Please be patient as the page loads
|
IPR1_MOUSE
|
||||||
| SwissProt Accessions | Q8BVK9, Q3UCV7, Q80V00 | Gene names | Ipr1, Ifi75 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intracellular pathogen resistance protein 1. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
IPR1_MOUSE
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q8BVK9, Q3UCV7, Q80V00 | Gene names | Ipr1, Ifi75 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intracellular pathogen resistance protein 1. | |||||
|
SP110_HUMAN
|
||||||
| θ value | 2.92676e-75 (rank : 2) | NC score | 0.765024 (rank : 2) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9HB58, Q14976, Q14977, Q8WUZ6, Q9HCT8 | Gene names | SP110 | |||
|
Domain Architecture |

|
|||||
| Description | Sp110 nuclear body protein (Speckled 110 kDa) (Transcriptional coactivator Sp110) (Interferon-induced protein 41/75). | |||||
|
SP100_HUMAN
|
||||||
| θ value | 7.56453e-23 (rank : 3) | NC score | 0.519737 (rank : 6) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P23497, O75450, Q13343, Q96F70, Q96T24, Q96T95, Q9NP33, Q9UE32 | Gene names | SP100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigen Sp-100 (Speckled 100 kDa) (Nuclear dot-associated Sp100 protein) (Lysp100b). | |||||
|
LY10_HUMAN
|
||||||
| θ value | 1.86321e-21 (rank : 4) | NC score | 0.616979 (rank : 5) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q13342, Q13341, Q92881, Q96TG3 | Gene names | SP140, LYSP100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear body protein SP140 (Nuclear autoantigen Sp-140) (Speckled 140 kDa) (LYSp100 protein) (Lymphoid-restricted homolog of Sp100). | |||||
|
SP100_MOUSE
|
||||||
| θ value | 1.13037e-18 (rank : 5) | NC score | 0.706085 (rank : 4) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O35892, O35897, O88392, O88395 | Gene names | Sp100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigen Sp-100 (Speckled 100 kDa) (Nuclear dot-associated Sp100 protein). | |||||
|
S100R_MOUSE
|
||||||
| θ value | 1.92812e-18 (rank : 6) | NC score | 0.725624 (rank : 3) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99388 | Gene names | D1Lub1, Sp100-rs | |||
|
Domain Architecture |
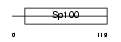
|
|||||
| Description | Putative Sp100-related protein. | |||||
|
DEAF1_HUMAN
|
||||||
| θ value | 1.06045e-08 (rank : 7) | NC score | 0.446233 (rank : 7) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O75398, O15152, O75399, O75510, O75511, O75512, O75513, Q9UET1 | Gene names | DEAF1, SPN, ZMYND5 | |||
|
Domain Architecture |

|
|||||
| Description | Deformed epidermal autoregulatory factor 1 homolog (Nuclear DEAF-1- related transcriptional regulator) (NUDR) (Suppressin) (Zinc finger MYND domain-containing protein 5). | |||||
|
DEAF1_MOUSE
|
||||||
| θ value | 1.06045e-08 (rank : 8) | NC score | 0.442624 (rank : 8) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Z1T5 | Gene names | Deaf1 | |||
|
Domain Architecture |

