Please be patient as the page loads
|
BAG4_HUMAN
|
||||||
| SwissProt Accessions | O95429, O95818 | Gene names | BAG4, SODD | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 4 (BCL2-associated athanogene 4) (BAG-4) (Silencer of death domains). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
BAG4_HUMAN
|
||||||
| θ value | 4.24358e-175 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 96 | |
| SwissProt Accessions | O95429, O95818 | Gene names | BAG4, SODD | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 4 (BCL2-associated athanogene 4) (BAG-4) (Silencer of death domains). | |||||
|
BAG4_MOUSE
|
||||||
| θ value | 2.12569e-142 (rank : 2) | NC score | 0.956571 (rank : 2) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8CI61, Q3TRL9, Q91VT5, Q9CWG2 | Gene names | Bag4, Sodd | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 4 (BCL2-associated athanogene 4) (BAG-4) (Silencer of death domains). | |||||
|
BAG3_HUMAN
|
||||||
| θ value | 6.62687e-19 (rank : 3) | NC score | 0.547145 (rank : 3) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O95817, Q9NT20, Q9P120 | Gene names | BAG3, BIS | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis) (Docking protein CAIR-1). | |||||
|
BAG3_MOUSE
|
||||||
| θ value | 6.62687e-19 (rank : 4) | NC score | 0.505563 (rank : 6) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9JLV1, Q9JJC7 | Gene names | Bag3, Bis | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis). | |||||
|
BAG5_HUMAN
|
||||||
| θ value | 1.9326e-10 (rank : 5) | NC score | 0.508522 (rank : 4) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UL15, O94950, Q86W59 | Gene names | BAG5, KIAA0873 | |||
|
Domain Architecture |
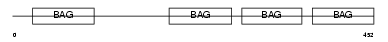
|
|||||
| Description | BAG family molecular chaperone regulator 5 (BCL2-associated athanogene 5) (BAG-5). | |||||
|
BAG5_MOUSE
|
||||||
| θ value | 7.34386e-10 (rank : 6) | NC score | 0.505999 (rank : 5) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CI32, Q3TVA8, Q8K175, Q9DAU0 | Gene names | Bag5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BAG family molecular chaperone regulator 5 (BCL2-associated athanogene 5) (BAG-5). | |||||
|
PRRT3_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 7) | NC score | 0.106771 (rank : 7) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
FMN2_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 8) | NC score | 0.058236 (rank : 19) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
MAGE1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 9) | NC score | 0.060300 (rank : 18) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
SPEG_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 10) | NC score | 0.024309 (rank : 66) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
CTND2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 11) | NC score | 0.033838 (rank : 50) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UQB3, O00379, O15390, O43206, O43840, Q13589, Q9UM66, Q9UPM3 | Gene names | CTNND2, NPRAP | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Delta-catenin) (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin) (GT24). | |||||
|
SSXT_HUMAN
|
||||||
| θ value | 0.163984 (rank : 12) | NC score | 0.103460 (rank : 8) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15532, Q16404, Q9BXC6 | Gene names | SS18, SSXT, SYT | |||
|
Domain Architecture |
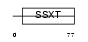
|
|||||
| Description | SSXT protein (Synovial sarcoma, translocated to X chromosome) (SYT protein). | |||||
|
CO4A3_HUMAN
|
||||||
| θ value | 0.21417 (rank : 13) | NC score | 0.023156 (rank : 68) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 421 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q01955, Q9BQT2 | Gene names | COL4A3 | |||
|
Domain Architecture |
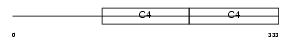
|
|||||
| Description | Collagen alpha-3(IV) chain precursor (Goodpasture antigen). | |||||
|
SSXT_MOUSE
|
||||||
| θ value | 0.21417 (rank : 14) | NC score | 0.102532 (rank : 9) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q62280 | Gene names | Ss18, Ssxt, Syt | |||
|
Domain Architecture |
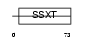
|
|||||
| Description | SSXT protein (SYT protein) (Synovial sarcoma-associated Ss18-alpha). | |||||
|
DAB1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 15) | NC score | 0.047192 (rank : 36) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75553, Q9NYA8 | Gene names | DAB1 | |||
|
Domain Architecture |
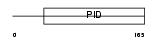
|
|||||
| Description | Disabled homolog 1. | |||||
|
IASPP_MOUSE
|
||||||
| θ value | 0.279714 (rank : 16) | NC score | 0.037957 (rank : 47) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1024 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q5I1X5 | Gene names | Ppp1r13l, Nkip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
REXO1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 17) | NC score | 0.050401 (rank : 32) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8N1G1, Q9ULT2 | Gene names | REXO1, ELOABP1, KIAA1138, TCEB3BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA exonuclease 1 homolog (EC 3.1.-.-) (Elongin A-binding protein 1) (EloA-BP1) (Transcription elongation factor B polypeptide 3-binding protein 1). | |||||
