Please be patient as the page loads
|
CIAS1_HUMAN
|
||||||
| SwissProt Accessions | Q96P20, O75434, Q59H68, Q5JQS8, Q5JQS9, Q6TG35, Q8TCW0, Q8TEU9, Q8WXH9 | Gene names | CIAS1, NALP3, PYPAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Cold autoinflammatory syndrome 1 protein (Cryopyrin) (NACHT-, LRR- and PYD-containing protein 3) (PYRIN-containing APAF1-like protein 1) (Angiotensin/vasopressin receptor AII/AVP-like). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CIAS1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q96P20, O75434, Q59H68, Q5JQS8, Q5JQS9, Q6TG35, Q8TCW0, Q8TEU9, Q8WXH9 | Gene names | CIAS1, NALP3, PYPAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Cold autoinflammatory syndrome 1 protein (Cryopyrin) (NACHT-, LRR- and PYD-containing protein 3) (PYRIN-containing APAF1-like protein 1) (Angiotensin/vasopressin receptor AII/AVP-like). | |||||
|
CIAS1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.996781 (rank : 2) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8R4B8, Q6JEL0 | Gene names | Cias1, Mmig1, Nalp3, Pypaf1 | |||
|
Domain Architecture |

|
|||||
| Description | Cold autoinflammatory syndrome 1 protein homolog (PYRIN-containing APAF1-like protein 1) (Mast cell maturation-associated-inducible protein 1) (Cryopyrin). | |||||
|
NAL12_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.987696 (rank : 3) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P59046, Q8NEU4, Q9BY26 | Gene names | NALP12, PYPAF7, RNO | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 12 (PYRIN-containing APAF1-like protein 7) (Monarch-1) (Regulated by nitric oxide). | |||||
|
NAL14_HUMAN
|
||||||
| θ value | 6.35588e-163 (rank : 4) | NC score | 0.974196 (rank : 4) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q86W24, Q7RTR6 | Gene names | NALP14, NOD5 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 14 (Nucleotide-binding oligomerization domain protein 5). | |||||
|
NALP4_HUMAN
|
||||||
| θ value | 2.689e-137 (rank : 5) | NC score | 0.971967 (rank : 5) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q96MN2, Q86W87, Q96AY6 | Gene names | NALP4, PAN2, PYPAF4, RNH2 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 4 (PYRIN-containing APAF1-like protein 4) (PAAD and NACHT-containing protein 2) (PYRIN and NACHT- containing protein 2) (Ribonuclease inhibitor 2). | |||||
|
NALP2_HUMAN
|
||||||
| θ value | 3.28708e-135 (rank : 6) | NC score | 0.967306 (rank : 6) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9NX02, Q53FL5, Q59G09, Q8IXT0, Q9BVN5, Q9H6G6, Q9HAV9, Q9NWK3 | Gene names | NALP2, NBS1, PAN1, PYPAF2 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 2 (PYRIN domain and NACHT domain-containing protein 1) (PYRIN-containing APAF1-like protein 2) (Nucleotide-binding site protein 1). | |||||
|
NAL13_HUMAN
|
||||||
| θ value | 1.33764e-128 (rank : 7) | NC score | 0.963862 (rank : 7) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q86W25, Q7RTR5 | Gene names | NALP13, NOD14 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 13 (Nucleotide-binding oligomerization domain protein 14). | |||||
|
NALP1_HUMAN
|
||||||
| θ value | 8.11517e-126 (rank : 8) | NC score | 0.925572 (rank : 13) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9C000, Q9BZZ8, Q9BZZ9, Q9HAV8, Q9UFT4, Q9Y2E0 | Gene names | NALP1, CARD7, DEFCAP, KIAA0926, NAC | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 1 (Death effector filament- forming ced-4-like apoptosis protein) (Nucleotide-binding domain and caspase recruitment domain) (Caspase recruitment domain protein 7). | |||||
|
NALP5_HUMAN
|
||||||
| θ value | 4.16784e-122 (rank : 9) | NC score | 0.958051 (rank : 9) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P59047, Q86W29 | Gene names | NALP5, MATER | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 5 (Mater protein homolog). | |||||
|
NALP5_MOUSE
|
||||||
| θ value | 9.30652e-114 (rank : 10) | NC score | 0.889632 (rank : 16) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9R1M5, Q9JLR2 | Gene names | Nalp5, Mater | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 5 (Maternal antigen that embryos require) (Mater protein) (Ooplasm-specific protein 1) (OP1). | |||||
|
NALP7_HUMAN
|
||||||
| θ value | 2.89973e-107 (rank : 11) | NC score | 0.960195 (rank : 8) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8WX94, Q7RTR1 | Gene names | NALP7, NOD12, PYPAF3 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 7 (PYRIN-containing APAF1-like protein 3). | |||||
|
NALP6_HUMAN
|
||||||
| θ value | 3.66817e-102 (rank : 12) | NC score | 0.950265 (rank : 10) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P59044 | Gene names | NALP6, PYPAF5 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 6 (PYRIN-containing APAF1-like protein 5). | |||||
|
NALP6_MOUSE
|
||||||
| θ value | 6.25694e-102 (rank : 13) | NC score | 0.948892 (rank : 11) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q91WS2, Q8K0L4 | Gene names | Nalp6, Pypaf5 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 6 (PYRIN-containing APAF1-like protein 5-like). | |||||
|
NAL11_HUMAN
|
||||||
| θ value | 1.88391e-98 (rank : 14) | NC score | 0.946033 (rank : 12) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P59045, Q53ZZ0, Q8NBF5 | Gene names | NALP11, NOD17, PYPAF6 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 11 (PYRIN-containing APAF1-like protein 6) (Nucleotide-binding oligomerization domain protein 17). | |||||
|
NAL10_HUMAN
|
||||||
| θ value | 6.27148e-94 (rank : 15) | NC score | 0.921601 (rank : 14) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q86W26, Q6JGT0 | Gene names | NALP10, NOD8, PYNOD | |||
|
Domain Architecture |
|
|||||
| Description | NACHT, LRR and PYD-containing protein 10 (Nucleotide-binding oligomerization domain protein 8). | |||||
|
NAL10_MOUSE
|
||||||
| θ value | 6.50502e-83 (rank : 16) | NC score | 0.915736 (rank : 15) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8CCN1 | Gene names | Nalp10, Pynod | |||
|
Domain Architecture |
|
|||||
| Description | NACHT, LRR and PYD-containing protein 10. | |||||
|
RINI_MOUSE
|
||||||
| θ value | 3.0428e-56 (rank : 17) | NC score | 0.869170 (rank : 18) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q91VI7, Q924P4 | Gene names | Rnh1, Rnh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribonuclease inhibitor (Ribonuclease/angiogenin inhibitor 1). | |||||
|
RINI_HUMAN
|
||||||
| θ value | 6.34462e-54 (rank : 18) | NC score | 0.872956 (rank : 17) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P13489 | Gene names | RNH1, PRI, RNH | |||
|
Domain Architecture |
|
|||||
| Description | Ribonuclease inhibitor (Ribonuclease/angiogenin inhibitor 1) (RAI) (Placental ribonuclease inhibitor) (RNase inhibitor) (RI). | |||||
|
CAR15_MOUSE
|
||||||
| θ value | 4.89182e-30 (rank : 19) | NC score | 0.758073 (rank : 19) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8K3Z0 | Gene names | Card15, Nod2 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 15 (Protein Nod2). | |||||
|
CARD4_HUMAN
|
||||||
| θ value | 5.06226e-27 (rank : 20) | NC score | 0.717949 (rank : 22) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9Y239, Q8IWF5 | Gene names | CARD4, NOD1 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 4 (Protein Nod1). | |||||
|
CARD4_MOUSE
|
||||||
| θ value | 6.61148e-27 (rank : 21) | NC score | 0.719134 (rank : 21) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8BHB0, Q8BUT6 | Gene names | Card4 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 4. | |||||
|
CAR15_HUMAN
|
||||||
| θ value | 1.92365e-26 (rank : 22) | NC score | 0.743559 (rank : 20) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9HC29, Q96RH5, Q96RH6, Q96RH8 | Gene names | CARD15, IBD1, NOD2 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 15 (Protein Nod2) (Inflammatory bowel disease protein 1). | |||||
|
C2TA_MOUSE
|
||||||
| θ value | 8.95645e-16 (rank : 23) | NC score | 0.645103 (rank : 23) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P79621, O46787, O78036, O78109, Q31115, Q9TPP1 | Gene names | Ciita, C2ta, Mhc2ta | |||
|
Domain Architecture |

