Please be patient as the page loads
|
EYA1_MOUSE
|
||||||
| SwissProt Accessions | P97767, O08818 | Gene names | Eya1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
EYA1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.989351 (rank : 2) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q99502 | Gene names | EYA1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
|
EYA1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 108 | |
| SwissProt Accessions | P97767, O08818 | Gene names | Eya1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
|
EYA4_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.969381 (rank : 6) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O95677, O95464, O95679, Q8IW39, Q9NTR7 | Gene names | EYA4 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 4 (EC 3.1.3.48). | |||||
|
EYA4_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.974209 (rank : 3) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Z191 | Gene names | Eya4 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 4 (EC 3.1.3.48). | |||||
|
EYA2_MOUSE
|
||||||
| θ value | 5.01279e-176 (rank : 5) | NC score | 0.973625 (rank : 4) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O08575, P97925 | Gene names | Eya2, Eab1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 2 (EC 3.1.3.48). | |||||
|
EYA2_HUMAN
|
||||||
| θ value | 3.5924e-174 (rank : 6) | NC score | 0.971986 (rank : 5) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O00167, Q5JSW8, Q96CV6, Q96H97, Q99503, Q99812, Q9BWF6, Q9H4S3, Q9H4S9, Q9NPZ4, Q9UIX7 | Gene names | EYA2, EAB1 | |||
|
Domain Architecture |
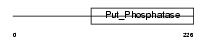
|
|||||
| Description | Eyes absent homolog 2 (EC 3.1.3.48). | |||||
|
EYA3_HUMAN
|
||||||
| θ value | 8.9516e-133 (rank : 7) | NC score | 0.962100 (rank : 8) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99504, O95463, Q99813 | Gene names | EYA3 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 3 (EC 3.1.3.48). | |||||
|
EYA3_MOUSE
|
||||||
| θ value | 9.94316e-116 (rank : 8) | NC score | 0.962209 (rank : 7) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P97480, P97768 | Gene names | Eya3 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 3 (EC 3.1.3.48). | |||||
|
EWS_HUMAN
|
||||||
| θ value | 3.77169e-06 (rank : 9) | NC score | 0.177632 (rank : 11) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q01844, Q92635 | Gene names | EWSR1, EWS | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein EWS (EWS oncogene) (Ewing sarcoma breakpoint region 1 protein). | |||||
|
EWS_MOUSE
|
||||||
| θ value | 4.92598e-06 (rank : 10) | NC score | 0.179623 (rank : 10) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q61545 | Gene names | Ewsr1, Ews, Ewsh | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein EWS. | |||||
|
FUS_MOUSE
|
||||||
| θ value | 9.29e-05 (rank : 11) | NC score | 0.180836 (rank : 9) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P56959 | Gene names | Fus | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein FUS (Pigpen protein). | |||||
|
FUS_HUMAN
|
||||||
| θ value | 0.000121331 (rank : 12) | NC score | 0.171648 (rank : 12) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P35637 | Gene names | FUS, TLS | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein FUS (Oncogene FUS) (Oncogene TLS) (Translocated in liposarcoma protein) (POMp75) (75 kDa DNA-pairing protein). | |||||
|
TRIM2_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 13) | NC score | 0.039154 (rank : 36) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9C040, O60272, Q9BSI9, Q9UFZ1 | Gene names | TRIM2, KIAA0517, RNF86 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 2 (RING finger protein 86). | |||||
|
TRIM2_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 14) | NC score | 0.039264 (rank : 35) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9ESN6, Q3UHP5, Q8C981 | Gene names | Trim2, Narf | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 2 (Neural activity-related RING finger protein). | |||||
|
SOX8_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 15) | NC score | 0.027916 (rank : 60) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P57073, Q9NZW2 | Gene names | SOX8 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-8. | |||||
|
MUCDL_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 16) | NC score | 0.039588 (rank : 32) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9HBB8, Q9H746, Q9HAU3, Q9HBB5, Q9HBB6, Q9HBB7, Q9NX86, Q9NXI9 | Gene names | MUCDHL | |||
|
Domain Architecture |

|
|||||
| Description | Mucin and cadherin-like protein precursor (Mu-protocadherin). | |||||
|
RBM14_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 17) | NC score | 0.045300 (rank : 27) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8C2Q3, Q3TJB6, Q91Z21, Q9DBI6 | Gene names | Rbm14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 14 (RNA-binding motif protein 14). | |||||
|
SOX8_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 18) | NC score | 0.038482 (rank : 38) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q04886 | Gene names | Sox8, Sox-8 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-8. | |||||
|
DDX17_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 19) | NC score | 0.038763 (rank : 37) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q92841, Q69YT1, Q6ICD6 | Gene names | DDX17 | |||
|
Domain Architecture |
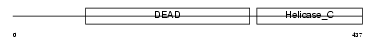
|
|||||
| Description | Probable ATP-dependent RNA helicase DDX17 (EC 3.6.1.-) (DEAD box protein 17) (RNA-dependent helicase p72) (DEAD box protein p72). | |||||
|
ZBT16_HUMAN
|
||||||
| θ value | 0.125558 (rank : 20) | NC score | 0.001693 (rank : 106) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1010 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q05516, Q8TAL4 | Gene names | ZBTB16, PLZF, ZNF145 | |||
|
Domain Architecture |
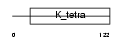
|
|||||
| Description | Zinc finger and BTB domain-containing protein 16 (Zinc finger protein PLZF) (Promyelocytic leukemia zinc finger protein) (Zinc finger protein 145). | |||||
|
ZIMP7_MOUSE
|
||||||
| θ value | 0.125558 (rank : 21) | NC score | 0.044137 (rank : 29) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 566 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8CIE2, Q6P558, Q6P9Q5, Q8BJ00 | Gene names | Zimp7, D11Bwg0280e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PIAS-like protein Zimp7. | |||||
|
BAG4_MOUSE
|
||||||
| θ value | 0.163984 (rank : 22) | NC score | 0.060937 (rank : 19) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8CI61, Q3TRL9, Q91VT5, Q9CWG2 | Gene names | Bag4, Sodd | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 4 (BCL2-associated athanogene 4) (BAG-4) (Silencer of death domains). | |||||
|
CNOT4_MOUSE
|
||||||
| θ value | 0.163984 (rank : 23) | NC score | 0.037790 (rank : 39) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BT14, Q8CCR4, Q9CV74, Q9Z1D0 | Gene names | Cnot4, Not4 | |||
|
Domain Architecture |
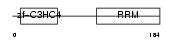
|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
NU214_HUMAN
|
||||||
| θ value | 0.21417 (rank : 24) | NC score | 0.074409 (rank : 14) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P35658 | Gene names | NUP214, CAIN, CAN | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup214 (Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein). | |||||
|
PPRB_HUMAN
|
||||||
| θ value | 0.21417 (rank : 25) | NC score | 0.039301 (rank : 33) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15648, O43810, O75447, Q9HD39 | Gene names | PPARBP, ARC205, DRIP205, TRAP220, TRIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2) (p53 regulatory protein RB18A) (Vitamin D receptor-interacting protein complex component DRIP205) (Activator- recruited cofactor 205 kDa component) (ARC205). | |||||
|
SSXT_MOUSE
|
||||||
| θ value | 0.21417 (rank : 26) | NC score | 0.047858 (rank : 24) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q62280 | Gene names | Ss18, Ssxt, Syt | |||
|
Domain Architecture |
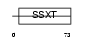
|
|||||
| Description | SSXT protein (SYT protein) (Synovial sarcoma-associated Ss18-alpha). | |||||
|
DDX17_MOUSE
|
||||||
| θ value | 0.279714 (rank : 27) | NC score | 0.033471 (rank : 48) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q501J6, Q6P5G1, Q8BIN2 | Gene names | Ddx17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX17 (EC 3.6.1.-) (DEAD box protein 17). | |||||
|
GATA4_HUMAN
|
||||||
| θ value | 0.365318 (rank : 28) | NC score | 0.027625 (rank : 61) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P43694 | Gene names | GATA4 | |||
|
Domain Architecture |
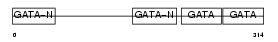
|
|||||
| Description | Transcription factor GATA-4 (GATA-binding factor 4). | |||||
|
DDX3Y_HUMAN
|
||||||
| θ value | 0.47712 (rank : 29) | NC score | 0.031576 (rank : 51) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O15523, Q8IYV7 | Gene names | DDX3Y, DBY | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX3Y (EC 3.6.1.-) (DEAD box protein 3, Y- chromosomal). | |||||
|
ILF3_MOUSE
|
||||||
| θ value | 0.47712 (rank : 30) | NC score | 0.045885 (rank : 25) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Z1X4, Q80VD5, Q812A1, Q8BP80, Q8C2H8, Q8K588 | Gene names | Ilf3 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin enhancer-binding factor 3. | |||||
|
ZIM10_HUMAN
|
||||||
| θ value | 0.47712 (rank : 31) | NC score | 0.045497 (rank : 26) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9ULJ6, Q7Z7E6 | Gene names | RAI17, KIAA1224, ZIMP10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 17 (PIAS-like protein Zimp10). | |||||
|
DDX3X_HUMAN
|
||||||
| θ value | 0.62314 (rank : 32) | NC score | 0.030033 (rank : 55) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O00571, O15536 | Gene names | DDX3X, DBX, DDX3 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX3X (EC 3.6.1.-) (DEAD box protein 3, X- chromosomal) (Helicase-like protein 2) (HLP2) (DEAD box, X isoform). | |||||
|
MCM3A_MOUSE
|
||||||
| θ value | 0.62314 (rank : 33) | NC score | 0.059528 (rank : 20) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9WUU9 | Gene names | Mcm3ap, Ganp, Map80 | |||
|
Domain Architecture |

