Please be patient as the page loads
|
SMAD4_MOUSE
|
||||||
| SwissProt Accessions | P97471, Q9CW56 | Gene names | Smad4, Dpc4, Madh4 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 4 (SMAD 4) (Mothers against DPP homolog 4) (Deletion target in pancreatic carcinoma 4 homolog) (Smad4). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SMAD4_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.985217 (rank : 2) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q13485 | Gene names | SMAD4, DPC4, MADH4 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 4 (SMAD 4) (Mothers against DPP homolog 4) (Deletion target in pancreatic carcinoma 4) (hSMAD4). | |||||
|
SMAD4_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 101 | |
| SwissProt Accessions | P97471, Q9CW56 | Gene names | Smad4, Dpc4, Madh4 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 4 (SMAD 4) (Mothers against DPP homolog 4) (Deletion target in pancreatic carcinoma 4 homolog) (Smad4). | |||||
|
SMAD9_HUMAN
|
||||||
| θ value | 1.00825e-67 (rank : 3) | NC score | 0.925348 (rank : 5) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O15198, O14989 | Gene names | SMAD9, MADH6, MADH9 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 9 (SMAD 9) (Mothers against DPP homolog 9) (Smad9) (Madh6). | |||||
|
SMAD5_MOUSE
|
||||||
| θ value | 1.71981e-67 (rank : 4) | NC score | 0.923047 (rank : 7) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P97454, P70341, Q810K0 | Gene names | Smad5, Madh5, Msmad5 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 5 (SMAD 5) (Mothers against DPP homolog 5) (Smad5) (mSmad5) (Dwarfin-C) (Dwf-C). | |||||
|
SMAD3_HUMAN
|
||||||
| θ value | 2.24614e-67 (rank : 5) | NC score | 0.926398 (rank : 3) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P84022, O09064, O09144, O14510, O35273, Q92940, Q93002, Q9GKR4 | Gene names | SMAD3, MADH3 | |||
|
Domain Architecture |
|
|||||
| Description | Mothers against decapentaplegic homolog 3 (SMAD 3) (Mothers against DPP homolog 3) (Mad3) (hMAD-3) (JV15-2) (hSMAD3). | |||||
|
SMAD3_MOUSE
|
||||||
| θ value | 2.24614e-67 (rank : 6) | NC score | 0.926398 (rank : 4) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BUN5, O09064, O09144, O14510, O35273, Q8BX84, Q92940, Q93002, Q9GKR4 | Gene names | Smad3, Madh3 | |||
|
Domain Architecture |
|
|||||
| Description | Mothers against decapentaplegic homolog 3 (SMAD 3) (Mothers against DPP homolog 3) (Mad3) (mMad3). | |||||
|
SMAD5_HUMAN
|
||||||
| θ value | 2.4834e-66 (rank : 7) | NC score | 0.923893 (rank : 6) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q99717, O14688, Q15798, Q9UQA1 | Gene names | SMAD5, MADH5 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 5 (SMAD 5) (Mothers against DPP homolog 5) (Smad5) (hSmad5) (JV5-1). | |||||
|
SMAD1_HUMAN
|
||||||
| θ value | 4.23606e-66 (rank : 8) | NC score | 0.922510 (rank : 8) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q15797, Q16636 | Gene names | SMAD1, BSP1, MADH1, MADR1 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 1 (SMAD 1) (Mothers against DPP homolog 1) (Mad-related protein 1) (Transforming growth factor- beta signaling protein 1) (BSP-1) (hSMAD1) (JV4-1). | |||||
|
SMAD1_MOUSE
|
||||||
| θ value | 1.36269e-64 (rank : 9) | NC score | 0.922113 (rank : 9) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P70340, P70442, Q9CYK6 | Gene names | Smad1, Madh1, Madr1 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 1 (SMAD 1) (Mothers against DPP homolog 1) (Mad-related protein 1) (Mothers-against-DPP-related 1) (mMad1) (Dwarfin-A) (Dwf-A). | |||||
|
SMAD9_MOUSE
|
||||||
| θ value | 4.38363e-63 (rank : 10) | NC score | 0.914955 (rank : 10) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9JIW5, Q8CFF9 | Gene names | Smad9, Madh8, Madh9, Smad8 | |||
|
Domain Architecture |
|
|||||
| Description | Mothers against decapentaplegic homolog 9 (SMAD 9) (Mothers against DPP homolog 9) (Smad9) (Smad8). | |||||
|
SMAD2_HUMAN
|
||||||
| θ value | 1.2411e-41 (rank : 11) | NC score | 0.906736 (rank : 12) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15796 | Gene names | SMAD2, MADH2, MADR2 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 2 (SMAD 2) (Mothers against DPP homolog 2) (Mad-related protein 2) (hMAD-2) (JV18-1) (hSMAD2). | |||||
|
SMAD2_MOUSE
|
||||||
| θ value | 1.2411e-41 (rank : 12) | NC score | 0.906762 (rank : 11) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q62432, Q9D8P6 | Gene names | Smad2, Madh2, Madr2 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 2 (SMAD 2) (Mothers against DPP homolog 2) (Mad-related protein 2) (mMad2). | |||||
|
SMAD6_HUMAN
|
||||||
| θ value | 1.2077e-20 (rank : 13) | NC score | 0.777164 (rank : 14) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O43541, O43654, Q15799, Q9UKZ3 | Gene names | SMAD6, MADH6 | |||
|
Domain Architecture |
|
|||||
| Description | Mothers against decapentaplegic homolog 6 (SMAD 6) (Mothers against DPP homolog 6) (Smad6) (hSMAD6). | |||||
|
SMAD6_MOUSE
|
||||||
| θ value | 2.06002e-20 (rank : 14) | NC score | 0.778363 (rank : 13) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O35182, Q9CW62 | Gene names | Smad6, Madh6, Madh7, Msmad6 | |||
|
Domain Architecture |
|
|||||
| Description | Mothers against decapentaplegic homolog 6 (SMAD 6) (Mothers against DPP homolog 6) (Smad6) (Mad homolog 7). | |||||
|
SMAD7_HUMAN
|
||||||
| θ value | 2.7842e-17 (rank : 15) | NC score | 0.761781 (rank : 16) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O15105, O14740, Q6DK23 | Gene names | SMAD7, MADH7, MADH8 | |||
|
Domain Architecture |
|
|||||
| Description | Mothers against decapentaplegic homolog 7 (SMAD 7) (Mothers against DPP homolog 7) (Smad7) (hSMAD7). | |||||
|
SMAD7_MOUSE
|
||||||
| θ value | 2.7842e-17 (rank : 16) | NC score | 0.762637 (rank : 15) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O35253, O88709 | Gene names | Smad7, Madh7 | |||
|
Domain Architecture |
|
|||||
| Description | Mothers against decapentaplegic homolog 7 (SMAD 7) (Mothers against DPP homolog 7) (Smad7) (Madh8). | |||||
|
FOXJ3_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 17) | NC score | 0.044533 (rank : 28) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UPW0, Q9NSS7 | Gene names | FOXJ3, KIAA1041 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein J3. | |||||
|
FOXJ3_MOUSE
|
||||||
| θ value | 0.00102713 (rank : 18) | NC score | 0.039514 (rank : 30) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BUR3, Q3UTD8 | Gene names | Foxj3, Kiaa1041 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein J3. | |||||
|
PORIM_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 19) | NC score | 0.096395 (rank : 17) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N131, Q8IWS2, Q96QV2 | Gene names | TMEM123, KCT3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Porimin precursor (Transmembrane protein 123) (Pro-oncosis receptor inducing membrane injury) (Keratinocytes-associated transmembrane protein 3) (KCT-3). | |||||
|
PO3F3_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 20) | NC score | 0.030866 (rank : 39) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P31361 | Gene names | Pou3f3, Brn-1, Brn1, Otf8 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 3, transcription factor 3 (Brain-specific homeobox/POU domain protein 1) (Brain-1) (Brn-1 protein). | |||||
|
PO3F3_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 21) | NC score | 0.030227 (rank : 40) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P20264, P78379 | Gene names | POU3F3, BRN1, OTF8 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 3, transcription factor 3 (Brain-specific homeobox/POU domain protein 1) (Brain-1) (Brn-1 protein). | |||||
|
BORG5_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 22) | NC score | 0.057159 (rank : 19) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q00587, Q96GN1 | Gene names | CDC42EP1, BORG5, CEP1, MSE55 | |||
|
Domain Architecture |
|
|||||
| Description | Cdc42 effector protein 1 (Binder of Rho GTPases 5) (Serum protein MSE55). | |||||
|
ARI1A_HUMAN
|
||||||
| θ value | 0.163984 (rank : 23) | NC score | 0.030946 (rank : 35) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
CREB5_HUMAN
|
||||||
| θ value | 0.163984 (rank : 24) | NC score | 0.030869 (rank : 38) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q02930, Q05886, Q06246 | Gene names | CREB5, CREBPA | |||
|
Domain Architecture |

