Please be patient as the page loads
|
ERB1_HUMAN
|
||||||
| SwissProt Accessions | P60509 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-R(b)_3p24.3 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (HERV-R(b) Env protein) [Includes: Surface protein domain (SU); Transmembrane protein domain (TM)]. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ERB1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P60509 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-R(b)_3p24.3 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (HERV-R(b) Env protein) [Includes: Surface protein domain (SU); Transmembrane protein domain (TM)]. | |||||
|
ENH1_HUMAN
|
||||||
| θ value | 7.09661e-13 (rank : 2) | NC score | 0.479353 (rank : 6) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9N2K0, O00354, Q96L63, Q9UNM3 | Gene names | ||||
|
Domain Architecture |
|
|||||
| Description | HERV-H_2q24.3 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (Env protein HERV-H/p62) (Env protein HERV-H19) (HERV- H/env62) (Env protein HERV-Hcl.3) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
EFR1_HUMAN
|
||||||
| θ value | 7.34386e-10 (rank : 3) | NC score | 0.543917 (rank : 2) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P60508 | Gene names | ERVFRDE1 | |||
|
Domain Architecture |

|
|||||
| Description | HERV-FRD_6p24.1 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (HERV-FRD) (Syncytin-2) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
ENR1_HUMAN
|
||||||
| θ value | 4.76016e-09 (rank : 4) | NC score | 0.474343 (rank : 8) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14264 | Gene names | ERV3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-R_7q21.2 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (ERV3 envelope protein) (ERV-3 envelope protein) (HERV-R envelope protein) (ERV-R envelope protein) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
ENH2_HUMAN
|
||||||
| θ value | 6.87365e-08 (rank : 5) | NC score | 0.443074 (rank : 10) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9N2J9 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | HERV-H_3q26 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (Env protein HERV-H/p60) (HERV-H/env60) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
ENV2_MOUSE
|
||||||
| θ value | 1.17247e-07 (rank : 6) | NC score | 0.514038 (rank : 4) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P11370, Q61529 | Gene names | Fv4, Env | |||
|
Domain Architecture |

|
|||||
| Description | Retrovirus-related Env polyprotein from Fv-4 locus. | |||||
|
EFC1_HUMAN
|
||||||
| θ value | 2.61198e-07 (rank : 7) | NC score | 0.506779 (rank : 5) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P60507 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-F(c)1_Xq21.33 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (Fc1env) (HERV-Fc1env) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
ENH3_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 8) | NC score | 0.425693 (rank : 11) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9N2J8 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | HERV-H_2q24.1 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (Env protein HERV-H/p59) (HERV-H/env59) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
EFC2_HUMAN
|
||||||
| θ value | 3.19293e-05 (rank : 9) | NC score | 0.477140 (rank : 7) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P60608 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-F(c)2_7q36.2 provirus ancestral Env polyprotein (Envelope polyprotein) (Fc2deltaenv) [Includes: Surface protein (SU); Truncated transmembrane protein (TM)]. | |||||
|
ENW1_HUMAN
|
||||||
| θ value | 3.19293e-05 (rank : 10) | NC score | 0.515543 (rank : 3) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UQF0, O95244, O95245, Q8NHY7, Q9NRZ2, Q9NZG3 | Gene names | ERVWE1 | |||
|
Domain Architecture |
|
|||||
| Description | HERV-W_7q21.2 provirus ancestral Env polyprotein precursor (Envelope polyprotein gPr73) (HERV-7q Envelope protein) (HERV-W envelope protein) (Syncytin) (Syncytin-1) (Enverin) (Env-W) [Contains: Surface protein (SU) (gp50); Transmembrane protein (TM) (gp24)]. | |||||
|
ENT1_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 11) | NC score | 0.469840 (rank : 9) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P61550 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-T_19q13.11 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (HERV-T Env protein) [Includes: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
