Please be patient as the page loads
|
MDM2_HUMAN
|
||||||
| SwissProt Accessions | Q00987, Q13226, Q13297, Q13298, Q13299, Q13300, Q13301, Q53XW0, Q71TW9, Q9UGI3, Q9UMT8 | Gene names | MDM2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3 Mdm2 (EC 6.3.2.-) (p53-binding protein Mdm2) (Oncoprotein Mdm2) (Double minute 2 protein) (Hdm2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
MDM2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 92 | |
| SwissProt Accessions | Q00987, Q13226, Q13297, Q13298, Q13299, Q13300, Q13301, Q53XW0, Q71TW9, Q9UGI3, Q9UMT8 | Gene names | MDM2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3 Mdm2 (EC 6.3.2.-) (p53-binding protein Mdm2) (Oncoprotein Mdm2) (Double minute 2 protein) (Hdm2). | |||||
|
MDM2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.975358 (rank : 2) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P23804, Q61040, Q64330 | Gene names | Mdm2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3 Mdm2 (EC 6.3.2.-) (p53-binding protein Mdm2) (Oncoprotein Mdm2) (Double minute 2 protein). | |||||
|
MDM4_HUMAN
|
||||||
| θ value | 8.55514e-59 (rank : 3) | NC score | 0.877328 (rank : 3) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O15151, Q6GS18, Q8IV83 | Gene names | MDM4, MDMX | |||
|
Domain Architecture |
|
|||||
| Description | Mdm4 protein (p53-binding protein Mdm4) (Mdm2-like p53-binding protein) (Mdmx protein) (Double minute 4 protein). | |||||
|
MDM4_MOUSE
|
||||||
| θ value | 2.10721e-57 (rank : 4) | NC score | 0.870463 (rank : 4) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O35618 | Gene names | Mdm4, Mdmx | |||
|
Domain Architecture |
|
|||||
| Description | Mdm4 protein (p53-binding protein Mdm4) (Mdm2-like p53-binding protein) (Mdmx protein) (Double minute 4 protein). | |||||
|
NEURL_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 5) | NC score | 0.263297 (rank : 5) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q923S6, Q91ZT6, Q9CWF1, Q9QWF1 | Gene names | Neurl, Neurl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuralized-like protein 1 (m-neuralized 1) (m-neu1). | |||||
|
NEUL1_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 6) | NC score | 0.259012 (rank : 6) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O76050, Q5TDR2, Q5TDR3, Q8TAN0, Q9H463 | Gene names | NEURL, NEURL1, RNF67 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuralized-like protein 1 (h-neuralized 1) (h-neu) (RING finger protein 67). | |||||
|
BIRC3_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 7) | NC score | 0.171199 (rank : 13) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q13489, Q16628, Q9HC27, Q9UP46 | Gene names | BIRC3, API2, IAP1, MIHC | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 3 (Inhibitor of apoptosis protein 1) (HIAP1) (HIAP-1) (C-IAP2) (TNFR2-TRAF signaling complex protein 1) (IAP homolog C) (Apoptosis inhibitor 2) (API2). | |||||
|
RNF34_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 8) | NC score | 0.208226 (rank : 7) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q99KR6, Q3UV45 | Gene names | Rnf34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 34 (RING finger protein RIFF) (Phafin-1). | |||||
|
RNF34_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 9) | NC score | 0.206869 (rank : 8) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q969K3, Q8NG47, Q9H6W8 | Gene names | RNF34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 34 (RING finger protein RIFF) (FYVE-RING finger protein Momo) (Human RING finger homologous to inhibitor of apoptosis protein) (hRFI) (Caspases-8 and -10-associated RING finger protein 1) (CARP-1) (Caspase regulator CARP1). | |||||
|
TOPRS_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 10) | NC score | 0.106765 (rank : 21) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q80Z37, Q3U5B5, Q3UPZ1, Q3USN3, Q3UZ55, Q8BXP2, Q8CFF5, Q8CGC8, Q920L3 | Gene names | Topors | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein E3 ligase Topors (EC 6.3.2.-) (SUMO1-protein E3 ligase Topors) (Topoisomerase I-binding RING finger protein) (Topoisomerase I-binding arginine/serine-rich protein) (Tumor suppressor p53-binding protein 3) (p53-binding protein 3) (p53BP3). | |||||
|
BIRC2_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 11) | NC score | 0.168592 (rank : 14) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q62210, O08864 | Gene names | Birc2, Birc3, Iap2 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 2 (Inhibitor of apoptosis protein 2) (MIAP2) (MIAP-2). | |||||
|
BIRC3_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 12) | NC score | 0.167103 (rank : 15) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O08863 | Gene names | Birc3, Birc2, Iap1 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 3 (Inhibitor of apoptosis protein 1) (MIAP1) (MIAP-1). | |||||
|
MGRN1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 13) | NC score | 0.179191 (rank : 11) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9D074, Q3U5V9, Q3UDA1, Q6ZQ97, Q8BZM9 | Gene names | Mgrn1, Kiaa0544 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase MGRN1 (EC 6.3.2.-) (Mahogunin ring finger protein 1). | |||||
|
FANCM_HUMAN
|
||||||
| θ value | 0.163984 (rank : 14) | NC score | 0.041645 (rank : 40) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IYD8, Q3YFH9, Q8N9X6, Q9HCH6 | Gene names | FANCM, KIAA1596 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group M protein (EC 3.6.1.-) (ATP-dependent RNA helicase FANCM) (Protein FACM) (Fanconi anemia-associated polypeptide of 250 kDa) (FAAP250) (Protein Hef ortholog). | |||||
|
LRSM1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 15) | NC score | 0.068589 (rank : 30) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1078 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q6UWE0, Q5VVV0, Q8NB40, Q96GT5, Q96MX5, Q96MZ7 | Gene names | LRSAM1, TAL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein LRSAM1 (EC 6.3.2.-) (Leucine-rich repeat and sterile alpha motif-containing protein 1) (Tsg101-associated ligase) (hTAL). | |||||
|
MGRN1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 16) | NC score | 0.173300 (rank : 12) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O60291, Q86W76 | Gene names | MGRN1, KIAA0544, RNF156 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase MGRN1 (EC 6.3.2.-) (Mahogunin ring finger protein 1) (RING finger protein 156). | |||||
|
RFFL_MOUSE
|
||||||
| θ value | 0.21417 (rank : 17) | NC score | 0.195692 (rank : 9) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6ZQM0, Q5SVC2, Q5SVC4, Q9D543, Q9D9B1 | Gene names | Rffl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rififylin (RING finger and FYVE-like domain-containing protein 1) (Fring). | |||||
|
DSPP_HUMAN
|
||||||
| θ value | 0.279714 (rank : 18) | NC score | 0.050954 (rank : 36) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
RFFL_HUMAN
|
||||||
| θ value | 0.279714 (rank : 19) | NC score | 0.191739 (rank : 10) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8WZ73, Q8NHW0, Q8TBY7, Q96BE6 | Gene names | RFFL, RNF189, RNF34L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rififylin (RING finger and FYVE-like domain-containing protein 1) (FYVE-RING finger protein Sakura) (Fring) (Caspases-8 and -10- associated RING finger protein 2) (CARP-2) (Caspase regulator CARP2) (RING finger protein 189) (RING finger protein 34-like). | |||||
|
BIRC2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 20) | NC score | 0.152249 (rank : 16) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13490, Q16516, Q4TTG0 | Gene names | BIRC2, API1, IAP2, MIHB | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 2 (Inhibitor of apoptosis protein 2) (HIAP2) (HIAP-2) (C-IAP1) (TNFR2-TRAF signaling complex protein 2) (IAP homolog B). | |||||
