Please be patient as the page loads
|
PLXA3_HUMAN
|
||||||
| SwissProt Accessions | P51805 | Gene names | PLXNA3, PLXN4, SEX | |||
|
Domain Architecture |

|
|||||
| Description | Plexin-A3 precursor (Plexin-4) (Semaphorin receptor SEX). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PLXA1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.996098 (rank : 2) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q9UIW2 | Gene names | PLXNA1, NOV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A1 precursor (Semaphorin receptor NOV). | |||||
|
PLXA1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.995738 (rank : 3) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | P70206, Q5DTR0 | Gene names | Plxna1, KIAA4053 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A1 precursor (Plexin-1) (Plex 1). | |||||
|
PLXA2_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.995070 (rank : 4) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | O75051, Q6UX61, Q96GN9, Q9BRL1, Q9UIW1 | Gene names | PLXNA2, KIAA0463, OCT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A2 precursor (Semaphorin receptor OCT). | |||||
|
PLXA2_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.994859 (rank : 5) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | P70207, Q6NVE6, Q6PHN4, Q80TZ7, Q8R1I4 | Gene names | Plxna2, Kiaa0463 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A2 precursor (Plexin-2) (Plex 2). | |||||
|
PLXA3_HUMAN
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 102 | |
| SwissProt Accessions | P51805 | Gene names | PLXNA3, PLXN4, SEX | |||
|
Domain Architecture |

