Please be patient as the page loads
|
ZC3H3_HUMAN
|
||||||
| SwissProt Accessions | Q8IXZ2, Q14163, Q8N4E2, Q9BUS4 | Gene names | ZC3H3, KIAA0150, ZC3HDC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 3. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ZC3H3_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 159 | |
| SwissProt Accessions | Q8IXZ2, Q14163, Q8N4E2, Q9BUS4 | Gene names | ZC3H3, KIAA0150, ZC3HDC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 3. | |||||
|
ZC3H3_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.941185 (rank : 2) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q8CHP0, Q80X78 | Gene names | Zc3h3, Kiaa0150, Zc3hdc3, Zfp623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 3. | |||||
|
CPSF4_HUMAN
|
||||||
| θ value | 1.47631e-18 (rank : 3) | NC score | 0.638360 (rank : 3) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O95639, Q86TF8, Q9BTW6 | Gene names | CPSF4, CPSF30, NAR, NEB1 | |||
|
Domain Architecture |
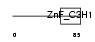
|
|||||
| Description | Cleavage and polyadenylation specificity factor 30 kDa subunit (CPSF 30 kDa subunit) (NS1 effector domain-binding protein 1) (Neb-1) (No arches homolog). | |||||
|
CPSF4_MOUSE
|
||||||
| θ value | 1.47974e-10 (rank : 4) | NC score | 0.580025 (rank : 4) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8BQZ5, O54930 | Gene names | Cpsf4, Cpsf30 | |||
|
Domain Architecture |
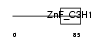
|
|||||
| Description | Cleavage and polyadenylation specificity factor 30 kDa subunit (CPSF 30 kDa subunit) (Clipper homolog) (Clipper/CPSF 30K). | |||||
|
ZC3H6_MOUSE
|
||||||
| θ value | 1.53129e-07 (rank : 5) | NC score | 0.394327 (rank : 7) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8BYK8, Q80UK3, Q8C1H1, Q9D604 | Gene names | Zc3h6, Zc3hdc6 | |||
|
Domain Architecture |
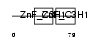
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
MKRN2_HUMAN
|
||||||
| θ value | 9.92553e-07 (rank : 6) | NC score | 0.335230 (rank : 12) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9H000, Q8N391, Q96BD4, Q9BUY2, Q9NRY1 | Gene names | MKRN2, RNF62 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-2 (RING finger protein 62). | |||||
|
ZC3H6_HUMAN
|
||||||
| θ value | 2.88788e-06 (rank : 7) | NC score | 0.387908 (rank : 8) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P61129 | Gene names | ZC3H6, KIAA2035, ZC3HDC6 | |||
|
Domain Architecture |
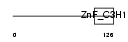
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
MKRN2_MOUSE
|
||||||
| θ value | 3.77169e-06 (rank : 8) | NC score | 0.335697 (rank : 11) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9ERV1, Q9D0L9 | Gene names | Mkrn2 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-2. | |||||
|
MBN3_HUMAN
|
||||||
| θ value | 8.40245e-06 (rank : 9) | NC score | 0.327497 (rank : 13) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9NUK0, Q5JXN8, Q5JXN9, Q5JXP4, Q6UDQ1, Q8TAD9, Q8TAF4, Q9H0Z7, Q9UF37 | Gene names | MBNL3, CHCR, MBLX39, MBXL | |||
|
Domain Architecture |

|
|||||
| Description | Muscleblind-like X-linked protein (Muscleblind-like protein 3) (Cys3His CCG1-required protein) (Protein HCHCR). | |||||
|
MBN3_MOUSE
|
||||||
| θ value | 1.09739e-05 (rank : 10) | NC score | 0.320450 (rank : 14) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8R003 | Gene names | Mbnl3, Chcr, Mbxl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Muscleblind-like X-linked protein (Muscleblind-like protein 3) (Cys3His CCG1-required protein) (Protein MCHCR). | |||||
|
CS007_MOUSE
|
||||||
| θ value | 1.87187e-05 (rank : 11) | NC score | 0.371085 (rank : 9) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 464 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q6ZPZ3 | Gene names | Kiaa1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7 homolog. | |||||
|
CS007_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 12) | NC score | 0.370634 (rank : 10) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9UPT8, Q9Y420 | Gene names | C19orf7, KIAA1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7. | |||||
|
ZC3H8_MOUSE
|
||||||
| θ value | 4.1701e-05 (rank : 13) | NC score | 0.409889 (rank : 6) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9JJ48, Q80X92 | Gene names | Zc3h8, Fliz1, Zc3hdc8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 8 (Fetal liver zinc finger protein 1). | |||||
|
ZC3H8_HUMAN
|
||||||
| θ value | 5.44631e-05 (rank : 14) | NC score | 0.412213 (rank : 5) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8N5P1, Q9BZ75 | Gene names | ZC3H8, ZC3HDC8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 8. | |||||
|
NFH_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 15) | NC score | 0.051262 (rank : 70) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
MBNL_HUMAN
|
||||||
| θ value | 9.29e-05 (rank : 16) | NC score | 0.293255 (rank : 18) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9NR56, O43311, O43797 | Gene names | MBNL1, EXP, KIAA0428, MBNL | |||
|
Domain Architecture |

|
|||||
| Description | Muscleblind-like protein (Triplet-expansion RNA-binding protein). | |||||
|
MBNL_MOUSE
|
||||||
| θ value | 9.29e-05 (rank : 17) | NC score | 0.295762 (rank : 17) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JKP5 | Gene names | Mbnl1, Exp, Mbnl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Muscleblind-like protein (Triplet-expansion RNA-binding protein). | |||||
|
MKRN1_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 18) | NC score | 0.301781 (rank : 16) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9UHC7, Q6GSF1, Q9H0G0, Q9UEZ7 | Gene names | MKRN1, RNF61 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-1 (RING finger protein 61). | |||||
|
MKRN1_MOUSE
|
||||||
| θ value | 0.00020696 (rank : 19) | NC score | 0.305817 (rank : 15) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9QXP6 | Gene names | Mkrn1 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-1. | |||||
|
RC3H1_MOUSE
|
||||||
| θ value | 0.000270298 (rank : 20) | NC score | 0.160297 (rank : 32) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q4VGL6, Q69Z31 | Gene names | Rc3h1, Gm551, Kiaa2025 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Roquin (RING finger and C3H zinc finger protein 1) (Sanroque protein). | |||||
|
SYNJ1_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 21) | NC score | 0.098699 (rank : 41) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O43426, O43425, O94984 | Gene names | SYNJ1, KIAA0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
RC3H1_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 22) | NC score | 0.163687 (rank : 31) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q5TC82, Q5W180, Q5W181, Q8IVE6, Q8N9V1 | Gene names | RC3H1, KIAA2025 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Roquin (RING finger and C3H zinc finger protein 1). | |||||
|
MKRN4_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 23) | NC score | 0.260940 (rank : 21) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q13434 | Gene names | MKRN4, RNF64, ZNF127L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Makorin-4 (Zinc finger protein 127-Xp) (ZNF127-Xp) (RING finger protein 64). | |||||
|
TTP_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 24) | NC score | 0.277060 (rank : 20) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P22893, P11520 | Gene names | Zfp36, Tis11, Tis11a | |||
|
Domain Architecture |
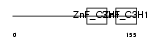
|
|||||
| Description | Tristetraproline (TTP) (Zinc finger protein 36) (Zfp-36) (TIS11A protein) (TIS11) (Growth factor-inducible nuclear protein NUP475) (TPA-induced sequence 11). | |||||
|
TTP_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 25) | NC score | 0.280170 (rank : 19) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P26651 | Gene names | ZFP36, G0S24, TIS11A, TTP | |||
|
Domain Architecture |
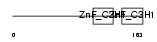
|
|||||
| Description | Tristetraproline (TTP) (Zinc finger protein 36 homolog) (Zfp-36) (TIS11A protein) (TIS11) (Growth factor-inducible nuclear protein NUP475) (G0/G1 switch regulatory protein 24). | |||||
|
ZC3H5_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 26) | NC score | 0.178773 (rank : 26) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8BL48, Q8BVI6, Q99JZ8, Q99LL9 | Gene names | Zc3h5, Kiaa1753, Zc3hdc5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 5. | |||||
|
CHD9_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 27) | NC score | 0.026256 (rank : 117) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BYH8, Q7TMV5, Q8BJG8, Q8BZJ2, Q8CHG8 | Gene names | Chd9, Kiaa0308, Pric320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha- interacting complex protein 320 kDa). | |||||
|
JPH2_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 28) | NC score | 0.050386 (rank : 73) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9BR39, O95913, Q5JY74, Q9UJN4 | Gene names | JPH2, JP2 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
TISB_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 29) | NC score | 0.241852 (rank : 23) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q07352, Q13851 | Gene names | ZFP36L1, BERG36, BRF1, ERF1, TIS11B | |||
|
Domain Architecture |
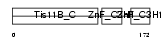
|
|||||
| Description | Butyrate response factor 1 (TIS11B protein) (EGF-response factor 1) (ERF-1). | |||||
|
TISD_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 30) | NC score | 0.240288 (rank : 25) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P47974, Q9BSJ3 | Gene names | ZFP36L2, BRF2, ERF2, TIS11D | |||
|
Domain Architecture |
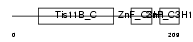
|
|||||
| Description | Butyrate response factor 2 (TIS11D protein) (EGF-response factor 2) (ERF-2). | |||||
|
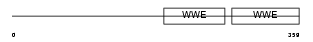
PAR12_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 31) | NC score | 0.138484 (rank : 35) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9H0J9, Q9H610, Q9NP36, Q9NTI3 | Gene names | PARP12, ZC3HDC1 | |||
|
Domain Architecture |
|
|||||
| Description | Poly [ADP-ribose] polymerase 12 (EC 2.4.2.30) (PARP-12) (Zinc finger CCCH domain-containing protein 1). | |||||
|
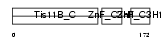
TISB_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 32) | NC score | 0.240294 (rank : 24) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P23950 | Gene names | Zfp36l1, Brf1, Tis11b | |||
|
Domain Architecture |
|
|||||
| Description | Butyrate response factor 1 (TIS11B protein). | |||||
|
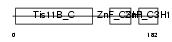
TISD_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 33) | NC score | 0.244239 (rank : 22) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P23949 | Gene names | Zfp36l2, Brf2, Tis11d | |||
|
Domain Architecture |
|
|||||
| Description | Butyrate response factor 2 (TIS11D protein). | |||||
|
SYNJ1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 34) | NC score | 0.087748 (rank : 44) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8CHC4 | Gene names | Synj1, Kiaa0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
MUC1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 35) | NC score | 0.055938 (rank : 64) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
PAR12_MOUSE
|
||||||
| θ value | 0.163984 (rank : 36) | NC score | 0.129935 (rank : 37) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8BZ20, Q80VL6, Q8K333, Q8R3U2 | Gene names | Parp12, Zc3hdc1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 12 (EC 2.4.2.30) (PARP-12) (Zinc finger CCCH domain-containing protein 1). | |||||
|
PARPT_HUMAN
|
||||||
| θ value | 0.163984 (rank : 37) | NC score | 0.106234 (rank : 38) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7Z3E1, Q68CY9, Q86VP4, Q9Y4P7 | Gene names | TIPARP, PARP7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TCDD-inducible poly [ADP-ribose] polymerase (EC 2.4.2.30) (Poly [ADP- ribose] polymerase 7) (PARP-7). | |||||
|
PARPT_MOUSE
|
||||||
| θ value | 0.163984 (rank : 38) | NC score | 0.106001 (rank : 39) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8C1B2, Q3UD50, Q8C032 | Gene names | Tiparp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TCDD-inducible poly [ADP-ribose] polymerase (EC 2.4.2.30). | |||||
|
ZC11A_HUMAN
|
||||||
| θ value | 0.21417 (rank : 39) | NC score | 0.172160 (rank : 29) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O75152, Q6AHY4, Q6AHY9, Q6AW79, Q6AWA1, Q6PJK4, Q86XZ7 | Gene names | ZC3H11A, KIAA0663 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 11A. | |||||
|
DOT1L_HUMAN
|
||||||
| θ value | 0.279714 (rank : 40) | NC score | 0.086716 (rank : 45) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8TEK3, O60379, Q96JL1 | Gene names | DOT1L, KIAA1814 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-79 specific (EC 2.1.1.43) (Histone H3-K79 methyltransferase) (H3-K79-HMTase) (DOT1-like protein). | |||||
|
SPEG_MOUSE
|
||||||
| θ value | 0.279714 (rank : 41) | NC score | 0.013867 (rank : 147) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
ZC3H5_HUMAN
|
||||||
| θ value | 0.279714 (rank : 42) | NC score | 0.170727 (rank : 30) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9C0B0 | Gene names | ZC3H5, KIAA1753, ZC3HDC5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 5. | |||||
|
ZEP1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 43) | NC score | 0.021027 (rank : 128) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1036 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P15822, Q14122 | Gene names | HIVEP1, ZNF40 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 40 (Human immunodeficiency virus type I enhancer- binding protein 1) (HIV-EP1) (Major histocompatibility complex-binding protein 1) (MBP-1) (Positive regulatory domain II-binding factor 1) (PRDII-BF1). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 0.365318 (rank : 44) | NC score | 0.065100 (rank : 52) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
MNAB_MOUSE
|
||||||
| θ value | 0.47712 (rank : 45) | NC score | 0.143578 (rank : 34) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P0C090 | Gene names | Mnab | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated nucleic acid-binding protein. | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 46) | NC score | 0.100921 (rank : 40) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
BAT2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 47) | NC score | 0.079059 (rank : 48) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
MNAB_HUMAN
|
||||||
| θ value | 0.62314 (rank : 48) | NC score | 0.130381 (rank : 36) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9HBD1, Q5JPD7, Q86ST6, Q8N3D6, Q96F27, Q9H5J2, Q9HBD2, Q9NWN9, Q9NXE1 | Gene names | MNAB, RNF164 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated nucleic acid-binding protein (RING finger protein 164). | |||||
|
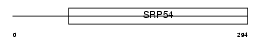
ABCG1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 49) | NC score | 0.023933 (rank : 121) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P45844, Q86SU8, Q96L76, Q9BXK6, Q9BXK7, Q9BXK8, Q9BXK9, Q9BXL0, Q9BXL1, Q9BXL2, Q9BXL3, Q9BXL4 | Gene names | ABCG1, ABC8, WHT1 | |||
|
Domain Architecture |
|
|||||
| Description | ATP-binding cassette sub-family G member 1 (White protein homolog) (ATP-binding cassette transporter 8). | |||||
|
CXAR_HUMAN
|
||||||
| θ value | 0.813845 (rank : 50) | NC score | 0.033533 (rank : 100) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P78310, O00694, Q8WWT6, Q8WWT7, Q8WWT8, Q9UKV4 | Gene names | CXADR, CAR | |||
|
Domain Architecture |
|
|||||
| Description | Coxsackievirus and adenovirus receptor precursor (Coxsackievirus B- adenovirus receptor) (hCAR) (CVB3-binding protein) (HCVADR). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 51) | NC score | 0.081648 (rank : 47) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
ZN750_HUMAN
|
||||||
| θ value | 0.813845 (rank : 52) | NC score | 0.047198 (rank : 79) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q32MQ0, Q9H899 | Gene names | ZNF750 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ZNF750. | |||||
|
CXAR_MOUSE
|
||||||
| θ value | 1.06291 (rank : 53) | NC score | 0.031965 (rank : 103) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P97792, O09052, Q3ULD0, Q91W66, Q99KG0, Q9DBJ8 | Gene names | Cxadr, Car | |||
|
Domain Architecture |
|
|||||
| Description | Coxsackievirus and adenovirus receptor homolog precursor (CAR) (mCAR). | |||||
|
DHX57_MOUSE
|
||||||
| θ value | 1.06291 (rank : 54) | NC score | 0.051468 (rank : 69) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6P5D3, Q3TS93, Q6NZK4, Q6P1B4, Q8BI63, Q8BIA2, Q8R360 | Gene names | Dhx57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX57 (EC 3.6.1.-) (DEAH box protein 57). | |||||
|
DOK7_MOUSE
|
||||||
| θ value | 1.06291 (rank : 55) | NC score | 0.047997 (rank : 77) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q18PE0, Q3TCZ6, Q5FW70, Q8C8U7 | Gene names | Dok7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Dok-7 (Downstream of tyrosine kinase 7). | |||||
|
EVL_MOUSE
|
||||||
| θ value | 1.06291 (rank : 56) | NC score | 0.051604 (rank : 68) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P70429, Q9ERU8 | Gene names | Evl | |||
|
Domain Architecture |
|
|||||
| Description | Ena/VASP-like protein (Ena/vasodilator-stimulated phosphoprotein- like). | |||||
|
FOG1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 57) | NC score | 0.025370 (rank : 118) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O35615 | Gene names | Zfpm1, Fog, Fog1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
RBM22_HUMAN
|
||||||
| θ value | 1.06291 (rank : 58) | NC score | 0.097613 (rank : 42) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NW64, O95607 | Gene names | RBM22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA-splicing factor RBM22 (RNA-binding motif protein 22). | |||||
|
RBM22_MOUSE
|
||||||
| θ value | 1.06291 (rank : 59) | NC score | 0.097613 (rank : 43) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BHS3, Q3TJB0, Q9CXA0 | Gene names | Rbm22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA-splicing factor RBM22 (RNA-binding motif protein 22). | |||||
|
TGON2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 60) | NC score | 0.059747 (rank : 59) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
ZC11A_MOUSE
|
||||||
| θ value | 1.06291 (rank : 61) | NC score | 0.158406 (rank : 33) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6NZF1, Q6NXW9, Q6PDR6, Q80TU7, Q8C5L5, Q99JN6 | Gene names | Zc3h11a, Kiaa0663 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 11A. | |||||
|
BAT2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 62) | NC score | 0.073735 (rank : 49) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
NALP1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 63) | NC score | 0.017054 (rank : 136) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9C000, Q9BZZ8, Q9BZZ9, Q9HAV8, Q9UFT4, Q9Y2E0 | Gene names | NALP1, CARD7, DEFCAP, KIAA0926, NAC | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 1 (Death effector filament- forming ced-4-like apoptosis protein) (Nucleotide-binding domain and caspase recruitment domain) (Caspase recruitment domain protein 7). | |||||
|
NANO1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 64) | NC score | 0.036724 (rank : 94) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8WY41 | Gene names | NANOS1, NOS1 | |||
|
Domain Architecture |

