Please be patient as the page loads
|
SETB1_MOUSE
|
||||||
| SwissProt Accessions | O88974, Q922K1 | Gene names | Setdb1, Eset | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 4 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 4) (H3-K9-HMTase 4) (SET domain bifurcated 1) (ERG-associated protein with SET domain) (ESET). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SETB1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.977838 (rank : 2) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q15047, Q96GM9 | Gene names | SETDB1, KIAA0067 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 4 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 4) (H3-K9-HMTase 4) (SET domain bifurcated 1) (ERG-associated protein with SET domain) (ESET). | |||||
|
SETB1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 150 | |
| SwissProt Accessions | O88974, Q922K1 | Gene names | Setdb1, Eset | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 4 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 4) (H3-K9-HMTase 4) (SET domain bifurcated 1) (ERG-associated protein with SET domain) (ESET). | |||||
|
SETB2_HUMAN
|
||||||
| θ value | 3.35642e-63 (rank : 3) | NC score | 0.857298 (rank : 3) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q96T68, Q86UD6, Q96AI6 | Gene names | SETDB2, CLLD8 | |||
|
Domain Architecture |

|
|||||
| Description | Probable histone-lysine N-methyltransferase, H3 lysine-9 specific (EC 2.1.1.43) (Histone H3-K9 methyltransferase) (H3-K9-HMTase) (SET domain bifurcated 2) (Chronic lymphocytic leukemia deletion region gene 8 protein). | |||||
|
EHMT2_HUMAN
|
||||||
| θ value | 1.05554e-24 (rank : 4) | NC score | 0.361204 (rank : 18) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q96KQ7, Q14349, Q6PK06, Q96MH5, Q96QD0, Q9UQL8, Q9Y331 | Gene names | EHMT2, BAT8, G9A, NG36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
EHMT2_MOUSE
|
||||||
| θ value | 4.01107e-24 (rank : 5) | NC score | 0.359751 (rank : 19) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9Z148, Q6PE08, Q8K4R6, Q8K4R7, Q9Z149 | Gene names | Ehmt2, Bat8, G9a, Ng36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
EHMT1_HUMAN
|
||||||
| θ value | 1.16704e-23 (rank : 6) | NC score | 0.352339 (rank : 20) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9H9B1, Q8TCN7, Q96F53, Q96JF1, Q96KH4 | Gene names | EHMT1, EUHMTASE1, KIAA1876 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 5 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 5) (H3-K9-HMTase 5) (Euchromatic histone-lysine N-methyltransferase 1) (Eu-HMTase1) (G9a- like protein 1) (GLP1). | |||||
|
SUV91_HUMAN
|
||||||
| θ value | 1.2077e-20 (rank : 7) | NC score | 0.654981 (rank : 4) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O43463, Q53G60, Q6FHK6 | Gene names | SUV39H1, SUV39H | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 1 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 1) (H3-K9-HMTase 1) (Suppressor of variegation 3-9 homolog 1) (Su(var)3-9 homolog 1). | |||||
|
SUV91_MOUSE
|
||||||
| θ value | 5.99374e-20 (rank : 8) | NC score | 0.646271 (rank : 5) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O54864, Q9JLC7, Q9JLP8 | Gene names | Suv39h1, Suv39h | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 1 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 1) (H3-K9-HMTase 1) (Suppressor of variegation 3-9 homolog 1) (Su(var)3-9 homolog 1) (Position-effect variegation 3-9 homolog). | |||||
|
SUV92_MOUSE
|
||||||
| θ value | 2.35696e-16 (rank : 9) | NC score | 0.628047 (rank : 6) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9EQQ0, Q9CUK3, Q9JLP7 | Gene names | Suv39h2 | |||
|
Domain Architecture |
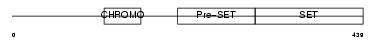
|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 2 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 2) (H3-K9-HMTase 2) (Suppressor of variegation 3-9 homolog 2) (Su(var)3-9 homolog 2). | |||||
|
SUV92_HUMAN
|
||||||
| θ value | 3.40345e-15 (rank : 10) | NC score | 0.627124 (rank : 7) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9H5I1 | Gene names | SUV39H2 | |||
|
Domain Architecture |
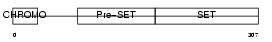
|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 2 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 2) (H3-K9-HMTase 2) (Suppressor of variegation 3-9 homolog 2) (Su(var)3-9 homolog 2). | |||||
|
MLL4_HUMAN
|
||||||
| θ value | 3.89403e-11 (rank : 11) | NC score | 0.436026 (rank : 14) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
SETD2_HUMAN
|
||||||
| θ value | 1.9326e-10 (rank : 12) | NC score | 0.520230 (rank : 8) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
HRX_HUMAN
|
||||||
| θ value | 9.59137e-10 (rank : 13) | NC score | 0.429521 (rank : 15) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 1.63604e-09 (rank : 14) | NC score | 0.342439 (rank : 21) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 87 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 3.08544e-08 (rank : 15) | NC score | 0.315423 (rank : 23) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
NSD1_HUMAN
|
||||||
| θ value | 3.08544e-08 (rank : 16) | NC score | 0.429519 (rank : 16) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
NSD1_MOUSE
|
||||||
| θ value | 3.08544e-08 (rank : 17) | NC score | 0.427981 (rank : 17) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 6.87365e-08 (rank : 18) | NC score | 0.328323 (rank : 22) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
SET1A_HUMAN
|
||||||
| θ value | 1.99992e-07 (rank : 19) | NC score | 0.464990 (rank : 12) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
EZH1_HUMAN
|
||||||
| θ value | 1.29631e-06 (rank : 20) | NC score | 0.474638 (rank : 9) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q92800, O43287, Q14459 | Gene names | EZH1, KIAA0388 | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of zeste homolog 1 (ENX-2). | |||||
|
EZH1_MOUSE
|
||||||
| θ value | 1.29631e-06 (rank : 21) | NC score | 0.474207 (rank : 10) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P70351, Q9R089 | Gene names | Ezh1, Enx2 | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of zeste homolog 1 (ENX-2). | |||||
|
EZH2_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 22) | NC score | 0.469258 (rank : 11) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q15910, Q15755, Q92857 | Gene names | EZH2 | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of zeste homolog 2 (ENX-1). | |||||
|
EZH2_MOUSE
|
||||||
| θ value | 3.19293e-05 (rank : 23) | NC score | 0.461803 (rank : 13) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q61188, Q9R090 | Gene names | Ezh2, Enx1h | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of zeste homolog 2 (ENX-1). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 24) | NC score | 0.047707 (rank : 86) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
PI5PA_HUMAN
|
||||||
| θ value | 0.125558 (rank : 25) | NC score | 0.056478 (rank : 57) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 749 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q15735, Q32M61, Q6ZTH6, Q8N902, Q9UDT9 | Gene names | PIB5PA, PIPP | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase A (EC 3.1.3.56). | |||||
|
SON_HUMAN
|
||||||
| θ value | 0.125558 (rank : 26) | NC score | 0.057271 (rank : 54) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 27) | NC score | 0.063321 (rank : 39) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
JPH3_HUMAN
|
||||||
| θ value | 0.163984 (rank : 28) | NC score | 0.025962 (rank : 124) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WXH2, Q8N471, Q9HDC3, Q9HDC4 | Gene names | JPH3, JP3 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-3 (Junctophilin type 3) (JP-3). | |||||
|
BAZ2B_HUMAN
|
||||||
| θ value | 0.21417 (rank : 29) | NC score | 0.076518 (rank : 28) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 803 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9UIF8, Q96EA1, Q96SQ8, Q9P252, Q9Y4N8 | Gene names | BAZ2B, KIAA1476 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2B (hWALp4). | |||||
|
EPAS1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 30) | NC score | 0.022215 (rank : 146) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
|
GOGA1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 31) | NC score | 0.031949 (rank : 108) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1222 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q92805, Q8IYZ9 | Gene names | GOLGA1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 1 (Golgin-97). | |||||
|
LTBP3_MOUSE
|
||||||
| θ value | 0.21417 (rank : 32) | NC score | 0.032594 (rank : 106) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 475 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q61810, Q8BNQ6 | Gene names | Ltbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 3 precursor (LTBP-3). | |||||
|
TARSH_HUMAN
|
||||||
| θ value | 0.21417 (rank : 33) | NC score | 0.073849 (rank : 29) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q7Z7G0, Q6ZW20, Q6ZW22, Q9C082, Q9UFI6 | Gene names | ABI3BP, NESHBP, TARSH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Target of Nesh-SH3 precursor (Tarsh) (Nesh-binding protein) (NeshBP) (ABI gene family member 3-binding protein). | |||||
|
VPS72_MOUSE
|
||||||
| θ value | 0.21417 (rank : 34) | NC score | 0.082024 (rank : 27) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q62481, Q3U2N4, Q810A9, Q99K81 | Gene names | Vps72, Tcfl1, Yl1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
MYH3_HUMAN
|
||||||
| θ value | 0.279714 (rank : 35) | NC score | 0.025067 (rank : 128) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1660 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P11055, Q15492 | Gene names | MYH3 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, fast skeletal muscle, embryonic (Muscle embryonic myosin heavy chain) (SMHCE). | |||||
|
TTF2_MOUSE
|
||||||
| θ value | 0.279714 (rank : 36) | NC score | 0.027323 (rank : 120) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5NC05, Q5M924 | Gene names | Ttf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 2 (EC 3.6.1.-) (RNA polymerase II termination factor) (Transcription release factor 2). | |||||
|
CEL_HUMAN
|
||||||
| θ value | 0.365318 (rank : 37) | NC score | 0.029180 (rank : 117) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
SPTN4_HUMAN
|
||||||
| θ value | 0.365318 (rank : 38) | NC score | 0.033585 (rank : 105) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 570 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9H254, Q9H1K7, Q9H1K8, Q9H1K9, Q9H3G8, Q9HCD0 | Gene names | SPTBN4, KIAA1642, SPTBN3 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 3 (Spectrin, non-erythroid beta chain 3) (Beta-IV spectrin). | |||||
|
CIC_MOUSE
|
||||||
| θ value | 0.47712 (rank : 39) | NC score | 0.047287 (rank : 87) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 742 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q924A2, Q6PDJ8, Q8CGE4, Q8CHH0, Q9CW61 | Gene names | Cic, Kiaa0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
LTBP3_HUMAN
|
||||||
| θ value | 0.47712 (rank : 40) | NC score | 0.030038 (rank : 113) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 502 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9NS15, O15107, Q96HB9, Q9H7K2, Q9UFN4 | Gene names | LTBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 3 precursor (LTBP-3). | |||||
|
MAP4_HUMAN
|
||||||
| θ value | 0.47712 (rank : 41) | NC score | 0.053575 (rank : 64) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 428 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P27816, Q13082, Q96A76 | Gene names | MAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
RRBP1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 42) | NC score | 0.031892 (rank : 109) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
CT2NL_MOUSE
|
||||||
| θ value | 0.62314 (rank : 43) | NC score | 0.041434 (rank : 94) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 902 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q99LJ0, Q6ZPR0, Q8BSV1 | Gene names | Cttnbp2nl, Kiaa1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
NOTC2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 44) | NC score | 0.039645 (rank : 97) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1113 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q04721, Q99734, Q9H240 | Gene names | NOTCH2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 2 precursor (Notch 2) (hN2) [Contains: Notch 2 extracellular truncation; Notch 2 intracellular domain]. | |||||
|
OR2J1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 45) | NC score | 0.002127 (rank : 196) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 758 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9GZK6, Q9GZK1 | Gene names | OR2J1 | |||
|
Domain Architecture |
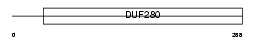
|
|||||
| Description | Olfactory receptor 2J1 (Olfactory receptor 6-5) (OR6-5) (Hs6M1-4). | |||||
|
KIF4A_MOUSE
|
||||||
| θ value | 0.813845 (rank : 46) | NC score | 0.024230 (rank : 135) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1033 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P33174 | Gene names | Kif4a, Kif4, Kns4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
LTBP4_MOUSE
|
||||||
| θ value | 0.813845 (rank : 47) | NC score | 0.025138 (rank : 127) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8K4G1, Q8K4G0 | Gene names | Ltbp4 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 4 precursor (LTBP-4). | |||||
|
AKAP2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 48) | NC score | 0.031211 (rank : 112) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9Y2D5, Q9UG26 | Gene names | AKAP2, KIAA0920 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2). | |||||
|
CENPF_HUMAN
|
||||||
| θ value | 1.06291 (rank : 49) | NC score | 0.025398 (rank : 126) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
CRSP7_HUMAN
|
||||||
| θ value | 1.06291 (rank : 50) | NC score | 0.045205 (rank : 90) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O95402 | Gene names | CRSP7, ARC70 | |||
|
Domain Architecture |

