Please be patient as the page loads
|
ST5_MOUSE
|
||||||
| SwissProt Accessions | Q924W7, Q78H54, Q8K2P3, Q924W8 | Gene names | St5, Dennd2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity 5 (DENN domain-containing protein 2B). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
DEN2A_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.966000 (rank : 6) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9ULE3, Q1RMD5, Q86XY0 | Gene names | DENND2A, KIAA1277 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 2A. | |||||
|
DEN2A_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.969802 (rank : 3) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8C4S8, Q3TJU2, Q3TUG3, Q3UXA3 | Gene names | Dennd2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 2A. | |||||
|
DEN2C_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.966924 (rank : 5) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q68D51, Q5TCX6, Q6P3R3 | Gene names | DENND2C | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 2C. | |||||
|
DEN2C_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.967556 (rank : 4) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6P9P8, Q3T9X2, Q3UV30, Q80V46 | Gene names | Dennd2c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 2C. | |||||
|
ST5_HUMAN
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.985741 (rank : 2) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P78524, P78523, Q16492, Q7KYY2, Q7KZ12, Q8NE12, Q9BQQ6 | Gene names | ST5, DENND2B, HTS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity 5 (HeLa tumor suppression 1) (DENN domain-containing protein 2B). | |||||
|
ST5_MOUSE
|
||||||
| θ value | 0 (rank : 6) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 126 | |
| SwissProt Accessions | Q924W7, Q78H54, Q8K2P3, Q924W8 | Gene names | St5, Dennd2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity 5 (DENN domain-containing protein 2B). | |||||
|
DEN2D_HUMAN
|
||||||
| θ value | 2.00348e-116 (rank : 7) | NC score | 0.959126 (rank : 8) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9H6A0, Q5T5V6, Q9BSU0 | Gene names | DENND2D | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 2D. | |||||
|
DEN2D_MOUSE
|
||||||
| θ value | 3.77839e-115 (rank : 8) | NC score | 0.959529 (rank : 7) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q91VV4, Q3U028, Q9D831 | Gene names | Dennd2d | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 2D. | |||||
|
DEN1A_MOUSE
|
||||||
| θ value | 1.08726e-37 (rank : 9) | NC score | 0.790143 (rank : 9) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8K382 | Gene names | Dennd1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 1A. | |||||
|
DEN1A_HUMAN
|
||||||
| θ value | 5.39604e-37 (rank : 10) | NC score | 0.784333 (rank : 10) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8TEH3, Q5VWF0, Q8IVD6, Q9H796 | Gene names | DENND1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 1A. | |||||
|
RA6I1_HUMAN
|
||||||
| θ value | 3.07829e-16 (rank : 11) | NC score | 0.562141 (rank : 12) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6IQ26, Q96GN3, Q9H6U7, Q9UFV0, Q9UPR1 | Gene names | RAB6IP1, KIAA1091 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab6-interacting protein 1 (Rab6IP1). | |||||
|
RA6I1_MOUSE
|
||||||
| θ value | 3.07829e-16 (rank : 12) | NC score | 0.562796 (rank : 11) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6PAL8, Q62146, Q8C829, Q8VDF6, Q9QYZ2 | Gene names | Rab6ip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab6-interacting protein 1 (Rab6IP1). | |||||
|
MTMR5_HUMAN
|
||||||
| θ value | 7.09661e-13 (rank : 13) | NC score | 0.417092 (rank : 15) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O95248, O60228, Q5JXD8, Q5PPM2, Q96GR9, Q9UGB8 | Gene names | SBF1, MTMR5 | |||
|
Domain Architecture |

