Please be patient as the page loads
|
AATF_MOUSE
|
||||||
| SwissProt Accessions | Q9JKX4, Q7TQN1, Q8C5Q2, Q99P89 | Gene names | Aatf, Che1, Trb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein AATF (Apoptosis-antagonizing transcription factor) (Rb-binding protein Che-1) (Traube protein). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
AATF_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.975449 (rank : 2) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9NY61, Q9P0A4, Q9UNX5 | Gene names | AATF, CHE1, DED | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein AATF (Apoptosis-antagonizing transcription factor) (Rb-binding protein Che-1). | |||||
|
AATF_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9JKX4, Q7TQN1, Q8C5Q2, Q99P89 | Gene names | Aatf, Che1, Trb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein AATF (Apoptosis-antagonizing transcription factor) (Rb-binding protein Che-1) (Traube protein). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 3) | NC score | 0.091668 (rank : 5) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
LEO1_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 4) | NC score | 0.105962 (rank : 3) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 5) | NC score | 0.082870 (rank : 7) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
DMD_HUMAN
|
||||||
| θ value | 0.125558 (rank : 6) | NC score | 0.028942 (rank : 27) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P11532, Q02295, Q14169, Q14170, Q5JYU0, Q7KZ48 | Gene names | DMD | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 7) | NC score | 0.096687 (rank : 4) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
ERIC1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 8) | NC score | 0.089004 (rank : 6) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
NIBA_HUMAN
|
||||||
| θ value | 0.279714 (rank : 9) | NC score | 0.058363 (rank : 12) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BZQ8, Q5TEM8, Q8TEI5, Q9H593, Q9H9Y8, Q9HCB9 | Gene names | C1orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Niban protein. | |||||
|
ATRX_HUMAN
|
||||||
| θ value | 0.47712 (rank : 10) | NC score | 0.047020 (rank : 16) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P46100, P51068, Q15886, Q59FB5, Q59H31, Q5H9A2, Q5JWI4, Q7Z2J1, Q9H0Z1, Q9NTS3 | Gene names | ATRX, RAD54L, XH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked helicase II) (X-linked nuclear protein) (XNP) (Znf- HX). | |||||
|
NSBP1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 11) | NC score | 0.062018 (rank : 10) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
SAFB2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 12) | NC score | 0.034835 (rank : 23) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 720 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q14151, Q8TB13 | Gene names | SAFB2, KIAA0138 | |||
|
Domain Architecture |

|
|||||
| Description | Scaffold attachment factor B2. | |||||
|
CP135_HUMAN
|
||||||
| θ value | 1.38821 (rank : 13) | NC score | 0.024799 (rank : 30) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1290 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q66GS9, O75130, Q9H8H7 | Gene names | CEP135, CEP4, KIAA0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
F10A1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 14) | NC score | 0.040250 (rank : 18) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P50502, O14999 | Gene names | ST13, FAM10A1, HIP, SNC6 | |||
|
Domain Architecture |
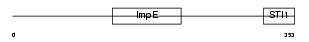
|
|||||
| Description | Hsc70-interacting protein (Hip) (Suppression of tumorigenicity protein 13) (Putative tumor suppressor ST13) (Protein FAM10A1) (Progesterone receptor-associated p48 protein) (NY-REN-33 antigen). | |||||
|
LAMA2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 15) | NC score | 0.015979 (rank : 47) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1198 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q60675, Q05003, Q64061 | Gene names | Lama2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
JIP4_MOUSE
|
||||||
| θ value | 2.36792 (rank : 16) | NC score | 0.027360 (rank : 28) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q58A65, Q3UH77, Q3UHF0, Q58VQ4, Q5NC70, Q5NC78, Q6A057, Q6PAS3, Q8CJC2 | Gene names | Spag9, Jip4, Jsap2, Kiaa0516, Mapk8ip4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 4 (JNK-interacting protein 4) (JIP-4) (JNK-associated leucine-zipper protein) (JLP) (Sperm-associated antigen 9) (Mitogen-activated protein kinase 8- interacting protein 4) (JNK/SAPK-associated protein 2) (JSAP2). | |||||