NMDE1_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 12) | NC score | 0.139891 (rank : 15) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q12879, O00669 | Gene names | GRIN2A | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 1 precursor (N-methyl D- aspartate receptor subtype 2A) (NR2A) (NMDAR2A) (hNR2A). | |||||
|
NMDE1_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 13) | NC score | 0.139982 (rank : 14) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P35436 | Gene names | Grin2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 1 precursor (N-methyl D- aspartate receptor subtype 2A) (NR2A) (NMDAR2A). | |||||
|
NMDE2_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 14) | NC score | 0.139651 (rank : 16) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q13224, Q12919, Q13220, Q13225, Q9UM56 | Gene names | GRIN2B | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 2 precursor (N-methyl D- aspartate receptor subtype 2B) (NR2B) (NMDAR2B) (N-methyl-D-aspartate receptor subunit 3) (NR3) (hNR3). | |||||
|
NMDE2_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 15) | NC score | 0.139606 (rank : 17) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q01097, Q9DCB2 | Gene names | Grin2b | |||
|
Domain Architecture |
|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 2 precursor (N-methyl D- aspartate receptor subtype 2B) (NR2B) (NMDAR2B). | |||||
|
NMDE3_MOUSE
|
||||||
| θ value | 0.163984 (rank : 16) | NC score | 0.136239 (rank : 18) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q01098 | Gene names | Grin2c | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 3 precursor (N-methyl D- aspartate receptor subtype 2C) (NR2C) (NMDAR2C). | |||||
|
NMDE3_HUMAN
|
||||||
| θ value | 0.21417 (rank : 17) | NC score | 0.134967 (rank : 19) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14957 | Gene names | GRIN2C | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 3 precursor (N-methyl D- aspartate receptor subtype 2C) (NR2C) (NMDAR2C). | |||||
|
NMDE4_HUMAN
|
||||||
| θ value | 0.21417 (rank : 18) | NC score | 0.133162 (rank : 20) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O15399 | Gene names | GRIN2D | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 4 precursor (N-methyl D- aspartate receptor subtype 2D) (NR2D) (NMDAR2D) (EB11). | |||||
|
NMDE4_MOUSE
|
||||||
| θ value | 0.21417 (rank : 19) | NC score | 0.132871 (rank : 21) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q03391 | Gene names | Grin2d | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 4 precursor (N-methyl D- aspartate receptor subtype 2D) (NR2D) (NMDAR2D). | |||||
|
COT1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 20) | NC score | 0.008215 (rank : 50) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P10589 | Gene names | NR2F1, EAR3, ERBAL3, TFCOUP1 | |||
|
Domain Architecture |
|
|||||
| Description | COUP transcription factor 1 (COUP-TF1) (COUP-TF I) (V-ERBA-related protein EAR-3). | |||||
|
COT1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 21) | NC score | 0.008220 (rank : 49) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60632, Q61438 | Gene names | Nr2f1, Erbal3, Tfcoup1 | |||
|
Domain Architecture |
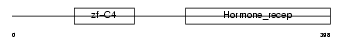
|
|||||
| Description | COUP transcription factor 1 (COUP-TF1) (COUP-TF I). | |||||
|
GNPTA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 22) | NC score | 0.030413 (rank : 47) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q3T906, Q3ZQK2, Q6IPW5, Q86TQ2, Q96N13, Q9ULL2 | Gene names | GNPTAB, GNPTA, KIAA1208 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylglucosamine-1-phosphotransferase subunits alpha/beta precursor (EC 2.7.8.17) (GlcNAc-1-phosphotransferase alpha/beta subunits) (UDP- N-acetylglucosamine-1-phosphotransferase alpha/beta subunits) (Stealth protein GNPTAB) [Contains: N-acetylglucosamine-1-phosphotransferase subunit alpha; N-acetylglucosamine-1-phosphotransferase subunit beta]. | |||||
|
GNPTA_MOUSE
|
||||||
| θ value | 6.88961 (rank : 23) | NC score | 0.030210 (rank : 48) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q69ZN6, Q3U3K6, Q3US34 | Gene names | Gnptab, Gnpta, Kiaa1208 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylglucosamine-1-phosphotransferase subunits alpha/beta precursor (EC 2.7.8.17) (GlcNAc-1-phosphotransferase alpha/beta subunits) (UDP- N-acetylglucosamine-1-phosphotransferase alpha/beta subunits) (Stealth protein GNPTAB) [Contains: N-acetylglucosamine-1-phosphotransferase subunit alpha; N-acetylglucosamine-1-phosphotransferase subunit beta]. | |||||
|