|
|||||
| Description | Nanos homolog 1 (NOS-1) (EC_Rep1a). | |||||
|
U2AFL_MOUSE
|
||||||
| θ value | 1.38821 (rank : 65) | NC score | 0.033445 (rank : 101) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 389 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q64707 | Gene names | Zrsr1, Sp2, Sp2-7, U2af1-rs1, U2af1l1 | |||
|
Domain Architecture |

|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 1 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 1) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 1) (SP2). | |||||
|
U383_MOUSE
|
||||||
| θ value | 1.38821 (rank : 66) | NC score | 0.035329 (rank : 97) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9D2Q2, Q3V1K3 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative methyltransferase UPF0383 (EC 2.1.1.-). | |||||
|
ABCG1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 67) | NC score | 0.021842 (rank : 123) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q64343 | Gene names | Abcg1, Abc8, Wht1 | |||
|
Domain Architecture |
|
|||||
| Description | ATP-binding cassette sub-family G member 1 (White protein homolog) (ATP-binding cassette transporter 8). | |||||
|
ADA15_HUMAN
|
||||||
| θ value | 1.81305 (rank : 68) | NC score | 0.010142 (rank : 156) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q13444, Q13493, Q96C78 | Gene names | ADAM15, MDC15 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 15 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 15) (Metalloproteinase-like, disintegrin-like, and cysteine- rich protein 15) (MDC-15) (Metalloprotease RGD disintegrin protein) (Metargidin). | |||||
|
CAMKV_HUMAN
|
||||||
| θ value | 1.81305 (rank : 69) | NC score | 0.006242 (rank : 161) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1084 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8NCB2, Q6FIB8, Q8NBS8, Q8NC85, Q8NDU4, Q8WTT8, Q9BQC9, Q9H0Q5 | Gene names | CAMKV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CaM kinase-like vesicle-associated protein. | |||||
|
CSF1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 70) | NC score | 0.049967 (rank : 75) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P09603, Q13130, Q14086, Q14806, Q9UQR8 | Gene names | CSF1 | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage colony-stimulating factor 1 precursor (CSF-1) (MCSF) (M- CSF) (Lanimostim). | |||||
|
DHX57_HUMAN
|
||||||
| θ value | 1.81305 (rank : 71) | NC score | 0.057297 (rank : 62) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q6P158, Q53SI4, Q6P9G1, Q7Z6H3, Q8NG17, Q96M33 | Gene names | DHX57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX57 (EC 3.6.1.-) (DEAH box protein 57). | |||||
|
INVS_MOUSE
|
||||||
| θ value | 1.81305 (rank : 72) | NC score | 0.015812 (rank : 142) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O89019, O88849 | Gene names | Invs, Inv, Nphp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning protein) (Nephrocystin-2). | |||||
|
K1802_MOUSE
|
||||||
| θ value | 1.81305 (rank : 73) | NC score | 0.041123 (rank : 87) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 1.81305 (rank : 74) | NC score | 0.062643 (rank : 55) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
PDLI5_HUMAN
|
||||||
| θ value | 1.81305 (rank : 75) | NC score | 0.016109 (rank : 140) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 322 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96HC4, O60705 | Gene names | PDLIM5, ENH | |||
|
Domain Architecture |

|
|||||
| Description | PDZ and LIM domain protein 5 (Enigma homolog) (Enigma-like PDZ and LIM domains protein). | |||||
|
SEPT9_HUMAN
|
||||||
| θ value | 1.81305 (rank : 76) | NC score | 0.035329 (rank : 98) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 706 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9UHD8, Q96QF3, Q96QF4, Q96QF5, Q9HA04, Q9UG40, Q9Y5W4 | Gene names | SEPT9, KIAA0991, MSF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Septin-9 (MLL septin-like fusion protein) (MLL septin-like fusion protein MSF-A) (Ovarian/Breast septin) (Ov/Br septin) (Septin D1). | |||||
|
TARA_HUMAN
|
||||||
| θ value | 1.81305 (rank : 77) | NC score | 0.044288 (rank : 83) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
ZN330_HUMAN
|
||||||
| θ value | 1.81305 (rank : 78) | NC score | 0.062804 (rank : 53) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y3S2 | Gene names | ZNF330, NOA36 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 330 (Nucleolar cysteine-rich protein) (Nucleolar autoantigen 36). | |||||
|
ZN330_MOUSE
|
||||||
| θ value | 1.81305 (rank : 79) | NC score | 0.062743 (rank : 54) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q922H9, Q8C389, Q8K2M4 | Gene names | Znf330, Noa36, Zfp330 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 330 (Nucleolar cysteine-rich protein) (Nucleolar autoantigen 36). | |||||
|
BIN1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 80) | NC score | 0.030883 (rank : 106) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O00499, O00297, O00545, O43867, O60552, O60553, O60554, O60555, O75514, O75515, O75516, O75517, O75518, Q92944, Q99688 | Gene names | BIN1, AMPHL | |||
|
Domain Architecture |

|
|||||
| Description | Myc box-dependent-interacting protein 1 (Bridging integrator 1) (Amphiphysin-like protein) (Amphiphysin II) (Box-dependent myc- interacting protein 1). | |||||
|
CAC1A_MOUSE
|
||||||
| θ value | 2.36792 (rank : 81) | NC score | 0.013120 (rank : 148) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P97445 | Gene names | Cacna1a, Cach4, Cacn3, Cacnl1a4, Ccha1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
EVL_HUMAN
|
||||||
| θ value | 2.36792 (rank : 82) | NC score | 0.052694 (rank : 66) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UI08, O95884, Q7Z522, Q8TBV1, Q9UF25, Q9UIC2 | Gene names | EVL, RNB6 | |||
|
Domain Architecture |

