Please be patient as the page loads
|
SEM3A_MOUSE
|
||||||
| SwissProt Accessions | O08665, Q5BL08, Q62180, Q62215 | Gene names | Sema3a, Semad, SemD | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3A precursor (Semaphorin III) (Sema III) (Semaphorin D) (Sema D). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SEM3A_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999158 (rank : 2) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q14563 | Gene names | SEMA3A | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3A precursor (Semaphorin III) (Sema III). | |||||
|
SEM3A_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 92 | |
| SwissProt Accessions | O08665, Q5BL08, Q62180, Q62215 | Gene names | Sema3a, Semad, SemD | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3A precursor (Semaphorin III) (Sema III) (Semaphorin D) (Sema D). | |||||
|
SEM3B_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.993476 (rank : 8) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q13214, Q8TB71, Q8TDV7, Q93018, Q96GX0 | Gene names | SEMA3B, SEMA5 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3B precursor (Semaphorin V) (Sema V) (Sema A(V)). | |||||
|
SEM3B_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.994191 (rank : 7) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q62177 | Gene names | Sema3b, Sema, Semaa | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3B precursor (Semaphorin A) (Sema A). | |||||
|
SEM3C_HUMAN
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.991719 (rank : 11) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q99985 | Gene names | SEMA3C, SEMAE | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3C precursor (Semaphorin E) (Sema E). | |||||
|
SEM3C_MOUSE
|
||||||
| θ value | 0 (rank : 6) | NC score | 0.991355 (rank : 12) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q62181 | Gene names | Sema3c, Semae, SemE | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3C precursor (Semaphorin E) (Sema E). | |||||
|
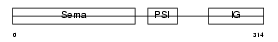
SEM3D_HUMAN
|
||||||
| θ value | 0 (rank : 7) | NC score | 0.994532 (rank : 3) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O95025, Q6UW77, Q8NCQ1 | Gene names | SEMA3D | |||
|
Domain Architecture |
|
|||||
| Description | Semaphorin-3D precursor. | |||||
|
SEM3D_MOUSE
|
||||||
| θ value | 0 (rank : 8) | NC score | 0.994398 (rank : 4) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8BH34 | Gene names | Sema3d | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-3D precursor. | |||||
|
SEM3E_HUMAN
|
||||||
| θ value | 0 (rank : 9) | NC score | 0.994388 (rank : 5) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | O15041, Q75M94, Q75M97 | Gene names | SEMA3E, KIAA0331 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-3E precursor. | |||||
|
SEM3E_MOUSE
|
||||||
| θ value | 0 (rank : 10) | NC score | 0.994309 (rank : 6) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P70275, O09078, O09079 | Gene names | Sema3e, Semah, Semh | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3E precursor (Semaphorin H) (Sema H). | |||||
|
SEM3F_HUMAN
|
||||||
| θ value | 0 (rank : 11) | NC score | 0.991777 (rank : 10) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q13275, Q13274, Q13372, Q15704 | Gene names | SEMA3F | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3F precursor (Semaphorin IV) (Sema IV) (Sema III/F). | |||||
|
SEM3F_MOUSE
|
||||||
| θ value | 0 (rank : 12) | NC score | 0.992449 (rank : 9) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O88632, O88633 | Gene names | Sema3f | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3F precursor (Semaphorin IV) (Sema IV). | |||||
|
SEM4D_HUMAN
|
||||||
| θ value | 4.05562e-101 (rank : 13) | NC score | 0.965191 (rank : 15) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q92854, Q7Z5S4 | Gene names | SEMA4D, CD100 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4D precursor (Leukocyte activation antigen CD100) (BB18) (A8) (GR3). | |||||
|
SEM4D_MOUSE
|
||||||
| θ value | 5.29683e-101 (rank : 14) | NC score | 0.965437 (rank : 14) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O09126 | Gene names | Sema4d, Semacl2, Semaj | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4D precursor (Semaphorin J) (Sema J) (Semaphorin C-like 2) (M-Sema G). | |||||
|
SEM4C_MOUSE
|
||||||
| θ value | 1.70393e-99 (rank : 15) | NC score | 0.967236 (rank : 13) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q64151 | Gene names | Sema4c, Semacl1, Semai | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4C precursor (Semaphorin I) (Sema I) (Semaphorin C-like 1) (M-Sema F). | |||||
|
SEM4G_MOUSE
|
||||||
| θ value | 8.4565e-99 (rank : 16) | NC score | 0.963665 (rank : 17) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9WUH7 | Gene names | Sema4g | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4G precursor. | |||||
|
SEM4C_HUMAN
|
||||||
| θ value | 1.59483e-97 (rank : 17) | NC score | 0.962752 (rank : 18) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9C0C4, Q7Z5X0 | Gene names | SEMA4C, KIAA1739 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4C precursor. | |||||
|
SEM4G_HUMAN
|
||||||
| θ value | 1.3501e-96 (rank : 18) | NC score | 0.963685 (rank : 16) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9NTN9, Q9HCF3 | Gene names | SEMA4G, KIAA1619 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4G precursor. | |||||
|
SEM4B_HUMAN
|
||||||
| θ value | 8.47616e-91 (rank : 19) | NC score | 0.959727 (rank : 21) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9NPR2, Q6UXE3, Q8WVP9, Q96FK5, Q9C0B8, Q9H691, Q9NPM8, Q9NPN0 | Gene names | SEMA4B, KIAA1745 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4B precursor. | |||||
|
SEM4B_MOUSE
|
||||||
| θ value | 1.44581e-90 (rank : 20) | NC score | 0.957957 (rank : 23) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q62179, Q4PKI6, Q69ZB7 | Gene names | Sema4b, Kiaa1745, Semac, SemC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-4B precursor (Semaphorin C) (Sema C). | |||||
|
SEM6A_HUMAN
|
||||||
| θ value | 1.07224e-85 (rank : 21) | NC score | 0.944306 (rank : 27) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9H2E6, Q9P2H9 | Gene names | SEMA6A, KIAA1368 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6A precursor (Semaphorin VIA) (Sema VIA) (Semaphorin 6A-1) (SEMA6A-1). | |||||
|
SEM6A_MOUSE
|
||||||
| θ value | 3.44927e-84 (rank : 22) | NC score | 0.943340 (rank : 29) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O35464, Q6P5A8, Q6PCN9, Q9EQ71 | Gene names | Sema6a, Semaq | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6A precursor (Semaphorin VIA) (Sema VIA) (Semaphorin 6A-1) (SEMA6A-1) (Semaphorin Q) (Sema Q). | |||||
|
SEM4A_HUMAN
|
||||||
| θ value | 1.44917e-82 (rank : 23) | NC score | 0.958143 (rank : 22) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9H3S1, Q8WUA9 | Gene names | SEMA4A, SEMB | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4A precursor (Semaphorin B) (Sema B). | |||||
|
SEM6D_MOUSE
|
||||||
| θ value | 1.44917e-82 (rank : 24) | NC score | 0.941613 (rank : 30) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q76KF0, Q76KF1, Q76KF2, Q76KF3, Q76KF4, Q80TD0 | Gene names | Sema6d, Kiaa1479 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-6D precursor. | |||||
|
SEM6D_HUMAN
|
||||||
| θ value | 2.47191e-82 (rank : 25) | NC score | 0.940768 (rank : 31) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8NFY4, Q8NFY3, Q8NFY5, Q8NFY6, Q8NFY7, Q9P249 | Gene names | SEMA6D, KIAA1479 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-6D precursor. | |||||
|
SEM4A_MOUSE
|
||||||
| θ value | 4.66183e-81 (rank : 26) | NC score | 0.956793 (rank : 24) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q62178 | Gene names | Sema4a, Semab, SemB | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4A precursor (Semaphorin B) (Sema B). | |||||
|
SEM5B_HUMAN
|
||||||
| θ value | 1.07473e-77 (rank : 27) | NC score | 0.793102 (rank : 35) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9P283, Q6UY12 | Gene names | SEMA5B, KIAA1445 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-5B precursor. | |||||
|
SEM5B_MOUSE
|
||||||
| θ value | 5.8972e-76 (rank : 28) | NC score | 0.789951 (rank : 37) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q60519 | Gene names | Sema5b, Semag, SemG | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-5B precursor (Semaphorin G) (Sema G). | |||||
|
SEM6C_HUMAN
|
||||||
| θ value | 6.52014e-75 (rank : 29) | NC score | 0.943900 (rank : 28) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9H3T2, Q5JR71, Q8WXT8, Q8WXT9, Q8WXU0, Q96JF8 | Gene names | SEMA6C, KIAA1869, SEMAY | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6C precursor (Semaphorin Y) (Sema Y). | |||||
|
SEM4F_MOUSE
|
||||||
| θ value | 9.41505e-74 (rank : 30) | NC score | 0.961172 (rank : 19) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9Z123, Q9R1Y1 | Gene names | Sema4f, Semaw | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4F precursor (Semaphorin W) (Sema W). | |||||
|
SEM5A_HUMAN
|
||||||
| θ value | 5.1662e-72 (rank : 31) | NC score | 0.788006 (rank : 38) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q13591, O60408, Q1RLL9 | Gene names | SEMA5A, SEMAF | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-5A precursor (Semaphorin F) (Sema F). | |||||
|
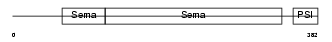
SEM4F_HUMAN
|
||||||
| θ value | 6.7473e-72 (rank : 32) | NC score | 0.960658 (rank : 20) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O95754, Q9NS35 | Gene names | SEMA4F, SEMAM, SEMAW | |||
|
Domain Architecture |
|
|||||
| Description | Semaphorin-4F precursor (Semaphorin W) (Sema W) (Semaphorin M) (Sema M). | |||||
|
SEM6C_MOUSE
|
||||||
| θ value | 6.7473e-72 (rank : 33) | NC score | 0.929542 (rank : 34) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9WTM3 | Gene names | Sema6c, Semay | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6C precursor (Semaphorin Y) (Sema Y). | |||||
|
SEM6B_MOUSE
|
||||||
| θ value | 1.96316e-71 (rank : 34) | NC score | 0.936164 (rank : 33) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O54951 | Gene names | Sema6b, Seman | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6B precursor (Semaphorin VIB) (Sema VIB) (Semaphorin N) (Sema N). | |||||
|
SEM5A_MOUSE
|
||||||
| θ value | 4.37348e-71 (rank : 35) | NC score | 0.792643 (rank : 36) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q62217 | Gene names | Sema5a, Semaf, SemF | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-5A precursor (Semaphorin F) (Sema F). | |||||
|
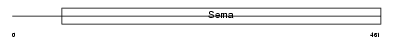
SEM6B_HUMAN
|
||||||
| θ value | 4.83544e-70 (rank : 36) | NC score | 0.937772 (rank : 32) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9H3T3, Q9NRK9 | Gene names | SEMA6B, SEMAZ | |||
|
Domain Architecture |
|
|||||
| Description | Semaphorin-6B precursor (Semaphorin Z) (Sema Z). | |||||
|
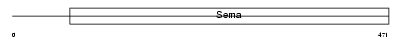
SEM7A_HUMAN
|
||||||
| θ value | 2.7521e-57 (rank : 37) | NC score | 0.948138 (rank : 25) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O75326 | Gene names | SEMA7A, CD108, SEMAL | |||
|
Domain Architecture |
|
|||||
| Description | Semaphorin-7A precursor (Semaphorin L) (Sema L) (Semaphorin K1) (Sema K1) (John-Milton-Hargen human blood group Ag) (JMH blood group antigen) (CD108 antigen) (CDw108). | |||||
|
SEM7A_MOUSE
|
||||||
| θ value | 7.49467e-55 (rank : 38) | NC score | 0.946119 (rank : 26) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9QUR8, O88371 | Gene names | Sema7a, Cd108, Semal, Semk1 | |||
|
Domain Architecture |
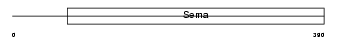
|
|||||
| Description | Semaphorin-7A precursor (Semaphorin L) (Sema L) (Semaphorin K1) (Sema K1) (CD108 antigen) (CDw108). | |||||
|
PLXA2_MOUSE
|
||||||
| θ value | 2.21117e-06 (rank : 39) | NC score | 0.307336 (rank : 44) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P70207, Q6NVE6, Q6PHN4, Q80TZ7, Q8R1I4 | Gene names | Plxna2, Kiaa0463 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A2 precursor (Plexin-2) (Plex 2). | |||||
|
PLXC1_MOUSE
|
||||||
| θ value | 1.43324e-05 (rank : 40) | NC score | 0.327541 (rank : 40) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9QZC2, Q8CGW1 | Gene names | Plxnc1, Vespr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-C1 precursor (Virus-encoded semaphorin protein receptor) (CD232 antigen). | |||||
|
PLXB2_HUMAN
|
||||||
| θ value | 3.19293e-05 (rank : 41) | NC score | 0.339747 (rank : 39) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O15031, Q7KZU3, Q9BSU7 | Gene names | PLXNB2, KIAA0315 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B2 precursor (MM1). | |||||
|
PLXA2_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 42) | NC score | 0.306365 (rank : 45) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O75051, Q6UX61, Q96GN9, Q9BRL1, Q9UIW1 | Gene names | PLXNA2, KIAA0463, OCT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A2 precursor (Semaphorin receptor OCT). | |||||
|
PLXA4_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 43) | NC score | 0.315221 (rank : 42) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q80UG2, Q5DTW8, Q8BKK9 | Gene names | Plxna4, Kiaa1550 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A4 precursor. | |||||
|
PLXA3_HUMAN
|
||||||
| θ value | 9.29e-05 (rank : 44) | NC score | 0.287244 (rank : 47) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P51805 | Gene names | PLXNA3, PLXN4, SEX | |||
|
Domain Architecture |

