Please be patient as the page loads
|
NIBA_HUMAN
|
||||||
| SwissProt Accessions | Q9BZQ8, Q5TEM8, Q8TEI5, Q9H593, Q9H9Y8, Q9HCB9 | Gene names | C1orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Niban protein. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
NIBA_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q9BZQ8, Q5TEM8, Q8TEI5, Q9H593, Q9H9Y8, Q9HCB9 | Gene names | C1orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Niban protein. | |||||
|
NIBL_MOUSE
|
||||||
| θ value | 9.92011e-124 (rank : 2) | NC score | 0.900491 (rank : 2) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8R1F1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Niban-like protein. | |||||
|
NIBL_HUMAN
|
||||||
| θ value | 3.30237e-119 (rank : 3) | NC score | 0.899873 (rank : 3) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96TA1, Q9BUS2, Q9NT35 | Gene names | C9orf88 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Niban-like protein (Meg-3). | |||||
|
FLIP1_HUMAN
|
||||||
| θ value | 9.29e-05 (rank : 4) | NC score | 0.064824 (rank : 11) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q7Z7B0, Q5VUL6, Q86TC3, Q8N8B9, Q96SK6, Q9NVI8, Q9ULE5 | Gene names | FILIP1, KIAA1275 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
K0329_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 5) | NC score | 0.134708 (rank : 4) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O15040, Q9UEG6 | Gene names | KIAA0329, KIAA0297 | |||
|
Domain Architecture |

|
|||||
| Description | Protein KIAA0329/KIAA0297. | |||||
|
GOGA4_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 6) | NC score | 0.057585 (rank : 14) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
GP179_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 7) | NC score | 0.066420 (rank : 10) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6PRD1 | Gene names | GPR179, GPR158L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 179 precursor (Probable G-protein coupled receptor 158-like 1). | |||||
|
CABYR_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 8) | NC score | 0.086149 (rank : 5) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D424, Q91Y41, Q91Y42 | Gene names | Cabyr, Cbp86 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-binding tyrosine phosphorylation-regulated protein (Calcium- binding protein 86) (Testis-specific calcium-binding protein CBP86). | |||||
|
MAP1B_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 9) | NC score | 0.079964 (rank : 6) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
TXLNA_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 10) | NC score | 0.074933 (rank : 9) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 982 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P40222, Q66K62, Q86T54, Q86T85, Q86T86, Q86Y86, Q86YW3, Q8N2Y3 | Gene names | TXLNA, TXLN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-taxilin. | |||||
|
TXLNA_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 11) | NC score | 0.079358 (rank : 7) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 846 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6PAM1, Q6P1E5 | Gene names | Txlna, Txln | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-taxilin. | |||||
|
TRIPB_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 12) | NC score | 0.053807 (rank : 17) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1587 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q15643, O14689, O15154, O95949 | Gene names | TRIP11, CEV14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid receptor-interacting protein 11 (TRIP-11) (Golgi-associated microtubule-binding protein 210) (GMAP-210) (Trip230) (Clonal evolution-related gene on chromosome 14). | |||||
|
SSPO_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 13) | NC score | 0.023235 (rank : 52) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 800 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CG65 | Gene names | Sspo | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SCO-spondin precursor. | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 14) | NC score | 0.039471 (rank : 31) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
MAP1B_HUMAN
|
||||||
| θ value | 0.163984 (rank : 15) | NC score | 0.076210 (rank : 8) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
NCOA6_HUMAN
|
||||||
| θ value | 0.21417 (rank : 16) | NC score | 0.036194 (rank : 38) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
UTRO_HUMAN
|
||||||
| θ value | 0.21417 (rank : 17) | NC score | 0.044628 (rank : 28) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P46939 | Gene names | UTRN, DMDL | |||
|
Domain Architecture |

|
|||||
| Description | Utrophin (Dystrophin-related protein 1) (DRP1) (DRP). | |||||
|
AATF_MOUSE
|
||||||
| θ value | 0.279714 (rank : 18) | NC score | 0.058363 (rank : 13) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JKX4, Q7TQN1, Q8C5Q2, Q99P89 | Gene names | Aatf, Che1, Trb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein AATF (Apoptosis-antagonizing transcription factor) (Rb-binding protein Che-1) (Traube protein). | |||||
|
GOGB1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 19) | NC score | 0.048290 (rank : 24) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
ANR18_HUMAN
|
||||||
| θ value | 0.813845 (rank : 20) | NC score | 0.032170 (rank : 43) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1106 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8IVF6, Q7Z468 | Gene names | ANKRD18A, KIAA2015 | |||
|
Domain Architecture |
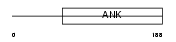
|
|||||
| Description | Ankyrin repeat domain-containing protein 18A. | |||||
|
ATBF1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 21) | NC score | 0.013621 (rank : 76) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1547 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15911, O15101, Q13719 | Gene names | ATBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 22) | NC score | 0.036518 (rank : 37) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
NFM_HUMAN
|
||||||
| θ value | 0.813845 (rank : 23) | NC score | 0.025967 (rank : 48) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P07197 | Gene names | NEF3, NEFM, NFM | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M) (Neurofilament 3). | |||||
|
TACC3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 24) | NC score | 0.055258 (rank : 16) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9Y6A5, Q9UMQ1 | Gene names | TACC3, ERIC1 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 3 (ERIC-1). | |||||
|
HEMGN_MOUSE
|
||||||
| θ value | 1.06291 (rank : 25) | NC score | 0.046006 (rank : 25) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ERZ0, Q3TAE6, Q80YT2, Q8CF29 | Gene names | Hemgn, Ndr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hemogen (Hemopoietic gene protein) (Negative differentiation regulator protein) (mNDR). | |||||
|
NID2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 26) | NC score | 0.012679 (rank : 77) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O88322 | Gene names | Nid2 | |||
|
Domain Architecture |