|
|||||
| Description | MHC class II transactivator (CIITA). | |||||
|
C2TA_HUMAN
|
||||||
| θ value | 5.43371e-13 (rank : 24) | NC score | 0.619055 (rank : 24) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P33076 | Gene names | CIITA, MHC2TA | |||
|
Domain Architecture |

|
|||||
| Description | MHC class II transactivator (CIITA). | |||||
|
RGP1_HUMAN
|
||||||
| θ value | 1.25267e-09 (rank : 25) | NC score | 0.391840 (rank : 29) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P46060 | Gene names | RANGAP1 | |||
|
Domain Architecture |
|
|||||
| Description | Ran GTPase-activating protein 1. | |||||
|
LRC34_HUMAN
|
||||||
| θ value | 8.97725e-08 (rank : 26) | NC score | 0.545100 (rank : 25) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8IZ02 | Gene names | LRRC34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 34. | |||||
|
RGP1_MOUSE
|
||||||
| θ value | 4.45536e-07 (rank : 27) | NC score | 0.373298 (rank : 30) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P46061, Q60801 | Gene names | Rangap1, Fug1 | |||
|
Domain Architecture |
|
|||||
| Description | Ran GTPase-activating protein 1. | |||||
|
MEFV_MOUSE
|
||||||
| θ value | 5.81887e-07 (rank : 28) | NC score | 0.192048 (rank : 40) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JJ26 | Gene names | Mefv | |||
|
Domain Architecture |

|
|||||
| Description | Pyrin (Marenostrin). | |||||
|
MEFV_HUMAN
|
||||||
| θ value | 9.92553e-07 (rank : 29) | NC score | 0.104522 (rank : 44) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O15553, Q96PN4, Q96PN5 | Gene names | MEFV, MEF | |||
|
Domain Architecture |