ENH4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 24) | NC score | 0.278431 (rank : 12) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96L62 | Gene names | ||||
|
Domain Architecture |
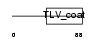
|
|||||
| Description | HERV-H_1q41 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (Env protein HERV-Hcl.7) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
ENH5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 25) | NC score | 0.276355 (rank : 13) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P61549 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-H_19p13.11 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (Env protein RGH1) (Env protein RTLV-H) [Includes: Surface protein (SU)]. | |||||
|
GRIA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 26) | NC score | 0.053248 (rank : 40) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P42261 | Gene names | GRIA1, GLUH1, GLUR1 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor 1 precursor (GluR-1) (GluR-A) (GluR-K1) (Glutamate receptor ionotropic, AMPA 1) (AMPA-selective glutamate receptor 1). | |||||
|
GRIA1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 27) | NC score | 0.053644 (rank : 38) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P23818 | Gene names | Gria1, Glur1 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor 1 precursor (GluR-1) (GluR-A) (GluR-K1) (Glutamate receptor ionotropic, AMPA 1) (AMPA-selective glutamate receptor 1). | |||||
|
GRIA2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 28) | NC score | 0.052377 (rank : 43) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P42262, Q96FP6 | Gene names | GRIA2, GLUR2 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor 2 precursor (GluR-2) (GluR-B) (GluR-K2) (Glutamate receptor ionotropic, AMPA 2) (AMPA-selective glutamate receptor 2). | |||||
|
GRIA2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 29) | NC score | 0.052220 (rank : 45) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P23819, Q61604, Q61605, Q8BG69, Q8BXU3, Q9D6D3 | Gene names | Gria2, Glur2 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor 2 precursor (GluR-2) (GluR-B) (GluR-K2) (Glutamate receptor ionotropic, AMPA 2) (AMPA-selective glutamate receptor 2). | |||||
|
GRIA3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 30) | NC score | 0.052396 (rank : 42) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P42263, Q4VXD5, Q4VXD6, Q9HDA0, Q9HDA1, Q9HDA2 | Gene names | GRIA3, GLUR3, GLURC | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor 3 precursor (GluR-3) (GluR-C) (GluR-K3) (Glutamate receptor ionotropic, AMPA 3) (AMPA-selective glutamate receptor 3). | |||||
|
GRIA3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 31) | NC score | 0.052285 (rank : 44) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z2W9, Q5DTJ0 | Gene names | Gria3, Glur3, Kiaa4184 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate receptor 3 precursor (GluR-3) (GluR-C) (GluR-K3) (Glutamate receptor ionotropic, AMPA 3) (AMPA-selective glutamate receptor 3). | |||||
|
GRIA4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 32) | NC score | 0.053477 (rank : 39) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P48058 | Gene names | GRIA4, GLUR4 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor 4 precursor (GluR-4) (GluR4) (GluR-D) (Glutamate receptor ionotropic, AMPA 4) (AMPA-selective glutamate receptor 4). | |||||
|
GRIA4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 33) | NC score | 0.053046 (rank : 41) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z2W8 | Gene names | Gria4, Glur4 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor 4 precursor (GluR-4) (GluR4) (GluR-D) (Glutamate receptor ionotropic, AMPA 4) (AMPA-selective glutamate receptor 4). | |||||
|
GRID1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 34) | NC score | 0.056795 (rank : 30) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9ULK0, Q8IXT3 | Gene names | GRID1, KIAA1220 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor delta-1 subunit precursor (GluR delta-1). | |||||
|
GRID1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 35) | NC score | 0.057230 (rank : 29) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61627 | Gene names | Grid1 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor delta-1 subunit precursor (GluR delta-1). | |||||
|
GRID2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 36) | NC score | 0.057845 (rank : 28) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O43424 | Gene names | GRID2, GLURD2 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor delta-2 subunit precursor (GluR delta-2). | |||||