|
|||||
| Description | Nidogen-2 precursor (NID-2) (Entactin-2). | |||||
|
NIN_HUMAN
|
||||||
| θ value | 1.06291 (rank : 27) | NC score | 0.049591 (rank : 22) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1416 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8N4C6, Q6P0P6, Q9BWU6, Q9C012, Q9C013, Q9C014, Q9H5I6, Q9HAT7, Q9HBY5, Q9HCK7, Q9UH61 | Gene names | NIN, KIAA1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein (hNinein) (Glycogen synthase kinase 3 beta-interacting protein) (GSK3B-interacting protein). | |||||
|
FA8_HUMAN
|
||||||
| θ value | 1.38821 (rank : 28) | NC score | 0.019187 (rank : 62) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P00451 | Gene names | F8, F8C | |||
|
Domain Architecture |

|
|||||
| Description | Coagulation factor VIII precursor (Procoagulant component) (Antihemophilic factor) (AHF) [Contains: Factor VIIIa heavy chain, 200 kDa isoform; Factor VIIIa heavy chain, 92 kDa isoform; Factor VIII B chain; Factor VIIIa light chain]. | |||||
|
FIGN_HUMAN
|
||||||
| θ value | 1.38821 (rank : 29) | NC score | 0.022748 (rank : 53) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5HY92, Q9H6M5, Q9NVZ9 | Gene names | FIGN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fidgetin. | |||||
|
GOGA4_HUMAN
|
||||||
| θ value | 1.38821 (rank : 30) | NC score | 0.050300 (rank : 21) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
KMHN1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 31) | NC score | 0.045521 (rank : 26) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 950 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8TBZ0, Q86YI9, Q8N7W0 | Gene names | KMHN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cancer/testis antigen KM-HN-1. | |||||
|
RAD50_HUMAN
|
||||||
| θ value | 1.38821 (rank : 32) | NC score | 0.044991 (rank : 27) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q92878, O43254, Q6GMT7, Q6P5X3, Q9UP86 | Gene names | RAD50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (hRAD50). | |||||
|
SPTA1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 33) | NC score | 0.032895 (rank : 42) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 675 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P08032, P97502 | Gene names | Spta1, Spna1, Spta | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin alpha chain, erythrocyte (Erythroid alpha-spectrin). | |||||
|
TXLNB_HUMAN
|
||||||
| θ value | 1.81305 (rank : 34) | NC score | 0.064744 (rank : 12) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 912 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8N3L3, Q76L25, Q86T52, Q8N3S2 | Gene names | TXLNB, C6orf198, MDP77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77) (hMDP77). | |||||
|
DDEF1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.018309 (rank : 63) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9QWY8, O08612, Q80TG8, Q80UV6, Q99LV8, Q9Z2B6 | Gene names | Ddef1, Asap1, Kiaa1249, Shag1 | |||
|
Domain Architecture |

|
|||||
| Description | 130 kDa phosphatidylinositol 4,5-biphosphate-dependent ARF1 GTPase- activating protein (PIP2-dependent ARF1 GAP) (ADP-ribosylation factor- directed GTPase-activating protein 1) (ARF GTPase-activating protein 1) (Development and differentiation-enhancing factor 1) (Differentiation-enhancing factor 1) (DEF-1). | |||||
|
IKKB_MOUSE
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.006538 (rank : 82) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 984 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O88351, Q9R1J6 | Gene names | Ikbkb, Ikkb | |||
|
Domain Architecture |
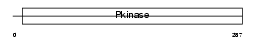
|
|||||
| Description | Inhibitor of nuclear factor kappa B kinase subunit beta (EC 2.7.11.10) (I-kappa-B-kinase beta) (IkBKB) (IKK-beta) (IKK-B) (I-kappa-B kinase 2) (IKK2) (Nuclear factor NF-kappa-B inhibitor kinase beta) (NFKBIKB). | |||||
|
MSH3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 37) | NC score | 0.021228 (rank : 55) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P13705 | Gene names | Msh3, Rep-3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein Msh3 (Repair-3 protein) (REP-1). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 2.36792 (rank : 38) | NC score | 0.036615 (rank : 36) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
RAD9A_MOUSE
|
||||||
| θ value | 2.36792 (rank : 39) | NC score | 0.033391 (rank : 41) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z0F6, Q8QZZ3 | Gene names | Rad9a, Rad9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell cycle checkpoint control protein RAD9A (EC 3.1.11.2) (DNA repair exonuclease rad9 homolog A) (Rad9-like protein) (mRAD9). | |||||
|
STAM1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 40) | NC score | 0.020544 (rank : 56) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P70297 | Gene names | Stam, Stam1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal transducing adapter molecule 1 (STAM-1). | |||||
|
SVIL_MOUSE
|
||||||
| θ value | 2.36792 (rank : 41) | NC score | 0.024110 (rank : 51) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K4L3 | Gene names | Svil | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Supervillin (Archvillin) (p205/p250). | |||||
|
CCD41_MOUSE
|
||||||
| θ value | 3.0926 (rank : 42) | NC score | 0.038262 (rank : 34) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9D5R3, Q3U7X7, Q3UX57, Q80VF0 | Gene names | Ccdc41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41. | |||||
|
CCD46_HUMAN
|
||||||
| θ value | 3.0926 (rank : 43) | NC score | 0.039332 (rank : 32) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1430 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8N8E3, Q6PIB5, Q8NCR4, Q8NFR4 | Gene names | CCDC46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 46. | |||||
|
NIN_MOUSE
|
||||||
| θ value | 3.0926 (rank : 44) | NC score | 0.044563 (rank : 29) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61043, Q6ZPM7 | Gene names | Nin, Kiaa1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein. | |||||
|
RYR2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 45) | NC score | 0.017905 (rank : 65) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92736, Q15411, Q546N8 | Gene names | RYR2 | |||
|
Domain Architecture |

