Please be patient as the page loads
|
DPOLZ_MOUSE
|
||||||
| SwissProt Accessions | Q61493, Q9JMD6, Q9QWX6 | Gene names | Rev3l, Polz, Sez4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (Seizure-related protein 4). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
DPOLZ_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.967263 (rank : 2) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O60673, O43214 | Gene names | REV3L, POLZ, REV3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (hREV3). | |||||
|
DPOLZ_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 169 | |
| SwissProt Accessions | Q61493, Q9JMD6, Q9QWX6 | Gene names | Rev3l, Polz, Sez4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (Seizure-related protein 4). | |||||
|
DPOD1_MOUSE
|
||||||
| θ value | 5.6855e-87 (rank : 3) | NC score | 0.808349 (rank : 3) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P52431, O54883 | Gene names | Pold1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase delta catalytic subunit (EC 2.7.7.7). | |||||
|
DPOD1_HUMAN
|
||||||
| θ value | 2.82168e-86 (rank : 4) | NC score | 0.808089 (rank : 4) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P28340, Q8NER3, Q96H98 | Gene names | POLD1, POLD | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase delta catalytic subunit (EC 2.7.7.7) (DNA polymerase subunit delta p125). | |||||
|
DPOLA_MOUSE
|
||||||
| θ value | 1.6247e-33 (rank : 5) | NC score | 0.639734 (rank : 6) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P33609 | Gene names | Pola1, Pola | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase alpha catalytic subunit (EC 2.7.7.7). | |||||
|
DPOLA_HUMAN
|
||||||
| θ value | 1.79631e-32 (rank : 6) | NC score | 0.640583 (rank : 5) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P09884, Q86UQ7 | Gene names | POLA1, POLA | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase alpha catalytic subunit (EC 2.7.7.7). | |||||
|
CENPE_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 7) | NC score | 0.028775 (rank : 64) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
PDLI1_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 8) | NC score | 0.050875 (rank : 20) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O00151, Q5VZH5, Q9BPZ9 | Gene names | PDLIM1, CLIM1, CLP36 | |||
|
Domain Architecture |
|
|||||
| Description | PDZ and LIM domain protein 1 (Elfin) (LIM domain protein CLP-36) (C- terminal LIM domain protein 1). | |||||
|
UBP1_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 9) | NC score | 0.067989 (rank : 12) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O94782, Q9UFR0, Q9UNJ3 | Gene names | USP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 1 (EC 3.1.2.15) (Ubiquitin thioesterase 1) (Ubiquitin-specific-processing protease 1) (Deubiquitinating enzyme 1) (hUBP). | |||||
|
CSPG2_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 10) | NC score | 0.020474 (rank : 87) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 711 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q62059, Q62058, Q9CUU0 | Gene names | Cspg2 | |||
|
Domain Architecture |

|
|||||
| Description | Versican core protein precursor (Large fibroblast proteoglycan) (Chondroitin sulfate proteoglycan core protein 2) (PG-M). | |||||
|
DNMT1_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 11) | NC score | 0.056662 (rank : 17) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P26358, Q9UHG5, Q9ULA2, Q9UMZ6 | Gene names | DNMT1, AIM, DNMT | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase HsaI) (DNA MTase HsaI) (MCMT) (M.HsaI). | |||||
|
NCOR1_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 12) | NC score | 0.058422 (rank : 16) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O75376, Q9UPV5, Q9UQ18 | Gene names | NCOR1, KIAA1047 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR). | |||||
|
SYCP2_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 13) | NC score | 0.068505 (rank : 11) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BX26, O75763, Q5JX11, Q9NTX8, Q9UG27 | Gene names | SYCP2, SCP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptonemal complex protein 2 (SCP-2) (Synaptonemal complex lateral element protein) (hsSCP2). | |||||
|
NU153_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 14) | NC score | 0.065326 (rank : 13) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P49790 | Gene names | NUP153 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup153 (Nucleoporin Nup153) (153 kDa nucleoporin). | |||||
|
ATX2_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 15) | NC score | 0.043943 (rank : 29) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 375 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99700, Q6ZQZ7, Q99493 | Gene names | ATXN2, ATX2, SCA2, TNRC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2 (Spinocerebellar ataxia type 2 protein) (Trinucleotide repeat-containing gene 13 protein). | |||||
|
PPRB_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 16) | NC score | 0.060477 (rank : 15) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q925J9, O88323, Q3UHV0, Q6AXD5, Q8BW37, Q8BX19, Q8VDQ7, Q925K0 | Gene names | Pparbp, Trip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2). | |||||
|
HRX_HUMAN
|
||||||
| θ value | 0.163984 (rank : 17) | NC score | 0.042627 (rank : 31) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
TBCD1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 18) | NC score | 0.032405 (rank : 54) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q86TI0, Q96K82, Q9UPP4 | Gene names | TBC1D1, KIAA1108 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 1. | |||||
|
DPOE1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 19) | NC score | 0.098604 (rank : 7) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9WVF7, Q9QX50 | Gene names | Pole, Pole1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase epsilon, catalytic subunit A (EC 2.7.7.7) (DNA polymerase II subunit A). | |||||
|
DSPP_MOUSE
|
||||||
| θ value | 0.21417 (rank : 20) | NC score | 0.052061 (rank : 19) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
OXR1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 21) | NC score | 0.042360 (rank : 32) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8N573, Q3LIB5, Q6ZVK9, Q8N8V0, Q9H266, Q9NWC7 | Gene names | OXR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1. | |||||
|
TAF3_MOUSE
|
||||||
| θ value | 0.21417 (rank : 22) | NC score | 0.048655 (rank : 23) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 463 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q5HZG4, Q3U490, Q3UWX2, Q8BIU8, Q99JH4 | Gene names | Taf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
CASG_MOUSE
|
||||||
| θ value | 0.279714 (rank : 23) | NC score | 0.083047 (rank : 10) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q02862 | Gene names | Csng | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gamma-casein precursor (PP22). | |||||
|
DLGP4_HUMAN
|
||||||
| θ value | 0.279714 (rank : 24) | NC score | 0.031610 (rank : 57) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y2H0, Q5T2Y4, Q5T2Y5, Q9H137, Q9H138, Q9H1L7 | Gene names | DLGAP4, DAP4, KIAA0964 | |||
|
Domain Architecture |

|
|||||
| Description | Disks large-associated protein 4 (DAP-4) (SAP90/PSD-95-associated protein 4) (SAPAP4) (PSD-95/SAP90-binding protein 4). | |||||
|
DSPP_HUMAN
|
||||||
| θ value | 0.279714 (rank : 25) | NC score | 0.064614 (rank : 14) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
AFF1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 26) | NC score | 0.042829 (rank : 30) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O88573 | Gene names | Aff1, Mllt2, Mllt2h | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4). | |||||
|
DPOE1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 27) | NC score | 0.098013 (rank : 8) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q07864, Q13533, Q86VH9 | Gene names | POLE, POLE1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase epsilon, catalytic subunit A (EC 2.7.7.7) (DNA polymerase II subunit A). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 28) | NC score | 0.037175 (rank : 40) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
CT032_HUMAN
|
||||||
| θ value | 0.47712 (rank : 29) | NC score | 0.031558 (rank : 58) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NQ75, Q96K09, Q9BYL5 | Gene names | HEFL, C20orf32 | |||
|
Domain Architecture |
|
|||||
| Description | HEF-like protein. | |||||
|
GP179_HUMAN
|
||||||
| θ value | 0.47712 (rank : 30) | NC score | 0.041159 (rank : 34) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6PRD1 | Gene names | GPR179, GPR158L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 179 precursor (Probable G-protein coupled receptor 158-like 1). | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 0.47712 (rank : 31) | NC score | 0.044139 (rank : 28) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
ZBED4_HUMAN
|
||||||
| θ value | 0.47712 (rank : 32) | NC score | 0.044864 (rank : 26) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75132, Q9UGG8 | Gene names | ZBED4, KIAA0637 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger BED domain-containing protein 4. | |||||
|
C8AP2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 33) | NC score | 0.042219 (rank : 33) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UKL3, Q6PH76, Q7LCQ7, Q86YD9, Q9NUQ4, Q9NZV9, Q9P2N1, Q9Y563 | Gene names | CASP8AP2, FLASH, KIAA1315, RIP25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
CPT1C_HUMAN
|
||||||
| θ value | 0.62314 (rank : 34) | NC score | 0.017778 (rank : 99) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8TCG5, Q8N6Q9, Q8NDS6, Q8TE84 | Gene names | CPT1C | |||
|
Domain Architecture |

|
|||||
| Description | Carnitine O-palmitoyltransferase I, brain isoform (EC 2.3.1.21) (Carnitine palmitoyltransferase 1C) (CPT IC). | |||||
|
NCOR1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 35) | NC score | 0.048881 (rank : 22) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q60974, Q60812 | Gene names | Ncor1, Rxrip13 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) (Retinoid X receptor- interacting protein 13) (RIP13). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 0.62314 (rank : 36) | NC score | 0.018029 (rank : 96) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
SFRS4_HUMAN
|
||||||
| θ value | 0.62314 (rank : 37) | NC score | 0.023381 (rank : 76) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q08170, Q5VXP1, Q9BUA4, Q9UEB5 | Gene names | SFRS4, SRP75 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4 (Pre-mRNA-splicing factor SRP75) (SRP001LB). | |||||
|
SP17_HUMAN
|
||||||
| θ value | 0.62314 (rank : 38) | NC score | 0.048651 (rank : 24) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15506, Q9BXF7 | Gene names | SPA17, SP17 | |||
|
Domain Architecture |
|
|||||
| Description | Sperm surface protein Sp17 (Sperm autoantigenic protein 17) (Sperm protein 17) (Sp17-1). | |||||
|
AKAP1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 39) | NC score | 0.034179 (rank : 50) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O08715, O08714, P97488 | Gene names | Akap1, Akap | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 1, mitochondrial precursor (Protein kinase A- anchoring protein 1) (PRKA1) (Dual specificity A-kinase-anchoring protein 1) (D-AKAP-1) (Spermatid A-kinase anchor protein) (S-AKAP). | |||||
|
SETD2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 40) | NC score | 0.032574 (rank : 53) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
MPDZ_MOUSE
|
||||||
| θ value | 1.06291 (rank : 41) | NC score | 0.016312 (rank : 109) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8VBX6, O08783, Q6P7U4, Q80ZY8, Q8BKJ1, Q8C0H8, Q8VBV5, Q8VBY0, Q9Z1K3 | Gene names | Mpdz, Mupp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multiple PDZ domain protein (Multi PDZ domain protein 1) (Multi-PDZ domain protein 1). | |||||
|
NAF1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 42) | NC score | 0.024136 (rank : 75) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 852 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9WUU8, Q922A9, Q922F7, Q9EPP8, Q9R0X3 | Gene names | Tnip1, Abin, Naf1 | |||
|
Domain Architecture |