|
MIB1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 21) | NC score | 0.068212 (rank : 31) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q86YT6, Q68D01, Q6YI51, Q8NBY0, Q8TCB5, Q8TCL7, Q9P2M3 | Gene names | MIB1, DIP1, KIAA1323, ZZANK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein MIB1 (EC 6.3.2.-) (Mind bomb homolog 1) (DAPK-interacting protein 1) (DIP-1) (Zinc finger ZZ type with ankyrin repeat domain protein 2). | |||||
|
MIB1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 22) | NC score | 0.069480 (rank : 29) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q80SY4, Q5XK51, Q6IS57, Q6YI52, Q6ZPT8, Q8BNR1, Q8C6W2, Q921Q1 | Gene names | Mib1, Dip1, Kiaa1323, Mib | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein MIB1 (EC 6.3.2.-) (Mind bomb homolog 1) (DAPK-interacting protein 1) (DIP-1). | |||||
|
CHFR_MOUSE
|
||||||
| θ value | 0.47712 (rank : 23) | NC score | 0.065611 (rank : 33) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q810L3, Q8BJZ9, Q8BWH4 | Gene names | Chfr | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase CHFR (EC 6.3.2.-) (Checkpoint with forkhead and RING finger domains protein). | |||||
|
AN32A_HUMAN
|
||||||
| θ value | 0.62314 (rank : 24) | NC score | 0.028943 (rank : 55) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P39687 | Gene names | ANP32A, C15orf1, LANP, MAPM, PHAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member A (Potent heat-stable protein phosphatase 2A inhibitor I1PP2A) (Acidic nuclear phosphoprotein pp32) (Leucine-rich acidic nuclear protein) (Lanp) (Putative HLA-DR-associated protein I) (PHAPI) (Mapmodulin). | |||||
|
BIRC4_MOUSE
|
||||||
| θ value | 0.62314 (rank : 25) | NC score | 0.149358 (rank : 17) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q60989, O08865 | Gene names | Birc4, Aipa, Api3, Miha, Xiap | |||
|
Domain Architecture |
|
|||||
| Description | Baculoviral IAP repeat-containing protein 4 (Inhibitor of apoptosis protein 3) (X-linked inhibitor of apoptosis protein) (X-linked IAP) (IAP homolog A) (MIAP3) (MIAP-3). | |||||
|
LRSM1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 26) | NC score | 0.073522 (rank : 28) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q80ZI6 | Gene names | Lrsam1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein LRSAM1 (EC 6.3.2.-) (Leucine-rich repeat and sterile alpha motif-containing protein 1) (Tsg101-associated ligase). | |||||
|
RBP2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 27) | NC score | 0.020241 (rank : 65) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9ERU9, Q61992, Q8C9K9 | Gene names | Ranbp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2 (RanBP2). | |||||
|
ZDH21_HUMAN
|
||||||
| θ value | 0.62314 (rank : 28) | NC score | 0.036799 (rank : 46) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IVQ6, Q5VWG1 | Gene names | ZDHHC21 | |||
|
Domain Architecture |
|
|||||
| Description | Probable palmitoyltransferase ZDHHC21 (EC 2.3.1.-) (Zinc finger DHHC domain-containing protein 21) (DHHC-21). | |||||
|
ZDH21_MOUSE
|
||||||
| θ value | 0.62314 (rank : 29) | NC score | 0.036860 (rank : 45) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D270, Q6EMK1, Q80XQ3 | Gene names | Zdhhc21, Gramp3 | |||
|
Domain Architecture |
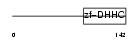
|
|||||
| Description | Probable palmitoyltransferase ZDHHC21 (EC 2.3.1.-) (Zinc finger DHHC domain-containing protein 21) (DHHC-21) (GABA-A receptor-associated membrane protein 3). | |||||
|
BIRC4_HUMAN
|
||||||
| θ value | 0.813845 (rank : 30) | NC score | 0.136958 (rank : 18) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P98170, Q9NQ14 | Gene names | BIRC4, API3, IAP3, XIAP | |||
|
Domain Architecture |
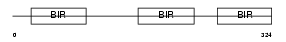
|
|||||
| Description | Baculoviral IAP repeat-containing protein 4 (Inhibitor of apoptosis protein 3) (X-linked inhibitor of apoptosis protein) (X-linked IAP) (IAP-like protein) (HILP). | |||||
|
DSPP_MOUSE
|
||||||
| θ value | 0.813845 (rank : 31) | NC score | 0.053137 (rank : 35) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
NALP5_MOUSE
|
||||||
| θ value | 0.813845 (rank : 32) | NC score | 0.022903 (rank : 61) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9R1M5, Q9JLR2 | Gene names | Nalp5, Mater | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 5 (Maternal antigen that embryos require) (Mater protein) (Ooplasm-specific protein 1) (OP1). | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 0.813845 (rank : 33) | NC score | 0.045149 (rank : 39) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
ZRAB2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 34) | NC score | 0.061035 (rank : 34) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O95218, Q9UP63 | Gene names | ZRANB2, ZIS, ZNF265 | |||
|
Domain Architecture |
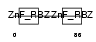
|
|||||
| Description | Zinc finger Ran-binding domain-containing protein 2 Zinc finger protein 265 (Zinc finger, splicing). | |||||
|
MYLIP_HUMAN
|
||||||
| θ value | 1.06291 (rank : 35) | NC score | 0.075786 (rank : 27) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8WY64, Q9BU73, Q9NRL9, Q9UHE7 | Gene names | MYLIP, BZF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase MYLIP (EC 6.3.2.-) (Myosin regulatory light chain- interacting protein) (MIR). | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 1.38821 (rank : 36) | NC score | 0.041007 (rank : 41) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
RPGR_HUMAN
|
||||||
| θ value | 1.38821 (rank : 37) | NC score | 0.029036 (rank : 54) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92834, O00702, O00737, Q8N5T6, Q93039, Q9HD29, Q9UMR1 | Gene names | RPGR, RP3, XLRP3 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator. | |||||
|
TOPRS_HUMAN
|
||||||
| θ value | 1.38821 (rank : 38) | NC score | 0.088152 (rank : 24) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9NS56, O43273, Q6P987, Q9NS55, Q9UNR9 | Gene names | TOPORS, LUN, TP53BPL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein E3 ligase Topors (EC 6.3.2.-) (SUMO1-protein E3 ligase Topors) (Topoisomerase I-binding RING finger protein) (Topoisomerase I-binding arginine/serine-rich protein) (Tumor suppressor p53-binding protein 3) (p53-binding protein 3) (p53BP3). | |||||
|
CR034_MOUSE
|
||||||
| θ value | 1.81305 (rank : 39) | NC score | 0.013957 (rank : 78) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 643 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CDV0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C18orf34 homolog. | |||||
|
MYLIP_MOUSE
|
||||||
| θ value | 1.81305 (rank : 40) | NC score | 0.067795 (rank : 32) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8BM54, Q3TX01, Q8BLY9, Q91Z47 | Gene names | Mylip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase MYLIP (EC 6.3.2.-) (Myosin regulatory light chain- interacting protein) (MIR). | |||||
|
ABTAP_MOUSE
|
||||||
| θ value | 2.36792 (rank : 41) | NC score | 0.032616 (rank : 49) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q3V1V3 | Gene names | Abtap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ABT1-associated protein. | |||||
|
FA13C_HUMAN
|
||||||
| θ value | 2.36792 (rank : 42) | NC score | 0.024060 (rank : 60) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NE31, Q99787 | Gene names | FAM13C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13C1. | |||||
|
PHF14_HUMAN
|
||||||
| θ value | 2.36792 (rank : 43) | NC score | 0.030254 (rank : 51) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
PHF14_MOUSE
|
||||||
| θ value | 2.36792 (rank : 44) | NC score | 0.030025 (rank : 52) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
BCL6_HUMAN
|
||||||