|
|||||
| Description | Ryanodine receptor 2 (Cardiac muscle-type ryanodine receptor) (RyR2) (RYR-2) (Cardiac muscle ryanodine receptor-calcium release channel) (hRYR-2). | |||||
|
UACA_HUMAN
|
||||||
| θ value | 3.0926 (rank : 46) | NC score | 0.035729 (rank : 40) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1622 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9BZF9, Q8N3B8, Q9HCL1, Q9NWC6 | Gene names | UACA, KIAA1561 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uveal autoantigen with coiled-coil domains and ankyrin repeats protein. | |||||
|
UBP24_HUMAN
|
||||||
| θ value | 3.0926 (rank : 47) | NC score | 0.013865 (rank : 75) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UPU5, Q6ZSY2, Q8N2Y4, Q9NXD1 | Gene names | USP24, KIAA1057 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 24 (EC 3.1.2.15) (Ubiquitin thioesterase 24) (Ubiquitin-specific-processing protease 24) (Deubiquitinating enzyme 24). | |||||
|
APOE_HUMAN
|
||||||
| θ value | 4.03905 (rank : 48) | NC score | 0.036101 (rank : 39) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P02649, Q9P2S4 | Gene names | APOE | |||
|
Domain Architecture |
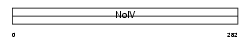
|
|||||
| Description | Apolipoprotein E precursor (Apo-E). | |||||
|
CAN6_HUMAN
|
||||||
| θ value | 4.03905 (rank : 49) | NC score | 0.015570 (rank : 69) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y6Q1, Q9UEQ1, Q9UJA8 | Gene names | CAPN6, CALPM, CANPX | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-6 (Calpamodulin) (CalpM) (Calpain-like protease X-linked). | |||||
|
DYH5_HUMAN
|
||||||
| θ value | 4.03905 (rank : 50) | NC score | 0.019512 (rank : 60) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8TE73, Q92860, Q96L74, Q9H5S7, Q9HCG9 | Gene names | DNAH5, DNAHC5, KIAA1603 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ciliary dynein heavy chain 5 (Axonemal beta dynein heavy chain 5) (HL1). | |||||
|
FIGN_MOUSE
|
||||||
| θ value | 4.03905 (rank : 51) | NC score | 0.019853 (rank : 59) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ERZ6, Q3TPB0, Q3UP57, Q6PCM0 | Gene names | Fign | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fidgetin. | |||||
|
FOXP1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 52) | NC score | 0.015363 (rank : 70) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9H334, Q9H332, Q9H333, Q9P0R1 | Gene names | FOXP1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P1. | |||||
|
MACF1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 53) | NC score | 0.037327 (rank : 35) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1341 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9UPN3, O75053, Q5VW20, Q8WXY2, Q9H540, Q9UKP0, Q9ULG9 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoforms 1/2/3/5 (Actin cross-linking family protein 7) (Macrophin-1) (Trabeculin-alpha) (620 kDa actin-binding protein) (ABP620). | |||||
|
MACF4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 54) | NC score | 0.043449 (rank : 30) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1321 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q96PK2, Q8WXY1 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoform 4. | |||||
|
MAST4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 55) | NC score | 0.004373 (rank : 88) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||
|
TACC3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 56) | NC score | 0.048317 (rank : 23) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9JJ11, Q9WVK9 | Gene names | Tacc3, Aint | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 3 (ARNT-interacting protein). | |||||
|
AMOL1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 57) | NC score | 0.031903 (rank : 44) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9D4H4, Q571F1 | Gene names | Amotl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
DDX54_HUMAN
|
||||||
| θ value | 5.27518 (rank : 58) | NC score | 0.007522 (rank : 81) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TDD1, Q86YT8, Q9BRZ1 | Gene names | DDX54 | |||
|
Domain Architecture |
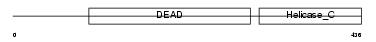
|
|||||
| Description | ATP-dependent RNA helicase DDX54 (EC 3.6.1.-) (DEAD box protein 54) (ATP-dependent RNA helicase DP97). | |||||
|
ERIC1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 59) | NC score | 0.026368 (rank : 45) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
FOXP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.014472 (rank : 71) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P58462 | Gene names | Foxp1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P1 (Forkhead-related transcription factor 1). | |||||
|
HSC20_HUMAN
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.025542 (rank : 49) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8IWL3, Q9BWS7 | Gene names | HSCB, HSC20 | |||
|
Domain Architecture |
|
|||||
| Description | Co-chaperone protein HscB, mitochondrial precursor (Hsc20). | |||||
|
K0247_HUMAN
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.026308 (rank : 46) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92537 | Gene names | KIAA0247 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0247 precursor. | |||||
|
RPGR1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 63) | NC score | 0.038675 (rank : 33) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 578 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q96KN7, Q7Z2W6, Q8IXV5, Q96QA8, Q9HB94, Q9HB95, Q9HBK6, Q9NR40 | Gene names | RPGRIP1 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator-interacting protein 1 (RPGR-interacting protein 1). | |||||
|
SEC63_HUMAN
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.022397 (rank : 54) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
SEC63_MOUSE
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.026060 (rank : 47) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
CAN6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 66) | NC score | 0.013999 (rank : 73) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35646 | Gene names | Capn6, Capa6 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-6. | |||||
|
CND1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.016086 (rank : 68) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15021, Q8N6U3 | Gene names | NCAPD2, CAPD2, CNAP1, KIAA0159 | |||
|
Domain Architecture |