|
|||||
| Description | Deformed epidermal autoregulatory factor 1 homolog (Nuclear DEAF-1- related transcriptional regulator) (NUDR). | |||||
|
GMEB2_HUMAN
|
||||||
| θ value | 1.29631e-06 (rank : 9) | NC score | 0.382304 (rank : 10) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UKD1, Q5TDS0, Q9H431, Q9H4X7, Q9H4X8, Q9UF78, Q9ULF1 | Gene names | GMEB2, KIAA1269 | |||
|
Domain Architecture |
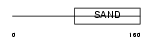
|
|||||
| Description | Glucocorticoid modulatory element-binding protein 2 (GMEB-2) (Parvovirus initiation factor p79) (PIF p79) (DNA-binding protein p79PIF). | |||||
|
GMEB2_MOUSE
|
||||||
| θ value | 1.29631e-06 (rank : 10) | NC score | 0.384662 (rank : 9) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P58929, Q6PCY0 | Gene names | Gmeb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucocorticoid modulatory element-binding protein 2 (GMEB-2). | |||||
|
GMEB1_MOUSE
|
||||||
| θ value | 2.44474e-05 (rank : 11) | NC score | 0.369848 (rank : 11) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9JL60 | Gene names | Gmeb1 | |||
|
Domain Architecture |
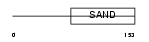
|
|||||
| Description | Glucocorticoid modulatory element-binding protein 1 (GMEB-1). | |||||
|
GMEB1_HUMAN
|
||||||
| θ value | 3.19293e-05 (rank : 12) | NC score | 0.369038 (rank : 12) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y692, Q9NWH1, Q9UKD0 | Gene names | GMEB1 | |||
|
Domain Architecture |
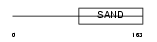
|
|||||
| Description | Glucocorticoid modulatory element-binding protein 1 (GMEB-1) (Parvovirus initiation factor p96) (PIF p96) (DNA-binding protein p96PIF). | |||||
|
PRDM2_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 13) | NC score | 0.028346 (rank : 51) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q13029, Q13149, Q14550, Q5VUL9 | Gene names | PRDM2, RIZ | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 2 (PR domain-containing protein 2) (Retinoblastoma protein-interacting zinc-finger protein) (Zinc finger protein RIZ) (MTE-binding protein) (MTB-ZF) (GATA-3-binding protein G3B). | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 0.125558 (rank : 14) | NC score | 0.042123 (rank : 41) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
AIRE_MOUSE
|
||||||
| θ value | 0.163984 (rank : 15) | NC score | 0.160537 (rank : 13) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Z0E3, Q9JLW0, Q9JLW1, Q9JLW2, Q9JLW3, Q9JLW4, Q9JLW5, Q9JLW6, Q9JLW7, Q9JLW8, Q9JLW9, Q9JLX0 | Gene names | Aire | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein homolog) (APECED protein homolog). | |||||
|
NOL8_HUMAN
|
||||||
| θ value | 0.21417 (rank : 16) | NC score | 0.049434 (rank : 36) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q76FK4, Q5TCC7, Q5TCC8, Q5TCD3, Q5TCD5, Q5TCD6, Q5TCD7, Q76D35, Q7L3E2, Q9H586, Q9H795, Q9H7W7, Q9H9J6, Q9NWA4, Q9NWM4 | Gene names | NOL8, NOP132 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar protein 8 (Nucleolar protein Nop132). | |||||
|
CD2L7_HUMAN
|
||||||
| θ value | 0.279714 (rank : 17) | NC score | 0.011196 (rank : 73) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1275 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NYV4, O94978 | Gene names | CRKRS, CRK7, KIAA0904 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle 2-related protein kinase 7 (EC 2.7.11.22) (CDC2- related protein kinase 7) (Cdc2-related kinase, arginine/serine-rich) (CrkRS). | |||||
|
4ET_MOUSE
|
||||||
| θ value | 0.47712 (rank : 18) | NC score | 0.048870 (rank : 37) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9EST3, Q9CSS3 | Gene names | Eif4enif1, Clast4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4E transporter (eIF4E transporter) (4E-T) (Eukaryotic translation initiation factor 4E nuclear import factor 1). | |||||
|
ELL_MOUSE
|
||||||
| θ value | 0.62314 (rank : 19) | NC score | 0.039553 (rank : 43) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O08856 | Gene names | Ell | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II elongation factor ELL (Eleven-nineteen lysine-rich leukemia protein). | |||||
|
TSH3_MOUSE
|
||||||
| θ value | 0.62314 (rank : 20) | NC score | 0.025916 (rank : 53) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 590 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CGV9, Q5DTX4 | Gene names | Tshz3, Kiaa1474, Tsh3, Zfp537, Znf537 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 3 (Zinc finger protein 537). | |||||
|
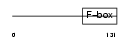
FBX16_MOUSE
|
||||||
| θ value | 0.813845 (rank : 21) | NC score | 0.043669 (rank : 39) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QZM9, Q3TP74, Q8C5V6 | Gene names | Fbxo16, Fbx16 | |||
|
Domain Architecture |
|
|||||
| Description | F-box only protein 16. | |||||
|
MARK2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 22) | NC score | 0.005686 (rank : 82) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1056 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7KZI7, Q15449, Q15524, Q5XGA3, Q68A18, Q96HB3, Q96RG0 | Gene names | MARK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL motif kinase) (EMK1) (PAR1 homolog). | |||||
|
WNK4_MOUSE
|
||||||
| θ value | 0.813845 (rank : 23) | NC score | 0.013100 (rank : 68) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 961 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80UE6, Q4VAC1, Q80XB5, Q80XN2, Q8R0N0, Q8R340 | Gene names | Wnk4, Prkwnk4 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK4 (EC 2.7.11.1) (Protein kinase with no lysine 4) (Protein kinase, lysine-deficient 4). | |||||
|
GGA1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 24) | NC score | 0.025037 (rank : 57) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8R0H9, Q3U2N1 | Gene names | Gga1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADP-ribosylation factor-binding protein GGA1 (Golgi-localized, gamma ear-containing, ARF-binding protein 1) (Gamma-adaptin-related protein 1). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 1.06291 (rank : 25) | NC score | 0.034076 (rank : 47) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
WNK4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 26) | NC score | 0.013047 (rank : 69) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1043 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96J92, Q8N8X3, Q8N8Z2, Q96DT8, Q9BYS5 | Gene names | WNK4, PRKWNK4 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK4 (EC 2.7.11.1) (Protein kinase with no lysine 4) (Protein kinase, lysine-deficient 4). | |||||
|
CKAP2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 27) | NC score | 0.038812 (rank : 44) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3V1H1, Q66LN2, Q8BSF0 | Gene names | Ckap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 2. | |||||
|
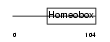
IRX1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 28) | NC score | 0.020487 (rank : 63) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P78414, Q7Z2F8, Q8N312 | Gene names | IRX1, IRXA1 | |||
|
Domain Architecture |
|
|||||
| Description | Iroquois-class homeodomain protein IRX-1 (Iroquois homeobox protein 1) (Homeodomain protein IRXA1). | |||||
|
AIRE_HUMAN
|
||||||
| θ value | 1.81305 (rank : 29) | NC score | 0.146591 (rank : 14) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O43918, O43922, O43932, O75745 | Gene names | AIRE, APECED | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein) (APECED protein). | |||||
|
CHD2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 30) | NC score | 0.014708 (rank : 67) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O14647 | Gene names | CHD2 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 2 (EC 3.6.1.-) (ATP- dependent helicase CHD2) (CHD-2). | |||||
|
NEIL1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 31) | NC score | 0.037852 (rank : 45) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96FI4, Q6ZRA7, Q86XW7, Q9H6C3 | Gene names | NEIL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endonuclease VIII-like 1 (EC 3.2.2.-) (EC 4.2.99.18) (Nei-like 1) (DNA glycosylase/AP lyase Neil1) (DNA-(apurinic or apyrimidinic site) lyase Neil1) (NEH1) (FPG1). | |||||
|
SNIP_HUMAN
|
||||||
| θ value | 1.81305 (rank : 32) | NC score | 0.031935 (rank : 49) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9C0H9, Q8N4W8 | Gene names | P140, KIAA1684 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | p130Cas-associated protein (p140Cap) (SNAP-25-interacting protein) (SNIP). | |||||
|
TARA_HUMAN
|
||||||
| θ value | 1.81305 (rank : 33) | NC score | 0.026322 (rank : 52) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 34) | NC score | 0.011959 (rank : 72) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
CABIN_HUMAN
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.037448 (rank : 46) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y6J0, Q9Y460 | Gene names | CABIN1, KIAA0330 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin-binding protein Cabin 1 (Calcineurin inhibitor) (CAIN). | |||||
|
CO002_HUMAN
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.032220 (rank : 48) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NZP6 | Gene names | C15orf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C15orf2. | |||||
|
JHD2C_MOUSE
|
||||||
| θ value | 2.36792 (rank : 37) | NC score | 0.023786 (rank : 58) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q69ZK6, Q6NV48, Q8BUF5, Q8C4I5, Q8C5Q9 | Gene names | Jmjd1c, Jhdm2c, Kiaa1380 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable JmjC domain-containing histone demethylation protein 2C (EC 1.14.11.-) (Jumonji domain-containing protein 1C). | |||||
|
KI67_HUMAN
|
||||||
| θ value | 2.36792 (rank : 38) | NC score | 0.039972 (rank : 42) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
NCOR2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 39) | NC score | 0.025231 (rank : 55) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
PACS1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 40) | NC score | 0.044620 (rank : 38) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8K212, Q6P5H8 | Gene names | Pacs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphofurin acidic cluster sorting protein 1 (PACS-1). | |||||
|
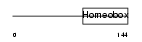
PHX2A_HUMAN
|
||||||
| θ value | 2.36792 (rank : 41) | NC score | 0.008006 (rank : 78) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O14813, Q8IVZ2 | Gene names | PHOX2A, ARIX, PMX2A | |||
|
Domain Architecture |
|
|||||
| Description | Paired mesoderm homeobox protein 2A (Paired-like homeobox 2A) (Aristaless homeobox protein homolog) (ARIX1 homeodomain protein). | |||||
|
YLPM1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 42) | NC score | 0.029512 (rank : 50) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
CCD96_MOUSE
|
||||||
| θ value | 3.0926 (rank : 43) | NC score | 0.016203 (rank : 65) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9CR92 | Gene names | Ccdc96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
PACS1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 44) | NC score | 0.042964 (rank : 40) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6VY07, Q6PJY6, Q6PKB6, Q7Z590, Q7Z5W4, Q8N8K6, Q96MW0, Q9NW92, Q9ULP5 | Gene names | PACS1, KIAA1175 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphofurin acidic cluster sorting protein 1 (PACS-1). | |||||
|
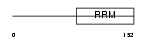
THOC4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 45) | NC score | 0.025897 (rank : 54) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q86V81, O43672 | Gene names | THOC4, ALY, BEF | |||
|
Domain Architecture |
|
|||||
| Description | THO complex subunit 4 (Tho4) (Ally of AML-1 and LEF-1) (Transcriptional coactivator Aly/REF) (bZIP-enhancing factor BEF). | |||||
|
DCC_MOUSE
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.009503 (rank : 75) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
ELK4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 47) | NC score | 0.012396 (rank : 70) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P41158 | Gene names | Elk4, Sap1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
CO039_HUMAN
|
||||||
| θ value | 5.27518 (rank : 48) | NC score | 0.023007 (rank : 60) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6ZRI6, Q71JB1, Q7L3S0, Q8N3F2, Q96FB6, Q9NTU5 | Gene names | C15orf39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf39. | |||||
|
PEG3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | 0.002936 (rank : 85) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
PTN12_HUMAN
|
||||||
| θ value | 5.27518 (rank : 50) | NC score | 0.006661 (rank : 80) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q05209, Q16130, Q59FD6, Q75MN8, Q86XU4 | Gene names | PTPN12 | |||
|
Domain Architecture |
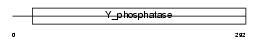
|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 12 (EC 3.1.3.48) (Protein-tyrosine phosphatase G1) (PTPG1). | |||||
|
REFP2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 51) | NC score | 0.025100 (rank : 56) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JJW6, Q8C869 | Gene names | Refbp2, Ref2 | |||
|
Domain Architecture |
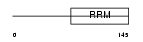
|
|||||
| Description | RNA and export factor-binding protein 2. | |||||
|
SVIL_MOUSE
|
||||||
| θ value | 5.27518 (rank : 52) | NC score | 0.017453 (rank : 64) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K4L3 | Gene names | Svil | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Supervillin (Archvillin) (p205/p250). | |||||
|
DLX3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 53) | NC score | 0.005114 (rank : 83) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 369 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60479 | Gene names | DLX3 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein DLX-3. | |||||
|
K1949_HUMAN
|
||||||
| θ value | 6.88961 (rank : 54) | NC score | 0.020543 (rank : 62) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6NYC8, Q68CK8, Q6ZUJ6, Q8NDQ4, Q8TF52, Q9BRL9 | Gene names | KIAA1949 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1949. | |||||
|
MINT_MOUSE
|
||||||
| θ value | 6.88961 (rank : 55) | NC score | 0.023266 (rank : 59) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
MYOM1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 56) | NC score | 0.008710 (rank : 76) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P52179 | Gene names | MYOM1 | |||
|
Domain Architecture |