|
|||||
| Description | 80 kDa MCM3-associated protein (GANP protein). | |||||
|
NUPL1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 34) | NC score | 0.037150 (rank : 40) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8R332, Q3UJF4, Q8BUA7, Q8BVG7, Q8C0W3 | Gene names | Nupl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin p58/p45 (Nucleoporin-like 1). | |||||
|
PL10_MOUSE
|
||||||
| θ value | 0.62314 (rank : 35) | NC score | 0.029840 (rank : 56) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P16381 | Gene names | D1Pas1, Pl10 | |||
|
Domain Architecture |

|
|||||
| Description | Putative ATP-dependent RNA helicase Pl10 (EC 3.6.1.-). | |||||
|
PUM2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 36) | NC score | 0.043486 (rank : 30) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TB72, O00234, Q53TV7, Q8WY43, Q9HAN2 | Gene names | PUM2, KIAA0235, PUMH2 | |||
|
Domain Architecture |

|
|||||
| Description | Pumilio homolog 2 (Pumilio-2). | |||||
|
SON_HUMAN
|
||||||
| θ value | 0.62314 (rank : 37) | NC score | 0.017141 (rank : 82) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
ZN560_HUMAN
|
||||||
| θ value | 0.62314 (rank : 38) | NC score | 0.000891 (rank : 109) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 751 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96MR9, Q495S9, Q495T1 | Gene names | ZNF560 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 560. | |||||
|
HNRL1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 39) | NC score | 0.065391 (rank : 15) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8VDM6, Q3U201, Q3UPB0, Q6AZA7, Q8BY45, Q8K365 | Gene names | Hnrpul1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein U-like protein 1. | |||||
|
ILF3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 40) | NC score | 0.035775 (rank : 44) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q12906, O43409, Q6P1X1, Q86XY7, Q99544, Q99545, Q9BZH4, Q9BZH5, Q9NQ95, Q9NQ96, Q9NQ97, Q9NQ98, Q9NQ99, Q9NQA0, Q9NQA1, Q9NQA2, Q9NRN2, Q9NRN3, Q9NRN4, Q9UMZ9, Q9UN00, Q9UN84, Q9UNA2 | Gene names | ILF3, DRBF, MPHOSPH4, NF90 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin enhancer-binding factor 3 (Nuclear factor of activated T- cells 90 kDa) (NF-AT-90) (Double-stranded RNA-binding protein 76) (DRBP76) (Translational control protein 80) (TCP80) (Nuclear factor associated with dsRNA) (NFAR) (M-phase phosphoprotein 4) (MPP4). | |||||
|
ARI1B_HUMAN
|
||||||
| θ value | 1.06291 (rank : 41) | NC score | 0.039268 (rank : 34) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 567 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8NFD5, Q5JRD1, Q5VYC4, Q8IZY8, Q8TEV0, Q8TF02, Q99491, Q9ULI5 | Gene names | ARID1B, DAN15, KIAA1235, OSA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-rich interactive domain-containing protein 1B (ARID domain- containing protein 1B) (Osa homolog 2) (hOsa2) (p250R) (BRG1-binding protein hELD/OSA1) (BRG1-associated factor 250b) (BAF250B). | |||||
|
HNRL1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 42) | NC score | 0.063898 (rank : 16) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9BUJ2, O76022, Q6ZSZ0, Q7L8P4, Q8N6Z4, Q96G37, Q9HAL3, Q9UG75 | Gene names | HNRPUL1, E1BAP5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein U-like protein 1 (Adenovirus early region 1B-associated protein 5) (E1B-55 kDa-associated protein 5) (E1B-AP5). | |||||
|
DAZ2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 43) | NC score | 0.036570 (rank : 43) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q13117, Q96P41, Q9NR91 | Gene names | DAZ2 | |||
|
Domain Architecture |
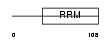
|
|||||
| Description | Deleted in azoospermia protein 2. | |||||
|
DDX3Y_MOUSE
|
||||||
| θ value | 1.38821 (rank : 44) | NC score | 0.028855 (rank : 57) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q62095, Q9QWS9 | Gene names | Ddx3y, D1Pas1-rs1, Dead2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX3Y (EC 3.6.1.-) (DEAD box protein 3, Y- chromosomal) (DEAD-box RNA helicase DEAD2) (mDEAD2) (D1Pas1-related sequence 1). | |||||
|
HORN_HUMAN
|
||||||
| θ value | 1.38821 (rank : 45) | NC score | 0.018159 (rank : 80) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q86YZ3 | Gene names | HRNR, S100A18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hornerin. | |||||
|
MUC1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 46) | NC score | 0.019499 (rank : 76) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
RUNX1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 47) | NC score | 0.016163 (rank : 84) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q03347, O08598, Q62049, Q9ESB9, Q9ET65 | Gene names | Runx1, Aml1, Cbfa2, Pebp2ab | |||
|
Domain Architecture |
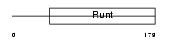
|
|||||
| Description | Runt-related transcription factor 1 (Core-binding factor, alpha 2 subunit) (CBF-alpha 2) (Acute myeloid leukemia 1 protein) (Oncogene AML-1) (Polyomavirus enhancer-binding protein 2 alpha B subunit) (PEBP2-alpha B) (PEA2-alpha B) (SL3-3 enhancer factor 1 alpha B subunit) (SL3/AKV core-binding factor alpha B subunit). | |||||
|
ANXA7_HUMAN
|
||||||
| θ value | 1.81305 (rank : 48) | NC score | 0.015425 (rank : 86) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P20073 | Gene names | ANXA7, ANX7, SNX | |||
|
Domain Architecture |
|
|||||
| Description | Annexin A7 (Annexin VII) (Synexin). | |||||
|
BAG4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 49) | NC score | 0.049833 (rank : 23) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O95429, O95818 | Gene names | BAG4, SODD | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 4 (BCL2-associated athanogene 4) (BAG-4) (Silencer of death domains). | |||||
|
DAZ1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 50) | NC score | 0.036601 (rank : 42) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9NQZ3, Q9NQZ4 | Gene names | DAZ1, DAZ | |||
|
Domain Architecture |
|
|||||
| Description | Deleted in azoospermia protein 1. | |||||
|
DAZ4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 51) | NC score | 0.036645 (rank : 41) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86SG3, Q9NR88, Q9NR89 | Gene names | DAZ4 | |||
|
Domain Architecture |