|
|||||
| Description | cAMP response element-binding protein 5 (CRE-BPa). | |||||
|
CXXC6_HUMAN
|
||||||
| θ value | 0.163984 (rank : 25) | NC score | 0.044894 (rank : 27) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
MAP4_MOUSE
|
||||||
| θ value | 0.163984 (rank : 26) | NC score | 0.039682 (rank : 29) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P27546 | Gene names | Map4, Mtap4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
SF01_HUMAN
|
||||||
| θ value | 0.163984 (rank : 27) | NC score | 0.046067 (rank : 26) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15637, Q14818, Q14819, Q15913, Q8IY00, Q92744, Q92745, Q969H7, Q9BW01, Q9UEI0 | Gene names | SF1, ZFM1, ZNF162 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor 1 (Zinc finger protein 162) (Transcription factor ZFM1) (Zinc finger gene in MEN1 locus) (Mammalian branch point-binding protein mBBP) (BBP). | |||||
|
SF01_MOUSE
|
||||||
| θ value | 0.163984 (rank : 28) | NC score | 0.046383 (rank : 25) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q64213, O08817, P70167, Q61454, Q921Z4 | Gene names | Sf1, Zfm1, Zfp162 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor 1 (Zinc finger protein 162) (Transcription factor ZFM1) (mZFM) (Zinc finger gene in MEN1 locus) (Mammalian branch point- binding protein mBBP) (BBP) (CW17). | |||||
|
IF4G1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 29) | NC score | 0.031006 (rank : 34) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q04637, O43177, O95066, Q5HYG0, Q6ZN21, Q8N102 | Gene names | EIF4G1, EIF4G, EIF4GI | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1) (p220). | |||||
|
BRPF1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 30) | NC score | 0.016988 (rank : 57) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P55201, Q9UHI0 | Gene names | BRPF1, BR140 | |||
|
Domain Architecture |

|
|||||
| Description | Peregrin (Bromodomain and PHD finger-containing protein 1) (BR140 protein). | |||||
|
CSPG3_HUMAN
|
||||||
| θ value | 0.279714 (rank : 31) | NC score | 0.017006 (rank : 56) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O14594, Q9UPK6 | Gene names | CSPG3, NCAN, NEUR | |||
|
Domain Architecture |

|
|||||
| Description | Neurocan core protein precursor (Chondroitin sulfate proteoglycan 3). | |||||
|
R3HD2_MOUSE
|
||||||
| θ value | 0.279714 (rank : 32) | NC score | 0.034548 (rank : 32) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
ABI1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 33) | NC score | 0.016806 (rank : 59) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8IZP0, O15147, O76049, O95060, Q5W070, Q5W072, Q8TB63, Q96S81, Q9NXZ9, Q9NYB8 | Gene names | ABI1, SSH3BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Abl interactor 1 (Abelson interactor 1) (Abi-1) (Spectrin SH3 domain- binding protein 1) (Eps8 SH3 domain-binding protein) (Eps8-binding protein) (e3B1) (Nap1-binding protein) (Nap1BP) (Abl-binding protein 4) (AblBP4). | |||||
|
ATX2L_HUMAN
|
||||||
| θ value | 0.47712 (rank : 34) | NC score | 0.052725 (rank : 20) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8WWM7, O95135, Q6NVJ8, Q6PJW6, Q8IU61, Q8IU95, Q8WWM3, Q8WWM4, Q8WWM5, Q8WWM6, Q99703 | Gene names | ATXN2L, A2D, A2LG, A2LP, A2RP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2-like protein (Ataxin-2 domain protein) (Ataxin-2-related protein). | |||||
|
ONEC2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 35) | NC score | 0.019896 (rank : 54) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6XBJ3 | Gene names | Onecut2, Oc2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | One cut domain family member 2 (Transcription factor ONECUT-2) (OC-2). | |||||
|
ATX2L_MOUSE
|
||||||
| θ value | 0.813845 (rank : 36) | NC score | 0.051267 (rank : 22) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q7TQH0, Q80XN9, Q8K059 | Gene names | Atxn2l, A2lp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2-like protein. | |||||
|
CD68_HUMAN
|
||||||
| θ value | 0.813845 (rank : 37) | NC score | 0.030883 (rank : 37) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P34810, Q96BI7 | Gene names | CD68 | |||
|
Domain Architecture |
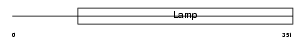
|
|||||
| Description | Macrosialin precursor (GP110) (CD68 antigen). | |||||
|
MAFB_MOUSE
|
||||||
| θ value | 0.813845 (rank : 38) | NC score | 0.018405 (rank : 55) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P54841 | Gene names | Mafb, Krml, Maf1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor MafB (V-maf musculoaponeurotic fibrosarcoma oncogene homolog B) (Transcription factor MAF1) (Segmentation protein KR) (Kreisler). | |||||
|
MUC2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 39) | NC score | 0.022938 (rank : 47) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q02817, Q14878 | Gene names | MUC2, SMUC | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-2 precursor (Intestinal mucin-2). | |||||
|
PODXL_MOUSE
|
||||||
| θ value | 0.813845 (rank : 40) | NC score | 0.047773 (rank : 24) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R0M4, Q9ESZ1 | Gene names | Podxl, Pclp1 | |||
|
Domain Architecture |

|
|||||
| Description | Podocalyxin-like protein 1 precursor. | |||||
|
WASIP_HUMAN
|
||||||
| θ value | 0.813845 (rank : 41) | NC score | 0.021382 (rank : 48) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
MUC7_HUMAN
|
||||||
| θ value | 1.06291 (rank : 42) | NC score | 0.024920 (rank : 45) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
SHAN1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 43) | NC score | 0.008116 (rank : 86) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y566, Q9NYW9 | Gene names | SHANK1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 1 (Shank1) (Somatostatin receptor-interacting protein) (SSTR-interacting protein) (SSTRIP). | |||||
|
TFG_HUMAN
|
||||||
| θ value | 1.06291 (rank : 44) | NC score | 0.038524 (rank : 31) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92734, Q15656 | Gene names | TFG | |||
|
Domain Architecture |