|
|||||
| Description | Pyrin (Marenostrin). | |||||
|
LRC34_MOUSE
|
||||||
| θ value | 2.88788e-06 (rank : 30) | NC score | 0.512051 (rank : 26) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9DAM1 | Gene names | Lrrc34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 34. | |||||
|
BIRC1_HUMAN
|
||||||
| θ value | 9.29e-05 (rank : 31) | NC score | 0.236951 (rank : 34) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q13075, O75857, Q13730, Q99796 | Gene names | BIRC1, NAIP | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1 (Neuronal apoptosis inhibitory protein). | |||||
|
BIR1A_MOUSE
|
||||||
| θ value | 0.000121331 (rank : 32) | NC score | 0.218889 (rank : 35) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9QWK5, Q9JIB5, Q9R017 | Gene names | Birc1a, Naip, Naip1 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1a (Neuronal apoptosis inhibitory protein 1). | |||||
|
LRC31_HUMAN
|
||||||
| θ value | 0.000121331 (rank : 33) | NC score | 0.500090 (rank : 27) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6UY01, Q8N848, Q9H5N5 | Gene names | LRRC31 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 31. | |||||
|
CAR12_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 34) | NC score | 0.330275 (rank : 31) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9NPP4, Q96J81, Q96J82, Q96J83 | Gene names | CARD12, CLAN, CLAN1 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 12 (Ice protease- activating factor) (Ipaf) (CARD, LRR, and NACHT-containing protein) (Clan protein). | |||||
|
LRC45_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 35) | NC score | 0.160071 (rank : 42) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 933 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96CN5 | Gene names | LRRC45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 45. | |||||
|
BIR1E_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 36) | NC score | 0.209084 (rank : 37) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9R016, O09121, O09122, P81703, Q9R029 | Gene names | Birc1e, Naip-rs3, Naip5 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1e (Neuronal apoptosis inhibitory protein 5). | |||||
|
PHLPP_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 37) | NC score | 0.040647 (rank : 49) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 604 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O60346, Q641Q7, Q6P4C4, Q6PJI6, Q86TN6, Q96FK2, Q9NUY1 | Gene names | PHLPP, KIAA0606, PLEKHE1, SCOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein) (hSCOP). | |||||
|
ASC_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 38) | NC score | 0.319060 (rank : 32) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9EPB4, Q9D2W9 | Gene names | Pycard, Asc | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-associated speck-like protein containing a CARD (mASC) (PYD and CARD domain-containing protein). | |||||
|
BIR1F_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 39) | NC score | 0.210635 (rank : 36) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9JIB6, O09121, O09122, P81704 | Gene names | Birc1f, Naip-rs4, Naip6 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1f (Neuronal apoptosis inhibitory protein 6). | |||||
|
BIR1B_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 40) | NC score | 0.197412 (rank : 39) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9QUK4, O09124, Q9R030 | Gene names | Birc1b, Naip-rs6, Naip2 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1b (Neuronal apoptosis inhibitory protein 2). | |||||
|
BIR1G_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 41) | NC score | 0.202591 (rank : 38) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9JIB3 | Gene names | Birc1g, Naip7 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1g (Neuronal apoptosis inhibitory protein 7). | |||||
|
LRC8D_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 42) | NC score | 0.032774 (rank : 51) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q7L1W4, Q6UWB2, Q9NVW3 | Gene names | LRRC8D, LRRC5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 8D. | |||||
|
LRC8D_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 43) | NC score | 0.029259 (rank : 52) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BGR2, Q3UVA9, Q8CI13 | Gene names | Lrrc8d, Lrrc5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 8D. | |||||
|
PHLPP_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 44) | NC score | 0.043312 (rank : 48) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8CHE4, Q6PCN0, Q8QZU8, Q8R5E5, Q99KL6 | Gene names | Phlpp, Kiaa0606, Plekhe1, Scop | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein). | |||||
|
CF154_HUMAN
|
||||||
| θ value | 0.125558 (rank : 45) | NC score | 0.482997 (rank : 28) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q5JTD7 | Gene names | C6orf154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf154. | |||||
|
ASC_HUMAN
|
||||||
| θ value | 0.163984 (rank : 46) | NC score | 0.264723 (rank : 33) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9ULZ3, Q96D12, Q9BSZ5, Q9HBD0, Q9NXJ8 | Gene names | PYCARD, ASC, CARD5, TMS1 | |||
|
Domain Architecture |
|
|||||
| Description | Apoptosis-associated speck-like protein containing a CARD (hASC) (PYD and CARD domain-containing protein) (Target of methylation-induced silencing 1) (Caspase recruitment domain-containing protein 5). | |||||
|
LRC45_MOUSE
|
||||||
| θ value | 0.279714 (rank : 47) | NC score | 0.170080 (rank : 41) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8CIM1, Q3U5Z2 | Gene names | Lrrc45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 45. | |||||
|
GP1BA_HUMAN
|
||||||
| θ value | 1.81305 (rank : 48) | NC score | 0.025486 (rank : 53) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 407 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P07359, Q14441, Q16469, Q8N1F3, Q8NG39, Q9HDC7, Q9UEK1, Q9UQS4 | Gene names | GP1BA | |||
|
Domain Architecture |

|
|||||
| Description | Platelet glycoprotein Ib alpha chain precursor (Glycoprotein Ibalpha) (GP-Ib alpha) (GPIbA) (GPIb-alpha) (Antigen CD42b-alpha) (CD42b antigen) [Contains: Glycocalicin]. | |||||
|
LRC15_MOUSE
|
||||||
| θ value | 1.81305 (rank : 49) | NC score | 0.023727 (rank : 54) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q80X72 | Gene names | Lrrc15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 15 precursor. | |||||
|
FLRT1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.010289 (rank : 66) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9NZU1 | Gene names | FLRT1 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat transmembrane protein FLRT1 precursor (Fibronectin-like domain-containing leucine-rich transmembrane protein 1). | |||||
|
MYST2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.011674 (rank : 63) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O95251 | Gene names | MYST2, HBO1, HBOa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST2 (EC 2.3.1.48) (MYST protein 2) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 2) (Histone acetyltransferase binding to ORC1). | |||||
|
MYST2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 52) | NC score | 0.011674 (rank : 62) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5SVQ0, Q5SVQ1, Q5SVQ2, Q5SVQ3, Q5SVQ7, Q6PGC6, Q80Y65 | Gene names | Myst2, Hbo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST2 (EC 2.3.1.48) (MYST protein 2) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 2) (Histone acetyltransferase binding to ORC1). | |||||
|
AN32B_HUMAN
|
||||||
| θ value | 3.0926 (rank : 53) | NC score | 0.011555 (rank : 65) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92688, O00655, P78458, P78459 | Gene names | ANP32B, APRIL, PHAPI2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member B (PHAPI2 protein) (Silver-stainable protein SSP29) (Acidic protein rich in leucines). | |||||
|
ATCAY_HUMAN
|
||||||
| θ value | 3.0926 (rank : 54) | NC score | 0.008358 (rank : 70) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q86WG3, Q8NAQ2, Q8TAQ3, Q96HC6, Q96JF5 | Gene names | ATCAY, KIAA1872 | |||
|
Domain Architecture |

|
|||||
| Description | Caytaxin (Ataxia Cayman type protein) (BNIP-H). | |||||
|
FBXL2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.009220 (rank : 68) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BH16, Q8BXM4 | Gene names | Fbxl2, Fbl2 | |||
|
Domain Architecture |
|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2). | |||||
|
ZC3H6_HUMAN
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.011597 (rank : 64) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P61129 | Gene names | ZC3H6, KIAA2035, ZC3HDC6 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
A20A1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 57) | NC score | 0.001918 (rank : 77) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 698 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5TYW2, Q9H0H6 | Gene names | ANKRD20A1, ANKRD20A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 20A1. | |||||
|
A20A2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 58) | NC score | 0.002018 (rank : 76) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 716 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q4UJ75 | Gene names | ANKRD20A2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 20A2. | |||||
|
LRC15_HUMAN
|
||||||
| θ value | 4.03905 (rank : 59) | NC score | 0.018669 (rank : 56) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8TF66, Q7RTN7 | Gene names | LRRC15, LIB | |||
|
Domain Architecture |
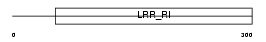
|
|||||
| Description | Leucine-rich repeat-containing protein 15 precursor (hLib). | |||||
|
ODBB_HUMAN
|
||||||
| θ value | 4.03905 (rank : 60) | NC score | 0.012687 (rank : 60) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P21953 | Gene names | BCKDHB | |||
|
Domain Architecture |
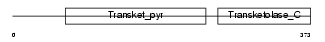
|
|||||
| Description | 2-oxoisovalerate dehydrogenase subunit beta, mitochondrial precursor (EC 1.2.4.4) (Branched-chain alpha-keto acid dehydrogenase E1 component beta chain) (BCKDH E1-beta). | |||||
|
CTBL1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.012295 (rank : 61) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WYA6, Q53HI8, Q5JWZ2, Q5JWZ3, Q5JWZ7, Q5JWZ8, Q8N454, Q8NCL2, Q8TBD6, Q96KD2, Q9H7A5, Q9NQF9, Q9NTX0, Q9Y3M7 | Gene names | CTNNBL1, C20orf33 | |||
|
Domain Architecture |
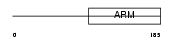
|
|||||
| Description | Beta-catenin-like protein 1 (Nuclear-associated protein) (NAP) (Testis development protein NYD-SP19). | |||||
|
FMOD_MOUSE
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.009875 (rank : 67) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P50608 | Gene names | Fmod | |||
|
Domain Architecture |

