE
|
||||||
| NC score | -0.000193 (rank : 133) | θ value | 3.0926 (rank : 98) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O08762 | Gene names | Prss12, Bssp3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurotrypsin precursor (EC 3.4.21.-) (Motopsin) (Brain-specific serine protease 3) (BSSP-3). | |||||
|
HMX2_MOUSE
|
||||||
| NC score | -0.001294 (rank : 134) | θ value | 8.99809 (rank : 120) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P43687, Q8R1Q2 | Gene names | Hmx2, Nkx-5.2, Nkx5-2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein HMX2 (Homeobox protein Nkx-5.2). | |||||
|
MARK2_MOUSE
|
||||||
| NC score | -0.004114 (rank : 135) | θ value | 8.99809 (rank : 122) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q05512, Q6PDR4, Q8BR95 | Gene names | Mark2, Emk | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL Motif Kinase) (EMK1). | |||||
|
IKKE_MOUSE
|
||||||
| NC score | -0.005146 (rank : 136) | θ value | 6.88961 (rank : 113) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 848 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9R0T8 | Gene names | Ikbke, Ikke, Ikki | |||
|
Domain Architecture |

|
|||||
| Description | Inhibitor of nuclear factor kappa-B kinase epsilon subunit (EC 2.7.11.10) (I kappa-B kinase epsilon) (IkBKE) (IKK-epsilon) (IKK- E) (Inducible I kappa-B kinase) (IKK-i). | |||||
|
TYK2_HUMAN
|
||||||
| NC score | -0.005721 (rank : 137) | θ value | 8.99809 (rank : 130) | |||
| Query Neighborhood Hits | 130 | Target Neighborhood Hits | 841 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P29597, Q6QB10, Q96CH0 | Gene names | TYK2 | |||
|
Domain Architecture |

|
|||||
| Description | Non-receptor tyrosine-protein kinase TYK2 (EC 2.7.10.2). | |||||