|
|||||
| Description | Plexin-A3 precursor (Plexin-4) (Semaphorin receptor SEX). | |||||
|
PLXC1_HUMAN
|
||||||
| θ value | 9.29e-05 (rank : 45) | NC score | 0.321588 (rank : 41) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O60486, Q59H25 | Gene names | PLXNC1, VESPR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-C1 precursor (Virus-encoded semaphorin protein receptor) (CD232 antigen). | |||||
|
PLXA4_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 46) | NC score | 0.314566 (rank : 43) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9HCM2, Q6UWC6, Q6ZW89, Q8N969, Q8ND00, Q8NEN3, Q9NTD4 | Gene names | PLXNA4, KIAA1550, PLXNA4A, PLXNA4B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A4 precursor. | |||||
|
PLXA1_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 47) | NC score | 0.293864 (rank : 46) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P70206, Q5DTR0 | Gene names | Plxna1, KIAA4053 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A1 precursor (Plexin-1) (Plex 1). | |||||
|
OBSCN_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 48) | NC score | 0.021245 (rank : 74) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
PLXA1_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 49) | NC score | 0.284082 (rank : 48) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9UIW2 | Gene names | PLXNA1, NOV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A1 precursor (Semaphorin receptor NOV). | |||||
|
PLXB1_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 50) | NC score | 0.275660 (rank : 49) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O43157, Q6NY20, Q9UIV7, Q9UJ92, Q9UJ93 | Gene names | PLXNB1, KIAA0407, SEP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B1 precursor (Semaphorin receptor SEP). | |||||
|
WDR35_HUMAN
|
||||||
| θ value | 0.125558 (rank : 51) | NC score | 0.028287 (rank : 70) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9P2L0, Q8NE11 | Gene names | WDR35, KIAA1336 | |||
|
Domain Architecture |
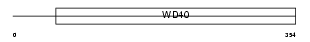
|
|||||
| Description | WD repeat protein 35. | |||||
|
WDR35_MOUSE
|
||||||
| θ value | 0.365318 (rank : 52) | NC score | 0.028099 (rank : 71) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BND3, Q8CDL4 | Gene names | Wdr35 | |||
|
Domain Architecture |
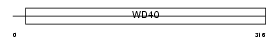
|
|||||
| Description | WD repeat protein 35. | |||||
|
AMGO1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 53) | NC score | 0.013950 (rank : 84) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 353 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q86WK6, Q8IW71, Q9ULQ7 | Gene names | AMIGO1, ALI2, AMIGO, KIAA1163 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amphoterin-induced protein 1 precursor (AMIGO-1) (Alivin-2). | |||||
|
DSCL1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 54) | NC score | 0.027673 (rank : 72) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8TD84, Q76MU9, Q8IZY3, Q8IZY4, Q8WXU7, Q9ULT7 | Gene names | DSCAML1, DSCAM2, KIAA1132 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Down syndrome cell adhesion molecule-like protein 1 precursor (Down syndrome cell adhesion molecule 2). | |||||
|
PLXB3_HUMAN
|
||||||
| θ value | 0.47712 (rank : 55) | NC score | 0.247148 (rank : 52) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9ULL4, Q9HDA4 | Gene names | PLXNB3, KIAA1206 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B3 precursor. | |||||
|
PLXB1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 56) | NC score | 0.274791 (rank : 50) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8CJH3, Q6ZQC3, Q80ZZ1 | Gene names | Plxnb1, Kiaa0407 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B1 precursor. | |||||
|
BTNL1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 57) | NC score | 0.015877 (rank : 80) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q7TST0, O70356 | Gene names | Btnl1, Btnl3, Gm316, Ng10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Butyrophilin-like protein 1 precursor. | |||||
|
PLXD1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 58) | NC score | 0.246477 (rank : 53) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9Y4D7, Q6PJS9, Q8IZJ2, Q9BTQ2 | Gene names | PLXND1, KIAA0620 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-D1 precursor. | |||||
|
JAML1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 59) | NC score | 0.021234 (rank : 75) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q86YT9, Q5DTC6, Q7Z499, Q8N9I7, Q8NF70 | Gene names | AMICA1, JAML | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctional adhesion molecule-like precursor (Dendritic cell-specific protein CREA7-1) (Adhesion molecule interacting with CXADR antigen 1). | |||||
|
C8AP2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 60) | NC score | 0.011691 (rank : 89) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9WUF3, Q9CSW2 | Gene names | Casp8ap2, Flash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
CNTN4_MOUSE
|
||||||
| θ value | 1.38821 (rank : 61) | NC score | 0.030912 (rank : 69) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q69Z26, Q8BRT6, Q8CBD3 | Gene names | Cntn4, Kiaa3024 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Contactin-4 precursor (Brain-derived immunoglobulin superfamily protein 2) (BIG-2). | |||||
|
IBP7_HUMAN
|
||||||
| θ value | 1.38821 (rank : 62) | NC score | 0.012702 (rank : 86) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q16270, Q07822 | Gene names | IGFBP7, MAC25, PSF | |||
|
Domain Architecture |
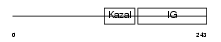
|
|||||
| Description | Insulin-like growth factor-binding protein 7 precursor (IGFBP-7) (IBP- 7) (IGF-binding protein 7) (MAC25 protein) (Prostacyclin-stimulating factor) (PGI2-stimulating factor) (IGFBP-rP1). | |||||
|
KIT_HUMAN
|
||||||
| θ value | 1.38821 (rank : 63) | NC score | -0.001418 (rank : 103) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 953 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P10721, Q9UM99 | Gene names | KIT | |||
|
Domain Architecture |