|
|||||
| Description | Ena/VASP-like protein (Ena/vasodilator-stimulated phosphoprotein- like). | |||||
|
MBB1A_HUMAN
|
||||||
| θ value | 2.36792 (rank : 83) | NC score | 0.047028 (rank : 80) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
SPIT1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 84) | NC score | 0.028195 (rank : 112) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43278, Q7Z7D2 | Gene names | SPINT1, HAI1 | |||
|
Domain Architecture |

|
|||||
| Description | Kunitz-type protease inhibitor 1 precursor (Hepatocyte growth factor activator inhibitor type 1) (HAI-1). | |||||
|
U2AF1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 85) | NC score | 0.046352 (rank : 81) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q01081 | Gene names | U2AF1, U2AF35 | |||
|
Domain Architecture |
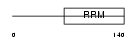
|
|||||
| Description | Splicing factor U2AF 35 kDa subunit (U2 auxiliary factor 35 kDa subunit) (U2 snRNP auxiliary factor small subunit). | |||||
|
U2AF1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 86) | NC score | 0.046335 (rank : 82) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9D883, Q564E4, Q99LX2, Q9CZ98 | Gene names | U2af1 | |||
|
Domain Architecture |
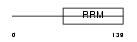
|
|||||
| Description | Splicing factor U2AF 35 kDa subunit (U2 auxiliary factor 35 kDa subunit) (U2 snRNP auxiliary factor small subunit). | |||||
|
ZN438_HUMAN
|
||||||
| θ value | 2.36792 (rank : 87) | NC score | 0.009573 (rank : 157) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q7Z4V0, Q5T426, Q658Q4, Q6ZN65 | Gene names | ZNF438 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 438. | |||||
|
ADA17_HUMAN
|
||||||
| θ value | 3.0926 (rank : 88) | NC score | 0.033717 (rank : 99) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P78536, O60226 | Gene names | ADAM17, CSVP, TACE | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 17 precursor (EC 3.4.24.86) (A disintegrin and metalloproteinase domain 17) (TNF-alpha-converting enzyme) (TNF-alpha convertase) (Snake venom-like protease) (CD156b antigen). | |||||
|
ALMS1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 89) | NC score | 0.030093 (rank : 109) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8TCU4, Q53S05, Q580Q8, Q86VP9, Q9Y4G4 | Gene names | ALMS1, KIAA0328 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alstrom syndrome protein 1. | |||||
|
BSN_HUMAN
|
||||||
| θ value | 3.0926 (rank : 90) | NC score | 0.068655 (rank : 50) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
GFI1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 91) | NC score | 0.001020 (rank : 169) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 736 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P70338, O09063 | Gene names | Gfi1, Gfi-1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein Gfi-1 (Growth factor independence 1). | |||||
|
H12_HUMAN
|
||||||
| θ value | 3.0926 (rank : 92) | NC score | 0.028709 (rank : 111) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P16403 | Gene names | HIST1H1C, H1F2 | |||
|
Domain Architecture |
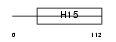
|
|||||
| Description | Histone H1.2 (Histone H1d). | |||||
|
MEFV_MOUSE
|
||||||
| θ value | 3.0926 (rank : 93) | NC score | 0.020804 (rank : 130) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JJ26 | Gene names | Mefv | |||
|
Domain Architecture |

|
|||||
| Description | Pyrin (Marenostrin). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 94) | NC score | 0.085016 (rank : 46) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
SSA27_MOUSE
|
||||||
| θ value | 3.0926 (rank : 95) | NC score | 0.032135 (rank : 102) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P56873 | Gene names | Sssca1, C184l | |||
|
Domain Architecture |
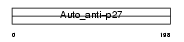
|
|||||
| Description | Sjoegren syndrome/scleroderma autoantigen 1 homolog (Autoantigen p27 homolog) (Protein C184L). | |||||
|
T22D2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 96) | NC score | 0.048288 (rank : 76) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O75157, Q6PI50, Q9H2Z6, Q9H2Z7, Q9H2Z8 | Gene names | TSC22D2, KIAA0669, TILZ4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TSC22 domain family protein 2 (TSC22-related-inducible leucine zipper protein 4). | |||||
|
ZN687_HUMAN
|
||||||
| θ value | 3.0926 (rank : 97) | NC score | 0.012092 (rank : 151) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8N1G0, Q68DQ8, Q9H937, Q9P2A7 | Gene names | ZNF687, KIAA1441 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 687. | |||||
|
AFF2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 98) | NC score | 0.031849 (rank : 104) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O55112 | Gene names | Aff2, Fmr2, Ox19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation protein 2 homolog) (Protein FMR-2) (FMR2P) (Protein Ox19). | |||||
|
AFF4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 99) | NC score | 0.038077 (rank : 92) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
BTBD9_MOUSE
|
||||||
| θ value | 4.03905 (rank : 100) | NC score | 0.008941 (rank : 158) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C726 | Gene names | Btbd9 | |||
|
Domain Architecture |
|
|||||
| Description | BTB/POZ domain-containing protein 9. | |||||
|
CCD45_MOUSE
|
||||||
| θ value | 4.03905 (rank : 101) | NC score | 0.020624 (rank : 131) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 663 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BVV7 | Gene names | Ccdc45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 45. | |||||
|
CD017_HUMAN
|
||||||
| θ value | 4.03905 (rank : 102) | NC score | 0.037217 (rank : 93) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q53FE4, Q6FI84, Q6IS77, Q9H0D9 | Gene names | C4orf17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C4orf17. | |||||
|
CTND2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 103) | NC score | 0.024124 (rank : 120) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UQB3, O00379, O15390, O43206, O43840, Q13589, Q9UM66, Q9UPM3 | Gene names | CTNND2, NPRAP | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Delta-catenin) (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin) (GT24). | |||||
|
CYTSA_HUMAN
|
||||||
| θ value | 4.03905 (rank : 104) | NC score | 0.011390 (rank : 154) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1138 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q69YQ0, O15081 | Gene names | CYTSA, KIAA0376 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytospin-A (NY-REN-22 antigen). | |||||
|
EGLN1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 105) | NC score | 0.019437 (rank : 133) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91YE3, Q8VHJ2, Q922P3 | Gene names | Egln1 | |||
|
Domain Architecture |
|
|||||
| Description | Egl nine homolog 1 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 2) (HIF-prolyl hydroxylase 2) (HIF-PH2) (HPH-2) (SM-20). | |||||
|
FBX42_HUMAN
|
||||||
| θ value | 4.03905 (rank : 106) | NC score | 0.021115 (rank : 126) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6P3S6, Q5TEU8, Q86XI0, Q8N3N4, Q8N5F8, Q9BRM0, Q9P2L4 | Gene names | FBXO42, FBX42, KIAA1332 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 42. | |||||
|
GRN_MOUSE
|
||||||
| θ value | 4.03905 (rank : 107) | NC score | 0.014197 (rank : 145) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P28798 | Gene names | Grn | |||
|
Domain Architecture |

|
|||||
| Description | Granulins precursor (Proepithelin) (PEPI) (PC cell-derived growth factor) (PCDGF) [Contains: Acrogranin; Granulin-1; Granulin-2; Granulin-3; Granulin-4; Granulin-5; Granulin-6; Granulin-7]. | |||||
|
IF16_HUMAN
|
||||||
| θ value | 4.03905 (rank : 108) | NC score | 0.026937 (rank : 114) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q16666, Q9UH78 | Gene names | IFI16, IFNGIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-interferon-inducible protein Ifi-16 (Interferon-inducible myeloid differentiation transcriptional activator) (IFI 16). | |||||
|
KCNB2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 109) | NC score | 0.007478 (rank : 159) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q92953, Q9BXD3 | Gene names | KCNB2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 2 (Voltage-gated potassium channel subunit Kv2.2). | |||||
|
MAGD2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 110) | NC score | 0.011579 (rank : 152) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UNF1, O76058, Q5BJF3, Q8NAL6, Q9H218, Q9P0U9, Q9UM52 | Gene names | MAGED2, BCG1 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen D2 (MAGE-D2 antigen) (MAGE-D) (Breast cancer-associated gene 1 protein) (BCG-1) (11B6) (Hepatocellular carcinoma-associated protein JCL-1). | |||||
|
RNZ2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 111) | NC score | 0.021418 (rank : 125) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80Y81, Q99MF0, Q99MF1, Q9CTA2, Q9D1A8, Q9EPZ2 | Gene names | Elac2 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc phosphodiesterase ELAC protein 2 (EC 3.1.26.11) (Ribonuclease Z 2) (RNase Z 2) (tRNase Z 2) (tRNA 3 endonuclease 2) (ElaC homolog protein 2). | |||||
|
SREC2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 112) | NC score | 0.012538 (rank : 149) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 605 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P59222 | Gene names | Scarf2, Srec2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor class F member 2 precursor (Scavenger receptor expressed by endothelial cells 2 protein) (SREC-II). | |||||
|
AMOT_HUMAN
|
||||||
| θ value | 5.27518 (rank : 113) | NC score | 0.026666 (rank : 116) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q4VCS5, Q504X5, Q9HD27, Q9UPT1 | Gene names | AMOT, KIAA1071 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin. | |||||
|
ATX2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 114) | NC score | 0.038803 (rank : 89) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O70305, P97421 | Gene names | Atxn2, Atx2, Sca2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2 (Spinocerebellar ataxia type 2 protein homolog). | |||||
|
EVX1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 115) | NC score | 0.003910 (rank : 165) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P23683 | Gene names | Evx1 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox even-skipped homolog protein 1 (EVX-1). | |||||
|
GTSE1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 116) | NC score | 0.038224 (rank : 90) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8R080, O89015, Q9CSG9 | Gene names | Gtse1, B99 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G2 and S phase expressed protein 1 (Gtse-1) (B99 protein). | |||||
|
MAP4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 117) | NC score | 0.042996 (rank : 86) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P27546 | Gene names | Map4, Mtap4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
MDC1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 118) | NC score | 0.047849 (rank : 78) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
NEST_HUMAN
|
||||||
| θ value | 5.27518 (rank : 119) | NC score | 0.012503 (rank : 150) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1305 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P48681, O00552 | Gene names | NES | |||
|
Domain Architecture |

|
|||||
| Description | Nestin. | |||||
|
PHF12_MOUSE
|
||||||
| θ value | 5.27518 (rank : 120) | NC score | 0.021066 (rank : 127) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5SPL2, Q5SPL3, Q6ZPN8, Q80W50 | Gene names | Phf12, Kiaa1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
PRB4S_HUMAN
|
||||||
| θ value | 5.27518 (rank : 121) | NC score | 0.044022 (rank : 85) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
REPS1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 122) | NC score | 0.018041 (rank : 135) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O54916, Q8C9J9, Q99LR8 | Gene names | Reps1 | |||
|
Domain Architecture |

|
|||||
| Description | RalBP1-associated Eps domain-containing protein 1 (RalBP1-interacting protein 1). | |||||
|
RFWD2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 123) | NC score | 0.016238 (rank : 139) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NHY2, Q6H103, Q9H6L7 | Gene names | RFWD2, COP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger and WD repeat domain protein 2 (EC 6.3.2.-) (Ubiquitin- protein ligase COP1) (Constitutive photomorphogenesis protein 1 homolog) (hCOP1). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 124) | NC score | 0.036358 (rank : 95) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
SYN2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 125) | NC score | 0.026771 (rank : 115) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q64332, Q6NZR0, Q9QWV7 | Gene names | Syn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synapsin-2 (Synapsin II). | |||||
|
TEP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 126) | NC score | 0.007295 (rank : 160) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P97499 | Gene names | Tep1, Tp1 | |||
|
Domain Architecture |

|
|||||
| Description | Telomerase protein component 1 (Telomerase-associated protein 1) (Telomerase protein 1) (p240) (p80 homolog). | |||||
|
TRI31_HUMAN
|
||||||
| θ value | 5.27518 (rank : 127) | NC score | 0.014274 (rank : 144) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 299 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9BZY9, Q53H52, Q5RI37, Q5SRJ7, Q5SRJ8, Q5SS28, Q96AK4, Q96AP8, Q99579, Q9BZY8 | Gene names | TRIM31, C6orf13, RNF | |||
|
Domain Architecture |
|
|||||
| Description | Tripartite motif-containing protein 31. | |||||
|
UBXD2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 128) | NC score | 0.016960 (rank : 137) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92575, Q8IYM5 | Gene names | UBXD2, KIAA0242, UBXDC1 | |||
|
Domain Architecture |