|
|||||
| Description | SET-binding factor 1 (Sbf1) (Myotubularin-related protein 5). | |||||
|
MTMRD_HUMAN
|
||||||
| θ value | 3.52202e-12 (rank : 14) | NC score | 0.416332 (rank : 16) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q86WG5, Q3MJF0, Q68DQ3, Q6P459, Q6PJD1, Q7Z325, Q7Z621, Q86VE2, Q96FE2, Q9C097 | Gene names | SBF2, CMT4B2, KIAA1766, MTMR13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myotubularin-related protein 13 (SET-binding factor 2). | |||||
|
MYCPP_HUMAN
|
||||||
| θ value | 6.00763e-12 (rank : 15) | NC score | 0.557988 (rank : 13) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q7Z401, Q14655, Q86T77, Q8IVX2, Q8NB93 | Gene names | MYCPBP, IRLB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-myc promoter-binding protein. | |||||
|
DEN4C_HUMAN
|
||||||
| θ value | 2.13673e-09 (rank : 16) | NC score | 0.534993 (rank : 14) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q5VZ89, Q6AI48, Q6ZUB3, Q8NCY7, Q9H6N4, Q9NUT1, Q9NWA5 | Gene names | DENND4C, C9orf55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 4C. | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 17) | NC score | 0.048369 (rank : 23) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 18) | NC score | 0.065746 (rank : 21) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 19) | NC score | 0.067376 (rank : 20) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
CNOT4_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 20) | NC score | 0.086922 (rank : 18) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BT14, Q8CCR4, Q9CV74, Q9Z1D0 | Gene names | Cnot4, Not4 | |||
|
Domain Architecture |
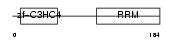
|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
CNOT4_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 21) | NC score | 0.089647 (rank : 17) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95628, O95339, O95627, Q8IYM7, Q8NCL0, Q9NPQ1, Q9NZN6 | Gene names | CNOT4, NOT4 | |||
|
Domain Architecture |
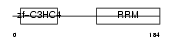
|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
GA2L3_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 22) | NC score | 0.044975 (rank : 26) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q86XJ1 | Gene names | GAS2L3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 3 (Growth arrest-specific 2-like 3). | |||||
|
DACT1_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 23) | NC score | 0.072985 (rank : 19) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NYF0, Q86TY0 | Gene names | DACT1, DPR1, HNG3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dapper homolog 1 (hDPR1) (Heptacellular carcinoma novel gene 3 protein). | |||||
|
MAP1A_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 24) | NC score | 0.040403 (rank : 29) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
ASPP1_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 25) | NC score | 0.023765 (rank : 59) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q62415 | Gene names | Ppp1r13b, Aspp1 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-stimulating of p53 protein 1 (Protein phosphatase 1 regulatory subunit 13B). | |||||
|
PTN23_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 26) | NC score | 0.031520 (rank : 43) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 726 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q6PB44, Q69ZJ0, Q8R1Z5, Q923E6 | Gene names | Ptpn23, Kiaa1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48). | |||||
|
DOCK4_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 27) | NC score | 0.031523 (rank : 42) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8N1I0, O14584, O94824, Q8NB45 | Gene names | DOCK4, KIAA0716 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 4. | |||||
|
DOCK4_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 28) | NC score | 0.032615 (rank : 41) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P59764 | Gene names | Dock4, Kiaa0716 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 4. | |||||
|
MDC1_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 29) | NC score | 0.051034 (rank : 22) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
MEGF9_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 30) | NC score | 0.020655 (rank : 69) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8BH27, Q8BWI4 | Gene names | Megf9, Egfl5, Kiaa0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
BCOR_MOUSE
|
||||||
| θ value | 0.163984 (rank : 31) | NC score | 0.019647 (rank : 72) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8CGN4, Q6PDK5, Q6ZPM3, Q8BKF5, Q8CGN1, Q8CGN2, Q8CGN3 | Gene names | Bcor, Kiaa1575 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BCoR protein (BCL-6 corepressor). | |||||
|
IF4B_MOUSE
|
||||||
| θ value | 0.163984 (rank : 32) | NC score | 0.038719 (rank : 32) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BGD9, Q3TD64 | Gene names | Eif4b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4B (eIF-4B). | |||||
|
RBMX2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 33) | NC score | 0.032618 (rank : 40) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y388, Q5JY82, Q9Y3I8 | Gene names | RBMX2 | |||
|
Domain Architecture |
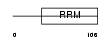
|
|||||
| Description | RNA-binding motif protein, X-linked 2. | |||||
|
DC1L2_MOUSE
|
||||||
| θ value | 0.21417 (rank : 34) | NC score | 0.035537 (rank : 35) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6PDL0 | Gene names | Dync1li2, Dncli2, Dnclic2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic dynein 1 light intermediate chain 2 (Dynein light intermediate chain 2, cytosolic). | |||||
|
PKN3_MOUSE
|
||||||
| θ value | 0.21417 (rank : 35) | NC score | 0.002746 (rank : 120) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1009 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K045 | Gene names | Pkn3, Pknbeta | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase N3 (EC 2.7.11.13) (Protein kinase PKN- beta) (Protein-kinase C-related kinase 3). | |||||
|
CI079_HUMAN
|
||||||
| θ value | 0.279714 (rank : 36) | NC score | 0.040672 (rank : 27) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6ZUB1, Q5SQC9, Q8NA41, Q8ND27 | Gene names | C9orf79 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C9orf79. | |||||
|
LEF1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 37) | NC score | 0.023344 (rank : 61) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P27782 | Gene names | Lef1, Lef-1 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphoid enhancer-binding factor 1 (LEF-1). | |||||
|
RC3H1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 38) | NC score | 0.022248 (rank : 64) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5TC82, Q5W180, Q5W181, Q8IVE6, Q8N9V1 | Gene names | RC3H1, KIAA2025 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Roquin (RING finger and C3H zinc finger protein 1). | |||||
|
TTBK1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 39) | NC score | 0.014922 (rank : 80) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1058 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q5TCY1, Q2L6C6, Q8N444, Q96JH2 | Gene names | TTBK1, BDTK, KIAA1855 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 1 (EC 2.7.11.1) (Brain-derived tau kinase). | |||||
|
LAD1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 40) | NC score | 0.040271 (rank : 30) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O00515, O95614 | Gene names | LAD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (120 kDa linear IgA bullous dermatosis antigen) (97 kDa linear IgA bullous dermatosis antigen) (Linear IgA disease antigen homolog) (LadA). | |||||
|
CRSP2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 41) | NC score | 0.036874 (rank : 33) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O60244, Q9UNB3 | Gene names | CRSP2, ARC150, DRIP150, EXLM1, TRAP170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CRSP complex subunit 2 (Cofactor required for Sp1 transcriptional activation subunit 2) (Transcriptional coactivator CRSP150) (Vitamin D3 receptor-interacting protein complex 150 kDa component) (DRIP150) (Thyroid hormone receptor-associated protein complex 170 kDa component) (Trap170) (Activator-recruited cofactor 150 kDa component) (ARC150). | |||||
|
PRRT3_HUMAN
|
||||||
| θ value | 0.62314 (rank : 42) | NC score | 0.034699 (rank : 38) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
SPAST_MOUSE
|
||||||
| θ value | 0.62314 (rank : 43) | NC score | 0.014855 (rank : 82) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9QYY8, Q6ZPY6, Q80VE0, Q9CVK0 | Gene names | Spast, Kiaa1083, Spg4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spastin. | |||||
|
BCL6_HUMAN
|
||||||
| θ value | 0.813845 (rank : 44) | NC score | -0.001059 (rank : 126) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P41182 | Gene names | BCL6, BCL5, LAZ3, ZBTB27, ZNF51 | |||
|
Domain Architecture |
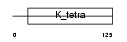
|
|||||
| Description | B-cell lymphoma 6 protein (BCL-6) (Zinc finger protein 51) (LAZ-3 protein) (BCL-5) (Zinc finger and BTB domain-containing protein 27). | |||||
|
CD2L7_HUMAN
|
||||||
| θ value | 0.813845 (rank : 45) | NC score | 0.008512 (rank : 102) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1275 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9NYV4, O94978 | Gene names | CRKRS, CRK7, KIAA0904 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle 2-related protein kinase 7 (EC 2.7.11.22) (CDC2- related protein kinase 7) (Cdc2-related kinase, arginine/serine-rich) (CrkRS). | |||||
|
GP152_MOUSE
|
||||||
| θ value | 0.813845 (rank : 46) | NC score | 0.018425 (rank : 76) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BXS7 | Gene names | Gpr152 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 152. | |||||
|
RBBP6_MOUSE
|
||||||
| θ value | 0.813845 (rank : 47) | NC score | 0.021568 (rank : 66) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
RBNS5_HUMAN
|
||||||
| θ value | 0.813845 (rank : 48) | NC score | 0.029374 (rank : 46) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H1K0, Q3KP30, Q59EY8, Q8NAQ1 | Gene names | ZFYVE20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rabenosyn-5 (FYVE finger-containing Rab5 effector protein rabenosyn-5) (Zinc finger FYVE domain-containing protein 20) (110 kDa protein). | |||||
|
SGK1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 49) | NC score | 0.001330 (rank : 121) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 848 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9WVC6 | Gene names | Sgk, Sgk1 | |||
|
Domain Architecture |
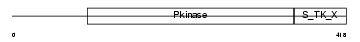
|
|||||
| Description | Serine/threonine-protein kinase Sgk1 (EC 2.7.11.1) (Serum/glucocorticoid-regulated kinase 1). | |||||
|
ALMS1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 50) | NC score | 0.023102 (rank : 63) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8K4E0, Q6A084, Q8C9N9 | Gene names | Alms1, Kiaa0328 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alstrom syndrome protein 1 homolog. | |||||
|
B4GN3_MOUSE
|
||||||
| θ value | 1.06291 (rank : 51) | NC score | 0.027895 (rank : 51) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6L8S8 | Gene names | B4galnt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 2 (EC 2.4.1.244) (NGalNAc-T2) (Beta- 1,4-N-acetylgalactosaminyltransferase III) (Beta4GalNAc-T3) (Beta4GalNAcT3). | |||||
|
CENPC_HUMAN
|
||||||
| θ value | 1.06291 (rank : 52) | NC score | 0.030397 (rank : 45) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q03188, Q9P0M5 | Gene names | CENPC1, CENPC, ICEN7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein C 1 (CENP-C) (Centromere autoantigen C) (Interphase centromere complex protein 7). | |||||
|
MBB1A_HUMAN
|
||||||
| θ value | 1.06291 (rank : 53) | NC score | 0.034883 (rank : 37) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
NFH_MOUSE
|
||||||
| θ value | 1.06291 (rank : 54) | NC score | 0.022099 (rank : 65) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
SH23A_HUMAN
|
||||||
| θ value | 1.06291 (rank : 55) | NC score | 0.016508 (rank : 79) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BRG2, Q9Y2X4 | Gene names | SH2D3A, NSP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH2 domain-containing protein 3A (Novel SH2-containing protein 1). | |||||
|
SON_HUMAN
|
||||||
| θ value | 1.06291 (rank : 56) | NC score | 0.033722 (rank : 39) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
WDR46_MOUSE
|
||||||
| θ value | 1.06291 (rank : 57) | NC score | 0.023294 (rank : 62) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Z0H1 | Gene names | Wdr46, Bing4 | |||
|
Domain Architecture |
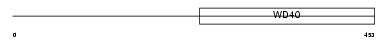
|
|||||
| Description | WD repeat protein 46 (WD repeat protein BING4). | |||||
|
FMN2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 58) | NC score | 0.028761 (rank : 47) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9JL04 | Gene names | Fmn2 | |||
|
Domain Architecture |

|
|||||
| Description | Formin-2. | |||||
|
GTSE1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 59) | NC score | 0.040606 (rank : 28) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9NYZ3, Q53GX5, Q5R3I6, Q6DHX4, Q9BRE0, Q9UGZ9, Q9Y557 | Gene names | GTSE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G2 and S phase expressed protein 1 (B99 homolog). | |||||
|
LEF1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 60) | NC score | 0.020320 (rank : 70) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UJU2, Q9HAZ0 | Gene names | LEF1 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphoid enhancer-binding factor 1 (LEF-1) (T cell-specific transcription factor 1-alpha) (TCF1-alpha). | |||||
|
MAGE1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 61) | NC score | 0.028758 (rank : 48) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
RAI1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 62) | NC score | 0.030921 (rank : 44) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q7Z5J4, Q8N3B4, Q8ND08, Q8WU64, Q96JK5, Q9H1C1, Q9H1C2, Q9UF69 | Gene names | RAI1, KIAA1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
TTLL5_HUMAN
|
||||||
| θ value | 1.38821 (rank : 63) | NC score | 0.013367 (rank : 87) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6EMB2, Q9P1V5, Q9UPZ4 | Gene names | TTLL5, KIAA0998, STAMP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tubulin--tyrosine ligase-like protein 5 (SRC1 and TIF2-associated modulatory protein). | |||||
|
CF152_MOUSE
|
||||||
| θ value | 1.81305 (rank : 64) | NC score | 0.025633 (rank : 53) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q80ST9, Q9CYM9, Q9D5J9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf152 homolog. | |||||
|
COIA1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | 0.013432 (rank : 86) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 481 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P39061, Q60672, Q61437, Q62001, Q62002, Q9JK63 | Gene names | Col18a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVIII) chain precursor [Contains: Endostatin]. | |||||
|
RECQ5_HUMAN
|
||||||
| θ value | 1.81305 (rank : 66) | NC score | 0.014505 (rank : 85) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O94762, Q9P1W7, Q9UNC8 | Gene names | RECQL5, RECQ5 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase Q5 (EC 3.6.1.-) (RecQ protein-like 5) (RecQ5). | |||||
|
STOX1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 67) | NC score | 0.034916 (rank : 36) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZVD7, Q4F8Q6, Q5I946, Q5I947, Q5I948, Q5VX38, Q5VX39, Q6ZRY3, Q96LR3, Q96LS0 | Gene names | STOX1, C10orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Storkhead-box protein 1 (Winged helix domain-containing protein). | |||||
|
CBP_MOUSE
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.023776 (rank : 58) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
DDX24_HUMAN
|
||||||
| θ value | 2.36792 (rank : 69) | NC score | 0.008560 (rank : 101) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9GZR7 | Gene names | DDX24 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX24 (EC 3.6.1.-) (DEAD box protein 24). | |||||
|
EHMT2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 70) | NC score | 0.010618 (rank : 94) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Z148, Q6PE08, Q8K4R6, Q8K4R7, Q9Z149 | Gene names | Ehmt2, Bat8, G9a, Ng36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
GAB1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 71) | NC score | 0.018243 (rank : 77) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QYY0, Q91VW7 | Gene names | Gab1 | |||
|
Domain Architecture |
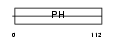
|
|||||
| Description | GRB2-associated-binding protein 1 (GRB2-associated binder 1). | |||||
|
PCIF1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 72) | NC score | 0.028181 (rank : 50) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H4Z3, Q9NT85 | Gene names | PCIF1, C20orf67 | |||
|
Domain Architecture |
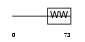
|
|||||
| Description | Phosphorylated CTD-interacting factor 1. | |||||
|
SEMG1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 73) | NC score | 0.016686 (rank : 78) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P04279, Q6X4I9, Q6Y809, Q6Y822, Q6Y823, Q86U64, Q96QM3 | Gene names | SEMG1, SEMG | |||
|
Domain Architecture |