|
GRID2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 37) | NC score | 0.058304 (rank : 27) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61625 | Gene names | Grid2 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor delta-2 subunit precursor (GluR delta-2). | |||||
|
GRIK1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 38) | NC score | 0.054717 (rank : 34) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P39086, Q13001, Q86SU9 | Gene names | GRIK1, GLUR5 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor, ionotropic kainate 1 precursor (Glutamate receptor 5) (GluR-5) (GluR5) (Excitatory amino acid receptor 3) (EAA3). | |||||
|
GRIK1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 39) | NC score | 0.051029 (rank : 46) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q60934 | Gene names | Grik1, Glur5 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor, ionotropic kainate 1 precursor (Glutamate receptor 5) (GluR-5) (GluR5). | |||||
|
GRIK2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 40) | NC score | 0.054300 (rank : 37) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q13002 | Gene names | GRIK2, GLUR6 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor, ionotropic kainate 2 precursor (Glutamate receptor 6) (GluR-6) (GluR6) (Excitatory amino acid receptor 4) (EAA4). | |||||
|
GRIK3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 41) | NC score | 0.054867 (rank : 32) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q13003, Q13004, Q16136 | Gene names | GRIK3, GLUR7 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor, ionotropic kainate 3 precursor (Glutamate receptor 7) (GluR-7) (GluR7) (Excitatory amino acid receptor 5) (EAA5). | |||||
|
GRIK4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 42) | NC score | 0.054875 (rank : 31) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q16099 | Gene names | GRIK4, GRIK | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor, ionotropic kainate 4 precursor (Glutamate receptor KA-1) (KA1) (Excitatory amino acid receptor 1) (EAA1). | |||||
|
GRIK4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 43) | NC score | 0.054821 (rank : 33) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BMF5 | Gene names | Grik4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate receptor, ionotropic kainate 4 precursor. | |||||
|
GRIK5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 44) | NC score | 0.054422 (rank : 35) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q16478 | Gene names | GRIK5, GRIK2 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor, ionotropic kainate 5 precursor (Glutamate receptor KA-2) (KA2) (Excitatory amino acid receptor 2) (EAA2). | |||||
|
GRIK5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 45) | NC score | 0.054367 (rank : 36) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61626 | Gene names | Grik5 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor, ionotropic kainate 5 precursor (Glutamate receptor KA-2) (KA2) (Glutamate receptor gamma-2) (GluR gamma-2). | |||||
|
NMD3A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 46) | NC score | 0.087530 (rank : 22) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8TCU5, Q8TF29, Q8WXI6 | Gene names | GRIN3A, KIAA1973 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate [NMDA] receptor subunit 3A precursor (N-methyl-D-aspartate receptor subtype NR3A) (NMDAR-L). | |||||
|
NMD3B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 47) | NC score | 0.084508 (rank : 24) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O60391, Q5EAK7, Q7RTW9 | Gene names | GRIN3B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate [NMDA] receptor subunit 3B precursor (N-methyl-D-aspartate receptor subtype NR3B) (NR3B) (NMDAR3B). | |||||
|
NMD3B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 48) | NC score | 0.085221 (rank : 23) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q91ZU9, Q8VHV0 | Gene names | Grin3b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate [NMDA] receptor subunit 3B precursor (N-methyl-D-aspartate receptor subunit NR3B) (NR3B) (NMDAR3B) (NMDA receptor NR4) (Nr4). | |||||
|
NMDZ1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 49) | NC score | 0.078250 (rank : 25) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q05586, P35437, Q12867, Q12868 | Gene names | GRIN1, NMDAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit zeta 1 precursor (N-methyl-D- aspartate receptor subunit NR1). | |||||