|
|||||
| Description | Condensin complex subunit 1 (Non-SMC condensin I complex subunit D2) (Chromosome condensation-related SMC-associated protein 1) (Chromosome-associated protein D2) (hCAP-D2) (XCAP-D2 homolog). | |||||
|
DDX18_MOUSE
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.006527 (rank : 84) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K363, Q3MIB0, Q8BVZ2, Q9D2E0 | Gene names | Ddx18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX18 (EC 3.6.1.-) (DEAD box protein 18). | |||||
|
IKKB_HUMAN
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.005075 (rank : 86) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 980 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14920, O75327 | Gene names | IKBKB, IKKB | |||
|
Domain Architecture |
|
|||||
| Description | Inhibitor of nuclear factor kappa B kinase subunit beta (EC 2.7.11.10) (I-kappa-B-kinase beta) (IkBKB) (IKK-beta) (IKK-B) (I-kappa-B kinase 2) (IKK2) (Nuclear factor NF-kappa-B inhibitor kinase beta) (NFKBIKB). | |||||
|
RAI2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | 0.018203 (rank : 64) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QVY8 | Gene names | Rai2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 2 (3f8). | |||||
|
RPKL1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.005783 (rank : 85) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 708 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y6S9, Q69YT9, Q6ZMQ6, Q96NR9, Q9BSU9 | Gene names | RPS6KL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribosomal protein S6 kinase-like 1 (EC 2.7.11.1). | |||||
|
S12A2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.008728 (rank : 80) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55012 | Gene names | Slc12a2, Nkcc1 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 12 member 2 (Bumetanide-sensitive sodium- (potassium)-chloride cotransporter 1) (Basolateral Na-K-Cl symporter). | |||||
|
STAM1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | 0.016734 (rank : 67) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q92783, Q8N6D9 | Gene names | STAM, STAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal transducing adapter molecule 1 (STAM-1). | |||||
|
TSC2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.014344 (rank : 72) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61037, P97723, P97724, P97725, P97727, Q61007, Q61008, Q9WUF6 | Gene names | Tsc2 | |||
|
Domain Architecture |

|
|||||
| Description | Tuberin (Tuberous sclerosis 2 homolog protein). | |||||
|
UT14B_MOUSE
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.020452 (rank : 57) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6EJB6, Q6EJB5, Q8BL60 | Gene names | Utp14b, Jsd | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U3 small nucleolar RNA-associated protein 14 homolog B (Juvenile spermatogonial depletion protein). | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.019383 (rank : 61) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
CTNL1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.013932 (rank : 74) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O88327, Q810J3 | Gene names | Ctnnal1, Catnal1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-catulin (Catenin alpha-like protein 1) (Alpha-catenin-related protein) (ACRP). | |||||
|
IL16_MOUSE
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.009196 (rank : 79) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O54824, O70236, Q3TEM4, Q8R0G5 | Gene names | Il16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-16 precursor (IL-16) (Lymphocyte chemoattractant factor) (LCF). | |||||
|
IMPA1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.012174 (rank : 78) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P29218 | Gene names | IMPA1, IMPA | |||
|
Domain Architecture |

|
|||||
| Description | Inositol monophosphatase (EC 3.1.3.25) (IMPase) (IMP) (Inositol-1(or 4)-monophosphatase) (Lithium-sensitive myo-inositol monophosphatase A1). | |||||
|
MTMR3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.006532 (rank : 83) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13615, Q9NYN5, Q9NYN6, Q9UDX6, Q9UEG3 | Gene names | MTMR3, KIAA0371, ZFYVE10 | |||
|
Domain Architecture |

|
|||||
| Description | Myotubularin-related protein 3 (EC 3.1.3.48) (FYVE domain-containing dual specificity protein phosphatase 1) (FYVE-DSP1) (Zinc finger FYVE domain-containing protein 10). | |||||
|
PTN22_MOUSE
|
||||||
| θ value | 8.99809 (rank : 81) | NC score | 0.004495 (rank : 87) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P29352, Q7TMP9 | Gene names | Ptpn22, Ptpn8 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 22 (EC 3.1.3.48) (Hematopoietic cell protein-tyrosine phosphatase 70Z-PEP). | |||||
|
RAD9A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.024817 (rank : 50) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99638, Q6FI29, Q96C41 | Gene names | RAD9A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell cycle checkpoint control protein RAD9A (EC 3.1.11.2) (DNA repair exonuclease rad9 homolog A) (hRAD9). | |||||
|
ROCK1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 83) | NC score | 0.020174 (rank : 58) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 2126 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P70335, Q8C3G4, Q8C7H0 | Gene names | Rock1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 1 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 1) (p160 ROCK-1) (p160ROCK). | |||||
|
TRM6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.016956 (rank : 66) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UJA5, Q76P92, Q9BQV5, Q9ULR7 | Gene names | TRM6, KIAA1153 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA (adenine-N(1)-)-methyltransferase non-catalytic subunit TRM6 (tRNA(m1A58)-methyltransferase subunit TRM6) (tRNA(m1A58)MTase subunit TRM6). | |||||
|
FLIP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.050361 (rank : 20) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1176 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9CS72 | Gene names | Filip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
TXLNB_MOUSE
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.057446 (rank : 15) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 640 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8VBT1, Q3UVB8, Q8BUK2 | Gene names | Txlnb, Mdp77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77). | |||||
|
TXLNG_HUMAN
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.052635 (rank : 19) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9NUQ3, Q9P0X1 | Gene names | TXLNG, CXorf15, LSR5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gamma-taxilin (Lipopolysaccharide-specific response protein 5). | |||||
|
TXLNG_MOUSE
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.053573 (rank : 18) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 552 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BHN1, Q8BP11 | Gene names | Txlng | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gamma-taxilin. | |||||
|
NIBA_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q9BZQ8, Q5TEM8, Q8TEI5, Q9H593, Q9H9Y8, Q9HCB9 | Gene names | C1orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Niban protein. | |||||
|
NIBL_MOUSE
|
||||||
| NC score | 0.900491 (rank : 2) | θ value | 9.92011e-124 (rank : 2) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8R1F1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Niban-like protein. | |||||
|
NIBL_HUMAN
|
||||||
| NC score | 0.899873 (rank : 3) | θ value | 3.30237e-119 (rank : 3) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96TA1, Q9BUS2, Q9NT35 | Gene names | C9orf88 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Niban-like protein (Meg-3). | |||||
|
K0329_HUMAN
|
||||||
| NC score | 0.134708 (rank : 4) | θ value | 0.00035302 (rank : 5) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O15040, Q9UEG6 | Gene names | KIAA0329, KIAA0297 | |||
|
Domain Architecture |