|
|||||
| Description | CRSP complex subunit 7 (Cofactor required for Sp1 transcriptional activation subunit 7) (Transcriptional coactivator CRSP70) (Activator- recruited cofactor 70 kDa component) (ARC70). | |||||
|
GRIN1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 51) | NC score | 0.040503 (rank : 96) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
MYCD_HUMAN
|
||||||
| θ value | 1.06291 (rank : 52) | NC score | 0.043569 (rank : 91) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8IZQ8, Q8N7Q1 | Gene names | MYOCD, MYCD | |||
|
Domain Architecture |

|
|||||
| Description | Myocardin. | |||||
|
NCOA6_HUMAN
|
||||||
| θ value | 1.06291 (rank : 53) | NC score | 0.054932 (rank : 60) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
SOX15_HUMAN
|
||||||
| θ value | 1.06291 (rank : 54) | NC score | 0.020263 (rank : 154) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60248, P35717, Q9Y6W7 | Gene names | SOX15, SOX12, SOX20, SOX26, SOX27 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-15 protein (SOX-20 protein) (SOX-12 protein). | |||||
|
ALMS1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 55) | NC score | 0.029786 (rank : 115) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K4E0, Q6A084, Q8C9N9 | Gene names | Alms1, Kiaa0328 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alstrom syndrome protein 1 homolog. | |||||
|
MILK2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 56) | NC score | 0.048181 (rank : 85) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8IY33, Q7RTP4, Q7Z655, Q8TEQ4, Q9H5F9 | Gene names | ||||
|
Domain Architecture |
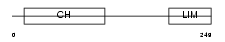
|
|||||
| Description | MICAL-like protein 2. | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 57) | NC score | 0.032456 (rank : 107) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
SHC2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 58) | NC score | 0.014417 (rank : 179) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BMC3 | Gene names | Shc2, Sck, ShcB | |||
|
Domain Architecture |

|
|||||
| Description | SHC-transforming protein 2 (SH2 domain protein C2) (Src homology 2 domain-containing-transforming protein C2) (Protein Sck) (Protein Sli). | |||||
|
FYCO1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 59) | NC score | 0.024126 (rank : 137) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8VDC1, Q3V385, Q7TMT0, Q8BJN2 | Gene names | Fyco1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1. | |||||
|
NELL2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 60) | NC score | 0.020938 (rank : 150) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 449 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99435 | Gene names | NELL2, NRP2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein NELL2 precursor (NEL-like protein 2) (Nel-related protein 2). | |||||
|
NELL2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 61) | NC score | 0.020892 (rank : 151) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61220 | Gene names | Nell2, Mel91 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein NELL2 precursor (NEL-like protein 2) (MEL91 protein). | |||||
|
NRX2A_HUMAN
|
||||||
| θ value | 1.81305 (rank : 62) | NC score | 0.022158 (rank : 147) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9P2S2, Q9Y2D6 | Gene names | NRXN2, KIAA0921 | |||
|
Domain Architecture |

|
|||||
| Description | Neurexin-2-alpha precursor (Neurexin II-alpha). | |||||
|
NRX2B_HUMAN
|
||||||
| θ value | 1.81305 (rank : 63) | NC score | 0.028960 (rank : 118) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 481 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P58401 | Gene names | NRXN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurexin-2-beta precursor (Neurexin II-beta). | |||||
|
SOX17_MOUSE
|
||||||
| θ value | 1.81305 (rank : 64) | NC score | 0.018862 (rank : 168) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61473, Q61472, Q62248 | Gene names | Sox17, Sox-17 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-17. | |||||
|
TTC16_HUMAN
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | 0.026951 (rank : 122) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8NEE8, Q5JU66, Q96M72 | Gene names | TTC16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 16 (TPR repeat protein 16). | |||||
|
VPS72_HUMAN
|
||||||
| θ value | 1.81305 (rank : 66) | NC score | 0.066822 (rank : 31) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15906, Q53GJ2, Q5U0R4 | Gene names | VPS72, TCFL1, YL1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
ASPH_HUMAN
|
||||||
| θ value | 2.36792 (rank : 67) | NC score | 0.029199 (rank : 116) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q12797 | Gene names | ASPH | |||
|
Domain Architecture |

|
|||||
| Description | Aspartyl/asparaginyl beta-hydroxylase (EC 1.14.11.16) (Aspartate beta- hydroxylase) (ASP beta-hydroxylase) (Peptide-aspartate beta- dioxygenase). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.049856 (rank : 83) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
CO002_HUMAN
|
||||||
| θ value | 2.36792 (rank : 69) | NC score | 0.047177 (rank : 88) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NZP6 | Gene names | C15orf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C15orf2. | |||||
|
ERRFI_HUMAN
|
||||||
| θ value | 2.36792 (rank : 70) | NC score | 0.024799 (rank : 130) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UJM3, Q9NTG9, Q9UD05 | Gene names | ERRFI1, MIG6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERBB receptor feedback inhibitor 1 (Mitogen-inducible gene 6 protein) (Mig-6). | |||||
|
RASF8_MOUSE
|
||||||
| θ value | 2.36792 (rank : 71) | NC score | 0.036024 (rank : 103) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8CJ96, Q8CCW4, Q9CU91 | Gene names | Rassf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras association domain-containing protein 8 (Carcinoma-associated protein HOJ-1 homolog). | |||||
|
ZF106_HUMAN
|
||||||
| θ value | 2.36792 (rank : 72) | NC score | 0.024841 (rank : 129) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H2Y7, Q6NSD9, Q6PEK1, Q86T43, Q86T45, Q86T50, Q86T58, Q86TA9, Q96M37, Q9H7B8 | Gene names | ZFP106, ZNF474 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 106 homolog (Zfp-106) (Zinc finger protein 474). | |||||
|
BTBD5_HUMAN
|
||||||
| θ value | 3.0926 (rank : 73) | NC score | 0.005004 (rank : 192) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NXS3 | Gene names | BTBD5 | |||
|
Domain Architecture |

|
|||||
| Description | BTB/POZ domain-containing protein 5. | |||||
|
CTGE5_MOUSE
|
||||||
| θ value | 3.0926 (rank : 74) | NC score | 0.024610 (rank : 132) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 928 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8R311, Q8CIE3 | Gene names | Ctage5, Mea6, Mgea6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cutaneous T-cell lymphoma-associated antigen 5 homolog (cTAGE-5 protein) (Meningioma-expressed antigen 6). | |||||
|
CUTL1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 75) | NC score | 0.028324 (rank : 119) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
FLNB_HUMAN
|
||||||
| θ value | 3.0926 (rank : 76) | NC score | 0.019329 (rank : 161) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75369, Q13706, Q8WXS9, Q8WXT0, Q8WXT1, Q8WXT2, Q9NRB5, Q9NT26, Q9UEV9 | Gene names | FLNB, FLN1L, FLN3, TABP, TAP | |||
|
Domain Architecture |

|
|||||
| Description | Filamin-B (FLN-B) (Beta-filamin) (Actin-binding-like protein) (Thyroid autoantigen) (Truncated actin-binding protein) (Truncated ABP) (ABP- 280 homolog) (ABP-278) (Filamin 3) (Filamin homolog 1) (Fh1). | |||||
|
MBB1A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 77) | NC score | 0.042364 (rank : 92) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
MYH10_HUMAN
|
||||||
| θ value | 3.0926 (rank : 78) | NC score | 0.020179 (rank : 155) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P35580, Q16087 | Gene names | MYH10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MYH10_MOUSE
|
||||||
| θ value | 3.0926 (rank : 79) | NC score | 0.020354 (rank : 153) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1612 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q61879, Q5SV63 | Gene names | Myh10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 3.0926 (rank : 80) | NC score | 0.063697 (rank : 38) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
NTNG1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 81) | NC score | 0.019120 (rank : 165) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Y2I2, Q5VU86, Q5VU87, Q5VU89, Q5VU90, Q5VU91, Q7Z2Y3, Q8N633 | Gene names | NTNG1, KIAA0976, LMNT1 | |||
|
Domain Architecture |
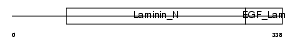
|
|||||
| Description | Netrin G1 precursor (Laminet-1). | |||||
|
PHAR3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 82) | NC score | 0.018792 (rank : 169) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BYK5, Q8BYS8, Q8C058, Q9DB87 | Gene names | Phactr3, Scapin1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatase and actin regulator 3 (Scapinin) (Scaffold-associated PP1- inhibiting protein). | |||||
|
PKHA6_HUMAN
|
||||||
| θ value | 3.0926 (rank : 83) | NC score | 0.023782 (rank : 140) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Y2H5 | Gene names | PLEKHA6, KIAA0969, PEPP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
RECQ4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 84) | NC score | 0.012453 (rank : 184) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q75NR7, Q76MT1, Q99PV9 | Gene names | Recql4, Recq4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent DNA helicase Q4 (EC 3.6.1.-) (RecQ protein-like 4). | |||||
|
SAPS1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 85) | NC score | 0.036500 (rank : 101) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UPN7, Q504V2, Q6NVJ6, Q9BU97 | Gene names | SAPS1, KIAA1115 | |||
|
Domain Architecture |

|
|||||
| Description | SAPS domain family member 1. | |||||
|
CBP_MOUSE
|
||||||
| θ value | 4.03905 (rank : 86) | NC score | 0.050161 (rank : 79) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
DACT1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 87) | NC score | 0.018313 (rank : 171) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8R4A3, Q80VG9, Q8BP49, Q9JK89 | Gene names | Dact1, Thyex3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dapper homolog 1 (MDpr1) (Frodo) (Thymus-expressed novel gene 3 protein). | |||||
|
GOGA4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 88) | NC score | 0.023792 (rank : 139) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
HS74L_HUMAN
|
||||||
| θ value | 4.03905 (rank : 89) | NC score | 0.014822 (rank : 176) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O95757, Q4W5M5, Q8IWA2 | Gene names | HSPA4L, APG1, OSP94 | |||
|
Domain Architecture |
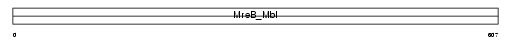
|
|||||
| Description | Heat shock 70 kDa protein 4L (Osmotic stress protein 94) (Heat shock 70-related protein APG-1). | |||||
|
IASPP_HUMAN
|
||||||
| θ value | 4.03905 (rank : 90) | NC score | 0.046385 (rank : 89) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8WUF5, Q2PNZ9, Q5DU71, Q5I1X4, Q6P1R7, Q6PKF8, Q9Y290 | Gene names | PPP1R13L, IASPP, NKIP1, PPP1R13BL, RAI | |||
|
Domain Architecture |
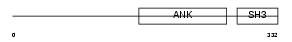
|
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
JAG1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 91) | NC score | 0.018906 (rank : 167) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9QXX0 | Gene names | Jag1 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-1 precursor (Jagged1) (CD339 antigen). | |||||
|
LEO1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 92) | NC score | 0.037256 (rank : 99) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
MNAB_MOUSE
|
||||||
| θ value | 4.03905 (rank : 93) | NC score | 0.023508 (rank : 142) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P0C090 | Gene names | Mnab | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated nucleic acid-binding protein. | |||||
|
MUC1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 94) | NC score | 0.038294 (rank : 98) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
OR2J3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 95) | NC score | -0.000030 (rank : 197) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 748 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O76001, Q96R15, Q9GZK5, Q9GZL4, Q9GZL5 | Gene names | OR2J3 | |||
|
Domain Architecture |
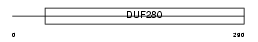
|
|||||
| Description | Olfactory receptor 2J3 (Olfactory receptor 6-6) (OR6-6) (Hs6M1-3). | |||||
|
STAB2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 96) | NC score | 0.019232 (rank : 162) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8R4U0, Q8BM87 | Gene names | Stab2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stabilin-2 precursor [Contains: Short form stabilin-2]. | |||||
|
SUHW4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 97) | NC score | 0.003428 (rank : 195) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 860 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6N043, Q6MZM6, Q6N085, Q9H0U5, Q9HCI8, Q9NXS0 | Gene names | SUHW4, KIAA1584 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppressor of hairy wing homolog 4. | |||||
|
SYNE1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 98) | NC score | 0.026316 (rank : 123) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
UBQL3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 99) | NC score | 0.012874 (rank : 183) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8C5U9 | Gene names | Ubqln3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquilin-3. | |||||
|
UT14A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 100) | NC score | 0.021682 (rank : 148) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9BVJ6, Q5JYF1 | Gene names | UTP14A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U3 small nucleolar RNA-associated protein 14 homolog A (Antigen NY-CO- 16). | |||||
|
WTAP_HUMAN
|
||||||
| θ value | 4.03905 (rank : 101) | NC score | 0.033831 (rank : 104) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q15007, Q96T28, Q9BYJ7, Q9H4E2 | Gene names | WTAP, KIAA0105 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wilms' tumor 1-associating protein (WT1-associated protein) (Putative pre-mRNA-splicing regulator female-lethal(2D) homolog). | |||||
|
ZN638_HUMAN
|
||||||
| θ value | 4.03905 (rank : 102) | NC score | 0.023999 (rank : 138) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q14966, Q53R34, Q5XJ05, Q68DP3, Q6P2H2, Q7Z3T7, Q8NF92, Q8TCA1, Q9H2G1, Q9NP37 | Gene names | ZNF638, NP220, ZFML | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein) (Cutaneous T-cell lymphoma-associated antigen se33-1) (CTCL tumor antigen se33-1). | |||||
|
FIBA_HUMAN
|
||||||
| θ value | 5.27518 (rank : 103) | NC score | 0.018927 (rank : 166) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P02671, Q9BX62, Q9UCH2 | Gene names | FGA | |||
|
Domain Architecture |