|
|||||
| Description | Plexin-A3 precursor (Plexin-4) (Semaphorin receptor SEX). | |||||
|
PLXA4_HUMAN
|
||||||
| θ value | 0 (rank : 6) | NC score | 0.993841 (rank : 7) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q9HCM2, Q6UWC6, Q6ZW89, Q8N969, Q8ND00, Q8NEN3, Q9NTD4 | Gene names | PLXNA4, KIAA1550, PLXNA4A, PLXNA4B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A4 precursor. | |||||
|
PLXA4_MOUSE
|
||||||
| θ value | 0 (rank : 7) | NC score | 0.994150 (rank : 6) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q80UG2, Q5DTW8, Q8BKK9 | Gene names | Plxna4, Kiaa1550 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A4 precursor. | |||||
|
PLXB1_HUMAN
|
||||||
| θ value | 0 (rank : 8) | NC score | 0.942922 (rank : 13) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | O43157, Q6NY20, Q9UIV7, Q9UJ92, Q9UJ93 | Gene names | PLXNB1, KIAA0407, SEP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B1 precursor (Semaphorin receptor SEP). | |||||
|
PLXB1_MOUSE
|
||||||
| θ value | 0 (rank : 9) | NC score | 0.953066 (rank : 12) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q8CJH3, Q6ZQC3, Q80ZZ1 | Gene names | Plxnb1, Kiaa0407 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B1 precursor. | |||||
|
PLXB2_HUMAN
|
||||||
| θ value | 0 (rank : 10) | NC score | 0.972879 (rank : 9) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | O15031, Q7KZU3, Q9BSU7 | Gene names | PLXNB2, KIAA0315 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B2 precursor (MM1). | |||||
|
PLXB3_HUMAN
|
||||||
| θ value | 0 (rank : 11) | NC score | 0.969545 (rank : 10) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9ULL4, Q9HDA4 | Gene names | PLXNB3, KIAA1206 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B3 precursor. | |||||
|
PLXB3_MOUSE
|
||||||
| θ value | 0 (rank : 12) | NC score | 0.968590 (rank : 11) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9QY40, Q80TH8 | Gene names | Plxnb3, Kiaa1206, Plxn6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B3 precursor (Plexin-6). | |||||
|
PLXD1_HUMAN
|
||||||
| θ value | 0 (rank : 13) | NC score | 0.974430 (rank : 8) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9Y4D7, Q6PJS9, Q8IZJ2, Q9BTQ2 | Gene names | PLXND1, KIAA0620 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-D1 precursor. | |||||
|
PLXC1_HUMAN
|
||||||
| θ value | 2.5344e-111 (rank : 14) | NC score | 0.925531 (rank : 15) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | O60486, Q59H25 | Gene names | PLXNC1, VESPR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-C1 precursor (Virus-encoded semaphorin protein receptor) (CD232 antigen). | |||||
|
PLXC1_MOUSE
|
||||||
| θ value | 9.63072e-111 (rank : 15) | NC score | 0.927467 (rank : 14) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9QZC2, Q8CGW1 | Gene names | Plxnc1, Vespr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-C1 precursor (Virus-encoded semaphorin protein receptor) (CD232 antigen). | |||||
|
MET_MOUSE
|
||||||
| θ value | 1.08979e-29 (rank : 16) | NC score | 0.206334 (rank : 55) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 844 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P16056, Q62125 | Gene names | Met | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte growth factor receptor precursor (EC 2.7.10.1) (HGF receptor) (Scatter factor receptor) (SF receptor) (HGF/SF receptor) (Met proto-oncogene tyrosine kinase) (c-Met). | |||||
|
MET_HUMAN
|
||||||
| θ value | 5.40856e-29 (rank : 17) | NC score | 0.206852 (rank : 54) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 831 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P08581, O60366, Q12875, Q9UDX7, Q9UPL8 | Gene names | MET | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte growth factor receptor precursor (EC 2.7.10.1) (HGF receptor) (Scatter factor receptor) (SF receptor) (HGF/SF receptor) (Met proto-oncogene tyrosine kinase) (c-Met). | |||||
|
RON_MOUSE
|
||||||
| θ value | 7.30988e-26 (rank : 18) | NC score | 0.180933 (rank : 58) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q62190, Q62555 | Gene names | Mst1r, Ron, Stk | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage-stimulating protein receptor precursor (EC 2.7.10.1) (MSP receptor) (p185-Ron) (Stem cell-derived tyrosine kinase) (CDw136 antigen) [Contains: Macrophage-stimulating protein receptor alpha chain; Macrophage-stimulating protein receptor beta chain]. | |||||
|
RON_HUMAN
|
||||||
| θ value | 2.12685e-25 (rank : 19) | NC score | 0.183462 (rank : 56) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q04912 | Gene names | MST1R, RON | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage-stimulating protein receptor precursor (EC 2.7.10.1) (MSP receptor) (p185-Ron) (CDw136 antigen) [Contains: Macrophage- stimulating protein receptor alpha chain; Macrophage-stimulating protein receptor beta chain]. | |||||
|
SEM4A_HUMAN
|
||||||
| θ value | 3.18553e-13 (rank : 20) | NC score | 0.406337 (rank : 17) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9H3S1, Q8WUA9 | Gene names | SEMA4A, SEMB | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4A precursor (Semaphorin B) (Sema B). | |||||
|
SEM4C_HUMAN
|
||||||
| θ value | 1.02475e-11 (rank : 21) | NC score | 0.379442 (rank : 20) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9C0C4, Q7Z5X0 | Gene names | SEMA4C, KIAA1739 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4C precursor. | |||||
|
SEM4A_MOUSE
|
||||||
| θ value | 3.89403e-11 (rank : 22) | NC score | 0.400311 (rank : 19) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q62178 | Gene names | Sema4a, Semab, SemB | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4A precursor (Semaphorin B) (Sema B). | |||||
|
SEM4B_MOUSE
|
||||||
| θ value | 2.79066e-09 (rank : 23) | NC score | 0.412617 (rank : 16) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q62179, Q4PKI6, Q69ZB7 | Gene names | Sema4b, Kiaa1745, Semac, SemC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-4B precursor (Semaphorin C) (Sema C). | |||||
|
SEM4G_MOUSE
|
||||||
| θ value | 3.64472e-09 (rank : 24) | NC score | 0.367820 (rank : 21) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9WUH7 | Gene names | Sema4g | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4G precursor. | |||||
|
SEM4B_HUMAN
|
||||||
| θ value | 6.21693e-09 (rank : 25) | NC score | 0.401799 (rank : 18) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9NPR2, Q6UXE3, Q8WVP9, Q96FK5, Q9C0B8, Q9H691, Q9NPM8, Q9NPN0 | Gene names | SEMA4B, KIAA1745 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4B precursor. | |||||
|
SEM4G_HUMAN
|
||||||
| θ value | 2.36244e-08 (rank : 26) | NC score | 0.366795 (rank : 22) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9NTN9, Q9HCF3 | Gene names | SEMA4G, KIAA1619 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4G precursor. | |||||
|
SEM4C_MOUSE
|
||||||
| θ value | 4.0297e-08 (rank : 27) | NC score | 0.347606 (rank : 24) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q64151 | Gene names | Sema4c, Semacl1, Semai | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4C precursor (Semaphorin I) (Sema I) (Semaphorin C-like 1) (M-Sema F). | |||||
|
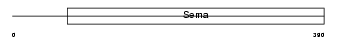
SEM7A_MOUSE
|
||||||
| θ value | 4.45536e-07 (rank : 28) | NC score | 0.352582 (rank : 23) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9QUR8, O88371 | Gene names | Sema7a, Cd108, Semal, Semk1 | |||
|
Domain Architecture |
|
|||||
| Description | Semaphorin-7A precursor (Semaphorin L) (Sema L) (Semaphorin K1) (Sema K1) (CD108 antigen) (CDw108). | |||||
|
SEM5A_MOUSE
|
||||||
| θ value | 9.92553e-07 (rank : 29) | NC score | 0.319209 (rank : 38) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q62217 | Gene names | Sema5a, Semaf, SemF | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-5A precursor (Semaphorin F) (Sema F). | |||||
|
SEM6D_HUMAN
|
||||||
| θ value | 1.69304e-06 (rank : 30) | NC score | 0.343729 (rank : 26) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8NFY4, Q8NFY3, Q8NFY5, Q8NFY6, Q8NFY7, Q9P249 | Gene names | SEMA6D, KIAA1479 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-6D precursor. | |||||
|
SEM6D_MOUSE
|
||||||
| θ value | 1.69304e-06 (rank : 31) | NC score | 0.345312 (rank : 25) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q76KF0, Q76KF1, Q76KF2, Q76KF3, Q76KF4, Q80TD0 | Gene names | Sema6d, Kiaa1479 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-6D precursor. | |||||
|
SEM5B_HUMAN
|
||||||
| θ value | 2.21117e-06 (rank : 32) | NC score | 0.321699 (rank : 33) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9P283, Q6UY12 | Gene names | SEMA5B, KIAA1445 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-5B precursor. | |||||
|
SEM5B_MOUSE
|
||||||
| θ value | 4.92598e-06 (rank : 33) | NC score | 0.320506 (rank : 35) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q60519 | Gene names | Sema5b, Semag, SemG | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-5B precursor (Semaphorin G) (Sema G). | |||||
|
SEM4D_MOUSE
|
||||||
| θ value | 6.43352e-06 (rank : 34) | NC score | 0.326620 (rank : 29) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O09126 | Gene names | Sema4d, Semacl2, Semaj | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4D precursor (Semaphorin J) (Sema J) (Semaphorin C-like 2) (M-Sema G). | |||||
|
SEM5A_HUMAN
|
||||||
| θ value | 8.40245e-06 (rank : 35) | NC score | 0.307739 (rank : 44) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q13591, O60408, Q1RLL9 | Gene names | SEMA5A, SEMAF | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-5A precursor (Semaphorin F) (Sema F). | |||||
|
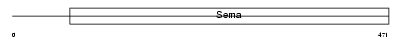
SEM7A_HUMAN
|
||||||
| θ value | 8.40245e-06 (rank : 36) | NC score | 0.340764 (rank : 27) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O75326 | Gene names | SEMA7A, CD108, SEMAL | |||
|
Domain Architecture |
|
|||||
| Description | Semaphorin-7A precursor (Semaphorin L) (Sema L) (Semaphorin K1) (Sema K1) (John-Milton-Hargen human blood group Ag) (JMH blood group antigen) (CD108 antigen) (CDw108). | |||||
|
SEM4F_MOUSE
|
||||||
| θ value | 5.44631e-05 (rank : 37) | NC score | 0.323752 (rank : 31) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9Z123, Q9R1Y1 | Gene names | Sema4f, Semaw | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4F precursor (Semaphorin W) (Sema W). | |||||
|
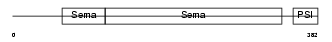