|
|||||
| Description | Fibromodulin precursor (FM) (Collagen-binding 59 kDa protein) (Keratan sulfate proteoglycan fibromodulin) (KSPG fibromodulin). | |||||
|
LRC10_MOUSE
|
||||||
| θ value | 5.27518 (rank : 63) | NC score | 0.016007 (rank : 57) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K3W2, Q8BUL2 | Gene names | Lrrc10, Hrlrrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 10 (Heart-restricted leucine- rich repeat protein). | |||||
|
LRC42_MOUSE
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.039746 (rank : 50) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8R2U7, Q3TE62, Q9CTP9 | Gene names | Lrrc42 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 42. | |||||
|
PIDD_HUMAN
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.021729 (rank : 55) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9HB75, Q59FD1, Q59H10, Q59HC7, Q7Z4P8, Q8NC89, Q8NDL2, Q96C25, Q9NRE6 | Gene names | LRDD, PIDD | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat and death domain-containing protein (p53-induced protein with a death domain). | |||||
|
WD51B_HUMAN
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | -0.000432 (rank : 80) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TC44 | Gene names | WDR51B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 51B. | |||||
|
AN32C_HUMAN
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.014090 (rank : 58) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43423 | Gene names | ANP32C, PP32R1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member C (Phosphoprotein 32-related protein 1) (Tumorigenic protein pp32r1). | |||||
|
CD180_HUMAN
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.008891 (rank : 69) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q99467 | Gene names | CD180, LY64, RP105 | |||
|
Domain Architecture |

|
|||||
| Description | CD180 antigen precursor (Lymphocyte antigen 64) (Radioprotective 105 kDa protein). | |||||
|
CEP35_HUMAN
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.003110 (rank : 73) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5VT06, O75068, Q8TDK3, Q8WY20 | Gene names | CEP350, CAP350, GM133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein 350 (Centrosome-associated protein of 350 kDa). | |||||
|
CTBL1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 70) | NC score | 0.013405 (rank : 59) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CWL8, Q3UL41, Q80X64, Q8VHE3 | Gene names | Ctnnbl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-catenin-like protein 1 (Nuclear-associated protein) (NAP). | |||||
|
EMR1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 71) | NC score | -0.000413 (rank : 79) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 382 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14246, Q8NGA7 | Gene names | EMR1, TM7LN3 | |||
|
Domain Architecture |

|
|||||
| Description | EGF-like module-containing mucin-like hormone receptor-like 1 precursor (Cell surface glycoprotein EMR1) (EMR1 hormone receptor). | |||||
|
FBXL2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.007308 (rank : 71) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UKC9, Q9NVQ8, Q9UK27, Q9UKA5, Q9Y3Y9 | Gene names | FBXL2, FBL2, FBL3 | |||
|
Domain Architecture |
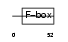
|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2) (F-box protein FBL2/FBL3). | |||||
|
GNPAT_MOUSE
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.005714 (rank : 72) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P98192, Q9WUT6 | Gene names | Gnpat, Dhapat | |||
|
Domain Architecture |
|
|||||
| Description | Dihydroxyacetone phosphate acyltransferase (EC 2.3.1.42) (DHAP-AT) (DAP-AT) (Glycerone-phosphate O-acyltransferase) (Acyl- CoA:dihydroxyacetonephosphateacyltransferase). | |||||
|
NMDE4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.002586 (rank : 74) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15399 | Gene names | GRIN2D | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 4 precursor (N-methyl D- aspartate receptor subtype 2D) (NR2D) (NMDAR2D) (EB11). | |||||
|
NMDE4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.002516 (rank : 75) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q03391 | Gene names | Grin2d | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 4 precursor (N-methyl D- aspartate receptor subtype 2D) (NR2D) (NMDAR2D). | |||||
|
TRPC7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.001832 (rank : 78) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9HCX4 | Gene names | TRPC7, TRP7 | |||
|
Domain Architecture |
|
|||||
| Description | Short transient receptor potential channel 7 (TrpC7) (TRP7 protein). | |||||
|
CARD8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 77) | NC score | 0.125653 (rank : 43) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y2G2, Q6PGP8, Q96P82 | Gene names | CARD8, KIAA0955, NDPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 8 (Apoptotic protein NDPP1) (DACAR) (CARD-inhibitor of NF-kappa-B-activating ligand) (CARDINAL) (Tumor-up-regulated CARD-containing antagonist of CASP9) (TUCAN). | |||||
|
CD14_MOUSE
|
||||||
| θ value | θ > 10 (rank : 78) | NC score | 0.052860 (rank : 46) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P10810 | Gene names | Cd14 | |||
|
Domain Architecture |
|
|||||
| Description | Monocyte differentiation antigen CD14 precursor (Myeloid cell-specific leucine-rich glycoprotein). | |||||
|
LRC14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 79) | NC score | 0.082781 (rank : 45) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15048 | Gene names | LRRC14, KIAA0014 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat-containing protein 14. | |||||
|
TMOD2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 80) | NC score | 0.052608 (rank : 47) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NZR1 | Gene names | TMOD2, NTMOD | |||
|
Domain Architecture |

|
|||||
| Description | Tropomodulin-2 (Neuronal tropomodulin) (N-Tmod). | |||||
|
CIAS1_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q96P20, O75434, Q59H68, Q5JQS8, Q5JQS9, Q6TG35, Q8TCW0, Q8TEU9, Q8WXH9 | Gene names | CIAS1, NALP3, PYPAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Cold autoinflammatory syndrome 1 protein (Cryopyrin) (NACHT-, LRR- and PYD-containing protein 3) (PYRIN-containing APAF1-like protein 1) (Angiotensin/vasopressin receptor AII/AVP-like). | |||||
|
CIAS1_MOUSE
|
||||||
| NC score | 0.996781 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8R4B8, Q6JEL0 | Gene names | Cias1, Mmig1, Nalp3, Pypaf1 | |||
|
Domain Architecture |