|
PRDM1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 17) | NC score | 0.002774 (rank : 55) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 766 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60636 | Gene names | Prdm1, Blimp1 | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 1 (PR domain-containing protein 1) (Beta-interferon gene positive-regulatory domain I-binding factor) (BLIMP-1). | |||||
|
SNPH_HUMAN
|
||||||
| θ value | 2.36792 (rank : 18) | NC score | 0.032092 (rank : 24) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O15079 | Gene names | SNPH, KIAA0374 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaphilin. | |||||
|
TAOK2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 19) | NC score | 0.007350 (rank : 52) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1172 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UL54, O94957, Q6UW73, Q7LC09, Q9NSW2 | Gene names | TAOK2, KIAA0881, MAP3K17, PSK, PSK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO2 (EC 2.7.11.1) (Thousand and one amino acid protein 2) (Prostate-derived STE20-like kinase 1) (PSK-1) (Kinase from chicken homolog C) (hKFC-C). | |||||
|
CJ064_HUMAN
|
||||||
| θ value | 3.0926 (rank : 20) | NC score | 0.030761 (rank : 25) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6ZS81, Q8N4A3 | Gene names | C10orf64 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C10orf64. | |||||
|
NEK1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 21) | NC score | 0.006105 (rank : 53) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P51954 | Gene names | Nek1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Nek1 (EC 2.7.11.1) (NimA-related protein kinase 1). | |||||
|
ANR21_HUMAN
|
||||||
| θ value | 4.03905 (rank : 22) | NC score | 0.016003 (rank : 46) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q86YR6 | Gene names | ANKRD21, POTE | |||
|
Domain Architecture |
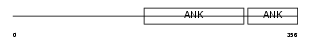
|
|||||
| Description | Ankyrin repeat domain-containing protein 21 (Protein POTE) (Prostate, ovary, testis-expressed protein). | |||||
|
BRDT_HUMAN
|
||||||
| θ value | 4.03905 (rank : 23) | NC score | 0.018310 (rank : 40) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q58F21, O14789, Q6P5T1, Q7Z4A6, Q8IWI6 | Gene names | BRDT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain testis-specific protein (RING3-like protein). | |||||
|
F10A5_HUMAN
|
||||||
| θ value | 4.03905 (rank : 24) | NC score | 0.035892 (rank : 21) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8NFI4 | Gene names | FAM10A5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM10A5. | |||||
|
MYH4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 25) | NC score | 0.018317 (rank : 38) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1377 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y623 | Gene names | MYH4 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-4 (Myosin heavy chain, skeletal muscle, fetal) (Myosin heavy chain IIb) (MyHC-IIb). | |||||
|
MYH8_HUMAN
|
||||||
| θ value | 4.03905 (rank : 26) | NC score | 0.017678 (rank : 41) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1610 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P13535, Q14910 | Gene names | MYH8 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
AFF2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 27) | NC score | 0.036690 (rank : 20) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P51816, O43786, O60215, P78407, Q13521, Q14323, Q9UNA5 | Gene names | AFF2, FMR2, OX19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation 2 protein) (Protein FMR-2) (FMR2P) (Protein Ox19) (Fragile X E mental retardation syndrome protein). | |||||
|
CAD22_MOUSE
|
||||||
| θ value | 5.27518 (rank : 28) | NC score | 0.004388 (rank : 54) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9WTP5 | Gene names | Cdh22 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-22 precursor (Pituitary and brain cadherin) (PB-cadherin). | |||||
|
DCDC2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 29) | NC score | 0.029586 (rank : 26) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q5DU00, Q5SZU0, Q80Y99 | Gene names | Dcdc2, Kiaa1154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublecortin domain-containing protein 2. | |||||