|
|||||
| Description | Semenogelin-1 precursor (Semenogelin I) (SGI) [Contains: Alpha- inhibin-92; Alpha-inhibin-31; Seminal basic protein]. | |||||
|
TARSH_HUMAN
|
||||||
| θ value | 2.36792 (rank : 74) | NC score | 0.036157 (rank : 34) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q7Z7G0, Q6ZW20, Q6ZW22, Q9C082, Q9UFI6 | Gene names | ABI3BP, NESHBP, TARSH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Target of Nesh-SH3 precursor (Tarsh) (Nesh-binding protein) (NeshBP) (ABI gene family member 3-binding protein). | |||||
|
MUC1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 75) | NC score | 0.023819 (rank : 57) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
PUM2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 76) | NC score | 0.010811 (rank : 92) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TB72, O00234, Q53TV7, Q8WY43, Q9HAN2 | Gene names | PUM2, KIAA0235, PUMH2 | |||
|
Domain Architecture |

|
|||||
| Description | Pumilio homolog 2 (Pumilio-2). | |||||
|
ZIMP7_MOUSE
|
||||||
| θ value | 3.0926 (rank : 77) | NC score | 0.020814 (rank : 68) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 566 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8CIE2, Q6P558, Q6P9Q5, Q8BJ00 | Gene names | Zimp7, D11Bwg0280e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PIAS-like protein Zimp7. | |||||
|
ASXL1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 78) | NC score | 0.025585 (rank : 54) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8IXJ9, Q5JWS9, Q8IYY7, Q9H466, Q9NQF8, Q9UFJ0, Q9UFP8, Q9Y2I4 | Gene names | ASXL1, KIAA0978 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative Polycomb group protein ASXL1 (Additional sex combs-like protein 1). | |||||
|
BIG1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 79) | NC score | 0.007286 (rank : 106) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y6D6, Q9NV46, Q9UFV2, Q9UNL0 | Gene names | ARFGEF1, ARFGEP1, BIG1 | |||
|
Domain Architecture |

|
|||||
| Description | Brefeldin A-inhibited guanine nucleotide-exchange protein 1 (Brefeldin A-inhibited GEP 1) (p200 ARF-GEP1) (p200 ARF guanine nucleotide exchange factor). | |||||
|
CRSP7_HUMAN
|
||||||
| θ value | 4.03905 (rank : 80) | NC score | 0.028727 (rank : 49) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95402 | Gene names | CRSP7, ARC70 | |||
|
Domain Architecture |

|
|||||
| Description | CRSP complex subunit 7 (Cofactor required for Sp1 transcriptional activation subunit 7) (Transcriptional coactivator CRSP70) (Activator- recruited cofactor 70 kDa component) (ARC70). | |||||
|
ETV3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 81) | NC score | 0.019502 (rank : 73) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
GON4L_MOUSE
|
||||||
| θ value | 4.03905 (rank : 82) | NC score | 0.018459 (rank : 75) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9DB00, Q80TB4, Q91YI9 | Gene names | Gon4l, Gon4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
JPH2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 83) | NC score | 0.014709 (rank : 83) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9BR39, O95913, Q5JY74, Q9UJN4 | Gene names | JPH2, JP2 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
NIN_MOUSE
|
||||||
| θ value | 4.03905 (rank : 84) | NC score | 0.004023 (rank : 116) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q61043, Q6ZPM7 | Gene names | Nin, Kiaa1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein. | |||||
|
PRP4B_MOUSE
|
||||||
| θ value | 4.03905 (rank : 85) | NC score | 0.006054 (rank : 110) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1040 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q61136, O88378, Q8BND8, Q8R4Y5, Q9CTL9, Q9CTT0 | Gene names | Prpf4b, Cbp143, Prp4h, Prp4k, Prp4m, Prpk | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PRP4 homolog (EC 2.7.11.1) (PRP4 pre- mRNA-processing factor 4 homolog) (Pre-mRNA protein kinase). | |||||
|
RNF8_HUMAN
|
||||||
| θ value | 4.03905 (rank : 86) | NC score | 0.006649 (rank : 107) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 622 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O76064 | Gene names | RNF8, KIAA0646 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase protein RNF8 (EC 6.3.2.-) (RING finger protein 8). | |||||
|
RPGF1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 87) | NC score | 0.011370 (rank : 90) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q13905 | Gene names | RAPGEF1, GRF2 | |||
|
Domain Architecture |

|
|||||
| Description | Rap guanine nucleotide exchange factor 1 (Guanine nucleotide-releasing factor 2) (C3G protein) (CRK SH3-binding GNRP). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 88) | NC score | 0.045927 (rank : 24) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
WDR20_MOUSE
|
||||||
| θ value | 4.03905 (rank : 89) | NC score | 0.007571 (rank : 105) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D5R2 | Gene names | Wdr20 | |||
|
Domain Architecture |
|
|||||
| Description | WD repeat protein 20. | |||||
|
WNK2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 90) | NC score | 0.009225 (rank : 98) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
YTHD2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 91) | NC score | 0.014623 (rank : 84) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y5A9, Q5VSZ9, Q8TDH0, Q9BUJ5 | Gene names | YTHDF2, HGRG8 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain family protein 2 (High-glucose-regulated protein 8) (NY- REN-2 antigen) (CLL-associated antigen KW-14). | |||||
|
BAT2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 92) | NC score | 0.045078 (rank : 25) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
BAT2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 93) | NC score | 0.039693 (rank : 31) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
CF152_HUMAN
|
||||||
| θ value | 5.27518 (rank : 94) | NC score | 0.008751 (rank : 100) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86VQ0, Q9BWX7 | Gene names | C6orf152 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf152. | |||||
|
GP179_HUMAN
|
||||||
| θ value | 5.27518 (rank : 95) | NC score | 0.021550 (rank : 67) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6PRD1 | Gene names | GPR179, GPR158L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 179 precursor (Probable G-protein coupled receptor 158-like 1). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 96) | NC score | 0.020194 (rank : 71) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
JHD2B_HUMAN
|
||||||
| θ value | 5.27518 (rank : 97) | NC score | 0.009236 (rank : 97) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q7LBC6, Q9BVH6, Q9BW93, Q9BZ52, Q9NYF4, Q9UPS0 | Gene names | JMJD1B, C5orf7, JHDM2B, KIAA1082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 2B (EC 1.14.11.-) (Jumonji domain-containing protein 1B) (Nuclear protein 5qNCA). | |||||
|
K1543_MOUSE
|
||||||
| θ value | 5.27518 (rank : 98) | NC score | 0.018897 (rank : 74) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q80VC9, Q5DTW9, Q8BUZ0 | Gene names | Kiaa1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 99) | NC score | 0.024686 (rank : 55) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
ROCK1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 100) | NC score | 0.001046 (rank : 122) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 2126 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P70335, Q8C3G4, Q8C7H0 | Gene names | Rock1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 1 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 1) (p160 ROCK-1) (p160ROCK). | |||||
|
USBP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 101) | NC score | 0.011470 (rank : 89) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R370 | Gene names | Ushbp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USH1C-binding protein 1 (Usher syndrome type-1C protein-binding protein 1). | |||||
|
ZBT16_HUMAN
|
||||||
| θ value | 5.27518 (rank : 102) | NC score | -0.000729 (rank : 125) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1010 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q05516, Q8TAL4 | Gene names | ZBTB16, PLZF, ZNF145 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger and BTB domain-containing protein 16 (Zinc finger protein PLZF) (Promyelocytic leukemia zinc finger protein) (Zinc finger protein 145). | |||||
|
AT1A2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 103) | NC score | 0.005402 (rank : 112) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P50993, Q07059, Q86UZ5, Q9UQ25 | Gene names | ATP1A2, KIAA0778 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2). | |||||
|
AT1A2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 104) | NC score | 0.005397 (rank : 113) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PIE5 | Gene names | Atp1a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2) (Alpha(+)). | |||||
|
DLG4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 105) | NC score | 0.003551 (rank : 118) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P78352, Q92941, Q9UKK8 | Gene names | DLG4, PSD95 | |||
|
Domain Architecture |