| θ value | 3.0926 (rank : 45) | NC score | 0.001364 (rank : 97) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P41182 | Gene names | BCL6, BCL5, LAZ3, ZBTB27, ZNF51 | |||
|
Domain Architecture |
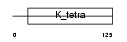
|
|||||
| Description | B-cell lymphoma 6 protein (BCL-6) (Zinc finger protein 51) (LAZ-3 protein) (BCL-5) (Zinc finger and BTB domain-containing protein 27). | |||||
|
BCL6_MOUSE
|
||||||
| θ value | 3.0926 (rank : 46) | NC score | 0.001461 (rank : 96) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 913 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P41183, Q61065 | Gene names | Bcl6, Bcl-6 | |||
|
Domain Architecture |
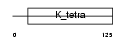
|
|||||
| Description | B-cell lymphoma 6 protein homolog. | |||||
|
CHD9_MOUSE
|
||||||
| θ value | 3.0926 (rank : 47) | NC score | 0.015882 (rank : 74) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BYH8, Q7TMV5, Q8BJG8, Q8BZJ2, Q8CHG8 | Gene names | Chd9, Kiaa0308, Pric320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha- interacting complex protein 320 kDa). | |||||
|
LEO1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 48) | NC score | 0.040884 (rank : 42) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
PRP4B_HUMAN
|
||||||
| θ value | 3.0926 (rank : 49) | NC score | 0.008470 (rank : 86) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1030 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q13523, Q8TDP2, Q96QT7, Q9UEE6 | Gene names | PRPF4B, KIAA0536, PRP4, PRP4H, PRP4K | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PRP4 homolog (EC 2.7.11.1) (PRP4 pre- mRNA-processing factor 4 homolog) (PRP4 kinase). | |||||
|
PRP4B_MOUSE
|
||||||
| θ value | 3.0926 (rank : 50) | NC score | 0.008304 (rank : 88) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1040 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q61136, O88378, Q8BND8, Q8R4Y5, Q9CTL9, Q9CTT0 | Gene names | Prpf4b, Cbp143, Prp4h, Prp4k, Prp4m, Prpk | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PRP4 homolog (EC 2.7.11.1) (PRP4 pre- mRNA-processing factor 4 homolog) (Pre-mRNA protein kinase). | |||||
|
RPGR1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 51) | NC score | 0.024892 (rank : 58) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 713 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9EPQ2, Q8CAC2, Q8CDJ9, Q91WE0, Q9CUK6, Q9D5Q1 | Gene names | Rpgrip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator-interacting protein 1 (RPGR-interacting protein 1). | |||||
|
BIRC7_HUMAN
|
||||||
| θ value | 4.03905 (rank : 52) | NC score | 0.114865 (rank : 19) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96CA5, Q9BQV0, Q9H2A8, Q9HAP7 | Gene names | BIRC7, KIAP, LIVIN, MLIAP | |||
|
Domain Architecture |
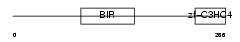
|
|||||
| Description | Baculoviral IAP repeat-containing protein 7 (Kidney inhibitor of apoptosis protein) (KIAP) (Melanoma inhibitor of apoptosis protein) (ML-IAP) (Livin). | |||||
|
CM007_MOUSE
|
||||||
| θ value | 4.03905 (rank : 53) | NC score | 0.034063 (rank : 48) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K2Y0, Q9CYU1, Q9D1Y0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C13orf7 homolog. | |||||
|
NP1L1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 54) | NC score | 0.020358 (rank : 63) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55209 | Gene names | NAP1L1, NRP | |||
|
Domain Architecture |
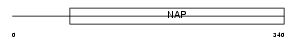
|
|||||
| Description | Nucleosome assembly protein 1-like 1 (NAP-1-related protein) (hNRP). | |||||
|
NP1L1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 55) | NC score | 0.020338 (rank : 64) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P28656, Q3UL14 | Gene names | Nap1l1, Nrp | |||
|
Domain Architecture |
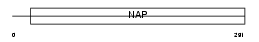
|
|||||
| Description | Nucleosome assembly protein 1-like 1 (NAP-1-related protein) (Brain protein DN38). | |||||
|
SESN1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 56) | NC score | 0.014357 (rank : 76) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P58006, Q7TNF3 | Gene names | Sesn1, Pa26, Sest1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sestrin-1 (p53-regulated protein PA26). | |||||
|
TRI37_HUMAN
|
||||||
| θ value | 4.03905 (rank : 57) | NC score | 0.037679 (rank : 44) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O94972, Q7Z3E6, Q8IYF7, Q8WYF7 | Gene names | TRIM37, KIAA0898, MUL, POB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 37 (Mulibrey nanism protein). | |||||
|
TRI37_MOUSE
|
||||||
| θ value | 4.03905 (rank : 58) | NC score | 0.037730 (rank : 43) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 316 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6PCX9, Q8CHC5 | Gene names | Trim37, Kiaa0898 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 37. | |||||
|
TXND2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 59) | NC score | 0.018572 (rank : 67) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
CHD1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.015783 (rank : 75) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
DDIT3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.022177 (rank : 62) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P35638 | Gene names | DDIT3, CHOP, GADD153 | |||
|
Domain Architecture |
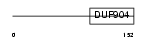
|
|||||
| Description | DNA damage-inducible transcript 3 (DDIT-3) (Growth arrest and DNA- damage-inducible protein GADD153) (C/EBP-homologous protein) (CHOP). | |||||
|
DPF3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.012106 (rank : 80) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P58269 | Gene names | Dpf3, Cerd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc-finger protein Dpf3 (cer-d4). | |||||
|
FA44A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 63) | NC score | 0.024900 (rank : 57) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8NFC6, Q6P0M8 | Gene names | FAM44A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM44A. | |||||
|
LEF1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.011583 (rank : 82) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P27782 | Gene names | Lef1, Lef-1 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphoid enhancer-binding factor 1 (LEF-1). | |||||
|
RBM16_HUMAN
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.018984 (rank : 66) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 300 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UPN6, Q5TBU6, Q9BQN8, Q9BX43 | Gene names | RBM16, KIAA1116 | |||
|
Domain Architecture |

|
|||||
| Description | Putative RNA-binding protein 16 (RNA-binding motif protein 16). | |||||
|
RN103_HUMAN
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.045771 (rank : 38) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O00237, Q8IVB9 | Gene names | RNF103 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 103 (Zinc finger protein 103 homolog) (Zfp-103) (KF-1) (hKF-1). | |||||
|
RN103_MOUSE
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.035852 (rank : 47) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R1W3, O08670, O08883 | Gene names | Rnf103, Zfp103 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 103 (Zinc finger protein 103) (Zfp-103) (KF-1) (mKF-1). | |||||
|
RN123_MOUSE
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.029894 (rank : 53) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q5XPI3, Q6PGE0 | Gene names | Rnf123, Kpc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase subunit KPC1 (EC 6.3.2.-) (Kip1 ubiquitination-promoting complex protein 1) (RING finger protein 123). | |||||
|
SLK_HUMAN