|
|||||
| Description | Nef-associated factor 1 (Naf1) (TNFAIP3-interacting protein 1) (A20- binding inhibitor of NF-kappa-B activation) (ABIN) (Virion-associated nuclear shuttling protein) (VAN) (mVAN). | |||||
|
PDLI1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 43) | NC score | 0.030513 (rank : 61) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O70400, Q99K93 | Gene names | Pdlim1, Clim1 | |||
|
Domain Architecture |
|
|||||
| Description | PDZ and LIM domain protein 1 (Elfin) (LIM domain protein CLP-36) (C- terminal LIM domain protein 1). | |||||
|
RAI2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 44) | NC score | 0.033032 (rank : 52) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y5P3, Q8N6X7 | Gene names | RAI2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 2. | |||||
|
CCD46_MOUSE
|
||||||
| θ value | 1.38821 (rank : 45) | NC score | 0.019875 (rank : 90) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q5PR68, Q80ZN6, Q99JS4, Q9D9T1 | Gene names | Ccdc46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled coil domain-containing protein 46. | |||||
|
CLAP1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 46) | NC score | 0.021941 (rank : 80) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7Z460, O75118, Q8N5B8, Q9BQT5 | Gene names | CLASP1, KIAA0622, MAST1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CLIP-associating protein 1 (Cytoplasmic linker-associated protein 1) (Multiple asters homolog 1). | |||||
|
F10A4_HUMAN
|
||||||
| θ value | 1.38821 (rank : 47) | NC score | 0.020764 (rank : 86) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8IZP2 | Gene names | FAM10A4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM10A4. | |||||
|
IRS1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 48) | NC score | 0.021238 (rank : 82) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P35569 | Gene names | Irs1, Irs-1 | |||
|
Domain Architecture |
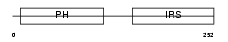
|
|||||
| Description | Insulin receptor substrate 1 (IRS-1). | |||||
|
MYH8_HUMAN
|
||||||
| θ value | 1.38821 (rank : 49) | NC score | 0.016305 (rank : 110) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1610 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P13535, Q14910 | Gene names | MYH8 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
RBM28_MOUSE
|
||||||
| θ value | 1.38821 (rank : 50) | NC score | 0.016760 (rank : 105) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8CGC6, Q8BG25, Q8VEJ8, Q9CS22, Q9CSE6 | Gene names | Rbm28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 28 (RNA-binding motif protein 28). | |||||
|
ABL2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 51) | NC score | 0.001281 (rank : 168) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1045 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P42684, Q5T0X6, Q6NZY6 | Gene names | ABL2, ABLL, ARG | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase ABL2 (EC 2.7.10.2) (Abelson murine leukemia viral oncogene homolog 2) (Tyrosine kinase ARG). | |||||
|
BARX2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 52) | NC score | 0.006556 (rank : 154) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UMQ3, O43518 | Gene names | BARX2 | |||
|
Domain Architecture |
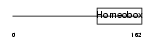
|
|||||
| Description | Homeobox protein BarH-like 2. | |||||
|
BRWD1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 53) | NC score | 0.012737 (rank : 128) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 478 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q921C3, Q921C2 | Gene names | Brwd1, Wdr9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and WD repeat domain-containing protein 1 (WD repeat protein 9). | |||||
|
CAC1H_MOUSE
|
||||||
| θ value | 1.81305 (rank : 54) | NC score | 0.010195 (rank : 142) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O88427, Q80TJ2, Q9JKU5 | Gene names | Cacna1h, Kiaa1120 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1H (Voltage- gated calcium channel subunit alpha Cav3.2). | |||||
|
CCNT2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 55) | NC score | 0.033761 (rank : 51) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O60583, O60582, Q5I1Y0 | Gene names | CCNT2 | |||
|
Domain Architecture |
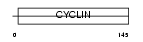
|
|||||
| Description | Cyclin-T2 (CycT2). | |||||
|
CT112_HUMAN
|
||||||
| θ value | 1.81305 (rank : 56) | NC score | 0.028841 (rank : 63) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96MY1, Q5JYB7, Q6P0Y4, Q9BR34, Q9NQF6 | Gene names | C20orf112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf112. | |||||
|
DGKG_MOUSE
|
||||||
| θ value | 1.81305 (rank : 57) | NC score | 0.011419 (rank : 133) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q91WG7 | Gene names | Dgkg, Dagk3 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase gamma (EC 2.7.1.107) (Diglyceride kinase gamma) (DGK-gamma) (DAG kinase gamma) (88 kDa diacylglycerol kinase). | |||||
|
KI18A_MOUSE
|
||||||
| θ value | 1.81305 (rank : 58) | NC score | 0.014125 (rank : 121) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91WD7, Q3TP77, Q8BLL1 | Gene names | Kif18a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 18A. | |||||
|
SNX11_MOUSE
|
||||||
| θ value | 1.81305 (rank : 59) | NC score | 0.038824 (rank : 37) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91WL6, Q3V3A6 | Gene names | Snx11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-11. | |||||
|
TTK_HUMAN
|
||||||
| θ value | 1.81305 (rank : 60) | NC score | 0.003182 (rank : 165) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 816 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P33981 | Gene names | TTK, MPS1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Dual specificity protein kinase TTK (EC 2.7.12.1) (Phosphotyrosine picked threonine-protein kinase) (PYT). | |||||
|
CCD62_HUMAN
|
||||||
| θ value | 2.36792 (rank : 61) | NC score | 0.023307 (rank : 77) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 631 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6P9F0, Q6ZVF2, Q86VJ0, Q9BYZ5 | Gene names | CCDC62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 60 (Aaa-protein) (Protein TSP- NY). | |||||
|
CHD1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 62) | NC score | 0.013349 (rank : 125) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
CXXC6_HUMAN
|
||||||
| θ value | 2.36792 (rank : 63) | NC score | 0.034964 (rank : 48) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
EEA1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 64) | NC score | 0.016009 (rank : 112) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1796 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q15075, Q14221 | Gene names | EEA1, ZFYVE2 | |||
|
Domain Architecture |

|
|||||
| Description | Early endosome antigen 1 (Endosome-associated protein p162) (Zinc finger FYVE domain-containing protein 2). | |||||
|
GOGB1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 65) | NC score | 0.017966 (rank : 98) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
LIPS_HUMAN
|
||||||
| θ value | 2.36792 (rank : 66) | NC score | 0.025958 (rank : 69) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q05469, Q3LRT2, Q6NSL7 | Gene names | LIPE | |||
|
Domain Architecture |
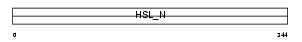
|
|||||
| Description | Hormone-sensitive lipase (EC 3.1.1.79) (HSL). | |||||
|
NOC3L_HUMAN
|
||||||
| θ value | 2.36792 (rank : 67) | NC score | 0.040855 (rank : 35) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8WTT2, Q9H5M6, Q9H9D8 | Gene names | NOC3L, AD24, C10orf117, FAD24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar complex protein 3 homolog (NOC3 protein homolog) (NOC3-like protein) (Nucleolar complex-associated protein 3-like protein) (Factor for adipocyte differentiation 24). | |||||
|
PHF3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.030312 (rank : 62) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q92576, Q5T1T6, Q9NQ16, Q9UI45 | Gene names | PHF3, KIAA0244 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 3. | |||||
|
SENP6_HUMAN
|
||||||
| θ value | 2.36792 (rank : 69) | NC score | 0.039699 (rank : 36) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9GZR1, O94891, Q8TBY4, Q9UJV5 | Gene names | SENP6, KIAA0797, SSP1, SUSP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 6 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP6) (SUMO-1-specific protease 1). | |||||
|
SN1L2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 70) | NC score | 0.000881 (rank : 169) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 964 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9H0K1, O94878, Q6AZE2, Q76N03, Q8NCV7, Q96CZ8 | Gene names | SNF1LK2, KIAA0781, QIK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SNF1-like kinase 2 (EC 2.7.11.1) (Qin- induced kinase). | |||||
|
TP53B_MOUSE
|
||||||
| θ value | 2.36792 (rank : 71) | NC score | 0.035481 (rank : 45) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 644 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P70399, Q68FD0, Q8CI97, Q91YC9 | Gene names | Tp53bp1, Trp53bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
TTF2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 72) | NC score | 0.020992 (rank : 84) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q5NC05, Q5M924 | Gene names | Ttf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 2 (EC 3.6.1.-) (RNA polymerase II termination factor) (Transcription release factor 2). | |||||
|
ADNP_HUMAN
|
||||||
| θ value | 3.0926 (rank : 73) | NC score | 0.035678 (rank : 43) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 265 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H2P0, O94881, Q9UG34 | Gene names | ADNP, KIAA0784 | |||
|
Domain Architecture |

|
|||||
| Description | Activity-dependent neuroprotector (Activity-dependent neuroprotective protein). | |||||
|
AHI1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 74) | NC score | 0.010677 (rank : 137) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 537 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8N157, Q504T3, Q5TCP9, Q6P098, Q6PIT6, Q8NDX0, Q9H0H2 | Gene names | AHI1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Jouberin (Abelson helper integration site 1 protein homolog) (AHI-1). | |||||
|
ANK1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 75) | NC score | 0.006050 (rank : 157) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 430 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q02357, P70440, P97446, P97941, Q3TZ35, Q3UH42, Q3UHP2, Q3UYY7, Q61302, Q61303, Q78E45 | Gene names | Ank1, Ank-1 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-1 (Erythrocyte ankyrin). | |||||
|
ASXL1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 76) | NC score | 0.022786 (rank : 78) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P59598 | Gene names | Asxl1, Kiaa0978 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative Polycomb group protein ASXL1 (Additional sex combs-like protein 1). | |||||
|
CT112_MOUSE
|
||||||
| θ value | 3.0926 (rank : 77) | NC score | 0.026076 (rank : 68) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6DIB4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf112 homolog. | |||||
|
LAP4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 78) | NC score | 0.007340 (rank : 151) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 590 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q80U72, Q6P5H7, Q7TPH8, Q80VB1, Q8CI48, Q8VII1, Q922S3 | Gene names | Scrib, Kiaa0147, Lap4, Scrib1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP4 (Protein scribble homolog). | |||||
|
RSF1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 79) | NC score | 0.050082 (rank : 21) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
SLC2B_HUMAN
|
||||||
| θ value | 3.0926 (rank : 80) | NC score | 0.032063 (rank : 56) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8NEV8, Q9Y4D6 | Gene names | EXPH5, KIAA0624, SLAC2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Slp homolog lacking C2 domains b (Exophilin-5). | |||||
|
TNIK_HUMAN
|
||||||
| θ value | 3.0926 (rank : 81) | NC score | 0.004911 (rank : 162) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1570 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9UKE5, O60298, Q8WUY7, Q9UKD8, Q9UKD9, Q9UKE0, Q9UKE1, Q9UKE2, Q9UKE3, Q9UKE4 | Gene names | TNIK, KIAA0551 | |||
|
Domain Architecture |

|
|||||
| Description | TRAF2 and NCK-interacting protein kinase (EC 2.7.11.1). | |||||
|
TTC3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 82) | NC score | 0.021134 (rank : 83) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 664 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P53804, O60767, P78476, P78477 | Gene names | TTC3, TPRD | |||
|
Domain Architecture |

|
|||||
| Description | Tetratricopeptide repeat protein 3 (TPR repeat protein 3) (TPR repeat protein D). | |||||
|
ANK1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 83) | NC score | 0.005515 (rank : 160) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P16157, O43400, Q13768, Q59FP2, Q8N604, Q99407 | Gene names | ANK1, ANK | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-1 (Erythrocyte ankyrin) (Ankyrin-R). | |||||
|
ATRX_HUMAN
|
||||||
| θ value | 4.03905 (rank : 84) | NC score | 0.024604 (rank : 73) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P46100, P51068, Q15886, Q59FB5, Q59H31, Q5H9A2, Q5JWI4, Q7Z2J1, Q9H0Z1, Q9NTS3 | Gene names | ATRX, RAD54L, XH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked helicase II) (X-linked nuclear protein) (XNP) (Znf- HX). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 4.03905 (rank : 85) | NC score | 0.028349 (rank : 65) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
CCNB3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 86) | NC score | 0.020795 (rank : 85) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8WWL7, Q96SB5, Q96SB6, Q96SB7, Q9NT38 | Gene names | CCNB3, CYCB3 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B3. | |||||
|
K0226_HUMAN
|
||||||
| θ value | 4.03905 (rank : 87) | NC score | 0.018012 (rank : 97) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92622 | Gene names | KIAA0226 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0226. | |||||
|
KCAB3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 88) | NC score | 0.012465 (rank : 129) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43448 | Gene names | KCNAB3, KCNA3B | |||
|
Domain Architecture |
|
|||||
| Description | Voltage-gated potassium channel subunit beta-3 (K(+) channel subunit beta-3) (Kv-beta-3). | |||||
|
MYO9B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 89) | NC score | 0.010797 (rank : 136) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q13459, O75314, Q9NUJ2, Q9UHN0 | Gene names | MYO9B, MYR5 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9B (Myosin IXb) (Unconventional myosin-9b). | |||||
|
NBR1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 90) | NC score | 0.017342 (rank : 100) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P97432, Q9JIH9 | Gene names | Nbr1, M17s2 | |||
|
Domain Architecture |