|
|||||
| Description | Myomesin-1 (190 kDa titin-associated protein) (190 kDa connectin- associated protein). | |||||
|
S12A5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 57) | NC score | 0.007671 (rank : 79) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H2X9, Q5VZ41, Q9H4Z0, Q9ULP4 | Gene names | SLC12A5, KCC2, KIAA1176 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 12 member 5 (Electroneutral potassium-chloride cotransporter 2) (Erythroid K-Cl cotransporter 2) (Neuronal K-Cl cotransporter) (hKCC2). | |||||
|
U2AFM_HUMAN
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | 0.011994 (rank : 71) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q15696 | Gene names | ZRSR2, U2AF1-RS2, U2AF1L2, U2AF1RS2 | |||
|
Domain Architecture |
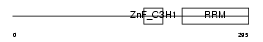
|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 2 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 2) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 2) (NY-REN-20 antigen). | |||||
|
ATAD2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 59) | NC score | 0.006537 (rank : 81) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 322 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6PL18, Q658P2, Q68CQ0, Q6PJV6, Q8N890, Q9UHS5 | Gene names | ATAD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATPase family AAA domain-containing protein 2. | |||||
|
ATF5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 60) | NC score | 0.015521 (rank : 66) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y2D1, Q9BSA1, Q9UNQ3 | Gene names | ATF5, ATFX | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-5 (Activating transcription factor 5) (Transcription factor ATFx). | |||||
|
DPOLZ_MOUSE
|
||||||
| θ value | 8.99809 (rank : 61) | NC score | 0.010458 (rank : 74) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61493, Q9JMD6, Q9QWX6 | Gene names | Rev3l, Polz, Sez4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (Seizure-related protein 4). | |||||
|
RBM19_HUMAN
|
||||||
| θ value | 8.99809 (rank : 62) | NC score | 0.008587 (rank : 77) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y4C8, Q9BPY6, Q9UFN5 | Gene names | RBM19, KIAA0682 | |||
|
Domain Architecture |