|
|||||
| Description | Mast/stem cell growth factor receptor precursor (EC 2.7.10.1) (SCFR) (Proto-oncogene tyrosine-protein kinase Kit) (c-kit) (CD117 antigen). | |||||
|
MINK1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 64) | NC score | -0.003120 (rank : 106) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N4C8, Q9P1X1, Q9P2R8 | Gene names | MINK1, MAP4K6, MINK | |||
|
Domain Architecture |

|
|||||
| Description | Misshapen-like kinase 1 (EC 2.7.11.1) (Mitogen-activated protein kinase kinase kinase kinase 6) (MAPK/ERK kinase kinase kinase 6) (MEK kinase kinase 6) (MEKKK 6) (Misshapen/NIK-related kinase) (GCK family kinase MiNK). | |||||
|
MINK1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | -0.002321 (rank : 105) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1451 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9JM52, Q921M6, Q9JM92 | Gene names | Mink1, Map4k6, Mink | |||
|
Domain Architecture |

|
|||||
| Description | Misshapen-like kinase 1 (EC 2.7.11.1) (Mitogen-activated protein kinase kinase kinase kinase 6) (MAPK/ERK kinase kinase kinase 6) (MEK kinase kinase 6) (MEKKK 6) (Misshapen/NIK-related kinase) (GCK family kinase MiNK). | |||||
|
NCA11_MOUSE
|
||||||
| θ value | 1.81305 (rank : 66) | NC score | 0.010711 (rank : 94) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 661 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P13595, Q61949 | Gene names | Ncam1, Ncam | |||
|
Domain Architecture |
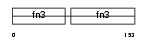
|
|||||
| Description | Neural cell adhesion molecule 1, 180 kDa isoform precursor (N-CAM 180) (NCAM-180). | |||||
|
NCA12_MOUSE
|
||||||
| θ value | 1.81305 (rank : 67) | NC score | 0.011627 (rank : 90) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 399 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P13594, Q61950 | Gene names | Ncam1, Ncam | |||
|
Domain Architecture |

|
|||||
| Description | Neural cell adhesion molecule 1, 120 kDa isoform precursor (N-CAM 120) (NCAM-120). | |||||
|
DUX1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.003337 (rank : 101) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43812 | Gene names | DUX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double homeobox protein 1. | |||||
|
DUX5_HUMAN
|
||||||
| θ value | 2.36792 (rank : 69) | NC score | 0.003469 (rank : 99) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96PT3 | Gene names | DUX5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double homeobox protein 5. | |||||
|
NCA11_HUMAN
|
||||||
| θ value | 2.36792 (rank : 70) | NC score | 0.011222 (rank : 93) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P13591, Q15829, Q16180 | Gene names | NCAM1, NCAM | |||
|
Domain Architecture |

|
|||||
| Description | Neural cell adhesion molecule 1, 140 kDa isoform precursor (N-CAM 140) (NCAM-140) (CD56 antigen). | |||||
|
NCA12_HUMAN
|
||||||
| θ value | 2.36792 (rank : 71) | NC score | 0.011424 (rank : 92) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 437 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P13592, P13593 | Gene names | NCAM1, NCAM | |||
|
Domain Architecture |

|
|||||
| Description | Neural cell adhesion molecule 1, 120 kDa isoform precursor (N-CAM 120) (NCAM-120) (CD56 antigen). | |||||
|
IBP7_MOUSE
|
||||||
| θ value | 3.0926 (rank : 72) | NC score | 0.011770 (rank : 88) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61581, O88812, Q9EQW0 | Gene names | Igfbp7, Mac25 | |||
|
Domain Architecture |
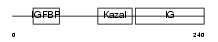
|
|||||
| Description | Insulin-like growth factor-binding protein 7 precursor (IGFBP-7) (IBP- 7) (IGF-binding protein 7) (MAC25 protein). | |||||
|
KIRR3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 73) | NC score | 0.013138 (rank : 85) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8IZU9, Q6UWJ9, Q6UWL5, Q96JG0 | Gene names | KIRREL3, KIAA1867, NEPH2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kin of IRRE-like protein 3 precursor (Kin of irregular chiasm-like protein 3) (Nephrin-like 2). | |||||
|
KIRR3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 74) | NC score | 0.012293 (rank : 87) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BR86, Q6P555, Q810H3, Q8BGQ5, Q8BRS8 | Gene names | Kirrel3, Kiaa1867 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kin of IRRE-like protein 3 precursor (Kin of irregular chiasm-like protein 3). | |||||
|
FGFR2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | -0.002106 (rank : 104) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1062 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P21802, P18443, Q01742, Q12922, Q14300, Q14301, Q14302, Q14303, Q14304, Q14305, Q14672, Q14718, Q14719, Q86YI4, Q96KL9, Q96KM0, Q96KM1, Q96KM2, Q9NZU2, Q9NZU3, Q9UD01, Q9UD02, Q9UIH3, Q9UIH4, Q9UIH5, Q9UIH6, Q9UIH7, Q9UIH8, Q9UM87, Q9UMC6, Q9UNS7, Q9UQH7, Q9UQH8, Q9UQH9, Q9UQI0 | Gene names | FGFR2, BEK, KSAM | |||
|
Domain Architecture |