|
|||||
| Description | Protein TFG (TRK-fused gene protein). | |||||
|
NCOA6_MOUSE
|
||||||
| θ value | 1.38821 (rank : 45) | NC score | 0.052407 (rank : 21) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
NEDD4_MOUSE
|
||||||
| θ value | 1.38821 (rank : 46) | NC score | 0.007415 (rank : 89) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P46935, O08758, Q3UZI2, Q8BGB3 | Gene names | Nedd4, Kiaa0093, Nedd-4, Nedd4a | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-) (Neural precursor cell expressed developmentally down-regulated protein 4). | |||||
|
GLIS3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 47) | NC score | -0.000887 (rank : 99) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 758 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NEA6 | Gene names | GLIS3, ZNF515 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLIS3 (GLI-similar 3) (Zinc finger protein 515). | |||||
|
MUC1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 48) | NC score | 0.024717 (rank : 46) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
RERE_MOUSE
|
||||||
| θ value | 1.81305 (rank : 49) | NC score | 0.025740 (rank : 44) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
ASPP2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.009206 (rank : 84) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q13625, Q12892, Q86X75, Q96KQ3 | Gene names | TP53BP2, ASPP2, BBP | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-stimulating of p53 protein 2 (Tumor suppressor p53-binding protein 2) (p53-binding protein 2) (p53BP2) (53BP2) (Bcl2-binding protein) (Bbp) (NY-REN-51 antigen). | |||||
|
FIGN_HUMAN
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.010294 (rank : 75) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5HY92, Q9H6M5, Q9NVZ9 | Gene names | FIGN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fidgetin. | |||||
|
LRC41_HUMAN
|
||||||
| θ value | 2.36792 (rank : 52) | NC score | 0.028439 (rank : 43) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15345, Q71RA8, Q9BSM0 | Gene names | LRRC41, MUF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 41 (Protein Muf1). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 2.36792 (rank : 53) | NC score | 0.028850 (rank : 41) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
WIRE_MOUSE
|
||||||
| θ value | 2.36792 (rank : 54) | NC score | 0.020984 (rank : 51) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6PEV3 | Gene names | Wire | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein). | |||||
|
BRAC_HUMAN
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.007596 (rank : 88) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15178 | Gene names | T | |||
|
Domain Architecture |
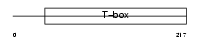
|
|||||
| Description | Brachyury protein (T protein). | |||||
|
EYA1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.011577 (rank : 70) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99502 | Gene names | EYA1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
|
GLP_MOUSE
|
||||||
| θ value | 3.0926 (rank : 57) | NC score | 0.049426 (rank : 23) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P14220 | Gene names | Gypa | |||
|
Domain Architecture |

|
|||||
| Description | Glycophorin. | |||||
|
MAP4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.034095 (rank : 33) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 428 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P27816, Q13082, Q96A76 | Gene names | MAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
MARK2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | -0.001788 (rank : 101) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1056 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7KZI7, Q15449, Q15524, Q5XGA3, Q68A18, Q96HB3, Q96RG0 | Gene names | MARK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL motif kinase) (EMK1) (PAR1 homolog). | |||||
|
RERE_HUMAN
|
||||||
| θ value | 3.0926 (rank : 60) | NC score | 0.021280 (rank : 49) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
ATN1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 61) | NC score | 0.028496 (rank : 42) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
IASPP_HUMAN
|
||||||
| θ value | 4.03905 (rank : 62) | NC score | 0.009796 (rank : 77) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8WUF5, Q2PNZ9, Q5DU71, Q5I1X4, Q6P1R7, Q6PKF8, Q9Y290 | Gene names | PPP1R13L, IASPP, NKIP1, PPP1R13BL, RAI | |||
|
Domain Architecture |
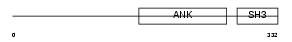
|
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
MUC5B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.016826 (rank : 58) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1020 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9HC84, O00447, O00573, O14985, O15494, O95291, O95451, Q14881, Q99552, Q9UE28 | Gene names | MUC5B, MUC5 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-5B precursor (Mucin-5 subtype B, tracheobronchial) (High molecular weight salivary mucin MG1) (Sublingual gland mucin). | |||||
|
ROBO1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.001184 (rank : 98) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 529 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y6N7, Q7Z300, Q9BUS7 | Gene names | ROBO1, DUTT1 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 1 precursor (H-Robo-1) (Deleted in U twenty twenty). | |||||
|
BPTF_HUMAN
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.012102 (rank : 68) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
CR001_HUMAN
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.009630 (rank : 79) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15165, O15166, O15167, O15168, Q6NXP3 | Gene names | C18orf1 | |||
|
Domain Architecture |
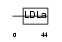
|
|||||
| Description | Protein C18orf1. | |||||
|
DLG2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.004923 (rank : 93) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q15700, Q59G57, Q68CQ8, Q6ZTA8 | Gene names | DLG2 | |||
|
Domain Architecture |