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.002669 (rank : 94) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1775 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9H2G2, O00211, Q6P1Z4, Q86WU7, Q86WW1, Q92603, Q9NQL0, Q9NQL1 | Gene names | SLK, KIAA0204, STK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | STE20-like serine/threonine-protein kinase (EC 2.7.11.1) (STE20-like kinase) (STE20-related serine/threonine-protein kinase) (STE20-related kinase) (hSLK) (Serine/threonine-protein kinase 2) (CTCL tumor antigen se20-9). | |||||
|
AATF_MOUSE
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | 0.018311 (rank : 68) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JKX4, Q7TQN1, Q8C5Q2, Q99P89 | Gene names | Aatf, Che1, Trb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein AATF (Apoptosis-antagonizing transcription factor) (Rb-binding protein Che-1) (Traube protein). | |||||
|
CGRF1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.096033 (rank : 23) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BMJ7, Q3U3F1, Q8CI24 | Gene names | Cgrrf1, Cgr19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with RING finger domain 1 (Cell growth regulatory gene 19 protein). | |||||
|
GIDRP_MOUSE
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.024226 (rank : 59) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BJM3, Q3UGJ8, Q5U5U8 | Gene names | Gidrp88, D19Ertd386e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Growth inhibition and differentiation-related protein 88 homolog. | |||||
|
NOP56_HUMAN
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | 0.018075 (rank : 69) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O00567, Q9NQ05 | Gene names | NOL5A, NOP56 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar protein Nop56 (Nucleolar protein 5A). | |||||
|
RBCC1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.008045 (rank : 89) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1048 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9ESK9, Q3TXX2, Q61384, Q8BRY9, Q9JK14 | Gene names | Rb1cc1, Cc1, Kiaa0203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RB1-inducible coiled-coil protein 1 (Coiled-coil-forming protein 1) (LaXp180 protein). | |||||
|
RBMX2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.012056 (rank : 81) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y388, Q5JY82, Q9Y3I8 | Gene names | RBMX2 | |||
|
Domain Architecture |
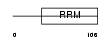
|
|||||
| Description | RNA-binding motif protein, X-linked 2. | |||||
|
RIMB1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.005617 (rank : 92) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
RPC5_MOUSE
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.015998 (rank : 73) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CZT4, Q8CI35, Q9DBD1 | Gene names | Polr3e, Sin | |||
|
Domain Architecture |
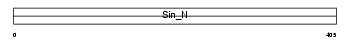
|
|||||
| Description | DNA-directed RNA polymerases III 80 kDa polypeptide (EC 2.7.7.6) (RNA polymerase III subunit 5) (RPC5) (Sex-lethal interactor homolog). | |||||
|
SLK_MOUSE
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.002313 (rank : 95) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1798 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O54988, Q80U65, Q8CAU2, Q8CDW2, Q9WU41 | Gene names | Slk, Kiaa0204, Stk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | STE20-like serine/threonine-protein kinase (EC 2.7.11.1) (STE20-like kinase) (STE20-related serine/threonine-protein kinase) (STE20-related kinase) (mSLK) (Serine/threonine-protein kinase 2) (STE20-related kinase SMAK) (Etk4). | |||||
|
TXND2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.030276 (rank : 50) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
ACHA6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.006844 (rank : 91) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15825 | Gene names | CHRNA6 | |||
|
Domain Architecture |
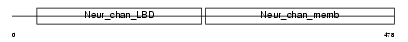
|
|||||
| Description | Neuronal acetylcholine receptor protein subunit alpha-6 precursor. | |||||
|
ASPX_MOUSE
|
||||||
| θ value | 8.99809 (rank : 81) | NC score | 0.017110 (rank : 70) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P50289, Q9DAM6 | Gene names | Acrv1 | |||
|
Domain Architecture |

|
|||||
| Description | Acrosomal protein SP-10 precursor (Acrosomal vesicle protein 1) (MSA- 63). | |||||
|
ATS18_MOUSE
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.002675 (rank : 93) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q4VC17, Q8BZD1 | Gene names | Adamts18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAMTS-18 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 18) (ADAM-TS 18) (ADAM-TS18). | |||||
|
CCAR1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 83) | NC score | 0.016677 (rank : 72) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8CH18, Q6AXC9, Q6PAR2, Q80XE4, Q8BJY0, Q8BVN2, Q8CGG1, Q9CSR5 | Gene names | Ccar1, Carp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle and apoptosis regulator protein 1 (Cell cycle and apoptosis regulatory protein 1) (CARP-1). | |||||
|
CHD1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.014238 (rank : 77) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
FXR2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.010697 (rank : 85) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9WVR4, Q9WVR5 | Gene names | Fxr2, Fxr2h | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation syndrome-related protein 2. | |||||
|
PKD2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.007534 (rank : 90) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13563, O60441, Q15764 | Gene names | PKD2 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystin-2 (Polycystic kidney disease 2 protein homolog) (Autosomal dominant polycystic kidney disease type II protein) (Polycystwin) (R48321). | |||||
|
RPGR_MOUSE
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.025699 (rank : 56) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9R0X5, O88408, Q9CU92 | Gene names | Rpgr | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator (mRpgr). | |||||
|
SFRS4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.016718 (rank : 71) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8VE97, Q9JJC3 | Gene names | Sfrs4 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4. | |||||
|
SOX6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.011376 (rank : 83) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P35712, Q9BXQ3, Q9BXQ4, Q9BXQ5, Q9H0I8 | Gene names | SOX6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-6. | |||||
|
SOX6_MOUSE
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.011106 (rank : 84) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P40645, Q62250, Q9QWS5 | Gene names | Sox6, Sox-6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-6 (SOX-LZ). | |||||
|
WDHD1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.008415 (rank : 87) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O75717 | Gene names | WDHD1, AND1 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat and HMG-box DNA-binding protein 1 (Acidic nucleoplasmic DNA- binding protein 1) (And-1). | |||||
|
ZN330_MOUSE
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.013472 (rank : 79) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q922H9, Q8C389, Q8K2M4 | Gene names | Znf330, Noa36, Zfp330 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 330 (Nucleolar cysteine-rich protein) (Nucleolar autoantigen 36). | |||||
|
BIRC5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.050011 (rank : 37) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O15392, Q2I3N8, Q4VGX0, Q53F61, Q5MGC6, Q6FHL2, Q75SP2, Q9P2W8 | Gene names | BIRC5, API4, IAP4 | |||
|
Domain Architecture |
|
|||||
| Description | Baculoviral IAP repeat-containing protein 5 (Apoptosis inhibitor survivin) (Apoptosis inhibitor 4). | |||||