|
|||||
| Description | Next to BRCA1 gene 1 protein (Neighbor of BRCA1 gene 1 protein) (Membrane component, chromosome 17, surface marker 2). | |||||
|
NOC3L_MOUSE
|
||||||
| θ value | 4.03905 (rank : 91) | NC score | 0.038000 (rank : 39) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8VI84, Q9JKM3 | Gene names | Noc3l, Ad24, Fad24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar complex protein 3 homolog (NOC3 protein homolog) (NOC3-like protein) (Nucleolar complex-associated protein 3-like protein) (Factor for adipocyte differentiation 24). | |||||
|
PPRB_HUMAN
|
||||||
| θ value | 4.03905 (rank : 92) | NC score | 0.045779 (rank : 25) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q15648, O43810, O75447, Q9HD39 | Gene names | PPARBP, ARC205, DRIP205, TRAP220, TRIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2) (p53 regulatory protein RB18A) (Vitamin D receptor-interacting protein complex component DRIP205) (Activator- recruited cofactor 205 kDa component) (ARC205). | |||||
|
RBBP6_HUMAN
|
||||||
| θ value | 4.03905 (rank : 93) | NC score | 0.044507 (rank : 27) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
RNF34_HUMAN
|
||||||
| θ value | 4.03905 (rank : 94) | NC score | 0.013893 (rank : 123) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q969K3, Q8NG47, Q9H6W8 | Gene names | RNF34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 34 (RING finger protein RIFF) (FYVE-RING finger protein Momo) (Human RING finger homologous to inhibitor of apoptosis protein) (hRFI) (Caspases-8 and -10-associated RING finger protein 1) (CARP-1) (Caspase regulator CARP1). | |||||
|
RRP5_HUMAN
|
||||||
| θ value | 4.03905 (rank : 95) | NC score | 0.037063 (rank : 41) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q14690, Q86VQ8 | Gene names | PDCD11, KIAA0185 | |||
|
Domain Architecture |

|
|||||
| Description | RRP5 protein homolog (Programmed cell death protein 11). | |||||
|
TM119_HUMAN
|
||||||
| θ value | 4.03905 (rank : 96) | NC score | 0.024225 (rank : 74) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q4V9L6, Q6UXE5, Q8N2F5 | Gene names | TMEM119 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 119 precursor. | |||||
|
TRM6_MOUSE
|
||||||
| θ value | 4.03905 (rank : 97) | NC score | 0.025128 (rank : 71) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CE96, Q3TJZ8, Q3TME7, Q6ZPW8, Q80Y59, Q8CEU0 | Gene names | Trm6, Kiaa1153 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA (adenine-N(1)-)-methyltransferase non-catalytic subunit TRM6 (tRNA(m1A58)-methyltransferase subunit TRM6) (tRNA(m1A58)MTase subunit TRM6). | |||||
|
YLPM1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 98) | NC score | 0.024648 (rank : 72) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
ZN167_HUMAN
|
||||||
| θ value | 4.03905 (rank : 99) | NC score | -0.001799 (rank : 171) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9P0L1, Q6WL09, Q96FQ2 | Gene names | ZNF167, ZNF448, ZNF64 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 167 (Zinc finger protein 64) (Zinc finger protein 448) (ZFP). | |||||
|
AF17_HUMAN
|
||||||
| θ value | 5.27518 (rank : 100) | NC score | 0.012060 (rank : 131) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P55198, Q9UF49 | Gene names | MLLT6, AF17 | |||
|
Domain Architecture |
|
|||||
| Description | Protein AF-17. | |||||
|
ARI4A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 101) | NC score | 0.027478 (rank : 66) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P29374, Q15991, Q15992, Q15993 | Gene names | ARID4A, RBBP1, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 4A (ARID domain- containing protein 4A) (Retinoblastoma-binding protein 1) (RBBP-1). | |||||
|
CSTN1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 102) | NC score | 0.006989 (rank : 152) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O94985 | Gene names | CLSTN1, CS1, KIAA0911 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calsyntenin-1 precursor. | |||||
|
CT075_MOUSE
|
||||||
| θ value | 5.27518 (rank : 103) | NC score | 0.007457 (rank : 150) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P59383 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Uncharacterized protein C20orf75 homolog precursor. | |||||
|
GCC2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 104) | NC score | 0.015673 (rank : 114) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1619 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8CHG3, Q8BR44, Q8R2Q5, Q9CT45 | Gene names | Gcc2, Kiaa0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185). | |||||
|
ITSN1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 105) | NC score | 0.013870 (rank : 124) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1757 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q15811, O95216, Q9UET5, Q9UK60, Q9UNK1, Q9UNK2, Q9UQ92 | Gene names | ITSN1, ITSN, SH3D1A | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (SH3 domain-containing protein 1A) (SH3P17). | |||||
|
MAP1B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 106) | NC score | 0.030636 (rank : 60) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
MCPH1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 107) | NC score | 0.016971 (rank : 103) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NEM0, Q66GU1, Q9H9C7 | Gene names | MCPH1 | |||
|
Domain Architecture |

|
|||||
| Description | Microcephalin. | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 108) | NC score | 0.021794 (rank : 81) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
MYRIP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 109) | NC score | 0.015669 (rank : 115) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8NFW9, Q8IUF5, Q9Y3V4 | Gene names | MYRIP, SLAC2C | |||
|
Domain Architecture |
|
|||||
| Description | Rab effector MyRIP (Myosin-VIIa- and Rab-interacting protein) (Exophilin-8) (Slp homolog lacking C2 domains c). | |||||
|
PACN2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 110) | NC score | 0.011034 (rank : 135) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 339 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UNF0, O95921, Q96HV9, Q9H0D3, Q9NPN1, Q9Y4V2 | Gene names | PACSIN2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C and casein kinase substrate in neurons protein 2. | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 5.27518 (rank : 111) | NC score | 0.034345 (rank : 49) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
PREP_MOUSE
|
||||||
| θ value | 5.27518 (rank : 112) | NC score | 0.015739 (rank : 113) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K411, Q3THC4, Q3TJI1, Q3TPP0, Q3URQ8, Q4KUG2, Q4KUG3, Q6ZPX6, Q922N1, Q9CV63 | Gene names | Pitrm1, Kiaa1104, Ntup1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Presequence protease, mitochondrial precursor (EC 3.4.24.-) (Pitrilysin metalloproteinase 1). | |||||
|
RIB2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 113) | NC score | 0.017009 (rank : 102) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P04844, Q6IBA5, Q96E21, Q9BUQ3, Q9UBE1 | Gene names | RPN2 | |||
|
Domain Architecture |

|
|||||
| Description | Dolichyl-diphosphooligosaccharide--protein glycosyltransferase 63 kDa subunit precursor (EC 2.4.1.119) (Ribophorin II) (RPN-II) (RIBIIR). | |||||
|
SRPK2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 114) | NC score | 0.005162 (rank : 161) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 664 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P78362, O75220, O75221, Q6NUL0, Q6V1X2, Q8IYQ3 | Gene names | SRPK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SRPK2 (EC 2.7.11.1) (Serine/arginine- rich protein-specific kinase 2) (SR-protein-specific kinase 2) (SFRS protein kinase 2). | |||||
|
TR4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 115) | NC score | 0.004493 (rank : 163) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P49116, P55092 | Gene names | NR2C2, TAK1, TR4 | |||
|
Domain Architecture |

|
|||||
| Description | Orphan nuclear receptor TR4 (Orphan nuclear receptor TAK1). | |||||
|
TRAK1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 116) | NC score | 0.017081 (rank : 101) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UPV9, Q96B69 | Gene names | TRAK1, KIAA1042, OIP106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trafficking kinesin-binding protein 1 (106 kDa O-GlcNAc transferase- interacting protein). | |||||
|
UBP2L_MOUSE
|
||||||
| θ value | 5.27518 (rank : 117) | NC score | 0.035542 (rank : 44) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q80X50, Q8BIT6, Q8BIW4, Q8BJ01, Q8CIG7, Q8K102, Q9CRT6 | Gene names | Ubap2l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated protein 2-like. | |||||
|
ARFG3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 118) | NC score | 0.026777 (rank : 67) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NP61, Q9BSC6, Q9H9J0, Q9NT10, Q9NUP5, Q9Y4V3, Q9Y4V4 | Gene names | ARFGAP3, ARFGAP1 | |||
|
Domain Architecture |
|
|||||
| Description | ADP-ribosylation factor GTPase-activating protein 3 (ARF GAP 3). | |||||
|
BRCA2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 119) | NC score | 0.016419 (rank : 108) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P51587, O00183, O15008, Q13879 | Gene names | BRCA2, FANCD1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 2 susceptibility protein (Fanconi anemia group D1 protein). | |||||
|
C8AP2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 120) | NC score | 0.030923 (rank : 59) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9WUF3, Q9CSW2 | Gene names | Casp8ap2, Flash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
CCD41_MOUSE
|
||||||
| θ value | 6.88961 (rank : 121) | NC score | 0.015355 (rank : 117) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9D5R3, Q3U7X7, Q3UX57, Q80VF0 | Gene names | Ccdc41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41. | |||||
|
CHST3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 122) | NC score | 0.008002 (rank : 147) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7LGC8, O75099 | Gene names | CHST3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carbohydrate sulfotransferase 3 (EC 2.8.2.17) (Chondroitin 6- sulfotransferase) (Chondroitin 6-O-sulfotransferase 1) (C6ST-1) (C6ST) (Galactose/N-acetylglucosamine/N-acetylglucosamine 6-O- sulfotransferase 0) (GST-0). | |||||
|
CK5P2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 123) | NC score | 0.016795 (rank : 104) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1433 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q96SN8, Q7Z3L4, Q7Z3U1, Q7Z7I6, Q9BSW0, Q9H6J6, Q9HCD9, Q9NV90, Q9UIW9 | Gene names | CDK5RAP2, CEP215, KIAA1633 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48) (Centrosome-associated protein 215). | |||||
|
COIA1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 124) | NC score | 0.005542 (rank : 159) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 481 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P39061, Q60672, Q61437, Q62001, Q62002, Q9JK63 | Gene names | Col18a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVIII) chain precursor [Contains: Endostatin]. | |||||
|
EXO1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 125) | NC score | 0.019588 (rank : 91) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9QZ11, Q3TLM4, Q923A5 | Gene names | Exo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Exonuclease 1 (EC 3.1.-.-) (mExo1) (Exonuclease I). | |||||
|
FACD2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 126) | NC score | 0.012424 (rank : 130) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80V62, Q9CWC7 | Gene names | Fancd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group D2 protein homolog (Protein FACD2). | |||||
|
GOGA4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 127) | NC score | 0.016503 (rank : 107) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
GRSF1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 128) | NC score | 0.008559 (rank : 146) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C5Q4, Q8BR05, Q8BRG7 | Gene names | Grsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-rich sequence factor 1 (GRSF-1). | |||||
|
ICAL_MOUSE
|
||||||
| θ value | 6.88961 (rank : 129) | NC score | 0.016644 (rank : 106) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P51125, Q9EQV4, Q9EQV5, Q9QXQ3, Q9QXQ4, Q9R0N1 | Gene names | Cast | |||
|
Domain Architecture |

|
|||||
| Description | Calpastatin (Calpain inhibitor). | |||||
|
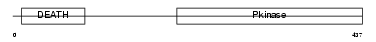
IRAK2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 130) | NC score | 0.004280 (rank : 164) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 714 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O43187 | Gene names | IRAK2 | |||
|
Domain Architecture |
|
|||||
| Description | Interleukin-1 receptor-associated kinase-like 2 (IRAK-2). | |||||
|
K1024_MOUSE
|
||||||
| θ value | 6.88961 (rank : 131) | NC score | 0.013066 (rank : 127) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8K3V7, Q69ZS9 | Gene names | Kiaa1024 | |||
|
Domain Architecture |

|
|||||
| Description | UPF0258 protein KIAA1024 (Protein DD1). | |||||
|
KTN1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 132) | NC score | 0.014029 (rank : 122) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1324 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q61595, Q8BG49, Q8BHF4, Q8BHM8, Q8C9Y5, Q8CG51, Q8CG52, Q8CG53, Q8CG54, Q8CG55, Q8CG56, Q8CG57, Q8CG58, Q8CG59, Q8CG60, Q8CG61, Q8CG62, Q8CG63 | Gene names | Ktn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin. | |||||
|
MX1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 133) | NC score | 0.007519 (rank : 149) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P09922 | Gene names | Mx1 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced GTP-binding protein Mx1 (Influenza resistance protein). | |||||
|
MYH11_MOUSE
|
||||||
| θ value | 6.88961 (rank : 134) | NC score | 0.011924 (rank : 132) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
NFASC_HUMAN
|
||||||
| θ value | 6.88961 (rank : 135) | NC score | 0.002108 (rank : 166) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O94856, Q5T2F0, Q5T2F1, Q5T2F2, Q5T2F3, Q5T2F4, Q5T2F5, Q5T2F6, Q5T2F7, Q5T2F9, Q5T2G0, Q5W9F8, Q68DH3, Q6ZQV6, Q7Z3K1, Q96HT1, Q96K50 | Gene names | NFASC, KIAA0756 | |||
|
Domain Architecture |