|
|||||
| Description | Discs large homolog 4 (Postsynaptic density protein 95) (PSD-95) (Synapse-associated protein 90) (SAP90). | |||||
|
DLG4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 106) | NC score | 0.003733 (rank : 117) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q62108, Q5NCV5, Q5NCV6, Q5NCV7, Q91WJ1 | Gene names | Dlg4, Dlgh4, Psd95 | |||
|
Domain Architecture |

|
|||||
| Description | Discs large homolog 4 (Postsynaptic density protein 95) (PSD-95) (Synapse-associated protein 90) (SAP90). | |||||
|
HTSF1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 107) | NC score | 0.024457 (rank : 56) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BGC0, Q1WWK0, Q9CT41, Q9DAU3 | Gene names | Htatsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 homolog. | |||||
|
IF4B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 108) | NC score | 0.023638 (rank : 60) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P23588 | Gene names | EIF4B | |||
|
Domain Architecture |
|
|||||
| Description | Eukaryotic translation initiation factor 4B (eIF-4B). | |||||
|
KI13A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 109) | NC score | 0.004282 (rank : 115) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H1H9, Q9H1H8 | Gene names | KIF13A, RBKIN | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13A (Kinesin-like protein RBKIN). | |||||
|
NFIC_MOUSE
|
||||||
| θ value | 6.88961 (rank : 110) | NC score | 0.014901 (rank : 81) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P70255, O09072, P70256, Q3U2I9, Q99MA3, Q99MA4, Q99MA5, Q99MA6, Q99MA7, Q99MA8, Q9R1G3 | Gene names | Nfic | |||
|
Domain Architecture |
|
|||||
| Description | Nuclear factor 1 C-type (Nuclear factor 1/C) (NF1-C) (NFI-C) (NF-I/C) (CCAAT-box-binding transcription factor) (CTF) (TGGCA-binding protein). | |||||
|
PHF8_HUMAN
|
||||||
| θ value | 6.88961 (rank : 111) | NC score | 0.007892 (rank : 104) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UPP1, Q5H9U5, Q5VUJ4, Q7Z6D4, Q9HAH2 | Gene names | PHF8, KIAA1111, ZNF422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
TRAK1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 112) | NC score | 0.005218 (rank : 114) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UPV9, Q96B69 | Gene names | TRAK1, KIAA1042, OIP106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trafficking kinesin-binding protein 1 (106 kDa O-GlcNAc transferase- interacting protein). | |||||
|
WEE1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 113) | NC score | 0.000938 (rank : 123) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 794 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P47810, Q9EPL7 | Gene names | Wee1 | |||
|
Domain Architecture |

|
|||||
| Description | Wee1-like protein kinase (EC 2.7.10.2) (Wee1A kinase). | |||||
|
GCP6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 114) | NC score | 0.011043 (rank : 91) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96RT7, Q5JZ80, Q6PJ40, Q86YE9, Q9BY91, Q9UGX3, Q9UGX4 | Gene names | TUBGCP6, GCP6, KIAA1669 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-tubulin complex component 6 (GCP-6). | |||||
|
GP158_MOUSE
|
||||||
| θ value | 8.99809 (rank : 115) | NC score | 0.010806 (rank : 93) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8C419, Q8CHB0 | Gene names | Gpr158, Kiaa1136 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 158 precursor. | |||||
|
HGS_MOUSE
|
||||||
| θ value | 8.99809 (rank : 116) | NC score | 0.008257 (rank : 103) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99LI8, Q61691, Q8BQW3 | Gene names | Hgs, Hrs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate. | |||||
|
KI13A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 117) | NC score | 0.003203 (rank : 119) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9EQW7, O35062 | Gene names | Kif13a | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13A. | |||||
|
LRBA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 118) | NC score | 0.006199 (rank : 109) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P50851, Q9H2U3, Q9H2U4 | Gene names | LRBA, BGL, CDC4L, LBA | |||
|
Domain Architecture |

|
|||||
| Description | Lipopolysaccharide-responsive and beige-like anchor protein (CDC4-like protein) (Beige-like protein). | |||||
|
MBIP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 119) | NC score | 0.008937 (rank : 99) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99LQ1, Q3TBM3 | Gene names | Mbip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MAP3K12-binding inhibitory protein 1 (MAPK upstream kinase-binding inhibitory protein) (MUK-binding inhibitory protein). | |||||
|
NPY4R_HUMAN
|
||||||
| θ value | 8.99809 (rank : 120) | NC score | -0.000329 (rank : 124) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P50391, Q13456 | Gene names | PPYR1, NPY4R | |||
|
Domain Architecture |
|
|||||
| Description | Neuropeptide Y receptor type 4 (NPY4-R) (Pancreatic polypeptide receptor 1) (PP1). | |||||
|
OBF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | 0.011940 (rank : 88) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q16633, Q14983 | Gene names | POU2AF1, OBF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | POU domain class 2-associating factor 1 (B-cell-specific coactivator OBF-1) (OCT-binding factor 1) (BOB-1) (OCA-B). | |||||
|
P73L_HUMAN
|
||||||
| θ value | 8.99809 (rank : 122) | NC score | 0.006025 (rank : 111) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H3D4, O75080, O75195, O75922, O76078, Q6VEG2, Q6VEG3, Q6VEG4, Q6VFJ1, Q6VFJ2, Q6VFJ3, Q6VH20, Q7LDI3, Q7LDI4, Q7LDI5, Q96KR0, Q9H3D2, Q9H3D3, Q9H3P8, Q9NPH7, Q9P1B4, Q9P1B5, Q9P1B6, Q9P1B7, Q9UBV9, Q9UE10, Q9UP26, Q9UP27, Q9UP28, Q9UP74 | Gene names | TP73L, KET, P63, P73H, P73L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor protein p73-like (p73L) (p63) (Tumor protein 63) (TP63) (p51) (p40) (Keratinocyte transcription factor KET) (Chronic ulcerative stomatitis protein) (CUSP). | |||||
|
RT34_HUMAN
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | 0.026759 (rank : 52) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P82930, Q9BVI7 | Gene names | MRPS34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial 28S ribosomal protein S34 (S34mt) (MRP-S34). | |||||
|
SIA7A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.010071 (rank : 95) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9QZ39, Q9JJP5 | Gene names | St6galnac1, Siat7a | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-N-acetylgalactosaminide alpha-2,6-sialyltransferase 1 (EC 2.4.99.3) (GalNAc alpha-2,6-sialyltransferase I) (ST6GalNAc I) (Sialyltransferase 7A). | |||||
|
SPR1B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 125) | NC score | 0.009629 (rank : 96) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q62267 | Gene names | Sprr1b | |||
|
Domain Architecture |