|
|||||
| Description | Deleted in azoospermia protein 4. | |||||
|
DDX3X_MOUSE
|
||||||
| θ value | 1.81305 (rank : 52) | NC score | 0.027481 (rank : 63) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q62167, O09060, O09143 | Gene names | Ddx3x, D1Pas1-rs2, Ddx3, Dead3, Erh | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX3X (EC 3.6.1.-) (DEAD box protein 3, X- chromosomal) (DEAD box RNA helicase DEAD3) (mDEAD3) (Embryonic RNA helicase) (D1Pas1-related sequence 2). | |||||
|
HTF4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 53) | NC score | 0.020627 (rank : 74) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q99081 | Gene names | TCF12, HEB, HTF4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 12 (Transcription factor HTF-4) (E-box-binding protein) (DNA-binding protein HTF4). | |||||
|
HTF4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 54) | NC score | 0.020511 (rank : 75) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q61286 | Gene names | Tcf12, Alf1, Meb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 12 (Transcription factor HTF-4) (E-box-binding protein) (DNA-binding protein HTF4) (Class A helix-loop-helix transcription factor ME1). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 55) | NC score | 0.026341 (rank : 65) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 56) | NC score | 0.023617 (rank : 70) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
NUP98_HUMAN
|
||||||
| θ value | 1.81305 (rank : 57) | NC score | 0.028426 (rank : 58) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P52948, Q8IUT2, Q8WYB0, Q96E54, Q9H3Q4, Q9NT02, Q9UF57, Q9UHX0, Q9Y6J4, Q9Y6J5 | Gene names | NUP98 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup98-Nup96 precursor [Contains: Nuclear pore complex protein Nup98 (Nucleoporin Nup98) (98 kDa nucleoporin); Nuclear pore complex protein Nup96 (Nucleoporin Nup96) (96 kDa nucleoporin)]. | |||||
|
PDC6I_HUMAN
|
||||||
| θ value | 1.81305 (rank : 58) | NC score | 0.024092 (rank : 69) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8WUM4, Q9BX86, Q9NUN0, Q9P2H2, Q9UKL5 | Gene names | PDCD6IP, AIP1, ALIX, KIAA1375 | |||
|
Domain Architecture |
|
|||||
| Description | Programmed cell death 6-interacting protein (PDCD6-interacting protein) (ALG-2-interacting protein 1) (Hp95). | |||||
|
PEF1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 59) | NC score | 0.012487 (rank : 90) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BFY6, Q8VCT5, Q9CYW8, Q9D934 | Gene names | Pef1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peflin (PEF protein with a long N-terminal hydrophobic domain) (Penta- EF hand domain-containing protein 1). | |||||
|
PPRB_MOUSE
|
||||||
| θ value | 1.81305 (rank : 60) | NC score | 0.035659 (rank : 45) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q925J9, O88323, Q3UHV0, Q6AXD5, Q8BW37, Q8BX19, Q8VDQ7, Q925K0 | Gene names | Pparbp, Trip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2). | |||||
|
PUM2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 61) | NC score | 0.039967 (rank : 31) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80U58, Q80UZ9, Q91YW4, Q925A0, Q9ERC7 | Gene names | Pum2, Kiaa0235 | |||
|
Domain Architecture |

|
|||||
| Description | Pumilio homolog 2. | |||||
|
RERE_MOUSE
|
||||||
| θ value | 1.81305 (rank : 62) | NC score | 0.023594 (rank : 71) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
RNF12_HUMAN
|
||||||
| θ value | 1.81305 (rank : 63) | NC score | 0.030142 (rank : 54) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NVW2, Q9Y598 | Gene names | RNF12, RLIM | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 12 (LIM domain-interacting RING finger protein) (RING finger LIM domain-binding protein) (R-LIM) (NY-REN-43 antigen). | |||||
|
ATN1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 64) | NC score | 0.044555 (rank : 28) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
CBP_MOUSE
|
||||||
| θ value | 2.36792 (rank : 65) | NC score | 0.025578 (rank : 67) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
RPB1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 66) | NC score | 0.063368 (rank : 18) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P24928 | Gene names | POLR2A | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase II largest subunit (EC 2.7.7.6) (RPB1). | |||||
|
RPB1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 67) | NC score | 0.063456 (rank : 17) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P08775 | Gene names | Polr2a, Rpii215, Rpo2-1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase II largest subunit (EC 2.7.7.6) (RPB1). | |||||
|
SOX30_MOUSE
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.027544 (rank : 62) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8CGW4 | Gene names | Sox30 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-30. | |||||
|
CDSN_HUMAN
|
||||||
| θ value | 3.0926 (rank : 69) | NC score | 0.028030 (rank : 59) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q15517, O43509, Q5SQ85, Q5STD2, Q7LA70, Q7LA71, Q86Z04, Q8IZU4, Q8IZU5, Q8IZU6, Q8N5P3, Q95IF9, Q9NP52, Q9NPE0, Q9NPG5, Q9NRH4, Q9NRH5, Q9NRH6, Q9NRH7, Q9NRH8, Q9UBH8, Q9UIN6, Q9UIN7, Q9UIN8, Q9UIN9, Q9UIP0 | Gene names | CDSN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Corneodesmosin precursor (S protein). | |||||
|
HXA5_HUMAN
|
||||||
| θ value | 3.0926 (rank : 70) | NC score | 0.004107 (rank : 102) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 336 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P20719, O43367, Q96CY6 | Gene names | HOXA5, HOX1C | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-A5 (Hox-1C). | |||||
|
S18L1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 71) | NC score | 0.033802 (rank : 46) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75177, Q5JXJ3, Q8NE69, Q9BR55, Q9H4K6 | Gene names | SS18L1, KIAA0693 | |||
|
Domain Architecture |
|
|||||
| Description | SS18-like protein 1 (SYT homolog 1). | |||||
|
SIRT3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 72) | NC score | 0.011224 (rank : 91) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NTG7, Q9Y6E8 | Gene names | SIRT3, SIR2L3 | |||
|
Domain Architecture |
|
|||||
| Description | NAD-dependent deacetylase sirtuin-3, mitochondrial precursor (EC 3.5.1.-) (SIR2-like protein 3) (hSIRT3). | |||||
|
TAB2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 73) | NC score | 0.014611 (rank : 88) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q99K90, Q3UGP1, Q8BTP4, Q8CHD3, Q99KP4 | Gene names | Map3k7ip2, Kiaa0733, Tab2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 7-interacting protein 2 (TAK1-binding protein 2) (TAB2). | |||||
|
YLPM1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 74) | NC score | 0.025570 (rank : 68) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
DAZ3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | 0.033743 (rank : 47) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NR90 | Gene names | DAZ3 | |||
|
Domain Architecture |
|
|||||
| Description | Deleted in azoospermia protein 3. | |||||
|
ETV3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 76) | NC score | 0.006900 (rank : 99) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
HRBL_HUMAN
|
||||||
| θ value | 4.03905 (rank : 77) | NC score | 0.020675 (rank : 73) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95081, O75429, Q96AB9, Q96GL4 | Gene names | HRBL, RABR | |||
|
Domain Architecture |
|
|||||
| Description | HIV-1 Rev-binding protein-like protein (Rev/Rex activation domain- binding protein related) (RAB-R). | |||||
|
K1210_HUMAN
|
||||||
| θ value | 4.03905 (rank : 78) | NC score | 0.018428 (rank : 79) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ULL0, Q5JPN4 | Gene names | KIAA1210 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1210. | |||||
|
K1C10_HUMAN
|
||||||
| θ value | 4.03905 (rank : 79) | NC score | 0.001038 (rank : 108) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 526 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P13645, Q14664, Q8N175 | Gene names | KRT10 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 10 (Cytokeratin-10) (CK-10) (Keratin-10) (K10). | |||||
|
NUPL1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 80) | NC score | 0.030278 (rank : 52) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BVL2, O43160 | Gene names | NUPL1, KIAA0410 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin p58/p45 (Nucleoporin-like 1). | |||||
|
PGCA_HUMAN
|
||||||
| θ value | 4.03905 (rank : 81) | NC score | 0.013045 (rank : 89) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P16112, Q13650, Q9UCP4, Q9UCP5, Q9UDE0 | Gene names | AGC1, CSPG1 | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP) (Chondroitin sulfate proteoglycan core protein 1) [Contains: Aggrecan core protein 2]. | |||||
|
SNPC4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 82) | NC score | 0.019255 (rank : 77) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q5SXM2, Q9Y6P7 | Gene names | SNAPC4, SNAP190 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | snRNA-activating protein complex subunit 4 (SNAPc subunit 4) (snRNA- activating protein complex 190 kDa subunit) (SNAPc 190 kDa subunit) (Proximal sequence element-binding transcription factor subunit alpha) (PSE-binding factor subunit alpha) (PTF subunit alpha). | |||||
|
AFF4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 83) | NC score | 0.007619 (rank : 97) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
ANX11_MOUSE
|
||||||
| θ value | 5.27518 (rank : 84) | NC score | 0.015945 (rank : 85) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P97384 | Gene names | Anxa11, Anx11 | |||
|
Domain Architecture |
|
|||||
| Description | Annexin A11 (Annexin XI) (Calcyclin-associated annexin 50) (CAP-50). | |||||
|
ARI1A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 85) | NC score | 0.052908 (rank : 22) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
HORN_MOUSE
|
||||||
| θ value | 5.27518 (rank : 86) | NC score | 0.014783 (rank : 87) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8VHD8 | Gene names | Hrnr | |||
|
Domain Architecture |