|
|||||
| Description | Neurofascin precursor. | |||||
|
NRBP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 136) | NC score | 0.005641 (rank : 158) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UHY1, Q53FZ5, Q96SU3 | Gene names | NRBP1, BCON3, NRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor-binding protein. | |||||
|
P80C_HUMAN
|
||||||
| θ value | 6.88961 (rank : 137) | NC score | 0.036750 (rank : 42) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P38432 | Gene names | COIL, CLN80 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coilin (p80). | |||||
|
PDZD2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 138) | NC score | 0.018403 (rank : 94) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O15018, Q9BXD4 | Gene names | PDZD2, AIPC, KIAA0300, PDZK3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 2 (PDZ domain-containing protein 3) (Activated in prostate cancer protein). | |||||
|
RBBP6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 139) | NC score | 0.038135 (rank : 38) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
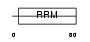
RBM7_HUMAN
|
||||||
| θ value | 6.88961 (rank : 140) | NC score | 0.010135 (rank : 143) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y580 | Gene names | RBM7 | |||
|
Domain Architecture |
|
|||||
| Description | RNA-binding protein 7 (RNA-binding motif protein 7). | |||||
|
RLF_HUMAN
|
||||||
| θ value | 6.88961 (rank : 141) | NC score | 0.007657 (rank : 148) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q13129, Q9NU60 | Gene names | RLF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein Rlf (Rearranged L-myc fusion gene protein) (Zn-15- related protein). | |||||
|
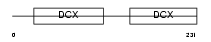
RP1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 142) | NC score | 0.011354 (rank : 134) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P56716 | Gene names | Rp1, Orp1, Rp1h | |||
|
Domain Architecture |
|
|||||
| Description | Oxygen-regulated protein 1 (Retinitis pigmentosa RP1 protein homolog). | |||||
|
SAFB2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 143) | NC score | 0.015515 (rank : 116) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q80YR5, Q8K153 | Gene names | Safb2 | |||
|
Domain Architecture |

|
|||||
| Description | Scaffold attachment factor B2. | |||||
|
SMG1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 144) | NC score | 0.014325 (rank : 119) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BKX6, Q5DU42, Q6ZQC0, Q8BLU4, Q8BWJ5, Q8BXD3 | Gene names | Smg1, Kiaa0421 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SMG1 (EC 2.7.11.1) (SMG-1). | |||||
|
SMYD1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 145) | NC score | 0.010412 (rank : 140) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P97443, P97442, P97444 | Gene names | Smyd1, Bop | |||
|
Domain Architecture |

|
|||||
| Description | SET and MYND domain-containing protein 1 (Zinc-finger protein BOP) (m- BOP) (CD8b-opposite). | |||||
|
SVIL_MOUSE
|
||||||
| θ value | 6.88961 (rank : 146) | NC score | 0.014264 (rank : 120) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8K4L3 | Gene names | Svil | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Supervillin (Archvillin) (p205/p250). | |||||
|
TRM6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 147) | NC score | 0.022603 (rank : 79) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UJA5, Q76P92, Q9BQV5, Q9ULR7 | Gene names | TRM6, KIAA1153 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA (adenine-N(1)-)-methyltransferase non-catalytic subunit TRM6 (tRNA(m1A58)-methyltransferase subunit TRM6) (tRNA(m1A58)MTase subunit TRM6). | |||||
|
CENPF_HUMAN
|
||||||
| θ value | 8.99809 (rank : 148) | NC score | 0.013235 (rank : 126) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
CHD4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 149) | NC score | 0.010309 (rank : 141) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6PDQ2 | Gene names | Chd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (CHD-4). | |||||
|
IPR1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 150) | NC score | 0.010458 (rank : 139) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BVK9, Q3UCV7, Q80V00 | Gene names | Ipr1, Ifi75 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intracellular pathogen resistance protein 1. | |||||
|
K0690_MOUSE
|
||||||
| θ value | 8.99809 (rank : 151) | NC score | 0.010633 (rank : 138) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6P5B0, Q7TMI5, Q80TU2 | Gene names | Kiaa0690 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0690. | |||||
|
LAD1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 152) | NC score | 0.019402 (rank : 92) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P57016 | Gene names | Lad1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (Linear IgA disease autoantigen). | |||||
|
MTF1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 153) | NC score | -0.001679 (rank : 170) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q07243 | Gene names | Mtf1, Mtf-1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-regulatory transcription factor 1 (Transcription factor MTF-1) (MRE-binding transcription factor). | |||||
|
MY18B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 154) | NC score | 0.014967 (rank : 118) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1299 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8IUG5, Q8NDI8, Q8TE65, Q8WWS0, Q96KH2, Q96KR8, Q96KR9 | Gene names | MYO18B | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18B (Myosin XVIIIb). | |||||
|
MYST4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 155) | NC score | 0.008760 (rank : 145) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8BRB7, Q7TNW5, Q8BG35, Q8C441, Q9JKX5 | Gene names | Myst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Querkopf protein). | |||||
|
NCOR2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 156) | NC score | 0.032195 (rank : 55) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
NPTN_MOUSE
|
||||||
| θ value | 8.99809 (rank : 157) | NC score | 0.006707 (rank : 153) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P97300, Q3U519, Q8C637, Q8C6H8 | Gene names | Nptn, Sdfr1, Sdr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuroplastin precursor (Stromal cell-derived receptor 1) (SDR-1). | |||||
|
PAR10_HUMAN
|
||||||
| θ value | 8.99809 (rank : 158) | NC score | 0.009400 (rank : 144) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q53GL7, Q8N2I0, Q8WV05, Q96CH7 | Gene names | PARP10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 10 (EC 2.4.2.30) (PARP-10). | |||||
|
RM03_HUMAN
|
||||||
| θ value | 8.99809 (rank : 159) | NC score | 0.019897 (rank : 89) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P09001 | Gene names | MRPL3, MRL3 | |||
|
Domain Architecture |
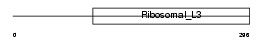
|
|||||
| Description | Mitochondrial 39S ribosomal protein L3 (L3mt) (MRP-L3). | |||||
|
ROBO1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 160) | NC score | 0.001499 (rank : 167) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 529 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y6N7, Q7Z300, Q9BUS7 | Gene names | ROBO1, DUTT1 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 1 precursor (H-Robo-1) (Deleted in U twenty twenty). | |||||
|
SL9A1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 161) | NC score | 0.006244 (rank : 156) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61165 | Gene names | Slc9a1, Nhe1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/hydrogen exchanger 1 (Na(+)/H(+) exchanger 1) (NHE-1) (Solute carrier family 9 member 1). | |||||
|
TCLB2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 162) | NC score | 0.020159 (rank : 88) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P56841 | Gene names | Tcl1b2 | |||
|
Domain Architecture |
|
|||||
| Description | TCL1B2 protein. | |||||
|
UBP2L_HUMAN
|
||||||
| θ value | 8.99809 (rank : 163) | NC score | 0.034975 (rank : 47) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q14157, Q5VU75, Q5VU76, Q9BTU3, Q9UGL5 | Gene names | UBAP2L, KIAA0144, NICE4 | |||
|
Domain Architecture |
|
|||||
| Description | Ubiquitin-associated protein 2-like (Protein NICE-4). | |||||
|
UBP3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 164) | NC score | 0.006530 (rank : 155) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y6I4 | Gene names | USP3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 3 (EC 3.1.2.15) (Ubiquitin thioesterase 3) (Ubiquitin-specific-processing protease 3) (Deubiquitinating enzyme 3). | |||||
|
UVRAG_HUMAN
|
||||||
| θ value | 8.99809 (rank : 165) | NC score | 0.016185 (rank : 111) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9P2Y5, O00392 | Gene names | UVRAG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UV radiation resistance-associated gene protein (p63). | |||||
|
VCXC_HUMAN
|
||||||
| θ value | 8.99809 (rank : 166) | NC score | 0.035351 (rank : 46) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9H321 | Gene names | VCXC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | VCX-C protein (Variably charged protein X-C). | |||||
|
VP13A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 167) | NC score | 0.018781 (rank : 93) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96RL7, Q5JSX9, Q5JSY0, Q5VYR5, Q702P4, Q709D0, Q86YF8, Q96S61, Q9H995, Q9Y2J1 | Gene names | VPS13A, CHAC, KIAA0986 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 13A (Chorein) (Chorea- acanthocytosis protein). | |||||
|
ZN207_HUMAN
|
||||||
| θ value | 8.99809 (rank : 168) | NC score | 0.025501 (rank : 70) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O43670, Q96HW5, Q9BUQ7 | Gene names | ZNF207 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 207. | |||||
|
ZN638_MOUSE
|
||||||
| θ value | 8.99809 (rank : 169) | NC score | 0.018125 (rank : 95) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q61464, Q6DFV9, Q8C941 | Gene names | Znf638, Np220, Zfml | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein). | |||||
|
SEMG2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 170) | NC score | 0.053028 (rank : 18) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q02383, Q6X2M6 | Gene names | SEMG2 | |||
|
Domain Architecture |

|
|||||
| Description | Semenogelin-2 precursor (Semenogelin II) (SGII). | |||||
|
SGP50_MOUSE
|
||||||
| θ value | θ > 10 (rank : 171) | NC score | 0.096443 (rank : 9) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q02596 | Gene names | Glycam1 | |||
|
Domain Architecture |

|
|||||
| Description | Sulfated 50 kDa glycoprotein precursor (SGP50) (Endothelial ligand FOR L-selectin) (Glycosylation-dependent cell adhesion molecule 1) (GlyCAM-1) (MC26). | |||||
|
DPOLZ_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 169 | |
| SwissProt Accessions | Q61493, Q9JMD6, Q9QWX6 | Gene names | Rev3l, Polz, Sez4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (Seizure-related protein 4). | |||||
|
DPOLZ_HUMAN
|
||||||
| NC score | 0.967263 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O60673, O43214 | Gene names | REV3L, POLZ, REV3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (hREV3). | |||||
|
DPOD1_MOUSE
|
||||||
| NC score | 0.808349 (rank : 3) | θ value | 5.6855e-87 (rank : 3) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P52431, O54883 | Gene names | Pold1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase delta catalytic subunit (EC 2.7.7.7). | |||||
|
DPOD1_HUMAN
|
||||||
| NC score | 0.808089 (rank : 4) | θ value | 2.82168e-86 (rank : 4) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P28340, Q8NER3, Q96H98 | Gene names | POLD1, POLD | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase delta catalytic subunit (EC 2.7.7.7) (DNA polymerase subunit delta p125). | |||||
|
DPOLA_HUMAN
|
||||||
| NC score | 0.640583 (rank : 5) | θ value | 1.79631e-32 (rank : 6) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P09884, Q86UQ7 | Gene names | POLA1, POLA | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase alpha catalytic subunit (EC 2.7.7.7). | |||||
|
DPOLA_MOUSE
|
||||||
| NC score | 0.639734 (rank : 6) | θ value | 1.6247e-33 (rank : 5) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P33609 | Gene names | Pola1, Pola | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase alpha catalytic subunit (EC 2.7.7.7). | |||||
|
DPOE1_MOUSE
|
||||||
| NC score | 0.098604 (rank : 7) | θ value | 0.21417 (rank : 19) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9WVF7, Q9QX50 | Gene names | Pole, Pole1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase epsilon, catalytic subunit A (EC 2.7.7.7) (DNA polymerase II subunit A). | |||||
|
DPOE1_HUMAN
|
||||||
| NC score | 0.098013 (rank : 8) | θ value | 0.365318 (rank : 27) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q07864, Q13533, Q86VH9 | Gene names | POLE, POLE1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase epsilon, catalytic subunit A (EC 2.7.7.7) (DNA polymerase II subunit A). | |||||
|
SGP50_MOUSE
|
||||||
| NC score | 0.096443 (rank : 9) | θ value | θ > 10 (rank : 171) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q02596 | Gene names | Glycam1 | |||
|
Domain Architecture |