|
|||||
| Description | Cornifin B (Small proline-rich protein 1B) (SPR1B) (SPR1 B). | |||||
|
TOM1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 126) | NC score | 0.006561 (rank : 108) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60784, Q5TIJ6 | Gene names | TOM1 | |||
|
Domain Architecture |
|
|||||
| Description | Target of Myb protein 1. | |||||
|
ST5_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 6) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 126 | |
| SwissProt Accessions | Q924W7, Q78H54, Q8K2P3, Q924W8 | Gene names | St5, Dennd2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity 5 (DENN domain-containing protein 2B). | |||||
|
ST5_HUMAN
|
||||||
| NC score | 0.985741 (rank : 2) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P78524, P78523, Q16492, Q7KYY2, Q7KZ12, Q8NE12, Q9BQQ6 | Gene names | ST5, DENND2B, HTS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity 5 (HeLa tumor suppression 1) (DENN domain-containing protein 2B). | |||||
|
DEN2A_MOUSE
|
||||||
| NC score | 0.969802 (rank : 3) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8C4S8, Q3TJU2, Q3TUG3, Q3UXA3 | Gene names | Dennd2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 2A. | |||||
|
DEN2C_MOUSE
|
||||||
| NC score | 0.967556 (rank : 4) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6P9P8, Q3T9X2, Q3UV30, Q80V46 | Gene names | Dennd2c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 2C. | |||||
|
DEN2C_HUMAN
|
||||||
| NC score | 0.966924 (rank : 5) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q68D51, Q5TCX6, Q6P3R3 | Gene names | DENND2C | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 2C. | |||||
|
DEN2A_HUMAN
|
||||||
| NC score | 0.966000 (rank : 6) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9ULE3, Q1RMD5, Q86XY0 | Gene names | DENND2A, KIAA1277 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 2A. | |||||
|
DEN2D_MOUSE
|
||||||
| NC score | 0.959529 (rank : 7) | θ value | 3.77839e-115 (rank : 8) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q91VV4, Q3U028, Q9D831 | Gene names | Dennd2d | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 2D. | |||||
|
DEN2D_HUMAN
|
||||||
| NC score | 0.959126 (rank : 8) | θ value | 2.00348e-116 (rank : 7) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9H6A0, Q5T5V6, Q9BSU0 | Gene names | DENND2D | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 2D. | |||||
|
DEN1A_MOUSE
|
||||||
| NC score | 0.790143 (rank : 9) | θ value | 1.08726e-37 (rank : 9) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8K382 | Gene names | Dennd1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 1A. | |||||
|
DEN1A_HUMAN
|
||||||
| NC score | 0.784333 (rank : 10) | θ value | 5.39604e-37 (rank : 10) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8TEH3, Q5VWF0, Q8IVD6, Q9H796 | Gene names | DENND1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 1A. | |||||
|
RA6I1_MOUSE
|
||||||
| NC score | 0.562796 (rank : 11) | θ value | 3.07829e-16 (rank : 12) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6PAL8, Q62146, Q8C829, Q8VDF6, Q9QYZ2 | Gene names | Rab6ip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab6-interacting protein 1 (Rab6IP1). | |||||
|
RA6I1_HUMAN
|
||||||
| NC score | 0.562141 (rank : 12) | θ value | 3.07829e-16 (rank : 11) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6IQ26, Q96GN3, Q9H6U7, Q9UFV0, Q9UPR1 | Gene names | RAB6IP1, KIAA1091 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab6-interacting protein 1 (Rab6IP1). | |||||
|
MYCPP_HUMAN
|
||||||
| NC score | 0.557988 (rank : 13) | θ value | 6.00763e-12 (rank : 15) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q7Z401, Q14655, Q86T77, Q8IVX2, Q8NB93 | Gene names | MYCPBP, IRLB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-myc promoter-binding protein. | |||||
|
DEN4C_HUMAN
|
||||||
| NC score | 0.534993 (rank : 14) | θ value | 2.13673e-09 (rank : 16) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q5VZ89, Q6AI48, Q6ZUB3, Q8NCY7, Q9H6N4, Q9NUT1, Q9NWA5 | Gene names | DENND4C, C9orf55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 4C. | |||||
|
MTMR5_HUMAN
|
||||||
| NC score | 0.417092 (rank : 15) | θ value | 7.09661e-13 (rank : 13) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O95248, O60228, Q5JXD8, Q5PPM2, Q96GR9, Q9UGB8 | Gene names | SBF1, MTMR5 | |||
|
Domain Architecture |

|
|||||
| Description | SET-binding factor 1 (Sbf1) (Myotubularin-related protein 5). | |||||
|
MTMRD_HUMAN
|
||||||
| NC score | 0.416332 (rank : 16) | θ value | 3.52202e-12 (rank : 14) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q86WG5, Q3MJF0, Q68DQ3, Q6P459, Q6PJD1, Q7Z325, Q7Z621, Q86VE2, Q96FE2, Q9C097 | Gene names | SBF2, CMT4B2, KIAA1766, MTMR13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myotubularin-related protein 13 (SET-binding factor 2). | |||||
|
CNOT4_HUMAN
|
||||||
| NC score | 0.089647 (rank : 17) | θ value | 0.0148317 (rank : 21) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95628, O95339, O95627, Q8IYM7, Q8NCL0, Q9NPQ1, Q9NZN6 | Gene names | CNOT4, NOT4 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
CNOT4_MOUSE
|
||||||
| NC score | 0.086922 (rank : 18) | θ value | 0.0113563 (rank : 20) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BT14, Q8CCR4, Q9CV74, Q9Z1D0 | Gene names | Cnot4, Not4 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
DACT1_HUMAN
|
||||||
| NC score | 0.072985 (rank : 19) | θ value | 0.0330416 (rank : 23) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NYF0, Q86TY0 | Gene names | DACT1, DPR1, HNG3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dapper homolog 1 (hDPR1) (Heptacellular carcinoma novel gene 3 protein). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.067376 (rank : 20) | θ value | 0.00665767 (rank : 19) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.065746 (rank : 21) | θ value | 0.00390308 (rank : 18) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.051034 (rank : 22) | θ value | 0.0961366 (rank : 29) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.048369 (rank : 23) | θ value | 0.00134147 (rank : 17) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.045927 (rank : 24) | θ value | 4.03905 (rank : 88) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
BAT2_HUMAN
|
||||||
| NC score | 0.045078 (rank : 25) | θ value | 5.27518 (rank : 92) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
GA2L3_HUMAN
|
||||||
| NC score | 0.044975 (rank : 26) | θ value | 0.0193708 (rank : 22) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q86XJ1 | Gene names | GAS2L3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 3 (Growth arrest-specific 2-like 3). | |||||
|
CI079_HUMAN
|
||||||
| NC score | 0.040672 (rank : 27) | θ value | 0.279714 (rank : 36) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6ZUB1, Q5SQC9, Q8NA41, Q8ND27 | Gene names | C9orf79 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C9orf79. | |||||
|
GTSE1_HUMAN
|
||||||
| NC score | 0.040606 (rank : 28) | θ value | 1.38821 (rank : 59) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9NYZ3, Q53GX5, Q5R3I6, Q6DHX4, Q9BRE0, Q9UGZ9, Q9Y557 | Gene names | GTSE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G2 and S phase expressed protein 1 (B99 homolog). | |||||
|
MAP1A_HUMAN
|
||||||
| NC score | 0.040403 (rank : 29) | θ value | 0.0431538 (rank : 24) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
LAD1_HUMAN
|
||||||
| NC score | 0.040271 (rank : 30) | θ value | 0.47712 (rank : 40) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O00515, O95614 | Gene names | LAD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (120 kDa linear IgA bullous dermatosis antigen) (97 kDa linear IgA bullous dermatosis antigen) (Linear IgA disease antigen homolog) (LadA). | |||||
|
BAT2_MOUSE
|
||||||
| NC score | 0.039693 (rank : 31) | θ value | 5.27518 (rank : 93) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
IF4B_MOUSE
|
||||||
| NC score | 0.038719 (rank : 32) | θ value | 0.163984 (rank : 32) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BGD9, Q3TD64 | Gene names | Eif4b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4B (eIF-4B). | |||||
|
CRSP2_HUMAN
|
||||||
| NC score | 0.036874 (rank : 33) | θ value | 0.62314 (rank : 41) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O60244, Q9UNB3 | Gene names | CRSP2, ARC150, DRIP150, EXLM1, TRAP170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CRSP complex subunit 2 (Cofactor required for Sp1 transcriptional activation subunit 2) (Transcriptional coactivator CRSP150) (Vitamin D3 receptor-interacting protein complex 150 kDa component) (DRIP150) (Thyroid hormone receptor-associated protein complex 170 kDa component) (Trap170) (Activator-recruited cofactor 150 kDa component) (ARC150). | |||||
|
TARSH_HUMAN
|
||||||
| NC score | 0.036157 (rank : 34) | θ value | 2.36792 (rank : 74) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q7Z7G0, Q6ZW20, Q6ZW22, Q9C082, Q9UFI6 | Gene names | ABI3BP, NESHBP, TARSH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Target of Nesh-SH3 precursor (Tarsh) (Nesh-binding protein) (NeshBP) (ABI gene family member 3-binding protein). | |||||
|
DC1L2_MOUSE
|
||||||
| NC score | 0.035537 (rank : 35) | θ value | 0.21417 (rank : 34) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6PDL0 | Gene names | Dync1li2, Dncli2, Dnclic2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic dynein 1 light intermediate chain 2 (Dynein light intermediate chain 2, cytosolic). | |||||
|
STOX1_HUMAN
|
||||||
| NC score | 0.034916 (rank : 36) | θ value | 1.81305 (rank : 67) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZVD7, Q4F8Q6, Q5I946, Q5I947, Q5I948, Q5VX38, Q5VX39, Q6ZRY3, Q96LR3, Q96LS0 | Gene names | STOX1, C10orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Storkhead-box protein 1 (Winged helix domain-containing protein). | |||||
|
MBB1A_HUMAN
|
||||||
| NC score | 0.034883 (rank : 37) | θ value | 1.06291 (rank : 53) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
PRRT3_HUMAN
|
||||||
| NC score | 0.034699 (rank : 38) | θ value | 0.62314 (rank : 42) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.033722 (rank : 39) | θ value | 1.06291 (rank : 56) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
RBMX2_HUMAN
|
||||||
| NC score | 0.032618 (rank : 40) | θ value | 0.163984 (rank : 33) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y388, Q5JY82, Q9Y3I8 | Gene names | RBMX2 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding motif protein, X-linked 2. | |||||
|
DOCK4_MOUSE
|
||||||
| NC score | 0.032615 (rank : 41) | θ value | 0.0961366 (rank : 28) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P59764 | Gene names | Dock4, Kiaa0716 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 4. | |||||
|
DOCK4_HUMAN
|
||||||
| NC score | 0.031523 (rank : 42) | θ value | 0.0961366 (rank : 27) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8N1I0, O14584, O94824, Q8NB45 | Gene names | DOCK4, KIAA0716 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 4. | |||||
|
PTN23_MOUSE
|
||||||
| NC score | 0.031520 (rank : 43) | θ value | 0.0736092 (rank : 26) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 726 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q6PB44, Q69ZJ0, Q8R1Z5, Q923E6 | Gene names | Ptpn23, Kiaa1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48). | |||||
|
RAI1_HUMAN
|
||||||
| NC score | 0.030921 (rank : 44) | θ value | 1.38821 (rank : 62) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q7Z5J4, Q8N3B4, Q8ND08, Q8WU64, Q96JK5, Q9H1C1, Q9H1C2, Q9UF69 | Gene names | RAI1, KIAA1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
CENPC_HUMAN
|
||||||
| NC score | 0.030397 (rank : 45) | θ value | 1.06291 (rank : 52) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q03188, Q9P0M5 | Gene names | CENPC1, CENPC, ICEN7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein C 1 (CENP-C) (Centromere autoantigen C) (Interphase centromere complex protein 7). | |||||
|
RBNS5_HUMAN
|
||||||
| NC score | 0.029374 (rank : 46) | θ value | 0.813845 (rank : 48) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H1K0, Q3KP30, Q59EY8, Q8NAQ1 | Gene names | ZFYVE20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rabenosyn-5 (FYVE finger-containing Rab5 effector protein rabenosyn-5) (Zinc finger FYVE domain-containing protein 20) (110 kDa protein). | |||||
|
FMN2_MOUSE
|
||||||
| NC score | 0.028761 (rank : 47) | θ value | 1.38821 (rank : 58) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9JL04 | Gene names | Fmn2 | |||
|
Domain Architecture |