|
|||||
| Description | Fibroblast growth factor receptor 2 precursor (EC 2.7.10.1) (FGFR-2) (Keratinocyte growth factor receptor 2) (CD332 antigen). | |||||
|
KIRR1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 76) | NC score | 0.015332 (rank : 82) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q96J84, Q5XKC6, Q7Z696, Q7Z7N8, Q8TB15, Q9H9N1, Q9NVA5 | Gene names | KIRREL, NEPH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kin of IRRE-like protein 1 precursor (Kin of irregular chiasm-like protein 1) (Nephrin-like protein 1). | |||||
|
DUX3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 77) | NC score | 0.002968 (rank : 102) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96PT4, Q9UND2 | Gene names | DUX3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative double homeobox protein 3. | |||||
|
KIRR1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 78) | NC score | 0.014796 (rank : 83) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q80W68, Q8CIJ4, Q8CJ59, Q923L4 | Gene names | Kirrel1, Kirrel, Neph1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kin of IRRE-like protein 1 precursor (Kin of irregular chiasm-like protein 1) (Nephrin-like protein 1). | |||||
|
TREM2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.019501 (rank : 78) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99NH8, Q8CGK4, Q8CIA6, Q99NH9, Q9JL34 | Gene names | Trem2, Trem2a, Trem2b, Trem2c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Triggering receptor expressed on myeloid cells 2 precursor (Triggering receptor expressed on monocytes 2) (TREM-2). | |||||
|
ZN596_HUMAN
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | -0.005593 (rank : 107) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 716 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TC21, O95015, Q8N9X0 | Gene names | ZNF596 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 596. | |||||
|
CC47_MOUSE
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.003456 (rank : 100) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D024, Q3TSB5, Q3V1S7, Q8C5D9, Q920S6 | Gene names | Ccdc47, Asp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 47 precursor (Adipocyte-specific protein 4). | |||||
|
CNTN4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.032996 (rank : 68) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 449 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8IWV2, Q8IX14, Q8TC35 | Gene names | CNTN4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Contactin-4 precursor (Brain-derived immunoglobulin superfamily protein 2) (BIG-2). | |||||
|
FGRL1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.009033 (rank : 95) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8N441, Q6PJN1, Q9BXN7, Q9H4D7 | Gene names | FGFRL1, FGFR5, FHFR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibroblast growth factor receptor-like 1 precursor (FGF receptor-like protein 1) (Fibroblast growth factor receptor 5) (FGFR-like protein) (FGF homologous factor receptor). | |||||
|
TREM2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.019686 (rank : 77) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NZC2, Q8N5H8, Q8WYN6 | Gene names | TREM2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Triggering receptor expressed on myeloid cells 2 precursor (Triggering receptor expressed on monocytes 2) (TREM-2). | |||||
|
BCLF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.003836 (rank : 97) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NYF8, Q14673, Q86WU6, Q86WY0 | Gene names | BCLAF1, BTF, KIAA0164 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bcl-2-associated transcription factor 1 (Btf). | |||||
|
BCLF1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.003676 (rank : 98) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K019, Q8BNZ0, Q8C2E9, Q9CSW5 | Gene names | Bclaf1, Btf, Kiaa0164 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bcl-2-associated transcription factor 1 (Btf). | |||||
|
CND2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.004458 (rank : 96) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15003, Q8TB87 | Gene names | NCAPH, BRRN, BRRN1, CAPH, KIAA0074 | |||
|
Domain Architecture |

|
|||||
| Description | Condensin complex subunit 2 (Non-SMC condensin I complex subunit H) (Barren homolog protein 1) (Chromosome-associated protein H) (hCAP-H) (XCAP-H homolog). | |||||
|
CNTN3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.020384 (rank : 76) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q07409 | Gene names | Cntn3, Pang, Pcs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Contactin-3 precursor (Brain-derived immunoglobulin superfamily protein 1) (BIG-1) (Plasmacytoma-associated neuronal glycoprotein). | |||||
|
IRPL2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.015860 (rank : 81) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NP60, Q9NZN0 | Gene names | IL1RAPL2, IL1R9 | |||
|
Domain Architecture |
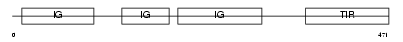
|
|||||
| Description | X-linked interleukin-1 receptor accessory protein-like 2 precursor (IL1RAPL-2-related protein) (Interleukin-1 receptor 9) (IL-1R9) (IL-1 receptor accessory protein-like 2) (Three immunoglobulin domain- containing IL-1 receptor-related 1) (TIGIRR-1). | |||||
|
IRPL2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.016534 (rank : 79) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9ERS6, Q9ER66 | Gene names | Il1rapl2 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked interleukin-1 receptor accessory protein-like 2 precursor (IL1RAPL-2-related protein) (TIGIRR-1). | |||||
|
OBSL1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.027258 (rank : 73) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O75147, Q96IW3 | Gene names | OBSL1, KIAA0657 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin-like protein 1 precursor. | |||||
|
ROBO2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.011624 (rank : 91) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q7TPD3, Q8BJ59 | Gene names | Robo2, Kiaa1568 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 2 precursor. | |||||
|
BAI1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.050994 (rank : 65) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 375 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O14514 | Gene names | BAI1 | |||
|
Domain Architecture |