|
ARI1A_HUMAN
|
||||||
| θ value | 0.47712 (rank : 18) | NC score | 0.070488 (rank : 15) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
LEF1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 19) | NC score | 0.041027 (rank : 41) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UJU2, Q9HAZ0 | Gene names | LEF1 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphoid enhancer-binding factor 1 (LEF-1) (T cell-specific transcription factor 1-alpha) (TCF1-alpha). | |||||
|
LEF1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 20) | NC score | 0.040942 (rank : 42) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P27782 | Gene names | Lef1, Lef-1 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphoid enhancer-binding factor 1 (LEF-1). | |||||
|
TULP4_HUMAN
|
||||||
| θ value | 0.62314 (rank : 21) | NC score | 0.042833 (rank : 40) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9NRJ4, Q9HD22, Q9P2F0 | Gene names | TULP4, KIAA1397, TUSP | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-like protein 4 (Tubby superfamily protein). | |||||
|
CEL_HUMAN
|
||||||
| θ value | 0.813845 (rank : 22) | NC score | 0.043540 (rank : 39) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
EYA1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 23) | NC score | 0.046936 (rank : 37) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99502 | Gene names | EYA1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
|
IL3RB_HUMAN
|
||||||
| θ value | 1.06291 (rank : 24) | NC score | 0.048055 (rank : 35) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P32927 | Gene names | CSF2RB, IL3RB, IL5RB | |||
|
Domain Architecture |

|
|||||
| Description | Cytokine receptor common beta chain precursor (GM-CSF/IL-3/IL-5 receptor common beta-chain) (CD131 antigen) (CDw131). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 25) | NC score | 0.052007 (rank : 29) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
YLPM1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 26) | NC score | 0.057536 (rank : 21) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
DYN1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 27) | NC score | 0.032841 (rank : 53) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P39053, Q61358, Q61359, Q61360, Q8JZZ4, Q9CSY7, Q9QXX1 | Gene names | Dnm1, Dnm | |||
|
Domain Architecture |

|
|||||
| Description | Dynamin-1 (EC 3.6.5.5). | |||||
|
SPEG_HUMAN
|
||||||
| θ value | 1.38821 (rank : 28) | NC score | 0.019586 (rank : 72) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
ZN750_HUMAN
|
||||||
| θ value | 1.38821 (rank : 29) | NC score | 0.037954 (rank : 48) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q32MQ0, Q9H899 | Gene names | ZNF750 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ZNF750. | |||||
|
EYA1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 30) | NC score | 0.049833 (rank : 33) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P97767, O08818 | Gene names | Eya1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
|
GGNB1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 31) | NC score | 0.076058 (rank : 13) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5YKI7, Q5YKI8 | Gene names | GGNBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin-binding protein 1. | |||||
|
PTN4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 32) | NC score | 0.013212 (rank : 89) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P29074 | Gene names | PTPN4 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 4 (EC 3.1.3.48) (Protein-tyrosine phosphatase MEG1) (PTPase-MEG1) (MEG). | |||||
|
TCGAP_HUMAN
|
||||||
| θ value | 1.81305 (rank : 33) | NC score | 0.026171 (rank : 62) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 872 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O14559, O14552, O14560, Q6ZSP6, Q96CP3, Q9NT23 | Gene names | SNX26, TCGAP | |||
|
Domain Architecture |

|
|||||
| Description | TC10/CDC42 GTPase-activating protein (Sorting nexin-26). | |||||
|
DYN1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 34) | NC score | 0.032887 (rank : 52) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q05193 | Gene names | DNM1, DNM | |||
|
Domain Architecture |

|
|||||
| Description | Dynamin-1 (EC 3.6.5.5). | |||||
|
GLI2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.006462 (rank : 101) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 971 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P10070, O60252, O60253, O60254, O60255, Q15590, Q15591 | Gene names | GLI2, THP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLI2 (Tax helper protein). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.040216 (rank : 43) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
CX020_HUMAN
|
||||||
| θ value | 3.0926 (rank : 37) | NC score | 0.038767 (rank : 45) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NDZ0, Q5JXE5 | Gene names | CXorf20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein CXorf20. | |||||