|
|||||
| Description | Protein KIAA0329/KIAA0297. | |||||
|
CABYR_MOUSE
|
||||||
| NC score | 0.086149 (rank : 5) | θ value | 0.0330416 (rank : 8) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D424, Q91Y41, Q91Y42 | Gene names | Cabyr, Cbp86 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-binding tyrosine phosphorylation-regulated protein (Calcium- binding protein 86) (Testis-specific calcium-binding protein CBP86). | |||||
|
MAP1B_MOUSE
|
||||||
| NC score | 0.079964 (rank : 6) | θ value | 0.0330416 (rank : 9) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
TXLNA_MOUSE
|
||||||
| NC score | 0.079358 (rank : 7) | θ value | 0.0330416 (rank : 11) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 846 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6PAM1, Q6P1E5 | Gene names | Txlna, Txln | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-taxilin. | |||||
|
MAP1B_HUMAN
|
||||||
| NC score | 0.076210 (rank : 8) | θ value | 0.163984 (rank : 15) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
TXLNA_HUMAN
|
||||||
| NC score | 0.074933 (rank : 9) | θ value | 0.0330416 (rank : 10) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 982 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P40222, Q66K62, Q86T54, Q86T85, Q86T86, Q86Y86, Q86YW3, Q8N2Y3 | Gene names | TXLNA, TXLN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-taxilin. | |||||
|
GP179_HUMAN
|
||||||
| NC score | 0.066420 (rank : 10) | θ value | 0.0252991 (rank : 7) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6PRD1 | Gene names | GPR179, GPR158L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 179 precursor (Probable G-protein coupled receptor 158-like 1). | |||||
|
FLIP1_HUMAN
|
||||||
| NC score | 0.064824 (rank : 11) | θ value | 9.29e-05 (rank : 4) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q7Z7B0, Q5VUL6, Q86TC3, Q8N8B9, Q96SK6, Q9NVI8, Q9ULE5 | Gene names | FILIP1, KIAA1275 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
TXLNB_HUMAN
|
||||||
| NC score | 0.064744 (rank : 12) | θ value | 1.81305 (rank : 34) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 912 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8N3L3, Q76L25, Q86T52, Q8N3S2 | Gene names | TXLNB, C6orf198, MDP77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77) (hMDP77). | |||||
|
AATF_MOUSE
|
||||||
| NC score | 0.058363 (rank : 13) | θ value | 0.279714 (rank : 18) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JKX4, Q7TQN1, Q8C5Q2, Q99P89 | Gene names | Aatf, Che1, Trb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein AATF (Apoptosis-antagonizing transcription factor) (Rb-binding protein Che-1) (Traube protein). | |||||
|
GOGA4_MOUSE
|
||||||
| NC score | 0.057585 (rank : 14) | θ value | 0.0113563 (rank : 6) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
TXLNB_MOUSE
|
||||||
| NC score | 0.057446 (rank : 15) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 640 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8VBT1, Q3UVB8, Q8BUK2 | Gene names | Txlnb, Mdp77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77). | |||||
|
TACC3_HUMAN
|
||||||
| NC score | 0.055258 (rank : 16) | θ value | 0.813845 (rank : 24) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9Y6A5, Q9UMQ1 | Gene names | TACC3, ERIC1 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 3 (ERIC-1). | |||||
|
TRIPB_HUMAN
|
||||||
| NC score | 0.053807 (rank : 17) | θ value | 0.0563607 (rank : 12) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1587 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q15643, O14689, O15154, O95949 | Gene names | TRIP11, CEV14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid receptor-interacting protein 11 (TRIP-11) (Golgi-associated microtubule-binding protein 210) (GMAP-210) (Trip230) (Clonal evolution-related gene on chromosome 14). | |||||
|
TXLNG_MOUSE
|
||||||
| NC score | 0.053573 (rank : 18) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 552 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BHN1, Q8BP11 | Gene names | Txlng | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gamma-taxilin. | |||||
|
TXLNG_HUMAN
|
||||||
| NC score | 0.052635 (rank : 19) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9NUQ3, Q9P0X1 | Gene names | TXLNG, CXorf15, LSR5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gamma-taxilin (Lipopolysaccharide-specific response protein 5). | |||||
|
FLIP1_MOUSE
|
||||||
| NC score | 0.050361 (rank : 20) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1176 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9CS72 | Gene names | Filip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
GOGA4_HUMAN
|
||||||
| NC score | 0.050300 (rank : 21) | θ value | 1.38821 (rank : 30) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
NIN_HUMAN
|
||||||
| NC score | 0.049591 (rank : 22) | θ value | 1.06291 (rank : 27) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1416 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8N4C6, Q6P0P6, Q9BWU6, Q9C012, Q9C013, Q9C014, Q9H5I6, Q9HAT7, Q9HBY5, Q9HCK7, Q9UH61 | Gene names | NIN, KIAA1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein (hNinein) (Glycogen synthase kinase 3 beta-interacting protein) (GSK3B-interacting protein). | |||||
|
TACC3_MOUSE
|
||||||
| NC score | 0.048317 (rank : 23) | θ value | 4.03905 (rank : 56) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9JJ11, Q9WVK9 | Gene names | Tacc3, Aint | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 3 (ARNT-interacting protein). | |||||
|
GOGB1_HUMAN
|
||||||
| NC score | 0.048290 (rank : 24) | θ value | 0.62314 (rank : 19) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
HEMGN_MOUSE
|
||||||
| NC score | 0.046006 (rank : 25) | θ value | 1.06291 (rank : 25) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ERZ0, Q3TAE6, Q80YT2, Q8CF29 | Gene names | Hemgn, Ndr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hemogen (Hemopoietic gene protein) (Negative differentiation regulator protein) (mNDR). | |||||
|
KMHN1_HUMAN
|
||||||
| NC score | 0.045521 (rank : 26) | θ value | 1.38821 (rank : 31) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 950 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8TBZ0, Q86YI9, Q8N7W0 | Gene names | KMHN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cancer/testis antigen KM-HN-1. | |||||
|
RAD50_HUMAN
|
||||||
| NC score | 0.044991 (rank : 27) | θ value | 1.38821 (rank : 32) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q92878, O43254, Q6GMT7, Q6P5X3, Q9UP86 | Gene names | RAD50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (hRAD50). | |||||
|
UTRO_HUMAN
|
||||||
| NC score | 0.044628 (rank : 28) | θ value | 0.21417 (rank : 17) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P46939 | Gene names | UTRN, DMDL | |||
|
Domain Architecture |