|
|||||
| Description | Brain-specific angiogenesis inhibitor 1 precursor. | |||||
|
BAI1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.050651 (rank : 66) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 340 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3UHD1, Q3UH36, Q8CGM0 | Gene names | Bai1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain-specific angiogenesis inhibitor 1 precursor. | |||||
|
HMCN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.050439 (rank : 67) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 983 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q96RW7, Q5TYR7, Q96DN3, Q96DN8, Q96SC3 | Gene names | HMCN1, FIBL6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hemicentin-1 precursor (Fibulin-6) (FIBL-6). | |||||
|
PLXB3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.250204 (rank : 51) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9QY40, Q80TH8 | Gene names | Plxnb3, Kiaa1206, Plxn6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B3 precursor (Plexin-6). | |||||
|
PROP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.089252 (rank : 54) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P27918, O15134, O15135, O15136, O75826 | Gene names | CFP, PFC | |||
|
Domain Architecture |
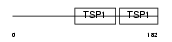
|
|||||
| Description | Properdin precursor (Factor P). | |||||
|
PROP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.086750 (rank : 55) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P11680, Q3TB98, Q3U779 | Gene names | Cfp, Pfc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Properdin precursor (Factor P). | |||||
|
SPON1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.064598 (rank : 59) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9HCB6, O94862, Q8NCD7, Q8WUR5 | Gene names | SPON1, KIAA0762, VSGP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spondin-1 precursor (F-spondin) (Vascular smooth muscle cell growth- promoting factor). | |||||
|
SPON1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.066110 (rank : 58) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VCC9, Q8K2Q8 | Gene names | Spon1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spondin-1 precursor (F-spondin). | |||||
|
UNC5A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.055072 (rank : 62) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8K1S4, Q6PEF7, Q80T71 | Gene names | Unc5a, Kiaa1976, Unc5h1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin receptor UNC5A precursor (Unc-5 homolog A) (Unc-5 homolog 1). | |||||
|
UNC5B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.058916 (rank : 60) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8IZJ1, Q86SN3, Q8N1Y2, Q9H9F3 | Gene names | UNC5B, P53RDL1, UNC5H2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin receptor UNC5B precursor (Unc-5 homolog B) (Unc-5 homolog 2) (p53-regulated receptor for death and life protein 1). | |||||
|
UNC5B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.058708 (rank : 61) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8K1S3, Q3U4F2, Q6PFH0, Q80Y85, Q9D398 | Gene names | Unc5b, Unc5h2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin receptor UNC5B precursor (Unc-5 homolog B) (Unc-5 homolog 2). | |||||
|
UNC5C_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.053708 (rank : 64) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O95185, Q8IUT0 | Gene names | UNC5C, UNC5H3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin receptor UNC5C precursor (Unc-5 homolog C) (Unc-5 homolog 3). | |||||
|
UNC5C_MOUSE
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.054735 (rank : 63) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O08747, Q8CD16 | Gene names | Unc5c, Rcm, Unc5h3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin receptor UNC5C precursor (Unc-5 homolog C) (Unc-5 homolog 3) (Rostral cerebellar malformation protein). | |||||
|
UNC5D_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.069698 (rank : 56) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6UXZ4, Q8WYP7 | Gene names | UNC5D, KIAA1777, UNC5H4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin receptor UNC5D precursor (Unc-5 homolog D) (Unc-5 homolog 4). | |||||
|
UNC5D_MOUSE
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.069580 (rank : 57) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8K1S2 | Gene names | Unc5d, Unc5h4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin receptor UNC5D precursor (Unc-5 homolog D) (Unc-5 homolog 4). | |||||
|
SEM3A_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 92 | |
| SwissProt Accessions | O08665, Q5BL08, Q62180, Q62215 | Gene names | Sema3a, Semad, SemD | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3A precursor (Semaphorin III) (Sema III) (Semaphorin D) (Sema D). | |||||
|
SEM3A_HUMAN
|
||||||
| NC score | 0.999158 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q14563 | Gene names | SEMA3A | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3A precursor (Semaphorin III) (Sema III). | |||||
|
SEM3D_HUMAN
|
||||||
| NC score | 0.994532 (rank : 3) | θ value | 0 (rank : 7) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O95025, Q6UW77, Q8NCQ1 | Gene names | SEMA3D | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3D precursor. | |||||
|
SEM3D_MOUSE
|
||||||
| NC score | 0.994398 (rank : 4) | θ value | 0 (rank : 8) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8BH34 | Gene names | Sema3d | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-3D precursor. | |||||
|
SEM3E_HUMAN
|
||||||
| NC score | 0.994388 (rank : 5) | θ value | 0 (rank : 9) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | O15041, Q75M94, Q75M97 | Gene names | SEMA3E, KIAA0331 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-3E precursor. | |||||
|
SEM3E_MOUSE
|
||||||
| NC score | 0.994309 (rank : 6) | θ value | 0 (rank : 10) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P70275, O09078, O09079 | Gene names | Sema3e, Semah, Semh | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3E precursor (Semaphorin H) (Sema H). | |||||
|
SEM3B_MOUSE
|
||||||
| NC score | 0.994191 (rank : 7) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q62177 | Gene names | Sema3b, Sema, Semaa | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3B precursor (Semaphorin A) (Sema A). | |||||
|
SEM3B_HUMAN
|
||||||
| NC score | 0.993476 (rank : 8) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q13214, Q8TB71, Q8TDV7, Q93018, Q96GX0 | Gene names | SEMA3B, SEMA5 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3B precursor (Semaphorin V) (Sema V) (Sema A(V)). | |||||
|
SEM3F_MOUSE
|
||||||
| NC score | 0.992449 (rank : 9) | θ value | 0 (rank : 12) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O88632, O88633 | Gene names | Sema3f | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3F precursor (Semaphorin IV) (Sema IV). | |||||
|
SEM3F_HUMAN
|
||||||
| NC score | 0.991777 (rank : 10) | θ value | 0 (rank : 11) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q13275, Q13274, Q13372, Q15704 | Gene names | SEMA3F | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3F precursor (Semaphorin IV) (Sema IV) (Sema III/F). | |||||
|
SEM3C_HUMAN
|
||||||
| NC score | 0.991719 (rank : 11) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q99985 | Gene names | SEMA3C, SEMAE | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3C precursor (Semaphorin E) (Sema E). | |||||
|
SEM3C_MOUSE
|
||||||
| NC score | 0.991355 (rank : 12) | θ value | 0 (rank : 6) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q62181 | Gene names | Sema3c, Semae, SemE | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-3C precursor (Semaphorin E) (Sema E). | |||||
|
SEM4C_MOUSE
|
||||||
| NC score | 0.967236 (rank : 13) | θ value | 1.70393e-99 (rank : 15) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q64151 | Gene names | Sema4c, Semacl1, Semai | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4C precursor (Semaphorin I) (Sema I) (Semaphorin C-like 1) (M-Sema F). | |||||
|
SEM4D_MOUSE
|
||||||
| NC score | 0.965437 (rank : 14) | θ value | 5.29683e-101 (rank : 14) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O09126 | Gene names | Sema4d, Semacl2, Semaj | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4D precursor (Semaphorin J) (Sema J) (Semaphorin C-like 2) (M-Sema G). | |||||
|
SEM4D_HUMAN
|
||||||
| NC score | 0.965191 (rank : 15) | θ value | 4.05562e-101 (rank : 13) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q92854, Q7Z5S4 | Gene names | SEMA4D, CD100 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4D precursor (Leukocyte activation antigen CD100) (BB18) (A8) (GR3). | |||||
|
SEM4G_HUMAN
|
||||||
| NC score | 0.963685 (rank : 16) | θ value | 1.3501e-96 (rank : 18) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9NTN9, Q9HCF3 | Gene names | SEMA4G, KIAA1619 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4G precursor. | |||||
|
SEM4G_MOUSE
|
||||||
| NC score | 0.963665 (rank : 17) | θ value | 8.4565e-99 (rank : 16) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9WUH7 | Gene names | Sema4g | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4G precursor. | |||||
|
SEM4C_HUMAN
|
||||||
| NC score | 0.962752 (rank : 18) | θ value | 1.59483e-97 (rank : 17) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9C0C4, Q7Z5X0 | Gene names | SEMA4C, KIAA1739 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4C precursor. | |||||
|
SEM4F_MOUSE
|
||||||
| NC score | 0.961172 (rank : 19) | θ value | 9.41505e-74 (rank : 30) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9Z123, Q9R1Y1 | Gene names | Sema4f, Semaw | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4F precursor (Semaphorin W) (Sema W). | |||||
|
SEM4F_HUMAN
|
||||||
| NC score | 0.960658 (rank : 20) | θ value | 6.7473e-72 (rank : 32) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O95754, Q9NS35 | Gene names | SEMA4F, SEMAM, SEMAW | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4F precursor (Semaphorin W) (Sema W) (Semaphorin M) (Sema M). | |||||
|
SEM4B_HUMAN
|
||||||
| NC score | 0.959727 (rank : 21) | θ value | 8.47616e-91 (rank : 19) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9NPR2, Q6UXE3, Q8WVP9, Q96FK5, Q9C0B8, Q9H691, Q9NPM8, Q9NPN0 | Gene names | SEMA4B, KIAA1745 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4B precursor. | |||||
|
SEM4A_HUMAN
|
||||||
| NC score | 0.958143 (rank : 22) | θ value | 1.44917e-82 (rank : 23) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9H3S1, Q8WUA9 | Gene names | SEMA4A, SEMB | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4A precursor (Semaphorin B) (Sema B). | |||||
|
SEM4B_MOUSE
|
||||||
| NC score | 0.957957 (rank : 23) | θ value | 1.44581e-90 (rank : 20) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q62179, Q4PKI6, Q69ZB7 | Gene names | Sema4b, Kiaa1745, Semac, SemC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-4B precursor (Semaphorin C) (Sema C). | |||||
|
SEM4A_MOUSE
|
||||||
| NC score | 0.956793 (rank : 24) | θ value | 4.66183e-81 (rank : 26) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q62178 | Gene names | Sema4a, Semab, SemB | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4A precursor (Semaphorin B) (Sema B). | |||||
|
SEM7A_HUMAN
|
||||||
| NC score | 0.948138 (rank : 25) | θ value | 2.7521e-57 (rank : 37) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O75326 | Gene names | SEMA7A, CD108, SEMAL | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-7A precursor (Semaphorin L) (Sema L) (Semaphorin K1) (Sema K1) (John-Milton-Hargen human blood group Ag) (JMH blood group antigen) (CD108 antigen) (CDw108). | |||||
|
SEM7A_MOUSE
|
||||||
| NC score | 0.946119 (rank : 26) | θ value | 7.49467e-55 (rank : 38) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9QUR8, O88371 | Gene names | Sema7a, Cd108, Semal, Semk1 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-7A precursor (Semaphorin L) (Sema L) (Semaphorin K1) (Sema K1) (CD108 antigen) (CDw108). | |||||
|
SEM6A_HUMAN
|
||||||
| NC score | 0.944306 (rank : 27) | θ value | 1.07224e-85 (rank : 21) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9H2E6, Q9P2H9 | Gene names | SEMA6A, KIAA1368 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6A precursor (Semaphorin VIA) (Sema VIA) (Semaphorin 6A-1) (SEMA6A-1). | |||||
|
SEM6C_HUMAN
|
||||||
| NC score | 0.943900 (rank : 28) | θ value | 6.52014e-75 (rank : 29) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9H3T2, Q5JR71, Q8WXT8, Q8WXT9, Q8WXU0, Q96JF8 | Gene names | SEMA6C, KIAA1869, SEMAY | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6C precursor (Semaphorin Y) (Sema Y). | |||||
|
SEM6A_MOUSE
|
||||||
| NC score | 0.943340 (rank : 29) | θ value | 3.44927e-84 (rank : 22) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O35464, Q6P5A8, Q6PCN9, Q9EQ71 | Gene names | Sema6a, Semaq | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6A precursor (Semaphorin VIA) (Sema VIA) (Semaphorin 6A-1) (SEMA6A-1) (Semaphorin Q) (Sema Q). | |||||
|
SEM6D_MOUSE
|
||||||
| NC score | 0.941613 (rank : 30) | θ value | 1.44917e-82 (rank : 24) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q76KF0, Q76KF1, Q76KF2, Q76KF3, Q76KF4, Q80TD0 | Gene names | Sema6d, Kiaa1479 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-6D precursor. | |||||
|
SEM6D_HUMAN
|
||||||
| NC score | 0.940768 (rank : 31) | θ value | 2.47191e-82 (rank : 25) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8NFY4, Q8NFY3, Q8NFY5, Q8NFY6, Q8NFY7, Q9P249 | Gene names | SEMA6D, KIAA1479 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Semaphorin-6D precursor. | |||||
|
SEM6B_HUMAN
|
||||||
| NC score | 0.937772 (rank : 32) | θ value | 4.83544e-70 (rank : 36) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9H3T3, Q9NRK9 | Gene names | SEMA6B, SEMAZ | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6B precursor (Semaphorin Z) (Sema Z). | |||||
|
SEM6B_MOUSE
|
||||||
| NC score | 0.936164 (rank : 33) | θ value | 1.96316e-71 (rank : 34) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O54951 | Gene names | Sema6b, Seman | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6B precursor (Semaphorin VIB) (Sema VIB) (Semaphorin N) (Sema N). | |||||
|
SEM6C_MOUSE
|
||||||
| NC score | 0.929542 (rank : 34) | θ value | 6.7473e-72 (rank : 33) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9WTM3 | Gene names | Sema6c, Semay | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6C precursor (Semaphorin Y) (Sema Y). | |||||
|
SEM5B_HUMAN
|
||||||
| NC score | 0.793102 (rank : 35) | θ value | 1.07473e-77 (rank : 27) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9P283, Q6UY12 | Gene names | SEMA5B, KIAA1445 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-5B precursor. | |||||
|
SEM5A_MOUSE
|
||||||
| NC score | 0.792643 (rank : 36) | θ value | 4.37348e-71 (rank : 35) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q62217 | Gene names | Sema5a, Semaf, SemF | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-5A precursor (Semaphorin F) (Sema F). | |||||
|
SEM5B_MOUSE
|
||||||
| NC score | 0.789951 (rank : 37) | θ value | 5.8972e-76 (rank : 28) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q60519 | Gene names | Sema5b, Semag, SemG | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-5B precursor (Semaphorin G) (Sema G). | |||||
|
SEM5A_HUMAN
|
||||||
| NC score | 0.788006 (rank : 38) | θ value | 5.1662e-72 (rank : 31) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q13591, O60408, Q1RLL9 | Gene names | SEMA5A, SEMAF | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-5A precursor (Semaphorin F) (Sema F). | |||||
|
PLXB2_HUMAN
|
||||||
| NC score | 0.339747 (rank : 39) | θ value | 3.19293e-05 (rank : 41) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O15031, Q7KZU3, Q9BSU7 | Gene names | PLXNB2, KIAA0315 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B2 precursor (MM1). | |||||
|
PLXC1_MOUSE
|
||||||
| NC score | 0.327541 (rank : 40) | θ value | 1.43324e-05 (rank : 40) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9QZC2, Q8CGW1 | Gene names | Plxnc1, Vespr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-C1 precursor (Virus-encoded semaphorin protein receptor) (CD232 antigen). | |||||
|
PLXC1_HUMAN
|
||||||
| NC score | 0.321588 (rank : 41) | θ value | 9.29e-05 (rank : 45) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O60486, Q59H25 | Gene names | PLXNC1, VESPR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-C1 precursor (Virus-encoded semaphorin protein receptor) (CD232 antigen). | |||||
|
PLXA4_MOUSE
|
||||||
| NC score | 0.315221 (rank : 42) | θ value | 7.1131e-05 (rank : 43) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q80UG2, Q5DTW8, Q8BKK9 | Gene names | Plxna4, Kiaa1550 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A4 precursor. | |||||
|
PLXA4_HUMAN
|
||||||
| NC score | 0.314566 (rank : 43) | θ value | 0.000461057 (rank : 46) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9HCM2, Q6UWC6, Q6ZW89, Q8N969, Q8ND00, Q8NEN3, Q9NTD4 | Gene names | PLXNA4, KIAA1550, PLXNA4A, PLXNA4B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A4 precursor. | |||||
|
PLXA2_MOUSE
|
||||||
| NC score | 0.307336 (rank : 44) | θ value | 2.21117e-06 (rank : 39) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P70207, Q6NVE6, Q6PHN4, Q80TZ7, Q8R1I4 | Gene names | Plxna2, Kiaa0463 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A2 precursor (Plexin-2) (Plex 2). | |||||
|
PLXA2_HUMAN
|
||||||
| NC score | 0.306365 (rank : 45) | θ value | 4.1701e-05 (rank : 42) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O75051, Q6UX61, Q96GN9, Q9BRL1, Q9UIW1 | Gene names | PLXNA2, KIAA0463, OCT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A2 precursor (Semaphorin receptor OCT). | |||||
|
PLXA1_MOUSE
|
||||||
| NC score | 0.293864 (rank : 46) | θ value | 0.00390308 (rank : 47) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P70206, Q5DTR0 | Gene names | Plxna1, KIAA4053 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A1 precursor (Plexin-1) (Plex 1). | |||||
|
PLXA3_HUMAN
|
||||||
| NC score | 0.287244 (rank : 47) | θ value | 9.29e-05 (rank : 44) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P51805 | Gene names | PLXNA3, PLXN4, SEX | |||
|
Domain Architecture |