|
|||||
| Description | Fibrinogen alpha chain precursor [Contains: Fibrinopeptide A]. | |||||
|
JADE2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 104) | NC score | 0.040610 (rank : 95) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q6ZQF7, Q3UHD5, Q6IE83 | Gene names | Phf15, Jade2, Kiaa0239 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-2 (PHD finger protein 15). | |||||
|
JHD2B_HUMAN
|
||||||
| θ value | 5.27518 (rank : 105) | NC score | 0.013580 (rank : 182) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7LBC6, Q9BVH6, Q9BW93, Q9BZ52, Q9NYF4, Q9UPS0 | Gene names | JMJD1B, C5orf7, JHDM2B, KIAA1082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 2B (EC 1.14.11.-) (Jumonji domain-containing protein 1B) (Nuclear protein 5qNCA). | |||||
|
MCPH1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 106) | NC score | 0.019218 (rank : 163) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7TT79, Q8BNI9, Q8C9I7 | Gene names | Mcph1 | |||
|
Domain Architecture |

|
|||||
| Description | Microcephalin. | |||||
|
MYH11_MOUSE
|
||||||
| θ value | 5.27518 (rank : 107) | NC score | 0.019124 (rank : 164) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
NID1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 108) | NC score | 0.014585 (rank : 178) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P10493, Q8BQI3, Q8C3U8, Q8C9P6 | Gene names | Nid1, Ent | |||
|
Domain Architecture |

|
|||||
| Description | Nidogen-1 precursor (Entactin). | |||||
|
SENP2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 109) | NC score | 0.011588 (rank : 185) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91ZX6, Q544T8, Q9D4Z0 | Gene names | Senp2, Smt3ip2 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 2 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP2) (SUMO-1/Smt3-specific isopeptidase 2) (Smt3ip2) (Axam2). | |||||
|
SMC4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 110) | NC score | 0.019422 (rank : 160) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9NTJ3, O95752, Q8NDL4, Q9UNT9 | Gene names | SMC4L1, CAPC, SMC4 | |||
|
Domain Architecture |
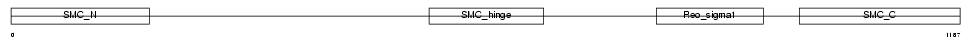
|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (hCAP-C) (XCAP-C homolog). | |||||
|
SSH3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 111) | NC score | 0.014305 (rank : 180) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8TE77, Q6PK42, Q76I75, Q8N9L8, Q8WYL0, Q9NV45, Q9NWZ7 | Gene names | SSH3, SSH3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase Slingshot homolog 3 (EC 3.1.3.48) (EC 3.1.3.16) (SSH-3L) (hSSH-3L). | |||||
|
TCGAP_MOUSE
|
||||||
| θ value | 5.27518 (rank : 112) | NC score | 0.017337 (rank : 174) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q80YF9, Q6P7W6 | Gene names | Snx26, Tcgap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TC10/CDC42 GTPase-activating protein (Sorting nexin-26). | |||||
|
TRIPB_HUMAN
|
||||||
| θ value | 5.27518 (rank : 113) | NC score | 0.024373 (rank : 134) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1587 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q15643, O14689, O15154, O95949 | Gene names | TRIP11, CEV14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid receptor-interacting protein 11 (TRIP-11) (Golgi-associated microtubule-binding protein 210) (GMAP-210) (Trip230) (Clonal evolution-related gene on chromosome 14). | |||||
|
UTRO_HUMAN
|
||||||
| θ value | 5.27518 (rank : 114) | NC score | 0.022674 (rank : 145) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P46939 | Gene names | UTRN, DMDL | |||
|
Domain Architecture |

|
|||||
| Description | Utrophin (Dystrophin-related protein 1) (DRP1) (DRP). | |||||
|
ANR11_HUMAN
|
||||||
| θ value | 6.88961 (rank : 115) | NC score | 0.053204 (rank : 65) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
APBA2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 116) | NC score | 0.006988 (rank : 190) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P98084, Q6PAJ2 | Gene names | Apba2, X11l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family A member 2 (Neuron- specific X11L protein) (Neuronal Munc18-1-interacting protein 2) (Mint-2) (Adapter protein X11beta). | |||||
|
BPA1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 117) | NC score | 0.024703 (rank : 131) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
BTBD5_MOUSE
|
||||||
| θ value | 6.88961 (rank : 118) | NC score | 0.003927 (rank : 194) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CR40, Q9CYV0 | Gene names | Btbd5 | |||
|
Domain Architecture |