|
MAP4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 30) | NC score | 0.016084 (rank : 45) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P27546 | Gene names | Map4, Mtap4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
WAPL_MOUSE
|
||||||
| θ value | 5.27518 (rank : 31) | NC score | 0.024924 (rank : 29) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q65Z40, Q6ZQE9 | Gene names | Wapal, Kiaa0261, Wapl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wings apart-like protein homolog (Dioxin-inducible factor 2) (DIF-2). | |||||
|
CP135_MOUSE
|
||||||
| θ value | 6.88961 (rank : 32) | NC score | 0.020031 (rank : 35) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1278 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6P5D4, Q6A030 | Gene names | Cep135, Cep4, Kiaa0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
MDM2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 33) | NC score | 0.018311 (rank : 39) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q00987, Q13226, Q13297, Q13298, Q13299, Q13300, Q13301, Q53XW0, Q71TW9, Q9UGI3, Q9UMT8 | Gene names | MDM2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3 Mdm2 (EC 6.3.2.-) (p53-binding protein Mdm2) (Oncoprotein Mdm2) (Double minute 2 protein) (Hdm2). | |||||
|
MYEF2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 34) | NC score | 0.011680 (rank : 49) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C854, Q60690, Q6P930, Q6ZPT3, Q8QZZ1 | Gene names | Myef2, Kiaa1341, Mef2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin expression factor 2 (MyEF-2) (MEF-2). | |||||
|
MYH6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 35) | NC score | 0.017386 (rank : 43) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH8_MOUSE
|
||||||
| θ value | 6.88961 (rank : 36) | NC score | 0.016769 (rank : 44) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1600 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P13542, Q5SX36 | Gene names | Myh8, Myhsp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
RHG12_MOUSE
|
||||||
| θ value | 6.88961 (rank : 37) | NC score | 0.017425 (rank : 42) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C0D4, Q8BVP8 | Gene names | Arhgap12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
SPHK1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 38) | NC score | 0.019230 (rank : 37) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NYA1, Q9HD92, Q9NY70, Q9NYL3 | Gene names | SPHK1, SPHK, SPK | |||
|
Domain Architecture |
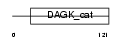
|
|||||
| Description | Sphingosine kinase 1 (EC 2.7.1.-) (SK 1) (SPK 1). | |||||
|
TESC_MOUSE
|
||||||
| θ value | 6.88961 (rank : 39) | NC score | 0.024062 (rank : 31) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JKL5 | Gene names | Tesc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tescalcin (TSC). | |||||
|
AFF2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 40) | NC score | 0.035446 (rank : 22) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O55112 | Gene names | Aff2, Fmr2, Ox19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation protein 2 homolog) (Protein FMR-2) (FMR2P) (Protein Ox19). | |||||
|
ANR26_MOUSE
|
||||||
| θ value | 8.99809 (rank : 41) | NC score | 0.015792 (rank : 48) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1554 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q811D2, Q69ZS2, Q9CS61 | Gene names | Ankrd26, Kiaa1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
CC47_MOUSE
|
||||||
| θ value | 8.99809 (rank : 42) | NC score | 0.023052 (rank : 32) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9D024, Q3TSB5, Q3V1S7, Q8C5D9, Q920S6 | Gene names | Ccdc47, Asp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 47 precursor (Adipocyte-specific protein 4). | |||||
|
KCNK5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 43) | NC score | 0.009890 (rank : 50) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95279 | Gene names | KCNK5, TASK2 | |||
|
Domain Architecture |
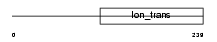
|
|||||
| Description | Potassium channel subfamily K member 5 (Acid-sensitive potassium channel protein TASK-2) (TWIK-related acid-sensitive K(+) channel 2). | |||||
|
LUZP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 44) | NC score | 0.019658 (rank : 36) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 940 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8R4U7, Q3UV18, Q7TS71, Q8BQW1, Q99NG3, Q9CSL6 | Gene names | Luzp1, Luzp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper protein 1 (Leucine zipper motif-containing protein). | |||||