|
|||||
| Description | Utrophin (Dystrophin-related protein 1) (DRP1) (DRP). | |||||
|
NIN_MOUSE
|
||||||
| NC score | 0.044563 (rank : 29) | θ value | 3.0926 (rank : 44) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61043, Q6ZPM7 | Gene names | Nin, Kiaa1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein. | |||||
|
MACF4_HUMAN
|
||||||
| NC score | 0.043449 (rank : 30) | θ value | 4.03905 (rank : 54) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1321 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q96PK2, Q8WXY1 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoform 4. | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.039471 (rank : 31) | θ value | 0.125558 (rank : 14) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
CCD46_HUMAN
|
||||||
| NC score | 0.039332 (rank : 32) | θ value | 3.0926 (rank : 43) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1430 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8N8E3, Q6PIB5, Q8NCR4, Q8NFR4 | Gene names | CCDC46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 46. | |||||
|
RPGR1_HUMAN
|
||||||
| NC score | 0.038675 (rank : 33) | θ value | 5.27518 (rank : 63) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 578 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q96KN7, Q7Z2W6, Q8IXV5, Q96QA8, Q9HB94, Q9HB95, Q9HBK6, Q9NR40 | Gene names | RPGRIP1 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator-interacting protein 1 (RPGR-interacting protein 1). | |||||
|
CCD41_MOUSE
|
||||||
| NC score | 0.038262 (rank : 34) | θ value | 3.0926 (rank : 42) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9D5R3, Q3U7X7, Q3UX57, Q80VF0 | Gene names | Ccdc41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41. | |||||
|
MACF1_HUMAN
|
||||||
| NC score | 0.037327 (rank : 35) | θ value | 4.03905 (rank : 53) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1341 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9UPN3, O75053, Q5VW20, Q8WXY2, Q9H540, Q9UKP0, Q9ULG9 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoforms 1/2/3/5 (Actin cross-linking family protein 7) (Macrophin-1) (Trabeculin-alpha) (620 kDa actin-binding protein) (ABP620). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.036615 (rank : 36) | θ value | 2.36792 (rank : 38) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.036518 (rank : 37) | θ value | 0.813845 (rank : 22) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
NCOA6_HUMAN
|
||||||
| NC score | 0.036194 (rank : 38) | θ value | 0.21417 (rank : 16) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
APOE_HUMAN
|
||||||
| NC score | 0.036101 (rank : 39) | θ value | 4.03905 (rank : 48) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P02649, Q9P2S4 | Gene names | APOE | |||
|
Domain Architecture |

|
|||||
| Description | Apolipoprotein E precursor (Apo-E). | |||||
|
UACA_HUMAN
|
||||||
| NC score | 0.035729 (rank : 40) | θ value | 3.0926 (rank : 46) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1622 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9BZF9, Q8N3B8, Q9HCL1, Q9NWC6 | Gene names | UACA, KIAA1561 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uveal autoantigen with coiled-coil domains and ankyrin repeats protein. | |||||
|
RAD9A_MOUSE
|
||||||
| NC score | 0.033391 (rank : 41) | θ value | 2.36792 (rank : 39) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z0F6, Q8QZZ3 | Gene names | Rad9a, Rad9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell cycle checkpoint control protein RAD9A (EC 3.1.11.2) (DNA repair exonuclease rad9 homolog A) (Rad9-like protein) (mRAD9). | |||||
|
SPTA1_MOUSE
|
||||||
| NC score | 0.032895 (rank : 42) | θ value | 1.38821 (rank : 33) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 675 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P08032, P97502 | Gene names | Spta1, Spna1, Spta | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin alpha chain, erythrocyte (Erythroid alpha-spectrin). | |||||
|
ANR18_HUMAN
|
||||||
| NC score | 0.032170 (rank : 43) | θ value | 0.813845 (rank : 20) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1106 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8IVF6, Q7Z468 | Gene names | ANKRD18A, KIAA2015 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat domain-containing protein 18A. | |||||
|
AMOL1_MOUSE
|
||||||
| NC score | 0.031903 (rank : 44) | θ value | 5.27518 (rank : 57) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9D4H4, Q571F1 | Gene names | Amotl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
ERIC1_HUMAN
|
||||||
| NC score | 0.026368 (rank : 45) | θ value | 5.27518 (rank : 59) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
K0247_HUMAN
|
||||||
| NC score | 0.026308 (rank : 46) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92537 | Gene names | KIAA0247 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0247 precursor. | |||||
|
SEC63_MOUSE
|
||||||
| NC score | 0.026060 (rank : 47) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
NFM_HUMAN
|
||||||
| NC score | 0.025967 (rank : 48) | θ value | 0.813845 (rank : 23) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P07197 | Gene names | NEF3, NEFM, NFM | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M) (Neurofilament 3). | |||||
|
HSC20_HUMAN
|
||||||
| NC score | 0.025542 (rank : 49) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8IWL3, Q9BWS7 | Gene names | HSCB, HSC20 | |||
|
Domain Architecture |