|
|||||
| Description | BTB/POZ domain-containing protein 5. | |||||
|
CBP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 119) | NC score | 0.042224 (rank : 93) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
DIAP1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 120) | NC score | 0.029983 (rank : 114) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O60610, Q9UC76 | Gene names | DIAPH1, DIAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1). | |||||
|
EFS_MOUSE
|
||||||
| θ value | 6.88961 (rank : 121) | NC score | 0.022847 (rank : 144) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 389 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q64355 | Gene names | Efs, Sin | |||
|
Domain Architecture |
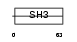
|
|||||
| Description | Embryonal Fyn-associated substrate (SRC-interacting protein) (Signal- integrating protein). | |||||
|
ITSN2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 122) | NC score | 0.010097 (rank : 187) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1371 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9Z0R6, Q9Z0R5 | Gene names | Itsn2, Ese2, Sh3d1B | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-2 (SH3 domain-containing protein 1B) (EH and SH3 domains protein 2) (EH domain and SH3 domain regulator of endocytosis 2). | |||||
|
JAG1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 123) | NC score | 0.017399 (rank : 173) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 514 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P78504, O14902, O15122, Q15816 | Gene names | JAG1, JAGL1 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-1 precursor (Jagged1) (hJ1) (CD339 antigen). | |||||
|
K1802_HUMAN
|
||||||
| θ value | 6.88961 (rank : 124) | NC score | 0.036044 (rank : 102) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
MAVS_MOUSE
|
||||||
| θ value | 6.88961 (rank : 125) | NC score | 0.020083 (rank : 157) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8VCF0 | Gene names | Mavs, Ips1, Visa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial antiviral signaling protein (Interferon-beta promoter stimulator protein 1) (IPS-1) (Virus-induced signaling adapter) (CARD adapter inducing interferon-beta) (Cardif). | |||||
|
MKL1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 126) | NC score | 0.036988 (rank : 100) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8K4J6 | Gene names | Mkl1, Bsac | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 1 (Myocardin-related transcription factor A) (MRTF-A) (Megakaryoblastic leukemia 1 protein homolog) (Basic SAP coiled-coil transcription activator). | |||||
|
MYH11_HUMAN
|
||||||
| θ value | 6.88961 (rank : 127) | NC score | 0.020107 (rank : 156) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
PDZD8_HUMAN
|
||||||
| θ value | 6.88961 (rank : 128) | NC score | 0.018709 (rank : 170) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8NEN9, Q86WE0, Q86WE5, Q9UFF1 | Gene names | PDZD8, PDZK8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 8 (Sarcoma antigen NY-SAR-84/NY-SAR- 104). | |||||
|
RPGR_HUMAN
|
||||||
| θ value | 6.88961 (rank : 129) | NC score | 0.010085 (rank : 188) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q92834, O00702, O00737, Q8N5T6, Q93039, Q9HD29, Q9UMR1 | Gene names | RPGR, RP3, XLRP3 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator. | |||||
|
RUFY2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 130) | NC score | 0.020463 (rank : 152) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1208 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8WXA3, Q96P51, Q9P1Z1 | Gene names | RUFY2, KIAA1537, RABIP4R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 2 (Rab4-interacting protein related). | |||||
|
RYBP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 131) | NC score | 0.019916 (rank : 158) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N488, Q9P2W5, Q9UMW4 | Gene names | RYBP, DEDAF, YEAF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING1 and YY1-binding protein (Death effector domain-associated factor) (DED-associated factor) (YY1 and E4TF1-associated factor 1) (Apoptin-associating protein 1) (APAP-1). | |||||
|
SOX30_MOUSE
|
||||||
| θ value | 6.88961 (rank : 132) | NC score | 0.031329 (rank : 111) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8CGW4 | Gene names | Sox30 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-30. | |||||
|
SSH1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 133) | NC score | 0.011167 (rank : 186) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8WYL5, Q6P6C0, Q8N9A7, Q8WYL3, Q8WYL4, Q9P2P8 | Gene names | SSH1, KIAA1298, SSH1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase Slingshot homolog 1 (EC 3.1.3.48) (EC 3.1.3.16) (SSH-1L) (hSSH-1L). | |||||
|
SYNE2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 134) | NC score | 0.027013 (rank : 121) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1323 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8WXH0, Q8N1S3, Q8NF49, Q8TER7, Q8WWW3, Q8WWW4, Q8WWW5, Q8WXH1, Q9NU50, Q9UFQ4, Q9Y2L4, Q9Y4R1 | Gene names | SYNE2, KIAA1011, NUA | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-2 (Nuclear envelope spectrin repeat protein 2) (Syne-2) (Synaptic nuclear envelope protein 2) (Nucleus and actin connecting element protein) (Protein NUANCE). | |||||
|
TMPS2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 135) | NC score | 0.004655 (rank : 193) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O15393, Q9BXX1 | Gene names | TMPRSS2, PRSS10 | |||
|
Domain Architecture |
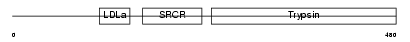
|
|||||
| Description | Transmembrane protease, serine 2 precursor (EC 3.4.21.-) [Contains: Transmembrane protease, serine 2 non-catalytic chain; Transmembrane protease, serine 2 catalytic chain]. | |||||
|
AZI1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 136) | NC score | 0.021382 (rank : 149) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1000 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UPN4, Q96F50 | Gene names | AZI1, CEP131, KIAA1118 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1) (Centrosomal protein of 131 kDa) (Cep131 protein). | |||||
|
CO020_HUMAN
|
||||||
| θ value | 8.99809 (rank : 137) | NC score | 0.024408 (rank : 133) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H611 | Gene names | C15orf20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf20. | |||||
|
CT2NL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 138) | NC score | 0.031618 (rank : 110) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9P2B4, Q96B40 | Gene names | CTTNBP2NL, KIAA1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
HMMR_HUMAN
|
||||||
| θ value | 8.99809 (rank : 139) | NC score | 0.017622 (rank : 172) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1082 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O75330, Q92767 | Gene names | HMMR, IHABP, RHAMM | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronan mediated motility receptor (Intracellular hyaluronic acid- binding protein) (Receptor for hyaluronan-mediated motility) (CD168 antigen). | |||||
|
LAMC2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 140) | NC score | 0.013861 (rank : 181) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q61092 | Gene names | Lamc2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-2 chain precursor (Laminin 5 gamma 2 subunit) (Kalinin/nicein/epiligrin 100 kDa subunit) (Laminin B2t chain). | |||||
|
LPP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 141) | NC score | 0.023658 (rank : 141) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q93052, Q8NFX5 | Gene names | LPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipoma-preferred partner (LIM domain-containing preferred translocation partner in lipoma). | |||||
|
MY18A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 142) | NC score | 0.019449 (rank : 159) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1989 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9JMH9, Q811D7 | Gene names | Myo18a, Myspdz | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
NFRKB_MOUSE
|
||||||
| θ value | 8.99809 (rank : 143) | NC score | 0.025811 (rank : 125) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q6PIJ4, Q8BWV5, Q8K0X6 | Gene names | Nfrkb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear factor related to kappa-B-binding protein (DNA-binding protein R kappa-B). | |||||
|
NTNG1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 144) | NC score | 0.014675 (rank : 177) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8R4G0, Q68FE5, Q69ZU3, Q8R4F3, Q8R4F4, Q8R4F5, Q8R4F6, Q8R4F7, Q8R4F8, Q8R4F9, Q9ESR3, Q9ESR4, Q9ESR5, Q9ESR6, Q9ESR7, Q9ESR8 | Gene names | Ntng1, Kiaa0976, Lmnt1 | |||
|
Domain Architecture |
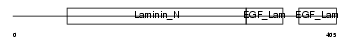
|
|||||
| Description | Netrin G1 precursor (Laminet-1). | |||||
|
RTP3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 145) | NC score | 0.049385 (rank : 84) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q5QGU6, Q8VDC2 | Gene names | Rtp3, Tmem7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor-transporting protein 3 (Transmembrane protein 7). | |||||
|
RYBP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 146) | NC score | 0.016441 (rank : 175) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CCI5, Q9WVK2 | Gene names | Rybp, Dedaf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING1 and YY1-binding protein (Death effector domain-associated factor) (DED-associated factor). | |||||
|
SMCE1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 147) | NC score | 0.024213 (rank : 136) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O54941, Q8BPD9 | Gene names | Smarce1, Baf57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator chromatin subfamily E member 1 (BRG1-associated factor 57). | |||||
|
SRBP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 148) | NC score | 0.022894 (rank : 143) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
TRI62_HUMAN
|
||||||
| θ value | 8.99809 (rank : 149) | NC score | 0.006286 (rank : 191) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9BVG3, Q9NVG0 | Gene names | TRIM62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 62. | |||||
|
UTX_HUMAN
|
||||||
| θ value | 8.99809 (rank : 150) | NC score | 0.009578 (rank : 189) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O15550 | Gene names | UTX | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitously transcribed X chromosome tetratricopeptide repeat protein (Ubiquitously transcribed TPR protein on the X chromosome). | |||||
|
ANKR6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.050012 (rank : 82) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y2G4, Q9NU24, Q9UFQ9 | Gene names | ANKRD6, KIAA0957 | |||
|
Domain Architecture |
|
|||||
| Description | Ankyrin repeat domain-containing protein 6. | |||||
|
ANR22_HUMAN
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.054035 (rank : 62) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5VYY1, Q8WU06 | Gene names | ANKRD22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 22. | |||||
|
ANR49_MOUSE
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.050091 (rank : 81) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8VE42, Q9Z2S1 | Gene names | Ankrd49, Fgif, Gbif | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 49 (Fetal globin-inducing factor) (Fetal globin-increasing factor). | |||||
|
BASP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.053760 (rank : 63) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
CBX1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.062384 (rank : 43) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P83916, P23197 | Gene names | CBX1, CBX | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 1 (Heterochromatin protein 1 homolog beta) (HP1 beta) (Modifier 1 protein) (M31) (Heterochromatin protein p25) (HP1Hsbeta) (p25beta). | |||||
|
CBX1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.062384 (rank : 44) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P83917, P23197 | Gene names | Cbx1, Cbx | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 1 (Heterochromatin protein 1 homolog beta) (HP1 beta) (Modifier 1 protein) (M31) (Heterochromatin protein p25). | |||||
|
CBX3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.063829 (rank : 37) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13185, Q96CD7, Q99409, Q9BVS3, Q9P0Z6 | Gene names | CBX3 | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 3 (Heterochromatin protein 1 homolog gamma) (HP1 gamma) (Modifier 2 protein) (HECH). | |||||
|
CBX3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.064469 (rank : 35) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P23198, Q921C4 | Gene names | Cbx3 | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 3 (Heterochromatin protein 1 homolog gamma) (HP1 gamma) (Modifier 2 protein) (M32). | |||||
|
CBX5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.059348 (rank : 50) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P45973 | Gene names | CBX5, HP1A | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 5 (Heterochromatin protein 1 homolog alpha) (HP1 alpha) (Antigen p25). | |||||
|
CBX5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.058901 (rank : 52) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61686, Q9CS35, Q9CXD1 | Gene names | Cbx5, Hp1a | |||
|
Domain Architecture |
|
|||||
| Description | Chromobox protein homolog 5 (Heterochromatin protein 1 homolog alpha) (HP1 alpha). | |||||
|
CSKI1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.050500 (rank : 76) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 997 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8WXD9, Q9P2P0 | Gene names | CASKIN1, KIAA1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
CTTB2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.050105 (rank : 80) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1061 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8WZ74, O43389, Q7LG11, Q9C0A5 | Gene names | CTTNBP2, C7orf8, CORTBP2, KIAA1758 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cortactin-binding protein 2 (CortBP2). | |||||
|
DPF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 163) | NC score | 0.055271 (rank : 59) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92782 | Gene names | DPF1, NEUD4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
GSCR1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 164) | NC score | 0.053201 (rank : 66) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
INVS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 165) | NC score | 0.050462 (rank : 77) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O89019, O88849 | Gene names | Invs, Inv, Nphp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning protein) (Nephrocystin-2). | |||||
|
LAD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 166) | NC score | 0.052393 (rank : 69) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P57016 | Gene names | Lad1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (Linear IgA disease autoantigen). | |||||
|
MARCS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 167) | NC score | 0.050509 (rank : 75) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P29966, Q2LA83, Q5TDB7 | Gene names | MARCKS, MACS, PRKCSL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS) (Protein kinase C substrate, 80 kDa protein, light chain) (PKCSL) (80K-L protein). | |||||
|
MDC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 168) | NC score | 0.059806 (rank : 48) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
MDC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 169) | NC score | 0.065955 (rank : 33) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
MPP8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 170) | NC score | 0.066082 (rank : 32) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q99549, Q86TK3, Q9BTP1 | Gene names | MPHOSPH8, MPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | M-phase phosphoprotein 8. | |||||
|
MYST3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 171) | NC score | 0.051796 (rank : 71) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
MYST4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 172) | NC score | 0.052489 (rank : 68) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
PHF10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 173) | NC score | 0.064442 (rank : 36) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8WUB8, Q2YDA3, Q9BXD2, Q9NV26 | Gene names | PHF10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10 (XAP135). | |||||
|
PHF10_MOUSE
|
||||||
| θ value | θ > 10 (rank : 174) | NC score | 0.062466 (rank : 42) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9D8M7, Q99LV5 | Gene names | Phf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10. | |||||
|
PHF11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 175) | NC score | 0.085380 (rank : 26) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UIL8, Q9Y5A2 | Gene names | PHF11, BCAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 11 (BRCA1 C-terminus-associated protein) (NY-REN-34 antigen). | |||||
|
PHF6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 176) | NC score | 0.065293 (rank : 34) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IWS0, Q96JK3, Q9BRU0 | Gene names | PHF6, KIAA1823 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 6 (PHD-like zinc finger protein). | |||||
|
PHF6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 177) | NC score | 0.063163 (rank : 40) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D4J7, Q80T86, Q8BYT8, Q8C1B5 | Gene names | Phf6, Kiaa1823 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 6. | |||||
|
PHF7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 178) | NC score | 0.051629 (rank : 72) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BWX1 | Gene names | PHF7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 7 (Testis development protein NYD-SP6). | |||||
|
PRP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 179) | NC score | 0.056982 (rank : 55) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
PRP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 180) | NC score | 0.059378 (rank : 49) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
PRR12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 181) | NC score | 0.051900 (rank : 70) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
RAI1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 182) | NC score | 0.061269 (rank : 45) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q7Z5J4, Q8N3B4, Q8ND08, Q8WU64, Q96JK5, Q9H1C1, Q9H1C2, Q9UF69 | Gene names | RAI1, KIAA1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
RAI1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 183) | NC score | 0.068838 (rank : 30) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61818, Q6ZPH7, Q8CEV1 | Gene names | Rai1, Kiaa1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
RSF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 184) | NC score | 0.050432 (rank : 78) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
SETD7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 185) | NC score | 0.060133 (rank : 47) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8WTS6, Q9C0E6 | Gene names | SETD7, KIAA1717, SET7 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET7 (EC 2.1.1.43) (Histone H3-K4 methyltransferase) (H3-K4-HMTase) (SET domain-containing protein 7) (Set9) (SET7/9). | |||||
|
SETD7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 186) | NC score | 0.061225 (rank : 46) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8VHL1 | Gene names | Setd7, Set7 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET7 (EC 2.1.1.43) (Histone H3-K4 methyltransferase) (H3-K4-HMTase) (SET domain-containing protein 7). | |||||
|
SETD8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 187) | NC score | 0.312043 (rank : 25) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9NQR1, Q86W83, Q8TD09 | Gene names | SETD8, PRSET7, SET07, SET8 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8) (PR/SET domain-containing protein 07) (PR/SET07) (PR-Set7). | |||||
|
SETD8_MOUSE
|
||||||
| θ value | θ > 10 (rank : 188) | NC score | 0.313260 (rank : 24) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q2YDW7, Q8C0J9 | Gene names | SETD8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8). | |||||
|
SHAN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 189) | NC score | 0.055285 (rank : 58) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Y566, Q9NYW9 | Gene names | SHANK1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 1 (Shank1) (Somatostatin receptor-interacting protein) (SSTR-interacting protein) (SSTRIP). | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 190) | NC score | 0.054675 (rank : 61) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 191) | NC score | 0.059074 (rank : 51) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
TCF20_HUMAN
|
||||||
| θ value | θ > 10 (rank : 192) | NC score | 0.063062 (rank : 41) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UGU0, O14528, Q13078, Q4V353, Q9H4M0 | Gene names | TCF20, KIAA0292, SPBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP) (AR1). | |||||
|
TCF20_MOUSE
|
||||||
| θ value | θ > 10 (rank : 193) | NC score | 0.057375 (rank : 53) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9EPQ8, Q60792 | Gene names | Tcf20, Spbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP). | |||||
|
TXND2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 194) | NC score | 0.051148 (rank : 74) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
WASIP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 195) | NC score | 0.052956 (rank : 67) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
WASIP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 196) | NC score | 0.056687 (rank : 56) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
WIRE_HUMAN
|
||||||
| θ value | θ > 10 (rank : 197) | NC score | 0.051215 (rank : 73) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8TF74, Q658J8, Q71RE1, Q8TE44 | Gene names | WIRE, WICH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein) (WIP- and CR16-homologous protein). | |||||
|
SETB1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 150 | |
| SwissProt Accessions | O88974, Q922K1 | Gene names | Setdb1, Eset | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 4 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 4) (H3-K9-HMTase 4) (SET domain bifurcated 1) (ERG-associated protein with SET domain) (ESET). | |||||
|
SETB1_HUMAN
|
||||||
| NC score | 0.977838 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q15047, Q96GM9 | Gene names | SETDB1, KIAA0067 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 4 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 4) (H3-K9-HMTase 4) (SET domain bifurcated 1) (ERG-associated protein with SET domain) (ESET). | |||||
|
SETB2_HUMAN
|
||||||
| NC score | 0.857298 (rank : 3) | θ value | 3.35642e-63 (rank : 3) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q96T68, Q86UD6, Q96AI6 | Gene names | SETDB2, CLLD8 | |||
|
Domain Architecture |