|
|||||
| Description | UBX domain-containing protein 2. | |||||
|
CT174_HUMAN
|
||||||
| θ value | 6.88961 (rank : 129) | NC score | 0.055010 (rank : 65) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q5JPB2, Q5TDR4, Q8TCP0 | Gene names | C20orf174 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf174. | |||||
|
HMGN3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 130) | NC score | 0.025177 (rank : 119) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15651, Q7RTT0, Q969M5, Q9BZT7 | Gene names | HMGN3, TRIP7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group nucleosome-binding domain-containing protein 3 (Thyroid receptor-interacting protein 7) (TRIP7). | |||||
|
KKCC1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 131) | NC score | -0.000280 (rank : 170) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 866 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8VBY2, Q9R054 | Gene names | Camkk1, Camkk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium/calmodulin-dependent protein kinase kinase 1 (EC 2.7.11.17) (Calcium/calmodulin-dependent protein kinase kinase alpha) (CaM-kinase kinase alpha) (CaM-KK alpha) (CaMKK alpha) (CaMKK 1) (CaM-kinase IV kinase). | |||||
|
M3K14_HUMAN
|
||||||
| θ value | 6.88961 (rank : 132) | NC score | 0.002056 (rank : 166) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 887 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q99558, Q8IYN1 | Gene names | MAP3K14, NIK | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 14 (EC 2.7.11.25) (NF- kappa beta-inducing kinase) (Serine/threonine-protein kinase NIK) (HsNIK). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 133) | NC score | 0.035918 (rank : 96) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
SALL2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 134) | NC score | 0.006019 (rank : 162) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9QX96 | Gene names | Sall2, Sal2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 2 (Spalt-like protein 2) (MSal-2). | |||||
|
SPHK2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 135) | NC score | 0.013965 (rank : 146) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NRA0, Q9BRN1, Q9H0Q2, Q9NWU7 | Gene names | SPHK2 | |||
|
Domain Architecture |
|
|||||
| Description | Sphingosine kinase 2 (EC 2.7.1.-) (SK 2) (SPK 2). | |||||
|
STRAD_MOUSE
|
||||||
| θ value | 6.88961 (rank : 136) | NC score | 0.023382 (rank : 122) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q60924 | Gene names | Stra13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-inducible E3 protein (Hematopoietic-specific protein E3). | |||||
|
ZN217_HUMAN
|
||||||
| θ value | 6.88961 (rank : 137) | NC score | 0.004821 (rank : 164) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 905 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O75362 | Gene names | ZNF217, ZABC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 217. | |||||
|
ZN297_MOUSE
|
||||||
| θ value | 6.88961 (rank : 138) | NC score | 0.005918 (rank : 163) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 816 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Z0G7 | Gene names | Znf297, Bing1, Zfp297 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 297 (BING1 protein). | |||||
|
ZN557_HUMAN
|
||||||
| θ value | 6.88961 (rank : 139) | NC score | -0.001022 (rank : 171) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 752 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8N988, Q6PEJ3, Q9BTZ1 | Gene names | ZNF557 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 557. | |||||
|
ZN703_HUMAN
|
||||||
| θ value | 6.88961 (rank : 140) | NC score | 0.030932 (rank : 105) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9H7S9, Q5XG76 | Gene names | ZNF703 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 703. | |||||
|
3BP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 141) | NC score | 0.016796 (rank : 138) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P55194, Q99KK8 | Gene names | Sh3bp1, 3bp1 | |||
|
Domain Architecture |
|
|||||
| Description | SH3 domain-binding protein 1 (3BP-1). | |||||
|
ADA17_MOUSE
|
||||||
| θ value | 8.99809 (rank : 142) | NC score | 0.027903 (rank : 113) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z0F8, O88726, Q9R1U4, Q9Z0K3 | Gene names | Adam17, Tace | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 17 precursor (EC 3.4.24.86) (A disintegrin and metalloproteinase domain 17) (TNF-alpha-converting enzyme) (TNF-alpha convertase) (CD156b antigen). | |||||
|
CAC1G_HUMAN
|
||||||
| θ value | 8.99809 (rank : 143) | NC score | 0.021780 (rank : 124) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 587 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O43497, O43498, O94770, Q9NYU4, Q9NYU5, Q9NYU6, Q9NYU7, Q9NYU8, Q9NYU9, Q9NYV0, Q9NYV1, Q9UHN9, Q9UHP0, Q9ULU6, Q9UNG7, Q9Y5T2, Q9Y5T3 | Gene names | CACNA1G, KIAA1123 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1G (Voltage- gated calcium channel subunit alpha Cav3.1) (Cav3.1c) (NBR13). | |||||
|
DBPA_MOUSE
|
||||||
| θ value | 8.99809 (rank : 144) | NC score | 0.015900 (rank : 141) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9JKB3, Q80WG4, Q9EQF7, Q9EQF8 | Gene names | Csda, Msy4, Ybx3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA-binding protein A (Cold shock domain-containing protein A) (Y-box protein 3). | |||||
|
DPOLL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 145) | NC score | 0.011519 (rank : 153) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9QXE2, Q9CTJ1 | Gene names | Poll, Polk | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase lambda (EC 2.7.7.7) (EC 4.2.99.-) (Pol Lambda) (DNA polymerase kappa). | |||||
|
M3K1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 146) | NC score | 0.001727 (rank : 167) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P53349, Q60831, Q9R0U3, Q9R256 | Gene names | Map3k1, Mekk, Mekk1 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 1 (EC 2.7.11.25) (MAPK/ERK kinase kinase 1) (MEK kinase 1) (MEKK 1). | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 147) | NC score | 0.029383 (rank : 110) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
MUC1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 148) | NC score | 0.067016 (rank : 51) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
PER3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 149) | NC score | 0.019917 (rank : 132) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
PHF2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 150) | NC score | 0.015077 (rank : 143) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
PRB3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 151) | NC score | 0.041056 (rank : 88) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
PSL1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 152) | NC score | 0.030182 (rank : 108) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8TCT7, O60365, Q567S3, Q8IUH9, Q9BUY6, Q9H3M4, Q9NPN2, Q9P1Z6 | Gene names | SPPL2B, KIAA1532, PSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal peptide peptidase-like 2B (EC 3.4.23.-) (Protein SPP-like 2B) (Protein SPPL2b) (Intramembrane protease 4) (IMP4) (Presenilin-like protein 1). | |||||
|
RP1L1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 153) | NC score | 0.030335 (rank : 107) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
SET1A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 154) | NC score | 0.044079 (rank : 84) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
SF3A1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 155) | NC score | 0.018901 (rank : 134) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8K4Z5, Q8C0M7, Q8C128, Q8C175, Q921T3 | Gene names | Sf3a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 3 subunit 1 (SF3a120). | |||||
|
SYBU_MOUSE
|
||||||
| θ value | 8.99809 (rank : 156) | NC score | 0.010249 (rank : 155) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BHS8, Q3TQN7, Q401F3, Q401F4, Q571C1, Q6P1J2, Q80XH0, Q8BHS7, Q8BI27 | Gene names | Sybu, Golsyn, Kiaa1472 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntabulin (Syntaxin-1-binding protein) (Golgi-localized syntaphilin- related protein) (m-Golsyn). | |||||
|
TNIK_HUMAN
|
||||||
| θ value | 8.99809 (rank : 157) | NC score | 0.001237 (rank : 168) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1570 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UKE5, O60298, Q8WUY7, Q9UKD8, Q9UKD9, Q9UKE0, Q9UKE1, Q9UKE2, Q9UKE3, Q9UKE4 | Gene names | TNIK, KIAA0551 | |||
|
Domain Architecture |

|
|||||
| Description | TRAF2 and NCK-interacting protein kinase (EC 2.7.11.1). | |||||
|
VASP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 158) | NC score | 0.038201 (rank : 91) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P50552, Q6PIZ1, Q93035 | Gene names | VASP | |||
|
Domain Architecture |
|
|||||
| Description | Vasodilator-stimulated phosphoprotein (VASP). | |||||
|
ZEP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 159) | NC score | 0.020816 (rank : 129) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1063 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q03172 | Gene names | Hivep1, Cryabp1, Znf40 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 40 (Transcription factor alphaA-CRYBP1) (Alpha A- crystallin-binding protein I) (Alpha A-CRYBP1). | |||||
|
AFF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.061534 (rank : 57) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
BASP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.050518 (rank : 72) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
BASP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.050705 (rank : 71) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
CEND_HUMAN
|
||||||
| θ value | θ > 10 (rank : 163) | NC score | 0.056200 (rank : 63) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8N111, Q9NYM6 | Gene names | CEND1, BM88 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell cycle exit and neuronal differentiation protein 1 (BM88 antigen). | |||||
|
CK024_HUMAN
|
||||||
| θ value | θ > 10 (rank : 164) | NC score | 0.058654 (rank : 60) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96F05, Q9H2K4 | Gene names | C11orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf24 precursor (Protein DM4E3). | |||||
|
LEUK_HUMAN
|
||||||
| θ value | θ > 10 (rank : 165) | NC score | 0.050242 (rank : 74) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P16150 | Gene names | SPN, CD43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leukosialin precursor (Leukocyte sialoglycoprotein) (Sialophorin) (Galactoglycoprotein) (GALGP) (CD43 antigen). | |||||
|
MKRN3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 166) | NC score | 0.177514 (rank : 27) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q13064 | Gene names | MKRN3, RNF63, ZNF127 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-3 (Zinc finger protein 127) (RING finger protein 63). | |||||
|
MKRN3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 167) | NC score | 0.173950 (rank : 28) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q60764, Q7TQE4 | Gene names | Mkrn3, Zfp127, Znf127 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-3 (Zinc finger protein 127). | |||||
|
MUC7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 168) | NC score | 0.060210 (rank : 58) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
NOLC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 169) | NC score | 0.058521 (rank : 61) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
PRP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 170) | NC score | 0.052293 (rank : 67) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 171) | NC score | 0.061976 (rank : 56) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
ZC3H3_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 159 | |
| SwissProt Accessions | Q8IXZ2, Q14163, Q8N4E2, Q9BUS4 | Gene names | ZC3H3, KIAA0150, ZC3HDC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 3. | |||||
|
ZC3H3_MOUSE
|
||||||
| NC score | 0.941185 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q8CHP0, Q80X78 | Gene names | Zc3h3, Kiaa0150, Zc3hdc3, Zfp623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 3. | |||||
|
CPSF4_HUMAN
|
||||||
| NC score | 0.638360 (rank : 3) | θ value | 1.47631e-18 (rank : 3) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O95639, Q86TF8, Q9BTW6 | Gene names | CPSF4, CPSF30, NAR, NEB1 | |||
|
Domain Architecture |

|
|||||
| Description | Cleavage and polyadenylation specificity factor 30 kDa subunit (CPSF 30 kDa subunit) (NS1 effector domain-binding protein 1) (Neb-1) (No arches homolog). | |||||
|
CPSF4_MOUSE
|
||||||
| NC score | 0.580025 (rank : 4) | θ value | 1.47974e-10 (rank : 4) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8BQZ5, O54930 | Gene names | Cpsf4, Cpsf30 | |||
|
Domain Architecture |