SEM4F_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 38) | NC score | 0.322544 (rank : 32) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O95754, Q9NS35 | Gene names | SEMA4F, SEMAM, SEMAW | |||
|
Domain Architecture |
|
|||||
| Description | Semaphorin-4F precursor (Semaphorin W) (Sema W) (Semaphorin M) (Sema M). | |||||
|
SEM3A_MOUSE
|
||||||
| θ value | 9.29e-05 (rank : 39) | NC score | 0.287244 (rank : 48) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O08665, Q5BL08, Q62180, Q62215 | Gene names | Sema3a, Semad, SemD | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3A precursor (Semaphorin III) (Sema III) (Semaphorin D) (Sema D). | |||||
|
SEM3E_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 40) | NC score | 0.288132 (rank : 47) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O15041, Q75M94, Q75M97 | Gene names | SEMA3E, KIAA0331 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-3E precursor. | |||||
|
SEM3E_MOUSE
|
||||||
| θ value | 0.000158464 (rank : 41) | NC score | 0.284384 (rank : 50) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P70275, O09078, O09079 | Gene names | Sema3e, Semah, Semh | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3E precursor (Semaphorin H) (Sema H). | |||||
|
SEM3F_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 42) | NC score | 0.320350 (rank : 36) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q13275, Q13274, Q13372, Q15704 | Gene names | SEMA3F | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3F precursor (Semaphorin IV) (Sema IV) (Sema III/F). | |||||
|
SEM3A_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 43) | NC score | 0.283450 (rank : 51) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q14563 | Gene names | SEMA3A | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3A precursor (Semaphorin III) (Sema III). | |||||
|
SEM3F_MOUSE
|
||||||
| θ value | 0.000270298 (rank : 44) | NC score | 0.320015 (rank : 37) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O88632, O88633 | Gene names | Sema3f | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3F precursor (Semaphorin IV) (Sema IV). | |||||
|
SEM3C_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 45) | NC score | 0.305918 (rank : 46) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q99985 | Gene names | SEMA3C, SEMAE | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3C precursor (Semaphorin E) (Sema E). | |||||
|
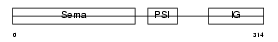
SEM3D_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 46) | NC score | 0.284778 (rank : 49) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | O95025, Q6UW77, Q8NCQ1 | Gene names | SEMA3D | |||
|
Domain Architecture |
|
|||||
| Description | Semaphorin-3D precursor. | |||||
|
SEM3B_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 47) | NC score | 0.313767 (rank : 39) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q62177 | Gene names | Sema3b, Sema, Semaa | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3B precursor (Semaphorin A) (Sema A). | |||||
|
RASA3_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 48) | NC score | 0.096277 (rank : 63) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q14644 | Gene names | RASA3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 3 (GAP1(IP4BP)) (Ins P4-binding protein). | |||||
|
SEM6A_MOUSE
|
||||||
| θ value | 0.00102713 (rank : 49) | NC score | 0.321588 (rank : 34) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O35464, Q6P5A8, Q6PCN9, Q9EQ71 | Gene names | Sema6a, Semaq | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6A precursor (Semaphorin VIA) (Sema VIA) (Semaphorin 6A-1) (SEMA6A-1) (Semaphorin Q) (Sema Q). | |||||
|
SEM3C_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 50) | NC score | 0.307464 (rank : 45) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q62181 | Gene names | Sema3c, Semae, SemE | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3C precursor (Semaphorin E) (Sema E). | |||||
|
SEM4D_HUMAN
|
||||||
| θ value | 0.00134147 (rank : 51) | NC score | 0.313066 (rank : 40) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q92854, Q7Z5S4 | Gene names | SEMA4D, CD100 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4D precursor (Leukocyte activation antigen CD100) (BB18) (A8) (GR3). | |||||
|
SEM6A_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 52) | NC score | 0.324707 (rank : 30) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9H2E6, Q9P2H9 | Gene names | SEMA6A, KIAA1368 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6A precursor (Semaphorin VIA) (Sema VIA) (Semaphorin 6A-1) (SEMA6A-1). | |||||
|
RASA3_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 53) | NC score | 0.084166 (rank : 64) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q60790 | Gene names | Rasa3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 3 (GAP1(IP4BP)) (Ins P4-binding protein) (GapIII). | |||||
|
SEM6C_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 54) | NC score | 0.310670 (rank : 41) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9H3T2, Q5JR71, Q8WXT8, Q8WXT9, Q8WXU0, Q96JF8 | Gene names | SEMA6C, KIAA1869, SEMAY | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6C precursor (Semaphorin Y) (Sema Y). | |||||
|
SEM6B_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 55) | NC score | 0.327027 (rank : 28) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9H3T3, Q9NRK9 | Gene names | SEMA6B, SEMAZ | |||
|
Domain Architecture |
|
|||||
| Description | Semaphorin-6B precursor (Semaphorin Z) (Sema Z). | |||||
|
SEM3B_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 56) | NC score | 0.308127 (rank : 43) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q13214, Q8TB71, Q8TDV7, Q93018, Q96GX0 | Gene names | SEMA3B, SEMA5 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3B precursor (Semaphorin V) (Sema V) (Sema A(V)). | |||||
|
EXOC2_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 57) | NC score | 0.119999 (rank : 59) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96KP1, Q5JPC8, Q96AN6, Q9NUZ8, Q9UJM7 | Gene names | EXOC2, SEC5, SEC5L1 | |||
|
Domain Architecture |
|
|||||
| Description | Exocyst complex component 2 (Exocyst complex component Sec5). | |||||
|
EXOC2_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 58) | NC score | 0.115290 (rank : 60) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9D4H1 | Gene names | Exoc2, Sec5, Sec5l1 | |||
|
Domain Architecture |
|
|||||
| Description | Exocyst complex component 2 (Exocyst complex component Sec5). | |||||
|
SEM3D_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 59) | NC score | 0.275198 (rank : 53) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8BH34 | Gene names | Sema3d | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-3D precursor. | |||||
|
ATRN_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 60) | NC score | 0.075035 (rank : 65) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 525 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O75882, O60295, O95414, Q3MIT3, Q5TDA2, Q5TDA4, Q5VYW3, Q9NTQ3, Q9NTQ4, Q9NU01, Q9NZ57, Q9NZ58 | Gene names | ATRN, KIAA0548, MGCA | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany homolog) (DPPT-L). | |||||
|
SYGP1_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 61) | NC score | 0.074518 (rank : 66) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96PV0, Q8TCS2, Q9UGE2 | Gene names | SYNGAP1, KIAA1938 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein SynGAP (Synaptic Ras-GTPase-activating protein 1) (Synaptic Ras-GAP 1) (Neuronal RasGAP). | |||||
|
SEM6B_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 62) | NC score | 0.309867 (rank : 42) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O54951 | Gene names | Sema6b, Seman | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6B precursor (Semaphorin VIB) (Sema VIB) (Semaphorin N) (Sema N). | |||||
|
SEM6C_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 63) | NC score | 0.281649 (rank : 52) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9WTM3 | Gene names | Sema6c, Semay | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6C precursor (Semaphorin Y) (Sema Y). | |||||
|
PKHD1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 64) | NC score | 0.181448 (rank : 57) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8TCZ9 | Gene names | PKHD1, FCYT, TIGM1 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney and hepatic disease 1 precursor (Fibrocystin) (Polyductin) (Tigmin). | |||||
|
PXDC1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 65) | NC score | 0.101660 (rank : 62) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8IUK5, Q5QCZ7, Q5QCZ8, Q5QCZ9, Q9HCT9 | Gene names | PLXDC1, TEM3, TEM7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin domain-containing protein 1 precursor (Tumor endothelial marker 7) (Tumor endothelial marker 3). | |||||
|
MEGF8_HUMAN
|
||||||
| θ value | 0.279714 (rank : 66) | NC score | 0.058648 (rank : 70) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q7Z7M0, O75097 | Gene names | MEGF8, EGFL4, KIAA0817 | |||
|
Domain Architecture |
|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
NGAP_HUMAN
|
||||||
| θ value | 0.279714 (rank : 67) | NC score | 0.043500 (rank : 74) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UJF2, O95174, Q5TFU9 | Gene names | RASAL2, NGAP | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein nGAP (RAS protein activator-like 1). | |||||
|
PXDC1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 68) | NC score | 0.105102 (rank : 61) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q91ZV7, Q8BM20, Q9CWV5 | Gene names | Plxdc1, Tem7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin domain-containing protein 1 precursor (Tumor endothelial marker 7). | |||||
|
RASA2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 69) | NC score | 0.042703 (rank : 75) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q15283, O00695, Q15284, Q92594, Q99577, Q9UEQ2 | Gene names | RASA2, GAP1M, RASGAP | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 2 (GAP1m). | |||||
|
ATRN_MOUSE
|
||||||
| θ value | 0.47712 (rank : 70) | NC score | 0.066805 (rank : 68) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9WU60, Q9R263, Q9WU77 | Gene names | Atrn, mg, Mgca | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany protein). | |||||
|
DAAM1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 71) | NC score | 0.006395 (rank : 88) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BPM0, Q80Y68, Q9CQQ2 | Gene names | Daam1 | |||
|
Domain Architecture |