|
|||||
| Description | Probable RNA-binding protein 19 (RNA-binding motif protein 19). | |||||
|
TTF1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 63) | NC score | 0.021847 (rank : 61) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q62187, Q9JKK5 | Gene names | Ttf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 1 (TTF-1) (TTF-I) (mTFF-I) (RNA polymerase I termination factor). | |||||
|
ZN687_HUMAN
|
||||||
| θ value | 8.99809 (rank : 64) | NC score | 0.003249 (rank : 84) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8N1G0, Q68DQ8, Q9H937, Q9P2A7 | Gene names | ZNF687, KIAA1441 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 687. | |||||
|
ZN690_HUMAN
|
||||||
| θ value | 8.99809 (rank : 65) | NC score | -0.000627 (rank : 86) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IWY8, Q32M75, Q32M76, Q8NA40 | Gene names | ZNF690 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 690. | |||||
|
HMG1X_HUMAN
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.071490 (rank : 17) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | Q9UGV6 | Gene names | HMG1L10 | |||
|
Domain Architecture |
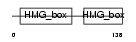
|
|||||
| Description | High mobility group protein 1-like 10 (HMG-1L10). | |||||
|
HMGB1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 67) | NC score | 0.068026 (rank : 21) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | P09429, Q6IBE1 | Gene names | HMGB1, HMG1 | |||
|
Domain Architecture |
|
|||||
| Description | High mobility group protein B1 (High mobility group protein 1) (HMG- 1). | |||||
|
HMGB1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 68) | NC score | 0.068038 (rank : 20) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | P63158, P07155, P27109, P27428 | Gene names | Hmgb1, Hmg-1, Hmg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein B1 (High mobility group protein 1) (HMG- 1). | |||||
|
HMGB2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 69) | NC score | 0.064092 (rank : 26) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P26583 | Gene names | HMGB2, HMG2 | |||
|
Domain Architecture |
|
|||||
| Description | High mobility group protein B2 (High mobility group protein 2) (HMG- 2). | |||||
|
HMGB2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.064563 (rank : 25) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P30681, Q3UXT1, Q9EQD5 | Gene names | Hmgb2, Hmg2 | |||
|
Domain Architecture |
|
|||||
| Description | High mobility group protein B2 (High mobility group protein 2) (HMG- 2). | |||||
|
HMGB3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.055948 (rank : 32) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | O15347, O95556, Q6NS40 | Gene names | HMGB3, HMG2A, HMG4 | |||
|
Domain Architecture |
|
|||||
| Description | High mobility group protein B3 (High mobility group protein 4) (HMG-4) (High mobility group protein 2a) (HMG-2a). | |||||
|
HMGB3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.056210 (rank : 31) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | O54879 | Gene names | Hmgb3, Hmg2a, Hmg4 | |||
|
Domain Architecture |
|
|||||
| Description | High mobility group protein B3 (High mobility group protein 4) (HMG-4) (High mobility group protein 2a) (HMG-2a). | |||||
|
PF21A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 73) | NC score | 0.056936 (rank : 29) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96BD5, Q6AWA2, Q9C0G7, Q9H8V9, Q9HAK6, Q9NZE9 | Gene names | PHF21A, BHC80, KIAA1696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (BHC80a). | |||||
|
PF21A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 74) | NC score | 0.056394 (rank : 30) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6ZPK0, Q6XVG0, Q80Z33, Q8CAZ4, Q8VEC8 | Gene names | Phf21a, Bhc80, Kiaa1696, Pftf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (mBHC80) (BHC80a). | |||||
|
PF21B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 75) | NC score | 0.050419 (rank : 35) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96EK2, Q5TFL2, Q6ICC0 | Gene names | PHF21B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
PHF12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 76) | NC score | 0.065286 (rank : 24) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96QT6, Q6ZML2, Q9BV34, Q9H7U9, Q9P205 | Gene names | PHF12, KIAA1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
PHF12_MOUSE
|
||||||
| θ value | θ > 10 (rank : 77) | NC score | 0.065772 (rank : 23) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5SPL2, Q5SPL3, Q6ZPN8, Q80W50 | Gene names | Phf12, Kiaa1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
PHF22_MOUSE
|
||||||
| θ value | θ > 10 (rank : 78) | NC score | 0.053527 (rank : 34) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D168, Q3U2Q5, Q921U2 | Gene names | Ints12, Phf22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 12 (PHD finger protein 22). | |||||
|
PKCB1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 79) | NC score | 0.072201 (rank : 16) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
SFR11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 80) | NC score | 0.055153 (rank : 33) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
TIF1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.066921 (rank : 22) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O15164, O95854 | Gene names | TRIM24, RNF82, TIF1, TIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24) (RING finger protein 82). | |||||
|
TIF1A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.070900 (rank : 18) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q64127, Q64126 | Gene names | Trim24, Tif1, Tif1a | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24). | |||||
|
TIF1G_HUMAN
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.058780 (rank : 28) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 410 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UPN9, O95855, Q9C017, Q9UJ79 | Gene names | TRIM33, KIAA1113, RFG7, TIF1G | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33) (RET-fused gene 7 protein) (Rfg7 protein). | |||||
|
TIF1G_MOUSE
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.060118 (rank : 27) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99PP7, Q6SI71, Q6ZPX5 | Gene names | Trim33, Kiaa1113 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33). | |||||
|
TRI66_HUMAN
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.075010 (rank : 15) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O15016, Q9BQQ4 | Gene names | TRIM66, KIAA0298 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
TRI66_MOUSE
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.069708 (rank : 19) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q924W6 | Gene names | Trim66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
IPR1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q8BVK9, Q3UCV7, Q80V00 | Gene names | Ipr1, Ifi75 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intracellular pathogen resistance protein 1. | |||||
|
SP110_HUMAN
|
||||||
| NC score | 0.765024 (rank : 2) | θ value | 2.92676e-75 (rank : 2) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9HB58, Q14976, Q14977, Q8WUZ6, Q9HCT8 | Gene names | SP110 | |||
|
Domain Architecture |

|
|||||
| Description | Sp110 nuclear body protein (Speckled 110 kDa) (Transcriptional coactivator Sp110) (Interferon-induced protein 41/75). | |||||
|
S100R_MOUSE
|
||||||
| NC score | 0.725624 (rank : 3) | θ value | 1.92812e-18 (rank : 6) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99388 | Gene names | D1Lub1, Sp100-rs | |||
|
Domain Architecture |