|
|||||
| Description | Sulfated 50 kDa glycoprotein precursor (SGP50) (Endothelial ligand FOR L-selectin) (Glycosylation-dependent cell adhesion molecule 1) (GlyCAM-1) (MC26). | |||||
|
CASG_MOUSE
|
||||||
| NC score | 0.083047 (rank : 10) | θ value | 0.279714 (rank : 23) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q02862 | Gene names | Csng | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gamma-casein precursor (PP22). | |||||
|
SYCP2_HUMAN
|
||||||
| NC score | 0.068505 (rank : 11) | θ value | 0.0431538 (rank : 13) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BX26, O75763, Q5JX11, Q9NTX8, Q9UG27 | Gene names | SYCP2, SCP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptonemal complex protein 2 (SCP-2) (Synaptonemal complex lateral element protein) (hsSCP2). | |||||
|
UBP1_HUMAN
|
||||||
| NC score | 0.067989 (rank : 12) | θ value | 0.0148317 (rank : 9) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O94782, Q9UFR0, Q9UNJ3 | Gene names | USP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 1 (EC 3.1.2.15) (Ubiquitin thioesterase 1) (Ubiquitin-specific-processing protease 1) (Deubiquitinating enzyme 1) (hUBP). | |||||
|
NU153_HUMAN
|
||||||
| NC score | 0.065326 (rank : 13) | θ value | 0.0563607 (rank : 14) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P49790 | Gene names | NUP153 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup153 (Nucleoporin Nup153) (153 kDa nucleoporin). | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.064614 (rank : 14) | θ value | 0.279714 (rank : 25) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
PPRB_MOUSE
|
||||||
| NC score | 0.060477 (rank : 15) | θ value | 0.0736092 (rank : 16) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q925J9, O88323, Q3UHV0, Q6AXD5, Q8BW37, Q8BX19, Q8VDQ7, Q925K0 | Gene names | Pparbp, Trip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2). | |||||
|
NCOR1_HUMAN
|
||||||
| NC score | 0.058422 (rank : 16) | θ value | 0.0431538 (rank : 12) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O75376, Q9UPV5, Q9UQ18 | Gene names | NCOR1, KIAA1047 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR). | |||||
|
DNMT1_HUMAN
|
||||||
| NC score | 0.056662 (rank : 17) | θ value | 0.0431538 (rank : 11) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P26358, Q9UHG5, Q9ULA2, Q9UMZ6 | Gene names | DNMT1, AIM, DNMT | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase HsaI) (DNA MTase HsaI) (MCMT) (M.HsaI). | |||||
|
SEMG2_HUMAN
|
||||||
| NC score | 0.053028 (rank : 18) | θ value | θ > 10 (rank : 170) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q02383, Q6X2M6 | Gene names | SEMG2 | |||
|
Domain Architecture |

|
|||||
| Description | Semenogelin-2 precursor (Semenogelin II) (SGII). | |||||
|
DSPP_MOUSE
|
||||||
| NC score | 0.052061 (rank : 19) | θ value | 0.21417 (rank : 20) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
PDLI1_HUMAN
|
||||||
| NC score | 0.050875 (rank : 20) | θ value | 0.0148317 (rank : 8) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O00151, Q5VZH5, Q9BPZ9 | Gene names | PDLIM1, CLIM1, CLP36 | |||
|
Domain Architecture |

|
|||||
| Description | PDZ and LIM domain protein 1 (Elfin) (LIM domain protein CLP-36) (C- terminal LIM domain protein 1). | |||||
|
RSF1_HUMAN
|
||||||
| NC score | 0.050082 (rank : 21) | θ value | 3.0926 (rank : 79) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
NCOR1_MOUSE
|
||||||
| NC score | 0.048881 (rank : 22) | θ value | 0.62314 (rank : 35) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q60974, Q60812 | Gene names | Ncor1, Rxrip13 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) (Retinoid X receptor- interacting protein 13) (RIP13). | |||||
|
TAF3_MOUSE
|
||||||
| NC score | 0.048655 (rank : 23) | θ value | 0.21417 (rank : 22) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 463 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q5HZG4, Q3U490, Q3UWX2, Q8BIU8, Q99JH4 | Gene names | Taf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
SP17_HUMAN
|
||||||
| NC score | 0.048651 (rank : 24) | θ value | 0.62314 (rank : 38) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15506, Q9BXF7 | Gene names | SPA17, SP17 | |||
|
Domain Architecture |

|
|||||
| Description | Sperm surface protein Sp17 (Sperm autoantigenic protein 17) (Sperm protein 17) (Sp17-1). | |||||
|
PPRB_HUMAN
|
||||||
| NC score | 0.045779 (rank : 25) | θ value | 4.03905 (rank : 92) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q15648, O43810, O75447, Q9HD39 | Gene names | PPARBP, ARC205, DRIP205, TRAP220, TRIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2) (p53 regulatory protein RB18A) (Vitamin D receptor-interacting protein complex component DRIP205) (Activator- recruited cofactor 205 kDa component) (ARC205). | |||||
|
ZBED4_HUMAN
|
||||||
| NC score | 0.044864 (rank : 26) | θ value | 0.47712 (rank : 32) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75132, Q9UGG8 | Gene names | ZBED4, KIAA0637 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger BED domain-containing protein 4. | |||||
|
RBBP6_HUMAN
|
||||||
| NC score | 0.044507 (rank : 27) | θ value | 4.03905 (rank : 93) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.044139 (rank : 28) | θ value | 0.47712 (rank : 31) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
ATX2_HUMAN
|
||||||
| NC score | 0.043943 (rank : 29) | θ value | 0.0736092 (rank : 15) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 375 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99700, Q6ZQZ7, Q99493 | Gene names | ATXN2, ATX2, SCA2, TNRC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2 (Spinocerebellar ataxia type 2 protein) (Trinucleotide repeat-containing gene 13 protein). | |||||
|
AFF1_MOUSE
|
||||||
| NC score | 0.042829 (rank : 30) | θ value | 0.365318 (rank : 26) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O88573 | Gene names | Aff1, Mllt2, Mllt2h | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.042627 (rank : 31) | θ value | 0.163984 (rank : 17) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
OXR1_HUMAN
|
||||||
| NC score | 0.042360 (rank : 32) | θ value | 0.21417 (rank : 21) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8N573, Q3LIB5, Q6ZVK9, Q8N8V0, Q9H266, Q9NWC7 | Gene names | OXR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1. | |||||
|
C8AP2_HUMAN
|
||||||
| NC score | 0.042219 (rank : 33) | θ value | 0.62314 (rank : 33) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UKL3, Q6PH76, Q7LCQ7, Q86YD9, Q9NUQ4, Q9NZV9, Q9P2N1, Q9Y563 | Gene names | CASP8AP2, FLASH, KIAA1315, RIP25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
GP179_HUMAN
|
||||||
| NC score | 0.041159 (rank : 34) | θ value | 0.47712 (rank : 30) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6PRD1 | Gene names | GPR179, GPR158L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 179 precursor (Probable G-protein coupled receptor 158-like 1). | |||||
|
NOC3L_HUMAN
|
||||||
| NC score | 0.040855 (rank : 35) | θ value | 2.36792 (rank : 67) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8WTT2, Q9H5M6, Q9H9D8 | Gene names | NOC3L, AD24, C10orf117, FAD24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar complex protein 3 homolog (NOC3 protein homolog) (NOC3-like protein) (Nucleolar complex-associated protein 3-like protein) (Factor for adipocyte differentiation 24). | |||||
|
SENP6_HUMAN
|
||||||
| NC score | 0.039699 (rank : 36) | θ value | 2.36792 (rank : 69) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9GZR1, O94891, Q8TBY4, Q9UJV5 | Gene names | SENP6, KIAA0797, SSP1, SUSP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 6 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP6) (SUMO-1-specific protease 1). | |||||
|
SNX11_MOUSE
|
||||||
| NC score | 0.038824 (rank : 37) | θ value | 1.81305 (rank : 59) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91WL6, Q3V3A6 | Gene names | Snx11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-11. | |||||
|
RBBP6_MOUSE
|
||||||
| NC score | 0.038135 (rank : 38) | θ value | 6.88961 (rank : 139) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
NOC3L_MOUSE
|
||||||
| NC score | 0.038000 (rank : 39) | θ value | 4.03905 (rank : 91) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8VI84, Q9JKM3 | Gene names | Noc3l, Ad24, Fad24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar complex protein 3 homolog (NOC3 protein homolog) (NOC3-like protein) (Nucleolar complex-associated protein 3-like protein) (Factor for adipocyte differentiation 24). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.037175 (rank : 40) | θ value | 0.365318 (rank : 28) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
RRP5_HUMAN
|
||||||
| NC score | 0.037063 (rank : 41) | θ value | 4.03905 (rank : 95) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q14690, Q86VQ8 | Gene names | PDCD11, KIAA0185 | |||
|
Domain Architecture |

|
|||||
| Description | RRP5 protein homolog (Programmed cell death protein 11). | |||||
|
P80C_HUMAN
|
||||||
| NC score | 0.036750 (rank : 42) | θ value | 6.88961 (rank : 137) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P38432 | Gene names | COIL, CLN80 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coilin (p80). | |||||
|
ADNP_HUMAN
|
||||||
| NC score | 0.035678 (rank : 43) | θ value | 3.0926 (rank : 73) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 265 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H2P0, O94881, Q9UG34 | Gene names | ADNP, KIAA0784 | |||
|
Domain Architecture |

|
|||||
| Description | Activity-dependent neuroprotector (Activity-dependent neuroprotective protein). | |||||
|
UBP2L_MOUSE
|
||||||
| NC score | 0.035542 (rank : 44) | θ value | 5.27518 (rank : 117) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q80X50, Q8BIT6, Q8BIW4, Q8BJ01, Q8CIG7, Q8K102, Q9CRT6 | Gene names | Ubap2l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated protein 2-like. | |||||
|
TP53B_MOUSE
|
||||||
| NC score | 0.035481 (rank : 45) | θ value | 2.36792 (rank : 71) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 644 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P70399, Q68FD0, Q8CI97, Q91YC9 | Gene names | Tp53bp1, Trp53bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
VCXC_HUMAN
|
||||||
| NC score | 0.035351 (rank : 46) | θ value | 8.99809 (rank : 166) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9H321 | Gene names | VCXC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | VCX-C protein (Variably charged protein X-C). | |||||
|
UBP2L_HUMAN
|
||||||
| NC score | 0.034975 (rank : 47) | θ value | 8.99809 (rank : 163) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q14157, Q5VU75, Q5VU76, Q9BTU3, Q9UGL5 | Gene names | UBAP2L, KIAA0144, NICE4 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-associated protein 2-like (Protein NICE-4). | |||||
|
CXXC6_HUMAN
|
||||||
| NC score | 0.034964 (rank : 48) | θ value | 2.36792 (rank : 63) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.034345 (rank : 49) | θ value | 5.27518 (rank : 111) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
AKAP1_MOUSE
|
||||||
| NC score | 0.034179 (rank : 50) | θ value | 0.813845 (rank : 39) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O08715, O08714, P97488 | Gene names | Akap1, Akap | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 1, mitochondrial precursor (Protein kinase A- anchoring protein 1) (PRKA1) (Dual specificity A-kinase-anchoring protein 1) (D-AKAP-1) (Spermatid A-kinase anchor protein) (S-AKAP). | |||||
|
CCNT2_HUMAN
|
||||||
| NC score | 0.033761 (rank : 51) | θ value | 1.81305 (rank : 55) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O60583, O60582, Q5I1Y0 | Gene names | CCNT2 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-T2 (CycT2). | |||||
|
RAI2_HUMAN
|
||||||
| NC score | 0.033032 (rank : 52) | θ value | 1.06291 (rank : 44) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y5P3, Q8N6X7 | Gene names | RAI2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 2. | |||||
|
SETD2_HUMAN
|
||||||
| NC score | 0.032574 (rank : 53) | θ value | 0.813845 (rank : 40) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
TBCD1_HUMAN
|
||||||
| NC score | 0.032405 (rank : 54) | θ value | 0.163984 (rank : 18) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q86TI0, Q96K82, Q9UPP4 | Gene names | TBC1D1, KIAA1108 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 1. | |||||
|
NCOR2_HUMAN
|
||||||
| NC score | 0.032195 (rank : 55) | θ value | 8.99809 (rank : 156) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
SLC2B_HUMAN
|
||||||
| NC score | 0.032063 (rank : 56) | θ value | 3.0926 (rank : 80) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8NEV8, Q9Y4D6 | Gene names | EXPH5, KIAA0624, SLAC2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Slp homolog lacking C2 domains b (Exophilin-5). | |||||
|
DLGP4_HUMAN
|
||||||
| NC score | 0.031610 (rank : 57) | θ value | 0.279714 (rank : 24) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y2H0, Q5T2Y4, Q5T2Y5, Q9H137, Q9H138, Q9H1L7 | Gene names | DLGAP4, DAP4, KIAA0964 | |||
|
Domain Architecture |