|
DX26B_MOUSE
|
||||||
| θ value | 3.0926 (rank : 38) | NC score | 0.027221 (rank : 60) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BND4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DDX26B. | |||||
|
EST2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 39) | NC score | 0.017699 (rank : 77) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O00748, Q16859, Q5MAB8, Q7Z366, Q8IUP4, Q8TCP8 | Gene names | CES2, ICE | |||
|
Domain Architecture |

|
|||||
| Description | Carboxylesterase 2 precursor (EC 3.1.1.1) (CE-2) (hCE-2). | |||||
|
PHLB1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 40) | NC score | 0.027474 (rank : 59) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1038 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q86UU1, O75133, Q4KMF8, Q8TEQ2 | Gene names | PHLDB1, DLNB07, KIAA0638, LL5A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 1 (Protein LL5-alpha). | |||||
|
RT30_HUMAN
|
||||||
| θ value | 3.0926 (rank : 41) | NC score | 0.030638 (rank : 57) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NP92, Q96I91, Q96Q19, Q9H0P8, Q9NSF9, Q9NZ76 | Gene names | MRPS30, PDCD9 | |||
|
Domain Architecture |
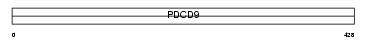
|
|||||
| Description | Mitochondrial 28S ribosomal protein S30 (S30mt) (MRP-S30) (Programmed cell death protein 9). | |||||
|
RUNX2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 42) | NC score | 0.031913 (rank : 54) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q13950, O14614, O14615, O95181 | Gene names | RUNX2, AML3, CBFA1, OSF2, PEBP2A | |||
|
Domain Architecture |
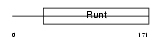
|
|||||
| Description | Runt-related transcription factor 2 (Core-binding factor, alpha 1 subunit) (CBF-alpha 1) (Acute myeloid leukemia 3 protein) (Oncogene AML-3) (Polyomavirus enhancer-binding protein 2 alpha A subunit) (PEBP2-alpha A) (PEA2-alpha A) (SL3-3 enhancer factor 1 alpha A subunit) (SL3/AKV core-binding factor alpha A subunit) (Osteoblast- specific transcription factor 2) (OSF-2). | |||||
|
RUNX2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 43) | NC score | 0.033397 (rank : 51) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q08775, O35183, Q08776, Q9JLN0, Q9QUQ6, Q9QY29, Q9R0U4, Q9Z2J7 | Gene names | Runx2, Aml3, Cbfa1, Osf2, Pebp2a | |||
|
Domain Architecture |
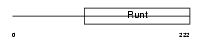
|
|||||
| Description | Runt-related transcription factor 2 (Core-binding factor, alpha 1 subunit) (CBF-alpha 1) (Acute myeloid leukemia 3 protein) (Oncogene AML-3) (Polyomavirus enhancer-binding protein 2 alpha A subunit) (PEBP2-alpha A) (PEA2-alpha A) (SL3-3 enhancer factor 1 alpha A subunit) (SL3/AKV core-binding factor alpha A subunit) (Osteoblast- specific transcription factor 2) (OSF-2). | |||||
|
SMCA4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 44) | NC score | 0.021904 (rank : 70) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P51532, O95052, Q9HBD3 | Gene names | SMARCA4, BRG1, SNF2B, SNF2L4 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L4 (EC 3.6.1.-) (ATP- dependent helicase SMARCA4) (SNF2-beta) (BRG-1 protein) (Mitotic growth and transcription activator) (Brahma protein homolog 1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 4). | |||||
|
STRAD_MOUSE
|
||||||
| θ value | 3.0926 (rank : 45) | NC score | 0.045300 (rank : 38) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q60924 | Gene names | Stra13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-inducible E3 protein (Hematopoietic-specific protein E3). | |||||
|
TRIP6_HUMAN
|
||||||
| θ value | 3.0926 (rank : 46) | NC score | 0.016458 (rank : 83) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q15654, O15170, O15275, Q9BTB2, Q9UNT4 | Gene names | TRIP6, OIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 6 (TRIP6) (OPA-interacting protein 1) (Zyxin-related protein 1) (ZRP-1). | |||||
|
UBP2L_HUMAN
|
||||||
| θ value | 3.0926 (rank : 47) | NC score | 0.052434 (rank : 27) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14157, Q5VU75, Q5VU76, Q9BTU3, Q9UGL5 | Gene names | UBAP2L, KIAA0144, NICE4 | |||
|
Domain Architecture |
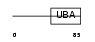
|
|||||
| Description | Ubiquitin-associated protein 2-like (Protein NICE-4). | |||||
|
UBP2L_MOUSE
|
||||||
| θ value | 3.0926 (rank : 48) | NC score | 0.053577 (rank : 24) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q80X50, Q8BIT6, Q8BIW4, Q8BJ01, Q8CIG7, Q8K102, Q9CRT6 | Gene names | Ubap2l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated protein 2-like. | |||||
|
UBQL3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 49) | NC score | 0.025356 (rank : 63) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9H347, Q9NRE0 | Gene names | UBQLN3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquilin-3. | |||||
|
ANTR1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 50) | NC score | 0.024741 (rank : 65) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9CZ52 | Gene names | Antxr1, Atr, Tem8 | |||
|
Domain Architecture |
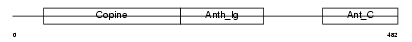