|
NMDZ1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 50) | NC score | 0.078166 (rank : 26) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P35438, Q8CFS4 | Gene names | Grin1, Glurz1 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit zeta 1 precursor (N-methyl-D- aspartate receptor subunit NR1). | |||||
|
ERB1_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P60509 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-R(b)_3p24.3 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (HERV-R(b) Env protein) [Includes: Surface protein domain (SU); Transmembrane protein domain (TM)]. | |||||
|
EFR1_HUMAN
|
||||||
| NC score | 0.543917 (rank : 2) | θ value | 7.34386e-10 (rank : 3) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P60508 | Gene names | ERVFRDE1 | |||
|
Domain Architecture |

|
|||||
| Description | HERV-FRD_6p24.1 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (HERV-FRD) (Syncytin-2) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
ENW1_HUMAN
|
||||||
| NC score | 0.515543 (rank : 3) | θ value | 3.19293e-05 (rank : 10) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UQF0, O95244, O95245, Q8NHY7, Q9NRZ2, Q9NZG3 | Gene names | ERVWE1 | |||
|
Domain Architecture |

|
|||||
| Description | HERV-W_7q21.2 provirus ancestral Env polyprotein precursor (Envelope polyprotein gPr73) (HERV-7q Envelope protein) (HERV-W envelope protein) (Syncytin) (Syncytin-1) (Enverin) (Env-W) [Contains: Surface protein (SU) (gp50); Transmembrane protein (TM) (gp24)]. | |||||
|
ENV2_MOUSE
|
||||||
| NC score | 0.514038 (rank : 4) | θ value | 1.17247e-07 (rank : 6) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P11370, Q61529 | Gene names | Fv4, Env | |||
|
Domain Architecture |

|
|||||
| Description | Retrovirus-related Env polyprotein from Fv-4 locus. | |||||
|
EFC1_HUMAN
|
||||||
| NC score | 0.506779 (rank : 5) | θ value | 2.61198e-07 (rank : 7) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P60507 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-F(c)1_Xq21.33 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (Fc1env) (HERV-Fc1env) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
ENH1_HUMAN
|
||||||
| NC score | 0.479353 (rank : 6) | θ value | 7.09661e-13 (rank : 2) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9N2K0, O00354, Q96L63, Q9UNM3 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | HERV-H_2q24.3 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (Env protein HERV-H/p62) (Env protein HERV-H19) (HERV- H/env62) (Env protein HERV-Hcl.3) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
EFC2_HUMAN
|
||||||
| NC score | 0.477140 (rank : 7) | θ value | 3.19293e-05 (rank : 9) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P60608 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-F(c)2_7q36.2 provirus ancestral Env polyprotein (Envelope polyprotein) (Fc2deltaenv) [Includes: Surface protein (SU); Truncated transmembrane protein (TM)]. | |||||
|
ENR1_HUMAN
|
||||||
| NC score | 0.474343 (rank : 8) | θ value | 4.76016e-09 (rank : 4) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14264 | Gene names | ERV3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-R_7q21.2 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (ERV3 envelope protein) (ERV-3 envelope protein) (HERV-R envelope protein) (ERV-R envelope protein) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
ENT1_HUMAN
|
||||||
| NC score | 0.469840 (rank : 9) | θ value | 0.000158464 (rank : 11) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P61550 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-T_19q13.11 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (HERV-T Env protein) [Includes: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
ENH2_HUMAN
|
||||||
| NC score | 0.443074 (rank : 10) | θ value | 6.87365e-08 (rank : 5) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9N2J9 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | HERV-H_3q26 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (Env protein HERV-H/p60) (HERV-H/env60) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
ENH3_HUMAN
|
||||||
| NC score | 0.425693 (rank : 11) | θ value | 6.43352e-06 (rank : 8) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9N2J8 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | HERV-H_2q24.1 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (Env protein HERV-H/p59) (HERV-H/env59) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
ENH4_HUMAN
|
||||||