|
|||||
| Description | Cold autoinflammatory syndrome 1 protein homolog (PYRIN-containing APAF1-like protein 1) (Mast cell maturation-associated-inducible protein 1) (Cryopyrin). | |||||
|
NAL12_HUMAN
|
||||||
| NC score | 0.987696 (rank : 3) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P59046, Q8NEU4, Q9BY26 | Gene names | NALP12, PYPAF7, RNO | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 12 (PYRIN-containing APAF1-like protein 7) (Monarch-1) (Regulated by nitric oxide). | |||||
|
NAL14_HUMAN
|
||||||
| NC score | 0.974196 (rank : 4) | θ value | 6.35588e-163 (rank : 4) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q86W24, Q7RTR6 | Gene names | NALP14, NOD5 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 14 (Nucleotide-binding oligomerization domain protein 5). | |||||
|
NALP4_HUMAN
|
||||||
| NC score | 0.971967 (rank : 5) | θ value | 2.689e-137 (rank : 5) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q96MN2, Q86W87, Q96AY6 | Gene names | NALP4, PAN2, PYPAF4, RNH2 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 4 (PYRIN-containing APAF1-like protein 4) (PAAD and NACHT-containing protein 2) (PYRIN and NACHT- containing protein 2) (Ribonuclease inhibitor 2). | |||||
|
NALP2_HUMAN
|
||||||
| NC score | 0.967306 (rank : 6) | θ value | 3.28708e-135 (rank : 6) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9NX02, Q53FL5, Q59G09, Q8IXT0, Q9BVN5, Q9H6G6, Q9HAV9, Q9NWK3 | Gene names | NALP2, NBS1, PAN1, PYPAF2 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 2 (PYRIN domain and NACHT domain-containing protein 1) (PYRIN-containing APAF1-like protein 2) (Nucleotide-binding site protein 1). | |||||
|
NAL13_HUMAN
|
||||||
| NC score | 0.963862 (rank : 7) | θ value | 1.33764e-128 (rank : 7) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q86W25, Q7RTR5 | Gene names | NALP13, NOD14 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 13 (Nucleotide-binding oligomerization domain protein 14). | |||||
|
NALP7_HUMAN
|
||||||
| NC score | 0.960195 (rank : 8) | θ value | 2.89973e-107 (rank : 11) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8WX94, Q7RTR1 | Gene names | NALP7, NOD12, PYPAF3 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 7 (PYRIN-containing APAF1-like protein 3). | |||||
|
NALP5_HUMAN
|
||||||
| NC score | 0.958051 (rank : 9) | θ value | 4.16784e-122 (rank : 9) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P59047, Q86W29 | Gene names | NALP5, MATER | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 5 (Mater protein homolog). | |||||
|
NALP6_HUMAN
|
||||||
| NC score | 0.950265 (rank : 10) | θ value | 3.66817e-102 (rank : 12) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P59044 | Gene names | NALP6, PYPAF5 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 6 (PYRIN-containing APAF1-like protein 5). | |||||
|
NALP6_MOUSE
|
||||||
| NC score | 0.948892 (rank : 11) | θ value | 6.25694e-102 (rank : 13) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q91WS2, Q8K0L4 | Gene names | Nalp6, Pypaf5 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 6 (PYRIN-containing APAF1-like protein 5-like). | |||||
|
NAL11_HUMAN
|
||||||
| NC score | 0.946033 (rank : 12) | θ value | 1.88391e-98 (rank : 14) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P59045, Q53ZZ0, Q8NBF5 | Gene names | NALP11, NOD17, PYPAF6 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 11 (PYRIN-containing APAF1-like protein 6) (Nucleotide-binding oligomerization domain protein 17). | |||||
|
NALP1_HUMAN
|
||||||
| NC score | 0.925572 (rank : 13) | θ value | 8.11517e-126 (rank : 8) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9C000, Q9BZZ8, Q9BZZ9, Q9HAV8, Q9UFT4, Q9Y2E0 | Gene names | NALP1, CARD7, DEFCAP, KIAA0926, NAC | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 1 (Death effector filament- forming ced-4-like apoptosis protein) (Nucleotide-binding domain and caspase recruitment domain) (Caspase recruitment domain protein 7). | |||||
|
NAL10_HUMAN
|
||||||
| NC score | 0.921601 (rank : 14) | θ value | 6.27148e-94 (rank : 15) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q86W26, Q6JGT0 | Gene names | NALP10, NOD8, PYNOD | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 10 (Nucleotide-binding oligomerization domain protein 8). | |||||
|
NAL10_MOUSE
|
||||||
| NC score | 0.915736 (rank : 15) | θ value | 6.50502e-83 (rank : 16) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8CCN1 | Gene names | Nalp10, Pynod | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 10. | |||||
|
NALP5_MOUSE
|
||||||
| NC score | 0.889632 (rank : 16) | θ value | 9.30652e-114 (rank : 10) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9R1M5, Q9JLR2 | Gene names | Nalp5, Mater | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 5 (Maternal antigen that embryos require) (Mater protein) (Ooplasm-specific protein 1) (OP1). | |||||
|
RINI_HUMAN
|
||||||
| NC score | 0.872956 (rank : 17) | θ value | 6.34462e-54 (rank : 18) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P13489 | Gene names | RNH1, PRI, RNH | |||
|
Domain Architecture |