|
|||||
| Description | Disks large-associated protein 4 (DAP-4) (SAP90/PSD-95-associated protein 4) (SAPAP4) (PSD-95/SAP90-binding protein 4). | |||||
|
CT032_HUMAN
|
||||||
| NC score | 0.031558 (rank : 58) | θ value | 0.47712 (rank : 29) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NQ75, Q96K09, Q9BYL5 | Gene names | HEFL, C20orf32 | |||
|
Domain Architecture |

|
|||||
| Description | HEF-like protein. | |||||
|
C8AP2_MOUSE
|
||||||
| NC score | 0.030923 (rank : 59) | θ value | 6.88961 (rank : 120) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9WUF3, Q9CSW2 | Gene names | Casp8ap2, Flash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
MAP1B_MOUSE
|
||||||
| NC score | 0.030636 (rank : 60) | θ value | 5.27518 (rank : 106) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
PDLI1_MOUSE
|
||||||
| NC score | 0.030513 (rank : 61) | θ value | 1.06291 (rank : 43) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O70400, Q99K93 | Gene names | Pdlim1, Clim1 | |||
|
Domain Architecture |

|
|||||
| Description | PDZ and LIM domain protein 1 (Elfin) (LIM domain protein CLP-36) (C- terminal LIM domain protein 1). | |||||
|
PHF3_HUMAN
|
||||||
| NC score | 0.030312 (rank : 62) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q92576, Q5T1T6, Q9NQ16, Q9UI45 | Gene names | PHF3, KIAA0244 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 3. | |||||
|
CT112_HUMAN
|
||||||
| NC score | 0.028841 (rank : 63) | θ value | 1.81305 (rank : 56) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96MY1, Q5JYB7, Q6P0Y4, Q9BR34, Q9NQF6 | Gene names | C20orf112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf112. | |||||
|
CENPE_HUMAN
|
||||||
| NC score | 0.028775 (rank : 64) | θ value | 0.00228821 (rank : 7) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.028349 (rank : 65) | θ value | 4.03905 (rank : 85) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
ARI4A_HUMAN
|
||||||
| NC score | 0.027478 (rank : 66) | θ value | 5.27518 (rank : 101) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P29374, Q15991, Q15992, Q15993 | Gene names | ARID4A, RBBP1, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 4A (ARID domain- containing protein 4A) (Retinoblastoma-binding protein 1) (RBBP-1). | |||||
|
ARFG3_HUMAN
|
||||||
| NC score | 0.026777 (rank : 67) | θ value | 6.88961 (rank : 118) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NP61, Q9BSC6, Q9H9J0, Q9NT10, Q9NUP5, Q9Y4V3, Q9Y4V4 | Gene names | ARFGAP3, ARFGAP1 | |||
|
Domain Architecture |

|
|||||
| Description | ADP-ribosylation factor GTPase-activating protein 3 (ARF GAP 3). | |||||
|
CT112_MOUSE
|
||||||
| NC score | 0.026076 (rank : 68) | θ value | 3.0926 (rank : 77) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6DIB4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf112 homolog. | |||||
|
LIPS_HUMAN
|
||||||
| NC score | 0.025958 (rank : 69) | θ value | 2.36792 (rank : 66) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q05469, Q3LRT2, Q6NSL7 | Gene names | LIPE | |||
|
Domain Architecture |

|
|||||
| Description | Hormone-sensitive lipase (EC 3.1.1.79) (HSL). | |||||
|
ZN207_HUMAN
|
||||||
| NC score | 0.025501 (rank : 70) | θ value | 8.99809 (rank : 168) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O43670, Q96HW5, Q9BUQ7 | Gene names | ZNF207 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 207. | |||||
|
TRM6_MOUSE
|
||||||
| NC score | 0.025128 (rank : 71) | θ value | 4.03905 (rank : 97) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CE96, Q3TJZ8, Q3TME7, Q6ZPW8, Q80Y59, Q8CEU0 | Gene names | Trm6, Kiaa1153 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA (adenine-N(1)-)-methyltransferase non-catalytic subunit TRM6 (tRNA(m1A58)-methyltransferase subunit TRM6) (tRNA(m1A58)MTase subunit TRM6). | |||||
|
YLPM1_HUMAN
|
||||||
| NC score | 0.024648 (rank : 72) | θ value | 4.03905 (rank : 98) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
ATRX_HUMAN
|
||||||
| NC score | 0.024604 (rank : 73) | θ value | 4.03905 (rank : 84) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P46100, P51068, Q15886, Q59FB5, Q59H31, Q5H9A2, Q5JWI4, Q7Z2J1, Q9H0Z1, Q9NTS3 | Gene names | ATRX, RAD54L, XH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked helicase II) (X-linked nuclear protein) (XNP) (Znf- HX). | |||||
|
TM119_HUMAN
|
||||||
| NC score | 0.024225 (rank : 74) | θ value | 4.03905 (rank : 96) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q4V9L6, Q6UXE5, Q8N2F5 | Gene names | TMEM119 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 119 precursor. | |||||
|
NAF1_MOUSE
|
||||||
| NC score | 0.024136 (rank : 75) | θ value | 1.06291 (rank : 42) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 852 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9WUU8, Q922A9, Q922F7, Q9EPP8, Q9R0X3 | Gene names | Tnip1, Abin, Naf1 | |||
|
Domain Architecture |

|
|||||
| Description | Nef-associated factor 1 (Naf1) (TNFAIP3-interacting protein 1) (A20- binding inhibitor of NF-kappa-B activation) (ABIN) (Virion-associated nuclear shuttling protein) (VAN) (mVAN). | |||||
|
SFRS4_HUMAN
|
||||||
| NC score | 0.023381 (rank : 76) | θ value | 0.62314 (rank : 37) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q08170, Q5VXP1, Q9BUA4, Q9UEB5 | Gene names | SFRS4, SRP75 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4 (Pre-mRNA-splicing factor SRP75) (SRP001LB). | |||||
|
CCD62_HUMAN
|
||||||
| NC score | 0.023307 (rank : 77) | θ value | 2.36792 (rank : 61) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 631 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6P9F0, Q6ZVF2, Q86VJ0, Q9BYZ5 | Gene names | CCDC62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 60 (Aaa-protein) (Protein TSP- NY). | |||||
|
ASXL1_MOUSE
|
||||||
| NC score | 0.022786 (rank : 78) | θ value | 3.0926 (rank : 76) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P59598 | Gene names | Asxl1, Kiaa0978 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative Polycomb group protein ASXL1 (Additional sex combs-like protein 1). | |||||
|
TRM6_HUMAN
|
||||||
| NC score | 0.022603 (rank : 79) | θ value | 6.88961 (rank : 147) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UJA5, Q76P92, Q9BQV5, Q9ULR7 | Gene names | TRM6, KIAA1153 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA (adenine-N(1)-)-methyltransferase non-catalytic subunit TRM6 (tRNA(m1A58)-methyltransferase subunit TRM6) (tRNA(m1A58)MTase subunit TRM6). | |||||
|
CLAP1_HUMAN
|
||||||
| NC score | 0.021941 (rank : 80) | θ value | 1.38821 (rank : 46) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7Z460, O75118, Q8N5B8, Q9BQT5 | Gene names | CLASP1, KIAA0622, MAST1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CLIP-associating protein 1 (Cytoplasmic linker-associated protein 1) (Multiple asters homolog 1). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.021794 (rank : 81) | θ value | 5.27518 (rank : 108) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
IRS1_MOUSE
|
||||||
| NC score | 0.021238 (rank : 82) | θ value | 1.38821 (rank : 48) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P35569 | Gene names | Irs1, Irs-1 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin receptor substrate 1 (IRS-1). | |||||
|
TTC3_HUMAN
|
||||||
| NC score | 0.021134 (rank : 83) | θ value | 3.0926 (rank : 82) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 664 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P53804, O60767, P78476, P78477 | Gene names | TTC3, TPRD | |||
|
Domain Architecture |

|
|||||
| Description | Tetratricopeptide repeat protein 3 (TPR repeat protein 3) (TPR repeat protein D). | |||||
|
TTF2_MOUSE
|
||||||
| NC score | 0.020992 (rank : 84) | θ value | 2.36792 (rank : 72) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q5NC05, Q5M924 | Gene names | Ttf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 2 (EC 3.6.1.-) (RNA polymerase II termination factor) (Transcription release factor 2). | |||||
|
CCNB3_HUMAN
|
||||||
| NC score | 0.020795 (rank : 85) | θ value | 4.03905 (rank : 86) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8WWL7, Q96SB5, Q96SB6, Q96SB7, Q9NT38 | Gene names | CCNB3, CYCB3 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B3. | |||||
|
F10A4_HUMAN
|
||||||
| NC score | 0.020764 (rank : 86) | θ value | 1.38821 (rank : 47) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8IZP2 | Gene names | FAM10A4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM10A4. | |||||
|
CSPG2_MOUSE
|
||||||
| NC score | 0.020474 (rank : 87) | θ value | 0.0330416 (rank : 10) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 711 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q62059, Q62058, Q9CUU0 | Gene names | Cspg2 | |||
|
Domain Architecture |

|
|||||
| Description | Versican core protein precursor (Large fibroblast proteoglycan) (Chondroitin sulfate proteoglycan core protein 2) (PG-M). | |||||
|
TCLB2_MOUSE
|
||||||
| NC score | 0.020159 (rank : 88) | θ value | 8.99809 (rank : 162) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P56841 | Gene names | Tcl1b2 | |||
|
Domain Architecture |

|
|||||
| Description | TCL1B2 protein. | |||||
|
RM03_HUMAN
|
||||||
| NC score | 0.019897 (rank : 89) | θ value | 8.99809 (rank : 159) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P09001 | Gene names | MRPL3, MRL3 | |||
|
Domain Architecture |

|
|||||
| Description | Mitochondrial 39S ribosomal protein L3 (L3mt) (MRP-L3). | |||||
|
CCD46_MOUSE
|
||||||
| NC score | 0.019875 (rank : 90) | θ value | 1.38821 (rank : 45) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q5PR68, Q80ZN6, Q99JS4, Q9D9T1 | Gene names | Ccdc46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled coil domain-containing protein 46. | |||||
|
EXO1_MOUSE
|
||||||
| NC score | 0.019588 (rank : 91) | θ value | 6.88961 (rank : 125) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9QZ11, Q3TLM4, Q923A5 | Gene names | Exo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Exonuclease 1 (EC 3.1.-.-) (mExo1) (Exonuclease I). | |||||
|
LAD1_MOUSE
|
||||||
| NC score | 0.019402 (rank : 92) | θ value | 8.99809 (rank : 152) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P57016 | Gene names | Lad1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (Linear IgA disease autoantigen). | |||||
|
VP13A_HUMAN
|
||||||
| NC score | 0.018781 (rank : 93) | θ value | 8.99809 (rank : 167) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96RL7, Q5JSX9, Q5JSY0, Q5VYR5, Q702P4, Q709D0, Q86YF8, Q96S61, Q9H995, Q9Y2J1 | Gene names | VPS13A, CHAC, KIAA0986 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 13A (Chorein) (Chorea- acanthocytosis protein). | |||||
|
PDZD2_HUMAN
|
||||||
| NC score | 0.018403 (rank : 94) | θ value | 6.88961 (rank : 138) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O15018, Q9BXD4 | Gene names | PDZD2, AIPC, KIAA0300, PDZK3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 2 (PDZ domain-containing protein 3) (Activated in prostate cancer protein). | |||||
|
ZN638_MOUSE
|
||||||
| NC score | 0.018125 (rank : 95) | θ value | 8.99809 (rank : 169) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q61464, Q6DFV9, Q8C941 | Gene names | Znf638, Np220, Zfml | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.018029 (rank : 96) | θ value | 0.62314 (rank : 36) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
K0226_HUMAN
|
||||||
| NC score | 0.018012 (rank : 97) | θ value | 4.03905 (rank : 87) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92622 | Gene names | KIAA0226 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0226. | |||||
|
GOGB1_HUMAN
|
||||||
| NC score | 0.017966 (rank : 98) | θ value | 2.36792 (rank : 65) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
CPT1C_HUMAN
|
||||||
| NC score | 0.017778 (rank : 99) | θ value | 0.62314 (rank : 34) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8TCG5, Q8N6Q9, Q8NDS6, Q8TE84 | Gene names | CPT1C | |||
|
Domain Architecture |