|
BIRC8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.109674 (rank : 20) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q96P09, Q96RW5 | Gene names | BIRC8, ILP2 | |||
|
Domain Architecture |
|
|||||
| Description | Baculoviral IAP repeat-containing protein 8 (Inhibitor of apoptosis- like protein 2) (IAP-like protein 2) (ILP-2) (Testis-specific inhibitor of apoptosis). | |||||
|
CGRF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.082422 (rank : 25) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q99675, Q96BX2 | Gene names | CGRRF1, CGR19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with RING finger domain 1 (Cell growth regulatory gene 19 protein). | |||||
|
RKHD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.081867 (rank : 26) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q86XN8 | Gene names | RKHD1, KIAA2031 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger and KH domain-containing protein 1. | |||||
|
RNF26_HUMAN
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.097626 (rank : 22) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9BY78 | Gene names | RNF26 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 26. | |||||
|
MDM2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 92 | |
| SwissProt Accessions | Q00987, Q13226, Q13297, Q13298, Q13299, Q13300, Q13301, Q53XW0, Q71TW9, Q9UGI3, Q9UMT8 | Gene names | MDM2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3 Mdm2 (EC 6.3.2.-) (p53-binding protein Mdm2) (Oncoprotein Mdm2) (Double minute 2 protein) (Hdm2). | |||||
|
MDM2_MOUSE
|
||||||
| NC score | 0.975358 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P23804, Q61040, Q64330 | Gene names | Mdm2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3 Mdm2 (EC 6.3.2.-) (p53-binding protein Mdm2) (Oncoprotein Mdm2) (Double minute 2 protein). | |||||
|
MDM4_HUMAN
|
||||||
| NC score | 0.877328 (rank : 3) | θ value | 8.55514e-59 (rank : 3) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O15151, Q6GS18, Q8IV83 | Gene names | MDM4, MDMX | |||
|
Domain Architecture |

|
|||||
| Description | Mdm4 protein (p53-binding protein Mdm4) (Mdm2-like p53-binding protein) (Mdmx protein) (Double minute 4 protein). | |||||
|
MDM4_MOUSE
|
||||||
| NC score | 0.870463 (rank : 4) | θ value | 2.10721e-57 (rank : 4) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O35618 | Gene names | Mdm4, Mdmx | |||
|
Domain Architecture |

|
|||||
| Description | Mdm4 protein (p53-binding protein Mdm4) (Mdm2-like p53-binding protein) (Mdmx protein) (Double minute 4 protein). | |||||
|
NEURL_MOUSE
|
||||||
| NC score | 0.263297 (rank : 5) | θ value | 0.00134147 (rank : 5) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q923S6, Q91ZT6, Q9CWF1, Q9QWF1 | Gene names | Neurl, Neurl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuralized-like protein 1 (m-neuralized 1) (m-neu1). | |||||
|
NEUL1_HUMAN
|
||||||
| NC score | 0.259012 (rank : 6) | θ value | 0.00175202 (rank : 6) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O76050, Q5TDR2, Q5TDR3, Q8TAN0, Q9H463 | Gene names | NEURL, NEURL1, RNF67 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuralized-like protein 1 (h-neuralized 1) (h-neu) (RING finger protein 67). | |||||
|
RNF34_MOUSE
|
||||||
| NC score | 0.208226 (rank : 7) | θ value | 0.0330416 (rank : 8) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q99KR6, Q3UV45 | Gene names | Rnf34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 34 (RING finger protein RIFF) (Phafin-1). | |||||
|
RNF34_HUMAN
|
||||||
| NC score | 0.206869 (rank : 8) | θ value | 0.0431538 (rank : 9) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q969K3, Q8NG47, Q9H6W8 | Gene names | RNF34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 34 (RING finger protein RIFF) (FYVE-RING finger protein Momo) (Human RING finger homologous to inhibitor of apoptosis protein) (hRFI) (Caspases-8 and -10-associated RING finger protein 1) (CARP-1) (Caspase regulator CARP1). | |||||
|
RFFL_MOUSE
|
||||||
| NC score | 0.195692 (rank : 9) | θ value | 0.21417 (rank : 17) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6ZQM0, Q5SVC2, Q5SVC4, Q9D543, Q9D9B1 | Gene names | Rffl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rififylin (RING finger and FYVE-like domain-containing protein 1) (Fring). | |||||
|
RFFL_HUMAN
|
||||||
| NC score | 0.191739 (rank : 10) | θ value | 0.279714 (rank : 19) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8WZ73, Q8NHW0, Q8TBY7, Q96BE6 | Gene names | RFFL, RNF189, RNF34L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rififylin (RING finger and FYVE-like domain-containing protein 1) (FYVE-RING finger protein Sakura) (Fring) (Caspases-8 and -10- associated RING finger protein 2) (CARP-2) (Caspase regulator CARP2) (RING finger protein 189) (RING finger protein 34-like). | |||||
|
MGRN1_MOUSE
|
||||||
| NC score | 0.179191 (rank : 11) | θ value | 0.125558 (rank : 13) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9D074, Q3U5V9, Q3UDA1, Q6ZQ97, Q8BZM9 | Gene names | Mgrn1, Kiaa0544 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase MGRN1 (EC 6.3.2.-) (Mahogunin ring finger protein 1). | |||||
|
MGRN1_HUMAN
|
||||||
| NC score | 0.173300 (rank : 12) | θ value | 0.21417 (rank : 16) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O60291, Q86W76 | Gene names | MGRN1, KIAA0544, RNF156 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase MGRN1 (EC 6.3.2.-) (Mahogunin ring finger protein 1) (RING finger protein 156). | |||||
|
BIRC3_HUMAN
|
||||||
| NC score | 0.171199 (rank : 13) | θ value | 0.0148317 (rank : 7) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q13489, Q16628, Q9HC27, Q9UP46 | Gene names | BIRC3, API2, IAP1, MIHC | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 3 (Inhibitor of apoptosis protein 1) (HIAP1) (HIAP-1) (C-IAP2) (TNFR2-TRAF signaling complex protein 1) (IAP homolog C) (Apoptosis inhibitor 2) (API2). | |||||
|
BIRC2_MOUSE
|
||||||
| NC score | 0.168592 (rank : 14) | θ value | 0.0961366 (rank : 11) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q62210, O08864 | Gene names | Birc2, Birc3, Iap2 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 2 (Inhibitor of apoptosis protein 2) (MIAP2) (MIAP-2). | |||||
|
BIRC3_MOUSE
|
||||||
| NC score | 0.167103 (rank : 15) | θ value | 0.0961366 (rank : 12) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O08863 | Gene names | Birc3, Birc2, Iap1 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 3 (Inhibitor of apoptosis protein 1) (MIAP1) (MIAP-1). | |||||
|
BIRC2_HUMAN
|
||||||
| NC score | 0.152249 (rank : 16) | θ value | 0.365318 (rank : 20) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13490, Q16516, Q4TTG0 | Gene names | BIRC2, API1, IAP2, MIHB | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 2 (Inhibitor of apoptosis protein 2) (HIAP2) (HIAP-2) (C-IAP1) (TNFR2-TRAF signaling complex protein 2) (IAP homolog B). | |||||
|
BIRC4_MOUSE
|
||||||
| NC score | 0.149358 (rank : 17) | θ value | 0.62314 (rank : 25) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q60989, O08865 | Gene names | Birc4, Aipa, Api3, Miha, Xiap | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 4 (Inhibitor of apoptosis protein 3) (X-linked inhibitor of apoptosis protein) (X-linked IAP) (IAP homolog A) (MIAP3) (MIAP-3). | |||||
|
BIRC4_HUMAN
|
||||||
| NC score | 0.136958 (rank : 18) | θ value | 0.813845 (rank : 30) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P98170, Q9NQ14 | Gene names | BIRC4, API3, IAP3, XIAP | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 4 (Inhibitor of apoptosis protein 3) (X-linked inhibitor of apoptosis protein) (X-linked IAP) (IAP-like protein) (HILP). | |||||
|
BIRC7_HUMAN
|
||||||