|
|||||
| Description | Plexin-A3 precursor (Plexin-4) (Semaphorin receptor SEX). | |||||
|
PLXA1_HUMAN
|
||||||
| NC score | 0.284082 (rank : 48) | θ value | 0.0252991 (rank : 49) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9UIW2 | Gene names | PLXNA1, NOV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-A1 precursor (Semaphorin receptor NOV). | |||||
|
PLXB1_HUMAN
|
||||||
| NC score | 0.275660 (rank : 49) | θ value | 0.0252991 (rank : 50) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O43157, Q6NY20, Q9UIV7, Q9UJ92, Q9UJ93 | Gene names | PLXNB1, KIAA0407, SEP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B1 precursor (Semaphorin receptor SEP). | |||||
|
PLXB1_MOUSE
|
||||||
| NC score | 0.274791 (rank : 50) | θ value | 0.62314 (rank : 56) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8CJH3, Q6ZQC3, Q80ZZ1 | Gene names | Plxnb1, Kiaa0407 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B1 precursor. | |||||
|
PLXB3_MOUSE
|
||||||
| NC score | 0.250204 (rank : 51) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9QY40, Q80TH8 | Gene names | Plxnb3, Kiaa1206, Plxn6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B3 precursor (Plexin-6). | |||||
|
PLXB3_HUMAN
|
||||||
| NC score | 0.247148 (rank : 52) | θ value | 0.47712 (rank : 55) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9ULL4, Q9HDA4 | Gene names | PLXNB3, KIAA1206 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B3 precursor. | |||||
|
PLXD1_HUMAN
|
||||||
| NC score | 0.246477 (rank : 53) | θ value | 0.813845 (rank : 58) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9Y4D7, Q6PJS9, Q8IZJ2, Q9BTQ2 | Gene names | PLXND1, KIAA0620 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-D1 precursor. | |||||
|
PROP_HUMAN
|
||||||
| NC score | 0.089252 (rank : 54) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P27918, O15134, O15135, O15136, O75826 | Gene names | CFP, PFC | |||
|
Domain Architecture |