|
|||||
| Description | Formin-2. | |||||
|
MAGE1_HUMAN
|
||||||
| NC score | 0.028758 (rank : 48) | θ value | 1.38821 (rank : 61) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
CRSP7_HUMAN
|
||||||
| NC score | 0.028727 (rank : 49) | θ value | 4.03905 (rank : 80) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95402 | Gene names | CRSP7, ARC70 | |||
|
Domain Architecture |

|
|||||
| Description | CRSP complex subunit 7 (Cofactor required for Sp1 transcriptional activation subunit 7) (Transcriptional coactivator CRSP70) (Activator- recruited cofactor 70 kDa component) (ARC70). | |||||
|
PCIF1_HUMAN
|
||||||
| NC score | 0.028181 (rank : 50) | θ value | 2.36792 (rank : 72) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H4Z3, Q9NT85 | Gene names | PCIF1, C20orf67 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphorylated CTD-interacting factor 1. | |||||
|
B4GN3_MOUSE
|
||||||
| NC score | 0.027895 (rank : 51) | θ value | 1.06291 (rank : 51) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6L8S8 | Gene names | B4galnt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 2 (EC 2.4.1.244) (NGalNAc-T2) (Beta- 1,4-N-acetylgalactosaminyltransferase III) (Beta4GalNAc-T3) (Beta4GalNAcT3). | |||||
|
RT34_HUMAN
|
||||||
| NC score | 0.026759 (rank : 52) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P82930, Q9BVI7 | Gene names | MRPS34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial 28S ribosomal protein S34 (S34mt) (MRP-S34). | |||||
|
CF152_MOUSE
|
||||||
| NC score | 0.025633 (rank : 53) | θ value | 1.81305 (rank : 64) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q80ST9, Q9CYM9, Q9D5J9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf152 homolog. | |||||
|
ASXL1_HUMAN
|
||||||
| NC score | 0.025585 (rank : 54) | θ value | 4.03905 (rank : 78) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8IXJ9, Q5JWS9, Q8IYY7, Q9H466, Q9NQF8, Q9UFJ0, Q9UFP8, Q9Y2I4 | Gene names | ASXL1, KIAA0978 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative Polycomb group protein ASXL1 (Additional sex combs-like protein 1). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.024686 (rank : 55) | θ value | 5.27518 (rank : 99) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
HTSF1_MOUSE
|
||||||
| NC score | 0.024457 (rank : 56) | θ value | 6.88961 (rank : 107) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BGC0, Q1WWK0, Q9CT41, Q9DAU3 | Gene names | Htatsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 homolog. | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.023819 (rank : 57) | θ value | 3.0926 (rank : 75) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
CBP_MOUSE
|
||||||
| NC score | 0.023776 (rank : 58) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
ASPP1_MOUSE
|
||||||
| NC score | 0.023765 (rank : 59) | θ value | 0.0563607 (rank : 25) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q62415 | Gene names | Ppp1r13b, Aspp1 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-stimulating of p53 protein 1 (Protein phosphatase 1 regulatory subunit 13B). | |||||
|
IF4B_HUMAN
|
||||||
| NC score | 0.023638 (rank : 60) | θ value | 6.88961 (rank : 108) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P23588 | Gene names | EIF4B | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4B (eIF-4B). | |||||
|
LEF1_MOUSE
|
||||||
| NC score | 0.023344 (rank : 61) | θ value | 0.279714 (rank : 37) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P27782 | Gene names | Lef1, Lef-1 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphoid enhancer-binding factor 1 (LEF-1). | |||||
|
WDR46_MOUSE
|
||||||
| NC score | 0.023294 (rank : 62) | θ value | 1.06291 (rank : 57) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Z0H1 | Gene names | Wdr46, Bing4 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 46 (WD repeat protein BING4). | |||||
|
ALMS1_MOUSE
|
||||||
| NC score | 0.023102 (rank : 63) | θ value | 1.06291 (rank : 50) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8K4E0, Q6A084, Q8C9N9 | Gene names | Alms1, Kiaa0328 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alstrom syndrome protein 1 homolog. | |||||
|
RC3H1_HUMAN
|
||||||
| NC score | 0.022248 (rank : 64) | θ value | 0.279714 (rank : 38) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5TC82, Q5W180, Q5W181, Q8IVE6, Q8N9V1 | Gene names | RC3H1, KIAA2025 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Roquin (RING finger and C3H zinc finger protein 1). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.022099 (rank : 65) | θ value | 1.06291 (rank : 54) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
RBBP6_MOUSE
|
||||||
| NC score | 0.021568 (rank : 66) | θ value | 0.813845 (rank : 47) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
GP179_HUMAN
|
||||||
| NC score | 0.021550 (rank : 67) | θ value | 5.27518 (rank : 95) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6PRD1 | Gene names | GPR179, GPR158L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 179 precursor (Probable G-protein coupled receptor 158-like 1). | |||||
|
ZIMP7_MOUSE
|
||||||
| NC score | 0.020814 (rank : 68) | θ value | 3.0926 (rank : 77) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 566 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8CIE2, Q6P558, Q6P9Q5, Q8BJ00 | Gene names | Zimp7, D11Bwg0280e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PIAS-like protein Zimp7. | |||||
|
MEGF9_MOUSE
|
||||||
| NC score | 0.020655 (rank : 69) | θ value | 0.0961366 (rank : 30) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8BH27, Q8BWI4 | Gene names | Megf9, Egfl5, Kiaa0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
LEF1_HUMAN
|
||||||
| NC score | 0.020320 (rank : 70) | θ value | 1.38821 (rank : 60) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UJU2, Q9HAZ0 | Gene names | LEF1 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphoid enhancer-binding factor 1 (LEF-1) (T cell-specific transcription factor 1-alpha) (TCF1-alpha). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.020194 (rank : 71) | θ value | 5.27518 (rank : 96) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
BCOR_MOUSE
|
||||||
| NC score | 0.019647 (rank : 72) | θ value | 0.163984 (rank : 31) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8CGN4, Q6PDK5, Q6ZPM3, Q8BKF5, Q8CGN1, Q8CGN2, Q8CGN3 | Gene names | Bcor, Kiaa1575 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BCoR protein (BCL-6 corepressor). | |||||
|
ETV3_HUMAN
|
||||||
| NC score | 0.019502 (rank : 73) | θ value | 4.03905 (rank : 81) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
K1543_MOUSE
|
||||||
| NC score | 0.018897 (rank : 74) | θ value | 5.27518 (rank : 98) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q80VC9, Q5DTW9, Q8BUZ0 | Gene names | Kiaa1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
GON4L_MOUSE
|
||||||
| NC score | 0.018459 (rank : 75) | θ value | 4.03905 (rank : 82) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9DB00, Q80TB4, Q91YI9 | Gene names | Gon4l, Gon4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
GP152_MOUSE
|
||||||
| NC score | 0.018425 (rank : 76) | θ value | 0.813845 (rank : 46) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BXS7 | Gene names | Gpr152 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 152. | |||||
|
GAB1_MOUSE
|
||||||
| NC score | 0.018243 (rank : 77) | θ value | 2.36792 (rank : 71) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QYY0, Q91VW7 | Gene names | Gab1 | |||
|
Domain Architecture |