|
|||||
| Description | Disheveled-associated activator of morphogenesis 1. | |||||
|
MEGF8_MOUSE
|
||||||
| θ value | 1.06291 (rank : 72) | NC score | 0.052392 (rank : 73) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 520 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P60882, Q80TR3, Q80V41, Q8BMN9, Q8JZW7, Q8K0J3 | Gene names | Megf8, Egfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
DAB2P_MOUSE
|
||||||
| θ value | 1.38821 (rank : 73) | NC score | 0.053933 (rank : 71) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q3UHC7, Q3TPD5, Q3UH44, Q6JTV1, Q80T97 | Gene names | Dab2ip, Kiaa1743 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Disabled homolog 2-interacting protein (DAB2-interacting protein). | |||||
|
DAB2P_HUMAN
|
||||||
| θ value | 1.81305 (rank : 74) | NC score | 0.053584 (rank : 72) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q5VWQ8, Q8TDL2, Q96SE1, Q9C0C0 | Gene names | DAB2IP, AF9Q34, AIP1, KIAA1743 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Disabled homolog 2-interacting protein (DAB2-interacting protein) (DAB2 interaction protein) (ASK-interacting protein 1). | |||||
|
ITB6_MOUSE
|
||||||
| θ value | 1.81305 (rank : 75) | NC score | 0.018564 (rank : 79) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Z0T9 | Gene names | Itgb6 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-6 precursor. | |||||
|
ZN292_HUMAN
|
||||||
| θ value | 1.81305 (rank : 76) | NC score | 0.004688 (rank : 91) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 918 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60281, Q9H8G3, Q9H8J4 | Gene names | ZNF292, KIAA0530 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 292. | |||||
|
AKAP6_HUMAN
|
||||||
| θ value | 2.36792 (rank : 77) | NC score | 0.010225 (rank : 85) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13023, O15028 | Gene names | AKAP6, AKAP100, KIAA0311 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 6 (Protein kinase A-anchoring protein 6) (PRKA6) (A-kinase anchor protein 100 kDa) (AKAP 100) (mAKAP). | |||||
|
DAXX_MOUSE
|
||||||
| θ value | 2.36792 (rank : 78) | NC score | 0.010611 (rank : 84) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35613, Q9QWT8, Q9QWV3 | Gene names | Daxx | |||
|
Domain Architecture |