|
MIA2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 45) | NC score | 0.020461 (rank : 34) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q91ZV0 | Gene names | Mia2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma inhibitory activity protein 2 precursor. | |||||
|
PKN3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 46) | NC score | 0.001643 (rank : 56) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6P5Z2, Q9UM03 | Gene names | PKN3, PKNBETA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase N3 (EC 2.7.11.13) (Protein kinase PKN- beta) (Protein-kinase C-related kinase 3). | |||||
|
PRRT3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 47) | NC score | 0.007484 (rank : 51) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
SEC63_HUMAN
|
||||||
| θ value | 8.99809 (rank : 48) | NC score | 0.041198 (rank : 17) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
SEC63_MOUSE
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.037534 (rank : 19) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
TESC_HUMAN
|
||||||
| θ value | 8.99809 (rank : 50) | NC score | 0.021462 (rank : 33) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96BS2 | Gene names | TESC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tescalcin (TSC). | |||||
|
NSBP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.058994 (rank : 11) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
NUCKS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.052666 (rank : 15) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
NUCKS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 53) | NC score | 0.058074 (rank : 14) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80XU3, Q8BVD8 | Gene names | Nucks1, Nucks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (JC7). | |||||
|
PTMA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 54) | NC score | 0.080354 (rank : 9) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P06454, Q15249, Q15592 | Gene names | PTMA, TMSA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha-1]. | |||||
|
PTMA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.082103 (rank : 8) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P26350, Q3UQV6 | Gene names | Ptma | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha]. | |||||
|
PTMS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 56) | NC score | 0.058361 (rank : 13) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9D0J8 | Gene names | Ptms | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
AATF_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9JKX4, Q7TQN1, Q8C5Q2, Q99P89 | Gene names | Aatf, Che1, Trb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein AATF (Apoptosis-antagonizing transcription factor) (Rb-binding protein Che-1) (Traube protein). | |||||
|
AATF_HUMAN
|
||||||
| NC score | 0.975449 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9NY61, Q9P0A4, Q9UNX5 | Gene names | AATF, CHE1, DED | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein AATF (Apoptosis-antagonizing transcription factor) (Rb-binding protein Che-1). | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.105962 (rank : 3) | θ value | 0.0252991 (rank : 4) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.096687 (rank : 4) | θ value | 0.163984 (rank : 7) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.091668 (rank : 5) | θ value | 0.0113563 (rank : 3) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
ERIC1_HUMAN
|
||||||
| NC score | 0.089004 (rank : 6) | θ value | 0.21417 (rank : 8) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.082870 (rank : 7) | θ value | 0.0961366 (rank : 5) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
PTMA_MOUSE
|
||||||
| NC score | 0.082103 (rank : 8) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P26350, Q3UQV6 | Gene names | Ptma | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha]. | |||||
|
PTMA_HUMAN
|
||||||
| NC score | 0.080354 (rank : 9) | θ value | θ > 10 (rank : 54) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P06454, Q15249, Q15592 | Gene names | PTMA, TMSA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha-1]. | |||||
|
NSBP1_MOUSE
|
||||||
| NC score | 0.062018 (rank : 10) | θ value | 0.62314 (rank : 11) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