|
|||||
| Description | Putative Sp100-related protein. | |||||
|
SP100_MOUSE
|
||||||
| NC score | 0.706085 (rank : 4) | θ value | 1.13037e-18 (rank : 5) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O35892, O35897, O88392, O88395 | Gene names | Sp100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigen Sp-100 (Speckled 100 kDa) (Nuclear dot-associated Sp100 protein). | |||||
|
LY10_HUMAN
|
||||||
| NC score | 0.616979 (rank : 5) | θ value | 1.86321e-21 (rank : 4) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q13342, Q13341, Q92881, Q96TG3 | Gene names | SP140, LYSP100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear body protein SP140 (Nuclear autoantigen Sp-140) (Speckled 140 kDa) (LYSp100 protein) (Lymphoid-restricted homolog of Sp100). | |||||
|
SP100_HUMAN
|
||||||
| NC score | 0.519737 (rank : 6) | θ value | 7.56453e-23 (rank : 3) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P23497, O75450, Q13343, Q96F70, Q96T24, Q96T95, Q9NP33, Q9UE32 | Gene names | SP100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigen Sp-100 (Speckled 100 kDa) (Nuclear dot-associated Sp100 protein) (Lysp100b). | |||||
|
DEAF1_HUMAN
|
||||||
| NC score | 0.446233 (rank : 7) | θ value | 1.06045e-08 (rank : 7) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O75398, O15152, O75399, O75510, O75511, O75512, O75513, Q9UET1 | Gene names | DEAF1, SPN, ZMYND5 | |||
|
Domain Architecture |

|
|||||
| Description | Deformed epidermal autoregulatory factor 1 homolog (Nuclear DEAF-1- related transcriptional regulator) (NUDR) (Suppressin) (Zinc finger MYND domain-containing protein 5). | |||||
|
DEAF1_MOUSE
|
||||||
| NC score | 0.442624 (rank : 8) | θ value | 1.06045e-08 (rank : 8) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Z1T5 | Gene names | Deaf1 | |||
|
Domain Architecture |

|
|||||
| Description | Deformed epidermal autoregulatory factor 1 homolog (Nuclear DEAF-1- related transcriptional regulator) (NUDR). | |||||
|
GMEB2_MOUSE
|
||||||
| NC score | 0.384662 (rank : 9) | θ value | 1.29631e-06 (rank : 10) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P58929, Q6PCY0 | Gene names | Gmeb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucocorticoid modulatory element-binding protein 2 (GMEB-2). | |||||
|
GMEB2_HUMAN
|
||||||
| NC score | 0.382304 (rank : 10) | θ value | 1.29631e-06 (rank : 9) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UKD1, Q5TDS0, Q9H431, Q9H4X7, Q9H4X8, Q9UF78, Q9ULF1 | Gene names | GMEB2, KIAA1269 | |||
|
Domain Architecture |

|
|||||
| Description | Glucocorticoid modulatory element-binding protein 2 (GMEB-2) (Parvovirus initiation factor p79) (PIF p79) (DNA-binding protein p79PIF). | |||||
|
GMEB1_MOUSE
|
||||||
| NC score | 0.369848 (rank : 11) | θ value | 2.44474e-05 (rank : 11) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9JL60 | Gene names | Gmeb1 | |||
|
Domain Architecture |

|
|||||
| Description | Glucocorticoid modulatory element-binding protein 1 (GMEB-1). | |||||
|
GMEB1_HUMAN
|
||||||
| NC score | 0.369038 (rank : 12) | θ value | 3.19293e-05 (rank : 12) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y692, Q9NWH1, Q9UKD0 | Gene names | GMEB1 | |||
|
Domain Architecture |

|
|||||
| Description | Glucocorticoid modulatory element-binding protein 1 (GMEB-1) (Parvovirus initiation factor p96) (PIF p96) (DNA-binding protein p96PIF). | |||||
|
AIRE_MOUSE
|
||||||
| NC score | 0.160537 (rank : 13) | θ value | 0.163984 (rank : 15) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Z0E3, Q9JLW0, Q9JLW1, Q9JLW2, Q9JLW3, Q9JLW4, Q9JLW5, Q9JLW6, Q9JLW7, Q9JLW8, Q9JLW9, Q9JLX0 | Gene names | Aire | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein homolog) (APECED protein homolog). | |||||
|
AIRE_HUMAN
|
||||||
| NC score | 0.146591 (rank : 14) | θ value | 1.81305 (rank : 29) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O43918, O43922, O43932, O75745 | Gene names | AIRE, APECED | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein) (APECED protein). | |||||
|
TRI66_HUMAN
|
||||||
| NC score | 0.075010 (rank : 15) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O15016, Q9BQQ4 | Gene names | TRIM66, KIAA0298 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
PKCB1_HUMAN
|
||||||
| NC score | 0.072201 (rank : 16) | θ value | θ > 10 (rank : 79) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
HMG1X_HUMAN
|
||||||
| NC score | 0.071490 (rank : 17) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | Q9UGV6 | Gene names | HMG1L10 | |||
|
Domain Architecture |

|
|||||
| Description | High mobility group protein 1-like 10 (HMG-1L10). | |||||
|
TIF1A_MOUSE
|
||||||
| NC score | 0.070900 (rank : 18) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q64127, Q64126 | Gene names | Trim24, Tif1, Tif1a | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24). | |||||
|
TRI66_MOUSE
|
||||||
| NC score | 0.069708 (rank : 19) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q924W6 | Gene names | Trim66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
HMGB1_MOUSE
|
||||||
| NC score | 0.068038 (rank : 20) | θ value | θ > 10 (rank : 68) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | P63158, P07155, P27109, P27428 | Gene names | Hmgb1, Hmg-1, Hmg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein B1 (High mobility group protein 1) (HMG- 1). | |||||
|
HMGB1_HUMAN
|
||||||
| NC score | 0.068026 (rank : 21) | θ value | θ > 10 (rank : 67) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | P09429, Q6IBE1 | Gene names | HMGB1, HMG1 | |||
|
Domain Architecture |