|
|||||
| Description | Properdin precursor (Factor P). | |||||
|
PROP_MOUSE
|
||||||
| NC score | 0.086750 (rank : 55) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P11680, Q3TB98, Q3U779 | Gene names | Cfp, Pfc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Properdin precursor (Factor P). | |||||
|
UNC5D_HUMAN
|
||||||
| NC score | 0.069698 (rank : 56) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6UXZ4, Q8WYP7 | Gene names | UNC5D, KIAA1777, UNC5H4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin receptor UNC5D precursor (Unc-5 homolog D) (Unc-5 homolog 4). | |||||
|
UNC5D_MOUSE
|
||||||
| NC score | 0.069580 (rank : 57) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8K1S2 | Gene names | Unc5d, Unc5h4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin receptor UNC5D precursor (Unc-5 homolog D) (Unc-5 homolog 4). | |||||
|
SPON1_MOUSE
|
||||||
| NC score | 0.066110 (rank : 58) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VCC9, Q8K2Q8 | Gene names | Spon1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spondin-1 precursor (F-spondin). | |||||
|
SPON1_HUMAN
|
||||||
| NC score | 0.064598 (rank : 59) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9HCB6, O94862, Q8NCD7, Q8WUR5 | Gene names | SPON1, KIAA0762, VSGP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spondin-1 precursor (F-spondin) (Vascular smooth muscle cell growth- promoting factor). | |||||
|
UNC5B_HUMAN
|
||||||
| NC score | 0.058916 (rank : 60) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8IZJ1, Q86SN3, Q8N1Y2, Q9H9F3 | Gene names | UNC5B, P53RDL1, UNC5H2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin receptor UNC5B precursor (Unc-5 homolog B) (Unc-5 homolog 2) (p53-regulated receptor for death and life protein 1). | |||||
|
UNC5B_MOUSE
|
||||||
| NC score | 0.058708 (rank : 61) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8K1S3, Q3U4F2, Q6PFH0, Q80Y85, Q9D398 | Gene names | Unc5b, Unc5h2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin receptor UNC5B precursor (Unc-5 homolog B) (Unc-5 homolog 2). | |||||
|
UNC5A_MOUSE
|
||||||
| NC score | 0.055072 (rank : 62) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8K1S4, Q6PEF7, Q80T71 | Gene names | Unc5a, Kiaa1976, Unc5h1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin receptor UNC5A precursor (Unc-5 homolog A) (Unc-5 homolog 1). | |||||
|
UNC5C_MOUSE
|
||||||
| NC score | 0.054735 (rank : 63) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O08747, Q8CD16 | Gene names | Unc5c, Rcm, Unc5h3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin receptor UNC5C precursor (Unc-5 homolog C) (Unc-5 homolog 3) (Rostral cerebellar malformation protein). | |||||
|
UNC5C_HUMAN
|
||||||
| NC score | 0.053708 (rank : 64) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O95185, Q8IUT0 | Gene names | UNC5C, UNC5H3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin receptor UNC5C precursor (Unc-5 homolog C) (Unc-5 homolog 3). | |||||
|
BAI1_HUMAN
|
||||||
| NC score | 0.050994 (rank : 65) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 375 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O14514 | Gene names | BAI1 | |||
|
Domain Architecture |

|
|||||
| Description | Brain-specific angiogenesis inhibitor 1 precursor. | |||||
|
BAI1_MOUSE
|
||||||
| NC score | 0.050651 (rank : 66) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 340 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3UHD1, Q3UH36, Q8CGM0 | Gene names | Bai1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain-specific angiogenesis inhibitor 1 precursor. | |||||
|
HMCN1_HUMAN
|
||||||
| NC score | 0.050439 (rank : 67) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 983 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q96RW7, Q5TYR7, Q96DN3, Q96DN8, Q96SC3 | Gene names | HMCN1, FIBL6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hemicentin-1 precursor (Fibulin-6) (FIBL-6). | |||||
|
CNTN4_HUMAN
|
||||||
| NC score | 0.032996 (rank : 68) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 449 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8IWV2, Q8IX14, Q8TC35 | Gene names | CNTN4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Contactin-4 precursor (Brain-derived immunoglobulin superfamily protein 2) (BIG-2). | |||||
|
CNTN4_MOUSE
|
||||||
| NC score | 0.030912 (rank : 69) | θ value | 1.38821 (rank : 61) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q69Z26, Q8BRT6, Q8CBD3 | Gene names | Cntn4, Kiaa3024 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Contactin-4 precursor (Brain-derived immunoglobulin superfamily protein 2) (BIG-2). | |||||
|
WDR35_HUMAN
|
||||||
| NC score | 0.028287 (rank : 70) | θ value | 0.125558 (rank : 51) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9P2L0, Q8NE11 | Gene names | WDR35, KIAA1336 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 35. | |||||
|
WDR35_MOUSE
|
||||||
| NC score | 0.028099 (rank : 71) | θ value | 0.365318 (rank : 52) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BND3, Q8CDL4 | Gene names | Wdr35 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 35. | |||||
|
DSCL1_HUMAN
|
||||||
| NC score | 0.027673 (rank : 72) | θ value | 0.47712 (rank : 54) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8TD84, Q76MU9, Q8IZY3, Q8IZY4, Q8WXU7, Q9ULT7 | Gene names | DSCAML1, DSCAM2, KIAA1132 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Down syndrome cell adhesion molecule-like protein 1 precursor (Down syndrome cell adhesion molecule 2). | |||||
|
OBSL1_HUMAN
|
||||||
| NC score | 0.027258 (rank : 73) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O75147, Q96IW3 | Gene names | OBSL1, KIAA0657 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin-like protein 1 precursor. | |||||
|
OBSCN_HUMAN
|
||||||
| NC score | 0.021245 (rank : 74) | θ value | 0.0252991 (rank : 48) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
JAML1_HUMAN
|
||||||
| NC score | 0.021234 (rank : 75) | θ value | 1.06291 (rank : 59) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q86YT9, Q5DTC6, Q7Z499, Q8N9I7, Q8NF70 | Gene names | AMICA1, JAML | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctional adhesion molecule-like precursor (Dendritic cell-specific protein CREA7-1) (Adhesion molecule interacting with CXADR antigen 1). | |||||
|
CNTN3_MOUSE
|
||||||
| NC score | 0.020384 (rank : 76) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q07409 | Gene names | Cntn3, Pang, Pcs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Contactin-3 precursor (Brain-derived immunoglobulin superfamily protein 1) (BIG-1) (Plasmacytoma-associated neuronal glycoprotein). | |||||
|
TREM2_HUMAN
|
||||||
| NC score | 0.019686 (rank : 77) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NZC2, Q8N5H8, Q8WYN6 | Gene names | TREM2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Triggering receptor expressed on myeloid cells 2 precursor (Triggering receptor expressed on monocytes 2) (TREM-2). | |||||
|
TREM2_MOUSE
|
||||||
| NC score | 0.019501 (rank : 78) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99NH8, Q8CGK4, Q8CIA6, Q99NH9, Q9JL34 | Gene names | Trem2, Trem2a, Trem2b, Trem2c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Triggering receptor expressed on myeloid cells 2 precursor (Triggering receptor expressed on monocytes 2) (TREM-2). | |||||
|
IRPL2_MOUSE
|
||||||
| NC score | 0.016534 (rank : 79) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9ERS6, Q9ER66 | Gene names | Il1rapl2 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked interleukin-1 receptor accessory protein-like 2 precursor (IL1RAPL-2-related protein) (TIGIRR-1). | |||||
|
BTNL1_MOUSE
|
||||||
| NC score | 0.015877 (rank : 80) | θ value | 0.813845 (rank : 57) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q7TST0, O70356 | Gene names | Btnl1, Btnl3, Gm316, Ng10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Butyrophilin-like protein 1 precursor. | |||||
|
IRPL2_HUMAN
|
||||||
| NC score | 0.015860 (rank : 81) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NP60, Q9NZN0 | Gene names | IL1RAPL2, IL1R9 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked interleukin-1 receptor accessory protein-like 2 precursor (IL1RAPL-2-related protein) (Interleukin-1 receptor 9) (IL-1R9) (IL-1 receptor accessory protein-like 2) (Three immunoglobulin domain- containing IL-1 receptor-related 1) (TIGIRR-1). | |||||
|
KIRR1_HUMAN
|
||||||
| NC score | 0.015332 (rank : 82) | θ value | 4.03905 (rank : 76) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q96J84, Q5XKC6, Q7Z696, Q7Z7N8, Q8TB15, Q9H9N1, Q9NVA5 | Gene names | KIRREL, NEPH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kin of IRRE-like protein 1 precursor (Kin of irregular chiasm-like protein 1) (Nephrin-like protein 1). | |||||
|
KIRR1_MOUSE
|
||||||
| NC score | 0.014796 (rank : 83) | θ value | 5.27518 (rank : 78) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q80W68, Q8CIJ4, Q8CJ59, Q923L4 | Gene names | Kirrel1, Kirrel, Neph1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kin of IRRE-like protein 1 precursor (Kin of irregular chiasm-like protein 1) (Nephrin-like protein 1). | |||||
|
AMGO1_HUMAN
|
||||||
| NC score | 0.013950 (rank : 84) | θ value | 0.47712 (rank : 53) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 353 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q86WK6, Q8IW71, Q9ULQ7 | Gene names | AMIGO1, ALI2, AMIGO, KIAA1163 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amphoterin-induced protein 1 precursor (AMIGO-1) (Alivin-2). | |||||
|
KIRR3_HUMAN
|
||||||
| NC score | 0.013138 (rank : 85) | θ value | 3.0926 (rank : 73) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8IZU9, Q6UWJ9, Q6UWL5, Q96JG0 | Gene names | KIRREL3, KIAA1867, NEPH2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kin of IRRE-like protein 3 precursor (Kin of irregular chiasm-like protein 3) (Nephrin-like 2). | |||||
|
IBP7_HUMAN
|
||||||
| NC score | 0.012702 (rank : 86) | θ value | 1.38821 (rank : 62) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q16270, Q07822 | Gene names | IGFBP7, MAC25, PSF | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 7 precursor (IGFBP-7) (IBP- 7) (IGF-binding protein 7) (MAC25 protein) (Prostacyclin-stimulating factor) (PGI2-stimulating factor) (IGFBP-rP1). | |||||
|
KIRR3_MOUSE
|
||||||
| NC score | 0.012293 (rank : 87) | θ value | 3.0926 (rank : 74) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BR86, Q6P555, Q810H3, Q8BGQ5, Q8BRS8 | Gene names | Kirrel3, Kiaa1867 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kin of IRRE-like protein 3 precursor (Kin of irregular chiasm-like protein 3). | |||||
|
IBP7_MOUSE
|
||||||
| NC score | 0.011770 (rank : 88) | θ value | 3.0926 (rank : 72) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61581, O88812, Q9EQW0 | Gene names | Igfbp7, Mac25 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 7 precursor (IGFBP-7) (IBP- 7) (IGF-binding protein 7) (MAC25 protein). | |||||
|
C8AP2_MOUSE
|
||||||
| NC score | 0.011691 (rank : 89) | θ value | 1.38821 (rank : 60) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9WUF3, Q9CSW2 | Gene names | Casp8ap2, Flash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
NCA12_MOUSE
|
||||||
| NC score | 0.011627 (rank : 90) | θ value | 1.81305 (rank : 67) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 399 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P13594, Q61950 | Gene names | Ncam1, Ncam | |||
|
Domain Architecture |