|
|||||
| Description | Hornerin. | |||||
|
NCOA3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 87) | NC score | 0.009108 (rank : 95) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
PO6F2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 88) | NC score | 0.007063 (rank : 98) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 618 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P78424, P78425, Q75ME8, Q86UM6, Q9UDS7 | Gene names | POU6F2, RPF1 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 6, transcription factor 2 (Retina-derived POU-domain factor 1) (RPF-1). | |||||
|
RBM10_HUMAN
|
||||||
| θ value | 5.27518 (rank : 89) | NC score | 0.021080 (rank : 72) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P98175, Q14136 | Gene names | RBM10, DXS8237E, KIAA0122 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 10 (RNA-binding motif protein 10). | |||||
|
SALL2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 90) | NC score | 0.001658 (rank : 107) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 835 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y467, Q9Y4G1 | Gene names | SALL2, KIAA0360, SAL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 2 (Zinc finger protein SALL2) (HSal2). | |||||
|
TTDN1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 91) | NC score | 0.017171 (rank : 81) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D011 | Gene names | Ttdn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TTD non-photosensitive 1 protein homolog. | |||||
|
ZIC1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 92) | NC score | 0.000540 (rank : 110) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 716 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P46684 | Gene names | Zic1, Zic | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZIC 1 (Zinc finger protein of the cerebellum 1). | |||||
|
BCL9_HUMAN
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | 0.025690 (rank : 66) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
CBP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 94) | NC score | 0.018512 (rank : 78) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CUZD1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 95) | NC score | 0.006524 (rank : 100) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P70412, Q9CTZ7 | Gene names | Cuzd1, Itmap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CUB and zona pellucida-like domain-containing protein 1 precursor (Integral membrane-associated protein 1). | |||||
|
GP112_HUMAN
|
||||||
| θ value | 6.88961 (rank : 96) | NC score | 0.007745 (rank : 96) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8IZF6, Q86SM6 | Gene names | GPR112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 112. | |||||
|
OTOF_HUMAN
|
||||||
| θ value | 6.88961 (rank : 97) | NC score | 0.009321 (rank : 93) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9HC10, Q9HC08, Q9HC09, Q9Y650 | Gene names | OTOF, FER1L2 | |||
|
Domain Architecture |

|
|||||
| Description | Otoferlin (Fer-1-like protein 2). | |||||
|
OTOF_MOUSE
|
||||||
| θ value | 6.88961 (rank : 98) | NC score | 0.009235 (rank : 94) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ESF1, Q9ESF2 | Gene names | Otof, Fer1l2 | |||
|
Domain Architecture |

|
|||||
| Description | Otoferlin (Fer-1-like protein 2). | |||||
|
RGS6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 99) | NC score | 0.004182 (rank : 101) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z2H2, Q8BGI8 | Gene names | Rgs6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 6 (RGS6). | |||||
|
SYNPO_MOUSE
|
||||||
| θ value | 6.88961 (rank : 100) | NC score | 0.016495 (rank : 83) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8CC35, Q99JI0 | Gene names | Synpo, Kiaa1029 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin. | |||||
|
CDX1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 101) | NC score | 0.001832 (rank : 105) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 382 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P47902, Q4VAU4, Q9NYK8 | Gene names | CDX1 | |||
|
Domain Architecture |
|
|||||
| Description | Homeobox protein CDX-1 (Caudal-type homeobox protein 1). | |||||
|
CG1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 102) | NC score | 0.026642 (rank : 64) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q13495 | Gene names | CXorf6, CG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CG1 protein (F18). | |||||
|
CNOT4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 103) | NC score | 0.030248 (rank : 53) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O95628, O95339, O95627, Q8IYM7, Q8NCL0, Q9NPQ1, Q9NZN6 | Gene names | CNOT4, NOT4 | |||
|
Domain Architecture |
|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
GRIP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 104) | NC score | 0.002953 (rank : 104) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q925T6, Q8BLQ3, Q8C0T3, Q925T5, Q925T7 | Gene names | Grip1 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor-interacting protein 1 (GRIP1 protein). | |||||
|
MUC13_MOUSE
|
||||||
| θ value | 8.99809 (rank : 105) | NC score | 0.032820 (rank : 49) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P19467 | Gene names | Muc13, Ly64 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-13 precursor (Cell surface antigen 114/A10) (Lymphocyte antigen 64). | |||||
|
NMDE4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 106) | NC score | 0.003483 (rank : 103) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q03391 | Gene names | Grin2d | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 4 precursor (N-methyl D- aspartate receptor subtype 2D) (NR2D) (NMDAR2D). | |||||
|
SMAD4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 107) | NC score | 0.009431 (rank : 92) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P97471, Q9CW56 | Gene names | Smad4, Dpc4, Madh4 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 4 (SMAD 4) (Mothers against DPP homolog 4) (Deletion target in pancreatic carcinoma 4 homolog) (Smad4). | |||||
|
ZIM10_MOUSE
|
||||||
| θ value | 8.99809 (rank : 108) | NC score | 0.032167 (rank : 50) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 430 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6P1E1, Q6PDK9, Q6PF85, Q8BW47 | Gene names | Rai17, Zimp10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 17 (PIAS-like protein Zimp10). | |||||
|
RBP56_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.074553 (rank : 13) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q92804, Q92751 | Gene names | TAF15, RBP56, TAF2N | |||
|
Domain Architecture |

|
|||||
| Description | TATA-binding protein-associated factor 2N (RNA-binding protein 56) (TAFII68) (TAF(II)68). | |||||
|
TFG_HUMAN
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.057927 (rank : 21) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q92734, Q15656 | Gene names | TFG | |||
|
Domain Architecture |