| NC score | 0.114865 (rank : 19) | θ value | 4.03905 (rank : 52) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96CA5, Q9BQV0, Q9H2A8, Q9HAP7 | Gene names | BIRC7, KIAP, LIVIN, MLIAP | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 7 (Kidney inhibitor of apoptosis protein) (KIAP) (Melanoma inhibitor of apoptosis protein) (ML-IAP) (Livin). | |||||
|
BIRC8_HUMAN
|
||||||
| NC score | 0.109674 (rank : 20) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q96P09, Q96RW5 | Gene names | BIRC8, ILP2 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 8 (Inhibitor of apoptosis- like protein 2) (IAP-like protein 2) (ILP-2) (Testis-specific inhibitor of apoptosis). | |||||
|
TOPRS_MOUSE
|
||||||
| NC score | 0.106765 (rank : 21) | θ value | 0.0431538 (rank : 10) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q80Z37, Q3U5B5, Q3UPZ1, Q3USN3, Q3UZ55, Q8BXP2, Q8CFF5, Q8CGC8, Q920L3 | Gene names | Topors | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein E3 ligase Topors (EC 6.3.2.-) (SUMO1-protein E3 ligase Topors) (Topoisomerase I-binding RING finger protein) (Topoisomerase I-binding arginine/serine-rich protein) (Tumor suppressor p53-binding protein 3) (p53-binding protein 3) (p53BP3). | |||||
|
RNF26_HUMAN
|
||||||
| NC score | 0.097626 (rank : 22) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9BY78 | Gene names | RNF26 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 26. | |||||
|
CGRF1_MOUSE
|
||||||
| NC score | 0.096033 (rank : 23) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BMJ7, Q3U3F1, Q8CI24 | Gene names | Cgrrf1, Cgr19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with RING finger domain 1 (Cell growth regulatory gene 19 protein). | |||||
|
TOPRS_HUMAN
|
||||||
| NC score | 0.088152 (rank : 24) | θ value | 1.38821 (rank : 38) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9NS56, O43273, Q6P987, Q9NS55, Q9UNR9 | Gene names | TOPORS, LUN, TP53BPL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein E3 ligase Topors (EC 6.3.2.-) (SUMO1-protein E3 ligase Topors) (Topoisomerase I-binding RING finger protein) (Topoisomerase I-binding arginine/serine-rich protein) (Tumor suppressor p53-binding protein 3) (p53-binding protein 3) (p53BP3). | |||||
|
CGRF1_HUMAN
|
||||||
| NC score | 0.082422 (rank : 25) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q99675, Q96BX2 | Gene names | CGRRF1, CGR19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with RING finger domain 1 (Cell growth regulatory gene 19 protein). | |||||
|
RKHD1_HUMAN
|
||||||
| NC score | 0.081867 (rank : 26) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q86XN8 | Gene names | RKHD1, KIAA2031 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger and KH domain-containing protein 1. | |||||
|
MYLIP_HUMAN
|
||||||
| NC score | 0.075786 (rank : 27) | θ value | 1.06291 (rank : 35) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8WY64, Q9BU73, Q9NRL9, Q9UHE7 | Gene names | MYLIP, BZF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase MYLIP (EC 6.3.2.-) (Myosin regulatory light chain- interacting protein) (MIR). | |||||
|
LRSM1_MOUSE
|
||||||
| NC score | 0.073522 (rank : 28) | θ value | 0.62314 (rank : 26) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q80ZI6 | Gene names | Lrsam1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein LRSAM1 (EC 6.3.2.-) (Leucine-rich repeat and sterile alpha motif-containing protein 1) (Tsg101-associated ligase). | |||||
|
MIB1_MOUSE
|
||||||
| NC score | 0.069480 (rank : 29) | θ value | 0.365318 (rank : 22) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q80SY4, Q5XK51, Q6IS57, Q6YI52, Q6ZPT8, Q8BNR1, Q8C6W2, Q921Q1 | Gene names | Mib1, Dip1, Kiaa1323, Mib | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein MIB1 (EC 6.3.2.-) (Mind bomb homolog 1) (DAPK-interacting protein 1) (DIP-1). | |||||
|
LRSM1_HUMAN
|
||||||
| NC score | 0.068589 (rank : 30) | θ value | 0.21417 (rank : 15) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1078 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q6UWE0, Q5VVV0, Q8NB40, Q96GT5, Q96MX5, Q96MZ7 | Gene names | LRSAM1, TAL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein LRSAM1 (EC 6.3.2.-) (Leucine-rich repeat and sterile alpha motif-containing protein 1) (Tsg101-associated ligase) (hTAL). | |||||
|
MIB1_HUMAN
|
||||||
| NC score | 0.068212 (rank : 31) | θ value | 0.365318 (rank : 21) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q86YT6, Q68D01, Q6YI51, Q8NBY0, Q8TCB5, Q8TCL7, Q9P2M3 | Gene names | MIB1, DIP1, KIAA1323, ZZANK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein MIB1 (EC 6.3.2.-) (Mind bomb homolog 1) (DAPK-interacting protein 1) (DIP-1) (Zinc finger ZZ type with ankyrin repeat domain protein 2). | |||||
|
MYLIP_MOUSE
|
||||||
| NC score | 0.067795 (rank : 32) | θ value | 1.81305 (rank : 40) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8BM54, Q3TX01, Q8BLY9, Q91Z47 | Gene names | Mylip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase MYLIP (EC 6.3.2.-) (Myosin regulatory light chain- interacting protein) (MIR). | |||||
|
CHFR_MOUSE
|
||||||
| NC score | 0.065611 (rank : 33) | θ value | 0.47712 (rank : 23) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q810L3, Q8BJZ9, Q8BWH4 | Gene names | Chfr | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase CHFR (EC 6.3.2.-) (Checkpoint with forkhead and RING finger domains protein). | |||||
|
ZRAB2_HUMAN
|
||||||
| NC score | 0.061035 (rank : 34) | θ value | 0.813845 (rank : 34) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O95218, Q9UP63 | Gene names | ZRANB2, ZIS, ZNF265 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger Ran-binding domain-containing protein 2 Zinc finger protein 265 (Zinc finger, splicing). | |||||
|
DSPP_MOUSE
|
||||||
| NC score | 0.053137 (rank : 35) | θ value | 0.813845 (rank : 31) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.050954 (rank : 36) | θ value | 0.279714 (rank : 18) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
BIRC5_HUMAN
|
||||||
| NC score | 0.050011 (rank : 37) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O15392, Q2I3N8, Q4VGX0, Q53F61, Q5MGC6, Q6FHL2, Q75SP2, Q9P2W8 | Gene names | BIRC5, API4, IAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 5 (Apoptosis inhibitor survivin) (Apoptosis inhibitor 4). | |||||
|
RN103_HUMAN
|
||||||
| NC score | 0.045771 (rank : 38) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O00237, Q8IVB9 | Gene names | RNF103 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 103 (Zinc finger protein 103 homolog) (Zfp-103) (KF-1) (hKF-1). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.045149 (rank : 39) | θ value | 0.813845 (rank : 33) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
FANCM_HUMAN
|
||||||
| NC score | 0.041645 (rank : 40) | θ value | 0.163984 (rank : 14) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IYD8, Q3YFH9, Q8N9X6, Q9HCH6 | Gene names | FANCM, KIAA1596 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group M protein (EC 3.6.1.-) (ATP-dependent RNA helicase FANCM) (Protein FACM) (Fanconi anemia-associated polypeptide of 250 kDa) (FAAP250) (Protein Hef ortholog). | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.041007 (rank : 41) | θ value | 1.38821 (rank : 36) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.040884 (rank : 42) | θ value | 3.0926 (rank : 48) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
TRI37_MOUSE
|
||||||
| NC score | 0.037730 (rank : 43) | θ value | 4.03905 (rank : 58) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 316 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6PCX9, Q8CHC5 | Gene names | Trim37, Kiaa0898 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 37. | |||||
|
TRI37_HUMAN
|
||||||
| NC score | 0.037679 (rank : 44) | θ value | 4.03905 (rank : 57) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O94972, Q7Z3E6, Q8IYF7, Q8WYF7 | Gene names | TRIM37, KIAA0898, MUL, POB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 37 (Mulibrey nanism protein). | |||||
