Please be patient as the page loads
|
GNPAT_MOUSE
|
||||||
| SwissProt Accessions | P98192, Q9WUT6 | Gene names | Gnpat, Dhapat | |||
|
Domain Architecture |
|
|||||
| Description | Dihydroxyacetone phosphate acyltransferase (EC 2.3.1.42) (DHAP-AT) (DAP-AT) (Glycerone-phosphate O-acyltransferase) (Acyl- CoA:dihydroxyacetonephosphateacyltransferase). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
GNPAT_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.997123 (rank : 2) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O15228 | Gene names | GNPAT, DAPAT, DHAPAT | |||
|
Domain Architecture |
|
|||||
| Description | Dihydroxyacetone phosphate acyltransferase (EC 2.3.1.42) (DHAP-AT) (DAP-AT) (Glycerone-phosphate O-acyltransferase) (Acyl- CoA:dihydroxyacetonephosphateacyltransferase). | |||||
|
GNPAT_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P98192, Q9WUT6 | Gene names | Gnpat, Dhapat | |||
|
Domain Architecture |

|
|||||
| Description | Dihydroxyacetone phosphate acyltransferase (EC 2.3.1.42) (DHAP-AT) (DAP-AT) (Glycerone-phosphate O-acyltransferase) (Acyl- CoA:dihydroxyacetonephosphateacyltransferase). | |||||
|
PLSB_MOUSE
|
||||||
| θ value | 1.6247e-33 (rank : 3) | NC score | 0.705728 (rank : 3) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61586 | Gene names | Gpam | |||
|
Domain Architecture |

|
|||||
| Description | Glycerol-3-phosphate acyltransferase, mitochondrial precursor (EC 2.3.1.15) (GPAT) (P90). | |||||
|
PLSB_HUMAN
|
||||||
| θ value | 6.17384e-33 (rank : 4) | NC score | 0.704596 (rank : 4) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9HCL2, Q5VW51 | Gene names | GPAM, KIAA1560 | |||
|
Domain Architecture |
|
|||||
| Description | Glycerol-3-phosphate acyltransferase, mitochondrial precursor (EC 2.3.1.15) (GPAT). | |||||
|
FHOD1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 5) | NC score | 0.043016 (rank : 5) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6P9Q4, Q8BMK2 | Gene names | Fhod1, Fhos1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
KCMB2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 6) | NC score | 0.029955 (rank : 6) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y691 | Gene names | KCNMB2 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-activated potassium channel subunit beta 2 (Calcium-activated potassium channel, subfamily M subunit beta 2) (Maxi K channel subunit beta 2) (BK channel subunit beta 2) (Slo-beta 2) (K(VCA)beta 2) (Charybdotoxin receptor subunit beta 2) (BKbeta2) (Hbeta2) (Hbeta3). | |||||
|
LRSM1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 7) | NC score | 0.013358 (rank : 14) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 1078 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6UWE0, Q5VVV0, Q8NB40, Q96GT5, Q96MX5, Q96MZ7 | Gene names | LRSAM1, TAL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein LRSAM1 (EC 6.3.2.-) (Leucine-rich repeat and sterile alpha motif-containing protein 1) (Tsg101-associated ligase) (hTAL). | |||||
|
PLXA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 8) | NC score | 0.023419 (rank : 7) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UIW2 | Gene names | PLXNA1, NOV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A1 precursor (Semaphorin receptor NOV). | |||||
|
PLXA1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 9) | NC score | 0.023354 (rank : 8) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P70206, Q5DTR0 | Gene names | Plxna1, KIAA4053 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A1 precursor (Plexin-1) (Plex 1). | |||||
|
CIAS1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 10) | NC score | 0.005714 (rank : 16) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96P20, O75434, Q59H68, Q5JQS8, Q5JQS9, Q6TG35, Q8TCW0, Q8TEU9, Q8WXH9 | Gene names | CIAS1, NALP3, PYPAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Cold autoinflammatory syndrome 1 protein (Cryopyrin) (NACHT-, LRR- and PYD-containing protein 3) (PYRIN-containing APAF1-like protein 1) (Angiotensin/vasopressin receptor AII/AVP-like). | |||||
|
PLCA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 11) | NC score | 0.012664 (rank : 15) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99943, Q5BL03 | Gene names | AGPAT1, G15 | |||
|
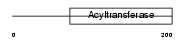
Domain Architecture |
|
|||||
| Description | 1-acyl-sn-glycerol-3-phosphate acyltransferase alpha (EC 2.3.1.51) (1- AGP acyltransferase 1) (1-AGPAT 1) (Lysophosphatidic acid acyltransferase-alpha) (LPAAT-alpha) (1-acylglycerol-3-phosphate O- acyltransferase 1) (Protein G15). | |||||
|
PLXA2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 12) | NC score | 0.022696 (rank : 11) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O75051, Q6UX61, Q96GN9, Q9BRL1, Q9UIW1 | Gene names | PLXNA2, KIAA0463, OCT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A2 precursor (Semaphorin receptor OCT). | |||||
|
PLXA2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 13) | NC score | 0.022706 (rank : 10) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P70207, Q6NVE6, Q6PHN4, Q80TZ7, Q8R1I4 | Gene names | Plxna2, Kiaa0463 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A2 precursor (Plexin-2) (Plex 2). | |||||
|
PLXA3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 14) | NC score | 0.022783 (rank : 9) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P51805 | Gene names | PLXNA3, PLXN4, SEX | |||
|
Domain Architecture |