|
|||||
| Description | Protein TFG (TRK-fused gene protein). | |||||
|
EYA1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 108 | |
| SwissProt Accessions | P97767, O08818 | Gene names | Eya1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
|
EYA1_HUMAN
|
||||||
| NC score | 0.989351 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q99502 | Gene names | EYA1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
|
EYA4_MOUSE
|
||||||
| NC score | 0.974209 (rank : 3) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Z191 | Gene names | Eya4 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 4 (EC 3.1.3.48). | |||||
|
EYA2_MOUSE
|
||||||
| NC score | 0.973625 (rank : 4) | θ value | 5.01279e-176 (rank : 5) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O08575, P97925 | Gene names | Eya2, Eab1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 2 (EC 3.1.3.48). | |||||
|
EYA2_HUMAN
|
||||||
| NC score | 0.971986 (rank : 5) | θ value | 3.5924e-174 (rank : 6) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O00167, Q5JSW8, Q96CV6, Q96H97, Q99503, Q99812, Q9BWF6, Q9H4S3, Q9H4S9, Q9NPZ4, Q9UIX7 | Gene names | EYA2, EAB1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 2 (EC 3.1.3.48). | |||||
|
EYA4_HUMAN
|
||||||
| NC score | 0.969381 (rank : 6) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O95677, O95464, O95679, Q8IW39, Q9NTR7 | Gene names | EYA4 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 4 (EC 3.1.3.48). | |||||
|
EYA3_MOUSE
|
||||||
| NC score | 0.962209 (rank : 7) | θ value | 9.94316e-116 (rank : 8) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P97480, P97768 | Gene names | Eya3 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 3 (EC 3.1.3.48). | |||||
|
EYA3_HUMAN
|
||||||
| NC score | 0.962100 (rank : 8) | θ value | 8.9516e-133 (rank : 7) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99504, O95463, Q99813 | Gene names | EYA3 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 3 (EC 3.1.3.48). | |||||
|
FUS_MOUSE
|
||||||
| NC score | 0.180836 (rank : 9) | θ value | 9.29e-05 (rank : 11) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P56959 | Gene names | Fus | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein FUS (Pigpen protein). | |||||
|
EWS_MOUSE
|
||||||
| NC score | 0.179623 (rank : 10) | θ value | 4.92598e-06 (rank : 10) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q61545 | Gene names | Ewsr1, Ews, Ewsh | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein EWS. | |||||
|
EWS_HUMAN
|
||||||
| NC score | 0.177632 (rank : 11) | θ value | 3.77169e-06 (rank : 9) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q01844, Q92635 | Gene names | EWSR1, EWS | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein EWS (EWS oncogene) (Ewing sarcoma breakpoint region 1 protein). | |||||
|
FUS_HUMAN
|
||||||
| NC score | 0.171648 (rank : 12) | θ value | 0.000121331 (rank : 12) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P35637 | Gene names | FUS, TLS | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein FUS (Oncogene FUS) (Oncogene TLS) (Translocated in liposarcoma protein) (POMp75) (75 kDa DNA-pairing protein). | |||||
|
RBP56_HUMAN
|
||||||
| NC score | 0.074553 (rank : 13) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q92804, Q92751 | Gene names | TAF15, RBP56, TAF2N | |||
|
Domain Architecture |

|
|||||
| Description | TATA-binding protein-associated factor 2N (RNA-binding protein 56) (TAFII68) (TAF(II)68). | |||||
|
NU214_HUMAN
|
||||||
| NC score | 0.074409 (rank : 14) | θ value | 0.21417 (rank : 24) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P35658 | Gene names | NUP214, CAIN, CAN | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup214 (Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein). | |||||
|
HNRL1_MOUSE
|
||||||
| NC score | 0.065391 (rank : 15) | θ value | 0.813845 (rank : 39) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8VDM6, Q3U201, Q3UPB0, Q6AZA7, Q8BY45, Q8K365 | Gene names | Hnrpul1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein U-like protein 1. | |||||
|
HNRL1_HUMAN
|
||||||
| NC score | 0.063898 (rank : 16) | θ value | 1.06291 (rank : 42) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9BUJ2, O76022, Q6ZSZ0, Q7L8P4, Q8N6Z4, Q96G37, Q9HAL3, Q9UG75 | Gene names | HNRPUL1, E1BAP5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein U-like protein 1 (Adenovirus early region 1B-associated protein 5) (E1B-55 kDa-associated protein 5) (E1B-AP5). | |||||
|
RPB1_MOUSE
|
||||||
| NC score | 0.063456 (rank : 17) | θ value | 2.36792 (rank : 67) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P08775 | Gene names | Polr2a, Rpii215, Rpo2-1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase II largest subunit (EC 2.7.7.6) (RPB1). | |||||
|
RPB1_HUMAN
|
||||||
| NC score | 0.063368 (rank : 18) | θ value | 2.36792 (rank : 66) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P24928 | Gene names | POLR2A | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase II largest subunit (EC 2.7.7.6) (RPB1). | |||||
|
BAG4_MOUSE
|
||||||
| NC score | 0.060937 (rank : 19) | θ value | 0.163984 (rank : 22) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8CI61, Q3TRL9, Q91VT5, Q9CWG2 | Gene names | Bag4, Sodd | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 4 (BCL2-associated athanogene 4) (BAG-4) (Silencer of death domains). | |||||
|
MCM3A_MOUSE
|
||||||
| NC score | 0.059528 (rank : 20) | θ value | 0.62314 (rank : 33) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9WUU9 | Gene names | Mcm3ap, Ganp, Map80 | |||
|
Domain Architecture |

|
|||||
| Description | 80 kDa MCM3-associated protein (GANP protein). | |||||
|
TFG_HUMAN
|
||||||
| NC score | 0.057927 (rank : 21) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q92734, Q15656 | Gene names | TFG | |||
|
Domain Architecture |

|
|||||
| Description | Protein TFG (TRK-fused gene protein). | |||||
|
ARI1A_HUMAN
|
||||||
| NC score | 0.052908 (rank : 22) | θ value | 5.27518 (rank : 85) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
BAG4_HUMAN
|
||||||
| NC score | 0.049833 (rank : 23) | θ value | 1.81305 (rank : 49) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O95429, O95818 | Gene names | BAG4, SODD | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 4 (BCL2-associated athanogene 4) (BAG-4) (Silencer of death domains). | |||||
|
SSXT_MOUSE
|
||||||
| NC score | 0.047858 (rank : 24) | θ value | 0.21417 (rank : 26) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q62280 | Gene names | Ss18, Ssxt, Syt | |||
|
Domain Architecture |

|
|||||
| Description | SSXT protein (SYT protein) (Synovial sarcoma-associated Ss18-alpha). | |||||
|
ILF3_MOUSE
|
||||||
| NC score | 0.045885 (rank : 25) | θ value | 0.47712 (rank : 30) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Z1X4, Q80VD5, Q812A1, Q8BP80, Q8C2H8, Q8K588 | Gene names | Ilf3 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin enhancer-binding factor 3. | |||||
|
ZIM10_HUMAN
|
||||||
| NC score | 0.045497 (rank : 26) | θ value | 0.47712 (rank : 31) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9ULJ6, Q7Z7E6 | Gene names | RAI17, KIAA1224, ZIMP10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 17 (PIAS-like protein Zimp10). | |||||
|
RBM14_MOUSE
|
||||||
| NC score | 0.045300 (rank : 27) | θ value | 0.0431538 (rank : 17) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8C2Q3, Q3TJB6, Q91Z21, Q9DBI6 | Gene names | Rbm14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 14 (RNA-binding motif protein 14). | |||||
|
ATN1_HUMAN
|
||||||
| NC score | 0.044555 (rank : 28) | θ value | 2.36792 (rank : 64) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
ZIMP7_MOUSE
|
||||||
| NC score | 0.044137 (rank : 29) | θ value | 0.125558 (rank : 21) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 566 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8CIE2, Q6P558, Q6P9Q5, Q8BJ00 | Gene names | Zimp7, D11Bwg0280e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PIAS-like protein Zimp7. | |||||
|
PUM2_HUMAN
|
||||||
| NC score | 0.043486 (rank : 30) | θ value | 0.62314 (rank : 36) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TB72, O00234, Q53TV7, Q8WY43, Q9HAN2 | Gene names | PUM2, KIAA0235, PUMH2 | |||
|
Domain Architecture |

|
|||||
| Description | Pumilio homolog 2 (Pumilio-2). | |||||
|
PUM2_MOUSE
|
||||||
| NC score | 0.039967 (rank : 31) | θ value | 1.81305 (rank : 61) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80U58, Q80UZ9, Q91YW4, Q925A0, Q9ERC7 | Gene names | Pum2, Kiaa0235 | |||
|
Domain Architecture |

|
|||||
| Description | Pumilio homolog 2. | |||||
|
MUCDL_HUMAN
|
||||||
| NC score | 0.039588 (rank : 32) | θ value | 0.0431538 (rank : 16) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9HBB8, Q9H746, Q9HAU3, Q9HBB5, Q9HBB6, Q9HBB7, Q9NX86, Q9NXI9 | Gene names | MUCDHL | |||
|
Domain Architecture |