|
|||||
| Description | Death domain-associated protein 6 (Daxx). | |||||
|
CENPF_HUMAN
|
||||||
| θ value | 3.0926 (rank : 79) | NC score | -0.002494 (rank : 102) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
GPM6A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 80) | NC score | 0.011706 (rank : 81) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P51674, Q92602 | Gene names | GPM6A, M6A | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal membrane glycoprotein M6-a (M6a). | |||||
|
GPM6A_MOUSE
|
||||||
| θ value | 3.0926 (rank : 81) | NC score | 0.011696 (rank : 82) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P35802 | Gene names | Gpm6a, M6a | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal membrane glycoprotein M6-a (M6a). | |||||
|
FRAS1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 82) | NC score | 0.008701 (rank : 86) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 759 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q86XX4, Q86UZ4, Q8N3U9, Q8NAU7, Q96JW7, Q9H6N9, Q9P228 | Gene names | FRAS1, KIAA1500 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Extracellular matrix protein FRAS1 precursor. | |||||
|
ITB1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 83) | NC score | 0.021277 (rank : 77) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P05556, P78466, P78467, Q13089, Q13090, Q13091, Q13212, Q14622, Q14647 | Gene names | ITGB1, FNRB | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-1 precursor (Fibronectin receptor subunit beta) (Integrin VLA-4 subunit beta) (CD29 antigen). | |||||
|
ITB1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 84) | NC score | 0.021015 (rank : 78) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P09055 | Gene names | Itgb1 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-1 precursor (Fibronectin receptor subunit beta) (Integrin VLA-4 subunit beta) (CD29 antigen). | |||||
|
LOXL1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 85) | NC score | 0.005674 (rank : 89) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q08397, Q96BW7 | Gene names | LOXL1, LOXL | |||
|
Domain Architecture |

|
|||||
| Description | Lysyl oxidase homolog 1 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 1) (LOL). | |||||
|
PXDC2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 86) | NC score | 0.073529 (rank : 67) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6UX71, Q96E59, Q96PD9, Q96SU9 | Gene names | PLXDC2, TEM7R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin domain-containing protein 2 precursor (Tumor endothelial marker 7-related protein). | |||||
|
PXDC2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 87) | NC score | 0.064133 (rank : 69) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9DC11, Q6PET5, Q91ZV6 | Gene names | Plxdc2, Tem7r | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin domain-containing protein 2 precursor (Tumor endothelial marker 7-related protein). | |||||
|
HGFA_HUMAN
|
||||||
| θ value | 5.27518 (rank : 88) | NC score | -0.000950 (rank : 99) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q04756, Q14726 | Gene names | HGFAC | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte growth factor activator precursor (EC 3.4.21.-) (HGF activator) (HGFA) [Contains: Hepatocyte growth factor activator short chain; Hepatocyte growth factor activator long chain]. | |||||
|
ITB8_HUMAN
|
||||||
| θ value | 5.27518 (rank : 89) | NC score | 0.017565 (rank : 80) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P26012 | Gene names | ITGB8 | |||
|
Domain Architecture |
|
|||||
| Description | Integrin beta-8 precursor. | |||||
|
KIF5A_MOUSE
|
||||||
| θ value | 5.27518 (rank : 90) | NC score | -0.000708 (rank : 98) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P33175, Q5DTP1, Q6PDY7, Q9Z2F9 | Gene names | Kif5a, Kiaa4086, Kif5, Nkhc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin heavy chain isoform 5A (Neuronal kinesin heavy chain) (NKHC) (Kinesin heavy chain neuron-specific 1). | |||||
|
MYT1L_MOUSE
|
||||||
| θ value | 5.27518 (rank : 91) | NC score | 0.004616 (rank : 92) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P97500, O08996, Q8C643, Q8C7L4, Q8CHB4 | Gene names | Myt1l, Kiaa1106, Nzf1, Png1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1-like protein (MyT1L protein) (Postmitotic neural gene 1 protein) (Zinc finger protein Png-1) (Neural zinc finger factor 1) (NZF-1). | |||||
|
NOC3L_MOUSE
|
||||||
| θ value | 5.27518 (rank : 92) | NC score | 0.006491 (rank : 87) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8VI84, Q9JKM3 | Gene names | Noc3l, Ad24, Fad24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar complex protein 3 homolog (NOC3 protein homolog) (NOC3-like protein) (Nucleolar complex-associated protein 3-like protein) (Factor for adipocyte differentiation 24). | |||||
|
CTRO_HUMAN
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | -0.000951 (rank : 100) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 2092 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O14578, Q6XUH8, Q86UQ9, Q9UPZ7 | Gene names | CIT, KIAA0949, STK21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
CTRO_MOUSE
|
||||||
| θ value | 6.88961 (rank : 94) | NC score | -0.001283 (rank : 101) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 2344 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P49025, O88528, O88937, O88938, Q3UM99, Q8CIJ1 | Gene names | Cit | |||
|
Domain Architecture |

|
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
RPGR1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 95) | NC score | 0.005026 (rank : 90) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 578 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96KN7, Q7Z2W6, Q8IXV5, Q96QA8, Q9HB94, Q9HB95, Q9HBK6, Q9NR40 | Gene names | RPGRIP1 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator-interacting protein 1 (RPGR-interacting protein 1). | |||||
|
AMPB_MOUSE
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | 0.002768 (rank : 96) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VCT3 | Gene names | Rnpep | |||
|
Domain Architecture |

|
|||||
| Description | Aminopeptidase B (EC 3.4.11.6) (Ap-B) (Arginyl aminopeptidase) (Arginine aminopeptidase) (Cytosol aminopeptidase IV). | |||||
|
ATM_HUMAN
|
||||||
| θ value | 8.99809 (rank : 97) | NC score | 0.003198 (rank : 94) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13315, O15429, Q12758, Q16551, Q93007, Q9NP02, Q9UCX7 | Gene names | ATM | |||
|
Domain Architecture |

|
|||||
| Description | Serine-protein kinase ATM (EC 2.7.11.1) (Ataxia telangiectasia mutated) (A-T, mutated). | |||||
|
FAT2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 98) | NC score | 0.001102 (rank : 97) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NYQ8, O75091, Q9NSR7 | Gene names | FAT2, CDHF8, KIAA0811, MEGF1 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin Fat 2 precursor (hFat2) (Multiple epidermal growth factor-like domains 1). | |||||
|
FBX5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 99) | NC score | 0.011438 (rank : 83) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UKT4, Q8WV29, Q9UGC8 | Gene names | FBXO5, EMI1, FBX5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 5 (Early mitotic inhibitor 1). | |||||
|
GNPAT_MOUSE
|
||||||
| θ value | 8.99809 (rank : 100) | NC score | 0.022783 (rank : 76) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P98192, Q9WUT6 | Gene names | Gnpat, Dhapat | |||
|
Domain Architecture |
|
|||||
| Description | Dihydroxyacetone phosphate acyltransferase (EC 2.3.1.42) (DHAP-AT) (DAP-AT) (Glycerone-phosphate O-acyltransferase) (Acyl- CoA:dihydroxyacetonephosphateacyltransferase). | |||||
|
GRIN1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 101) | NC score | 0.002926 (rank : 95) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3UNH4, Q6PGF9, Q9QZY2 | Gene names | Gprin1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
MYT1L_HUMAN
|
||||||
| θ value | 8.99809 (rank : 102) | NC score | 0.003696 (rank : 93) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UL68, Q6IQ17, Q9UPP6 | Gene names | MYT1L, KIAA1106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1-like protein (MyT1L protein) (MyT1-L). | |||||
|
PLXA3_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 102 | |
| SwissProt Accessions | P51805 | Gene names | PLXNA3, PLXN4, SEX | |||
|
Domain Architecture |