|
|||||
| Description | Plexin-A3 precursor (Plexin-4) (Semaphorin receptor SEX). | |||||
|
PLXA4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 15) | NC score | 0.022559 (rank : 13) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9HCM2, Q6UWC6, Q6ZW89, Q8N969, Q8ND00, Q8NEN3, Q9NTD4 | Gene names | PLXNA4, KIAA1550, PLXNA4A, PLXNA4B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A4 precursor. | |||||
|
PLXA4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 16) | NC score | 0.022583 (rank : 12) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80UG2, Q5DTW8, Q8BKK9 | Gene names | Plxna4, Kiaa1550 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A4 precursor. | |||||
|
GNPAT_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P98192, Q9WUT6 | Gene names | Gnpat, Dhapat | |||
|
Domain Architecture |

|
|||||
| Description | Dihydroxyacetone phosphate acyltransferase (EC 2.3.1.42) (DHAP-AT) (DAP-AT) (Glycerone-phosphate O-acyltransferase) (Acyl- CoA:dihydroxyacetonephosphateacyltransferase). | |||||
|
GNPAT_HUMAN
|
||||||
| NC score | 0.997123 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O15228 | Gene names | GNPAT, DAPAT, DHAPAT | |||
|
Domain Architecture |

|
|||||
| Description | Dihydroxyacetone phosphate acyltransferase (EC 2.3.1.42) (DHAP-AT) (DAP-AT) (Glycerone-phosphate O-acyltransferase) (Acyl- CoA:dihydroxyacetonephosphateacyltransferase). | |||||
|
PLSB_MOUSE
|
||||||
| NC score | 0.705728 (rank : 3) | θ value | 1.6247e-33 (rank : 3) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61586 | Gene names | Gpam | |||
|
Domain Architecture |

|
|||||
| Description | Glycerol-3-phosphate acyltransferase, mitochondrial precursor (EC 2.3.1.15) (GPAT) (P90). | |||||
|
PLSB_HUMAN
|
||||||
| NC score | 0.704596 (rank : 4) | θ value | 6.17384e-33 (rank : 4) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9HCL2, Q5VW51 | Gene names | GPAM, KIAA1560 | |||
|
Domain Architecture |