|
|||||
| Description | Cleavage and polyadenylation specificity factor 30 kDa subunit (CPSF 30 kDa subunit) (Clipper homolog) (Clipper/CPSF 30K). | |||||
|
ZC3H8_HUMAN
|
||||||
| NC score | 0.412213 (rank : 5) | θ value | 5.44631e-05 (rank : 14) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8N5P1, Q9BZ75 | Gene names | ZC3H8, ZC3HDC8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 8. | |||||
|
ZC3H8_MOUSE
|
||||||
| NC score | 0.409889 (rank : 6) | θ value | 4.1701e-05 (rank : 13) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9JJ48, Q80X92 | Gene names | Zc3h8, Fliz1, Zc3hdc8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 8 (Fetal liver zinc finger protein 1). | |||||
|
ZC3H6_MOUSE
|
||||||
| NC score | 0.394327 (rank : 7) | θ value | 1.53129e-07 (rank : 5) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8BYK8, Q80UK3, Q8C1H1, Q9D604 | Gene names | Zc3h6, Zc3hdc6 | |||
|
Domain Architecture |
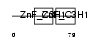
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
ZC3H6_HUMAN
|
||||||
| NC score | 0.387908 (rank : 8) | θ value | 2.88788e-06 (rank : 7) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P61129 | Gene names | ZC3H6, KIAA2035, ZC3HDC6 | |||
|
Domain Architecture |
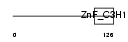
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
CS007_MOUSE
|
||||||
| NC score | 0.371085 (rank : 9) | θ value | 1.87187e-05 (rank : 11) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 464 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q6ZPZ3 | Gene names | Kiaa1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7 homolog. | |||||
|
CS007_HUMAN
|
||||||
| NC score | 0.370634 (rank : 10) | θ value | 4.1701e-05 (rank : 12) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9UPT8, Q9Y420 | Gene names | C19orf7, KIAA1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7. | |||||
|
MKRN2_MOUSE
|
||||||
| NC score | 0.335697 (rank : 11) | θ value | 3.77169e-06 (rank : 8) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9ERV1, Q9D0L9 | Gene names | Mkrn2 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-2. | |||||
|
MKRN2_HUMAN
|
||||||
| NC score | 0.335230 (rank : 12) | θ value | 9.92553e-07 (rank : 6) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9H000, Q8N391, Q96BD4, Q9BUY2, Q9NRY1 | Gene names | MKRN2, RNF62 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-2 (RING finger protein 62). | |||||
|
MBN3_HUMAN
|
||||||
| NC score | 0.327497 (rank : 13) | θ value | 8.40245e-06 (rank : 9) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9NUK0, Q5JXN8, Q5JXN9, Q5JXP4, Q6UDQ1, Q8TAD9, Q8TAF4, Q9H0Z7, Q9UF37 | Gene names | MBNL3, CHCR, MBLX39, MBXL | |||
|
Domain Architecture |

|
|||||
| Description | Muscleblind-like X-linked protein (Muscleblind-like protein 3) (Cys3His CCG1-required protein) (Protein HCHCR). | |||||
|
MBN3_MOUSE
|
||||||
| NC score | 0.320450 (rank : 14) | θ value | 1.09739e-05 (rank : 10) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8R003 | Gene names | Mbnl3, Chcr, Mbxl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Muscleblind-like X-linked protein (Muscleblind-like protein 3) (Cys3His CCG1-required protein) (Protein MCHCR). | |||||
|
MKRN1_MOUSE
|
||||||
| NC score | 0.305817 (rank : 15) | θ value | 0.00020696 (rank : 19) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9QXP6 | Gene names | Mkrn1 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-1. | |||||
|
MKRN1_HUMAN
|
||||||
| NC score | 0.301781 (rank : 16) | θ value | 0.00020696 (rank : 18) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9UHC7, Q6GSF1, Q9H0G0, Q9UEZ7 | Gene names | MKRN1, RNF61 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-1 (RING finger protein 61). | |||||
|
MBNL_MOUSE
|
||||||
| NC score | 0.295762 (rank : 17) | θ value | 9.29e-05 (rank : 17) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JKP5 | Gene names | Mbnl1, Exp, Mbnl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Muscleblind-like protein (Triplet-expansion RNA-binding protein). | |||||
|
MBNL_HUMAN
|
||||||
| NC score | 0.293255 (rank : 18) | θ value | 9.29e-05 (rank : 16) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9NR56, O43311, O43797 | Gene names | MBNL1, EXP, KIAA0428, MBNL | |||
|
Domain Architecture |

|
|||||
| Description | Muscleblind-like protein (Triplet-expansion RNA-binding protein). | |||||
|
TTP_HUMAN
|
||||||
| NC score | 0.280170 (rank : 19) | θ value | 0.0431538 (rank : 25) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P26651 | Gene names | ZFP36, G0S24, TIS11A, TTP | |||
|
Domain Architecture |

|
|||||
| Description | Tristetraproline (TTP) (Zinc finger protein 36 homolog) (Zfp-36) (TIS11A protein) (TIS11) (Growth factor-inducible nuclear protein NUP475) (G0/G1 switch regulatory protein 24). | |||||
|
TTP_MOUSE
|
||||||
| NC score | 0.277060 (rank : 20) | θ value | 0.0148317 (rank : 24) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P22893, P11520 | Gene names | Zfp36, Tis11, Tis11a | |||
|
Domain Architecture |

|
|||||
| Description | Tristetraproline (TTP) (Zinc finger protein 36) (Zfp-36) (TIS11A protein) (TIS11) (Growth factor-inducible nuclear protein NUP475) (TPA-induced sequence 11). | |||||
|
MKRN4_HUMAN
|
||||||
| NC score | 0.260940 (rank : 21) | θ value | 0.00665767 (rank : 23) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q13434 | Gene names | MKRN4, RNF64, ZNF127L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Makorin-4 (Zinc finger protein 127-Xp) (ZNF127-Xp) (RING finger protein 64). | |||||
|
TISD_MOUSE
|
||||||
| NC score | 0.244239 (rank : 22) | θ value | 0.0961366 (rank : 33) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P23949 | Gene names | Zfp36l2, Brf2, Tis11d | |||
|
Domain Architecture |

|
|||||
| Description | Butyrate response factor 2 (TIS11D protein). | |||||
|
TISB_HUMAN
|
||||||
| NC score | 0.241852 (rank : 23) | θ value | 0.0736092 (rank : 29) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q07352, Q13851 | Gene names | ZFP36L1, BERG36, BRF1, ERF1, TIS11B | |||
|
Domain Architecture |

|
|||||
| Description | Butyrate response factor 1 (TIS11B protein) (EGF-response factor 1) (ERF-1). | |||||
|
TISB_MOUSE
|
||||||
| NC score | 0.240294 (rank : 24) | θ value | 0.0961366 (rank : 32) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P23950 | Gene names | Zfp36l1, Brf1, Tis11b | |||
|
Domain Architecture |

|
|||||
| Description | Butyrate response factor 1 (TIS11B protein). | |||||
|
TISD_HUMAN
|
||||||
| NC score | 0.240288 (rank : 25) | θ value | 0.0736092 (rank : 30) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P47974, Q9BSJ3 | Gene names | ZFP36L2, BRF2, ERF2, TIS11D | |||
|
Domain Architecture |

|
|||||
| Description | Butyrate response factor 2 (TIS11D protein) (EGF-response factor 2) (ERF-2). | |||||
|
ZC3H5_MOUSE
|
||||||
| NC score | 0.178773 (rank : 26) | θ value | 0.0563607 (rank : 26) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8BL48, Q8BVI6, Q99JZ8, Q99LL9 | Gene names | Zc3h5, Kiaa1753, Zc3hdc5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 5. | |||||
|
MKRN3_HUMAN
|
||||||
| NC score | 0.177514 (rank : 27) | θ value | θ > 10 (rank : 166) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q13064 | Gene names | MKRN3, RNF63, ZNF127 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-3 (Zinc finger protein 127) (RING finger protein 63). | |||||
|
MKRN3_MOUSE
|
||||||
| NC score | 0.173950 (rank : 28) | θ value | θ > 10 (rank : 167) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q60764, Q7TQE4 | Gene names | Mkrn3, Zfp127, Znf127 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-3 (Zinc finger protein 127). | |||||
|
ZC11A_HUMAN
|
||||||
| NC score | 0.172160 (rank : 29) | θ value | 0.21417 (rank : 39) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O75152, Q6AHY4, Q6AHY9, Q6AW79, Q6AWA1, Q6PJK4, Q86XZ7 | Gene names | ZC3H11A, KIAA0663 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 11A. | |||||
|
ZC3H5_HUMAN
|
||||||
| NC score | 0.170727 (rank : 30) | θ value | 0.279714 (rank : 42) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9C0B0 | Gene names | ZC3H5, KIAA1753, ZC3HDC5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 5. | |||||
|
RC3H1_HUMAN
|
||||||
| NC score | 0.163687 (rank : 31) | θ value | 0.000786445 (rank : 22) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q5TC82, Q5W180, Q5W181, Q8IVE6, Q8N9V1 | Gene names | RC3H1, KIAA2025 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Roquin (RING finger and C3H zinc finger protein 1). | |||||
|
RC3H1_MOUSE
|
||||||
| NC score | 0.160297 (rank : 32) | θ value | 0.000270298 (rank : 20) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q4VGL6, Q69Z31 | Gene names | Rc3h1, Gm551, Kiaa2025 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Roquin (RING finger and C3H zinc finger protein 1) (Sanroque protein). | |||||
|
ZC11A_MOUSE
|
||||||
| NC score | 0.158406 (rank : 33) | θ value | 1.06291 (rank : 61) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6NZF1, Q6NXW9, Q6PDR6, Q80TU7, Q8C5L5, Q99JN6 | Gene names | Zc3h11a, Kiaa0663 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 11A. | |||||
|
MNAB_MOUSE
|
||||||
| NC score | 0.143578 (rank : 34) | θ value | 0.47712 (rank : 45) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P0C090 | Gene names | Mnab | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated nucleic acid-binding protein. | |||||
|
PAR12_HUMAN
|
||||||
| NC score | 0.138484 (rank : 35) | θ value | 0.0961366 (rank : 31) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9H0J9, Q9H610, Q9NP36, Q9NTI3 | Gene names | PARP12, ZC3HDC1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 12 (EC 2.4.2.30) (PARP-12) (Zinc finger CCCH domain-containing protein 1). | |||||
|
MNAB_HUMAN
|
||||||
| NC score | 0.130381 (rank : 36) | θ value | 0.62314 (rank : 48) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9HBD1, Q5JPD7, Q86ST6, Q8N3D6, Q96F27, Q9H5J2, Q9HBD2, Q9NWN9, Q9NXE1 | Gene names | MNAB, RNF164 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated nucleic acid-binding protein (RING finger protein 164). | |||||
|
PAR12_MOUSE
|
||||||
| NC score | 0.129935 (rank : 37) | θ value | 0.163984 (rank : 36) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8BZ20, Q80VL6, Q8K333, Q8R3U2 | Gene names | Parp12, Zc3hdc1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 12 (EC 2.4.2.30) (PARP-12) (Zinc finger CCCH domain-containing protein 1). | |||||
|
PARPT_HUMAN
|
||||||
| NC score | 0.106234 (rank : 38) | θ value | 0.163984 (rank : 37) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7Z3E1, Q68CY9, Q86VP4, Q9Y4P7 | Gene names | TIPARP, PARP7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TCDD-inducible poly [ADP-ribose] polymerase (EC 2.4.2.30) (Poly [ADP- ribose] polymerase 7) (PARP-7). | |||||
|
PARPT_MOUSE
|
||||||
| NC score | 0.106001 (rank : 39) | θ value | 0.163984 (rank : 38) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8C1B2, Q3UD50, Q8C032 | Gene names | Tiparp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TCDD-inducible poly [ADP-ribose] polymerase (EC 2.4.2.30). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.100921 (rank : 40) | θ value | 0.47712 (rank : 46) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
SYNJ1_HUMAN
|
||||||
| NC score | 0.098699 (rank : 41) | θ value | 0.000602161 (rank : 21) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O43426, O43425, O94984 | Gene names | SYNJ1, KIAA0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
RBM22_HUMAN
|
||||||
| NC score | 0.097613 (rank : 42) | θ value | 1.06291 (rank : 58) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NW64, O95607 | Gene names | RBM22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA-splicing factor RBM22 (RNA-binding motif protein 22). | |||||
|
RBM22_MOUSE
|
||||||
| NC score | 0.097613 (rank : 43) | θ value | 1.06291 (rank : 59) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BHS3, Q3TJB0, Q9CXA0 | Gene names | Rbm22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA-splicing factor RBM22 (RNA-binding motif protein 22). | |||||
|
SYNJ1_MOUSE
|
||||||
| NC score | 0.087748 (rank : 44) | θ value | 0.125558 (rank : 34) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8CHC4 | Gene names | Synj1, Kiaa0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
DOT1L_HUMAN
|
||||||
| NC score | 0.086716 (rank : 45) | θ value | 0.279714 (rank : 40) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8TEK3, O60379, Q96JL1 | Gene names | DOT1L, KIAA1814 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-79 specific (EC 2.1.1.43) (Histone H3-K79 methyltransferase) (H3-K79-HMTase) (DOT1-like protein). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.085016 (rank : 46) | θ value | 3.0926 (rank : 94) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.081648 (rank : 47) | θ value | 0.813845 (rank : 51) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
BAT2_HUMAN
|
||||||
| NC score | 0.079059 (rank : 48) | θ value | 0.62314 (rank : 47) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
BAT2_MOUSE
|
||||||
| NC score | 0.073735 (rank : 49) | θ value | 1.38821 (rank : 62) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.068655 (rank : 50) | θ value | 3.0926 (rank : 90) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
MUC1_MOUSE
|
||||||
| NC score | 0.067016 (rank : 51) | θ value | 8.99809 (rank : 148) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.065100 (rank : 52) | θ value | 0.365318 (rank : 44) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
ZN330_HUMAN
|
||||||
| NC score | 0.062804 (rank : 53) | θ value | 1.81305 (rank : 78) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y3S2 | Gene names | ZNF330, NOA36 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 330 (Nucleolar cysteine-rich protein) (Nucleolar autoantigen 36). | |||||
|
ZN330_MOUSE
|
||||||
| NC score | 0.062743 (rank : 54) | θ value | 1.81305 (rank : 79) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q922H9, Q8C389, Q8K2M4 | Gene names | Znf330, Noa36, Zfp330 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 330 (Nucleolar cysteine-rich protein) (Nucleolar autoantigen 36). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.062643 (rank : 55) | θ value | 1.81305 (rank : 74) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.061976 (rank : 56) | θ value | θ > 10 (rank : 171) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
AFF1_HUMAN
|
||||||
| NC score | 0.061534 (rank : 57) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
MUC7_HUMAN
|
||||||
| NC score | 0.060210 (rank : 58) | θ value | θ > 10 (rank : 168) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
TGON2_HUMAN
|
||||||
| NC score | 0.059747 (rank : 59) | θ value | 1.06291 (rank : 60) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
CK024_HUMAN
|
||||||
| NC score | 0.058654 (rank : 60) | θ value | θ > 10 (rank : 164) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96F05, Q9H2K4 | Gene names | C11orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf24 precursor (Protein DM4E3). | |||||
|
NOLC1_HUMAN
|
||||||
| NC score | 0.058521 (rank : 61) | θ value | θ > 10 (rank : 169) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
DHX57_HUMAN
|
||||||
| NC score | 0.057297 (rank : 62) | θ value | 1.81305 (rank : 71) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q6P158, Q53SI4, Q6P9G1, Q7Z6H3, Q8NG17, Q96M33 | Gene names | DHX57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX57 (EC 3.6.1.-) (DEAH box protein 57). | |||||
|
CEND_HUMAN
|
||||||
| NC score | 0.056200 (rank : 63) | θ value | θ > 10 (rank : 163) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8N111, Q9NYM6 | Gene names | CEND1, BM88 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell cycle exit and neuronal differentiation protein 1 (BM88 antigen). | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.055938 (rank : 64) | θ value | 0.163984 (rank : 35) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
CT174_HUMAN
|
||||||
| NC score | 0.055010 (rank : 65) | θ value | 6.88961 (rank : 129) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q5JPB2, Q5TDR4, Q8TCP0 | Gene names | C20orf174 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf174. | |||||
|
EVL_HUMAN
|
||||||
| NC score | 0.052694 (rank : 66) | θ value | 2.36792 (rank : 82) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UI08, O95884, Q7Z522, Q8TBV1, Q9UF25, Q9UIC2 | Gene names | EVL, RNB6 | |||
|
Domain Architecture |