|
|||||
| Description | Carnitine O-palmitoyltransferase I, brain isoform (EC 2.3.1.21) (Carnitine palmitoyltransferase 1C) (CPT IC). | |||||
|
NBR1_MOUSE
|
||||||
| NC score | 0.017342 (rank : 100) | θ value | 4.03905 (rank : 90) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P97432, Q9JIH9 | Gene names | Nbr1, M17s2 | |||
|
Domain Architecture |

|
|||||
| Description | Next to BRCA1 gene 1 protein (Neighbor of BRCA1 gene 1 protein) (Membrane component, chromosome 17, surface marker 2). | |||||
|
TRAK1_HUMAN
|
||||||
| NC score | 0.017081 (rank : 101) | θ value | 5.27518 (rank : 116) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UPV9, Q96B69 | Gene names | TRAK1, KIAA1042, OIP106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trafficking kinesin-binding protein 1 (106 kDa O-GlcNAc transferase- interacting protein). | |||||
|
RIB2_HUMAN
|
||||||
| NC score | 0.017009 (rank : 102) | θ value | 5.27518 (rank : 113) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P04844, Q6IBA5, Q96E21, Q9BUQ3, Q9UBE1 | Gene names | RPN2 | |||
|
Domain Architecture |

|
|||||
| Description | Dolichyl-diphosphooligosaccharide--protein glycosyltransferase 63 kDa subunit precursor (EC 2.4.1.119) (Ribophorin II) (RPN-II) (RIBIIR). | |||||
|
MCPH1_HUMAN
|
||||||
| NC score | 0.016971 (rank : 103) | θ value | 5.27518 (rank : 107) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NEM0, Q66GU1, Q9H9C7 | Gene names | MCPH1 | |||
|
Domain Architecture |

|
|||||
| Description | Microcephalin. | |||||
|
CK5P2_HUMAN
|
||||||
| NC score | 0.016795 (rank : 104) | θ value | 6.88961 (rank : 123) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1433 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q96SN8, Q7Z3L4, Q7Z3U1, Q7Z7I6, Q9BSW0, Q9H6J6, Q9HCD9, Q9NV90, Q9UIW9 | Gene names | CDK5RAP2, CEP215, KIAA1633 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48) (Centrosome-associated protein 215). | |||||
|
RBM28_MOUSE
|
||||||
| NC score | 0.016760 (rank : 105) | θ value | 1.38821 (rank : 50) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8CGC6, Q8BG25, Q8VEJ8, Q9CS22, Q9CSE6 | Gene names | Rbm28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 28 (RNA-binding motif protein 28). | |||||
|
ICAL_MOUSE
|
||||||
| NC score | 0.016644 (rank : 106) | θ value | 6.88961 (rank : 129) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P51125, Q9EQV4, Q9EQV5, Q9QXQ3, Q9QXQ4, Q9R0N1 | Gene names | Cast | |||
|
Domain Architecture |

|
|||||
| Description | Calpastatin (Calpain inhibitor). | |||||
|
GOGA4_HUMAN
|
||||||
| NC score | 0.016503 (rank : 107) | θ value | 6.88961 (rank : 127) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
BRCA2_HUMAN
|
||||||
| NC score | 0.016419 (rank : 108) | θ value | 6.88961 (rank : 119) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P51587, O00183, O15008, Q13879 | Gene names | BRCA2, FANCD1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 2 susceptibility protein (Fanconi anemia group D1 protein). | |||||
|
MPDZ_MOUSE
|
||||||
| NC score | 0.016312 (rank : 109) | θ value | 1.06291 (rank : 41) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8VBX6, O08783, Q6P7U4, Q80ZY8, Q8BKJ1, Q8C0H8, Q8VBV5, Q8VBY0, Q9Z1K3 | Gene names | Mpdz, Mupp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multiple PDZ domain protein (Multi PDZ domain protein 1) (Multi-PDZ domain protein 1). | |||||
|
MYH8_HUMAN
|
||||||
| NC score | 0.016305 (rank : 110) | θ value | 1.38821 (rank : 49) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1610 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P13535, Q14910 | Gene names | MYH8 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
UVRAG_HUMAN
|
||||||
| NC score | 0.016185 (rank : 111) | θ value | 8.99809 (rank : 165) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9P2Y5, O00392 | Gene names | UVRAG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UV radiation resistance-associated gene protein (p63). | |||||
|
EEA1_HUMAN
|
||||||
| NC score | 0.016009 (rank : 112) | θ value | 2.36792 (rank : 64) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1796 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q15075, Q14221 | Gene names | EEA1, ZFYVE2 | |||
|
Domain Architecture |

|
|||||
| Description | Early endosome antigen 1 (Endosome-associated protein p162) (Zinc finger FYVE domain-containing protein 2). | |||||
|
PREP_MOUSE
|
||||||
| NC score | 0.015739 (rank : 113) | θ value | 5.27518 (rank : 112) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K411, Q3THC4, Q3TJI1, Q3TPP0, Q3URQ8, Q4KUG2, Q4KUG3, Q6ZPX6, Q922N1, Q9CV63 | Gene names | Pitrm1, Kiaa1104, Ntup1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Presequence protease, mitochondrial precursor (EC 3.4.24.-) (Pitrilysin metalloproteinase 1). | |||||
|
GCC2_MOUSE
|
||||||
| NC score | 0.015673 (rank : 114) | θ value | 5.27518 (rank : 104) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1619 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8CHG3, Q8BR44, Q8R2Q5, Q9CT45 | Gene names | Gcc2, Kiaa0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185). | |||||
|
MYRIP_HUMAN
|
||||||
| NC score | 0.015669 (rank : 115) | θ value | 5.27518 (rank : 109) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8NFW9, Q8IUF5, Q9Y3V4 | Gene names | MYRIP, SLAC2C | |||
|
Domain Architecture |

|
|||||
| Description | Rab effector MyRIP (Myosin-VIIa- and Rab-interacting protein) (Exophilin-8) (Slp homolog lacking C2 domains c). | |||||
|
SAFB2_MOUSE
|
||||||
| NC score | 0.015515 (rank : 116) | θ value | 6.88961 (rank : 143) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q80YR5, Q8K153 | Gene names | Safb2 | |||
|
Domain Architecture |

|
|||||
| Description | Scaffold attachment factor B2. | |||||
|
CCD41_MOUSE
|
||||||
| NC score | 0.015355 (rank : 117) | θ value | 6.88961 (rank : 121) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9D5R3, Q3U7X7, Q3UX57, Q80VF0 | Gene names | Ccdc41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41. | |||||
|
MY18B_HUMAN
|
||||||
| NC score | 0.014967 (rank : 118) | θ value | 8.99809 (rank : 154) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1299 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8IUG5, Q8NDI8, Q8TE65, Q8WWS0, Q96KH2, Q96KR8, Q96KR9 | Gene names | MYO18B | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18B (Myosin XVIIIb). | |||||
|
SMG1_MOUSE
|
||||||
| NC score | 0.014325 (rank : 119) | θ value | 6.88961 (rank : 144) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BKX6, Q5DU42, Q6ZQC0, Q8BLU4, Q8BWJ5, Q8BXD3 | Gene names | Smg1, Kiaa0421 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SMG1 (EC 2.7.11.1) (SMG-1). | |||||
|
SVIL_MOUSE
|
||||||
| NC score | 0.014264 (rank : 120) | θ value | 6.88961 (rank : 146) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8K4L3 | Gene names | Svil | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Supervillin (Archvillin) (p205/p250). | |||||
|
KI18A_MOUSE
|
||||||
| NC score | 0.014125 (rank : 121) | θ value | 1.81305 (rank : 58) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91WD7, Q3TP77, Q8BLL1 | Gene names | Kif18a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 18A. | |||||
|
KTN1_MOUSE
|
||||||
| NC score | 0.014029 (rank : 122) | θ value | 6.88961 (rank : 132) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1324 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q61595, Q8BG49, Q8BHF4, Q8BHM8, Q8C9Y5, Q8CG51, Q8CG52, Q8CG53, Q8CG54, Q8CG55, Q8CG56, Q8CG57, Q8CG58, Q8CG59, Q8CG60, Q8CG61, Q8CG62, Q8CG63 | Gene names | Ktn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin. | |||||
|
RNF34_HUMAN
|
||||||
| NC score | 0.013893 (rank : 123) | θ value | 4.03905 (rank : 94) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q969K3, Q8NG47, Q9H6W8 | Gene names | RNF34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 34 (RING finger protein RIFF) (FYVE-RING finger protein Momo) (Human RING finger homologous to inhibitor of apoptosis protein) (hRFI) (Caspases-8 and -10-associated RING finger protein 1) (CARP-1) (Caspase regulator CARP1). | |||||
|
ITSN1_HUMAN
|
||||||
| NC score | 0.013870 (rank : 124) | θ value | 5.27518 (rank : 105) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1757 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q15811, O95216, Q9UET5, Q9UK60, Q9UNK1, Q9UNK2, Q9UQ92 | Gene names | ITSN1, ITSN, SH3D1A | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (SH3 domain-containing protein 1A) (SH3P17). | |||||
|
CHD1_HUMAN
|
||||||
| NC score | 0.013349 (rank : 125) | θ value | 2.36792 (rank : 62) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
CENPF_HUMAN
|
||||||
| NC score | 0.013235 (rank : 126) | θ value | 8.99809 (rank : 148) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
K1024_MOUSE
|
||||||
| NC score | 0.013066 (rank : 127) | θ value | 6.88961 (rank : 131) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8K3V7, Q69ZS9 | Gene names | Kiaa1024 | |||
|
Domain Architecture |

|
|||||
| Description | UPF0258 protein KIAA1024 (Protein DD1). | |||||
|
BRWD1_MOUSE
|
||||||
| NC score | 0.012737 (rank : 128) | θ value | 1.81305 (rank : 53) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 478 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q921C3, Q921C2 | Gene names | Brwd1, Wdr9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and WD repeat domain-containing protein 1 (WD repeat protein 9). | |||||
|
KCAB3_HUMAN
|
||||||
| NC score | 0.012465 (rank : 129) | θ value | 4.03905 (rank : 88) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43448 | Gene names | KCNAB3, KCNA3B | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-gated potassium channel subunit beta-3 (K(+) channel subunit beta-3) (Kv-beta-3). | |||||
|
FACD2_MOUSE
|
||||||
| NC score | 0.012424 (rank : 130) | θ value | 6.88961 (rank : 126) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80V62, Q9CWC7 | Gene names | Fancd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group D2 protein homolog (Protein FACD2). | |||||
|
AF17_HUMAN
|
||||||
| NC score | 0.012060 (rank : 131) | θ value | 5.27518 (rank : 100) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P55198, Q9UF49 | Gene names | MLLT6, AF17 | |||
|
Domain Architecture |

|
|||||
| Description | Protein AF-17. | |||||
|
MYH11_MOUSE
|
||||||
| NC score | 0.011924 (rank : 132) | θ value | 6.88961 (rank : 134) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
DGKG_MOUSE
|
||||||
| NC score | 0.011419 (rank : 133) | θ value | 1.81305 (rank : 57) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q91WG7 | Gene names | Dgkg, Dagk3 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase gamma (EC 2.7.1.107) (Diglyceride kinase gamma) (DGK-gamma) (DAG kinase gamma) (88 kDa diacylglycerol kinase). | |||||
|
RP1_MOUSE
|
||||||
| NC score | 0.011354 (rank : 134) | θ value | 6.88961 (rank : 142) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P56716 | Gene names | Rp1, Orp1, Rp1h | |||
|
Domain Architecture |