|
|||||
| Description | Co-chaperone protein HscB, mitochondrial precursor (Hsc20). | |||||
|
RAD9A_HUMAN
|
||||||
| NC score | 0.024817 (rank : 50) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99638, Q6FI29, Q96C41 | Gene names | RAD9A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell cycle checkpoint control protein RAD9A (EC 3.1.11.2) (DNA repair exonuclease rad9 homolog A) (hRAD9). | |||||
|
SVIL_MOUSE
|
||||||
| NC score | 0.024110 (rank : 51) | θ value | 2.36792 (rank : 41) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K4L3 | Gene names | Svil | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Supervillin (Archvillin) (p205/p250). | |||||
|
SSPO_MOUSE
|
||||||
| NC score | 0.023235 (rank : 52) | θ value | 0.0736092 (rank : 13) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 800 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CG65 | Gene names | Sspo | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SCO-spondin precursor. | |||||
|
FIGN_HUMAN
|
||||||
| NC score | 0.022748 (rank : 53) | θ value | 1.38821 (rank : 29) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5HY92, Q9H6M5, Q9NVZ9 | Gene names | FIGN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fidgetin. | |||||
|
SEC63_HUMAN
|
||||||
| NC score | 0.022397 (rank : 54) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
MSH3_MOUSE
|
||||||
| NC score | 0.021228 (rank : 55) | θ value | 2.36792 (rank : 37) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P13705 | Gene names | Msh3, Rep-3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein Msh3 (Repair-3 protein) (REP-1). | |||||
|
STAM1_MOUSE
|
||||||
| NC score | 0.020544 (rank : 56) | θ value | 2.36792 (rank : 40) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P70297 | Gene names | Stam, Stam1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal transducing adapter molecule 1 (STAM-1). | |||||
|
UT14B_MOUSE
|
||||||
| NC score | 0.020452 (rank : 57) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6EJB6, Q6EJB5, Q8BL60 | Gene names | Utp14b, Jsd | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U3 small nucleolar RNA-associated protein 14 homolog B (Juvenile spermatogonial depletion protein). | |||||
|
ROCK1_MOUSE
|
||||||
| NC score | 0.020174 (rank : 58) | θ value | 8.99809 (rank : 83) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 2126 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P70335, Q8C3G4, Q8C7H0 | Gene names | Rock1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 1 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 1) (p160 ROCK-1) (p160ROCK). | |||||
|
FIGN_MOUSE
|
||||||
| NC score | 0.019853 (rank : 59) | θ value | 4.03905 (rank : 51) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ERZ6, Q3TPB0, Q3UP57, Q6PCM0 | Gene names | Fign | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fidgetin. | |||||
|
DYH5_HUMAN
|
||||||
| NC score | 0.019512 (rank : 60) | θ value | 4.03905 (rank : 50) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8TE73, Q92860, Q96L74, Q9H5S7, Q9HCG9 | Gene names | DNAH5, DNAHC5, KIAA1603 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ciliary dynein heavy chain 5 (Axonemal beta dynein heavy chain 5) (HL1). | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.019383 (rank : 61) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
FA8_HUMAN
|
||||||
| NC score | 0.019187 (rank : 62) | θ value | 1.38821 (rank : 28) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P00451 | Gene names | F8, F8C | |||
|
Domain Architecture |

|
|||||
| Description | Coagulation factor VIII precursor (Procoagulant component) (Antihemophilic factor) (AHF) [Contains: Factor VIIIa heavy chain, 200 kDa isoform; Factor VIIIa heavy chain, 92 kDa isoform; Factor VIII B chain; Factor VIIIa light chain]. | |||||
|
DDEF1_MOUSE
|
||||||
| NC score | 0.018309 (rank : 63) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9QWY8, O08612, Q80TG8, Q80UV6, Q99LV8, Q9Z2B6 | Gene names | Ddef1, Asap1, Kiaa1249, Shag1 | |||
|
Domain Architecture |

|
|||||
| Description | 130 kDa phosphatidylinositol 4,5-biphosphate-dependent ARF1 GTPase- activating protein (PIP2-dependent ARF1 GAP) (ADP-ribosylation factor- directed GTPase-activating protein 1) (ARF GTPase-activating protein 1) (Development and differentiation-enhancing factor 1) (Differentiation-enhancing factor 1) (DEF-1). | |||||
|
RAI2_MOUSE
|
||||||
| NC score | 0.018203 (rank : 64) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QVY8 | Gene names | Rai2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 2 (3f8). | |||||
|
RYR2_HUMAN
|
||||||
| NC score | 0.017905 (rank : 65) | θ value | 3.0926 (rank : 45) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92736, Q15411, Q546N8 | Gene names | RYR2 | |||
|
Domain Architecture |

|
|||||
| Description | Ryanodine receptor 2 (Cardiac muscle-type ryanodine receptor) (RyR2) (RYR-2) (Cardiac muscle ryanodine receptor-calcium release channel) (hRYR-2). | |||||
|
TRM6_HUMAN
|
||||||
| NC score | 0.016956 (rank : 66) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UJA5, Q76P92, Q9BQV5, Q9ULR7 | Gene names | TRM6, KIAA1153 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA (adenine-N(1)-)-methyltransferase non-catalytic subunit TRM6 (tRNA(m1A58)-methyltransferase subunit TRM6) (tRNA(m1A58)MTase subunit TRM6). | |||||
|
STAM1_HUMAN
|
||||||
| NC score | 0.016734 (rank : 67) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q92783, Q8N6D9 | Gene names | STAM, STAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal transducing adapter molecule 1 (STAM-1). | |||||
|
CND1_HUMAN
|
||||||
| NC score | 0.016086 (rank : 68) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15021, Q8N6U3 | Gene names | NCAPD2, CAPD2, CNAP1, KIAA0159 | |||
|
Domain Architecture |

|
|||||
| Description | Condensin complex subunit 1 (Non-SMC condensin I complex subunit D2) (Chromosome condensation-related SMC-associated protein 1) (Chromosome-associated protein D2) (hCAP-D2) (XCAP-D2 homolog). | |||||
|
CAN6_HUMAN
|
||||||
| NC score | 0.015570 (rank : 69) | θ value | 4.03905 (rank : 49) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y6Q1, Q9UEQ1, Q9UJA8 | Gene names | CAPN6, CALPM, CANPX | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-6 (Calpamodulin) (CalpM) (Calpain-like protease X-linked). | |||||
|
FOXP1_HUMAN
|
||||||
| NC score | 0.015363 (rank : 70) | θ value | 4.03905 (rank : 52) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9H334, Q9H332, Q9H333, Q9P0R1 | Gene names | FOXP1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P1. | |||||
|
FOXP1_MOUSE
|
||||||
| NC score | 0.014472 (rank : 71) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P58462 | Gene names | Foxp1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P1 (Forkhead-related transcription factor 1). | |||||
|
TSC2_MOUSE
|
||||||
| NC score | 0.014344 (rank : 72) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61037, P97723, P97724, P97725, P97727, Q61007, Q61008, Q9WUF6 | Gene names | Tsc2 | |||
|
Domain Architecture |