|
|||||
| Description | Ribonuclease inhibitor (Ribonuclease/angiogenin inhibitor 1) (RAI) (Placental ribonuclease inhibitor) (RNase inhibitor) (RI). | |||||
|
RINI_MOUSE
|
||||||
| NC score | 0.869170 (rank : 18) | θ value | 3.0428e-56 (rank : 17) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q91VI7, Q924P4 | Gene names | Rnh1, Rnh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribonuclease inhibitor (Ribonuclease/angiogenin inhibitor 1). | |||||
|
CAR15_MOUSE
|
||||||
| NC score | 0.758073 (rank : 19) | θ value | 4.89182e-30 (rank : 19) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8K3Z0 | Gene names | Card15, Nod2 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 15 (Protein Nod2). | |||||
|
CAR15_HUMAN
|
||||||
| NC score | 0.743559 (rank : 20) | θ value | 1.92365e-26 (rank : 22) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9HC29, Q96RH5, Q96RH6, Q96RH8 | Gene names | CARD15, IBD1, NOD2 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 15 (Protein Nod2) (Inflammatory bowel disease protein 1). | |||||
|
CARD4_MOUSE
|
||||||
| NC score | 0.719134 (rank : 21) | θ value | 6.61148e-27 (rank : 21) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8BHB0, Q8BUT6 | Gene names | Card4 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 4. | |||||
|
CARD4_HUMAN
|
||||||
| NC score | 0.717949 (rank : 22) | θ value | 5.06226e-27 (rank : 20) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9Y239, Q8IWF5 | Gene names | CARD4, NOD1 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 4 (Protein Nod1). | |||||
|
C2TA_MOUSE
|
||||||
| NC score | 0.645103 (rank : 23) | θ value | 8.95645e-16 (rank : 23) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P79621, O46787, O78036, O78109, Q31115, Q9TPP1 | Gene names | Ciita, C2ta, Mhc2ta | |||
|
Domain Architecture |

|
|||||
| Description | MHC class II transactivator (CIITA). | |||||
|
C2TA_HUMAN
|
||||||
| NC score | 0.619055 (rank : 24) | θ value | 5.43371e-13 (rank : 24) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P33076 | Gene names | CIITA, MHC2TA | |||
|
Domain Architecture |

|
|||||
| Description | MHC class II transactivator (CIITA). | |||||
|
LRC34_HUMAN
|
||||||
| NC score | 0.545100 (rank : 25) | θ value | 8.97725e-08 (rank : 26) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8IZ02 | Gene names | LRRC34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 34. | |||||
|
LRC34_MOUSE
|
||||||
| NC score | 0.512051 (rank : 26) | θ value | 2.88788e-06 (rank : 30) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9DAM1 | Gene names | Lrrc34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 34. | |||||
|
LRC31_HUMAN
|
||||||
| NC score | 0.500090 (rank : 27) | θ value | 0.000121331 (rank : 33) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6UY01, Q8N848, Q9H5N5 | Gene names | LRRC31 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 31. | |||||
|
CF154_HUMAN
|
||||||
| NC score | 0.482997 (rank : 28) | θ value | 0.125558 (rank : 45) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q5JTD7 | Gene names | C6orf154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf154. | |||||
|
RGP1_HUMAN
|
||||||
| NC score | 0.391840 (rank : 29) | θ value | 1.25267e-09 (rank : 25) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P46060 | Gene names | RANGAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ran GTPase-activating protein 1. | |||||
|
RGP1_MOUSE
|
||||||
| NC score | 0.373298 (rank : 30) | θ value | 4.45536e-07 (rank : 27) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P46061, Q60801 | Gene names | Rangap1, Fug1 | |||
|
Domain Architecture |

|
|||||
| Description | Ran GTPase-activating protein 1. | |||||
|
CAR12_HUMAN
|
||||||
| NC score | 0.330275 (rank : 31) | θ value | 0.000786445 (rank : 34) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9NPP4, Q96J81, Q96J82, Q96J83 | Gene names | CARD12, CLAN, CLAN1 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 12 (Ice protease- activating factor) (Ipaf) (CARD, LRR, and NACHT-containing protein) (Clan protein). | |||||
|
ASC_MOUSE
|
||||||
| NC score | 0.319060 (rank : 32) | θ value | 0.0148317 (rank : 38) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9EPB4, Q9D2W9 | Gene names | Pycard, Asc | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-associated speck-like protein containing a CARD (mASC) (PYD and CARD domain-containing protein). | |||||
|
ASC_HUMAN
|
||||||
| NC score | 0.264723 (rank : 33) | θ value | 0.163984 (rank : 46) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9ULZ3, Q96D12, Q9BSZ5, Q9HBD0, Q9NXJ8 | Gene names | PYCARD, ASC, CARD5, TMS1 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-associated speck-like protein containing a CARD (hASC) (PYD and CARD domain-containing protein) (Target of methylation-induced silencing 1) (Caspase recruitment domain-containing protein 5). | |||||
|
BIRC1_HUMAN
|
||||||
| NC score | 0.236951 (rank : 34) | θ value | 9.29e-05 (rank : 31) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q13075, O75857, Q13730, Q99796 | Gene names | BIRC1, NAIP | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1 (Neuronal apoptosis inhibitory protein). | |||||
|
BIR1A_MOUSE
|
||||||
| NC score | 0.218889 (rank : 35) | θ value | 0.000121331 (rank : 32) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9QWK5, Q9JIB5, Q9R017 | Gene names | Birc1a, Naip, Naip1 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1a (Neuronal apoptosis inhibitory protein 1). | |||||
|
BIR1F_MOUSE
|
||||||
| NC score | 0.210635 (rank : 36) | θ value | 0.0148317 (rank : 39) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9JIB6, O09121, O09122, P81704 | Gene names | Birc1f, Naip-rs4, Naip6 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1f (Neuronal apoptosis inhibitory protein 6). | |||||
|
BIR1E_MOUSE
|
||||||
| NC score | 0.209084 (rank : 37) | θ value | 0.00665767 (rank : 36) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9R016, O09121, O09122, P81703, Q9R029 | Gene names | Birc1e, Naip-rs3, Naip5 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1e (Neuronal apoptosis inhibitory protein 5). | |||||
|
BIR1G_MOUSE
|
||||||
| NC score | 0.202591 (rank : 38) | θ value | 0.0193708 (rank : 41) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9JIB3 | Gene names | Birc1g, Naip7 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1g (Neuronal apoptosis inhibitory protein 7). | |||||
|
BIR1B_MOUSE
|
||||||
| NC score | 0.197412 (rank : 39) | θ value | 0.0193708 (rank : 40) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9QUK4, O09124, Q9R030 | Gene names | Birc1b, Naip-rs6, Naip2 | |||
|
Domain Architecture |

|
|||||
| Description | Baculoviral IAP repeat-containing protein 1b (Neuronal apoptosis inhibitory protein 2). | |||||
|
MEFV_MOUSE
|
||||||
| NC score | 0.192048 (rank : 40) | θ value | 5.81887e-07 (rank : 28) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JJ26 | Gene names | Mefv | |||
|
Domain Architecture |