|
|||||
| Description | Plexin-A3 precursor (Plexin-4) (Semaphorin receptor SEX). | |||||
|
PLXA1_HUMAN
|
||||||
| NC score | 0.996098 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q9UIW2 | Gene names | PLXNA1, NOV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A1 precursor (Semaphorin receptor NOV). | |||||
|
PLXA1_MOUSE
|
||||||
| NC score | 0.995738 (rank : 3) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | P70206, Q5DTR0 | Gene names | Plxna1, KIAA4053 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A1 precursor (Plexin-1) (Plex 1). | |||||
|
PLXA2_HUMAN
|
||||||
| NC score | 0.995070 (rank : 4) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | O75051, Q6UX61, Q96GN9, Q9BRL1, Q9UIW1 | Gene names | PLXNA2, KIAA0463, OCT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A2 precursor (Semaphorin receptor OCT). | |||||
|
PLXA2_MOUSE
|
||||||
| NC score | 0.994859 (rank : 5) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | P70207, Q6NVE6, Q6PHN4, Q80TZ7, Q8R1I4 | Gene names | Plxna2, Kiaa0463 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A2 precursor (Plexin-2) (Plex 2). | |||||
|
PLXA4_MOUSE
|
||||||
| NC score | 0.994150 (rank : 6) | θ value | 0 (rank : 7) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q80UG2, Q5DTW8, Q8BKK9 | Gene names | Plxna4, Kiaa1550 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A4 precursor. | |||||
|
PLXA4_HUMAN
|
||||||
| NC score | 0.993841 (rank : 7) | θ value | 0 (rank : 6) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q9HCM2, Q6UWC6, Q6ZW89, Q8N969, Q8ND00, Q8NEN3, Q9NTD4 | Gene names | PLXNA4, KIAA1550, PLXNA4A, PLXNA4B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A4 precursor. | |||||
|
PLXD1_HUMAN
|
||||||
| NC score | 0.974430 (rank : 8) | θ value | 0 (rank : 13) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9Y4D7, Q6PJS9, Q8IZJ2, Q9BTQ2 | Gene names | PLXND1, KIAA0620 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-D1 precursor. | |||||
|
PLXB2_HUMAN
|
||||||
| NC score | 0.972879 (rank : 9) | θ value | 0 (rank : 10) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | O15031, Q7KZU3, Q9BSU7 | Gene names | PLXNB2, KIAA0315 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B2 precursor (MM1). | |||||
|
PLXB3_HUMAN
|
||||||
| NC score | 0.969545 (rank : 10) | θ value | 0 (rank : 11) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9ULL4, Q9HDA4 | Gene names | PLXNB3, KIAA1206 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B3 precursor. | |||||
|
PLXB3_MOUSE
|
||||||
| NC score | 0.968590 (rank : 11) | θ value | 0 (rank : 12) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9QY40, Q80TH8 | Gene names | Plxnb3, Kiaa1206, Plxn6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B3 precursor (Plexin-6). | |||||
|
PLXB1_MOUSE
|
||||||
| NC score | 0.953066 (rank : 12) | θ value | 0 (rank : 9) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q8CJH3, Q6ZQC3, Q80ZZ1 | Gene names | Plxnb1, Kiaa0407 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B1 precursor. | |||||
|
PLXB1_HUMAN
|
||||||
| NC score | 0.942922 (rank : 13) | θ value | 0 (rank : 8) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | O43157, Q6NY20, Q9UIV7, Q9UJ92, Q9UJ93 | Gene names | PLXNB1, KIAA0407, SEP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B1 precursor (Semaphorin receptor SEP). | |||||
|
PLXC1_MOUSE
|
||||||
| NC score | 0.927467 (rank : 14) | θ value | 9.63072e-111 (rank : 15) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9QZC2, Q8CGW1 | Gene names | Plxnc1, Vespr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-C1 precursor (Virus-encoded semaphorin protein receptor) (CD232 antigen). | |||||
|
PLXC1_HUMAN
|
||||||
| NC score | 0.925531 (rank : 15) | θ value | 2.5344e-111 (rank : 14) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | O60486, Q59H25 | Gene names | PLXNC1, VESPR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-C1 precursor (Virus-encoded semaphorin protein receptor) (CD232 antigen). | |||||
|
SEM4B_MOUSE
|
||||||
| NC score | 0.412617 (rank : 16) | θ value | 2.79066e-09 (rank : 23) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q62179, Q4PKI6, Q69ZB7 | Gene names | Sema4b, Kiaa1745, Semac, SemC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-4B precursor (Semaphorin C) (Sema C). | |||||
|
SEM4A_HUMAN
|
||||||
| NC score | 0.406337 (rank : 17) | θ value | 3.18553e-13 (rank : 20) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9H3S1, Q8WUA9 | Gene names | SEMA4A, SEMB | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4A precursor (Semaphorin B) (Sema B). | |||||
|
SEM4B_HUMAN
|
||||||
| NC score | 0.401799 (rank : 18) | θ value | 6.21693e-09 (rank : 25) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9NPR2, Q6UXE3, Q8WVP9, Q96FK5, Q9C0B8, Q9H691, Q9NPM8, Q9NPN0 | Gene names | SEMA4B, KIAA1745 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4B precursor. | |||||
|
SEM4A_MOUSE
|
||||||
| NC score | 0.400311 (rank : 19) | θ value | 3.89403e-11 (rank : 22) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q62178 | Gene names | Sema4a, Semab, SemB | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4A precursor (Semaphorin B) (Sema B). | |||||
|
SEM4C_HUMAN
|
||||||
| NC score | 0.379442 (rank : 20) | θ value | 1.02475e-11 (rank : 21) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9C0C4, Q7Z5X0 | Gene names | SEMA4C, KIAA1739 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4C precursor. | |||||
|
SEM4G_MOUSE
|
||||||
| NC score | 0.367820 (rank : 21) | θ value | 3.64472e-09 (rank : 24) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9WUH7 | Gene names | Sema4g | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4G precursor. | |||||
|
SEM4G_HUMAN
|
||||||
| NC score | 0.366795 (rank : 22) | θ value | 2.36244e-08 (rank : 26) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9NTN9, Q9HCF3 | Gene names | SEMA4G, KIAA1619 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4G precursor. | |||||
|
SEM7A_MOUSE
|
||||||
| NC score | 0.352582 (rank : 23) | θ value | 4.45536e-07 (rank : 28) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9QUR8, O88371 | Gene names | Sema7a, Cd108, Semal, Semk1 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-7A precursor (Semaphorin L) (Sema L) (Semaphorin K1) (Sema K1) (CD108 antigen) (CDw108). | |||||
|
SEM4C_MOUSE
|
||||||
| NC score | 0.347606 (rank : 24) | θ value | 4.0297e-08 (rank : 27) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q64151 | Gene names | Sema4c, Semacl1, Semai | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4C precursor (Semaphorin I) (Sema I) (Semaphorin C-like 1) (M-Sema F). | |||||
|
SEM6D_MOUSE
|
||||||
| NC score | 0.345312 (rank : 25) | θ value | 1.69304e-06 (rank : 31) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q76KF0, Q76KF1, Q76KF2, Q76KF3, Q76KF4, Q80TD0 | Gene names | Sema6d, Kiaa1479 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-6D precursor. | |||||
|
SEM6D_HUMAN
|
||||||
| NC score | 0.343729 (rank : 26) | θ value | 1.69304e-06 (rank : 30) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8NFY4, Q8NFY3, Q8NFY5, Q8NFY6, Q8NFY7, Q9P249 | Gene names | SEMA6D, KIAA1479 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-6D precursor. | |||||
|
SEM7A_HUMAN
|
||||||
| NC score | 0.340764 (rank : 27) | θ value | 8.40245e-06 (rank : 36) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O75326 | Gene names | SEMA7A, CD108, SEMAL | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-7A precursor (Semaphorin L) (Sema L) (Semaphorin K1) (Sema K1) (John-Milton-Hargen human blood group Ag) (JMH blood group antigen) (CD108 antigen) (CDw108). | |||||
|
SEM6B_HUMAN
|
||||||
| NC score | 0.327027 (rank : 28) | θ value | 0.00869519 (rank : 55) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9H3T3, Q9NRK9 | Gene names | SEMA6B, SEMAZ | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6B precursor (Semaphorin Z) (Sema Z). | |||||
|
SEM4D_MOUSE
|
||||||
| NC score | 0.326620 (rank : 29) | θ value | 6.43352e-06 (rank : 34) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O09126 | Gene names | Sema4d, Semacl2, Semaj | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4D precursor (Semaphorin J) (Sema J) (Semaphorin C-like 2) (M-Sema G). | |||||
|
SEM6A_HUMAN
|
||||||
| NC score | 0.324707 (rank : 30) | θ value | 0.00175202 (rank : 52) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9H2E6, Q9P2H9 | Gene names | SEMA6A, KIAA1368 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6A precursor (Semaphorin VIA) (Sema VIA) (Semaphorin 6A-1) (SEMA6A-1). | |||||
|
SEM4F_MOUSE
|
||||||
| NC score | 0.323752 (rank : 31) | θ value | 5.44631e-05 (rank : 37) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9Z123, Q9R1Y1 | Gene names | Sema4f, Semaw | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4F precursor (Semaphorin W) (Sema W). | |||||
|
SEM4F_HUMAN
|
||||||
| NC score | 0.322544 (rank : 32) | θ value | 7.1131e-05 (rank : 38) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O95754, Q9NS35 | Gene names | SEMA4F, SEMAM, SEMAW | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4F precursor (Semaphorin W) (Sema W) (Semaphorin M) (Sema M). | |||||
|
SEM5B_HUMAN
|
||||||
| NC score | 0.321699 (rank : 33) | θ value | 2.21117e-06 (rank : 32) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9P283, Q6UY12 | Gene names | SEMA5B, KIAA1445 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-5B precursor. | |||||
|
SEM6A_MOUSE
|
||||||
| NC score | 0.321588 (rank : 34) | θ value | 0.00102713 (rank : 49) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O35464, Q6P5A8, Q6PCN9, Q9EQ71 | Gene names | Sema6a, Semaq | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6A precursor (Semaphorin VIA) (Sema VIA) (Semaphorin 6A-1) (SEMA6A-1) (Semaphorin Q) (Sema Q). | |||||
|
SEM5B_MOUSE
|
||||||
| NC score | 0.320506 (rank : 35) | θ value | 4.92598e-06 (rank : 33) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q60519 | Gene names | Sema5b, Semag, SemG | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-5B precursor (Semaphorin G) (Sema G). | |||||
|
SEM3F_HUMAN
|
||||||
| NC score | 0.320350 (rank : 36) | θ value | 0.000158464 (rank : 42) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q13275, Q13274, Q13372, Q15704 | Gene names | SEMA3F | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3F precursor (Semaphorin IV) (Sema IV) (Sema III/F). | |||||
|
SEM3F_MOUSE
|
||||||
| NC score | 0.320015 (rank : 37) | θ value | 0.000270298 (rank : 44) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O88632, O88633 | Gene names | Sema3f | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3F precursor (Semaphorin IV) (Sema IV). | |||||
|
SEM5A_MOUSE
|
||||||
| NC score | 0.319209 (rank : 38) | θ value | 9.92553e-07 (rank : 29) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q62217 | Gene names | Sema5a, Semaf, SemF | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-5A precursor (Semaphorin F) (Sema F). | |||||
|
SEM3B_MOUSE
|
||||||
| NC score | 0.313767 (rank : 39) | θ value | 0.000786445 (rank : 47) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q62177 | Gene names | Sema3b, Sema, Semaa | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3B precursor (Semaphorin A) (Sema A). | |||||
|
SEM4D_HUMAN
|
||||||
| NC score | 0.313066 (rank : 40) | θ value | 0.00134147 (rank : 51) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q92854, Q7Z5S4 | Gene names | SEMA4D, CD100 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4D precursor (Leukocyte activation antigen CD100) (BB18) (A8) (GR3). | |||||
|
SEM6C_HUMAN
|
||||||
| NC score | 0.310670 (rank : 41) | θ value | 0.00665767 (rank : 54) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9H3T2, Q5JR71, Q8WXT8, Q8WXT9, Q8WXU0, Q96JF8 | Gene names | SEMA6C, KIAA1869, SEMAY | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6C precursor (Semaphorin Y) (Sema Y). | |||||
|
SEM6B_MOUSE
|
||||||
| NC score | 0.309867 (rank : 42) | θ value | 0.0563607 (rank : 62) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O54951 | Gene names | Sema6b, Seman | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6B precursor (Semaphorin VIB) (Sema VIB) (Semaphorin N) (Sema N). | |||||
|
SEM3B_HUMAN
|
||||||
| NC score | 0.308127 (rank : 43) | θ value | 0.0113563 (rank : 56) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q13214, Q8TB71, Q8TDV7, Q93018, Q96GX0 | Gene names | SEMA3B, SEMA5 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3B precursor (Semaphorin V) (Sema V) (Sema A(V)). | |||||
|
SEM5A_HUMAN
|
||||||
| NC score | 0.307739 (rank : 44) | θ value | 8.40245e-06 (rank : 35) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q13591, O60408, Q1RLL9 | Gene names | SEMA5A, SEMAF | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-5A precursor (Semaphorin F) (Sema F). | |||||
|
SEM3C_MOUSE
|
||||||
| NC score | 0.307464 (rank : 45) | θ value | 0.00134147 (rank : 50) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q62181 | Gene names | Sema3c, Semae, SemE | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3C precursor (Semaphorin E) (Sema E). | |||||
|
SEM3C_HUMAN
|
||||||
| NC score | 0.305918 (rank : 46) | θ value | 0.000461057 (rank : 45) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q99985 | Gene names | SEMA3C, SEMAE | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3C precursor (Semaphorin E) (Sema E). | |||||
|
SEM3E_HUMAN
|
||||||
| NC score | 0.288132 (rank : 47) | θ value | 0.000158464 (rank : 40) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O15041, Q75M94, Q75M97 | Gene names | SEMA3E, KIAA0331 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-3E precursor. | |||||
|
SEM3A_MOUSE
|
||||||
| NC score | 0.287244 (rank : 48) | θ value | 9.29e-05 (rank : 39) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O08665, Q5BL08, Q62180, Q62215 | Gene names | Sema3a, Semad, SemD | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3A precursor (Semaphorin III) (Sema III) (Semaphorin D) (Sema D). | |||||
|
SEM3D_HUMAN
|
||||||
| NC score | 0.284778 (rank : 49) | θ value | 0.000602161 (rank : 46) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | O95025, Q6UW77, Q8NCQ1 | Gene names | SEMA3D | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3D precursor. | |||||
|
SEM3E_MOUSE
|
||||||
| NC score | 0.284384 (rank : 50) | θ value | 0.000158464 (rank : 41) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P70275, O09078, O09079 | Gene names | Sema3e, Semah, Semh | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3E precursor (Semaphorin H) (Sema H). | |||||
|
SEM3A_HUMAN
|
||||||
| NC score | 0.283450 (rank : 51) | θ value | 0.00020696 (rank : 43) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q14563 | Gene names | SEMA3A | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3A precursor (Semaphorin III) (Sema III). | |||||
|
SEM6C_MOUSE
|
||||||
| NC score | 0.281649 (rank : 52) | θ value | 0.0736092 (rank : 63) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9WTM3 | Gene names | Sema6c, Semay | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6C precursor (Semaphorin Y) (Sema Y). | |||||
|
SEM3D_MOUSE
|
||||||
| NC score | 0.275198 (rank : 53) | θ value | 0.0193708 (rank : 59) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8BH34 | Gene names | Sema3d | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-3D precursor. | |||||
|
MET_HUMAN
|
||||||
| NC score | 0.206852 (rank : 54) | θ value | 5.40856e-29 (rank : 17) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 831 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P08581, O60366, Q12875, Q9UDX7, Q9UPL8 | Gene names | MET | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte growth factor receptor precursor (EC 2.7.10.1) (HGF receptor) (Scatter factor receptor) (SF receptor) (HGF/SF receptor) (Met proto-oncogene tyrosine kinase) (c-Met). | |||||
|
MET_MOUSE
|
||||||
| NC score | 0.206334 (rank : 55) | θ value | 1.08979e-29 (rank : 16) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 844 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P16056, Q62125 | Gene names | Met | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte growth factor receptor precursor (EC 2.7.10.1) (HGF receptor) (Scatter factor receptor) (SF receptor) (HGF/SF receptor) (Met proto-oncogene tyrosine kinase) (c-Met). | |||||
|
RON_HUMAN
|
||||||
| NC score | 0.183462 (rank : 56) | θ value | 2.12685e-25 (rank : 19) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q04912 | Gene names | MST1R, RON | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage-stimulating protein receptor precursor (EC 2.7.10.1) (MSP receptor) (p185-Ron) (CDw136 antigen) [Contains: Macrophage- stimulating protein receptor alpha chain; Macrophage-stimulating protein receptor beta chain]. | |||||
|
PKHD1_HUMAN
|
||||||
| NC score | 0.181448 (rank : 57) | θ value | 0.163984 (rank : 64) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8TCZ9 | Gene names | PKHD1, FCYT, TIGM1 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney and hepatic disease 1 precursor (Fibrocystin) (Polyductin) (Tigmin). | |||||
|
RON_MOUSE
|
||||||
| NC score | 0.180933 (rank : 58) | θ value | 7.30988e-26 (rank : 18) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q62190, Q62555 | Gene names | Mst1r, Ron, Stk | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage-stimulating protein receptor precursor (EC 2.7.10.1) (MSP receptor) (p185-Ron) (Stem cell-derived tyrosine kinase) (CDw136 antigen) [Contains: Macrophage-stimulating protein receptor alpha chain; Macrophage-stimulating protein receptor beta chain]. | |||||
|
EXOC2_HUMAN
|
||||||
| NC score | 0.119999 (rank : 59) | θ value | 0.0193708 (rank : 57) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96KP1, Q5JPC8, Q96AN6, Q9NUZ8, Q9UJM7 | Gene names | EXOC2, SEC5, SEC5L1 | |||
|
Domain Architecture |