| NC score | 0.278431 (rank : 12) | θ value | θ > 10 (rank : 24) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96L62 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | HERV-H_1q41 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (Env protein HERV-Hcl.7) [Contains: Surface protein (SU); Transmembrane protein (TM)]. | |||||
|
ENH5_HUMAN
|
||||||
| NC score | 0.276355 (rank : 13) | θ value | θ > 10 (rank : 25) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P61549 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-H_19p13.11 provirus ancestral Env polyprotein precursor (Envelope polyprotein) (Env protein RGH1) (Env protein RTLV-H) [Includes: Surface protein (SU)]. | |||||
|
NMDE1_MOUSE
|
||||||
| NC score | 0.139982 (rank : 14) | θ value | 0.0563607 (rank : 13) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P35436 | Gene names | Grin2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 1 precursor (N-methyl D- aspartate receptor subtype 2A) (NR2A) (NMDAR2A). | |||||
|
NMDE1_HUMAN
|
||||||
| NC score | 0.139891 (rank : 15) | θ value | 0.0563607 (rank : 12) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q12879, O00669 | Gene names | GRIN2A | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 1 precursor (N-methyl D- aspartate receptor subtype 2A) (NR2A) (NMDAR2A) (hNR2A). | |||||
|
NMDE2_HUMAN
|
||||||
| NC score | 0.139651 (rank : 16) | θ value | 0.0563607 (rank : 14) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q13224, Q12919, Q13220, Q13225, Q9UM56 | Gene names | GRIN2B | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 2 precursor (N-methyl D- aspartate receptor subtype 2B) (NR2B) (NMDAR2B) (N-methyl-D-aspartate receptor subunit 3) (NR3) (hNR3). | |||||
|
NMDE2_MOUSE
|
||||||
| NC score | 0.139606 (rank : 17) | θ value | 0.0563607 (rank : 15) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q01097, Q9DCB2 | Gene names | Grin2b | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 2 precursor (N-methyl D- aspartate receptor subtype 2B) (NR2B) (NMDAR2B). | |||||
|
NMDE3_MOUSE
|
||||||
| NC score | 0.136239 (rank : 18) | θ value | 0.163984 (rank : 16) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q01098 | Gene names | Grin2c | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 3 precursor (N-methyl D- aspartate receptor subtype 2C) (NR2C) (NMDAR2C). | |||||
|
NMDE3_HUMAN
|
||||||
| NC score | 0.134967 (rank : 19) | θ value | 0.21417 (rank : 17) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14957 | Gene names | GRIN2C | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 3 precursor (N-methyl D- aspartate receptor subtype 2C) (NR2C) (NMDAR2C). | |||||
|
NMDE4_HUMAN
|
||||||
| NC score | 0.133162 (rank : 20) | θ value | 0.21417 (rank : 18) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O15399 | Gene names | GRIN2D | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 4 precursor (N-methyl D- aspartate receptor subtype 2D) (NR2D) (NMDAR2D) (EB11). | |||||
|
NMDE4_MOUSE
|
||||||
| NC score | 0.132871 (rank : 21) | θ value | 0.21417 (rank : 19) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q03391 | Gene names | Grin2d | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 4 precursor (N-methyl D- aspartate receptor subtype 2D) (NR2D) (NMDAR2D). | |||||
|
NMD3A_HUMAN
|
||||||
| NC score | 0.087530 (rank : 22) | θ value | θ > 10 (rank : 46) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8TCU5, Q8TF29, Q8WXI6 | Gene names | GRIN3A, KIAA1973 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate [NMDA] receptor subunit 3A precursor (N-methyl-D-aspartate receptor subtype NR3A) (NMDAR-L). | |||||
|
NMD3B_MOUSE
|
||||||
| NC score | 0.085221 (rank : 23) | θ value | θ > 10 (rank : 48) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q91ZU9, Q8VHV0 | Gene names | Grin3b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate [NMDA] receptor subunit 3B precursor (N-methyl-D-aspartate receptor subunit NR3B) (NR3B) (NMDAR3B) (NMDA receptor NR4) (Nr4). | |||||
|
NMD3B_HUMAN
|
||||||
| NC score | 0.084508 (rank : 24) | θ value | θ > 10 (rank : 47) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O60391, Q5EAK7, Q7RTW9 | Gene names | GRIN3B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate [NMDA] receptor subunit 3B precursor (N-methyl-D-aspartate receptor subtype NR3B) (NR3B) (NMDAR3B). | |||||
|
NMDZ1_HUMAN
|
||||||
| NC score | 0.078250 (rank : 25) | θ value | θ > 10 (rank : 49) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q05586, P35437, Q12867, Q12868 | Gene names | GRIN1, NMDAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit zeta 1 precursor (N-methyl-D- aspartate receptor subunit NR1). | |||||