|
|||||
| Description | Mucin and cadherin-like protein precursor (Mu-protocadherin). | |||||
|
PPRB_HUMAN
|
||||||
| NC score | 0.039301 (rank : 33) | θ value | 0.21417 (rank : 25) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15648, O43810, O75447, Q9HD39 | Gene names | PPARBP, ARC205, DRIP205, TRAP220, TRIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2) (p53 regulatory protein RB18A) (Vitamin D receptor-interacting protein complex component DRIP205) (Activator- recruited cofactor 205 kDa component) (ARC205). | |||||
|
ARI1B_HUMAN
|
||||||
| NC score | 0.039268 (rank : 34) | θ value | 1.06291 (rank : 41) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 567 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8NFD5, Q5JRD1, Q5VYC4, Q8IZY8, Q8TEV0, Q8TF02, Q99491, Q9ULI5 | Gene names | ARID1B, DAN15, KIAA1235, OSA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-rich interactive domain-containing protein 1B (ARID domain- containing protein 1B) (Osa homolog 2) (hOsa2) (p250R) (BRG1-binding protein hELD/OSA1) (BRG1-associated factor 250b) (BAF250B). | |||||
|
TRIM2_MOUSE
|
||||||
| NC score | 0.039264 (rank : 35) | θ value | 0.0252991 (rank : 14) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9ESN6, Q3UHP5, Q8C981 | Gene names | Trim2, Narf | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 2 (Neural activity-related RING finger protein). | |||||
|
TRIM2_HUMAN
|
||||||
| NC score | 0.039154 (rank : 36) | θ value | 0.0252991 (rank : 13) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9C040, O60272, Q9BSI9, Q9UFZ1 | Gene names | TRIM2, KIAA0517, RNF86 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 2 (RING finger protein 86). | |||||
|
DDX17_HUMAN
|
||||||
| NC score | 0.038763 (rank : 37) | θ value | 0.0736092 (rank : 19) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q92841, Q69YT1, Q6ICD6 | Gene names | DDX17 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DDX17 (EC 3.6.1.-) (DEAD box protein 17) (RNA-dependent helicase p72) (DEAD box protein p72). | |||||
|
SOX8_MOUSE
|
||||||
| NC score | 0.038482 (rank : 38) | θ value | 0.0563607 (rank : 18) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q04886 | Gene names | Sox8, Sox-8 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-8. | |||||
|
CNOT4_MOUSE
|
||||||
| NC score | 0.037790 (rank : 39) | θ value | 0.163984 (rank : 23) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BT14, Q8CCR4, Q9CV74, Q9Z1D0 | Gene names | Cnot4, Not4 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
NUPL1_MOUSE
|
||||||
| NC score | 0.037150 (rank : 40) | θ value | 0.62314 (rank : 34) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8R332, Q3UJF4, Q8BUA7, Q8BVG7, Q8C0W3 | Gene names | Nupl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin p58/p45 (Nucleoporin-like 1). | |||||
|
DAZ4_HUMAN
|
||||||
| NC score | 0.036645 (rank : 41) | θ value | 1.81305 (rank : 51) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86SG3, Q9NR88, Q9NR89 | Gene names | DAZ4 | |||
|
Domain Architecture |

|
|||||
| Description | Deleted in azoospermia protein 4. | |||||
|
DAZ1_HUMAN
|
||||||
| NC score | 0.036601 (rank : 42) | θ value | 1.81305 (rank : 50) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9NQZ3, Q9NQZ4 | Gene names | DAZ1, DAZ | |||
|
Domain Architecture |

|
|||||
| Description | Deleted in azoospermia protein 1. | |||||
|
DAZ2_HUMAN
|
||||||
| NC score | 0.036570 (rank : 43) | θ value | 1.38821 (rank : 43) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q13117, Q96P41, Q9NR91 | Gene names | DAZ2 | |||
|
Domain Architecture |

|
|||||
| Description | Deleted in azoospermia protein 2. | |||||
|
ILF3_HUMAN
|
||||||
| NC score | 0.035775 (rank : 44) | θ value | 0.813845 (rank : 40) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q12906, O43409, Q6P1X1, Q86XY7, Q99544, Q99545, Q9BZH4, Q9BZH5, Q9NQ95, Q9NQ96, Q9NQ97, Q9NQ98, Q9NQ99, Q9NQA0, Q9NQA1, Q9NQA2, Q9NRN2, Q9NRN3, Q9NRN4, Q9UMZ9, Q9UN00, Q9UN84, Q9UNA2 | Gene names | ILF3, DRBF, MPHOSPH4, NF90 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin enhancer-binding factor 3 (Nuclear factor of activated T- cells 90 kDa) (NF-AT-90) (Double-stranded RNA-binding protein 76) (DRBP76) (Translational control protein 80) (TCP80) (Nuclear factor associated with dsRNA) (NFAR) (M-phase phosphoprotein 4) (MPP4). | |||||
|
PPRB_MOUSE
|
||||||
| NC score | 0.035659 (rank : 45) | θ value | 1.81305 (rank : 60) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q925J9, O88323, Q3UHV0, Q6AXD5, Q8BW37, Q8BX19, Q8VDQ7, Q925K0 | Gene names | Pparbp, Trip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2). | |||||
|
S18L1_HUMAN
|
||||||
| NC score | 0.033802 (rank : 46) | θ value | 3.0926 (rank : 71) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75177, Q5JXJ3, Q8NE69, Q9BR55, Q9H4K6 | Gene names | SS18L1, KIAA0693 | |||
|
Domain Architecture |

|
|||||
| Description | SS18-like protein 1 (SYT homolog 1). | |||||
|
DAZ3_HUMAN
|
||||||
| NC score | 0.033743 (rank : 47) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NR90 | Gene names | DAZ3 | |||
|
Domain Architecture |

|
|||||
| Description | Deleted in azoospermia protein 3. | |||||
|
DDX17_MOUSE
|
||||||
| NC score | 0.033471 (rank : 48) | θ value | 0.279714 (rank : 27) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q501J6, Q6P5G1, Q8BIN2 | Gene names | Ddx17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX17 (EC 3.6.1.-) (DEAD box protein 17). | |||||
|
MUC13_MOUSE
|
||||||
| NC score | 0.032820 (rank : 49) | θ value | 8.99809 (rank : 105) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P19467 | Gene names | Muc13, Ly64 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-13 precursor (Cell surface antigen 114/A10) (Lymphocyte antigen 64). | |||||
|
ZIM10_MOUSE
|
||||||
| NC score | 0.032167 (rank : 50) | θ value | 8.99809 (rank : 108) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 430 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6P1E1, Q6PDK9, Q6PF85, Q8BW47 | Gene names | Rai17, Zimp10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 17 (PIAS-like protein Zimp10). | |||||
|
DDX3Y_HUMAN
|
||||||
| NC score | 0.031576 (rank : 51) | θ value | 0.47712 (rank : 29) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O15523, Q8IYV7 | Gene names | DDX3Y, DBY | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX3Y (EC 3.6.1.-) (DEAD box protein 3, Y- chromosomal). | |||||
|
NUPL1_HUMAN
|
||||||
| NC score | 0.030278 (rank : 52) | θ value | 4.03905 (rank : 80) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BVL2, O43160 | Gene names | NUPL1, KIAA0410 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin p58/p45 (Nucleoporin-like 1). | |||||
|
CNOT4_HUMAN
|
||||||
| NC score | 0.030248 (rank : 53) | θ value | 8.99809 (rank : 103) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O95628, O95339, O95627, Q8IYM7, Q8NCL0, Q9NPQ1, Q9NZN6 | Gene names | CNOT4, NOT4 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
RNF12_HUMAN
|
||||||
| NC score | 0.030142 (rank : 54) | θ value | 1.81305 (rank : 63) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NVW2, Q9Y598 | Gene names | RNF12, RLIM | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 12 (LIM domain-interacting RING finger protein) (RING finger LIM domain-binding protein) (R-LIM) (NY-REN-43 antigen). | |||||
|
DDX3X_HUMAN
|
||||||
| NC score | 0.030033 (rank : 55) | θ value | 0.62314 (rank : 32) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O00571, O15536 | Gene names | DDX3X, DBX, DDX3 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX3X (EC 3.6.1.-) (DEAD box protein 3, X- chromosomal) (Helicase-like protein 2) (HLP2) (DEAD box, X isoform). | |||||
|
PL10_MOUSE
|
||||||
| NC score | 0.029840 (rank : 56) | θ value | 0.62314 (rank : 35) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P16381 | Gene names | D1Pas1, Pl10 | |||
|
Domain Architecture |