|
|||||
| Description | Discs large homolog 2 (Postsynaptic density protein PSD-93) (Channel- associated protein of synapse-110) (Chapsyn-110). | |||||
|
NUMBL_MOUSE
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.009350 (rank : 81) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O08919, Q6NVG8 | Gene names | Numbl, Nbl | |||
|
Domain Architecture |
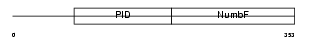
|
|||||
| Description | Numb-like protein. | |||||
|
PF21B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.013756 (rank : 66) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8C966 | Gene names | Phf21b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
PKHA4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 70) | NC score | 0.009644 (rank : 78) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H4M7, Q8N4M8, Q8N658 | Gene names | PLEKHA4, PEPP1 | |||
|
Domain Architecture |
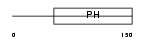
|
|||||
| Description | Pleckstrin homology domain-containing family A member 4 (Phosphoinositol 3-phosphate-binding protein 1) (PEPP-1). | |||||
|
PO3F2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 71) | NC score | 0.020496 (rank : 52) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P20265, Q14960, Q86V54, Q9UJL0 | Gene names | POU3F2, BRN2, OCT7, OTF7 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 3, transcription factor 2 (Nervous system-specific octamer-binding transcription factor N-Oct-3) (Brain-specific homeobox/POU domain protein 2) (Brain-2) (Protein Brn-2). | |||||
|
PO3F2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 72) | NC score | 0.020479 (rank : 53) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P31360 | Gene names | Pou3f2, Brn-2, Brn2, Otf7 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 3, transcription factor 2 (Nervous system-specific octamer-binding transcription factor N-Oct-3) (Brain-specific homeobox/POU domain protein 2) (Brain-2) (Protein Brn-2). | |||||
|
REXO1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 73) | NC score | 0.010356 (rank : 74) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N1G1, Q9ULT2 | Gene names | REXO1, ELOABP1, KIAA1138, TCEB3BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA exonuclease 1 homolog (EC 3.1.-.-) (Elongin A-binding protein 1) (EloA-BP1) (Transcription elongation factor B polypeptide 3-binding protein 1). | |||||
|
VGLL2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 74) | NC score | 0.015109 (rank : 63) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BGW8, Q8BWB4 | Gene names | Vgll2, Vgl2, Vito1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription cofactor vestigial-like protein 2 (Vgl-2) (Protein VITO1). | |||||
|
YAP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 75) | NC score | 0.009919 (rank : 76) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P46938, Q52KJ5 | Gene names | Yap1, Yap, Yap65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
ADRM1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.030915 (rank : 36) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q16186, Q9H1P2 | Gene names | ADRM1, GP110 | |||
|
Domain Architecture |
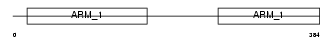
|
|||||
| Description | Adhesion-regulating molecule 1 precursor (110 kDa cell membrane glycoprotein) (Gp110). | |||||
|
ALMS1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.013497 (rank : 67) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8TCU4, Q53S05, Q580Q8, Q86VP9, Q9Y4G4 | Gene names | ALMS1, KIAA0328 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alstrom syndrome protein 1. | |||||
|
CBP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.015773 (rank : 62) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
DDEF2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.004010 (rank : 95) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 625 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7SIG6, Q501K1, Q66JN2 | Gene names | Ddef2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Development and differentiation-enhancing factor 2 (Pyk2 C-terminus- associated protein) (PAP) (Paxillin-associated protein with ARFGAP activity 3) (PAG3). | |||||
|
IQEC2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.014266 (rank : 64) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5JU85, O60275 | Gene names | IQSEC2, KIAA0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
IQEC2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.014212 (rank : 65) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5DU25 | Gene names | Iqsec2, Kiaa0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
LAMP3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.015860 (rank : 61) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UQV4, O94781, Q8NEC8 | Gene names | LAMP3, DCLAMP, TSC403 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lysosome-associated membrane glycoprotein 3 precursor (LAMP-3) (Lysosomal-associated membrane protein 3) (DC-lysosome-associated membrane glycoprotein) (DC LAMP) (Protein TSC403) (CD208 antigen). | |||||
|
MEGF9_HUMAN
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.003880 (rank : 96) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9H1U4, O75098 | Gene names | MEGF9, EGFL5, KIAA0818 | |||
|
Domain Architecture |
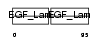
|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
MTG8_HUMAN
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.011555 (rank : 72) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q06455, Q06456, Q14873, Q16239, Q16346, Q16347, Q92479, Q9BRZ0 | Gene names | RUNX1T1, AML1T1, CBFA2T1, CDR, ETO, MTG8, ZMYND2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein CBFA2T1 (Protein MTG8) (Protein ETO) (Eight twenty one protein) (Cyclin-D-related protein) (Zinc finger MYND domain- containing protein 2). | |||||
|
MTG8_MOUSE
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.011562 (rank : 71) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61909 | Gene names | Runx1t1, Cbfa2t1, Cbfa2t1h, Mtg8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein CBFA2T1 (Protein MTG8). | |||||
|
NPAS4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.020999 (rank : 50) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
OTX1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | 0.004101 (rank : 94) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P80205 | Gene names | Otx1, Otx-1 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein OTX1. | |||||
|
PA24B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 88) | NC score | 0.009250 (rank : 83) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95712, Q59GF9, Q8TB10, Q9UKV7 | Gene names | PLA2G4B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytosolic phospholipase A2 beta (EC 3.1.1.4) (cPLA2-beta) (Phospholipase A2 group IVB). | |||||
|
RCOR3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 89) | NC score | 0.011016 (rank : 73) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9P2K3, Q5VT47, Q7L9I5, Q8N5U3, Q9NV83 | Gene names | RCOR3, KIAA1343 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
TBX21_MOUSE
|
||||||
| θ value | 6.88961 (rank : 90) | NC score | 0.005528 (rank : 91) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JKD8, Q9R0A6 | Gene names | Tbx21, Tbet, Tblym | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-box transcription factor TBX21 (T-box protein 21) (Transcription factor TBLYM) (T-cell-specific T-box transcription factor T-bet). | |||||
|
CJ047_MOUSE
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.016382 (rank : 60) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8C5R2, Q8R017, Q8R2Z9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf47 homolog. | |||||
|
CRF_HUMAN
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.011930 (rank : 69) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P06850 | Gene names | CRH | |||
|
Domain Architecture |

|
|||||
| Description | Corticoliberin precursor (Corticotropin-releasing factor) (CRF) (Corticotropin-releasing hormone). | |||||
|
EYA1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 93) | NC score | 0.009431 (rank : 80) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P97767, O08818 | Gene names | Eya1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
|
FIGN_MOUSE
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.007941 (rank : 87) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9ERZ6, Q3TPB0, Q3UP57, Q6PCM0 | Gene names | Fign | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fidgetin. | |||||
|
GASP2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 95) | NC score | 0.005455 (rank : 92) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96D09, Q8NAB4 | Gene names | GPRASP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor-associated sorting protein 2 (GASP-2). | |||||
|
HCFC1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | 0.009346 (rank : 82) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q61191, Q684R1, Q7TSB0, Q8C2D0, Q9QWH2 | Gene names | Hcfc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Host cell factor (HCF) (HCF-1) (C1 factor) [Contains: HCF N-terminal chain 1; HCF N-terminal chain 2; HCF N-terminal chain 3; HCF N- terminal chain 4; HCF N-terminal chain 5; HCF N-terminal chain 6; HCF C-terminal chain 1; HCF C-terminal chain 2; HCF C-terminal chain 3; HCF C-terminal chain 4; HCF C-terminal chain 5; HCF C-terminal chain 6]. | |||||
|
HIPK2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 97) | NC score | -0.001784 (rank : 100) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 871 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QZR5, O88905, Q99P45, Q99P46, Q9D2E6, Q9D474, Q9EQL2, Q9QZR4 | Gene names | Hipk2, Nbak1, Stank | |||
|
Domain Architecture |

|
|||||
| Description | Homeodomain-interacting protein kinase 2 (EC 2.7.11.1) (Nuclear body- associated kinase 1) (Sialophorin tail-associated nuclear serine/threonine-protein kinase). | |||||
|
MARK2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 98) | NC score | -0.002302 (rank : 102) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q05512, Q6PDR4, Q8BR95 | Gene names | Mark2, Emk | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL Motif Kinase) (EMK1). | |||||
|
OTX1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 99) | NC score | 0.003201 (rank : 97) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 379 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P32242, Q53TG6 | Gene names | OTX1 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein OTX1. | |||||
|
REL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 100) | NC score | 0.005926 (rank : 90) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q04864, Q2PNZ7, Q6LDY0 | Gene names | REL | |||
|
Domain Architecture |