|
|||||
| Description | Pyrin (Marenostrin). | |||||
|
LRC45_MOUSE
|
||||||
| NC score | 0.170080 (rank : 41) | θ value | 0.279714 (rank : 47) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8CIM1, Q3U5Z2 | Gene names | Lrrc45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 45. | |||||
|
LRC45_HUMAN
|
||||||
| NC score | 0.160071 (rank : 42) | θ value | 0.00390308 (rank : 35) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 933 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96CN5 | Gene names | LRRC45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 45. | |||||
|
CARD8_HUMAN
|
||||||
| NC score | 0.125653 (rank : 43) | θ value | θ > 10 (rank : 77) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y2G2, Q6PGP8, Q96P82 | Gene names | CARD8, KIAA0955, NDPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 8 (Apoptotic protein NDPP1) (DACAR) (CARD-inhibitor of NF-kappa-B-activating ligand) (CARDINAL) (Tumor-up-regulated CARD-containing antagonist of CASP9) (TUCAN). | |||||
|
MEFV_HUMAN
|
||||||
| NC score | 0.104522 (rank : 44) | θ value | 9.92553e-07 (rank : 29) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O15553, Q96PN4, Q96PN5 | Gene names | MEFV, MEF | |||
|
Domain Architecture |

|
|||||
| Description | Pyrin (Marenostrin). | |||||
|
LRC14_HUMAN
|
||||||
| NC score | 0.082781 (rank : 45) | θ value | θ > 10 (rank : 79) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15048 | Gene names | LRRC14, KIAA0014 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat-containing protein 14. | |||||
|
CD14_MOUSE
|
||||||
| NC score | 0.052860 (rank : 46) | θ value | θ > 10 (rank : 78) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P10810 | Gene names | Cd14 | |||
|
Domain Architecture |

|
|||||
| Description | Monocyte differentiation antigen CD14 precursor (Myeloid cell-specific leucine-rich glycoprotein). | |||||
|
TMOD2_HUMAN
|
||||||
| NC score | 0.052608 (rank : 47) | θ value | θ > 10 (rank : 80) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NZR1 | Gene names | TMOD2, NTMOD | |||
|
Domain Architecture |

|
|||||
| Description | Tropomodulin-2 (Neuronal tropomodulin) (N-Tmod). | |||||
|
PHLPP_MOUSE
|
||||||
| NC score | 0.043312 (rank : 48) | θ value | 0.0431538 (rank : 44) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8CHE4, Q6PCN0, Q8QZU8, Q8R5E5, Q99KL6 | Gene names | Phlpp, Kiaa0606, Plekhe1, Scop | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein). | |||||
|
PHLPP_HUMAN
|
||||||
| NC score | 0.040647 (rank : 49) | θ value | 0.0113563 (rank : 37) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 604 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O60346, Q641Q7, Q6P4C4, Q6PJI6, Q86TN6, Q96FK2, Q9NUY1 | Gene names | PHLPP, KIAA0606, PLEKHE1, SCOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein) (hSCOP). | |||||
|
LRC42_MOUSE
|
||||||
| NC score | 0.039746 (rank : 50) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8R2U7, Q3TE62, Q9CTP9 | Gene names | Lrrc42 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 42. | |||||
|
LRC8D_HUMAN
|
||||||
| NC score | 0.032774 (rank : 51) | θ value | 0.0193708 (rank : 42) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q7L1W4, Q6UWB2, Q9NVW3 | Gene names | LRRC8D, LRRC5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 8D. | |||||
|
LRC8D_MOUSE
|
||||||
| NC score | 0.029259 (rank : 52) | θ value | 0.0252991 (rank : 43) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BGR2, Q3UVA9, Q8CI13 | Gene names | Lrrc8d, Lrrc5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 8D. | |||||
|
GP1BA_HUMAN
|
||||||
| NC score | 0.025486 (rank : 53) | θ value | 1.81305 (rank : 48) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 407 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P07359, Q14441, Q16469, Q8N1F3, Q8NG39, Q9HDC7, Q9UEK1, Q9UQS4 | Gene names | GP1BA | |||
|
Domain Architecture |

|
|||||
| Description | Platelet glycoprotein Ib alpha chain precursor (Glycoprotein Ibalpha) (GP-Ib alpha) (GPIbA) (GPIb-alpha) (Antigen CD42b-alpha) (CD42b antigen) [Contains: Glycocalicin]. | |||||
|
LRC15_MOUSE
|
||||||
| NC score | 0.023727 (rank : 54) | θ value | 1.81305 (rank : 49) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q80X72 | Gene names | Lrrc15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 15 precursor. | |||||
|
PIDD_HUMAN
|
||||||
| NC score | 0.021729 (rank : 55) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9HB75, Q59FD1, Q59H10, Q59HC7, Q7Z4P8, Q8NC89, Q8NDL2, Q96C25, Q9NRE6 | Gene names | LRDD, PIDD | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat and death domain-containing protein (p53-induced protein with a death domain). | |||||
|
LRC15_HUMAN
|
||||||
| NC score | 0.018669 (rank : 56) | θ value | 4.03905 (rank : 59) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8TF66, Q7RTN7 | Gene names | LRRC15, LIB | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat-containing protein 15 precursor (hLib). | |||||
|
LRC10_MOUSE
|
||||||
| NC score | 0.016007 (rank : 57) | θ value | 5.27518 (rank : 63) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K3W2, Q8BUL2 | Gene names | Lrrc10, Hrlrrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 10 (Heart-restricted leucine- rich repeat protein). | |||||
|
AN32C_HUMAN
|
||||||
| NC score | 0.014090 (rank : 58) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43423 | Gene names | ANP32C, PP32R1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member C (Phosphoprotein 32-related protein 1) (Tumorigenic protein pp32r1). | |||||
|
CTBL1_MOUSE
|
||||||
| NC score | 0.013405 (rank : 59) | θ value | 8.99809 (rank : 70) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CWL8, Q3UL41, Q80X64, Q8VHE3 | Gene names | Ctnnbl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-catenin-like protein 1 (Nuclear-associated protein) (NAP). | |||||
|
ODBB_HUMAN
|
||||||
| NC score | 0.012687 (rank : 60) | θ value | 4.03905 (rank : 60) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P21953 | Gene names | BCKDHB | |||
|
Domain Architecture |