|
|||||
| Description | Putative ATP-dependent RNA helicase Pl10 (EC 3.6.1.-). | |||||
|
DDX3Y_MOUSE
|
||||||
| NC score | 0.028855 (rank : 57) | θ value | 1.38821 (rank : 44) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q62095, Q9QWS9 | Gene names | Ddx3y, D1Pas1-rs1, Dead2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX3Y (EC 3.6.1.-) (DEAD box protein 3, Y- chromosomal) (DEAD-box RNA helicase DEAD2) (mDEAD2) (D1Pas1-related sequence 1). | |||||
|
NUP98_HUMAN
|
||||||
| NC score | 0.028426 (rank : 58) | θ value | 1.81305 (rank : 57) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P52948, Q8IUT2, Q8WYB0, Q96E54, Q9H3Q4, Q9NT02, Q9UF57, Q9UHX0, Q9Y6J4, Q9Y6J5 | Gene names | NUP98 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup98-Nup96 precursor [Contains: Nuclear pore complex protein Nup98 (Nucleoporin Nup98) (98 kDa nucleoporin); Nuclear pore complex protein Nup96 (Nucleoporin Nup96) (96 kDa nucleoporin)]. | |||||
|
CDSN_HUMAN
|
||||||
| NC score | 0.028030 (rank : 59) | θ value | 3.0926 (rank : 69) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q15517, O43509, Q5SQ85, Q5STD2, Q7LA70, Q7LA71, Q86Z04, Q8IZU4, Q8IZU5, Q8IZU6, Q8N5P3, Q95IF9, Q9NP52, Q9NPE0, Q9NPG5, Q9NRH4, Q9NRH5, Q9NRH6, Q9NRH7, Q9NRH8, Q9UBH8, Q9UIN6, Q9UIN7, Q9UIN8, Q9UIN9, Q9UIP0 | Gene names | CDSN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Corneodesmosin precursor (S protein). | |||||
|
SOX8_HUMAN
|
||||||
| NC score | 0.027916 (rank : 60) | θ value | 0.0330416 (rank : 15) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P57073, Q9NZW2 | Gene names | SOX8 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-8. | |||||
|
GATA4_HUMAN
|
||||||
| NC score | 0.027625 (rank : 61) | θ value | 0.365318 (rank : 28) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P43694 | Gene names | GATA4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor GATA-4 (GATA-binding factor 4). | |||||
|
SOX30_MOUSE
|
||||||
| NC score | 0.027544 (rank : 62) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8CGW4 | Gene names | Sox30 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-30. | |||||
|
DDX3X_MOUSE
|
||||||
| NC score | 0.027481 (rank : 63) | θ value | 1.81305 (rank : 52) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q62167, O09060, O09143 | Gene names | Ddx3x, D1Pas1-rs2, Ddx3, Dead3, Erh | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX3X (EC 3.6.1.-) (DEAD box protein 3, X- chromosomal) (DEAD box RNA helicase DEAD3) (mDEAD3) (Embryonic RNA helicase) (D1Pas1-related sequence 2). | |||||
|
CG1_HUMAN
|
||||||
| NC score | 0.026642 (rank : 64) | θ value | 8.99809 (rank : 102) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q13495 | Gene names | CXorf6, CG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CG1 protein (F18). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.026341 (rank : 65) | θ value | 1.81305 (rank : 55) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
BCL9_HUMAN
|
||||||
| NC score | 0.025690 (rank : 66) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
CBP_MOUSE
|
||||||
| NC score | 0.025578 (rank : 67) | θ value | 2.36792 (rank : 65) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
YLPM1_MOUSE
|
||||||
| NC score | 0.025570 (rank : 68) | θ value | 3.0926 (rank : 74) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
PDC6I_HUMAN
|
||||||
| NC score | 0.024092 (rank : 69) | θ value | 1.81305 (rank : 58) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8WUM4, Q9BX86, Q9NUN0, Q9P2H2, Q9UKL5 | Gene names | PDCD6IP, AIP1, ALIX, KIAA1375 | |||
|
Domain Architecture |

|
|||||
| Description | Programmed cell death 6-interacting protein (PDCD6-interacting protein) (ALG-2-interacting protein 1) (Hp95). | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.023617 (rank : 70) | θ value | 1.81305 (rank : 56) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
RERE_MOUSE
|
||||||
| NC score | 0.023594 (rank : 71) | θ value | 1.81305 (rank : 62) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
RBM10_HUMAN
|
||||||
| NC score | 0.021080 (rank : 72) | θ value | 5.27518 (rank : 89) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P98175, Q14136 | Gene names | RBM10, DXS8237E, KIAA0122 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 10 (RNA-binding motif protein 10). | |||||
|
HRBL_HUMAN
|
||||||
| NC score | 0.020675 (rank : 73) | θ value | 4.03905 (rank : 77) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95081, O75429, Q96AB9, Q96GL4 | Gene names | HRBL, RABR | |||
|
Domain Architecture |

|
|||||
| Description | HIV-1 Rev-binding protein-like protein (Rev/Rex activation domain- binding protein related) (RAB-R). | |||||
|
HTF4_HUMAN
|
||||||
| NC score | 0.020627 (rank : 74) | θ value | 1.81305 (rank : 53) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q99081 | Gene names | TCF12, HEB, HTF4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 12 (Transcription factor HTF-4) (E-box-binding protein) (DNA-binding protein HTF4). | |||||
|
HTF4_MOUSE
|
||||||
| NC score | 0.020511 (rank : 75) | θ value | 1.81305 (rank : 54) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q61286 | Gene names | Tcf12, Alf1, Meb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 12 (Transcription factor HTF-4) (E-box-binding protein) (DNA-binding protein HTF4) (Class A helix-loop-helix transcription factor ME1). | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.019499 (rank : 76) | θ value | 1.38821 (rank : 46) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
SNPC4_HUMAN
|
||||||
| NC score | 0.019255 (rank : 77) | θ value | 4.03905 (rank : 82) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q5SXM2, Q9Y6P7 | Gene names | SNAPC4, SNAP190 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | snRNA-activating protein complex subunit 4 (SNAPc subunit 4) (snRNA- activating protein complex 190 kDa subunit) (SNAPc 190 kDa subunit) (Proximal sequence element-binding transcription factor subunit alpha) (PSE-binding factor subunit alpha) (PTF subunit alpha). | |||||
|
CBP_HUMAN
|
||||||
| NC score | 0.018512 (rank : 78) | θ value | 6.88961 (rank : 94) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
K1210_HUMAN
|
||||||
| NC score | 0.018428 (rank : 79) | θ value | 4.03905 (rank : 78) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ULL0, Q5JPN4 | Gene names | KIAA1210 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1210. | |||||
|
HORN_HUMAN
|
||||||
| NC score | 0.018159 (rank : 80) | θ value | 1.38821 (rank : 45) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q86YZ3 | Gene names | HRNR, S100A18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hornerin. | |||||
|
TTDN1_MOUSE
|
||||||
| NC score | 0.017171 (rank : 81) | θ value | 5.27518 (rank : 91) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D011 | Gene names | Ttdn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TTD non-photosensitive 1 protein homolog. | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.017141 (rank : 82) | θ value | 0.62314 (rank : 37) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
SYNPO_MOUSE
|
||||||
| NC score | 0.016495 (rank : 83) | θ value | 6.88961 (rank : 100) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8CC35, Q99JI0 | Gene names | Synpo, Kiaa1029 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin. | |||||
|
RUNX1_MOUSE
|
||||||
| NC score | 0.016163 (rank : 84) | θ value | 1.38821 (rank : 47) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q03347, O08598, Q62049, Q9ESB9, Q9ET65 | Gene names | Runx1, Aml1, Cbfa2, Pebp2ab | |||
|
Domain Architecture |