|
|||||
| Description | Neural cell adhesion molecule 1, 120 kDa isoform precursor (N-CAM 120) (NCAM-120). | |||||
|
ROBO2_MOUSE
|
||||||
| NC score | 0.011624 (rank : 91) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q7TPD3, Q8BJ59 | Gene names | Robo2, Kiaa1568 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 2 precursor. | |||||
|
NCA12_HUMAN
|
||||||
| NC score | 0.011424 (rank : 92) | θ value | 2.36792 (rank : 71) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 437 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P13592, P13593 | Gene names | NCAM1, NCAM | |||
|
Domain Architecture |

|
|||||
| Description | Neural cell adhesion molecule 1, 120 kDa isoform precursor (N-CAM 120) (NCAM-120) (CD56 antigen). | |||||
|
NCA11_HUMAN
|
||||||
| NC score | 0.011222 (rank : 93) | θ value | 2.36792 (rank : 70) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P13591, Q15829, Q16180 | Gene names | NCAM1, NCAM | |||
|
Domain Architecture |

|
|||||
| Description | Neural cell adhesion molecule 1, 140 kDa isoform precursor (N-CAM 140) (NCAM-140) (CD56 antigen). | |||||
|
NCA11_MOUSE
|
||||||
| NC score | 0.010711 (rank : 94) | θ value | 1.81305 (rank : 66) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 661 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P13595, Q61949 | Gene names | Ncam1, Ncam | |||
|
Domain Architecture |

|
|||||
| Description | Neural cell adhesion molecule 1, 180 kDa isoform precursor (N-CAM 180) (NCAM-180). | |||||
|
FGRL1_HUMAN
|
||||||
| NC score | 0.009033 (rank : 95) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8N441, Q6PJN1, Q9BXN7, Q9H4D7 | Gene names | FGFRL1, FGFR5, FHFR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibroblast growth factor receptor-like 1 precursor (FGF receptor-like protein 1) (Fibroblast growth factor receptor 5) (FGFR-like protein) (FGF homologous factor receptor). | |||||
|
CND2_HUMAN
|
||||||
| NC score | 0.004458 (rank : 96) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15003, Q8TB87 | Gene names | NCAPH, BRRN, BRRN1, CAPH, KIAA0074 | |||
|
Domain Architecture |

|
|||||
| Description | Condensin complex subunit 2 (Non-SMC condensin I complex subunit H) (Barren homolog protein 1) (Chromosome-associated protein H) (hCAP-H) (XCAP-H homolog). | |||||
|
BCLF1_HUMAN
|
||||||
| NC score | 0.003836 (rank : 97) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NYF8, Q14673, Q86WU6, Q86WY0 | Gene names | BCLAF1, BTF, KIAA0164 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bcl-2-associated transcription factor 1 (Btf). | |||||
|
BCLF1_MOUSE
|
||||||
| NC score | 0.003676 (rank : 98) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K019, Q8BNZ0, Q8C2E9, Q9CSW5 | Gene names | Bclaf1, Btf, Kiaa0164 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bcl-2-associated transcription factor 1 (Btf). | |||||
|
DUX5_HUMAN
|
||||||
| NC score | 0.003469 (rank : 99) | θ value | 2.36792 (rank : 69) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96PT3 | Gene names | DUX5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double homeobox protein 5. | |||||
|
CC47_MOUSE
|
||||||
| NC score | 0.003456 (rank : 100) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D024, Q3TSB5, Q3V1S7, Q8C5D9, Q920S6 | Gene names | Ccdc47, Asp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 47 precursor (Adipocyte-specific protein 4). | |||||
|
DUX1_HUMAN
|
||||||
| NC score | 0.003337 (rank : 101) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43812 | Gene names | DUX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double homeobox protein 1. | |||||
|
DUX3_HUMAN
|
||||||
| NC score | 0.002968 (rank : 102) | θ value | 5.27518 (rank : 77) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96PT4, Q9UND2 | Gene names | DUX3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative double homeobox protein 3. | |||||
|
KIT_HUMAN
|
||||||
| NC score | -0.001418 (rank : 103) | θ value | 1.38821 (rank : 63) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 953 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P10721, Q9UM99 | Gene names | KIT | |||
|
Domain Architecture |

|
|||||
| Description | Mast/stem cell growth factor receptor precursor (EC 2.7.10.1) (SCFR) (Proto-oncogene tyrosine-protein kinase Kit) (c-kit) (CD117 antigen). | |||||
|
FGFR2_HUMAN
|
||||||
| NC score | -0.002106 (rank : 104) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1062 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P21802, P18443, Q01742, Q12922, Q14300, Q14301, Q14302, Q14303, Q14304, Q14305, Q14672, Q14718, Q14719, Q86YI4, Q96KL9, Q96KM0, Q96KM1, Q96KM2, Q9NZU2, Q9NZU3, Q9UD01, Q9UD02, Q9UIH3, Q9UIH4, Q9UIH5, Q9UIH6, Q9UIH7, Q9UIH8, Q9UM87, Q9UMC6, Q9UNS7, Q9UQH7, Q9UQH8, Q9UQH9, Q9UQI0 | Gene names | FGFR2, BEK, KSAM | |||
|
Domain Architecture |

|
|||||
| Description | Fibroblast growth factor receptor 2 precursor (EC 2.7.10.1) (FGFR-2) (Keratinocyte growth factor receptor 2) (CD332 antigen). | |||||
|
MINK1_MOUSE
|
||||||
| NC score | -0.002321 (rank : 105) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1451 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9JM52, Q921M6, Q9JM92 | Gene names | Mink1, Map4k6, Mink | |||
|
Domain Architecture |

|
|||||
| Description | Misshapen-like kinase 1 (EC 2.7.11.1) (Mitogen-activated protein kinase kinase kinase kinase 6) (MAPK/ERK kinase kinase kinase 6) (MEK kinase kinase 6) (MEKKK 6) (Misshapen/NIK-related kinase) (GCK family kinase MiNK). | |||||
|
MINK1_HUMAN
|
||||||
| NC score | -0.003120 (rank : 106) | θ value | 1.81305 (rank : 64) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N4C8, Q9P1X1, Q9P2R8 | Gene names | MINK1, MAP4K6, MINK | |||
|
Domain Architecture |

|
|||||
| Description | Misshapen-like kinase 1 (EC 2.7.11.1) (Mitogen-activated protein kinase kinase kinase kinase 6) (MAPK/ERK kinase kinase kinase 6) (MEK kinase kinase 6) (MEKKK 6) (Misshapen/NIK-related kinase) (GCK family kinase MiNK). | |||||
|
ZN596_HUMAN
|
||||||
| NC score | -0.005593 (rank : 107) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 716 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TC21, O95015, Q8N9X0 | Gene names | ZNF596 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 596. | |||||