|
|||||
| Description | Ena/VASP-like protein (Ena/vasodilator-stimulated phosphoprotein- like). | |||||
|
PRP1_HUMAN
|
||||||
| NC score | 0.052293 (rank : 67) | θ value | θ > 10 (rank : 170) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
EVL_MOUSE
|
||||||
| NC score | 0.051604 (rank : 68) | θ value | 1.06291 (rank : 56) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P70429, Q9ERU8 | Gene names | Evl | |||
|
Domain Architecture |

|
|||||
| Description | Ena/VASP-like protein (Ena/vasodilator-stimulated phosphoprotein- like). | |||||
|
DHX57_MOUSE
|
||||||
| NC score | 0.051468 (rank : 69) | θ value | 1.06291 (rank : 54) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6P5D3, Q3TS93, Q6NZK4, Q6P1B4, Q8BI63, Q8BIA2, Q8R360 | Gene names | Dhx57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX57 (EC 3.6.1.-) (DEAH box protein 57). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.051262 (rank : 70) | θ value | 7.1131e-05 (rank : 15) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
BASP_MOUSE
|
||||||
| NC score | 0.050705 (rank : 71) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
BASP_HUMAN
|
||||||
| NC score | 0.050518 (rank : 72) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
JPH2_HUMAN
|
||||||
| NC score | 0.050386 (rank : 73) | θ value | 0.0736092 (rank : 28) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9BR39, O95913, Q5JY74, Q9UJN4 | Gene names | JPH2, JP2 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
LEUK_HUMAN
|
||||||
| NC score | 0.050242 (rank : 74) | θ value | θ > 10 (rank : 165) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P16150 | Gene names | SPN, CD43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leukosialin precursor (Leukocyte sialoglycoprotein) (Sialophorin) (Galactoglycoprotein) (GALGP) (CD43 antigen). | |||||
|
CSF1_HUMAN
|
||||||
| NC score | 0.049967 (rank : 75) | θ value | 1.81305 (rank : 70) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P09603, Q13130, Q14086, Q14806, Q9UQR8 | Gene names | CSF1 | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage colony-stimulating factor 1 precursor (CSF-1) (MCSF) (M- CSF) (Lanimostim). | |||||
|
T22D2_HUMAN
|
||||||
| NC score | 0.048288 (rank : 76) | θ value | 3.0926 (rank : 96) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O75157, Q6PI50, Q9H2Z6, Q9H2Z7, Q9H2Z8 | Gene names | TSC22D2, KIAA0669, TILZ4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TSC22 domain family protein 2 (TSC22-related-inducible leucine zipper protein 4). | |||||
|
DOK7_MOUSE
|
||||||
| NC score | 0.047997 (rank : 77) | θ value | 1.06291 (rank : 55) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q18PE0, Q3TCZ6, Q5FW70, Q8C8U7 | Gene names | Dok7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Dok-7 (Downstream of tyrosine kinase 7). | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.047849 (rank : 78) | θ value | 5.27518 (rank : 118) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
ZN750_HUMAN
|
||||||
| NC score | 0.047198 (rank : 79) | θ value | 0.813845 (rank : 52) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q32MQ0, Q9H899 | Gene names | ZNF750 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ZNF750. | |||||
|
MBB1A_HUMAN
|
||||||
| NC score | 0.047028 (rank : 80) | θ value | 2.36792 (rank : 83) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
U2AF1_HUMAN
|
||||||
| NC score | 0.046352 (rank : 81) | θ value | 2.36792 (rank : 85) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q01081 | Gene names | U2AF1, U2AF35 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor U2AF 35 kDa subunit (U2 auxiliary factor 35 kDa subunit) (U2 snRNP auxiliary factor small subunit). | |||||
|
U2AF1_MOUSE
|
||||||
| NC score | 0.046335 (rank : 82) | θ value | 2.36792 (rank : 86) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9D883, Q564E4, Q99LX2, Q9CZ98 | Gene names | U2af1 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor U2AF 35 kDa subunit (U2 auxiliary factor 35 kDa subunit) (U2 snRNP auxiliary factor small subunit). | |||||
|
TARA_HUMAN
|
||||||
| NC score | 0.044288 (rank : 83) | θ value | 1.81305 (rank : 77) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
SET1A_HUMAN
|
||||||
| NC score | 0.044079 (rank : 84) | θ value | 8.99809 (rank : 154) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
PRB4S_HUMAN
|
||||||
| NC score | 0.044022 (rank : 85) | θ value | 5.27518 (rank : 121) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
MAP4_MOUSE
|
||||||
| NC score | 0.042996 (rank : 86) | θ value | 5.27518 (rank : 117) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P27546 | Gene names | Map4, Mtap4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
K1802_MOUSE
|
||||||
| NC score | 0.041123 (rank : 87) | θ value | 1.81305 (rank : 73) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
PRB3_HUMAN
|
||||||
| NC score | 0.041056 (rank : 88) | θ value | 8.99809 (rank : 151) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
ATX2_MOUSE
|
||||||
| NC score | 0.038803 (rank : 89) | θ value | 5.27518 (rank : 114) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O70305, P97421 | Gene names | Atxn2, Atx2, Sca2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2 (Spinocerebellar ataxia type 2 protein homolog). | |||||
|
GTSE1_MOUSE
|
||||||
| NC score | 0.038224 (rank : 90) | θ value | 5.27518 (rank : 116) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8R080, O89015, Q9CSG9 | Gene names | Gtse1, B99 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G2 and S phase expressed protein 1 (Gtse-1) (B99 protein). | |||||
|
VASP_HUMAN
|
||||||
| NC score | 0.038201 (rank : 91) | θ value | 8.99809 (rank : 158) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P50552, Q6PIZ1, Q93035 | Gene names | VASP | |||
|
Domain Architecture |

|
|||||
| Description | Vasodilator-stimulated phosphoprotein (VASP). | |||||
|
AFF4_MOUSE
|
||||||
| NC score | 0.038077 (rank : 92) | θ value | 4.03905 (rank : 99) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
CD017_HUMAN
|
||||||
| NC score | 0.037217 (rank : 93) | θ value | 4.03905 (rank : 102) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q53FE4, Q6FI84, Q6IS77, Q9H0D9 | Gene names | C4orf17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C4orf17. | |||||
|
NANO1_HUMAN
|
||||||
| NC score | 0.036724 (rank : 94) | θ value | 1.38821 (rank : 64) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8WY41 | Gene names | NANOS1, NOS1 | |||
|
Domain Architecture |

|
|||||
| Description | Nanos homolog 1 (NOS-1) (EC_Rep1a). | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.036358 (rank : 95) | θ value | 5.27518 (rank : 124) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.035918 (rank : 96) | θ value | 6.88961 (rank : 133) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
U383_MOUSE
|
||||||
| NC score | 0.035329 (rank : 97) | θ value | 1.38821 (rank : 66) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9D2Q2, Q3V1K3 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative methyltransferase UPF0383 (EC 2.1.1.-). | |||||
|
SEPT9_HUMAN
|
||||||
| NC score | 0.035329 (rank : 98) | θ value | 1.81305 (rank : 76) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 706 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9UHD8, Q96QF3, Q96QF4, Q96QF5, Q9HA04, Q9UG40, Q9Y5W4 | Gene names | SEPT9, KIAA0991, MSF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Septin-9 (MLL septin-like fusion protein) (MLL septin-like fusion protein MSF-A) (Ovarian/Breast septin) (Ov/Br septin) (Septin D1). | |||||
|
ADA17_HUMAN
|
||||||
| NC score | 0.033717 (rank : 99) | θ value | 3.0926 (rank : 88) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P78536, O60226 | Gene names | ADAM17, CSVP, TACE | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 17 precursor (EC 3.4.24.86) (A disintegrin and metalloproteinase domain 17) (TNF-alpha-converting enzyme) (TNF-alpha convertase) (Snake venom-like protease) (CD156b antigen). | |||||
|
CXAR_HUMAN
|
||||||
| NC score | 0.033533 (rank : 100) | θ value | 0.813845 (rank : 50) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P78310, O00694, Q8WWT6, Q8WWT7, Q8WWT8, Q9UKV4 | Gene names | CXADR, CAR | |||
|
Domain Architecture |