|
|||||
| Description | Tuberin (Tuberous sclerosis 2 homolog protein). | |||||
|
CAN6_MOUSE
|
||||||
| NC score | 0.013999 (rank : 73) | θ value | 6.88961 (rank : 66) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35646 | Gene names | Capn6, Capa6 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-6. | |||||
|
CTNL1_MOUSE
|
||||||
| NC score | 0.013932 (rank : 74) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O88327, Q810J3 | Gene names | Ctnnal1, Catnal1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-catulin (Catenin alpha-like protein 1) (Alpha-catenin-related protein) (ACRP). | |||||
|
UBP24_HUMAN
|
||||||
| NC score | 0.013865 (rank : 75) | θ value | 3.0926 (rank : 47) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UPU5, Q6ZSY2, Q8N2Y4, Q9NXD1 | Gene names | USP24, KIAA1057 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 24 (EC 3.1.2.15) (Ubiquitin thioesterase 24) (Ubiquitin-specific-processing protease 24) (Deubiquitinating enzyme 24). | |||||
|
ATBF1_HUMAN
|
||||||
| NC score | 0.013621 (rank : 76) | θ value | 0.813845 (rank : 21) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1547 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15911, O15101, Q13719 | Gene names | ATBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
NID2_MOUSE
|
||||||
| NC score | 0.012679 (rank : 77) | θ value | 1.06291 (rank : 26) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O88322 | Gene names | Nid2 | |||
|
Domain Architecture |

|
|||||
| Description | Nidogen-2 precursor (NID-2) (Entactin-2). | |||||
|
IMPA1_HUMAN
|
||||||
| NC score | 0.012174 (rank : 78) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P29218 | Gene names | IMPA1, IMPA | |||
|
Domain Architecture |

|
|||||
| Description | Inositol monophosphatase (EC 3.1.3.25) (IMPase) (IMP) (Inositol-1(or 4)-monophosphatase) (Lithium-sensitive myo-inositol monophosphatase A1). | |||||
|
IL16_MOUSE
|
||||||
| NC score | 0.009196 (rank : 79) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O54824, O70236, Q3TEM4, Q8R0G5 | Gene names | Il16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-16 precursor (IL-16) (Lymphocyte chemoattractant factor) (LCF). | |||||
|
S12A2_MOUSE
|
||||||
| NC score | 0.008728 (rank : 80) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55012 | Gene names | Slc12a2, Nkcc1 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 12 member 2 (Bumetanide-sensitive sodium- (potassium)-chloride cotransporter 1) (Basolateral Na-K-Cl symporter). | |||||
|
DDX54_HUMAN
|
||||||
| NC score | 0.007522 (rank : 81) | θ value | 5.27518 (rank : 58) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TDD1, Q86YT8, Q9BRZ1 | Gene names | DDX54 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX54 (EC 3.6.1.-) (DEAD box protein 54) (ATP-dependent RNA helicase DP97). | |||||
|
IKKB_MOUSE
|
||||||
| NC score | 0.006538 (rank : 82) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 984 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O88351, Q9R1J6 | Gene names | Ikbkb, Ikkb | |||
|
Domain Architecture |

|
|||||
| Description | Inhibitor of nuclear factor kappa B kinase subunit beta (EC 2.7.11.10) (I-kappa-B-kinase beta) (IkBKB) (IKK-beta) (IKK-B) (I-kappa-B kinase 2) (IKK2) (Nuclear factor NF-kappa-B inhibitor kinase beta) (NFKBIKB). | |||||
|
MTMR3_HUMAN
|
||||||
| NC score | 0.006532 (rank : 83) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13615, Q9NYN5, Q9NYN6, Q9UDX6, Q9UEG3 | Gene names | MTMR3, KIAA0371, ZFYVE10 | |||
|
Domain Architecture |

|
|||||
| Description | Myotubularin-related protein 3 (EC 3.1.3.48) (FYVE domain-containing dual specificity protein phosphatase 1) (FYVE-DSP1) (Zinc finger FYVE domain-containing protein 10). | |||||
|
DDX18_MOUSE
|
||||||
| NC score | 0.006527 (rank : 84) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K363, Q3MIB0, Q8BVZ2, Q9D2E0 | Gene names | Ddx18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX18 (EC 3.6.1.-) (DEAD box protein 18). | |||||
|
RPKL1_HUMAN
|
||||||
| NC score | 0.005783 (rank : 85) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 708 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y6S9, Q69YT9, Q6ZMQ6, Q96NR9, Q9BSU9 | Gene names | RPS6KL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribosomal protein S6 kinase-like 1 (EC 2.7.11.1). | |||||
|
IKKB_HUMAN
|
||||||
| NC score | 0.005075 (rank : 86) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 980 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14920, O75327 | Gene names | IKBKB, IKKB | |||
|
Domain Architecture |

|
|||||
| Description | Inhibitor of nuclear factor kappa B kinase subunit beta (EC 2.7.11.10) (I-kappa-B-kinase beta) (IkBKB) (IKK-beta) (IKK-B) (I-kappa-B kinase 2) (IKK2) (Nuclear factor NF-kappa-B inhibitor kinase beta) (NFKBIKB). | |||||
|
PTN22_MOUSE
|
||||||
| NC score | 0.004495 (rank : 87) | θ value | 8.99809 (rank : 81) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P29352, Q7TMP9 | Gene names | Ptpn22, Ptpn8 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 22 (EC 3.1.3.48) (Hematopoietic cell protein-tyrosine phosphatase 70Z-PEP). | |||||
|
MAST4_HUMAN
|
||||||
| NC score | 0.004373 (rank : 88) | θ value | 4.03905 (rank : 55) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||