|
ZDH21_MOUSE
|
||||||
| NC score | 0.036860 (rank : 45) | θ value | 0.62314 (rank : 29) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D270, Q6EMK1, Q80XQ3 | Gene names | Zdhhc21, Gramp3 | |||
|
Domain Architecture |

|
|||||
| Description | Probable palmitoyltransferase ZDHHC21 (EC 2.3.1.-) (Zinc finger DHHC domain-containing protein 21) (DHHC-21) (GABA-A receptor-associated membrane protein 3). | |||||
|
ZDH21_HUMAN
|
||||||
| NC score | 0.036799 (rank : 46) | θ value | 0.62314 (rank : 28) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IVQ6, Q5VWG1 | Gene names | ZDHHC21 | |||
|
Domain Architecture |

|
|||||
| Description | Probable palmitoyltransferase ZDHHC21 (EC 2.3.1.-) (Zinc finger DHHC domain-containing protein 21) (DHHC-21). | |||||
|
RN103_MOUSE
|
||||||
| NC score | 0.035852 (rank : 47) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R1W3, O08670, O08883 | Gene names | Rnf103, Zfp103 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 103 (Zinc finger protein 103) (Zfp-103) (KF-1) (mKF-1). | |||||
|
CM007_MOUSE
|
||||||
| NC score | 0.034063 (rank : 48) | θ value | 4.03905 (rank : 53) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K2Y0, Q9CYU1, Q9D1Y0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C13orf7 homolog. | |||||
|
ABTAP_MOUSE
|
||||||
| NC score | 0.032616 (rank : 49) | θ value | 2.36792 (rank : 41) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q3V1V3 | Gene names | Abtap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ABT1-associated protein. | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.030276 (rank : 50) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
PHF14_HUMAN
|
||||||
| NC score | 0.030254 (rank : 51) | θ value | 2.36792 (rank : 43) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
PHF14_MOUSE
|
||||||
| NC score | 0.030025 (rank : 52) | θ value | 2.36792 (rank : 44) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
RN123_MOUSE
|
||||||
| NC score | 0.029894 (rank : 53) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q5XPI3, Q6PGE0 | Gene names | Rnf123, Kpc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase subunit KPC1 (EC 6.3.2.-) (Kip1 ubiquitination-promoting complex protein 1) (RING finger protein 123). | |||||
|
RPGR_HUMAN
|
||||||
| NC score | 0.029036 (rank : 54) | θ value | 1.38821 (rank : 37) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92834, O00702, O00737, Q8N5T6, Q93039, Q9HD29, Q9UMR1 | Gene names | RPGR, RP3, XLRP3 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator. | |||||
|
AN32A_HUMAN
|
||||||
| NC score | 0.028943 (rank : 55) | θ value | 0.62314 (rank : 24) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P39687 | Gene names | ANP32A, C15orf1, LANP, MAPM, PHAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member A (Potent heat-stable protein phosphatase 2A inhibitor I1PP2A) (Acidic nuclear phosphoprotein pp32) (Leucine-rich acidic nuclear protein) (Lanp) (Putative HLA-DR-associated protein I) (PHAPI) (Mapmodulin). | |||||
|
RPGR_MOUSE
|
||||||
| NC score | 0.025699 (rank : 56) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9R0X5, O88408, Q9CU92 | Gene names | Rpgr | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator (mRpgr). | |||||
|
FA44A_HUMAN
|
||||||
| NC score | 0.024900 (rank : 57) | θ value | 5.27518 (rank : 63) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8NFC6, Q6P0M8 | Gene names | FAM44A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM44A. | |||||
|
RPGR1_MOUSE
|
||||||
| NC score | 0.024892 (rank : 58) | θ value | 3.0926 (rank : 51) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 713 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9EPQ2, Q8CAC2, Q8CDJ9, Q91WE0, Q9CUK6, Q9D5Q1 | Gene names | Rpgrip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator-interacting protein 1 (RPGR-interacting protein 1). | |||||
|
GIDRP_MOUSE
|
||||||
| NC score | 0.024226 (rank : 59) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BJM3, Q3UGJ8, Q5U5U8 | Gene names | Gidrp88, D19Ertd386e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Growth inhibition and differentiation-related protein 88 homolog. | |||||
|
FA13C_HUMAN
|
||||||
| NC score | 0.024060 (rank : 60) | θ value | 2.36792 (rank : 42) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NE31, Q99787 | Gene names | FAM13C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13C1. | |||||
|
NALP5_MOUSE
|
||||||
| NC score | 0.022903 (rank : 61) | θ value | 0.813845 (rank : 32) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9R1M5, Q9JLR2 | Gene names | Nalp5, Mater | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 5 (Maternal antigen that embryos require) (Mater protein) (Ooplasm-specific protein 1) (OP1). | |||||
|
DDIT3_HUMAN
|
||||||
| NC score | 0.022177 (rank : 62) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P35638 | Gene names | DDIT3, CHOP, GADD153 | |||
|
Domain Architecture |

|
|||||
| Description | DNA damage-inducible transcript 3 (DDIT-3) (Growth arrest and DNA- damage-inducible protein GADD153) (C/EBP-homologous protein) (CHOP). | |||||
|
NP1L1_HUMAN
|
||||||
| NC score | 0.020358 (rank : 63) | θ value | 4.03905 (rank : 54) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55209 | Gene names | NAP1L1, NRP | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome assembly protein 1-like 1 (NAP-1-related protein) (hNRP). | |||||
|
NP1L1_MOUSE
|
||||||
| NC score | 0.020338 (rank : 64) | θ value | 4.03905 (rank : 55) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P28656, Q3UL14 | Gene names | Nap1l1, Nrp | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome assembly protein 1-like 1 (NAP-1-related protein) (Brain protein DN38). | |||||
|
RBP2_MOUSE
|
||||||
| NC score | 0.020241 (rank : 65) | θ value | 0.62314 (rank : 27) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9ERU9, Q61992, Q8C9K9 | Gene names | Ranbp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2 (RanBP2). | |||||
|
RBM16_HUMAN
|
||||||
| NC score | 0.018984 (rank : 66) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 300 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UPN6, Q5TBU6, Q9BQN8, Q9BX43 | Gene names | RBM16, KIAA1116 | |||
|
Domain Architecture |

|
|||||
| Description | Putative RNA-binding protein 16 (RNA-binding motif protein 16). | |||||
|
TXND2_HUMAN
|
||||||
| NC score | 0.018572 (rank : 67) | θ value | 4.03905 (rank : 59) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
AATF_MOUSE
|
||||||
| NC score | 0.018311 (rank : 68) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JKX4, Q7TQN1, Q8C5Q2, Q99P89 | Gene names | Aatf, Che1, Trb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein AATF (Apoptosis-antagonizing transcription factor) (Rb-binding protein Che-1) (Traube protein). | |||||
|
NOP56_HUMAN
|
||||||
| NC score | 0.018075 (rank : 69) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O00567, Q9NQ05 | Gene names | NOL5A, NOP56 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar protein Nop56 (Nucleolar protein 5A). | |||||
|
ASPX_MOUSE
|
||||||
| NC score | 0.017110 (rank : 70) | θ value | 8.99809 (rank : 81) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P50289, Q9DAM6 | Gene names | Acrv1 | |||
|
Domain Architecture |

|
|||||
| Description | Acrosomal protein SP-10 precursor (Acrosomal vesicle protein 1) (MSA- 63). | |||||
|
SFRS4_MOUSE
|
||||||
| NC score | 0.016718 (rank : 71) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8VE97, Q9JJC3 | Gene names | Sfrs4 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4. | |||||
|
CCAR1_MOUSE
|
||||||