|
|||||
| Description | Runt-related transcription factor 1 (Core-binding factor, alpha 2 subunit) (CBF-alpha 2) (Acute myeloid leukemia 1 protein) (Oncogene AML-1) (Polyomavirus enhancer-binding protein 2 alpha B subunit) (PEBP2-alpha B) (PEA2-alpha B) (SL3-3 enhancer factor 1 alpha B subunit) (SL3/AKV core-binding factor alpha B subunit). | |||||
|
ANX11_MOUSE
|
||||||
| NC score | 0.015945 (rank : 85) | θ value | 5.27518 (rank : 84) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P97384 | Gene names | Anxa11, Anx11 | |||
|
Domain Architecture |

|
|||||
| Description | Annexin A11 (Annexin XI) (Calcyclin-associated annexin 50) (CAP-50). | |||||
|
ANXA7_HUMAN
|
||||||
| NC score | 0.015425 (rank : 86) | θ value | 1.81305 (rank : 48) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P20073 | Gene names | ANXA7, ANX7, SNX | |||
|
Domain Architecture |

|
|||||
| Description | Annexin A7 (Annexin VII) (Synexin). | |||||
|
HORN_MOUSE
|
||||||
| NC score | 0.014783 (rank : 87) | θ value | 5.27518 (rank : 86) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8VHD8 | Gene names | Hrnr | |||
|
Domain Architecture |

|
|||||
| Description | Hornerin. | |||||
|
TAB2_MOUSE
|
||||||
| NC score | 0.014611 (rank : 88) | θ value | 3.0926 (rank : 73) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q99K90, Q3UGP1, Q8BTP4, Q8CHD3, Q99KP4 | Gene names | Map3k7ip2, Kiaa0733, Tab2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 7-interacting protein 2 (TAK1-binding protein 2) (TAB2). | |||||
|
PGCA_HUMAN
|
||||||
| NC score | 0.013045 (rank : 89) | θ value | 4.03905 (rank : 81) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P16112, Q13650, Q9UCP4, Q9UCP5, Q9UDE0 | Gene names | AGC1, CSPG1 | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP) (Chondroitin sulfate proteoglycan core protein 1) [Contains: Aggrecan core protein 2]. | |||||
|
PEF1_MOUSE
|
||||||
| NC score | 0.012487 (rank : 90) | θ value | 1.81305 (rank : 59) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BFY6, Q8VCT5, Q9CYW8, Q9D934 | Gene names | Pef1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peflin (PEF protein with a long N-terminal hydrophobic domain) (Penta- EF hand domain-containing protein 1). | |||||
|
SIRT3_HUMAN
|
||||||
| NC score | 0.011224 (rank : 91) | θ value | 3.0926 (rank : 72) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NTG7, Q9Y6E8 | Gene names | SIRT3, SIR2L3 | |||
|
Domain Architecture |

|
|||||
| Description | NAD-dependent deacetylase sirtuin-3, mitochondrial precursor (EC 3.5.1.-) (SIR2-like protein 3) (hSIRT3). | |||||
|
SMAD4_MOUSE
|
||||||
| NC score | 0.009431 (rank : 92) | θ value | 8.99809 (rank : 107) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P97471, Q9CW56 | Gene names | Smad4, Dpc4, Madh4 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 4 (SMAD 4) (Mothers against DPP homolog 4) (Deletion target in pancreatic carcinoma 4 homolog) (Smad4). | |||||
|
OTOF_HUMAN
|
||||||
| NC score | 0.009321 (rank : 93) | θ value | 6.88961 (rank : 97) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9HC10, Q9HC08, Q9HC09, Q9Y650 | Gene names | OTOF, FER1L2 | |||
|
Domain Architecture |

|
|||||
| Description | Otoferlin (Fer-1-like protein 2). | |||||
|
OTOF_MOUSE
|
||||||
| NC score | 0.009235 (rank : 94) | θ value | 6.88961 (rank : 98) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ESF1, Q9ESF2 | Gene names | Otof, Fer1l2 | |||
|
Domain Architecture |

|
|||||
| Description | Otoferlin (Fer-1-like protein 2). | |||||
|
NCOA3_MOUSE
|
||||||
| NC score | 0.009108 (rank : 95) | θ value | 5.27518 (rank : 87) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
GP112_HUMAN
|
||||||
| NC score | 0.007745 (rank : 96) | θ value | 6.88961 (rank : 96) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8IZF6, Q86SM6 | Gene names | GPR112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 112. | |||||
|
AFF4_HUMAN
|
||||||
| NC score | 0.007619 (rank : 97) | θ value | 5.27518 (rank : 83) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
PO6F2_HUMAN
|
||||||
| NC score | 0.007063 (rank : 98) | θ value | 5.27518 (rank : 88) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 618 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P78424, P78425, Q75ME8, Q86UM6, Q9UDS7 | Gene names | POU6F2, RPF1 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 6, transcription factor 2 (Retina-derived POU-domain factor 1) (RPF-1). | |||||
|
ETV3_HUMAN
|
||||||
| NC score | 0.006900 (rank : 99) | θ value | 4.03905 (rank : 76) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
CUZD1_MOUSE
|
||||||
| NC score | 0.006524 (rank : 100) | θ value | 6.88961 (rank : 95) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P70412, Q9CTZ7 | Gene names | Cuzd1, Itmap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CUB and zona pellucida-like domain-containing protein 1 precursor (Integral membrane-associated protein 1). | |||||
|
RGS6_MOUSE
|
||||||
| NC score | 0.004182 (rank : 101) | θ value | 6.88961 (rank : 99) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z2H2, Q8BGI8 | Gene names | Rgs6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 6 (RGS6). | |||||
|
HXA5_HUMAN
|
||||||
| NC score | 0.004107 (rank : 102) | θ value | 3.0926 (rank : 70) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 336 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P20719, O43367, Q96CY6 | Gene names | HOXA5, HOX1C | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-A5 (Hox-1C). | |||||
|
NMDE4_MOUSE
|
||||||
| NC score | 0.003483 (rank : 103) | θ value | 8.99809 (rank : 106) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q03391 | Gene names | Grin2d | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 4 precursor (N-methyl D- aspartate receptor subtype 2D) (NR2D) (NMDAR2D). | |||||
|
GRIP1_MOUSE
|
||||||
| NC score | 0.002953 (rank : 104) | θ value | 8.99809 (rank : 104) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q925T6, Q8BLQ3, Q8C0T3, Q925T5, Q925T7 | Gene names | Grip1 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor-interacting protein 1 (GRIP1 protein). | |||||
|
CDX1_HUMAN
|
||||||
| NC score | 0.001832 (rank : 105) | θ value | 8.99809 (rank : 101) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 382 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P47902, Q4VAU4, Q9NYK8 | Gene names | CDX1 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein CDX-1 (Caudal-type homeobox protein 1). | |||||
|
ZBT16_HUMAN
|
||||||
| NC score | 0.001693 (rank : 106) | θ value | 0.125558 (rank : 20) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 1010 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q05516, Q8TAL4 | Gene names | ZBTB16, PLZF, ZNF145 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 16 (Zinc finger protein PLZF) (Promyelocytic leukemia zinc finger protein) (Zinc finger protein 145). | |||||
|
SALL2_HUMAN
|
||||||
| NC score | 0.001658 (rank : 107) | θ value | 5.27518 (rank : 90) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 835 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y467, Q9Y4G1 | Gene names | SALL2, KIAA0360, SAL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 2 (Zinc finger protein SALL2) (HSal2). | |||||
|
K1C10_HUMAN
|
||||||
| NC score | 0.001038 (rank : 108) | θ value | 4.03905 (rank : 79) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 526 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P13645, Q14664, Q8N175 | Gene names | KRT10 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 10 (Cytokeratin-10) (CK-10) (Keratin-10) (K10). | |||||
|
ZN560_HUMAN
|
||||||
| NC score | 0.000891 (rank : 109) | θ value | 0.62314 (rank : 38) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 751 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96MR9, Q495S9, Q495T1 | Gene names | ZNF560 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 560. | |||||
|
ZIC1_MOUSE
|
||||||
| NC score | 0.000540 (rank : 110) | θ value | 5.27518 (rank : 92) | |||
| Query Neighborhood Hits | 108 | Target Neighborhood Hits | 716 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P46684 | Gene names | Zic1, Zic | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZIC 1 (Zinc finger protein of the cerebellum 1). | |||||