|
|||||
| Description | GRB2-associated-binding protein 1 (GRB2-associated binder 1). | |||||
|
SEMG1_HUMAN
|
||||||
| NC score | 0.016686 (rank : 78) | θ value | 2.36792 (rank : 73) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P04279, Q6X4I9, Q6Y809, Q6Y822, Q6Y823, Q86U64, Q96QM3 | Gene names | SEMG1, SEMG | |||
|
Domain Architecture |

|
|||||
| Description | Semenogelin-1 precursor (Semenogelin I) (SGI) [Contains: Alpha- inhibin-92; Alpha-inhibin-31; Seminal basic protein]. | |||||
|
SH23A_HUMAN
|
||||||
| NC score | 0.016508 (rank : 79) | θ value | 1.06291 (rank : 55) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BRG2, Q9Y2X4 | Gene names | SH2D3A, NSP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH2 domain-containing protein 3A (Novel SH2-containing protein 1). | |||||
|
TTBK1_HUMAN
|
||||||
| NC score | 0.014922 (rank : 80) | θ value | 0.279714 (rank : 39) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1058 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q5TCY1, Q2L6C6, Q8N444, Q96JH2 | Gene names | TTBK1, BDTK, KIAA1855 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 1 (EC 2.7.11.1) (Brain-derived tau kinase). | |||||
|
NFIC_MOUSE
|
||||||
| NC score | 0.014901 (rank : 81) | θ value | 6.88961 (rank : 110) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P70255, O09072, P70256, Q3U2I9, Q99MA3, Q99MA4, Q99MA5, Q99MA6, Q99MA7, Q99MA8, Q9R1G3 | Gene names | Nfic | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor 1 C-type (Nuclear factor 1/C) (NF1-C) (NFI-C) (NF-I/C) (CCAAT-box-binding transcription factor) (CTF) (TGGCA-binding protein). | |||||
|
SPAST_MOUSE
|
||||||
| NC score | 0.014855 (rank : 82) | θ value | 0.62314 (rank : 43) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9QYY8, Q6ZPY6, Q80VE0, Q9CVK0 | Gene names | Spast, Kiaa1083, Spg4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spastin. | |||||
|
JPH2_HUMAN
|
||||||
| NC score | 0.014709 (rank : 83) | θ value | 4.03905 (rank : 83) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9BR39, O95913, Q5JY74, Q9UJN4 | Gene names | JPH2, JP2 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
YTHD2_HUMAN
|
||||||
| NC score | 0.014623 (rank : 84) | θ value | 4.03905 (rank : 91) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y5A9, Q5VSZ9, Q8TDH0, Q9BUJ5 | Gene names | YTHDF2, HGRG8 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain family protein 2 (High-glucose-regulated protein 8) (NY- REN-2 antigen) (CLL-associated antigen KW-14). | |||||
|
RECQ5_HUMAN
|
||||||
| NC score | 0.014505 (rank : 85) | θ value | 1.81305 (rank : 66) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O94762, Q9P1W7, Q9UNC8 | Gene names | RECQL5, RECQ5 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase Q5 (EC 3.6.1.-) (RecQ protein-like 5) (RecQ5). | |||||
|
COIA1_MOUSE
|
||||||
| NC score | 0.013432 (rank : 86) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 481 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P39061, Q60672, Q61437, Q62001, Q62002, Q9JK63 | Gene names | Col18a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVIII) chain precursor [Contains: Endostatin]. | |||||
|
TTLL5_HUMAN
|
||||||
| NC score | 0.013367 (rank : 87) | θ value | 1.38821 (rank : 63) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6EMB2, Q9P1V5, Q9UPZ4 | Gene names | TTLL5, KIAA0998, STAMP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tubulin--tyrosine ligase-like protein 5 (SRC1 and TIF2-associated modulatory protein). | |||||
|
OBF1_HUMAN
|
||||||
| NC score | 0.011940 (rank : 88) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q16633, Q14983 | Gene names | POU2AF1, OBF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | POU domain class 2-associating factor 1 (B-cell-specific coactivator OBF-1) (OCT-binding factor 1) (BOB-1) (OCA-B). | |||||
|
USBP1_MOUSE
|
||||||
| NC score | 0.011470 (rank : 89) | θ value | 5.27518 (rank : 101) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R370 | Gene names | Ushbp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USH1C-binding protein 1 (Usher syndrome type-1C protein-binding protein 1). | |||||
|
RPGF1_HUMAN
|
||||||
| NC score | 0.011370 (rank : 90) | θ value | 4.03905 (rank : 87) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q13905 | Gene names | RAPGEF1, GRF2 | |||
|
Domain Architecture |

|
|||||
| Description | Rap guanine nucleotide exchange factor 1 (Guanine nucleotide-releasing factor 2) (C3G protein) (CRK SH3-binding GNRP). | |||||
|
GCP6_HUMAN
|
||||||
| NC score | 0.011043 (rank : 91) | θ value | 8.99809 (rank : 114) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96RT7, Q5JZ80, Q6PJ40, Q86YE9, Q9BY91, Q9UGX3, Q9UGX4 | Gene names | TUBGCP6, GCP6, KIAA1669 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-tubulin complex component 6 (GCP-6). | |||||
|
PUM2_HUMAN
|
||||||
| NC score | 0.010811 (rank : 92) | θ value | 3.0926 (rank : 76) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TB72, O00234, Q53TV7, Q8WY43, Q9HAN2 | Gene names | PUM2, KIAA0235, PUMH2 | |||
|
Domain Architecture |

|
|||||
| Description | Pumilio homolog 2 (Pumilio-2). | |||||
|
GP158_MOUSE
|
||||||
| NC score | 0.010806 (rank : 93) | θ value | 8.99809 (rank : 115) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8C419, Q8CHB0 | Gene names | Gpr158, Kiaa1136 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 158 precursor. | |||||
|
EHMT2_MOUSE
|
||||||
| NC score | 0.010618 (rank : 94) | θ value | 2.36792 (rank : 70) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Z148, Q6PE08, Q8K4R6, Q8K4R7, Q9Z149 | Gene names | Ehmt2, Bat8, G9a, Ng36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
SIA7A_MOUSE
|
||||||
| NC score | 0.010071 (rank : 95) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9QZ39, Q9JJP5 | Gene names | St6galnac1, Siat7a | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-N-acetylgalactosaminide alpha-2,6-sialyltransferase 1 (EC 2.4.99.3) (GalNAc alpha-2,6-sialyltransferase I) (ST6GalNAc I) (Sialyltransferase 7A). | |||||
|
SPR1B_MOUSE
|
||||||
| NC score | 0.009629 (rank : 96) | θ value | 8.99809 (rank : 125) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q62267 | Gene names | Sprr1b | |||
|
Domain Architecture |

|
|||||
| Description | Cornifin B (Small proline-rich protein 1B) (SPR1B) (SPR1 B). | |||||
|
JHD2B_HUMAN
|
||||||
| NC score | 0.009236 (rank : 97) | θ value | 5.27518 (rank : 97) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q7LBC6, Q9BVH6, Q9BW93, Q9BZ52, Q9NYF4, Q9UPS0 | Gene names | JMJD1B, C5orf7, JHDM2B, KIAA1082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 2B (EC 1.14.11.-) (Jumonji domain-containing protein 1B) (Nuclear protein 5qNCA). | |||||
|
WNK2_HUMAN
|
||||||
| NC score | 0.009225 (rank : 98) | θ value | 4.03905 (rank : 90) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
MBIP1_MOUSE
|
||||||
| NC score | 0.008937 (rank : 99) | θ value | 8.99809 (rank : 119) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99LQ1, Q3TBM3 | Gene names | Mbip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MAP3K12-binding inhibitory protein 1 (MAPK upstream kinase-binding inhibitory protein) (MUK-binding inhibitory protein). | |||||
|
CF152_HUMAN
|
||||||
| NC score | 0.008751 (rank : 100) | θ value | 5.27518 (rank : 94) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86VQ0, Q9BWX7 | Gene names | C6orf152 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf152. | |||||
|
DDX24_HUMAN
|
||||||
| NC score | 0.008560 (rank : 101) | θ value | 2.36792 (rank : 69) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9GZR7 | Gene names | DDX24 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX24 (EC 3.6.1.-) (DEAD box protein 24). | |||||
|
CD2L7_HUMAN
|
||||||
| NC score | 0.008512 (rank : 102) | θ value | 0.813845 (rank : 45) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1275 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9NYV4, O94978 | Gene names | CRKRS, CRK7, KIAA0904 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle 2-related protein kinase 7 (EC 2.7.11.22) (CDC2- related protein kinase 7) (Cdc2-related kinase, arginine/serine-rich) (CrkRS). | |||||
|
HGS_MOUSE
|
||||||
| NC score | 0.008257 (rank : 103) | θ value | 8.99809 (rank : 116) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99LI8, Q61691, Q8BQW3 | Gene names | Hgs, Hrs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate. | |||||
|
PHF8_HUMAN
|
||||||
| NC score | 0.007892 (rank : 104) | θ value | 6.88961 (rank : 111) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UPP1, Q5H9U5, Q5VUJ4, Q7Z6D4, Q9HAH2 | Gene names | PHF8, KIAA1111, ZNF422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
WDR20_MOUSE
|
||||||
| NC score | 0.007571 (rank : 105) | θ value | 4.03905 (rank : 89) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D5R2 | Gene names | Wdr20 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 20. | |||||
|
BIG1_HUMAN
|
||||||
| NC score | 0.007286 (rank : 106) | θ value | 4.03905 (rank : 79) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y6D6, Q9NV46, Q9UFV2, Q9UNL0 | Gene names | ARFGEF1, ARFGEP1, BIG1 | |||
|
Domain Architecture |