|
|||||
| Description | Probable histone-lysine N-methyltransferase, H3 lysine-9 specific (EC 2.1.1.43) (Histone H3-K9 methyltransferase) (H3-K9-HMTase) (SET domain bifurcated 2) (Chronic lymphocytic leukemia deletion region gene 8 protein). | |||||
|
SUV91_HUMAN
|
||||||
| NC score | 0.654981 (rank : 4) | θ value | 1.2077e-20 (rank : 7) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O43463, Q53G60, Q6FHK6 | Gene names | SUV39H1, SUV39H | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 1 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 1) (H3-K9-HMTase 1) (Suppressor of variegation 3-9 homolog 1) (Su(var)3-9 homolog 1). | |||||
|
SUV91_MOUSE
|
||||||
| NC score | 0.646271 (rank : 5) | θ value | 5.99374e-20 (rank : 8) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O54864, Q9JLC7, Q9JLP8 | Gene names | Suv39h1, Suv39h | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 1 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 1) (H3-K9-HMTase 1) (Suppressor of variegation 3-9 homolog 1) (Su(var)3-9 homolog 1) (Position-effect variegation 3-9 homolog). | |||||
|
SUV92_MOUSE
|
||||||
| NC score | 0.628047 (rank : 6) | θ value | 2.35696e-16 (rank : 9) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9EQQ0, Q9CUK3, Q9JLP7 | Gene names | Suv39h2 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 2 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 2) (H3-K9-HMTase 2) (Suppressor of variegation 3-9 homolog 2) (Su(var)3-9 homolog 2). | |||||
|
SUV92_HUMAN
|
||||||
| NC score | 0.627124 (rank : 7) | θ value | 3.40345e-15 (rank : 10) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9H5I1 | Gene names | SUV39H2 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 2 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 2) (H3-K9-HMTase 2) (Suppressor of variegation 3-9 homolog 2) (Su(var)3-9 homolog 2). | |||||
|
SETD2_HUMAN
|
||||||
| NC score | 0.520230 (rank : 8) | θ value | 1.9326e-10 (rank : 12) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
EZH1_HUMAN
|
||||||
| NC score | 0.474638 (rank : 9) | θ value | 1.29631e-06 (rank : 20) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q92800, O43287, Q14459 | Gene names | EZH1, KIAA0388 | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of zeste homolog 1 (ENX-2). | |||||
|
EZH1_MOUSE
|
||||||
| NC score | 0.474207 (rank : 10) | θ value | 1.29631e-06 (rank : 21) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P70351, Q9R089 | Gene names | Ezh1, Enx2 | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of zeste homolog 1 (ENX-2). | |||||
|
EZH2_HUMAN
|
||||||
| NC score | 0.469258 (rank : 11) | θ value | 6.43352e-06 (rank : 22) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q15910, Q15755, Q92857 | Gene names | EZH2 | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of zeste homolog 2 (ENX-1). | |||||
|
SET1A_HUMAN
|
||||||
| NC score | 0.464990 (rank : 12) | θ value | 1.99992e-07 (rank : 19) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
EZH2_MOUSE
|
||||||
| NC score | 0.461803 (rank : 13) | θ value | 3.19293e-05 (rank : 23) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q61188, Q9R090 | Gene names | Ezh2, Enx1h | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of zeste homolog 2 (ENX-1). | |||||
|
MLL4_HUMAN
|
||||||
| NC score | 0.436026 (rank : 14) | θ value | 3.89403e-11 (rank : 11) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.429521 (rank : 15) | θ value | 9.59137e-10 (rank : 13) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
NSD1_HUMAN
|
||||||
| NC score | 0.429519 (rank : 16) | θ value | 3.08544e-08 (rank : 16) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
NSD1_MOUSE
|
||||||
| NC score | 0.427981 (rank : 17) | θ value | 3.08544e-08 (rank : 17) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
EHMT2_HUMAN
|
||||||
| NC score | 0.361204 (rank : 18) | θ value | 1.05554e-24 (rank : 4) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q96KQ7, Q14349, Q6PK06, Q96MH5, Q96QD0, Q9UQL8, Q9Y331 | Gene names | EHMT2, BAT8, G9A, NG36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
EHMT2_MOUSE
|
||||||
| NC score | 0.359751 (rank : 19) | θ value | 4.01107e-24 (rank : 5) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9Z148, Q6PE08, Q8K4R6, Q8K4R7, Q9Z149 | Gene names | Ehmt2, Bat8, G9a, Ng36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
EHMT1_HUMAN
|
||||||
| NC score | 0.352339 (rank : 20) | θ value | 1.16704e-23 (rank : 6) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9H9B1, Q8TCN7, Q96F53, Q96JF1, Q96KH4 | Gene names | EHMT1, EUHMTASE1, KIAA1876 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 5 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 5) (H3-K9-HMTase 5) (Euchromatic histone-lysine N-methyltransferase 1) (Eu-HMTase1) (G9a- like protein 1) (GLP1). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.342439 (rank : 21) | θ value | 1.63604e-09 (rank : 14) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 87 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.328323 (rank : 22) | θ value | 6.87365e-08 (rank : 18) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.315423 (rank : 23) | θ value | 3.08544e-08 (rank : 15) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
SETD8_MOUSE
|
||||||
| NC score | 0.313260 (rank : 24) | θ value | θ > 10 (rank : 188) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q2YDW7, Q8C0J9 | Gene names | SETD8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8). | |||||
|
SETD8_HUMAN
|
||||||
| NC score | 0.312043 (rank : 25) | θ value | θ > 10 (rank : 187) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9NQR1, Q86W83, Q8TD09 | Gene names | SETD8, PRSET7, SET07, SET8 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8) (PR/SET domain-containing protein 07) (PR/SET07) (PR-Set7). | |||||
|
PHF11_HUMAN
|
||||||
| NC score | 0.085380 (rank : 26) | θ value | θ > 10 (rank : 175) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UIL8, Q9Y5A2 | Gene names | PHF11, BCAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 11 (BRCA1 C-terminus-associated protein) (NY-REN-34 antigen). | |||||
|
VPS72_MOUSE
|
||||||
| NC score | 0.082024 (rank : 27) | θ value | 0.21417 (rank : 34) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q62481, Q3U2N4, Q810A9, Q99K81 | Gene names | Vps72, Tcfl1, Yl1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
BAZ2B_HUMAN
|
||||||
| NC score | 0.076518 (rank : 28) | θ value | 0.21417 (rank : 29) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 803 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9UIF8, Q96EA1, Q96SQ8, Q9P252, Q9Y4N8 | Gene names | BAZ2B, KIAA1476 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2B (hWALp4). | |||||
|
TARSH_HUMAN
|
||||||
| NC score | 0.073849 (rank : 29) | θ value | 0.21417 (rank : 33) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q7Z7G0, Q6ZW20, Q6ZW22, Q9C082, Q9UFI6 | Gene names | ABI3BP, NESHBP, TARSH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Target of Nesh-SH3 precursor (Tarsh) (Nesh-binding protein) (NeshBP) (ABI gene family member 3-binding protein). | |||||
|
RAI1_MOUSE
|
||||||
| NC score | 0.068838 (rank : 30) | θ value | θ > 10 (rank : 183) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61818, Q6ZPH7, Q8CEV1 | Gene names | Rai1, Kiaa1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
VPS72_HUMAN
|
||||||
| NC score | 0.066822 (rank : 31) | θ value | 1.81305 (rank : 66) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15906, Q53GJ2, Q5U0R4 | Gene names | VPS72, TCFL1, YL1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
MPP8_HUMAN
|
||||||
| NC score | 0.066082 (rank : 32) | θ value | θ > 10 (rank : 170) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q99549, Q86TK3, Q9BTP1 | Gene names | MPHOSPH8, MPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | M-phase phosphoprotein 8. | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.065955 (rank : 33) | θ value | θ > 10 (rank : 169) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
PHF6_HUMAN
|
||||||
| NC score | 0.065293 (rank : 34) | θ value | θ > 10 (rank : 176) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IWS0, Q96JK3, Q9BRU0 | Gene names | PHF6, KIAA1823 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 6 (PHD-like zinc finger protein). | |||||
|
CBX3_MOUSE
|
||||||
| NC score | 0.064469 (rank : 35) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P23198, Q921C4 | Gene names | Cbx3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 3 (Heterochromatin protein 1 homolog gamma) (HP1 gamma) (Modifier 2 protein) (M32). | |||||
|
PHF10_HUMAN
|
||||||
| NC score | 0.064442 (rank : 36) | θ value | θ > 10 (rank : 173) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8WUB8, Q2YDA3, Q9BXD2, Q9NV26 | Gene names | PHF10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10 (XAP135). | |||||
|
CBX3_HUMAN
|
||||||
| NC score | 0.063829 (rank : 37) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13185, Q96CD7, Q99409, Q9BVS3, Q9P0Z6 | Gene names | CBX3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 3 (Heterochromatin protein 1 homolog gamma) (HP1 gamma) (Modifier 2 protein) (HECH). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.063697 (rank : 38) | θ value | 3.0926 (rank : 80) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.063321 (rank : 39) | θ value | 0.163984 (rank : 27) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
PHF6_MOUSE
|
||||||
| NC score | 0.063163 (rank : 40) | θ value | θ > 10 (rank : 177) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D4J7, Q80T86, Q8BYT8, Q8C1B5 | Gene names | Phf6, Kiaa1823 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 6. | |||||
|
TCF20_HUMAN
|
||||||
| NC score | 0.063062 (rank : 41) | θ value | θ > 10 (rank : 192) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UGU0, O14528, Q13078, Q4V353, Q9H4M0 | Gene names | TCF20, KIAA0292, SPBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP) (AR1). | |||||
|
PHF10_MOUSE
|
||||||
| NC score | 0.062466 (rank : 42) | θ value | θ > 10 (rank : 174) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9D8M7, Q99LV5 | Gene names | Phf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10. | |||||
|
CBX1_HUMAN
|
||||||
| NC score | 0.062384 (rank : 43) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P83916, P23197 | Gene names | CBX1, CBX | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 1 (Heterochromatin protein 1 homolog beta) (HP1 beta) (Modifier 1 protein) (M31) (Heterochromatin protein p25) (HP1Hsbeta) (p25beta). | |||||
|
CBX1_MOUSE
|
||||||
| NC score | 0.062384 (rank : 44) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P83917, P23197 | Gene names | Cbx1, Cbx | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 1 (Heterochromatin protein 1 homolog beta) (HP1 beta) (Modifier 1 protein) (M31) (Heterochromatin protein p25). | |||||
|
RAI1_HUMAN
|
||||||
| NC score | 0.061269 (rank : 45) | θ value | θ > 10 (rank : 182) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q7Z5J4, Q8N3B4, Q8ND08, Q8WU64, Q96JK5, Q9H1C1, Q9H1C2, Q9UF69 | Gene names | RAI1, KIAA1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
SETD7_MOUSE
|
||||||
| NC score | 0.061225 (rank : 46) | θ value | θ > 10 (rank : 186) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8VHL1 | Gene names | Setd7, Set7 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET7 (EC 2.1.1.43) (Histone H3-K4 methyltransferase) (H3-K4-HMTase) (SET domain-containing protein 7). | |||||
|
SETD7_HUMAN
|
||||||
| NC score | 0.060133 (rank : 47) | θ value | θ > 10 (rank : 185) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8WTS6, Q9C0E6 | Gene names | SETD7, KIAA1717, SET7 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET7 (EC 2.1.1.43) (Histone H3-K4 methyltransferase) (H3-K4-HMTase) (SET domain-containing protein 7) (Set9) (SET7/9). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.059806 (rank : 48) | θ value | θ > 10 (rank : 168) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
PRP2_MOUSE
|
||||||
| NC score | 0.059378 (rank : 49) | θ value | θ > 10 (rank : 180) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
CBX5_HUMAN
|
||||||
| NC score | 0.059348 (rank : 50) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P45973 | Gene names | CBX5, HP1A | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 5 (Heterochromatin protein 1 homolog alpha) (HP1 alpha) (Antigen p25). | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.059074 (rank : 51) | θ value | θ > 10 (rank : 191) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
CBX5_MOUSE
|
||||||
| NC score | 0.058901 (rank : 52) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61686, Q9CS35, Q9CXD1 | Gene names | Cbx5, Hp1a | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 5 (Heterochromatin protein 1 homolog alpha) (HP1 alpha). | |||||
|
TCF20_MOUSE
|
||||||
| NC score | 0.057375 (rank : 53) | θ value | θ > 10 (rank : 193) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9EPQ8, Q60792 | Gene names | Tcf20, Spbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP). | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.057271 (rank : 54) | θ value | 0.125558 (rank : 26) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
PRP1_HUMAN
|
||||||
| NC score | 0.056982 (rank : 55) | θ value | θ > 10 (rank : 179) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
WASIP_MOUSE
|
||||||
| NC score | 0.056687 (rank : 56) | θ value | θ > 10 (rank : 196) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
PI5PA_HUMAN
|
||||||
| NC score | 0.056478 (rank : 57) | θ value | 0.125558 (rank : 25) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 749 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q15735, Q32M61, Q6ZTH6, Q8N902, Q9UDT9 | Gene names | PIB5PA, PIPP | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase A (EC 3.1.3.56). | |||||
|
SHAN1_HUMAN
|
||||||
| NC score | 0.055285 (rank : 58) | θ value | θ > 10 (rank : 189) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Y566, Q9NYW9 | Gene names | SHANK1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 1 (Shank1) (Somatostatin receptor-interacting protein) (SSTR-interacting protein) (SSTRIP). | |||||
|
DPF1_HUMAN
|
||||||
| NC score | 0.055271 (rank : 59) | θ value | θ > 10 (rank : 163) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92782 | Gene names | DPF1, NEUD4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
NCOA6_HUMAN
|
||||||
| NC score | 0.054932 (rank : 60) | θ value | 1.06291 (rank : 53) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.054675 (rank : 61) | θ value | θ > 10 (rank : 190) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
ANR22_HUMAN
|
||||||
| NC score | 0.054035 (rank : 62) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5VYY1, Q8WU06 | Gene names | ANKRD22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 22. | |||||
|
BASP_HUMAN
|
||||||
| NC score | 0.053760 (rank : 63) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
MAP4_HUMAN
|
||||||
| NC score | 0.053575 (rank : 64) | θ value | 0.47712 (rank : 41) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 428 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P27816, Q13082, Q96A76 | Gene names | MAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
ANR11_HUMAN
|
||||||
| NC score | 0.053204 (rank : 65) | θ value | 6.88961 (rank : 115) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
GSCR1_HUMAN
|
||||||
| NC score | 0.053201 (rank : 66) | θ value | θ > 10 (rank : 164) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
WASIP_HUMAN
|
||||||
| NC score | 0.052956 (rank : 67) | θ value | θ > 10 (rank : 195) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
MYST4_HUMAN
|
||||||
| NC score | 0.052489 (rank : 68) | θ value | θ > 10 (rank : 172) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
LAD1_MOUSE
|
||||||
| NC score | 0.052393 (rank : 69) | θ value | θ > 10 (rank : 166) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P57016 | Gene names | Lad1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (Linear IgA disease autoantigen). | |||||
|
PRR12_HUMAN
|
||||||
| NC score | 0.051900 (rank : 70) | θ value | θ > 10 (rank : 181) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
MYST3_HUMAN
|
||||||
| NC score | 0.051796 (rank : 71) | θ value | θ > 10 (rank : 171) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
PHF7_HUMAN
|
||||||
| NC score | 0.051629 (rank : 72) | θ value | θ > 10 (rank : 178) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BWX1 | Gene names | PHF7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 7 (Testis development protein NYD-SP6). | |||||
|
WIRE_HUMAN
|
||||||
| NC score | 0.051215 (rank : 73) | θ value | θ > 10 (rank : 197) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8TF74, Q658J8, Q71RE1, Q8TE44 | Gene names | WIRE, WICH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein) (WIP- and CR16-homologous protein). | |||||
|
TXND2_HUMAN
|
||||||
| NC score | 0.051148 (rank : 74) | θ value | θ > 10 (rank : 194) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
MARCS_HUMAN
|
||||||
| NC score | 0.050509 (rank : 75) | θ value | θ > 10 (rank : 167) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P29966, Q2LA83, Q5TDB7 | Gene names | MARCKS, MACS, PRKCSL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS) (Protein kinase C substrate, 80 kDa protein, light chain) (PKCSL) (80K-L protein). | |||||
|
CSKI1_HUMAN
|
||||||
| NC score | 0.050500 (rank : 76) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 997 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8WXD9, Q9P2P0 | Gene names | CASKIN1, KIAA1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
INVS_MOUSE
|
||||||
| NC score | 0.050462 (rank : 77) | θ value | θ > 10 (rank : 165) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O89019, O88849 | Gene names | Invs, Inv, Nphp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning protein) (Nephrocystin-2). | |||||
|
RSF1_HUMAN
|
||||||
| NC score | 0.050432 (rank : 78) | θ value | θ > 10 (rank : 184) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
CBP_MOUSE
|
||||||
| NC score | 0.050161 (rank : 79) | θ value | 4.03905 (rank : 86) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CTTB2_HUMAN
|
||||||
| NC score | 0.050105 (rank : 80) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1061 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8WZ74, O43389, Q7LG11, Q9C0A5 | Gene names | CTTNBP2, C7orf8, CORTBP2, KIAA1758 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cortactin-binding protein 2 (CortBP2). | |||||
|
ANR49_MOUSE
|
||||||
| NC score | 0.050091 (rank : 81) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8VE42, Q9Z2S1 | Gene names | Ankrd49, Fgif, Gbif | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 49 (Fetal globin-inducing factor) (Fetal globin-increasing factor). | |||||
|
ANKR6_HUMAN
|
||||||
| NC score | 0.050012 (rank : 82) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y2G4, Q9NU24, Q9UFQ9 | Gene names | ANKRD6, KIAA0957 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat domain-containing protein 6. | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.049856 (rank : 83) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
RTP3_MOUSE
|
||||||
| NC score | 0.049385 (rank : 84) | θ value | 8.99809 (rank : 145) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q5QGU6, Q8VDC2 | Gene names | Rtp3, Tmem7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor-transporting protein 3 (Transmembrane protein 7). | |||||
|
MILK2_HUMAN
|
||||||
| NC score | 0.048181 (rank : 85) | θ value | 1.38821 (rank : 56) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8IY33, Q7RTP4, Q7Z655, Q8TEQ4, Q9H5F9 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | MICAL-like protein 2. | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.047707 (rank : 86) | θ value | 0.0563607 (rank : 24) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
CIC_MOUSE
|
||||||
| NC score | 0.047287 (rank : 87) | θ value | 0.47712 (rank : 39) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 742 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q924A2, Q6PDJ8, Q8CGE4, Q8CHH0, Q9CW61 | Gene names | Cic, Kiaa0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
CO002_HUMAN
|
||||||
| NC score | 0.047177 (rank : 88) | θ value | 2.36792 (rank : 69) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NZP6 | Gene names | C15orf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C15orf2. | |||||
|
IASPP_HUMAN
|
||||||
| NC score | 0.046385 (rank : 89) | θ value | 4.03905 (rank : 90) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8WUF5, Q2PNZ9, Q5DU71, Q5I1X4, Q6P1R7, Q6PKF8, Q9Y290 | Gene names | PPP1R13L, IASPP, NKIP1, PPP1R13BL, RAI | |||
|
Domain Architecture |