|
|||||
| Description | C-Rel proto-oncogene protein (C-Rel protein). | |||||
|
SRBP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 101) | NC score | 0.008895 (rank : 85) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
CRSP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.065701 (rank : 18) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O60244, Q9UNB3 | Gene names | CRSP2, ARC150, DRIP150, EXLM1, TRAP170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CRSP complex subunit 2 (Cofactor required for Sp1 transcriptional activation subunit 2) (Transcriptional coactivator CRSP150) (Vitamin D3 receptor-interacting protein complex 150 kDa component) (DRIP150) (Thyroid hormone receptor-associated protein complex 170 kDa component) (Trap170) (Activator-recruited cofactor 150 kDa component) (ARC150). | |||||
|
SMAD4_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 101 | |
| SwissProt Accessions | P97471, Q9CW56 | Gene names | Smad4, Dpc4, Madh4 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 4 (SMAD 4) (Mothers against DPP homolog 4) (Deletion target in pancreatic carcinoma 4 homolog) (Smad4). | |||||
|
SMAD4_HUMAN
|
||||||
| NC score | 0.985217 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q13485 | Gene names | SMAD4, DPC4, MADH4 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 4 (SMAD 4) (Mothers against DPP homolog 4) (Deletion target in pancreatic carcinoma 4) (hSMAD4). | |||||
|
SMAD3_HUMAN
|
||||||
| NC score | 0.926398 (rank : 3) | θ value | 2.24614e-67 (rank : 5) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P84022, O09064, O09144, O14510, O35273, Q92940, Q93002, Q9GKR4 | Gene names | SMAD3, MADH3 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 3 (SMAD 3) (Mothers against DPP homolog 3) (Mad3) (hMAD-3) (JV15-2) (hSMAD3). | |||||
|
SMAD3_MOUSE
|
||||||
| NC score | 0.926398 (rank : 4) | θ value | 2.24614e-67 (rank : 6) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BUN5, O09064, O09144, O14510, O35273, Q8BX84, Q92940, Q93002, Q9GKR4 | Gene names | Smad3, Madh3 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 3 (SMAD 3) (Mothers against DPP homolog 3) (Mad3) (mMad3). | |||||
|
SMAD9_HUMAN
|
||||||
| NC score | 0.925348 (rank : 5) | θ value | 1.00825e-67 (rank : 3) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O15198, O14989 | Gene names | SMAD9, MADH6, MADH9 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 9 (SMAD 9) (Mothers against DPP homolog 9) (Smad9) (Madh6). | |||||
|
SMAD5_HUMAN
|
||||||
| NC score | 0.923893 (rank : 6) | θ value | 2.4834e-66 (rank : 7) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q99717, O14688, Q15798, Q9UQA1 | Gene names | SMAD5, MADH5 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 5 (SMAD 5) (Mothers against DPP homolog 5) (Smad5) (hSmad5) (JV5-1). | |||||
|
SMAD5_MOUSE
|
||||||
| NC score | 0.923047 (rank : 7) | θ value | 1.71981e-67 (rank : 4) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P97454, P70341, Q810K0 | Gene names | Smad5, Madh5, Msmad5 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 5 (SMAD 5) (Mothers against DPP homolog 5) (Smad5) (mSmad5) (Dwarfin-C) (Dwf-C). | |||||
|
SMAD1_HUMAN
|
||||||
| NC score | 0.922510 (rank : 8) | θ value | 4.23606e-66 (rank : 8) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q15797, Q16636 | Gene names | SMAD1, BSP1, MADH1, MADR1 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 1 (SMAD 1) (Mothers against DPP homolog 1) (Mad-related protein 1) (Transforming growth factor- beta signaling protein 1) (BSP-1) (hSMAD1) (JV4-1). | |||||
|
SMAD1_MOUSE
|
||||||
| NC score | 0.922113 (rank : 9) | θ value | 1.36269e-64 (rank : 9) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P70340, P70442, Q9CYK6 | Gene names | Smad1, Madh1, Madr1 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 1 (SMAD 1) (Mothers against DPP homolog 1) (Mad-related protein 1) (Mothers-against-DPP-related 1) (mMad1) (Dwarfin-A) (Dwf-A). | |||||
|
SMAD9_MOUSE
|
||||||
| NC score | 0.914955 (rank : 10) | θ value | 4.38363e-63 (rank : 10) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9JIW5, Q8CFF9 | Gene names | Smad9, Madh8, Madh9, Smad8 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 9 (SMAD 9) (Mothers against DPP homolog 9) (Smad9) (Smad8). | |||||
|
SMAD2_MOUSE
|
||||||
| NC score | 0.906762 (rank : 11) | θ value | 1.2411e-41 (rank : 12) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q62432, Q9D8P6 | Gene names | Smad2, Madh2, Madr2 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 2 (SMAD 2) (Mothers against DPP homolog 2) (Mad-related protein 2) (mMad2). | |||||
|
SMAD2_HUMAN
|
||||||
| NC score | 0.906736 (rank : 12) | θ value | 1.2411e-41 (rank : 11) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15796 | Gene names | SMAD2, MADH2, MADR2 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 2 (SMAD 2) (Mothers against DPP homolog 2) (Mad-related protein 2) (hMAD-2) (JV18-1) (hSMAD2). | |||||
|
SMAD6_MOUSE
|
||||||
| NC score | 0.778363 (rank : 13) | θ value | 2.06002e-20 (rank : 14) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O35182, Q9CW62 | Gene names | Smad6, Madh6, Madh7, Msmad6 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 6 (SMAD 6) (Mothers against DPP homolog 6) (Smad6) (Mad homolog 7). | |||||
|
SMAD6_HUMAN
|
||||||
| NC score | 0.777164 (rank : 14) | θ value | 1.2077e-20 (rank : 13) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O43541, O43654, Q15799, Q9UKZ3 | Gene names | SMAD6, MADH6 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 6 (SMAD 6) (Mothers against DPP homolog 6) (Smad6) (hSMAD6). | |||||
|
SMAD7_MOUSE
|
||||||
| NC score | 0.762637 (rank : 15) | θ value | 2.7842e-17 (rank : 16) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O35253, O88709 | Gene names | Smad7, Madh7 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 7 (SMAD 7) (Mothers against DPP homolog 7) (Smad7) (Madh8). | |||||
|
SMAD7_HUMAN
|
||||||
| NC score | 0.761781 (rank : 16) | θ value | 2.7842e-17 (rank : 15) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O15105, O14740, Q6DK23 | Gene names | SMAD7, MADH7, MADH8 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 7 (SMAD 7) (Mothers against DPP homolog 7) (Smad7) (hSMAD7). | |||||
|
PORIM_HUMAN
|
||||||
| NC score | 0.096395 (rank : 17) | θ value | 0.0113563 (rank : 19) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N131, Q8IWS2, Q96QV2 | Gene names | TMEM123, KCT3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Porimin precursor (Transmembrane protein 123) (Pro-oncosis receptor inducing membrane injury) (Keratinocytes-associated transmembrane protein 3) (KCT-3). | |||||
|
CRSP2_HUMAN
|
||||||
| NC score | 0.065701 (rank : 18) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O60244, Q9UNB3 | Gene names | CRSP2, ARC150, DRIP150, EXLM1, TRAP170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CRSP complex subunit 2 (Cofactor required for Sp1 transcriptional activation subunit 2) (Transcriptional coactivator CRSP150) (Vitamin D3 receptor-interacting protein complex 150 kDa component) (DRIP150) (Thyroid hormone receptor-associated protein complex 170 kDa component) (Trap170) (Activator-recruited cofactor 150 kDa component) (ARC150). | |||||
|
BORG5_HUMAN
|
||||||
| NC score | 0.057159 (rank : 19) | θ value | 0.0431538 (rank : 22) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q00587, Q96GN1 | Gene names | CDC42EP1, BORG5, CEP1, MSE55 | |||
|
Domain Architecture |

|
|||||
| Description | Cdc42 effector protein 1 (Binder of Rho GTPases 5) (Serum protein MSE55). | |||||
|
ATX2L_HUMAN
|
||||||
| NC score | 0.052725 (rank : 20) | θ value | 0.47712 (rank : 34) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8WWM7, O95135, Q6NVJ8, Q6PJW6, Q8IU61, Q8IU95, Q8WWM3, Q8WWM4, Q8WWM5, Q8WWM6, Q99703 | Gene names | ATXN2L, A2D, A2LG, A2LP, A2RP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2-like protein (Ataxin-2 domain protein) (Ataxin-2-related protein). | |||||
|
NCOA6_MOUSE
|
||||||
| NC score | 0.052407 (rank : 21) | θ value | 1.38821 (rank : 45) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
ATX2L_MOUSE
|
||||||
| NC score | 0.051267 (rank : 22) | θ value | 0.813845 (rank : 36) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q7TQH0, Q80XN9, Q8K059 | Gene names | Atxn2l, A2lp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2-like protein. | |||||
|
GLP_MOUSE
|
||||||
| NC score | 0.049426 (rank : 23) | θ value | 3.0926 (rank : 57) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P14220 | Gene names | Gypa | |||
|
Domain Architecture |