NSBP1_HUMAN
|
||||||
| NC score | 0.058994 (rank : 11) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
NIBA_HUMAN
|
||||||
| NC score | 0.058363 (rank : 12) | θ value | 0.279714 (rank : 9) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BZQ8, Q5TEM8, Q8TEI5, Q9H593, Q9H9Y8, Q9HCB9 | Gene names | C1orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Niban protein. | |||||
|
PTMS_MOUSE
|
||||||
| NC score | 0.058361 (rank : 13) | θ value | θ > 10 (rank : 56) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9D0J8 | Gene names | Ptms | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
NUCKS_MOUSE
|
||||||
| NC score | 0.058074 (rank : 14) | θ value | θ > 10 (rank : 53) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80XU3, Q8BVD8 | Gene names | Nucks1, Nucks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (JC7). | |||||
|
NUCKS_HUMAN
|
||||||
| NC score | 0.052666 (rank : 15) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
ATRX_HUMAN
|
||||||
| NC score | 0.047020 (rank : 16) | θ value | 0.47712 (rank : 10) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P46100, P51068, Q15886, Q59FB5, Q59H31, Q5H9A2, Q5JWI4, Q7Z2J1, Q9H0Z1, Q9NTS3 | Gene names | ATRX, RAD54L, XH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked helicase II) (X-linked nuclear protein) (XNP) (Znf- HX). | |||||
|
SEC63_HUMAN
|
||||||
| NC score | 0.041198 (rank : 17) | θ value | 8.99809 (rank : 48) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
F10A1_HUMAN
|
||||||
| NC score | 0.040250 (rank : 18) | θ value | 1.38821 (rank : 14) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P50502, O14999 | Gene names | ST13, FAM10A1, HIP, SNC6 | |||
|
Domain Architecture |

|
|||||
| Description | Hsc70-interacting protein (Hip) (Suppression of tumorigenicity protein 13) (Putative tumor suppressor ST13) (Protein FAM10A1) (Progesterone receptor-associated p48 protein) (NY-REN-33 antigen). | |||||
|
SEC63_MOUSE
|
||||||
| NC score | 0.037534 (rank : 19) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
AFF2_HUMAN
|
||||||
| NC score | 0.036690 (rank : 20) | θ value | 5.27518 (rank : 27) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P51816, O43786, O60215, P78407, Q13521, Q14323, Q9UNA5 | Gene names | AFF2, FMR2, OX19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation 2 protein) (Protein FMR-2) (FMR2P) (Protein Ox19) (Fragile X E mental retardation syndrome protein). | |||||
|
F10A5_HUMAN
|
||||||
| NC score | 0.035892 (rank : 21) | θ value | 4.03905 (rank : 24) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8NFI4 | Gene names | FAM10A5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM10A5. | |||||
|
AFF2_MOUSE
|
||||||
| NC score | 0.035446 (rank : 22) | θ value | 8.99809 (rank : 40) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O55112 | Gene names | Aff2, Fmr2, Ox19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation protein 2 homolog) (Protein FMR-2) (FMR2P) (Protein Ox19). | |||||
|
SAFB2_HUMAN
|
||||||
| NC score | 0.034835 (rank : 23) | θ value | 0.813845 (rank : 12) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 720 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q14151, Q8TB13 | Gene names | SAFB2, KIAA0138 | |||
|
Domain Architecture |

|
|||||
| Description | Scaffold attachment factor B2. | |||||
|
SNPH_HUMAN
|
||||||
| NC score | 0.032092 (rank : 24) | θ value | 2.36792 (rank : 18) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O15079 | Gene names | SNPH, KIAA0374 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaphilin. | |||||
|
CJ064_HUMAN
|
||||||
| NC score | 0.030761 (rank : 25) | θ value | 3.0926 (rank : 20) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6ZS81, Q8N4A3 | Gene names | C10orf64 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C10orf64. | |||||
|
DCDC2_MOUSE
|
||||||
| NC score | 0.029586 (rank : 26) | θ value | 5.27518 (rank : 29) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q5DU00, Q5SZU0, Q80Y99 | Gene names | Dcdc2, Kiaa1154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublecortin domain-containing protein 2. | |||||
|
DMD_HUMAN
|
||||||
| NC score | 0.028942 (rank : 27) | θ value | 0.125558 (rank : 6) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P11532, Q02295, Q14169, Q14170, Q5JYU0, Q7KZ48 | Gene names | DMD | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
JIP4_MOUSE
|
||||||