|
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
CRSP7_HUMAN
|
||||||
| NC score | 0.045205 (rank : 90) | θ value | 1.06291 (rank : 50) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O95402 | Gene names | CRSP7, ARC70 | |||
|
Domain Architecture |

|
|||||
| Description | CRSP complex subunit 7 (Cofactor required for Sp1 transcriptional activation subunit 7) (Transcriptional coactivator CRSP70) (Activator- recruited cofactor 70 kDa component) (ARC70). | |||||
|
MYCD_HUMAN
|
||||||
| NC score | 0.043569 (rank : 91) | θ value | 1.06291 (rank : 52) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8IZQ8, Q8N7Q1 | Gene names | MYOCD, MYCD | |||
|
Domain Architecture |

|
|||||
| Description | Myocardin. | |||||
|
MBB1A_HUMAN
|
||||||
| NC score | 0.042364 (rank : 92) | θ value | 3.0926 (rank : 77) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
CBP_HUMAN
|
||||||
| NC score | 0.042224 (rank : 93) | θ value | 6.88961 (rank : 119) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CT2NL_MOUSE
|
||||||
| NC score | 0.041434 (rank : 94) | θ value | 0.62314 (rank : 43) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 902 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q99LJ0, Q6ZPR0, Q8BSV1 | Gene names | Cttnbp2nl, Kiaa1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
JADE2_MOUSE
|
||||||
| NC score | 0.040610 (rank : 95) | θ value | 5.27518 (rank : 104) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q6ZQF7, Q3UHD5, Q6IE83 | Gene names | Phf15, Jade2, Kiaa0239 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-2 (PHD finger protein 15). | |||||
|
GRIN1_HUMAN
|
||||||
| NC score | 0.040503 (rank : 96) | θ value | 1.06291 (rank : 51) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
NOTC2_HUMAN
|
||||||
| NC score | 0.039645 (rank : 97) | θ value | 0.62314 (rank : 44) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1113 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q04721, Q99734, Q9H240 | Gene names | NOTCH2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 2 precursor (Notch 2) (hN2) [Contains: Notch 2 extracellular truncation; Notch 2 intracellular domain]. | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.038294 (rank : 98) | θ value | 4.03905 (rank : 94) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.037256 (rank : 99) | θ value | 4.03905 (rank : 92) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
MKL1_MOUSE
|
||||||
| NC score | 0.036988 (rank : 100) | θ value | 6.88961 (rank : 126) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8K4J6 | Gene names | Mkl1, Bsac | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 1 (Myocardin-related transcription factor A) (MRTF-A) (Megakaryoblastic leukemia 1 protein homolog) (Basic SAP coiled-coil transcription activator). | |||||
|
SAPS1_HUMAN
|
||||||
| NC score | 0.036500 (rank : 101) | θ value | 3.0926 (rank : 85) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UPN7, Q504V2, Q6NVJ6, Q9BU97 | Gene names | SAPS1, KIAA1115 | |||
|
Domain Architecture |

|
|||||
| Description | SAPS domain family member 1. | |||||
|
K1802_HUMAN
|
||||||
| NC score | 0.036044 (rank : 102) | θ value | 6.88961 (rank : 124) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
RASF8_MOUSE
|
||||||
| NC score | 0.036024 (rank : 103) | θ value | 2.36792 (rank : 71) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8CJ96, Q8CCW4, Q9CU91 | Gene names | Rassf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras association domain-containing protein 8 (Carcinoma-associated protein HOJ-1 homolog). | |||||
|
WTAP_HUMAN
|
||||||
| NC score | 0.033831 (rank : 104) | θ value | 4.03905 (rank : 101) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q15007, Q96T28, Q9BYJ7, Q9H4E2 | Gene names | WTAP, KIAA0105 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wilms' tumor 1-associating protein (WT1-associated protein) (Putative pre-mRNA-splicing regulator female-lethal(2D) homolog). | |||||
|
SPTN4_HUMAN
|
||||||
| NC score | 0.033585 (rank : 105) | θ value | 0.365318 (rank : 38) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 570 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9H254, Q9H1K7, Q9H1K8, Q9H1K9, Q9H3G8, Q9HCD0 | Gene names | SPTBN4, KIAA1642, SPTBN3 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 3 (Spectrin, non-erythroid beta chain 3) (Beta-IV spectrin). | |||||
|
LTBP3_MOUSE
|
||||||
| NC score | 0.032594 (rank : 106) | θ value | 0.21417 (rank : 32) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 475 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q61810, Q8BNQ6 | Gene names | Ltbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 3 precursor (LTBP-3). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.032456 (rank : 107) | θ value | 1.38821 (rank : 57) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
GOGA1_HUMAN
|
||||||
| NC score | 0.031949 (rank : 108) | θ value | 0.21417 (rank : 31) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1222 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q92805, Q8IYZ9 | Gene names | GOLGA1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 1 (Golgin-97). | |||||
|
RRBP1_MOUSE
|
||||||
| NC score | 0.031892 (rank : 109) | θ value | 0.47712 (rank : 42) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
CT2NL_HUMAN
|
||||||
| NC score | 0.031618 (rank : 110) | θ value | 8.99809 (rank : 138) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9P2B4, Q96B40 | Gene names | CTTNBP2NL, KIAA1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
SOX30_MOUSE
|
||||||
| NC score | 0.031329 (rank : 111) | θ value | 6.88961 (rank : 132) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8CGW4 | Gene names | Sox30 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-30. | |||||
|
AKAP2_HUMAN
|
||||||
| NC score | 0.031211 (rank : 112) | θ value | 1.06291 (rank : 48) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9Y2D5, Q9UG26 | Gene names | AKAP2, KIAA0920 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2). | |||||
|
LTBP3_HUMAN
|
||||||
| NC score | 0.030038 (rank : 113) | θ value | 0.47712 (rank : 40) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 502 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9NS15, O15107, Q96HB9, Q9H7K2, Q9UFN4 | Gene names | LTBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 3 precursor (LTBP-3). | |||||
|
DIAP1_HUMAN
|
||||||
| NC score | 0.029983 (rank : 114) | θ value | 6.88961 (rank : 120) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O60610, Q9UC76 | Gene names | DIAPH1, DIAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1). | |||||
|
ALMS1_MOUSE
|
||||||
| NC score | 0.029786 (rank : 115) | θ value | 1.38821 (rank : 55) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K4E0, Q6A084, Q8C9N9 | Gene names | Alms1, Kiaa0328 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alstrom syndrome protein 1 homolog. | |||||
|
ASPH_HUMAN
|
||||||
| NC score | 0.029199 (rank : 116) | θ value | 2.36792 (rank : 67) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q12797 | Gene names | ASPH | |||
|
Domain Architecture |