|
|||||
| Description | Glycophorin. | |||||
|
PODXL_MOUSE
|
||||||
| NC score | 0.047773 (rank : 24) | θ value | 0.813845 (rank : 40) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R0M4, Q9ESZ1 | Gene names | Podxl, Pclp1 | |||
|
Domain Architecture |

|
|||||
| Description | Podocalyxin-like protein 1 precursor. | |||||
|
SF01_MOUSE
|
||||||
| NC score | 0.046383 (rank : 25) | θ value | 0.163984 (rank : 28) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q64213, O08817, P70167, Q61454, Q921Z4 | Gene names | Sf1, Zfm1, Zfp162 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 1 (Zinc finger protein 162) (Transcription factor ZFM1) (mZFM) (Zinc finger gene in MEN1 locus) (Mammalian branch point- binding protein mBBP) (BBP) (CW17). | |||||
|
SF01_HUMAN
|
||||||
| NC score | 0.046067 (rank : 26) | θ value | 0.163984 (rank : 27) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15637, Q14818, Q14819, Q15913, Q8IY00, Q92744, Q92745, Q969H7, Q9BW01, Q9UEI0 | Gene names | SF1, ZFM1, ZNF162 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 1 (Zinc finger protein 162) (Transcription factor ZFM1) (Zinc finger gene in MEN1 locus) (Mammalian branch point-binding protein mBBP) (BBP). | |||||
|
CXXC6_HUMAN
|
||||||
| NC score | 0.044894 (rank : 27) | θ value | 0.163984 (rank : 25) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
FOXJ3_HUMAN
|
||||||
| NC score | 0.044533 (rank : 28) | θ value | 0.000786445 (rank : 17) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UPW0, Q9NSS7 | Gene names | FOXJ3, KIAA1041 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein J3. | |||||
|
MAP4_MOUSE
|
||||||
| NC score | 0.039682 (rank : 29) | θ value | 0.163984 (rank : 26) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P27546 | Gene names | Map4, Mtap4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
FOXJ3_MOUSE
|
||||||
| NC score | 0.039514 (rank : 30) | θ value | 0.00102713 (rank : 18) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BUR3, Q3UTD8 | Gene names | Foxj3, Kiaa1041 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein J3. | |||||
|
TFG_HUMAN
|
||||||
| NC score | 0.038524 (rank : 31) | θ value | 1.06291 (rank : 44) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92734, Q15656 | Gene names | TFG | |||
|
Domain Architecture |

|
|||||
| Description | Protein TFG (TRK-fused gene protein). | |||||
|
R3HD2_MOUSE
|
||||||
| NC score | 0.034548 (rank : 32) | θ value | 0.279714 (rank : 32) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
MAP4_HUMAN
|
||||||
| NC score | 0.034095 (rank : 33) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 428 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P27816, Q13082, Q96A76 | Gene names | MAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
IF4G1_HUMAN
|
||||||
| NC score | 0.031006 (rank : 34) | θ value | 0.21417 (rank : 29) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q04637, O43177, O95066, Q5HYG0, Q6ZN21, Q8N102 | Gene names | EIF4G1, EIF4G, EIF4GI | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1) (p220). | |||||
|
ARI1A_HUMAN
|
||||||
| NC score | 0.030946 (rank : 35) | θ value | 0.163984 (rank : 23) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
ADRM1_HUMAN
|
||||||
| NC score | 0.030915 (rank : 36) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q16186, Q9H1P2 | Gene names | ADRM1, GP110 | |||
|
Domain Architecture |

|
|||||
| Description | Adhesion-regulating molecule 1 precursor (110 kDa cell membrane glycoprotein) (Gp110). | |||||
|
CD68_HUMAN
|
||||||
| NC score | 0.030883 (rank : 37) | θ value | 0.813845 (rank : 37) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P34810, Q96BI7 | Gene names | CD68 | |||
|
Domain Architecture |

|
|||||
| Description | Macrosialin precursor (GP110) (CD68 antigen). | |||||
|
CREB5_HUMAN
|
||||||
| NC score | 0.030869 (rank : 38) | θ value | 0.163984 (rank : 24) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q02930, Q05886, Q06246 | Gene names | CREB5, CREBPA | |||
|
Domain Architecture |

|
|||||
| Description | cAMP response element-binding protein 5 (CRE-BPa). | |||||
|
PO3F3_MOUSE
|
||||||
| NC score | 0.030866 (rank : 39) | θ value | 0.0148317 (rank : 20) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P31361 | Gene names | Pou3f3, Brn-1, Brn1, Otf8 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 3, transcription factor 3 (Brain-specific homeobox/POU domain protein 1) (Brain-1) (Brn-1 protein). | |||||
|
PO3F3_HUMAN
|
||||||
| NC score | 0.030227 (rank : 40) | θ value | 0.0252991 (rank : 21) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P20264, P78379 | Gene names | POU3F3, BRN1, OTF8 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 3, transcription factor 3 (Brain-specific homeobox/POU domain protein 1) (Brain-1) (Brn-1 protein). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.028850 (rank : 41) | θ value | 2.36792 (rank : 53) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
ATN1_HUMAN
|
||||||
| NC score | 0.028496 (rank : 42) | θ value | 4.03905 (rank : 61) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
LRC41_HUMAN
|
||||||
| NC score | 0.028439 (rank : 43) | θ value | 2.36792 (rank : 52) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15345, Q71RA8, Q9BSM0 | Gene names | LRRC41, MUF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 41 (Protein Muf1). | |||||
|
RERE_MOUSE
|
||||||
| NC score | 0.025740 (rank : 44) | θ value | 1.81305 (rank : 49) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
MUC7_HUMAN
|
||||||
| NC score | 0.024920 (rank : 45) | θ value | 1.06291 (rank : 42) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
MUC1_MOUSE
|
||||||
| NC score | 0.024717 (rank : 46) | θ value | 1.81305 (rank : 48) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
MUC2_HUMAN
|
||||||
| NC score | 0.022938 (rank : 47) | θ value | 0.813845 (rank : 39) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q02817, Q14878 | Gene names | MUC2, SMUC | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-2 precursor (Intestinal mucin-2). | |||||
|
WASIP_HUMAN
|
||||||
| NC score | 0.021382 (rank : 48) | θ value | 0.813845 (rank : 41) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
RERE_HUMAN
|
||||||
| NC score | 0.021280 (rank : 49) | θ value | 3.0926 (rank : 60) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
NPAS4_HUMAN
|
||||||
| NC score | 0.020999 (rank : 50) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
WIRE_MOUSE
|
||||||
| NC score | 0.020984 (rank : 51) | θ value | 2.36792 (rank : 54) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6PEV3 | Gene names | Wire | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein). | |||||
|
PO3F2_HUMAN
|
||||||
| NC score | 0.020496 (rank : 52) | θ value | 5.27518 (rank : 71) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P20265, Q14960, Q86V54, Q9UJL0 | Gene names | POU3F2, BRN2, OCT7, OTF7 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 3, transcription factor 2 (Nervous system-specific octamer-binding transcription factor N-Oct-3) (Brain-specific homeobox/POU domain protein 2) (Brain-2) (Protein Brn-2). | |||||
|
PO3F2_MOUSE
|
||||||
| NC score | 0.020479 (rank : 53) | θ value | 5.27518 (rank : 72) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P31360 | Gene names | Pou3f2, Brn-2, Brn2, Otf7 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 3, transcription factor 2 (Nervous system-specific octamer-binding transcription factor N-Oct-3) (Brain-specific homeobox/POU domain protein 2) (Brain-2) (Protein Brn-2). | |||||
|
ONEC2_MOUSE
|
||||||
| NC score | 0.019896 (rank : 54) | θ value | 0.62314 (rank : 35) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6XBJ3 | Gene names | Onecut2, Oc2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | One cut domain family member 2 (Transcription factor ONECUT-2) (OC-2). | |||||
|
MAFB_MOUSE
|
||||||
| NC score | 0.018405 (rank : 55) | θ value | 0.813845 (rank : 38) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P54841 | Gene names | Mafb, Krml, Maf1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor MafB (V-maf musculoaponeurotic fibrosarcoma oncogene homolog B) (Transcription factor MAF1) (Segmentation protein KR) (Kreisler). | |||||
|
CSPG3_HUMAN
|
||||||
| NC score | 0.017006 (rank : 56) | θ value | 0.279714 (rank : 31) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O14594, Q9UPK6 | Gene names | CSPG3, NCAN, NEUR | |||
|
Domain Architecture |