|
|||||
| Description | Anthrax toxin receptor 1 precursor (Tumor endothelial marker 8). | |||||
|
CECR2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 51) | NC score | 0.031028 (rank : 56) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9BXF3, Q658Z4, Q96P58, Q9C0C3 | Gene names | CECR2, KIAA1740 | |||
|
Domain Architecture |

|
|||||
| Description | Cat eye syndrome critical region protein 2. | |||||
|
DHX30_MOUSE
|
||||||
| θ value | 4.03905 (rank : 52) | NC score | 0.011401 (rank : 93) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99PU8, Q3U4Z4, Q3UFA0, Q3UJS4, Q91WA7, Q99KN7 | Gene names | Dhx30, Helg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX30 (EC 3.6.1.-) (DEAH box protein 30). | |||||
|
EP300_HUMAN
|
||||||
| θ value | 4.03905 (rank : 53) | NC score | 0.065057 (rank : 17) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
EPN2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 54) | NC score | 0.020369 (rank : 71) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8CHU3, O70447, Q8BZ85 | Gene names | Epn2 | |||
|
Domain Architecture |
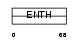
|
|||||
| Description | Epsin-2 (EPS-15-interacting protein 2) (Intersectin-EH-binding protein 2) (Ibp2). | |||||
|
JHD2A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 55) | NC score | 0.022691 (rank : 69) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6PCM1, Q2MJQ6, Q3TKW8, Q3UML3, Q6ZQ57, Q8K2J6, Q8K2K4, Q8R350 | Gene names | Jmjd1a, Jhdm2a, Kiaa0742 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 2A (EC 1.14.11.-) (Jumonji domain-containing protein 1A). | |||||
|
SEM6C_MOUSE
|
||||||
| θ value | 4.03905 (rank : 56) | NC score | 0.012817 (rank : 90) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9WTM3 | Gene names | Sema6c, Semay | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6C precursor (Semaphorin Y) (Sema Y). | |||||
|
SYCP1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 57) | NC score | 0.007972 (rank : 95) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1313 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q15431, O14963 | Gene names | SYCP1, SCP1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
T22D4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 58) | NC score | 0.034235 (rank : 49) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y3Q8 | Gene names | TSC22D4, TILZ2 | |||
|
Domain Architecture |

|
|||||
| Description | TSC22 domain family protein 4 (TSC22-related-inducible leucine zipper protein 2) (Tsc-22-like protein THG-1). | |||||
|
COCA1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 59) | NC score | 0.012708 (rank : 91) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99715, Q99716 | Gene names | COL12A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XII) chain precursor. | |||||
|
GCM1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.019220 (rank : 73) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P70348, O09103 | Gene names | Gcm1, Gcma | |||
|
Domain Architecture |
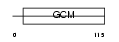
|
|||||
| Description | Chorion-specific transcription factor GCMa (Glial cells missing homolog 1) (GCM motif protein 1) (mGCMa) (mGCM1). | |||||
|
MAML1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.024775 (rank : 64) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q92585, Q9NZ12 | Gene names | MAML1, KIAA0200 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 1 (Mam-1). | |||||
|
NFAC3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.014022 (rank : 85) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P97305, Q60896 | Gene names | Nfatc3, Nfat4 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells, cytoplasmic 3 (NF-ATc3) (NFATc3) (T cell transcription factor NFAT4) (NF-AT4) (NFATx). | |||||
|
PDLI7_HUMAN
|
||||||
| θ value | 5.27518 (rank : 63) | NC score | 0.016604 (rank : 82) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 547 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9NR12, Q14250, Q5XG82, Q6NVZ5, Q96C91, Q9BXB8, Q9BXB9 | Gene names | PDLIM7, ENIGMA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ and LIM domain protein 7 (LIM mineralization protein) (LMP) (Protein enigma). | |||||
|
PRRT3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.068641 (rank : 16) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PE13 | Gene names | Prrt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
REST_MOUSE
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.003834 (rank : 102) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1506 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q922J3, Q571L7 | Gene names | Rsn, Kiaa4046 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Restin. | |||||
|
SOX15_MOUSE
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.013214 (rank : 88) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P43267, O70204, P70418, Q62246, Q91V00, Q91V43, Q920T1, Q9JLG2 | Gene names | Sox15, Sox-15 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-15 protein. | |||||
|