|
|||||
| Description | High mobility group protein B1 (High mobility group protein 1) (HMG- 1). | |||||
|
TIF1A_HUMAN
|
||||||
| NC score | 0.066921 (rank : 22) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O15164, O95854 | Gene names | TRIM24, RNF82, TIF1, TIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24) (RING finger protein 82). | |||||
|
PHF12_MOUSE
|
||||||
| NC score | 0.065772 (rank : 23) | θ value | θ > 10 (rank : 77) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5SPL2, Q5SPL3, Q6ZPN8, Q80W50 | Gene names | Phf12, Kiaa1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
PHF12_HUMAN
|
||||||
| NC score | 0.065286 (rank : 24) | θ value | θ > 10 (rank : 76) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96QT6, Q6ZML2, Q9BV34, Q9H7U9, Q9P205 | Gene names | PHF12, KIAA1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
HMGB2_MOUSE
|
||||||
| NC score | 0.064563 (rank : 25) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P30681, Q3UXT1, Q9EQD5 | Gene names | Hmgb2, Hmg2 | |||
|
Domain Architecture |

|
|||||
| Description | High mobility group protein B2 (High mobility group protein 2) (HMG- 2). | |||||
|
HMGB2_HUMAN
|
||||||
| NC score | 0.064092 (rank : 26) | θ value | θ > 10 (rank : 69) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P26583 | Gene names | HMGB2, HMG2 | |||
|
Domain Architecture |

|
|||||
| Description | High mobility group protein B2 (High mobility group protein 2) (HMG- 2). | |||||
|
TIF1G_MOUSE
|
||||||
| NC score | 0.060118 (rank : 27) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99PP7, Q6SI71, Q6ZPX5 | Gene names | Trim33, Kiaa1113 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33). | |||||
|
TIF1G_HUMAN
|
||||||
| NC score | 0.058780 (rank : 28) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 410 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UPN9, O95855, Q9C017, Q9UJ79 | Gene names | TRIM33, KIAA1113, RFG7, TIF1G | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33) (RET-fused gene 7 protein) (Rfg7 protein). | |||||
|
PF21A_HUMAN
|
||||||
| NC score | 0.056936 (rank : 29) | θ value | θ > 10 (rank : 73) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96BD5, Q6AWA2, Q9C0G7, Q9H8V9, Q9HAK6, Q9NZE9 | Gene names | PHF21A, BHC80, KIAA1696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (BHC80a). | |||||
|
PF21A_MOUSE
|
||||||
| NC score | 0.056394 (rank : 30) | θ value | θ > 10 (rank : 74) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6ZPK0, Q6XVG0, Q80Z33, Q8CAZ4, Q8VEC8 | Gene names | Phf21a, Bhc80, Kiaa1696, Pftf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (mBHC80) (BHC80a). | |||||
|
HMGB3_MOUSE
|
||||||
| NC score | 0.056210 (rank : 31) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | O54879 | Gene names | Hmgb3, Hmg2a, Hmg4 | |||
|
Domain Architecture |

|
|||||
| Description | High mobility group protein B3 (High mobility group protein 4) (HMG-4) (High mobility group protein 2a) (HMG-2a). | |||||
|
HMGB3_HUMAN
|
||||||
| NC score | 0.055948 (rank : 32) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | O15347, O95556, Q6NS40 | Gene names | HMGB3, HMG2A, HMG4 | |||
|
Domain Architecture |

|
|||||
| Description | High mobility group protein B3 (High mobility group protein 4) (HMG-4) (High mobility group protein 2a) (HMG-2a). | |||||
|
SFR11_HUMAN
|
||||||
| NC score | 0.055153 (rank : 33) | θ value | θ > 10 (rank : 80) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
PHF22_MOUSE
|
||||||
| NC score | 0.053527 (rank : 34) | θ value | θ > 10 (rank : 78) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D168, Q3U2Q5, Q921U2 | Gene names | Ints12, Phf22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 12 (PHD finger protein 22). | |||||
|
PF21B_HUMAN
|
||||||
| NC score | 0.050419 (rank : 35) | θ value | θ > 10 (rank : 75) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96EK2, Q5TFL2, Q6ICC0 | Gene names | PHF21B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
NOL8_HUMAN
|
||||||
| NC score | 0.049434 (rank : 36) | θ value | 0.21417 (rank : 16) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q76FK4, Q5TCC7, Q5TCC8, Q5TCD3, Q5TCD5, Q5TCD6, Q5TCD7, Q76D35, Q7L3E2, Q9H586, Q9H795, Q9H7W7, Q9H9J6, Q9NWA4, Q9NWM4 | Gene names | NOL8, NOP132 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar protein 8 (Nucleolar protein Nop132). | |||||
|
4ET_MOUSE
|
||||||
| NC score | 0.048870 (rank : 37) | θ value | 0.47712 (rank : 18) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9EST3, Q9CSS3 | Gene names | Eif4enif1, Clast4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4E transporter (eIF4E transporter) (4E-T) (Eukaryotic translation initiation factor 4E nuclear import factor 1). | |||||
|
PACS1_MOUSE
|
||||||
| NC score | 0.044620 (rank : 38) | θ value | 2.36792 (rank : 40) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8K212, Q6P5H8 | Gene names | Pacs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphofurin acidic cluster sorting protein 1 (PACS-1). | |||||
|
FBX16_MOUSE
|
||||||
| NC score | 0.043669 (rank : 39) | θ value | 0.813845 (rank : 21) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QZM9, Q3TP74, Q8C5V6 | Gene names | Fbxo16, Fbx16 | |||
|
Domain Architecture |