|
|||||
| Description | Neurocan core protein precursor (Chondroitin sulfate proteoglycan 3). | |||||
|
BRPF1_HUMAN
|
||||||
| NC score | 0.016988 (rank : 57) | θ value | 0.279714 (rank : 30) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P55201, Q9UHI0 | Gene names | BRPF1, BR140 | |||
|
Domain Architecture |

|
|||||
| Description | Peregrin (Bromodomain and PHD finger-containing protein 1) (BR140 protein). | |||||
|
MUC5B_HUMAN
|
||||||
| NC score | 0.016826 (rank : 58) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1020 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9HC84, O00447, O00573, O14985, O15494, O95291, O95451, Q14881, Q99552, Q9UE28 | Gene names | MUC5B, MUC5 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-5B precursor (Mucin-5 subtype B, tracheobronchial) (High molecular weight salivary mucin MG1) (Sublingual gland mucin). | |||||
|
ABI1_HUMAN
|
||||||
| NC score | 0.016806 (rank : 59) | θ value | 0.47712 (rank : 33) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8IZP0, O15147, O76049, O95060, Q5W070, Q5W072, Q8TB63, Q96S81, Q9NXZ9, Q9NYB8 | Gene names | ABI1, SSH3BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Abl interactor 1 (Abelson interactor 1) (Abi-1) (Spectrin SH3 domain- binding protein 1) (Eps8 SH3 domain-binding protein) (Eps8-binding protein) (e3B1) (Nap1-binding protein) (Nap1BP) (Abl-binding protein 4) (AblBP4). | |||||
|
CJ047_MOUSE
|
||||||
| NC score | 0.016382 (rank : 60) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8C5R2, Q8R017, Q8R2Z9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf47 homolog. | |||||
|
LAMP3_HUMAN
|
||||||
| NC score | 0.015860 (rank : 61) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UQV4, O94781, Q8NEC8 | Gene names | LAMP3, DCLAMP, TSC403 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lysosome-associated membrane glycoprotein 3 precursor (LAMP-3) (Lysosomal-associated membrane protein 3) (DC-lysosome-associated membrane glycoprotein) (DC LAMP) (Protein TSC403) (CD208 antigen). | |||||
|
CBP_HUMAN
|
||||||
| NC score | 0.015773 (rank : 62) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
VGLL2_MOUSE
|
||||||
| NC score | 0.015109 (rank : 63) | θ value | 5.27518 (rank : 74) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BGW8, Q8BWB4 | Gene names | Vgll2, Vgl2, Vito1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription cofactor vestigial-like protein 2 (Vgl-2) (Protein VITO1). | |||||
|
IQEC2_HUMAN
|
||||||
| NC score | 0.014266 (rank : 64) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5JU85, O60275 | Gene names | IQSEC2, KIAA0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
IQEC2_MOUSE
|
||||||
| NC score | 0.014212 (rank : 65) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5DU25 | Gene names | Iqsec2, Kiaa0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
PF21B_MOUSE
|
||||||
| NC score | 0.013756 (rank : 66) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8C966 | Gene names | Phf21b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
ALMS1_HUMAN
|
||||||
| NC score | 0.013497 (rank : 67) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8TCU4, Q53S05, Q580Q8, Q86VP9, Q9Y4G4 | Gene names | ALMS1, KIAA0328 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alstrom syndrome protein 1. | |||||
|
BPTF_HUMAN
|
||||||
| NC score | 0.012102 (rank : 68) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
CRF_HUMAN
|
||||||
| NC score | 0.011930 (rank : 69) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P06850 | Gene names | CRH | |||
|
Domain Architecture |

|
|||||
| Description | Corticoliberin precursor (Corticotropin-releasing factor) (CRF) (Corticotropin-releasing hormone). | |||||
|
EYA1_HUMAN
|
||||||
| NC score | 0.011577 (rank : 70) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99502 | Gene names | EYA1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
|
MTG8_MOUSE
|
||||||
| NC score | 0.011562 (rank : 71) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61909 | Gene names | Runx1t1, Cbfa2t1, Cbfa2t1h, Mtg8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein CBFA2T1 (Protein MTG8). | |||||
|
MTG8_HUMAN
|
||||||
| NC score | 0.011555 (rank : 72) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q06455, Q06456, Q14873, Q16239, Q16346, Q16347, Q92479, Q9BRZ0 | Gene names | RUNX1T1, AML1T1, CBFA2T1, CDR, ETO, MTG8, ZMYND2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein CBFA2T1 (Protein MTG8) (Protein ETO) (Eight twenty one protein) (Cyclin-D-related protein) (Zinc finger MYND domain- containing protein 2). | |||||
|
RCOR3_HUMAN
|
||||||
| NC score | 0.011016 (rank : 73) | θ value | 6.88961 (rank : 89) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9P2K3, Q5VT47, Q7L9I5, Q8N5U3, Q9NV83 | Gene names | RCOR3, KIAA1343 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
REXO1_HUMAN
|
||||||
| NC score | 0.010356 (rank : 74) | θ value | 5.27518 (rank : 73) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N1G1, Q9ULT2 | Gene names | REXO1, ELOABP1, KIAA1138, TCEB3BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA exonuclease 1 homolog (EC 3.1.-.-) (Elongin A-binding protein 1) (EloA-BP1) (Transcription elongation factor B polypeptide 3-binding protein 1). | |||||
|
FIGN_HUMAN
|
||||||
| NC score | 0.010294 (rank : 75) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5HY92, Q9H6M5, Q9NVZ9 | Gene names | FIGN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fidgetin. | |||||
|
YAP1_MOUSE
|
||||||
| NC score | 0.009919 (rank : 76) | θ value | 5.27518 (rank : 75) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P46938, Q52KJ5 | Gene names | Yap1, Yap, Yap65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
IASPP_HUMAN
|
||||||
| NC score | 0.009796 (rank : 77) | θ value | 4.03905 (rank : 62) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8WUF5, Q2PNZ9, Q5DU71, Q5I1X4, Q6P1R7, Q6PKF8, Q9Y290 | Gene names | PPP1R13L, IASPP, NKIP1, PPP1R13BL, RAI | |||
|
Domain Architecture |