| NC score | 0.027360 (rank : 28) | θ value | 2.36792 (rank : 16) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q58A65, Q3UH77, Q3UHF0, Q58VQ4, Q5NC70, Q5NC78, Q6A057, Q6PAS3, Q8CJC2 | Gene names | Spag9, Jip4, Jsap2, Kiaa0516, Mapk8ip4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 4 (JNK-interacting protein 4) (JIP-4) (JNK-associated leucine-zipper protein) (JLP) (Sperm-associated antigen 9) (Mitogen-activated protein kinase 8- interacting protein 4) (JNK/SAPK-associated protein 2) (JSAP2). | |||||
|
WAPL_MOUSE
|
||||||
| NC score | 0.024924 (rank : 29) | θ value | 5.27518 (rank : 31) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q65Z40, Q6ZQE9 | Gene names | Wapal, Kiaa0261, Wapl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wings apart-like protein homolog (Dioxin-inducible factor 2) (DIF-2). | |||||
|
CP135_HUMAN
|
||||||
| NC score | 0.024799 (rank : 30) | θ value | 1.38821 (rank : 13) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1290 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q66GS9, O75130, Q9H8H7 | Gene names | CEP135, CEP4, KIAA0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
TESC_MOUSE
|
||||||
| NC score | 0.024062 (rank : 31) | θ value | 6.88961 (rank : 39) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JKL5 | Gene names | Tesc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tescalcin (TSC). | |||||
|
CC47_MOUSE
|
||||||
| NC score | 0.023052 (rank : 32) | θ value | 8.99809 (rank : 42) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9D024, Q3TSB5, Q3V1S7, Q8C5D9, Q920S6 | Gene names | Ccdc47, Asp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 47 precursor (Adipocyte-specific protein 4). | |||||
|
TESC_HUMAN
|
||||||
| NC score | 0.021462 (rank : 33) | θ value | 8.99809 (rank : 50) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96BS2 | Gene names | TESC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tescalcin (TSC). | |||||
|
MIA2_MOUSE
|
||||||
| NC score | 0.020461 (rank : 34) | θ value | 8.99809 (rank : 45) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q91ZV0 | Gene names | Mia2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma inhibitory activity protein 2 precursor. | |||||
|
CP135_MOUSE
|
||||||
| NC score | 0.020031 (rank : 35) | θ value | 6.88961 (rank : 32) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1278 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6P5D4, Q6A030 | Gene names | Cep135, Cep4, Kiaa0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
LUZP1_MOUSE
|
||||||
| NC score | 0.019658 (rank : 36) | θ value | 8.99809 (rank : 44) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 940 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8R4U7, Q3UV18, Q7TS71, Q8BQW1, Q99NG3, Q9CSL6 | Gene names | Luzp1, Luzp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper protein 1 (Leucine zipper motif-containing protein). | |||||
|
SPHK1_HUMAN
|
||||||
| NC score | 0.019230 (rank : 37) | θ value | 6.88961 (rank : 38) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NYA1, Q9HD92, Q9NY70, Q9NYL3 | Gene names | SPHK1, SPHK, SPK | |||
|
Domain Architecture |

|
|||||
| Description | Sphingosine kinase 1 (EC 2.7.1.-) (SK 1) (SPK 1). | |||||
|
MYH4_HUMAN
|
||||||
| NC score | 0.018317 (rank : 38) | θ value | 4.03905 (rank : 25) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1377 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y623 | Gene names | MYH4 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-4 (Myosin heavy chain, skeletal muscle, fetal) (Myosin heavy chain IIb) (MyHC-IIb). | |||||
|
MDM2_HUMAN
|
||||||
| NC score | 0.018311 (rank : 39) | θ value | 6.88961 (rank : 33) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q00987, Q13226, Q13297, Q13298, Q13299, Q13300, Q13301, Q53XW0, Q71TW9, Q9UGI3, Q9UMT8 | Gene names | MDM2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3 Mdm2 (EC 6.3.2.-) (p53-binding protein Mdm2) (Oncoprotein Mdm2) (Double minute 2 protein) (Hdm2). | |||||
|
BRDT_HUMAN
|
||||||
| NC score | 0.018310 (rank : 40) | θ value | 4.03905 (rank : 23) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q58F21, O14789, Q6P5T1, Q7Z4A6, Q8IWI6 | Gene names | BRDT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain testis-specific protein (RING3-like protein). | |||||
|
MYH8_HUMAN
|
||||||
| NC score | 0.017678 (rank : 41) | θ value | 4.03905 (rank : 26) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1610 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P13535, Q14910 | Gene names | MYH8 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