|
|||||
| Description | F-box only protein 16. | |||||
|
PACS1_HUMAN
|
||||||
| NC score | 0.042964 (rank : 40) | θ value | 3.0926 (rank : 44) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6VY07, Q6PJY6, Q6PKB6, Q7Z590, Q7Z5W4, Q8N8K6, Q96MW0, Q9NW92, Q9ULP5 | Gene names | PACS1, KIAA1175 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphofurin acidic cluster sorting protein 1 (PACS-1). | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.042123 (rank : 41) | θ value | 0.125558 (rank : 14) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
KI67_HUMAN
|
||||||
| NC score | 0.039972 (rank : 42) | θ value | 2.36792 (rank : 38) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
ELL_MOUSE
|
||||||
| NC score | 0.039553 (rank : 43) | θ value | 0.62314 (rank : 19) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O08856 | Gene names | Ell | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II elongation factor ELL (Eleven-nineteen lysine-rich leukemia protein). | |||||
|
CKAP2_MOUSE
|
||||||
| NC score | 0.038812 (rank : 44) | θ value | 1.38821 (rank : 27) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3V1H1, Q66LN2, Q8BSF0 | Gene names | Ckap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 2. | |||||
|
NEIL1_HUMAN
|
||||||
| NC score | 0.037852 (rank : 45) | θ value | 1.81305 (rank : 31) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96FI4, Q6ZRA7, Q86XW7, Q9H6C3 | Gene names | NEIL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endonuclease VIII-like 1 (EC 3.2.2.-) (EC 4.2.99.18) (Nei-like 1) (DNA glycosylase/AP lyase Neil1) (DNA-(apurinic or apyrimidinic site) lyase Neil1) (NEH1) (FPG1). | |||||
|
CABIN_HUMAN
|
||||||
| NC score | 0.037448 (rank : 46) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y6J0, Q9Y460 | Gene names | CABIN1, KIAA0330 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin-binding protein Cabin 1 (Calcineurin inhibitor) (CAIN). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.034076 (rank : 47) | θ value | 1.06291 (rank : 25) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
CO002_HUMAN
|
||||||
| NC score | 0.032220 (rank : 48) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NZP6 | Gene names | C15orf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C15orf2. | |||||
|
SNIP_HUMAN
|
||||||
| NC score | 0.031935 (rank : 49) | θ value | 1.81305 (rank : 32) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9C0H9, Q8N4W8 | Gene names | P140, KIAA1684 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | p130Cas-associated protein (p140Cap) (SNAP-25-interacting protein) (SNIP). | |||||
|
YLPM1_MOUSE
|
||||||
| NC score | 0.029512 (rank : 50) | θ value | 2.36792 (rank : 42) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
PRDM2_HUMAN
|
||||||
| NC score | 0.028346 (rank : 51) | θ value | 0.0193708 (rank : 13) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q13029, Q13149, Q14550, Q5VUL9 | Gene names | PRDM2, RIZ | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 2 (PR domain-containing protein 2) (Retinoblastoma protein-interacting zinc-finger protein) (Zinc finger protein RIZ) (MTE-binding protein) (MTB-ZF) (GATA-3-binding protein G3B). | |||||
|
TARA_HUMAN
|
||||||
| NC score | 0.026322 (rank : 52) | θ value | 1.81305 (rank : 33) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
TSH3_MOUSE
|
||||||
| NC score | 0.025916 (rank : 53) | θ value | 0.62314 (rank : 20) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 590 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CGV9, Q5DTX4 | Gene names | Tshz3, Kiaa1474, Tsh3, Zfp537, Znf537 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 3 (Zinc finger protein 537). | |||||
|
THOC4_HUMAN
|
||||||
| NC score | 0.025897 (rank : 54) | θ value | 3.0926 (rank : 45) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q86V81, O43672 | Gene names | THOC4, ALY, BEF | |||
|
Domain Architecture |

|
|||||
| Description | THO complex subunit 4 (Tho4) (Ally of AML-1 and LEF-1) (Transcriptional coactivator Aly/REF) (bZIP-enhancing factor BEF). | |||||
|
NCOR2_MOUSE
|
||||||
| NC score | 0.025231 (rank : 55) | θ value | 2.36792 (rank : 39) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
REFP2_MOUSE
|
||||||
| NC score | 0.025100 (rank : 56) | θ value | 5.27518 (rank : 51) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JJW6, Q8C869 | Gene names | Refbp2, Ref2 | |||
|
Domain Architecture |

|
|||||
| Description | RNA and export factor-binding protein 2. | |||||
|
GGA1_MOUSE
|
||||||
| NC score | 0.025037 (rank : 57) | θ value | 1.06291 (rank : 24) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8R0H9, Q3U2N1 | Gene names | Gga1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADP-ribosylation factor-binding protein GGA1 (Golgi-localized, gamma ear-containing, ARF-binding protein 1) (Gamma-adaptin-related protein 1). | |||||
|
JHD2C_MOUSE
|
||||||
| NC score | 0.023786 (rank : 58) | θ value | 2.36792 (rank : 37) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q69ZK6, Q6NV48, Q8BUF5, Q8C4I5, Q8C5Q9 | Gene names | Jmjd1c, Jhdm2c, Kiaa1380 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable JmjC domain-containing histone demethylation protein 2C (EC 1.14.11.-) (Jumonji domain-containing protein 1C). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.023266 (rank : 59) | θ value | 6.88961 (rank : 55) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
CO039_HUMAN
|
||||||
| NC score | 0.023007 (rank : 60) | θ value | 5.27518 (rank : 48) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6ZRI6, Q71JB1, Q7L3S0, Q8N3F2, Q96FB6, Q9NTU5 | Gene names | C15orf39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf39. | |||||
|
TTF1_MOUSE
|
||||||
| NC score | 0.021847 (rank : 61) | θ value | 8.99809 (rank : 63) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q62187, Q9JKK5 | Gene names | Ttf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 1 (TTF-1) (TTF-I) (mTFF-I) (RNA polymerase I termination factor). | |||||
|
K1949_HUMAN
|
||||||
| NC score | 0.020543 (rank : 62) | θ value | 6.88961 (rank : 54) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6NYC8, Q68CK8, Q6ZUJ6, Q8NDQ4, Q8TF52, Q9BRL9 | Gene names | KIAA1949 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1949. | |||||
|
IRX1_HUMAN
|
||||||
| NC score | 0.020487 (rank : 63) | θ value | 1.38821 (rank : 28) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P78414, Q7Z2F8, Q8N312 | Gene names | IRX1, IRXA1 | |||
|
Domain Architecture |