|
|||||
| Description | Exocyst complex component 2 (Exocyst complex component Sec5). | |||||
|
EXOC2_MOUSE
|
||||||
| NC score | 0.115290 (rank : 60) | θ value | 0.0193708 (rank : 58) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9D4H1 | Gene names | Exoc2, Sec5, Sec5l1 | |||
|
Domain Architecture |

|
|||||
| Description | Exocyst complex component 2 (Exocyst complex component Sec5). | |||||
|
PXDC1_MOUSE
|
||||||
| NC score | 0.105102 (rank : 61) | θ value | 0.279714 (rank : 68) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q91ZV7, Q8BM20, Q9CWV5 | Gene names | Plxdc1, Tem7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin domain-containing protein 1 precursor (Tumor endothelial marker 7). | |||||
|
PXDC1_HUMAN
|
||||||
| NC score | 0.101660 (rank : 62) | θ value | 0.21417 (rank : 65) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8IUK5, Q5QCZ7, Q5QCZ8, Q5QCZ9, Q9HCT9 | Gene names | PLXDC1, TEM3, TEM7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin domain-containing protein 1 precursor (Tumor endothelial marker 7) (Tumor endothelial marker 3). | |||||
|
RASA3_HUMAN
|
||||||
| NC score | 0.096277 (rank : 63) | θ value | 0.00102713 (rank : 48) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q14644 | Gene names | RASA3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 3 (GAP1(IP4BP)) (Ins P4-binding protein). | |||||
|
RASA3_MOUSE
|
||||||
| NC score | 0.084166 (rank : 64) | θ value | 0.00665767 (rank : 53) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q60790 | Gene names | Rasa3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 3 (GAP1(IP4BP)) (Ins P4-binding protein) (GapIII). | |||||
|
ATRN_HUMAN
|
||||||
| NC score | 0.075035 (rank : 65) | θ value | 0.0252991 (rank : 60) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 525 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O75882, O60295, O95414, Q3MIT3, Q5TDA2, Q5TDA4, Q5VYW3, Q9NTQ3, Q9NTQ4, Q9NU01, Q9NZ57, Q9NZ58 | Gene names | ATRN, KIAA0548, MGCA | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany homolog) (DPPT-L). | |||||
|
SYGP1_HUMAN
|
||||||
| NC score | 0.074518 (rank : 66) | θ value | 0.0431538 (rank : 61) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96PV0, Q8TCS2, Q9UGE2 | Gene names | SYNGAP1, KIAA1938 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein SynGAP (Synaptic Ras-GTPase-activating protein 1) (Synaptic Ras-GAP 1) (Neuronal RasGAP). | |||||
|
PXDC2_HUMAN
|
||||||
| NC score | 0.073529 (rank : 67) | θ value | 4.03905 (rank : 86) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6UX71, Q96E59, Q96PD9, Q96SU9 | Gene names | PLXDC2, TEM7R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin domain-containing protein 2 precursor (Tumor endothelial marker 7-related protein). | |||||
|
ATRN_MOUSE
|
||||||
| NC score | 0.066805 (rank : 68) | θ value | 0.47712 (rank : 70) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9WU60, Q9R263, Q9WU77 | Gene names | Atrn, mg, Mgca | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany protein). | |||||
|
PXDC2_MOUSE
|
||||||
| NC score | 0.064133 (rank : 69) | θ value | 4.03905 (rank : 87) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9DC11, Q6PET5, Q91ZV6 | Gene names | Plxdc2, Tem7r | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin domain-containing protein 2 precursor (Tumor endothelial marker 7-related protein). | |||||
|
MEGF8_HUMAN
|
||||||
| NC score | 0.058648 (rank : 70) | θ value | 0.279714 (rank : 66) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q7Z7M0, O75097 | Gene names | MEGF8, EGFL4, KIAA0817 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
DAB2P_MOUSE
|
||||||
| NC score | 0.053933 (rank : 71) | θ value | 1.38821 (rank : 73) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q3UHC7, Q3TPD5, Q3UH44, Q6JTV1, Q80T97 | Gene names | Dab2ip, Kiaa1743 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Disabled homolog 2-interacting protein (DAB2-interacting protein). | |||||
|
DAB2P_HUMAN
|
||||||
| NC score | 0.053584 (rank : 72) | θ value | 1.81305 (rank : 74) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q5VWQ8, Q8TDL2, Q96SE1, Q9C0C0 | Gene names | DAB2IP, AF9Q34, AIP1, KIAA1743 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Disabled homolog 2-interacting protein (DAB2-interacting protein) (DAB2 interaction protein) (ASK-interacting protein 1). | |||||
|
MEGF8_MOUSE
|
||||||
| NC score | 0.052392 (rank : 73) | θ value | 1.06291 (rank : 72) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 520 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P60882, Q80TR3, Q80V41, Q8BMN9, Q8JZW7, Q8K0J3 | Gene names | Megf8, Egfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
NGAP_HUMAN
|
||||||
| NC score | 0.043500 (rank : 74) | θ value | 0.279714 (rank : 67) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UJF2, O95174, Q5TFU9 | Gene names | RASAL2, NGAP | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein nGAP (RAS protein activator-like 1). | |||||
|
RASA2_HUMAN
|
||||||
| NC score | 0.042703 (rank : 75) | θ value | 0.365318 (rank : 69) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q15283, O00695, Q15284, Q92594, Q99577, Q9UEQ2 | Gene names | RASA2, GAP1M, RASGAP | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 2 (GAP1m). | |||||
|
GNPAT_MOUSE
|
||||||
| NC score | 0.022783 (rank : 76) | θ value | 8.99809 (rank : 100) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P98192, Q9WUT6 | Gene names | Gnpat, Dhapat | |||
|
Domain Architecture |