RHG12_MOUSE
|
||||||
| NC score | 0.017425 (rank : 42) | θ value | 6.88961 (rank : 37) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C0D4, Q8BVP8 | Gene names | Arhgap12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
MYH6_HUMAN
|
||||||
| NC score | 0.017386 (rank : 43) | θ value | 6.88961 (rank : 35) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH8_MOUSE
|
||||||
| NC score | 0.016769 (rank : 44) | θ value | 6.88961 (rank : 36) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1600 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P13542, Q5SX36 | Gene names | Myh8, Myhsp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
MAP4_MOUSE
|
||||||
| NC score | 0.016084 (rank : 45) | θ value | 5.27518 (rank : 30) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P27546 | Gene names | Map4, Mtap4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
ANR21_HUMAN
|
||||||
| NC score | 0.016003 (rank : 46) | θ value | 4.03905 (rank : 22) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q86YR6 | Gene names | ANKRD21, POTE | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat domain-containing protein 21 (Protein POTE) (Prostate, ovary, testis-expressed protein). | |||||
|
LAMA2_MOUSE
|
||||||
| NC score | 0.015979 (rank : 47) | θ value | 1.38821 (rank : 15) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1198 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q60675, Q05003, Q64061 | Gene names | Lama2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
ANR26_MOUSE
|
||||||
| NC score | 0.015792 (rank : 48) | θ value | 8.99809 (rank : 41) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1554 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q811D2, Q69ZS2, Q9CS61 | Gene names | Ankrd26, Kiaa1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
MYEF2_MOUSE
|
||||||
| NC score | 0.011680 (rank : 49) | θ value | 6.88961 (rank : 34) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C854, Q60690, Q6P930, Q6ZPT3, Q8QZZ1 | Gene names | Myef2, Kiaa1341, Mef2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin expression factor 2 (MyEF-2) (MEF-2). | |||||
|
KCNK5_HUMAN
|
||||||
| NC score | 0.009890 (rank : 50) | θ value | 8.99809 (rank : 43) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95279 | Gene names | KCNK5, TASK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 5 (Acid-sensitive potassium channel protein TASK-2) (TWIK-related acid-sensitive K(+) channel 2). | |||||
|
PRRT3_HUMAN
|
||||||
| NC score | 0.007484 (rank : 51) | θ value | 8.99809 (rank : 47) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
TAOK2_HUMAN
|
||||||
| NC score | 0.007350 (rank : 52) | θ value | 2.36792 (rank : 19) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1172 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UL54, O94957, Q6UW73, Q7LC09, Q9NSW2 | Gene names | TAOK2, KIAA0881, MAP3K17, PSK, PSK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO2 (EC 2.7.11.1) (Thousand and one amino acid protein 2) (Prostate-derived STE20-like kinase 1) (PSK-1) (Kinase from chicken homolog C) (hKFC-C). | |||||
|
NEK1_MOUSE
|
||||||
| NC score | 0.006105 (rank : 53) | θ value | 3.0926 (rank : 21) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P51954 | Gene names | Nek1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Nek1 (EC 2.7.11.1) (NimA-related protein kinase 1). | |||||
|
CAD22_MOUSE
|
||||||
| NC score | 0.004388 (rank : 54) | θ value | 5.27518 (rank : 28) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9WTP5 | Gene names | Cdh22 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-22 precursor (Pituitary and brain cadherin) (PB-cadherin). | |||||
|
PRDM1_MOUSE
|
||||||
| NC score | 0.002774 (rank : 55) | θ value | 2.36792 (rank : 17) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 766 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60636 | Gene names | Prdm1, Blimp1 | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 1 (PR domain-containing protein 1) (Beta-interferon gene positive-regulatory domain I-binding factor) (BLIMP-1). | |||||
|
PKN3_HUMAN
|
||||||
| NC score | 0.001643 (rank : 56) | θ value | 8.99809 (rank : 46) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6P5Z2, Q9UM03 | Gene names | PKN3, PKNBETA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase N3 (EC 2.7.11.13) (Protein kinase PKN- beta) (Protein-kinase C-related kinase 3). | |||||