|
|||||
| Description | Iroquois-class homeodomain protein IRX-1 (Iroquois homeobox protein 1) (Homeodomain protein IRXA1). | |||||
|
SVIL_MOUSE
|
||||||
| NC score | 0.017453 (rank : 64) | θ value | 5.27518 (rank : 52) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K4L3 | Gene names | Svil | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Supervillin (Archvillin) (p205/p250). | |||||
|
CCD96_MOUSE
|
||||||
| NC score | 0.016203 (rank : 65) | θ value | 3.0926 (rank : 43) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9CR92 | Gene names | Ccdc96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
ATF5_HUMAN
|
||||||
| NC score | 0.015521 (rank : 66) | θ value | 8.99809 (rank : 60) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y2D1, Q9BSA1, Q9UNQ3 | Gene names | ATF5, ATFX | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-5 (Activating transcription factor 5) (Transcription factor ATFx). | |||||
|
CHD2_HUMAN
|
||||||
| NC score | 0.014708 (rank : 67) | θ value | 1.81305 (rank : 30) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O14647 | Gene names | CHD2 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 2 (EC 3.6.1.-) (ATP- dependent helicase CHD2) (CHD-2). | |||||
|
WNK4_MOUSE
|
||||||
| NC score | 0.013100 (rank : 68) | θ value | 0.813845 (rank : 23) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 961 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80UE6, Q4VAC1, Q80XB5, Q80XN2, Q8R0N0, Q8R340 | Gene names | Wnk4, Prkwnk4 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK4 (EC 2.7.11.1) (Protein kinase with no lysine 4) (Protein kinase, lysine-deficient 4). | |||||
|
WNK4_HUMAN
|
||||||
| NC score | 0.013047 (rank : 69) | θ value | 1.06291 (rank : 26) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1043 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96J92, Q8N8X3, Q8N8Z2, Q96DT8, Q9BYS5 | Gene names | WNK4, PRKWNK4 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK4 (EC 2.7.11.1) (Protein kinase with no lysine 4) (Protein kinase, lysine-deficient 4). | |||||
|
ELK4_MOUSE
|
||||||
| NC score | 0.012396 (rank : 70) | θ value | 4.03905 (rank : 47) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P41158 | Gene names | Elk4, Sap1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
U2AFM_HUMAN
|
||||||
| NC score | 0.011994 (rank : 71) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q15696 | Gene names | ZRSR2, U2AF1-RS2, U2AF1L2, U2AF1RS2 | |||
|
Domain Architecture |

|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 2 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 2) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 2) (NY-REN-20 antigen). | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.011959 (rank : 72) | θ value | 2.36792 (rank : 34) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
CD2L7_HUMAN
|
||||||
| NC score | 0.011196 (rank : 73) | θ value | 0.279714 (rank : 17) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1275 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NYV4, O94978 | Gene names | CRKRS, CRK7, KIAA0904 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle 2-related protein kinase 7 (EC 2.7.11.22) (CDC2- related protein kinase 7) (Cdc2-related kinase, arginine/serine-rich) (CrkRS). | |||||
|
DPOLZ_MOUSE
|
||||||
| NC score | 0.010458 (rank : 74) | θ value | 8.99809 (rank : 61) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61493, Q9JMD6, Q9QWX6 | Gene names | Rev3l, Polz, Sez4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (Seizure-related protein 4). | |||||
|
DCC_MOUSE
|
||||||
| NC score | 0.009503 (rank : 75) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
MYOM1_HUMAN
|
||||||
| NC score | 0.008710 (rank : 76) | θ value | 6.88961 (rank : 56) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P52179 | Gene names | MYOM1 | |||
|
Domain Architecture |

|
|||||
| Description | Myomesin-1 (190 kDa titin-associated protein) (190 kDa connectin- associated protein). | |||||
|
RBM19_HUMAN
|
||||||
| NC score | 0.008587 (rank : 77) | θ value | 8.99809 (rank : 62) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y4C8, Q9BPY6, Q9UFN5 | Gene names | RBM19, KIAA0682 | |||
|
Domain Architecture |

|
|||||
| Description | Probable RNA-binding protein 19 (RNA-binding motif protein 19). | |||||
|
PHX2A_HUMAN
|
||||||
| NC score | 0.008006 (rank : 78) | θ value | 2.36792 (rank : 41) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O14813, Q8IVZ2 | Gene names | PHOX2A, ARIX, PMX2A | |||
|
Domain Architecture |

|
|||||
| Description | Paired mesoderm homeobox protein 2A (Paired-like homeobox 2A) (Aristaless homeobox protein homolog) (ARIX1 homeodomain protein). | |||||
|
S12A5_HUMAN
|
||||||
| NC score | 0.007671 (rank : 79) | θ value | 6.88961 (rank : 57) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H2X9, Q5VZ41, Q9H4Z0, Q9ULP4 | Gene names | SLC12A5, KCC2, KIAA1176 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 12 member 5 (Electroneutral potassium-chloride cotransporter 2) (Erythroid K-Cl cotransporter 2) (Neuronal K-Cl cotransporter) (hKCC2). | |||||
|
PTN12_HUMAN
|
||||||
| NC score | 0.006661 (rank : 80) | θ value | 5.27518 (rank : 50) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q05209, Q16130, Q59FD6, Q75MN8, Q86XU4 | Gene names | PTPN12 | |||
|
Domain Architecture |
|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 12 (EC 3.1.3.48) (Protein-tyrosine phosphatase G1) (PTPG1). | |||||
|
ATAD2_HUMAN
|
||||||
| NC score | 0.006537 (rank : 81) | θ value | 8.99809 (rank : 59) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 322 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6PL18, Q658P2, Q68CQ0, Q6PJV6, Q8N890, Q9UHS5 | Gene names | ATAD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATPase family AAA domain-containing protein 2. | |||||
|
MARK2_HUMAN
|
||||||
| NC score | 0.005686 (rank : 82) | θ value | 0.813845 (rank : 22) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1056 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7KZI7, Q15449, Q15524, Q5XGA3, Q68A18, Q96HB3, Q96RG0 | Gene names | MARK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL motif kinase) (EMK1) (PAR1 homolog). | |||||
|
DLX3_HUMAN
|
||||||
| NC score | 0.005114 (rank : 83) | θ value | 6.88961 (rank : 53) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 369 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60479 | Gene names | DLX3 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein DLX-3. | |||||
|
ZN687_HUMAN
|
||||||
| NC score | 0.003249 (rank : 84) | θ value | 8.99809 (rank : 64) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8N1G0, Q68DQ8, Q9H937, Q9P2A7 | Gene names | ZNF687, KIAA1441 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 687. | |||||
|
PEG3_MOUSE
|
||||||
| NC score | 0.002936 (rank : 85) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
ZN690_HUMAN
|
||||||
| NC score | -0.000627 (rank : 86) | θ value | 8.99809 (rank : 65) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IWY8, Q32M75, Q32M76, Q8NA40 | Gene names | ZNF690 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 690. | |||||