|
|||||
| Description | Glycerol-3-phosphate acyltransferase, mitochondrial precursor (EC 2.3.1.15) (GPAT). | |||||
|
FHOD1_MOUSE
|
||||||
| NC score | 0.043016 (rank : 5) | θ value | 0.813845 (rank : 5) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6P9Q4, Q8BMK2 | Gene names | Fhod1, Fhos1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
KCMB2_HUMAN
|
||||||
| NC score | 0.029955 (rank : 6) | θ value | 1.38821 (rank : 6) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y691 | Gene names | KCNMB2 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit beta 2 (Calcium-activated potassium channel, subfamily M subunit beta 2) (Maxi K channel subunit beta 2) (BK channel subunit beta 2) (Slo-beta 2) (K(VCA)beta 2) (Charybdotoxin receptor subunit beta 2) (BKbeta2) (Hbeta2) (Hbeta3). | |||||
|
PLXA1_HUMAN
|
||||||
| NC score | 0.023419 (rank : 7) | θ value | 6.88961 (rank : 8) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UIW2 | Gene names | PLXNA1, NOV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A1 precursor (Semaphorin receptor NOV). | |||||
|
PLXA1_MOUSE
|
||||||
| NC score | 0.023354 (rank : 8) | θ value | 6.88961 (rank : 9) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P70206, Q5DTR0 | Gene names | Plxna1, KIAA4053 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A1 precursor (Plexin-1) (Plex 1). | |||||
|
PLXA3_HUMAN
|
||||||
| NC score | 0.022783 (rank : 9) | θ value | 8.99809 (rank : 14) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P51805 | Gene names | PLXNA3, PLXN4, SEX | |||
|
Domain Architecture |

|
|||||
| Description | Plexin-A3 precursor (Plexin-4) (Semaphorin receptor SEX). | |||||
|
PLXA2_MOUSE
|
||||||
| NC score | 0.022706 (rank : 10) | θ value | 8.99809 (rank : 13) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P70207, Q6NVE6, Q6PHN4, Q80TZ7, Q8R1I4 | Gene names | Plxna2, Kiaa0463 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A2 precursor (Plexin-2) (Plex 2). | |||||
|
PLXA2_HUMAN
|
||||||
| NC score | 0.022696 (rank : 11) | θ value | 8.99809 (rank : 12) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O75051, Q6UX61, Q96GN9, Q9BRL1, Q9UIW1 | Gene names | PLXNA2, KIAA0463, OCT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A2 precursor (Semaphorin receptor OCT). | |||||
|
PLXA4_MOUSE
|
||||||
| NC score | 0.022583 (rank : 12) | θ value | 8.99809 (rank : 16) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80UG2, Q5DTW8, Q8BKK9 | Gene names | Plxna4, Kiaa1550 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A4 precursor. | |||||
|
PLXA4_HUMAN
|
||||||
| NC score | 0.022559 (rank : 13) | θ value | 8.99809 (rank : 15) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9HCM2, Q6UWC6, Q6ZW89, Q8N969, Q8ND00, Q8NEN3, Q9NTD4 | Gene names | PLXNA4, KIAA1550, PLXNA4A, PLXNA4B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A4 precursor. | |||||
|
LRSM1_HUMAN
|
||||||
| NC score | 0.013358 (rank : 14) | θ value | 2.36792 (rank : 7) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 1078 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6UWE0, Q5VVV0, Q8NB40, Q96GT5, Q96MX5, Q96MZ7 | Gene names | LRSAM1, TAL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein LRSAM1 (EC 6.3.2.-) (Leucine-rich repeat and sterile alpha motif-containing protein 1) (Tsg101-associated ligase) (hTAL). | |||||
|
PLCA_HUMAN
|
||||||
| NC score | 0.012664 (rank : 15) | θ value | 8.99809 (rank : 11) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99943, Q5BL03 | Gene names | AGPAT1, G15 | |||
|
Domain Architecture |

|
|||||
| Description | 1-acyl-sn-glycerol-3-phosphate acyltransferase alpha (EC 2.3.1.51) (1- AGP acyltransferase 1) (1-AGPAT 1) (Lysophosphatidic acid acyltransferase-alpha) (LPAAT-alpha) (1-acylglycerol-3-phosphate O- acyltransferase 1) (Protein G15). | |||||
|
CIAS1_HUMAN
|
||||||
| NC score | 0.005714 (rank : 16) | θ value | 8.99809 (rank : 10) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96P20, O75434, Q59H68, Q5JQS8, Q5JQS9, Q6TG35, Q8TCW0, Q8TEU9, Q8WXH9 | Gene names | CIAS1, NALP3, PYPAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Cold autoinflammatory syndrome 1 protein (Cryopyrin) (NACHT-, LRR- and PYD-containing protein 3) (PYRIN-containing APAF1-like protein 1) (Angiotensin/vasopressin receptor AII/AVP-like). | |||||