|
|||||
| Description | Brefeldin A-inhibited guanine nucleotide-exchange protein 1 (Brefeldin A-inhibited GEP 1) (p200 ARF-GEP1) (p200 ARF guanine nucleotide exchange factor). | |||||
|
RNF8_HUMAN
|
||||||
| NC score | 0.006649 (rank : 107) | θ value | 4.03905 (rank : 86) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 622 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O76064 | Gene names | RNF8, KIAA0646 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase protein RNF8 (EC 6.3.2.-) (RING finger protein 8). | |||||
|
TOM1_HUMAN
|
||||||
| NC score | 0.006561 (rank : 108) | θ value | 8.99809 (rank : 126) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60784, Q5TIJ6 | Gene names | TOM1 | |||
|
Domain Architecture |

|
|||||
| Description | Target of Myb protein 1. | |||||
|
LRBA_HUMAN
|
||||||
| NC score | 0.006199 (rank : 109) | θ value | 8.99809 (rank : 118) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P50851, Q9H2U3, Q9H2U4 | Gene names | LRBA, BGL, CDC4L, LBA | |||
|
Domain Architecture |

|
|||||
| Description | Lipopolysaccharide-responsive and beige-like anchor protein (CDC4-like protein) (Beige-like protein). | |||||
|
PRP4B_MOUSE
|
||||||
| NC score | 0.006054 (rank : 110) | θ value | 4.03905 (rank : 85) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1040 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q61136, O88378, Q8BND8, Q8R4Y5, Q9CTL9, Q9CTT0 | Gene names | Prpf4b, Cbp143, Prp4h, Prp4k, Prp4m, Prpk | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PRP4 homolog (EC 2.7.11.1) (PRP4 pre- mRNA-processing factor 4 homolog) (Pre-mRNA protein kinase). | |||||
|
P73L_HUMAN
|
||||||
| NC score | 0.006025 (rank : 111) | θ value | 8.99809 (rank : 122) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H3D4, O75080, O75195, O75922, O76078, Q6VEG2, Q6VEG3, Q6VEG4, Q6VFJ1, Q6VFJ2, Q6VFJ3, Q6VH20, Q7LDI3, Q7LDI4, Q7LDI5, Q96KR0, Q9H3D2, Q9H3D3, Q9H3P8, Q9NPH7, Q9P1B4, Q9P1B5, Q9P1B6, Q9P1B7, Q9UBV9, Q9UE10, Q9UP26, Q9UP27, Q9UP28, Q9UP74 | Gene names | TP73L, KET, P63, P73H, P73L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor protein p73-like (p73L) (p63) (Tumor protein 63) (TP63) (p51) (p40) (Keratinocyte transcription factor KET) (Chronic ulcerative stomatitis protein) (CUSP). | |||||
|
AT1A2_HUMAN
|
||||||
| NC score | 0.005402 (rank : 112) | θ value | 6.88961 (rank : 103) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P50993, Q07059, Q86UZ5, Q9UQ25 | Gene names | ATP1A2, KIAA0778 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2). | |||||
|
AT1A2_MOUSE
|
||||||
| NC score | 0.005397 (rank : 113) | θ value | 6.88961 (rank : 104) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PIE5 | Gene names | Atp1a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2) (Alpha(+)). | |||||
|
TRAK1_HUMAN
|
||||||
| NC score | 0.005218 (rank : 114) | θ value | 6.88961 (rank : 112) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UPV9, Q96B69 | Gene names | TRAK1, KIAA1042, OIP106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trafficking kinesin-binding protein 1 (106 kDa O-GlcNAc transferase- interacting protein). | |||||
|
KI13A_HUMAN
|
||||||
| NC score | 0.004282 (rank : 115) | θ value | 6.88961 (rank : 109) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H1H9, Q9H1H8 | Gene names | KIF13A, RBKIN | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13A (Kinesin-like protein RBKIN). | |||||
|
NIN_MOUSE
|
||||||
| NC score | 0.004023 (rank : 116) | θ value | 4.03905 (rank : 84) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q61043, Q6ZPM7 | Gene names | Nin, Kiaa1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein. | |||||
|
DLG4_MOUSE
|
||||||
| NC score | 0.003733 (rank : 117) | θ value | 6.88961 (rank : 106) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q62108, Q5NCV5, Q5NCV6, Q5NCV7, Q91WJ1 | Gene names | Dlg4, Dlgh4, Psd95 | |||
|
Domain Architecture |

|
|||||
| Description | Discs large homolog 4 (Postsynaptic density protein 95) (PSD-95) (Synapse-associated protein 90) (SAP90). | |||||
|
DLG4_HUMAN
|
||||||
| NC score | 0.003551 (rank : 118) | θ value | 6.88961 (rank : 105) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P78352, Q92941, Q9UKK8 | Gene names | DLG4, PSD95 | |||
|
Domain Architecture |

|
|||||
| Description | Discs large homolog 4 (Postsynaptic density protein 95) (PSD-95) (Synapse-associated protein 90) (SAP90). | |||||
|
KI13A_MOUSE
|
||||||
| NC score | 0.003203 (rank : 119) | θ value | 8.99809 (rank : 117) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9EQW7, O35062 | Gene names | Kif13a | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13A. | |||||
|
PKN3_MOUSE
|
||||||
| NC score | 0.002746 (rank : 120) | θ value | 0.21417 (rank : 35) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1009 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K045 | Gene names | Pkn3, Pknbeta | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase N3 (EC 2.7.11.13) (Protein kinase PKN- beta) (Protein-kinase C-related kinase 3). | |||||
|
SGK1_MOUSE
|
||||||
| NC score | 0.001330 (rank : 121) | θ value | 0.813845 (rank : 49) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 848 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9WVC6 | Gene names | Sgk, Sgk1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Sgk1 (EC 2.7.11.1) (Serum/glucocorticoid-regulated kinase 1). | |||||
|
ROCK1_MOUSE
|
||||||
| NC score | 0.001046 (rank : 122) | θ value | 5.27518 (rank : 100) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 2126 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P70335, Q8C3G4, Q8C7H0 | Gene names | Rock1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 1 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 1) (p160 ROCK-1) (p160ROCK). | |||||
|
WEE1_MOUSE
|
||||||
| NC score | 0.000938 (rank : 123) | θ value | 6.88961 (rank : 113) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 794 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P47810, Q9EPL7 | Gene names | Wee1 | |||
|
Domain Architecture |

|
|||||
| Description | Wee1-like protein kinase (EC 2.7.10.2) (Wee1A kinase). | |||||
|
NPY4R_HUMAN
|
||||||
| NC score | -0.000329 (rank : 124) | θ value | 8.99809 (rank : 120) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P50391, Q13456 | Gene names | PPYR1, NPY4R | |||
|
Domain Architecture |

|
|||||
| Description | Neuropeptide Y receptor type 4 (NPY4-R) (Pancreatic polypeptide receptor 1) (PP1). | |||||
|
ZBT16_HUMAN
|
||||||
| NC score | -0.000729 (rank : 125) | θ value | 5.27518 (rank : 102) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1010 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q05516, Q8TAL4 | Gene names | ZBTB16, PLZF, ZNF145 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 16 (Zinc finger protein PLZF) (Promyelocytic leukemia zinc finger protein) (Zinc finger protein 145). | |||||
|
BCL6_HUMAN
|
||||||
| NC score | -0.001059 (rank : 126) | θ value | 0.813845 (rank : 44) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P41182 | Gene names | BCL6, BCL5, LAZ3, ZBTB27, ZNF51 | |||
|
Domain Architecture |

|
|||||
| Description | B-cell lymphoma 6 protein (BCL-6) (Zinc finger protein 51) (LAZ-3 protein) (BCL-5) (Zinc finger and BTB domain-containing protein 27). | |||||