|
|||||
| Description | Coxsackievirus and adenovirus receptor precursor (Coxsackievirus B- adenovirus receptor) (hCAR) (CVB3-binding protein) (HCVADR). | |||||
|
U2AFL_MOUSE
|
||||||
| NC score | 0.033445 (rank : 101) | θ value | 1.38821 (rank : 65) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 389 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q64707 | Gene names | Zrsr1, Sp2, Sp2-7, U2af1-rs1, U2af1l1 | |||
|
Domain Architecture |

|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 1 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 1) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 1) (SP2). | |||||
|
SSA27_MOUSE
|
||||||
| NC score | 0.032135 (rank : 102) | θ value | 3.0926 (rank : 95) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P56873 | Gene names | Sssca1, C184l | |||
|
Domain Architecture |
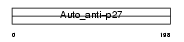
|
|||||
| Description | Sjoegren syndrome/scleroderma autoantigen 1 homolog (Autoantigen p27 homolog) (Protein C184L). | |||||
|
CXAR_MOUSE
|
||||||
| NC score | 0.031965 (rank : 103) | θ value | 1.06291 (rank : 53) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P97792, O09052, Q3ULD0, Q91W66, Q99KG0, Q9DBJ8 | Gene names | Cxadr, Car | |||
|
Domain Architecture |

|
|||||
| Description | Coxsackievirus and adenovirus receptor homolog precursor (CAR) (mCAR). | |||||
|
AFF2_MOUSE
|
||||||
| NC score | 0.031849 (rank : 104) | θ value | 4.03905 (rank : 98) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O55112 | Gene names | Aff2, Fmr2, Ox19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation protein 2 homolog) (Protein FMR-2) (FMR2P) (Protein Ox19). | |||||
|
ZN703_HUMAN
|
||||||
| NC score | 0.030932 (rank : 105) | θ value | 6.88961 (rank : 140) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9H7S9, Q5XG76 | Gene names | ZNF703 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 703. | |||||
|
BIN1_HUMAN
|
||||||
| NC score | 0.030883 (rank : 106) | θ value | 2.36792 (rank : 80) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O00499, O00297, O00545, O43867, O60552, O60553, O60554, O60555, O75514, O75515, O75516, O75517, O75518, Q92944, Q99688 | Gene names | BIN1, AMPHL | |||
|
Domain Architecture |

|
|||||
| Description | Myc box-dependent-interacting protein 1 (Bridging integrator 1) (Amphiphysin-like protein) (Amphiphysin II) (Box-dependent myc- interacting protein 1). | |||||
|
RP1L1_HUMAN
|
||||||
| NC score | 0.030335 (rank : 107) | θ value | 8.99809 (rank : 153) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
PSL1_HUMAN
|
||||||
| NC score | 0.030182 (rank : 108) | θ value | 8.99809 (rank : 152) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8TCT7, O60365, Q567S3, Q8IUH9, Q9BUY6, Q9H3M4, Q9NPN2, Q9P1Z6 | Gene names | SPPL2B, KIAA1532, PSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal peptide peptidase-like 2B (EC 3.4.23.-) (Protein SPP-like 2B) (Protein SPPL2b) (Intramembrane protease 4) (IMP4) (Presenilin-like protein 1). | |||||
|
ALMS1_HUMAN
|
||||||
| NC score | 0.030093 (rank : 109) | θ value | 3.0926 (rank : 89) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8TCU4, Q53S05, Q580Q8, Q86VP9, Q9Y4G4 | Gene names | ALMS1, KIAA0328 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alstrom syndrome protein 1. | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.029383 (rank : 110) | θ value | 8.99809 (rank : 147) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
H12_HUMAN
|
||||||
| NC score | 0.028709 (rank : 111) | θ value | 3.0926 (rank : 92) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P16403 | Gene names | HIST1H1C, H1F2 | |||
|
Domain Architecture |

|
|||||
| Description | Histone H1.2 (Histone H1d). | |||||
|
SPIT1_HUMAN
|
||||||
| NC score | 0.028195 (rank : 112) | θ value | 2.36792 (rank : 84) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43278, Q7Z7D2 | Gene names | SPINT1, HAI1 | |||
|
Domain Architecture |

|
|||||
| Description | Kunitz-type protease inhibitor 1 precursor (Hepatocyte growth factor activator inhibitor type 1) (HAI-1). | |||||
|
ADA17_MOUSE
|
||||||
| NC score | 0.027903 (rank : 113) | θ value | 8.99809 (rank : 142) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z0F8, O88726, Q9R1U4, Q9Z0K3 | Gene names | Adam17, Tace | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 17 precursor (EC 3.4.24.86) (A disintegrin and metalloproteinase domain 17) (TNF-alpha-converting enzyme) (TNF-alpha convertase) (CD156b antigen). | |||||
|
IF16_HUMAN
|
||||||
| NC score | 0.026937 (rank : 114) | θ value | 4.03905 (rank : 108) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q16666, Q9UH78 | Gene names | IFI16, IFNGIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-interferon-inducible protein Ifi-16 (Interferon-inducible myeloid differentiation transcriptional activator) (IFI 16). | |||||
|
SYN2_MOUSE
|
||||||
| NC score | 0.026771 (rank : 115) | θ value | 5.27518 (rank : 125) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q64332, Q6NZR0, Q9QWV7 | Gene names | Syn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synapsin-2 (Synapsin II). | |||||
|
AMOT_HUMAN
|
||||||
| NC score | 0.026666 (rank : 116) | θ value | 5.27518 (rank : 113) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q4VCS5, Q504X5, Q9HD27, Q9UPT1 | Gene names | AMOT, KIAA1071 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin. | |||||
|
CHD9_MOUSE
|
||||||
| NC score | 0.026256 (rank : 117) | θ value | 0.0736092 (rank : 27) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BYH8, Q7TMV5, Q8BJG8, Q8BZJ2, Q8CHG8 | Gene names | Chd9, Kiaa0308, Pric320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha- interacting complex protein 320 kDa). | |||||
|
FOG1_MOUSE
|
||||||
| NC score | 0.025370 (rank : 118) | θ value | 1.06291 (rank : 57) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O35615 | Gene names | Zfpm1, Fog, Fog1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
HMGN3_HUMAN
|
||||||
| NC score | 0.025177 (rank : 119) | θ value | 6.88961 (rank : 130) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15651, Q7RTT0, Q969M5, Q9BZT7 | Gene names | HMGN3, TRIP7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group nucleosome-binding domain-containing protein 3 (Thyroid receptor-interacting protein 7) (TRIP7). | |||||
|
CTND2_HUMAN
|
||||||
| NC score | 0.024124 (rank : 120) | θ value | 4.03905 (rank : 103) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UQB3, O00379, O15390, O43206, O43840, Q13589, Q9UM66, Q9UPM3 | Gene names | CTNND2, NPRAP | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Delta-catenin) (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin) (GT24). | |||||
|
ABCG1_HUMAN
|
||||||
| NC score | 0.023933 (rank : 121) | θ value | 0.813845 (rank : 49) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P45844, Q86SU8, Q96L76, Q9BXK6, Q9BXK7, Q9BXK8, Q9BXK9, Q9BXL0, Q9BXL1, Q9BXL2, Q9BXL3, Q9BXL4 | Gene names | ABCG1, ABC8, WHT1 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-binding cassette sub-family G member 1 (White protein homolog) (ATP-binding cassette transporter 8). | |||||
|
STRAD_MOUSE
|
||||||
| NC score | 0.023382 (rank : 122) | θ value | 6.88961 (rank : 136) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q60924 | Gene names | Stra13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-inducible E3 protein (Hematopoietic-specific protein E3). | |||||
|
ABCG1_MOUSE
|
||||||
| NC score | 0.021842 (rank : 123) | θ value | 1.81305 (rank : 67) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q64343 | Gene names | Abcg1, Abc8, Wht1 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-binding cassette sub-family G member 1 (White protein homolog) (ATP-binding cassette transporter 8). | |||||
|
CAC1G_HUMAN
|
||||||
| NC score | 0.021780 (rank : 124) | θ value | 8.99809 (rank : 143) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 587 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O43497, O43498, O94770, Q9NYU4, Q9NYU5, Q9NYU6, Q9NYU7, Q9NYU8, Q9NYU9, Q9NYV0, Q9NYV1, Q9UHN9, Q9UHP0, Q9ULU6, Q9UNG7, Q9Y5T2, Q9Y5T3 | Gene names | CACNA1G, KIAA1123 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1G (Voltage- gated calcium channel subunit alpha Cav3.1) (Cav3.1c) (NBR13). | |||||
|
RNZ2_MOUSE
|
||||||
| NC score | 0.021418 (rank : 125) | θ value | 4.03905 (rank : 111) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80Y81, Q99MF0, Q99MF1, Q9CTA2, Q9D1A8, Q9EPZ2 | Gene names | Elac2 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc phosphodiesterase ELAC protein 2 (EC 3.1.26.11) (Ribonuclease Z 2) (RNase Z 2) (tRNase Z 2) (tRNA 3 endonuclease 2) (ElaC homolog protein 2). | |||||
|
FBX42_HUMAN
|
||||||
| NC score | 0.021115 (rank : 126) | θ value | 4.03905 (rank : 106) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6P3S6, Q5TEU8, Q86XI0, Q8N3N4, Q8N5F8, Q9BRM0, Q9P2L4 | Gene names | FBXO42, FBX42, KIAA1332 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 42. | |||||
|
PHF12_MOUSE
|
||||||
| NC score | 0.021066 (rank : 127) | θ value | 5.27518 (rank : 120) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5SPL2, Q5SPL3, Q6ZPN8, Q80W50 | Gene names | Phf12, Kiaa1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
ZEP1_HUMAN
|
||||||
| NC score | 0.021027 (rank : 128) | θ value | 0.279714 (rank : 43) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1036 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P15822, Q14122 | Gene names | HIVEP1, ZNF40 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 40 (Human immunodeficiency virus type I enhancer- binding protein 1) (HIV-EP1) (Major histocompatibility complex-binding protein 1) (MBP-1) (Positive regulatory domain II-binding factor 1) (PRDII-BF1). | |||||
|
ZEP1_MOUSE
|
||||||
| NC score | 0.020816 (rank : 129) | θ value | 8.99809 (rank : 159) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1063 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q03172 | Gene names | Hivep1, Cryabp1, Znf40 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 40 (Transcription factor alphaA-CRYBP1) (Alpha A- crystallin-binding protein I) (Alpha A-CRYBP1). | |||||
|
MEFV_MOUSE
|
||||||
| NC score | 0.020804 (rank : 130) | θ value | 3.0926 (rank : 93) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JJ26 | Gene names | Mefv | |||
|
Domain Architecture |

|
|||||
| Description | Pyrin (Marenostrin). | |||||
|
CCD45_MOUSE
|
||||||
| NC score | 0.020624 (rank : 131) | θ value | 4.03905 (rank : 101) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 663 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BVV7 | Gene names | Ccdc45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 45. | |||||
|
PER3_HUMAN
|
||||||
| NC score | 0.019917 (rank : 132) | θ value | 8.99809 (rank : 149) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
EGLN1_MOUSE
|
||||||
| NC score | 0.019437 (rank : 133) | θ value | 4.03905 (rank : 105) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91YE3, Q8VHJ2, Q922P3 | Gene names | Egln1 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 1 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 2) (HIF-prolyl hydroxylase 2) (HIF-PH2) (HPH-2) (SM-20). | |||||
|
SF3A1_MOUSE
|
||||||
| NC score | 0.018901 (rank : 134) | θ value | 8.99809 (rank : 155) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8K4Z5, Q8C0M7, Q8C128, Q8C175, Q921T3 | Gene names | Sf3a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 3 subunit 1 (SF3a120). | |||||
|
REPS1_MOUSE
|
||||||
| NC score | 0.018041 (rank : 135) | θ value | 5.27518 (rank : 122) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O54916, Q8C9J9, Q99LR8 | Gene names | Reps1 | |||
|
Domain Architecture |