|
|||||
| Description | 2-oxoisovalerate dehydrogenase subunit beta, mitochondrial precursor (EC 1.2.4.4) (Branched-chain alpha-keto acid dehydrogenase E1 component beta chain) (BCKDH E1-beta). | |||||
|
CTBL1_HUMAN
|
||||||
| NC score | 0.012295 (rank : 61) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WYA6, Q53HI8, Q5JWZ2, Q5JWZ3, Q5JWZ7, Q5JWZ8, Q8N454, Q8NCL2, Q8TBD6, Q96KD2, Q9H7A5, Q9NQF9, Q9NTX0, Q9Y3M7 | Gene names | CTNNBL1, C20orf33 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-catenin-like protein 1 (Nuclear-associated protein) (NAP) (Testis development protein NYD-SP19). | |||||
|
MYST2_MOUSE
|
||||||
| NC score | 0.011674 (rank : 62) | θ value | 2.36792 (rank : 52) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5SVQ0, Q5SVQ1, Q5SVQ2, Q5SVQ3, Q5SVQ7, Q6PGC6, Q80Y65 | Gene names | Myst2, Hbo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST2 (EC 2.3.1.48) (MYST protein 2) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 2) (Histone acetyltransferase binding to ORC1). | |||||
|
MYST2_HUMAN
|
||||||
| NC score | 0.011674 (rank : 63) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O95251 | Gene names | MYST2, HBO1, HBOa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST2 (EC 2.3.1.48) (MYST protein 2) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 2) (Histone acetyltransferase binding to ORC1). | |||||
|
ZC3H6_HUMAN
|
||||||
| NC score | 0.011597 (rank : 64) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P61129 | Gene names | ZC3H6, KIAA2035, ZC3HDC6 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
AN32B_HUMAN
|
||||||
| NC score | 0.011555 (rank : 65) | θ value | 3.0926 (rank : 53) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92688, O00655, P78458, P78459 | Gene names | ANP32B, APRIL, PHAPI2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member B (PHAPI2 protein) (Silver-stainable protein SSP29) (Acidic protein rich in leucines). | |||||
|
FLRT1_HUMAN
|
||||||
| NC score | 0.010289 (rank : 66) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9NZU1 | Gene names | FLRT1 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat transmembrane protein FLRT1 precursor (Fibronectin-like domain-containing leucine-rich transmembrane protein 1). | |||||
|
FMOD_MOUSE
|
||||||
| NC score | 0.009875 (rank : 67) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P50608 | Gene names | Fmod | |||
|
Domain Architecture |

|
|||||
| Description | Fibromodulin precursor (FM) (Collagen-binding 59 kDa protein) (Keratan sulfate proteoglycan fibromodulin) (KSPG fibromodulin). | |||||
|
FBXL2_MOUSE
|
||||||
| NC score | 0.009220 (rank : 68) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BH16, Q8BXM4 | Gene names | Fbxl2, Fbl2 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2). | |||||
|
CD180_HUMAN
|
||||||
| NC score | 0.008891 (rank : 69) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q99467 | Gene names | CD180, LY64, RP105 | |||
|
Domain Architecture |

|
|||||
| Description | CD180 antigen precursor (Lymphocyte antigen 64) (Radioprotective 105 kDa protein). | |||||
|
ATCAY_HUMAN
|
||||||
| NC score | 0.008358 (rank : 70) | θ value | 3.0926 (rank : 54) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q86WG3, Q8NAQ2, Q8TAQ3, Q96HC6, Q96JF5 | Gene names | ATCAY, KIAA1872 | |||
|
Domain Architecture |

|
|||||
| Description | Caytaxin (Ataxia Cayman type protein) (BNIP-H). | |||||
|
FBXL2_HUMAN
|
||||||
| NC score | 0.007308 (rank : 71) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UKC9, Q9NVQ8, Q9UK27, Q9UKA5, Q9Y3Y9 | Gene names | FBXL2, FBL2, FBL3 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2) (F-box protein FBL2/FBL3). | |||||
|
GNPAT_MOUSE
|
||||||
| NC score | 0.005714 (rank : 72) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P98192, Q9WUT6 | Gene names | Gnpat, Dhapat | |||
|
Domain Architecture |

|
|||||
| Description | Dihydroxyacetone phosphate acyltransferase (EC 2.3.1.42) (DHAP-AT) (DAP-AT) (Glycerone-phosphate O-acyltransferase) (Acyl- CoA:dihydroxyacetonephosphateacyltransferase). | |||||
|
CEP35_HUMAN
|
||||||
| NC score | 0.003110 (rank : 73) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5VT06, O75068, Q8TDK3, Q8WY20 | Gene names | CEP350, CAP350, GM133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein 350 (Centrosome-associated protein of 350 kDa). | |||||
|
NMDE4_HUMAN
|
||||||
| NC score | 0.002586 (rank : 74) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15399 | Gene names | GRIN2D | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 4 precursor (N-methyl D- aspartate receptor subtype 2D) (NR2D) (NMDAR2D) (EB11). | |||||
|
NMDE4_MOUSE
|
||||||
| NC score | 0.002516 (rank : 75) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q03391 | Gene names | Grin2d | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 4 precursor (N-methyl D- aspartate receptor subtype 2D) (NR2D) (NMDAR2D). | |||||
|
A20A2_HUMAN
|
||||||
| NC score | 0.002018 (rank : 76) | θ value | 4.03905 (rank : 58) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 716 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q4UJ75 | Gene names | ANKRD20A2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 20A2. | |||||
|
A20A1_HUMAN
|
||||||
| NC score | 0.001918 (rank : 77) | θ value | 4.03905 (rank : 57) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 698 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5TYW2, Q9H0H6 | Gene names | ANKRD20A1, ANKRD20A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 20A1. | |||||
|
TRPC7_HUMAN
|
||||||
| NC score | 0.001832 (rank : 78) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9HCX4 | Gene names | TRPC7, TRP7 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 7 (TrpC7) (TRP7 protein). | |||||
|
EMR1_HUMAN
|
||||||
| NC score | -0.000413 (rank : 79) | θ value | 8.99809 (rank : 71) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 382 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14246, Q8NGA7 | Gene names | EMR1, TM7LN3 | |||
|
Domain Architecture |

|
|||||
| Description | EGF-like module-containing mucin-like hormone receptor-like 1 precursor (Cell surface glycoprotein EMR1) (EMR1 hormone receptor). | |||||
|
WD51B_HUMAN
|
||||||
| NC score | -0.000432 (rank : 80) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TC44 | Gene names | WDR51B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 51B. | |||||