|
|||||
| Description | Aspartyl/asparaginyl beta-hydroxylase (EC 1.14.11.16) (Aspartate beta- hydroxylase) (ASP beta-hydroxylase) (Peptide-aspartate beta- dioxygenase). | |||||
|
CEL_HUMAN
|
||||||
| NC score | 0.029180 (rank : 117) | θ value | 0.365318 (rank : 37) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
NRX2B_HUMAN
|
||||||
| NC score | 0.028960 (rank : 118) | θ value | 1.81305 (rank : 63) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 481 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P58401 | Gene names | NRXN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurexin-2-beta precursor (Neurexin II-beta). | |||||
|
CUTL1_MOUSE
|
||||||
| NC score | 0.028324 (rank : 119) | θ value | 3.0926 (rank : 75) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
TTF2_MOUSE
|
||||||
| NC score | 0.027323 (rank : 120) | θ value | 0.279714 (rank : 36) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5NC05, Q5M924 | Gene names | Ttf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 2 (EC 3.6.1.-) (RNA polymerase II termination factor) (Transcription release factor 2). | |||||
|
SYNE2_HUMAN
|
||||||
| NC score | 0.027013 (rank : 121) | θ value | 6.88961 (rank : 134) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1323 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8WXH0, Q8N1S3, Q8NF49, Q8TER7, Q8WWW3, Q8WWW4, Q8WWW5, Q8WXH1, Q9NU50, Q9UFQ4, Q9Y2L4, Q9Y4R1 | Gene names | SYNE2, KIAA1011, NUA | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-2 (Nuclear envelope spectrin repeat protein 2) (Syne-2) (Synaptic nuclear envelope protein 2) (Nucleus and actin connecting element protein) (Protein NUANCE). | |||||
|
TTC16_HUMAN
|
||||||
| NC score | 0.026951 (rank : 122) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8NEE8, Q5JU66, Q96M72 | Gene names | TTC16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 16 (TPR repeat protein 16). | |||||
|
SYNE1_HUMAN
|
||||||
| NC score | 0.026316 (rank : 123) | θ value | 4.03905 (rank : 98) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
JPH3_HUMAN
|
||||||
| NC score | 0.025962 (rank : 124) | θ value | 0.163984 (rank : 28) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WXH2, Q8N471, Q9HDC3, Q9HDC4 | Gene names | JPH3, JP3 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-3 (Junctophilin type 3) (JP-3). | |||||
|
NFRKB_MOUSE
|
||||||
| NC score | 0.025811 (rank : 125) | θ value | 8.99809 (rank : 143) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q6PIJ4, Q8BWV5, Q8K0X6 | Gene names | Nfrkb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear factor related to kappa-B-binding protein (DNA-binding protein R kappa-B). | |||||
|
CENPF_HUMAN
|
||||||
| NC score | 0.025398 (rank : 126) | θ value | 1.06291 (rank : 49) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
LTBP4_MOUSE
|
||||||
| NC score | 0.025138 (rank : 127) | θ value | 0.813845 (rank : 47) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8K4G1, Q8K4G0 | Gene names | Ltbp4 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 4 precursor (LTBP-4). | |||||
|
MYH3_HUMAN
|
||||||
| NC score | 0.025067 (rank : 128) | θ value | 0.279714 (rank : 35) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1660 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P11055, Q15492 | Gene names | MYH3 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, fast skeletal muscle, embryonic (Muscle embryonic myosin heavy chain) (SMHCE). | |||||
|
ZF106_HUMAN
|
||||||
| NC score | 0.024841 (rank : 129) | θ value | 2.36792 (rank : 72) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H2Y7, Q6NSD9, Q6PEK1, Q86T43, Q86T45, Q86T50, Q86T58, Q86TA9, Q96M37, Q9H7B8 | Gene names | ZFP106, ZNF474 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 106 homolog (Zfp-106) (Zinc finger protein 474). | |||||
|
ERRFI_HUMAN
|
||||||
| NC score | 0.024799 (rank : 130) | θ value | 2.36792 (rank : 70) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UJM3, Q9NTG9, Q9UD05 | Gene names | ERRFI1, MIG6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERBB receptor feedback inhibitor 1 (Mitogen-inducible gene 6 protein) (Mig-6). | |||||
|
BPA1_MOUSE
|
||||||
| NC score | 0.024703 (rank : 131) | θ value | 6.88961 (rank : 117) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
CTGE5_MOUSE
|
||||||
| NC score | 0.024610 (rank : 132) | θ value | 3.0926 (rank : 74) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 928 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8R311, Q8CIE3 | Gene names | Ctage5, Mea6, Mgea6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cutaneous T-cell lymphoma-associated antigen 5 homolog (cTAGE-5 protein) (Meningioma-expressed antigen 6). | |||||
|
CO020_HUMAN
|
||||||
| NC score | 0.024408 (rank : 133) | θ value | 8.99809 (rank : 137) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H611 | Gene names | C15orf20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf20. | |||||
|
TRIPB_HUMAN
|
||||||
| NC score | 0.024373 (rank : 134) | θ value | 5.27518 (rank : 113) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1587 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q15643, O14689, O15154, O95949 | Gene names | TRIP11, CEV14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid receptor-interacting protein 11 (TRIP-11) (Golgi-associated microtubule-binding protein 210) (GMAP-210) (Trip230) (Clonal evolution-related gene on chromosome 14). | |||||
|
KIF4A_MOUSE
|
||||||
| NC score | 0.024230 (rank : 135) | θ value | 0.813845 (rank : 46) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1033 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P33174 | Gene names | Kif4a, Kif4, Kns4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
SMCE1_MOUSE
|
||||||
| NC score | 0.024213 (rank : 136) | θ value | 8.99809 (rank : 147) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O54941, Q8BPD9 | Gene names | Smarce1, Baf57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator chromatin subfamily E member 1 (BRG1-associated factor 57). | |||||
|
FYCO1_MOUSE
|
||||||
| NC score | 0.024126 (rank : 137) | θ value | 1.81305 (rank : 59) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8VDC1, Q3V385, Q7TMT0, Q8BJN2 | Gene names | Fyco1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1. | |||||
|
ZN638_HUMAN
|
||||||
| NC score | 0.023999 (rank : 138) | θ value | 4.03905 (rank : 102) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q14966, Q53R34, Q5XJ05, Q68DP3, Q6P2H2, Q7Z3T7, Q8NF92, Q8TCA1, Q9H2G1, Q9NP37 | Gene names | ZNF638, NP220, ZFML | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein) (Cutaneous T-cell lymphoma-associated antigen se33-1) (CTCL tumor antigen se33-1). | |||||
|
GOGA4_MOUSE
|
||||||
| NC score | 0.023792 (rank : 139) | θ value | 4.03905 (rank : 88) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
PKHA6_HUMAN
|
||||||
| NC score | 0.023782 (rank : 140) | θ value | 3.0926 (rank : 83) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Y2H5 | Gene names | PLEKHA6, KIAA0969, PEPP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
LPP_HUMAN
|
||||||
| NC score | 0.023658 (rank : 141) | θ value | 8.99809 (rank : 141) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q93052, Q8NFX5 | Gene names | LPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipoma-preferred partner (LIM domain-containing preferred translocation partner in lipoma). | |||||
|
MNAB_MOUSE
|
||||||
| NC score | 0.023508 (rank : 142) | θ value | 4.03905 (rank : 93) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P0C090 | Gene names | Mnab | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated nucleic acid-binding protein. | |||||
|
SRBP1_MOUSE
|
||||||
| NC score | 0.022894 (rank : 143) | θ value | 8.99809 (rank : 148) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
EFS_MOUSE
|
||||||
| NC score | 0.022847 (rank : 144) | θ value | 6.88961 (rank : 121) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 389 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q64355 | Gene names | Efs, Sin | |||
|
Domain Architecture |

|
|||||
| Description | Embryonal Fyn-associated substrate (SRC-interacting protein) (Signal- integrating protein). | |||||
|
UTRO_HUMAN
|
||||||
| NC score | 0.022674 (rank : 145) | θ value | 5.27518 (rank : 114) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P46939 | Gene names | UTRN, DMDL | |||
|
Domain Architecture |

|
|||||
| Description | Utrophin (Dystrophin-related protein 1) (DRP1) (DRP). | |||||
|
EPAS1_MOUSE
|
||||||
| NC score | 0.022215 (rank : 146) | θ value | 0.21417 (rank : 30) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
|
NRX2A_HUMAN
|
||||||
| NC score | 0.022158 (rank : 147) | θ value | 1.81305 (rank : 62) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9P2S2, Q9Y2D6 | Gene names | NRXN2, KIAA0921 | |||
|
Domain Architecture |

|
|||||
| Description | Neurexin-2-alpha precursor (Neurexin II-alpha). | |||||
|
UT14A_HUMAN
|
||||||
| NC score | 0.021682 (rank : 148) | θ value | 4.03905 (rank : 100) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9BVJ6, Q5JYF1 | Gene names | UTP14A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U3 small nucleolar RNA-associated protein 14 homolog A (Antigen NY-CO- 16). | |||||
|
AZI1_HUMAN
|
||||||
| NC score | 0.021382 (rank : 149) | θ value | 8.99809 (rank : 136) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1000 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UPN4, Q96F50 | Gene names | AZI1, CEP131, KIAA1118 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1) (Centrosomal protein of 131 kDa) (Cep131 protein). | |||||
|
NELL2_HUMAN
|
||||||
| NC score | 0.020938 (rank : 150) | θ value | 1.81305 (rank : 60) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 449 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99435 | Gene names | NELL2, NRP2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein NELL2 precursor (NEL-like protein 2) (Nel-related protein 2). | |||||
|
NELL2_MOUSE
|
||||||
| NC score | 0.020892 (rank : 151) | θ value | 1.81305 (rank : 61) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61220 | Gene names | Nell2, Mel91 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein NELL2 precursor (NEL-like protein 2) (MEL91 protein). | |||||
|
RUFY2_HUMAN
|
||||||
| NC score | 0.020463 (rank : 152) | θ value | 6.88961 (rank : 130) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1208 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8WXA3, Q96P51, Q9P1Z1 | Gene names | RUFY2, KIAA1537, RABIP4R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 2 (Rab4-interacting protein related). | |||||
|
MYH10_MOUSE
|
||||||
| NC score | 0.020354 (rank : 153) | θ value | 3.0926 (rank : 79) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1612 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q61879, Q5SV63 | Gene names | Myh10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
SOX15_HUMAN
|
||||||
| NC score | 0.020263 (rank : 154) | θ value | 1.06291 (rank : 54) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60248, P35717, Q9Y6W7 | Gene names | SOX15, SOX12, SOX20, SOX26, SOX27 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-15 protein (SOX-20 protein) (SOX-12 protein). | |||||
|
MYH10_HUMAN
|
||||||
| NC score | 0.020179 (rank : 155) | θ value | 3.0926 (rank : 78) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P35580, Q16087 | Gene names | MYH10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MYH11_HUMAN
|
||||||
| NC score | 0.020107 (rank : 156) | θ value | 6.88961 (rank : 127) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
MAVS_MOUSE
|
||||||
| NC score | 0.020083 (rank : 157) | θ value | 6.88961 (rank : 125) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8VCF0 | Gene names | Mavs, Ips1, Visa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial antiviral signaling protein (Interferon-beta promoter stimulator protein 1) (IPS-1) (Virus-induced signaling adapter) (CARD adapter inducing interferon-beta) (Cardif). | |||||
|
RYBP_HUMAN
|
||||||
| NC score | 0.019916 (rank : 158) | θ value | 6.88961 (rank : 131) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N488, Q9P2W5, Q9UMW4 | Gene names | RYBP, DEDAF, YEAF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING1 and YY1-binding protein (Death effector domain-associated factor) (DED-associated factor) (YY1 and E4TF1-associated factor 1) (Apoptin-associating protein 1) (APAP-1). | |||||
|
MY18A_MOUSE
|
||||||
| NC score | 0.019449 (rank : 159) | θ value | 8.99809 (rank : 142) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1989 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9JMH9, Q811D7 | Gene names | Myo18a, Myspdz | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
SMC4_HUMAN
|
||||||
| NC score | 0.019422 (rank : 160) | θ value | 5.27518 (rank : 110) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9NTJ3, O95752, Q8NDL4, Q9UNT9 | Gene names | SMC4L1, CAPC, SMC4 | |||
|
Domain Architecture |
|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (hCAP-C) (XCAP-C homolog). | |||||
|
FLNB_HUMAN
|
||||||
| NC score | 0.019329 (rank : 161) | θ value | 3.0926 (rank : 76) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75369, Q13706, Q8WXS9, Q8WXT0, Q8WXT1, Q8WXT2, Q9NRB5, Q9NT26, Q9UEV9 | Gene names | FLNB, FLN1L, FLN3, TABP, TAP | |||
|
Domain Architecture |