|
|||||
| Description | RalBP1-associated Eps domain-containing protein 1 (RalBP1-interacting protein 1). | |||||
|
NALP1_HUMAN
|
||||||
| NC score | 0.017054 (rank : 136) | θ value | 1.38821 (rank : 63) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9C000, Q9BZZ8, Q9BZZ9, Q9HAV8, Q9UFT4, Q9Y2E0 | Gene names | NALP1, CARD7, DEFCAP, KIAA0926, NAC | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 1 (Death effector filament- forming ced-4-like apoptosis protein) (Nucleotide-binding domain and caspase recruitment domain) (Caspase recruitment domain protein 7). | |||||
|
UBXD2_HUMAN
|
||||||
| NC score | 0.016960 (rank : 137) | θ value | 5.27518 (rank : 128) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92575, Q8IYM5 | Gene names | UBXD2, KIAA0242, UBXDC1 | |||
|
Domain Architecture |

|
|||||
| Description | UBX domain-containing protein 2. | |||||
|
3BP1_MOUSE
|
||||||
| NC score | 0.016796 (rank : 138) | θ value | 8.99809 (rank : 141) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P55194, Q99KK8 | Gene names | Sh3bp1, 3bp1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain-binding protein 1 (3BP-1). | |||||
|
RFWD2_HUMAN
|
||||||
| NC score | 0.016238 (rank : 139) | θ value | 5.27518 (rank : 123) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NHY2, Q6H103, Q9H6L7 | Gene names | RFWD2, COP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger and WD repeat domain protein 2 (EC 6.3.2.-) (Ubiquitin- protein ligase COP1) (Constitutive photomorphogenesis protein 1 homolog) (hCOP1). | |||||
|
PDLI5_HUMAN
|
||||||
| NC score | 0.016109 (rank : 140) | θ value | 1.81305 (rank : 75) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 322 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96HC4, O60705 | Gene names | PDLIM5, ENH | |||
|
Domain Architecture |

|
|||||
| Description | PDZ and LIM domain protein 5 (Enigma homolog) (Enigma-like PDZ and LIM domains protein). | |||||
|
DBPA_MOUSE
|
||||||
| NC score | 0.015900 (rank : 141) | θ value | 8.99809 (rank : 144) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9JKB3, Q80WG4, Q9EQF7, Q9EQF8 | Gene names | Csda, Msy4, Ybx3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA-binding protein A (Cold shock domain-containing protein A) (Y-box protein 3). | |||||
|
INVS_MOUSE
|
||||||
| NC score | 0.015812 (rank : 142) | θ value | 1.81305 (rank : 72) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O89019, O88849 | Gene names | Invs, Inv, Nphp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning protein) (Nephrocystin-2). | |||||
|
PHF2_MOUSE
|
||||||
| NC score | 0.015077 (rank : 143) | θ value | 8.99809 (rank : 150) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
TRI31_HUMAN
|
||||||
| NC score | 0.014274 (rank : 144) | θ value | 5.27518 (rank : 127) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 299 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9BZY9, Q53H52, Q5RI37, Q5SRJ7, Q5SRJ8, Q5SS28, Q96AK4, Q96AP8, Q99579, Q9BZY8 | Gene names | TRIM31, C6orf13, RNF | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 31. | |||||
|
GRN_MOUSE
|
||||||
| NC score | 0.014197 (rank : 145) | θ value | 4.03905 (rank : 107) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P28798 | Gene names | Grn | |||
|
Domain Architecture |

|
|||||
| Description | Granulins precursor (Proepithelin) (PEPI) (PC cell-derived growth factor) (PCDGF) [Contains: Acrogranin; Granulin-1; Granulin-2; Granulin-3; Granulin-4; Granulin-5; Granulin-6; Granulin-7]. | |||||
|
SPHK2_HUMAN
|
||||||
| NC score | 0.013965 (rank : 146) | θ value | 6.88961 (rank : 135) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NRA0, Q9BRN1, Q9H0Q2, Q9NWU7 | Gene names | SPHK2 | |||
|
Domain Architecture |

|
|||||
| Description | Sphingosine kinase 2 (EC 2.7.1.-) (SK 2) (SPK 2). | |||||
|
SPEG_MOUSE
|
||||||
| NC score | 0.013867 (rank : 147) | θ value | 0.279714 (rank : 41) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
CAC1A_MOUSE
|
||||||
| NC score | 0.013120 (rank : 148) | θ value | 2.36792 (rank : 81) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P97445 | Gene names | Cacna1a, Cach4, Cacn3, Cacnl1a4, Ccha1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
SREC2_MOUSE
|
||||||
| NC score | 0.012538 (rank : 149) | θ value | 4.03905 (rank : 112) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 605 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P59222 | Gene names | Scarf2, Srec2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor class F member 2 precursor (Scavenger receptor expressed by endothelial cells 2 protein) (SREC-II). | |||||
|
NEST_HUMAN
|
||||||
| NC score | 0.012503 (rank : 150) | θ value | 5.27518 (rank : 119) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1305 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P48681, O00552 | Gene names | NES | |||
|
Domain Architecture |

|
|||||
| Description | Nestin. | |||||
|
ZN687_HUMAN
|
||||||
| NC score | 0.012092 (rank : 151) | θ value | 3.0926 (rank : 97) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8N1G0, Q68DQ8, Q9H937, Q9P2A7 | Gene names | ZNF687, KIAA1441 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 687. | |||||
|
MAGD2_HUMAN
|
||||||
| NC score | 0.011579 (rank : 152) | θ value | 4.03905 (rank : 110) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UNF1, O76058, Q5BJF3, Q8NAL6, Q9H218, Q9P0U9, Q9UM52 | Gene names | MAGED2, BCG1 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen D2 (MAGE-D2 antigen) (MAGE-D) (Breast cancer-associated gene 1 protein) (BCG-1) (11B6) (Hepatocellular carcinoma-associated protein JCL-1). | |||||
|
DPOLL_MOUSE
|
||||||
| NC score | 0.011519 (rank : 153) | θ value | 8.99809 (rank : 145) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9QXE2, Q9CTJ1 | Gene names | Poll, Polk | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase lambda (EC 2.7.7.7) (EC 4.2.99.-) (Pol Lambda) (DNA polymerase kappa). | |||||
|
CYTSA_HUMAN
|
||||||
| NC score | 0.011390 (rank : 154) | θ value | 4.03905 (rank : 104) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1138 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q69YQ0, O15081 | Gene names | CYTSA, KIAA0376 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytospin-A (NY-REN-22 antigen). | |||||
|
SYBU_MOUSE
|
||||||
| NC score | 0.010249 (rank : 155) | θ value | 8.99809 (rank : 156) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BHS8, Q3TQN7, Q401F3, Q401F4, Q571C1, Q6P1J2, Q80XH0, Q8BHS7, Q8BI27 | Gene names | Sybu, Golsyn, Kiaa1472 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntabulin (Syntaxin-1-binding protein) (Golgi-localized syntaphilin- related protein) (m-Golsyn). | |||||
|
ADA15_HUMAN
|
||||||
| NC score | 0.010142 (rank : 156) | θ value | 1.81305 (rank : 68) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q13444, Q13493, Q96C78 | Gene names | ADAM15, MDC15 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 15 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 15) (Metalloproteinase-like, disintegrin-like, and cysteine- rich protein 15) (MDC-15) (Metalloprotease RGD disintegrin protein) (Metargidin). | |||||
|
ZN438_HUMAN
|
||||||
| NC score | 0.009573 (rank : 157) | θ value | 2.36792 (rank : 87) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q7Z4V0, Q5T426, Q658Q4, Q6ZN65 | Gene names | ZNF438 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 438. | |||||
|
BTBD9_MOUSE
|
||||||
| NC score | 0.008941 (rank : 158) | θ value | 4.03905 (rank : 100) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C726 | Gene names | Btbd9 | |||
|
Domain Architecture |

|
|||||
| Description | BTB/POZ domain-containing protein 9. | |||||
|
KCNB2_HUMAN
|
||||||
| NC score | 0.007478 (rank : 159) | θ value | 4.03905 (rank : 109) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q92953, Q9BXD3 | Gene names | KCNB2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 2 (Voltage-gated potassium channel subunit Kv2.2). | |||||
|
TEP1_MOUSE
|
||||||
| NC score | 0.007295 (rank : 160) | θ value | 5.27518 (rank : 126) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P97499 | Gene names | Tep1, Tp1 | |||
|
Domain Architecture |

|
|||||
| Description | Telomerase protein component 1 (Telomerase-associated protein 1) (Telomerase protein 1) (p240) (p80 homolog). | |||||
|
CAMKV_HUMAN
|
||||||
| NC score | 0.006242 (rank : 161) | θ value | 1.81305 (rank : 69) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1084 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8NCB2, Q6FIB8, Q8NBS8, Q8NC85, Q8NDU4, Q8WTT8, Q9BQC9, Q9H0Q5 | Gene names | CAMKV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CaM kinase-like vesicle-associated protein. | |||||
|
SALL2_MOUSE
|
||||||
| NC score | 0.006019 (rank : 162) | θ value | 6.88961 (rank : 134) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9QX96 | Gene names | Sall2, Sal2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 2 (Spalt-like protein 2) (MSal-2). | |||||
|
ZN297_MOUSE
|
||||||
| NC score | 0.005918 (rank : 163) | θ value | 6.88961 (rank : 138) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 816 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Z0G7 | Gene names | Znf297, Bing1, Zfp297 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 297 (BING1 protein). | |||||
|
ZN217_HUMAN
|
||||||
| NC score | 0.004821 (rank : 164) | θ value | 6.88961 (rank : 137) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 905 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O75362 | Gene names | ZNF217, ZABC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 217. | |||||
|
EVX1_MOUSE
|
||||||
| NC score | 0.003910 (rank : 165) | θ value | 5.27518 (rank : 115) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P23683 | Gene names | Evx1 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox even-skipped homolog protein 1 (EVX-1). | |||||
|
M3K14_HUMAN
|
||||||
| NC score | 0.002056 (rank : 166) | θ value | 6.88961 (rank : 132) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 887 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q99558, Q8IYN1 | Gene names | MAP3K14, NIK | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 14 (EC 2.7.11.25) (NF- kappa beta-inducing kinase) (Serine/threonine-protein kinase NIK) (HsNIK). | |||||
|
M3K1_MOUSE
|
||||||
| NC score | 0.001727 (rank : 167) | θ value | 8.99809 (rank : 146) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P53349, Q60831, Q9R0U3, Q9R256 | Gene names | Map3k1, Mekk, Mekk1 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 1 (EC 2.7.11.25) (MAPK/ERK kinase kinase 1) (MEK kinase 1) (MEKK 1). | |||||
|
TNIK_HUMAN
|
||||||
| NC score | 0.001237 (rank : 168) | θ value | 8.99809 (rank : 157) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 1570 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UKE5, O60298, Q8WUY7, Q9UKD8, Q9UKD9, Q9UKE0, Q9UKE1, Q9UKE2, Q9UKE3, Q9UKE4 | Gene names | TNIK, KIAA0551 | |||
|
Domain Architecture |

|
|||||
| Description | TRAF2 and NCK-interacting protein kinase (EC 2.7.11.1). | |||||
|
GFI1_MOUSE
|
||||||
| NC score | 0.001020 (rank : 169) | θ value | 3.0926 (rank : 91) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 736 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P70338, O09063 | Gene names | Gfi1, Gfi-1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein Gfi-1 (Growth factor independence 1). | |||||
|
KKCC1_MOUSE
|
||||||
| NC score | -0.000280 (rank : 170) | θ value | 6.88961 (rank : 131) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 866 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8VBY2, Q9R054 | Gene names | Camkk1, Camkk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium/calmodulin-dependent protein kinase kinase 1 (EC 2.7.11.17) (Calcium/calmodulin-dependent protein kinase kinase alpha) (CaM-kinase kinase alpha) (CaM-KK alpha) (CaMKK alpha) (CaMKK 1) (CaM-kinase IV kinase). | |||||
|
ZN557_HUMAN
|
||||||
| NC score | -0.001022 (rank : 171) | θ value | 6.88961 (rank : 139) | |||
| Query Neighborhood Hits | 159 | Target Neighborhood Hits | 752 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8N988, Q6PEJ3, Q9BTZ1 | Gene names | ZNF557 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 557. | |||||