|
|||||
| Description | Oxygen-regulated protein 1 (Retinitis pigmentosa RP1 protein homolog). | |||||
|
PACN2_HUMAN
|
||||||
| NC score | 0.011034 (rank : 135) | θ value | 5.27518 (rank : 110) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 339 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UNF0, O95921, Q96HV9, Q9H0D3, Q9NPN1, Q9Y4V2 | Gene names | PACSIN2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C and casein kinase substrate in neurons protein 2. | |||||
|
MYO9B_HUMAN
|
||||||
| NC score | 0.010797 (rank : 136) | θ value | 4.03905 (rank : 89) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q13459, O75314, Q9NUJ2, Q9UHN0 | Gene names | MYO9B, MYR5 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9B (Myosin IXb) (Unconventional myosin-9b). | |||||
|
AHI1_HUMAN
|
||||||
| NC score | 0.010677 (rank : 137) | θ value | 3.0926 (rank : 74) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 537 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8N157, Q504T3, Q5TCP9, Q6P098, Q6PIT6, Q8NDX0, Q9H0H2 | Gene names | AHI1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Jouberin (Abelson helper integration site 1 protein homolog) (AHI-1). | |||||
|
K0690_MOUSE
|
||||||
| NC score | 0.010633 (rank : 138) | θ value | 8.99809 (rank : 151) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6P5B0, Q7TMI5, Q80TU2 | Gene names | Kiaa0690 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0690. | |||||
|
IPR1_MOUSE
|
||||||
| NC score | 0.010458 (rank : 139) | θ value | 8.99809 (rank : 150) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BVK9, Q3UCV7, Q80V00 | Gene names | Ipr1, Ifi75 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intracellular pathogen resistance protein 1. | |||||
|
SMYD1_MOUSE
|
||||||
| NC score | 0.010412 (rank : 140) | θ value | 6.88961 (rank : 145) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P97443, P97442, P97444 | Gene names | Smyd1, Bop | |||
|
Domain Architecture |

|
|||||
| Description | SET and MYND domain-containing protein 1 (Zinc-finger protein BOP) (m- BOP) (CD8b-opposite). | |||||
|
CHD4_MOUSE
|
||||||
| NC score | 0.010309 (rank : 141) | θ value | 8.99809 (rank : 149) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6PDQ2 | Gene names | Chd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (CHD-4). | |||||
|
CAC1H_MOUSE
|
||||||
| NC score | 0.010195 (rank : 142) | θ value | 1.81305 (rank : 54) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O88427, Q80TJ2, Q9JKU5 | Gene names | Cacna1h, Kiaa1120 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1H (Voltage- gated calcium channel subunit alpha Cav3.2). | |||||
|
RBM7_HUMAN
|
||||||
| NC score | 0.010135 (rank : 143) | θ value | 6.88961 (rank : 140) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y580 | Gene names | RBM7 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 7 (RNA-binding motif protein 7). | |||||
|
PAR10_HUMAN
|
||||||
| NC score | 0.009400 (rank : 144) | θ value | 8.99809 (rank : 158) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q53GL7, Q8N2I0, Q8WV05, Q96CH7 | Gene names | PARP10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 10 (EC 2.4.2.30) (PARP-10). | |||||
|
MYST4_MOUSE
|
||||||
| NC score | 0.008760 (rank : 145) | θ value | 8.99809 (rank : 155) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8BRB7, Q7TNW5, Q8BG35, Q8C441, Q9JKX5 | Gene names | Myst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Querkopf protein). | |||||
|
GRSF1_MOUSE
|
||||||
| NC score | 0.008559 (rank : 146) | θ value | 6.88961 (rank : 128) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C5Q4, Q8BR05, Q8BRG7 | Gene names | Grsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-rich sequence factor 1 (GRSF-1). | |||||
|
CHST3_HUMAN
|
||||||
| NC score | 0.008002 (rank : 147) | θ value | 6.88961 (rank : 122) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7LGC8, O75099 | Gene names | CHST3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carbohydrate sulfotransferase 3 (EC 2.8.2.17) (Chondroitin 6- sulfotransferase) (Chondroitin 6-O-sulfotransferase 1) (C6ST-1) (C6ST) (Galactose/N-acetylglucosamine/N-acetylglucosamine 6-O- sulfotransferase 0) (GST-0). | |||||
|
RLF_HUMAN
|
||||||
| NC score | 0.007657 (rank : 148) | θ value | 6.88961 (rank : 141) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q13129, Q9NU60 | Gene names | RLF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein Rlf (Rearranged L-myc fusion gene protein) (Zn-15- related protein). | |||||
|
MX1_MOUSE
|
||||||
| NC score | 0.007519 (rank : 149) | θ value | 6.88961 (rank : 133) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P09922 | Gene names | Mx1 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced GTP-binding protein Mx1 (Influenza resistance protein). | |||||
|
CT075_MOUSE
|
||||||
| NC score | 0.007457 (rank : 150) | θ value | 5.27518 (rank : 103) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P59383 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Uncharacterized protein C20orf75 homolog precursor. | |||||
|
LAP4_MOUSE
|
||||||
| NC score | 0.007340 (rank : 151) | θ value | 3.0926 (rank : 78) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 590 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q80U72, Q6P5H7, Q7TPH8, Q80VB1, Q8CI48, Q8VII1, Q922S3 | Gene names | Scrib, Kiaa0147, Lap4, Scrib1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP4 (Protein scribble homolog). | |||||
|
CSTN1_HUMAN
|
||||||
| NC score | 0.006989 (rank : 152) | θ value | 5.27518 (rank : 102) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O94985 | Gene names | CLSTN1, CS1, KIAA0911 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calsyntenin-1 precursor. | |||||
|
NPTN_MOUSE
|
||||||
| NC score | 0.006707 (rank : 153) | θ value | 8.99809 (rank : 157) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P97300, Q3U519, Q8C637, Q8C6H8 | Gene names | Nptn, Sdfr1, Sdr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuroplastin precursor (Stromal cell-derived receptor 1) (SDR-1). | |||||
|
BARX2_HUMAN
|
||||||
| NC score | 0.006556 (rank : 154) | θ value | 1.81305 (rank : 52) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UMQ3, O43518 | Gene names | BARX2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein BarH-like 2. | |||||
|
UBP3_HUMAN
|
||||||
| NC score | 0.006530 (rank : 155) | θ value | 8.99809 (rank : 164) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y6I4 | Gene names | USP3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 3 (EC 3.1.2.15) (Ubiquitin thioesterase 3) (Ubiquitin-specific-processing protease 3) (Deubiquitinating enzyme 3). | |||||
|
SL9A1_MOUSE
|
||||||
| NC score | 0.006244 (rank : 156) | θ value | 8.99809 (rank : 161) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61165 | Gene names | Slc9a1, Nhe1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/hydrogen exchanger 1 (Na(+)/H(+) exchanger 1) (NHE-1) (Solute carrier family 9 member 1). | |||||
|
ANK1_MOUSE
|
||||||
| NC score | 0.006050 (rank : 157) | θ value | 3.0926 (rank : 75) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 430 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q02357, P70440, P97446, P97941, Q3TZ35, Q3UH42, Q3UHP2, Q3UYY7, Q61302, Q61303, Q78E45 | Gene names | Ank1, Ank-1 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-1 (Erythrocyte ankyrin). | |||||
|
NRBP_HUMAN
|
||||||
| NC score | 0.005641 (rank : 158) | θ value | 6.88961 (rank : 136) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UHY1, Q53FZ5, Q96SU3 | Gene names | NRBP1, BCON3, NRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor-binding protein. | |||||
|
COIA1_MOUSE
|
||||||
| NC score | 0.005542 (rank : 159) | θ value | 6.88961 (rank : 124) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 481 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P39061, Q60672, Q61437, Q62001, Q62002, Q9JK63 | Gene names | Col18a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVIII) chain precursor [Contains: Endostatin]. | |||||
|
ANK1_HUMAN
|
||||||
| NC score | 0.005515 (rank : 160) | θ value | 4.03905 (rank : 83) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P16157, O43400, Q13768, Q59FP2, Q8N604, Q99407 | Gene names | ANK1, ANK | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-1 (Erythrocyte ankyrin) (Ankyrin-R). | |||||
|
SRPK2_HUMAN
|
||||||
| NC score | 0.005162 (rank : 161) | θ value | 5.27518 (rank : 114) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 664 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P78362, O75220, O75221, Q6NUL0, Q6V1X2, Q8IYQ3 | Gene names | SRPK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SRPK2 (EC 2.7.11.1) (Serine/arginine- rich protein-specific kinase 2) (SR-protein-specific kinase 2) (SFRS protein kinase 2). | |||||
|
TNIK_HUMAN
|
||||||
| NC score | 0.004911 (rank : 162) | θ value | 3.0926 (rank : 81) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1570 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9UKE5, O60298, Q8WUY7, Q9UKD8, Q9UKD9, Q9UKE0, Q9UKE1, Q9UKE2, Q9UKE3, Q9UKE4 | Gene names | TNIK, KIAA0551 | |||
|
Domain Architecture |

|
|||||
| Description | TRAF2 and NCK-interacting protein kinase (EC 2.7.11.1). | |||||
|
TR4_HUMAN
|
||||||
| NC score | 0.004493 (rank : 163) | θ value | 5.27518 (rank : 115) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P49116, P55092 | Gene names | NR2C2, TAK1, TR4 | |||
|
Domain Architecture |

|
|||||
| Description | Orphan nuclear receptor TR4 (Orphan nuclear receptor TAK1). | |||||
|
IRAK2_HUMAN
|
||||||
| NC score | 0.004280 (rank : 164) | θ value | 6.88961 (rank : 130) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 714 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O43187 | Gene names | IRAK2 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-1 receptor-associated kinase-like 2 (IRAK-2). | |||||
|
TTK_HUMAN
|
||||||
| NC score | 0.003182 (rank : 165) | θ value | 1.81305 (rank : 60) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 816 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P33981 | Gene names | TTK, MPS1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Dual specificity protein kinase TTK (EC 2.7.12.1) (Phosphotyrosine picked threonine-protein kinase) (PYT). | |||||
|
NFASC_HUMAN
|
||||||
| NC score | 0.002108 (rank : 166) | θ value | 6.88961 (rank : 135) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O94856, Q5T2F0, Q5T2F1, Q5T2F2, Q5T2F3, Q5T2F4, Q5T2F5, Q5T2F6, Q5T2F7, Q5T2F9, Q5T2G0, Q5W9F8, Q68DH3, Q6ZQV6, Q7Z3K1, Q96HT1, Q96K50 | Gene names | NFASC, KIAA0756 | |||
|
Domain Architecture |

|
|||||
| Description | Neurofascin precursor. | |||||
|
ROBO1_HUMAN
|
||||||
| NC score | 0.001499 (rank : 167) | θ value | 8.99809 (rank : 160) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 529 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y6N7, Q7Z300, Q9BUS7 | Gene names | ROBO1, DUTT1 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 1 precursor (H-Robo-1) (Deleted in U twenty twenty). | |||||
|
ABL2_HUMAN
|
||||||
| NC score | 0.001281 (rank : 168) | θ value | 1.81305 (rank : 51) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 1045 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P42684, Q5T0X6, Q6NZY6 | Gene names | ABL2, ABLL, ARG | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase ABL2 (EC 2.7.10.2) (Abelson murine leukemia viral oncogene homolog 2) (Tyrosine kinase ARG). | |||||
|
SN1L2_HUMAN
|
||||||
| NC score | 0.000881 (rank : 169) | θ value | 2.36792 (rank : 70) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 964 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9H0K1, O94878, Q6AZE2, Q76N03, Q8NCV7, Q96CZ8 | Gene names | SNF1LK2, KIAA0781, QIK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SNF1-like kinase 2 (EC 2.7.11.1) (Qin- induced kinase). | |||||
|
MTF1_MOUSE
|
||||||
| NC score | -0.001679 (rank : 170) | θ value | 8.99809 (rank : 153) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q07243 | Gene names | Mtf1, Mtf-1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-regulatory transcription factor 1 (Transcription factor MTF-1) (MRE-binding transcription factor). | |||||
|
ZN167_HUMAN
|
||||||
| NC score | -0.001799 (rank : 171) | θ value | 4.03905 (rank : 99) | |||
| Query Neighborhood Hits | 169 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9P0L1, Q6WL09, Q96FQ2 | Gene names | ZNF167, ZNF448, ZNF64 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 167 (Zinc finger protein 64) (Zinc finger protein 448) (ZFP). | |||||