|
NMDZ1_MOUSE
|
||||||
| NC score | 0.078166 (rank : 26) | θ value | θ > 10 (rank : 50) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P35438, Q8CFS4 | Gene names | Grin1, Glurz1 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit zeta 1 precursor (N-methyl-D- aspartate receptor subunit NR1). | |||||
|
GRID2_MOUSE
|
||||||
| NC score | 0.058304 (rank : 27) | θ value | θ > 10 (rank : 37) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61625 | Gene names | Grid2 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor delta-2 subunit precursor (GluR delta-2). | |||||
|
GRID2_HUMAN
|
||||||
| NC score | 0.057845 (rank : 28) | θ value | θ > 10 (rank : 36) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O43424 | Gene names | GRID2, GLURD2 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor delta-2 subunit precursor (GluR delta-2). | |||||
|
GRID1_MOUSE
|
||||||
| NC score | 0.057230 (rank : 29) | θ value | θ > 10 (rank : 35) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61627 | Gene names | Grid1 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor delta-1 subunit precursor (GluR delta-1). | |||||
|
GRID1_HUMAN
|
||||||
| NC score | 0.056795 (rank : 30) | θ value | θ > 10 (rank : 34) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9ULK0, Q8IXT3 | Gene names | GRID1, KIAA1220 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor delta-1 subunit precursor (GluR delta-1). | |||||
|
GRIK4_HUMAN
|
||||||
| NC score | 0.054875 (rank : 31) | θ value | θ > 10 (rank : 42) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q16099 | Gene names | GRIK4, GRIK | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor, ionotropic kainate 4 precursor (Glutamate receptor KA-1) (KA1) (Excitatory amino acid receptor 1) (EAA1). | |||||
|
GRIK3_HUMAN
|
||||||
| NC score | 0.054867 (rank : 32) | θ value | θ > 10 (rank : 41) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q13003, Q13004, Q16136 | Gene names | GRIK3, GLUR7 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor, ionotropic kainate 3 precursor (Glutamate receptor 7) (GluR-7) (GluR7) (Excitatory amino acid receptor 5) (EAA5). | |||||
|
GRIK4_MOUSE
|
||||||
| NC score | 0.054821 (rank : 33) | θ value | θ > 10 (rank : 43) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BMF5 | Gene names | Grik4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate receptor, ionotropic kainate 4 precursor. | |||||
|
GRIK1_HUMAN
|
||||||
| NC score | 0.054717 (rank : 34) | θ value | θ > 10 (rank : 38) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P39086, Q13001, Q86SU9 | Gene names | GRIK1, GLUR5 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor, ionotropic kainate 1 precursor (Glutamate receptor 5) (GluR-5) (GluR5) (Excitatory amino acid receptor 3) (EAA3). | |||||
|
GRIK5_HUMAN
|
||||||
| NC score | 0.054422 (rank : 35) | θ value | θ > 10 (rank : 44) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q16478 | Gene names | GRIK5, GRIK2 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor, ionotropic kainate 5 precursor (Glutamate receptor KA-2) (KA2) (Excitatory amino acid receptor 2) (EAA2). | |||||
|
GRIK5_MOUSE
|
||||||
| NC score | 0.054367 (rank : 36) | θ value | θ > 10 (rank : 45) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61626 | Gene names | Grik5 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor, ionotropic kainate 5 precursor (Glutamate receptor KA-2) (KA2) (Glutamate receptor gamma-2) (GluR gamma-2). | |||||
|
GRIK2_HUMAN
|
||||||
| NC score | 0.054300 (rank : 37) | θ value | θ > 10 (rank : 40) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q13002 | Gene names | GRIK2, GLUR6 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor, ionotropic kainate 2 precursor (Glutamate receptor 6) (GluR-6) (GluR6) (Excitatory amino acid receptor 4) (EAA4). | |||||
|
GRIA1_MOUSE
|
||||||
| NC score | 0.053644 (rank : 38) | θ value | θ > 10 (rank : 27) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P23818 | Gene names | Gria1, Glur1 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor 1 precursor (GluR-1) (GluR-A) (GluR-K1) (Glutamate receptor ionotropic, AMPA 1) (AMPA-selective glutamate receptor 1). | |||||
|
GRIA4_HUMAN
|
||||||
| NC score | 0.053477 (rank : 39) | θ value | θ > 10 (rank : 32) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P48058 | Gene names | GRIA4, GLUR4 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor 4 precursor (GluR-4) (GluR4) (GluR-D) (Glutamate receptor ionotropic, AMPA 4) (AMPA-selective glutamate receptor 4). | |||||