|
|||||
| Description | Filamin-B (FLN-B) (Beta-filamin) (Actin-binding-like protein) (Thyroid autoantigen) (Truncated actin-binding protein) (Truncated ABP) (ABP- 280 homolog) (ABP-278) (Filamin 3) (Filamin homolog 1) (Fh1). | |||||
|
STAB2_MOUSE
|
||||||
| NC score | 0.019232 (rank : 162) | θ value | 4.03905 (rank : 96) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8R4U0, Q8BM87 | Gene names | Stab2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stabilin-2 precursor [Contains: Short form stabilin-2]. | |||||
|
MCPH1_MOUSE
|
||||||
| NC score | 0.019218 (rank : 163) | θ value | 5.27518 (rank : 106) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7TT79, Q8BNI9, Q8C9I7 | Gene names | Mcph1 | |||
|
Domain Architecture |

|
|||||
| Description | Microcephalin. | |||||
|
MYH11_MOUSE
|
||||||
| NC score | 0.019124 (rank : 164) | θ value | 5.27518 (rank : 107) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
NTNG1_HUMAN
|
||||||
| NC score | 0.019120 (rank : 165) | θ value | 3.0926 (rank : 81) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Y2I2, Q5VU86, Q5VU87, Q5VU89, Q5VU90, Q5VU91, Q7Z2Y3, Q8N633 | Gene names | NTNG1, KIAA0976, LMNT1 | |||
|
Domain Architecture |

|
|||||
| Description | Netrin G1 precursor (Laminet-1). | |||||
|
FIBA_HUMAN
|
||||||
| NC score | 0.018927 (rank : 166) | θ value | 5.27518 (rank : 103) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P02671, Q9BX62, Q9UCH2 | Gene names | FGA | |||
|
Domain Architecture |

|
|||||
| Description | Fibrinogen alpha chain precursor [Contains: Fibrinopeptide A]. | |||||
|
JAG1_MOUSE
|
||||||
| NC score | 0.018906 (rank : 167) | θ value | 4.03905 (rank : 91) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9QXX0 | Gene names | Jag1 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-1 precursor (Jagged1) (CD339 antigen). | |||||
|
SOX17_MOUSE
|
||||||
| NC score | 0.018862 (rank : 168) | θ value | 1.81305 (rank : 64) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61473, Q61472, Q62248 | Gene names | Sox17, Sox-17 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-17. | |||||
|
PHAR3_MOUSE
|
||||||
| NC score | 0.018792 (rank : 169) | θ value | 3.0926 (rank : 82) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BYK5, Q8BYS8, Q8C058, Q9DB87 | Gene names | Phactr3, Scapin1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatase and actin regulator 3 (Scapinin) (Scaffold-associated PP1- inhibiting protein). | |||||
|
PDZD8_HUMAN
|
||||||
| NC score | 0.018709 (rank : 170) | θ value | 6.88961 (rank : 128) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8NEN9, Q86WE0, Q86WE5, Q9UFF1 | Gene names | PDZD8, PDZK8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 8 (Sarcoma antigen NY-SAR-84/NY-SAR- 104). | |||||
|
DACT1_MOUSE
|
||||||
| NC score | 0.018313 (rank : 171) | θ value | 4.03905 (rank : 87) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8R4A3, Q80VG9, Q8BP49, Q9JK89 | Gene names | Dact1, Thyex3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dapper homolog 1 (MDpr1) (Frodo) (Thymus-expressed novel gene 3 protein). | |||||
|
HMMR_HUMAN
|
||||||
| NC score | 0.017622 (rank : 172) | θ value | 8.99809 (rank : 139) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1082 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O75330, Q92767 | Gene names | HMMR, IHABP, RHAMM | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronan mediated motility receptor (Intracellular hyaluronic acid- binding protein) (Receptor for hyaluronan-mediated motility) (CD168 antigen). | |||||
|
JAG1_HUMAN
|
||||||
| NC score | 0.017399 (rank : 173) | θ value | 6.88961 (rank : 123) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 514 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P78504, O14902, O15122, Q15816 | Gene names | JAG1, JAGL1 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-1 precursor (Jagged1) (hJ1) (CD339 antigen). | |||||
|
TCGAP_MOUSE
|
||||||
| NC score | 0.017337 (rank : 174) | θ value | 5.27518 (rank : 112) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q80YF9, Q6P7W6 | Gene names | Snx26, Tcgap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TC10/CDC42 GTPase-activating protein (Sorting nexin-26). | |||||
|
RYBP_MOUSE
|
||||||
| NC score | 0.016441 (rank : 175) | θ value | 8.99809 (rank : 146) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CCI5, Q9WVK2 | Gene names | Rybp, Dedaf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING1 and YY1-binding protein (Death effector domain-associated factor) (DED-associated factor). | |||||
|
HS74L_HUMAN
|
||||||
| NC score | 0.014822 (rank : 176) | θ value | 4.03905 (rank : 89) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O95757, Q4W5M5, Q8IWA2 | Gene names | HSPA4L, APG1, OSP94 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock 70 kDa protein 4L (Osmotic stress protein 94) (Heat shock 70-related protein APG-1). | |||||
|
NTNG1_MOUSE
|
||||||
| NC score | 0.014675 (rank : 177) | θ value | 8.99809 (rank : 144) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8R4G0, Q68FE5, Q69ZU3, Q8R4F3, Q8R4F4, Q8R4F5, Q8R4F6, Q8R4F7, Q8R4F8, Q8R4F9, Q9ESR3, Q9ESR4, Q9ESR5, Q9ESR6, Q9ESR7, Q9ESR8 | Gene names | Ntng1, Kiaa0976, Lmnt1 | |||
|
Domain Architecture |

|
|||||
| Description | Netrin G1 precursor (Laminet-1). | |||||
|
NID1_MOUSE
|
||||||
| NC score | 0.014585 (rank : 178) | θ value | 5.27518 (rank : 108) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P10493, Q8BQI3, Q8C3U8, Q8C9P6 | Gene names | Nid1, Ent | |||
|
Domain Architecture |

|
|||||
| Description | Nidogen-1 precursor (Entactin). | |||||
|
SHC2_MOUSE
|
||||||
| NC score | 0.014417 (rank : 179) | θ value | 1.38821 (rank : 58) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BMC3 | Gene names | Shc2, Sck, ShcB | |||
|
Domain Architecture |

|
|||||
| Description | SHC-transforming protein 2 (SH2 domain protein C2) (Src homology 2 domain-containing-transforming protein C2) (Protein Sck) (Protein Sli). | |||||
|
SSH3_HUMAN
|
||||||
| NC score | 0.014305 (rank : 180) | θ value | 5.27518 (rank : 111) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8TE77, Q6PK42, Q76I75, Q8N9L8, Q8WYL0, Q9NV45, Q9NWZ7 | Gene names | SSH3, SSH3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase Slingshot homolog 3 (EC 3.1.3.48) (EC 3.1.3.16) (SSH-3L) (hSSH-3L). | |||||
|
LAMC2_MOUSE
|
||||||
| NC score | 0.013861 (rank : 181) | θ value | 8.99809 (rank : 140) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q61092 | Gene names | Lamc2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-2 chain precursor (Laminin 5 gamma 2 subunit) (Kalinin/nicein/epiligrin 100 kDa subunit) (Laminin B2t chain). | |||||
|
JHD2B_HUMAN
|
||||||
| NC score | 0.013580 (rank : 182) | θ value | 5.27518 (rank : 105) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7LBC6, Q9BVH6, Q9BW93, Q9BZ52, Q9NYF4, Q9UPS0 | Gene names | JMJD1B, C5orf7, JHDM2B, KIAA1082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 2B (EC 1.14.11.-) (Jumonji domain-containing protein 1B) (Nuclear protein 5qNCA). | |||||
|
UBQL3_MOUSE
|
||||||
| NC score | 0.012874 (rank : 183) | θ value | 4.03905 (rank : 99) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8C5U9 | Gene names | Ubqln3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquilin-3. | |||||
|
RECQ4_MOUSE
|
||||||
| NC score | 0.012453 (rank : 184) | θ value | 3.0926 (rank : 84) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q75NR7, Q76MT1, Q99PV9 | Gene names | Recql4, Recq4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent DNA helicase Q4 (EC 3.6.1.-) (RecQ protein-like 4). | |||||
|
SENP2_MOUSE
|
||||||
| NC score | 0.011588 (rank : 185) | θ value | 5.27518 (rank : 109) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91ZX6, Q544T8, Q9D4Z0 | Gene names | Senp2, Smt3ip2 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 2 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP2) (SUMO-1/Smt3-specific isopeptidase 2) (Smt3ip2) (Axam2). | |||||
|
SSH1_HUMAN
|
||||||
| NC score | 0.011167 (rank : 186) | θ value | 6.88961 (rank : 133) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8WYL5, Q6P6C0, Q8N9A7, Q8WYL3, Q8WYL4, Q9P2P8 | Gene names | SSH1, KIAA1298, SSH1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase Slingshot homolog 1 (EC 3.1.3.48) (EC 3.1.3.16) (SSH-1L) (hSSH-1L). | |||||
|
ITSN2_MOUSE
|
||||||
| NC score | 0.010097 (rank : 187) | θ value | 6.88961 (rank : 122) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 1371 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9Z0R6, Q9Z0R5 | Gene names | Itsn2, Ese2, Sh3d1B | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-2 (SH3 domain-containing protein 1B) (EH and SH3 domains protein 2) (EH domain and SH3 domain regulator of endocytosis 2). | |||||
|
RPGR_HUMAN
|
||||||
| NC score | 0.010085 (rank : 188) | θ value | 6.88961 (rank : 129) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q92834, O00702, O00737, Q8N5T6, Q93039, Q9HD29, Q9UMR1 | Gene names | RPGR, RP3, XLRP3 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator. | |||||
|
UTX_HUMAN
|
||||||
| NC score | 0.009578 (rank : 189) | θ value | 8.99809 (rank : 150) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O15550 | Gene names | UTX | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitously transcribed X chromosome tetratricopeptide repeat protein (Ubiquitously transcribed TPR protein on the X chromosome). | |||||
|
APBA2_MOUSE
|
||||||
| NC score | 0.006988 (rank : 190) | θ value | 6.88961 (rank : 116) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P98084, Q6PAJ2 | Gene names | Apba2, X11l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family A member 2 (Neuron- specific X11L protein) (Neuronal Munc18-1-interacting protein 2) (Mint-2) (Adapter protein X11beta). | |||||
|
TRI62_HUMAN
|
||||||
| NC score | 0.006286 (rank : 191) | θ value | 8.99809 (rank : 149) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9BVG3, Q9NVG0 | Gene names | TRIM62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 62. | |||||
|
BTBD5_HUMAN
|
||||||
| NC score | 0.005004 (rank : 192) | θ value | 3.0926 (rank : 73) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NXS3 | Gene names | BTBD5 | |||
|
Domain Architecture |

|
|||||
| Description | BTB/POZ domain-containing protein 5. | |||||
|
TMPS2_HUMAN
|
||||||
| NC score | 0.004655 (rank : 193) | θ value | 6.88961 (rank : 135) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O15393, Q9BXX1 | Gene names | TMPRSS2, PRSS10 | |||
|
Domain Architecture |

|
|||||
| Description | Transmembrane protease, serine 2 precursor (EC 3.4.21.-) [Contains: Transmembrane protease, serine 2 non-catalytic chain; Transmembrane protease, serine 2 catalytic chain]. | |||||
|
BTBD5_MOUSE
|
||||||
| NC score | 0.003927 (rank : 194) | θ value | 6.88961 (rank : 118) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CR40, Q9CYV0 | Gene names | Btbd5 | |||
|
Domain Architecture |

|
|||||
| Description | BTB/POZ domain-containing protein 5. | |||||
|
SUHW4_HUMAN
|
||||||
| NC score | 0.003428 (rank : 195) | θ value | 4.03905 (rank : 97) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 860 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6N043, Q6MZM6, Q6N085, Q9H0U5, Q9HCI8, Q9NXS0 | Gene names | SUHW4, KIAA1584 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppressor of hairy wing homolog 4. | |||||
|
OR2J1_HUMAN
|
||||||
| NC score | 0.002127 (rank : 196) | θ value | 0.62314 (rank : 45) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 758 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9GZK6, Q9GZK1 | Gene names | OR2J1 | |||
|
Domain Architecture |

|
|||||
| Description | Olfactory receptor 2J1 (Olfactory receptor 6-5) (OR6-5) (Hs6M1-4). | |||||
|
OR2J3_HUMAN
|
||||||
| NC score | -0.000030 (rank : 197) | θ value | 4.03905 (rank : 95) | |||
| Query Neighborhood Hits | 150 | Target Neighborhood Hits | 748 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O76001, Q96R15, Q9GZK5, Q9GZL4, Q9GZL5 | Gene names | OR2J3 | |||
|
Domain Architecture |

|
|||||
| Description | Olfactory receptor 2J3 (Olfactory receptor 6-6) (OR6-6) (Hs6M1-3). | |||||