|
|||||
| Description | Dihydroxyacetone phosphate acyltransferase (EC 2.3.1.42) (DHAP-AT) (DAP-AT) (Glycerone-phosphate O-acyltransferase) (Acyl- CoA:dihydroxyacetonephosphateacyltransferase). | |||||
|
ITB1_HUMAN
|
||||||
| NC score | 0.021277 (rank : 77) | θ value | 4.03905 (rank : 83) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P05556, P78466, P78467, Q13089, Q13090, Q13091, Q13212, Q14622, Q14647 | Gene names | ITGB1, FNRB | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-1 precursor (Fibronectin receptor subunit beta) (Integrin VLA-4 subunit beta) (CD29 antigen). | |||||
|
ITB1_MOUSE
|
||||||
| NC score | 0.021015 (rank : 78) | θ value | 4.03905 (rank : 84) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P09055 | Gene names | Itgb1 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-1 precursor (Fibronectin receptor subunit beta) (Integrin VLA-4 subunit beta) (CD29 antigen). | |||||
|
ITB6_MOUSE
|
||||||
| NC score | 0.018564 (rank : 79) | θ value | 1.81305 (rank : 75) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Z0T9 | Gene names | Itgb6 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-6 precursor. | |||||
|
ITB8_HUMAN
|
||||||
| NC score | 0.017565 (rank : 80) | θ value | 5.27518 (rank : 89) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P26012 | Gene names | ITGB8 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-8 precursor. | |||||
|
GPM6A_HUMAN
|
||||||
| NC score | 0.011706 (rank : 81) | θ value | 3.0926 (rank : 80) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P51674, Q92602 | Gene names | GPM6A, M6A | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal membrane glycoprotein M6-a (M6a). | |||||
|
GPM6A_MOUSE
|
||||||
| NC score | 0.011696 (rank : 82) | θ value | 3.0926 (rank : 81) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P35802 | Gene names | Gpm6a, M6a | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal membrane glycoprotein M6-a (M6a). | |||||
|
FBX5_HUMAN
|
||||||
| NC score | 0.011438 (rank : 83) | θ value | 8.99809 (rank : 99) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UKT4, Q8WV29, Q9UGC8 | Gene names | FBXO5, EMI1, FBX5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 5 (Early mitotic inhibitor 1). | |||||
|
DAXX_MOUSE
|
||||||
| NC score | 0.010611 (rank : 84) | θ value | 2.36792 (rank : 78) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35613, Q9QWT8, Q9QWV3 | Gene names | Daxx | |||
|
Domain Architecture |

|
|||||
| Description | Death domain-associated protein 6 (Daxx). | |||||
|
AKAP6_HUMAN
|
||||||
| NC score | 0.010225 (rank : 85) | θ value | 2.36792 (rank : 77) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13023, O15028 | Gene names | AKAP6, AKAP100, KIAA0311 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 6 (Protein kinase A-anchoring protein 6) (PRKA6) (A-kinase anchor protein 100 kDa) (AKAP 100) (mAKAP). | |||||
|
FRAS1_HUMAN
|
||||||
| NC score | 0.008701 (rank : 86) | θ value | 4.03905 (rank : 82) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 759 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q86XX4, Q86UZ4, Q8N3U9, Q8NAU7, Q96JW7, Q9H6N9, Q9P228 | Gene names | FRAS1, KIAA1500 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Extracellular matrix protein FRAS1 precursor. | |||||
|
NOC3L_MOUSE
|
||||||
| NC score | 0.006491 (rank : 87) | θ value | 5.27518 (rank : 92) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8VI84, Q9JKM3 | Gene names | Noc3l, Ad24, Fad24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar complex protein 3 homolog (NOC3 protein homolog) (NOC3-like protein) (Nucleolar complex-associated protein 3-like protein) (Factor for adipocyte differentiation 24). | |||||
|
DAAM1_MOUSE
|
||||||
| NC score | 0.006395 (rank : 88) | θ value | 1.06291 (rank : 71) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BPM0, Q80Y68, Q9CQQ2 | Gene names | Daam1 | |||
|
Domain Architecture |

|
|||||
| Description | Disheveled-associated activator of morphogenesis 1. | |||||
|
LOXL1_HUMAN
|
||||||
| NC score | 0.005674 (rank : 89) | θ value | 4.03905 (rank : 85) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q08397, Q96BW7 | Gene names | LOXL1, LOXL | |||
|
Domain Architecture |

|
|||||
| Description | Lysyl oxidase homolog 1 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 1) (LOL). | |||||
|
RPGR1_HUMAN
|
||||||
| NC score | 0.005026 (rank : 90) | θ value | 6.88961 (rank : 95) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 578 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96KN7, Q7Z2W6, Q8IXV5, Q96QA8, Q9HB94, Q9HB95, Q9HBK6, Q9NR40 | Gene names | RPGRIP1 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator-interacting protein 1 (RPGR-interacting protein 1). | |||||
|
ZN292_HUMAN
|
||||||
| NC score | 0.004688 (rank : 91) | θ value | 1.81305 (rank : 76) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 918 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60281, Q9H8G3, Q9H8J4 | Gene names | ZNF292, KIAA0530 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 292. | |||||
|
MYT1L_MOUSE
|
||||||
| NC score | 0.004616 (rank : 92) | θ value | 5.27518 (rank : 91) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P97500, O08996, Q8C643, Q8C7L4, Q8CHB4 | Gene names | Myt1l, Kiaa1106, Nzf1, Png1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1-like protein (MyT1L protein) (Postmitotic neural gene 1 protein) (Zinc finger protein Png-1) (Neural zinc finger factor 1) (NZF-1). | |||||
|
MYT1L_HUMAN
|
||||||
| NC score | 0.003696 (rank : 93) | θ value | 8.99809 (rank : 102) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UL68, Q6IQ17, Q9UPP6 | Gene names | MYT1L, KIAA1106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1-like protein (MyT1L protein) (MyT1-L). | |||||
|
ATM_HUMAN
|
||||||
| NC score | 0.003198 (rank : 94) | θ value | 8.99809 (rank : 97) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13315, O15429, Q12758, Q16551, Q93007, Q9NP02, Q9UCX7 | Gene names | ATM | |||
|
Domain Architecture |

|
|||||
| Description | Serine-protein kinase ATM (EC 2.7.11.1) (Ataxia telangiectasia mutated) (A-T, mutated). | |||||
|
GRIN1_MOUSE
|
||||||
| NC score | 0.002926 (rank : 95) | θ value | 8.99809 (rank : 101) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3UNH4, Q6PGF9, Q9QZY2 | Gene names | Gprin1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
AMPB_MOUSE
|
||||||
| NC score | 0.002768 (rank : 96) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VCT3 | Gene names | Rnpep | |||
|
Domain Architecture |

|
|||||
| Description | Aminopeptidase B (EC 3.4.11.6) (Ap-B) (Arginyl aminopeptidase) (Arginine aminopeptidase) (Cytosol aminopeptidase IV). | |||||
|
FAT2_HUMAN
|
||||||
| NC score | 0.001102 (rank : 97) | θ value | 8.99809 (rank : 98) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NYQ8, O75091, Q9NSR7 | Gene names | FAT2, CDHF8, KIAA0811, MEGF1 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin Fat 2 precursor (hFat2) (Multiple epidermal growth factor-like domains 1). | |||||
|
KIF5A_MOUSE
|
||||||
| NC score | -0.000708 (rank : 98) | θ value | 5.27518 (rank : 90) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P33175, Q5DTP1, Q6PDY7, Q9Z2F9 | Gene names | Kif5a, Kiaa4086, Kif5, Nkhc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin heavy chain isoform 5A (Neuronal kinesin heavy chain) (NKHC) (Kinesin heavy chain neuron-specific 1). | |||||
|
HGFA_HUMAN
|
||||||
| NC score | -0.000950 (rank : 99) | θ value | 5.27518 (rank : 88) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q04756, Q14726 | Gene names | HGFAC | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte growth factor activator precursor (EC 3.4.21.-) (HGF activator) (HGFA) [Contains: Hepatocyte growth factor activator short chain; Hepatocyte growth factor activator long chain]. | |||||
|
CTRO_HUMAN
|
||||||
| NC score | -0.000951 (rank : 100) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 2092 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O14578, Q6XUH8, Q86UQ9, Q9UPZ7 | Gene names | CIT, KIAA0949, STK21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
CTRO_MOUSE
|
||||||
| NC score | -0.001283 (rank : 101) | θ value | 6.88961 (rank : 94) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 2344 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P49025, O88528, O88937, O88938, Q3UM99, Q8CIJ1 | Gene names | Cit | |||
|
Domain Architecture |

|
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
CENPF_HUMAN
|
||||||
| NC score | -0.002494 (rank : 102) | θ value | 3.0926 (rank : 79) | |||
| Query Neighborhood Hits | 102 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||