|
GRIA1_HUMAN
|
||||||
| NC score | 0.053248 (rank : 40) | θ value | θ > 10 (rank : 26) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P42261 | Gene names | GRIA1, GLUH1, GLUR1 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor 1 precursor (GluR-1) (GluR-A) (GluR-K1) (Glutamate receptor ionotropic, AMPA 1) (AMPA-selective glutamate receptor 1). | |||||
|
GRIA4_MOUSE
|
||||||
| NC score | 0.053046 (rank : 41) | θ value | θ > 10 (rank : 33) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z2W8 | Gene names | Gria4, Glur4 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor 4 precursor (GluR-4) (GluR4) (GluR-D) (Glutamate receptor ionotropic, AMPA 4) (AMPA-selective glutamate receptor 4). | |||||
|
GRIA3_HUMAN
|
||||||
| NC score | 0.052396 (rank : 42) | θ value | θ > 10 (rank : 30) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P42263, Q4VXD5, Q4VXD6, Q9HDA0, Q9HDA1, Q9HDA2 | Gene names | GRIA3, GLUR3, GLURC | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor 3 precursor (GluR-3) (GluR-C) (GluR-K3) (Glutamate receptor ionotropic, AMPA 3) (AMPA-selective glutamate receptor 3). | |||||
|
GRIA2_HUMAN
|
||||||
| NC score | 0.052377 (rank : 43) | θ value | θ > 10 (rank : 28) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P42262, Q96FP6 | Gene names | GRIA2, GLUR2 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor 2 precursor (GluR-2) (GluR-B) (GluR-K2) (Glutamate receptor ionotropic, AMPA 2) (AMPA-selective glutamate receptor 2). | |||||
|
GRIA3_MOUSE
|
||||||
| NC score | 0.052285 (rank : 44) | θ value | θ > 10 (rank : 31) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z2W9, Q5DTJ0 | Gene names | Gria3, Glur3, Kiaa4184 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate receptor 3 precursor (GluR-3) (GluR-C) (GluR-K3) (Glutamate receptor ionotropic, AMPA 3) (AMPA-selective glutamate receptor 3). | |||||
|
GRIA2_MOUSE
|
||||||
| NC score | 0.052220 (rank : 45) | θ value | θ > 10 (rank : 29) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P23819, Q61604, Q61605, Q8BG69, Q8BXU3, Q9D6D3 | Gene names | Gria2, Glur2 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor 2 precursor (GluR-2) (GluR-B) (GluR-K2) (Glutamate receptor ionotropic, AMPA 2) (AMPA-selective glutamate receptor 2). | |||||
|
GRIK1_MOUSE
|
||||||
| NC score | 0.051029 (rank : 46) | θ value | θ > 10 (rank : 39) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q60934 | Gene names | Grik1, Glur5 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor, ionotropic kainate 1 precursor (Glutamate receptor 5) (GluR-5) (GluR5). | |||||
|
GNPTA_HUMAN
|
||||||
| NC score | 0.030413 (rank : 47) | θ value | 6.88961 (rank : 22) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q3T906, Q3ZQK2, Q6IPW5, Q86TQ2, Q96N13, Q9ULL2 | Gene names | GNPTAB, GNPTA, KIAA1208 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylglucosamine-1-phosphotransferase subunits alpha/beta precursor (EC 2.7.8.17) (GlcNAc-1-phosphotransferase alpha/beta subunits) (UDP- N-acetylglucosamine-1-phosphotransferase alpha/beta subunits) (Stealth protein GNPTAB) [Contains: N-acetylglucosamine-1-phosphotransferase subunit alpha; N-acetylglucosamine-1-phosphotransferase subunit beta]. | |||||
|
GNPTA_MOUSE
|
||||||
| NC score | 0.030210 (rank : 48) | θ value | 6.88961 (rank : 23) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q69ZN6, Q3U3K6, Q3US34 | Gene names | Gnptab, Gnpta, Kiaa1208 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylglucosamine-1-phosphotransferase subunits alpha/beta precursor (EC 2.7.8.17) (GlcNAc-1-phosphotransferase alpha/beta subunits) (UDP- N-acetylglucosamine-1-phosphotransferase alpha/beta subunits) (Stealth protein GNPTAB) [Contains: N-acetylglucosamine-1-phosphotransferase subunit alpha; N-acetylglucosamine-1-phosphotransferase subunit beta]. | |||||
|
COT1_MOUSE
|
||||||
| NC score | 0.008220 (rank : 49) | θ value | 6.88961 (rank : 21) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60632, Q61438 | Gene names | Nr2f1, Erbal3, Tfcoup1 | |||
|
Domain Architecture |

|
|||||
| Description | COUP transcription factor 1 (COUP-TF1) (COUP-TF I). | |||||
|
COT1_HUMAN
|
||||||
| NC score | 0.008215 (rank : 50) | θ value | 6.88961 (rank : 20) | |||
| Query Neighborhood Hits | 23 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P10589 | Gene names | NR2F1, EAR3, ERBAL3, TFCOUP1 | |||
|
Domain Architecture |

|
|||||
| Description | COUP transcription factor 1 (COUP-TF1) (COUP-TF I) (V-ERBA-related protein EAR-3). | |||||