|
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
PKHA4_HUMAN
|
||||||
| NC score | 0.009644 (rank : 78) | θ value | 5.27518 (rank : 70) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H4M7, Q8N4M8, Q8N658 | Gene names | PLEKHA4, PEPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 4 (Phosphoinositol 3-phosphate-binding protein 1) (PEPP-1). | |||||
|
CR001_HUMAN
|
||||||
| NC score | 0.009630 (rank : 79) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15165, O15166, O15167, O15168, Q6NXP3 | Gene names | C18orf1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein C18orf1. | |||||
|
EYA1_MOUSE
|
||||||
| NC score | 0.009431 (rank : 80) | θ value | 8.99809 (rank : 93) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P97767, O08818 | Gene names | Eya1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
|
NUMBL_MOUSE
|
||||||
| NC score | 0.009350 (rank : 81) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O08919, Q6NVG8 | Gene names | Numbl, Nbl | |||
|
Domain Architecture |

|
|||||
| Description | Numb-like protein. | |||||
|
HCFC1_MOUSE
|
||||||
| NC score | 0.009346 (rank : 82) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q61191, Q684R1, Q7TSB0, Q8C2D0, Q9QWH2 | Gene names | Hcfc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Host cell factor (HCF) (HCF-1) (C1 factor) [Contains: HCF N-terminal chain 1; HCF N-terminal chain 2; HCF N-terminal chain 3; HCF N- terminal chain 4; HCF N-terminal chain 5; HCF N-terminal chain 6; HCF C-terminal chain 1; HCF C-terminal chain 2; HCF C-terminal chain 3; HCF C-terminal chain 4; HCF C-terminal chain 5; HCF C-terminal chain 6]. | |||||
|
PA24B_HUMAN
|
||||||
| NC score | 0.009250 (rank : 83) | θ value | 6.88961 (rank : 88) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95712, Q59GF9, Q8TB10, Q9UKV7 | Gene names | PLA2G4B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytosolic phospholipase A2 beta (EC 3.1.1.4) (cPLA2-beta) (Phospholipase A2 group IVB). | |||||
|
ASPP2_HUMAN
|
||||||
| NC score | 0.009206 (rank : 84) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q13625, Q12892, Q86X75, Q96KQ3 | Gene names | TP53BP2, ASPP2, BBP | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-stimulating of p53 protein 2 (Tumor suppressor p53-binding protein 2) (p53-binding protein 2) (p53BP2) (53BP2) (Bcl2-binding protein) (Bbp) (NY-REN-51 antigen). | |||||
|
SRBP1_MOUSE
|
||||||
| NC score | 0.008895 (rank : 85) | θ value | 8.99809 (rank : 101) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
SHAN1_HUMAN
|
||||||
| NC score | 0.008116 (rank : 86) | θ value | 1.06291 (rank : 43) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y566, Q9NYW9 | Gene names | SHANK1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 1 (Shank1) (Somatostatin receptor-interacting protein) (SSTR-interacting protein) (SSTRIP). | |||||
|
FIGN_MOUSE
|
||||||
| NC score | 0.007941 (rank : 87) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9ERZ6, Q3TPB0, Q3UP57, Q6PCM0 | Gene names | Fign | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fidgetin. | |||||
|
BRAC_HUMAN
|
||||||
| NC score | 0.007596 (rank : 88) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15178 | Gene names | T | |||
|
Domain Architecture |

|
|||||
| Description | Brachyury protein (T protein). | |||||
|
NEDD4_MOUSE
|
||||||
| NC score | 0.007415 (rank : 89) | θ value | 1.38821 (rank : 46) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P46935, O08758, Q3UZI2, Q8BGB3 | Gene names | Nedd4, Kiaa0093, Nedd-4, Nedd4a | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-) (Neural precursor cell expressed developmentally down-regulated protein 4). | |||||
|
REL_HUMAN
|
||||||
| NC score | 0.005926 (rank : 90) | θ value | 8.99809 (rank : 100) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q04864, Q2PNZ7, Q6LDY0 | Gene names | REL | |||
|
Domain Architecture |

|
|||||
| Description | C-Rel proto-oncogene protein (C-Rel protein). | |||||
|
TBX21_MOUSE
|
||||||
| NC score | 0.005528 (rank : 91) | θ value | 6.88961 (rank : 90) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JKD8, Q9R0A6 | Gene names | Tbx21, Tbet, Tblym | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-box transcription factor TBX21 (T-box protein 21) (Transcription factor TBLYM) (T-cell-specific T-box transcription factor T-bet). | |||||
|
GASP2_HUMAN
|
||||||
| NC score | 0.005455 (rank : 92) | θ value | 8.99809 (rank : 95) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96D09, Q8NAB4 | Gene names | GPRASP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor-associated sorting protein 2 (GASP-2). | |||||
|
DLG2_HUMAN
|
||||||
| NC score | 0.004923 (rank : 93) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q15700, Q59G57, Q68CQ8, Q6ZTA8 | Gene names | DLG2 | |||
|
Domain Architecture |

|
|||||
| Description | Discs large homolog 2 (Postsynaptic density protein PSD-93) (Channel- associated protein of synapse-110) (Chapsyn-110). | |||||
|
OTX1_MOUSE
|
||||||
| NC score | 0.004101 (rank : 94) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P80205 | Gene names | Otx1, Otx-1 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein OTX1. | |||||
|
DDEF2_MOUSE
|
||||||
| NC score | 0.004010 (rank : 95) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 625 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7SIG6, Q501K1, Q66JN2 | Gene names | Ddef2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Development and differentiation-enhancing factor 2 (Pyk2 C-terminus- associated protein) (PAP) (Paxillin-associated protein with ARFGAP activity 3) (PAG3). | |||||
|
MEGF9_HUMAN
|
||||||
| NC score | 0.003880 (rank : 96) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9H1U4, O75098 | Gene names | MEGF9, EGFL5, KIAA0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
OTX1_HUMAN
|
||||||
| NC score | 0.003201 (rank : 97) | θ value | 8.99809 (rank : 99) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 379 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P32242, Q53TG6 | Gene names | OTX1 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein OTX1. | |||||
|
ROBO1_HUMAN
|
||||||
| NC score | 0.001184 (rank : 98) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 529 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y6N7, Q7Z300, Q9BUS7 | Gene names | ROBO1, DUTT1 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 1 precursor (H-Robo-1) (Deleted in U twenty twenty). | |||||
|
GLIS3_HUMAN
|
||||||
| NC score | -0.000887 (rank : 99) | θ value | 1.81305 (rank : 47) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 758 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NEA6 | Gene names | GLIS3, ZNF515 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLIS3 (GLI-similar 3) (Zinc finger protein 515). | |||||
|
HIPK2_MOUSE
|
||||||
| NC score | -0.001784 (rank : 100) | θ value | 8.99809 (rank : 97) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 871 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QZR5, O88905, Q99P45, Q99P46, Q9D2E6, Q9D474, Q9EQL2, Q9QZR4 | Gene names | Hipk2, Nbak1, Stank | |||
|
Domain Architecture |

|
|||||
| Description | Homeodomain-interacting protein kinase 2 (EC 2.7.11.1) (Nuclear body- associated kinase 1) (Sialophorin tail-associated nuclear serine/threonine-protein kinase). | |||||
|
MARK2_HUMAN
|
||||||
| NC score | -0.001788 (rank : 101) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1056 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7KZI7, Q15449, Q15524, Q5XGA3, Q68A18, Q96HB3, Q96RG0 | Gene names | MARK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL motif kinase) (EMK1) (PAR1 homolog). | |||||
|
MARK2_MOUSE
|
||||||
| NC score | -0.002302 (rank : 102) | θ value | 8.99809 (rank : 98) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q05512, Q6PDR4, Q8BR95 | Gene names | Mark2, Emk | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL Motif Kinase) (EMK1). | |||||