| NC score | 0.016677 (rank : 72) | θ value | 8.99809 (rank : 83) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8CH18, Q6AXC9, Q6PAR2, Q80XE4, Q8BJY0, Q8BVN2, Q8CGG1, Q9CSR5 | Gene names | Ccar1, Carp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle and apoptosis regulator protein 1 (Cell cycle and apoptosis regulatory protein 1) (CARP-1). | |||||
|
RPC5_MOUSE
|
||||||
| NC score | 0.015998 (rank : 73) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CZT4, Q8CI35, Q9DBD1 | Gene names | Polr3e, Sin | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerases III 80 kDa polypeptide (EC 2.7.7.6) (RNA polymerase III subunit 5) (RPC5) (Sex-lethal interactor homolog). | |||||
|
CHD9_MOUSE
|
||||||
| NC score | 0.015882 (rank : 74) | θ value | 3.0926 (rank : 47) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BYH8, Q7TMV5, Q8BJG8, Q8BZJ2, Q8CHG8 | Gene names | Chd9, Kiaa0308, Pric320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha- interacting complex protein 320 kDa). | |||||
|
CHD1_HUMAN
|
||||||
| NC score | 0.015783 (rank : 75) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
SESN1_MOUSE
|
||||||
| NC score | 0.014357 (rank : 76) | θ value | 4.03905 (rank : 56) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P58006, Q7TNF3 | Gene names | Sesn1, Pa26, Sest1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sestrin-1 (p53-regulated protein PA26). | |||||
|
CHD1_MOUSE
|
||||||
| NC score | 0.014238 (rank : 77) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
CR034_MOUSE
|
||||||
| NC score | 0.013957 (rank : 78) | θ value | 1.81305 (rank : 39) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 643 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CDV0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C18orf34 homolog. | |||||
|
ZN330_MOUSE
|
||||||
| NC score | 0.013472 (rank : 79) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q922H9, Q8C389, Q8K2M4 | Gene names | Znf330, Noa36, Zfp330 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 330 (Nucleolar cysteine-rich protein) (Nucleolar autoantigen 36). | |||||
|
DPF3_MOUSE
|
||||||
| NC score | 0.012106 (rank : 80) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P58269 | Gene names | Dpf3, Cerd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc-finger protein Dpf3 (cer-d4). | |||||
|
RBMX2_HUMAN
|
||||||
| NC score | 0.012056 (rank : 81) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y388, Q5JY82, Q9Y3I8 | Gene names | RBMX2 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding motif protein, X-linked 2. | |||||
|
LEF1_MOUSE
|
||||||
| NC score | 0.011583 (rank : 82) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P27782 | Gene names | Lef1, Lef-1 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphoid enhancer-binding factor 1 (LEF-1). | |||||
|
SOX6_HUMAN
|
||||||
| NC score | 0.011376 (rank : 83) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P35712, Q9BXQ3, Q9BXQ4, Q9BXQ5, Q9H0I8 | Gene names | SOX6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-6. | |||||
|
SOX6_MOUSE
|
||||||
| NC score | 0.011106 (rank : 84) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P40645, Q62250, Q9QWS5 | Gene names | Sox6, Sox-6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-6 (SOX-LZ). | |||||
|
FXR2_MOUSE
|
||||||
| NC score | 0.010697 (rank : 85) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9WVR4, Q9WVR5 | Gene names | Fxr2, Fxr2h | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation syndrome-related protein 2. | |||||
|
PRP4B_HUMAN
|
||||||
| NC score | 0.008470 (rank : 86) | θ value | 3.0926 (rank : 49) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1030 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q13523, Q8TDP2, Q96QT7, Q9UEE6 | Gene names | PRPF4B, KIAA0536, PRP4, PRP4H, PRP4K | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PRP4 homolog (EC 2.7.11.1) (PRP4 pre- mRNA-processing factor 4 homolog) (PRP4 kinase). | |||||
|
WDHD1_HUMAN
|
||||||
| NC score | 0.008415 (rank : 87) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O75717 | Gene names | WDHD1, AND1 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat and HMG-box DNA-binding protein 1 (Acidic nucleoplasmic DNA- binding protein 1) (And-1). | |||||
|
PRP4B_MOUSE
|
||||||
| NC score | 0.008304 (rank : 88) | θ value | 3.0926 (rank : 50) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1040 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q61136, O88378, Q8BND8, Q8R4Y5, Q9CTL9, Q9CTT0 | Gene names | Prpf4b, Cbp143, Prp4h, Prp4k, Prp4m, Prpk | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PRP4 homolog (EC 2.7.11.1) (PRP4 pre- mRNA-processing factor 4 homolog) (Pre-mRNA protein kinase). | |||||
|
RBCC1_MOUSE
|
||||||
| NC score | 0.008045 (rank : 89) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1048 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9ESK9, Q3TXX2, Q61384, Q8BRY9, Q9JK14 | Gene names | Rb1cc1, Cc1, Kiaa0203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RB1-inducible coiled-coil protein 1 (Coiled-coil-forming protein 1) (LaXp180 protein). | |||||
|
PKD2_HUMAN
|
||||||
| NC score | 0.007534 (rank : 90) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13563, O60441, Q15764 | Gene names | PKD2 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystin-2 (Polycystic kidney disease 2 protein homolog) (Autosomal dominant polycystic kidney disease type II protein) (Polycystwin) (R48321). | |||||
|
ACHA6_HUMAN
|
||||||
| NC score | 0.006844 (rank : 91) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15825 | Gene names | CHRNA6 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal acetylcholine receptor protein subunit alpha-6 precursor. | |||||
|
RIMB1_MOUSE
|
||||||
| NC score | 0.005617 (rank : 92) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
ATS18_MOUSE
|
||||||
| NC score | 0.002675 (rank : 93) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q4VC17, Q8BZD1 | Gene names | Adamts18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAMTS-18 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 18) (ADAM-TS 18) (ADAM-TS18). | |||||
|
SLK_HUMAN
|
||||||
| NC score | 0.002669 (rank : 94) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1775 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9H2G2, O00211, Q6P1Z4, Q86WU7, Q86WW1, Q92603, Q9NQL0, Q9NQL1 | Gene names | SLK, KIAA0204, STK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | STE20-like serine/threonine-protein kinase (EC 2.7.11.1) (STE20-like kinase) (STE20-related serine/threonine-protein kinase) (STE20-related kinase) (hSLK) (Serine/threonine-protein kinase 2) (CTCL tumor antigen se20-9). | |||||
|
SLK_MOUSE
|
||||||
| NC score | 0.002313 (rank : 95) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1798 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O54988, Q80U65, Q8CAU2, Q8CDW2, Q9WU41 | Gene names | Slk, Kiaa0204, Stk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | STE20-like serine/threonine-protein kinase (EC 2.7.11.1) (STE20-like kinase) (STE20-related serine/threonine-protein kinase) (STE20-related kinase) (mSLK) (Serine/threonine-protein kinase 2) (STE20-related kinase SMAK) (Etk4). | |||||
|
BCL6_MOUSE
|
||||||
| NC score | 0.001461 (rank : 96) | θ value | 3.0926 (rank : 46) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 913 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P41183, Q61065 | Gene names | Bcl6, Bcl-6 | |||
|
Domain Architecture |

|
|||||
| Description | B-cell lymphoma 6 protein homolog. | |||||
|
BCL6_HUMAN
|
||||||
| NC score | 0.001364 (rank : 97) | θ value | 3.0926 (rank : 45) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P41182 | Gene names | BCL6, BCL5, LAZ3, ZBTB27, ZNF51 | |||
|
Domain Architecture |

|
|||||
| Description | B-cell lymphoma 6 protein (BCL-6) (Zinc finger protein 51) (LAZ-3 protein) (BCL-5) (Zinc finger and BTB domain-containing protein 27). | |||||