TMC2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.016809 (rank : 81) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8TDI7, Q5JXT0, Q5JXT1, Q6UWW4, Q6ZS41, Q8N9F3, Q9BYN2, Q9BYN3, Q9BYN4, Q9BYN5 | Gene names | TMC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane cochlear-expressed protein 2. | |||||
|
TRAF7_HUMAN
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.007645 (rank : 97) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6Q0C0, Q9H073 | Gene names | TRAF7, RFWD1, RNF119 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin protein ligase TRAF7 (EC 6.3.2.-) (TNF receptor- associated factor 7) (RING finger and WD repeat domain protein 1) (RING finger protein 119). | |||||
|
UBIP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.018752 (rank : 74) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NZI7, Q68CT0, Q86Y57, Q9H8V0, Q9UD76, Q9UD78 | Gene names | UBP1, LBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Upstream-binding protein 1 (LBP-1). | |||||
|
ZNF8_HUMAN
|
||||||
| θ value | 5.27518 (rank : 70) | NC score | -0.000370 (rank : 107) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 748 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P17098, Q6PI99 | Gene names | ZNF8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 8 (Zinc finger protein HF.18). | |||||
|
AMOL1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.012605 (rank : 92) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9D4H4, Q571F1 | Gene names | Amotl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
ARI1B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.088056 (rank : 11) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 567 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8NFD5, Q5JRD1, Q5VYC4, Q8IZY8, Q8TEV0, Q8TF02, Q99491, Q9ULI5 | Gene names | ARID1B, DAN15, KIAA1235, OSA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-rich interactive domain-containing protein 1B (ARID domain- containing protein 1B) (Osa homolog 2) (hOsa2) (p250R) (BRG1-binding protein hELD/OSA1) (BRG1-associated factor 250b) (BAF250B). | |||||
|
ARRD1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | 0.015292 (rank : 84) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N5I2 | Gene names | ARRDC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arrestin domain-containing protein 1. | |||||
|
CBL_HUMAN
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.016815 (rank : 80) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 353 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P22681 | Gene names | CBL, CBL2, RNF55 | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase CBL (EC 6.3.2.-) (Signal transduction protein CBL) (Proto-oncogene c-CBL) (Casitas B-lineage lymphoma proto- oncogene) (RING finger protein 55). | |||||
|
CN032_HUMAN
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.048077 (rank : 34) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8NDC0, Q96BG5 | Gene names | C14orf32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32. | |||||
|
CNOT4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.023462 (rank : 67) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BT14, Q8CCR4, Q9CV74, Q9Z1D0 | Gene names | Cnot4, Not4 | |||
|
Domain Architecture |
|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.026978 (rank : 61) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.029387 (rank : 58) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
OTU7B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.017960 (rank : 76) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6GQQ9, Q8WWA7, Q9NQ53, Q9UFF4 | Gene names | OTUD7B, ZA20D1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | OTU domain-containing protein 7B (EC 3.-.-.-) (Zinc finger protein Cezanne) (Zinc finger A20 domain-containing protein 1) (Cellular zinc finger anti-NF-kappa B protein). | |||||
|
PER2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.013778 (rank : 87) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
R3HD2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.039724 (rank : 44) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
SFPQ_MOUSE
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.038603 (rank : 46) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8VIJ6, Q9ERW2 | Gene names | Sfpq | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit). | |||||
|
SP5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.003555 (rank : 104) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 832 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6BEB4 | Gene names | SP5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Sp5. | |||||
|
SP5_MOUSE
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.003702 (rank : 103) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9JHX2 | Gene names | Sp5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor Sp5. | |||||
|
TRIM8_MOUSE
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.007851 (rank : 96) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99PJ2, Q8C508, Q8C700, Q8CGI2, Q99PQ4 | Gene names | Trim8, Gerp, Rnf27 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 8 (RING finger protein 27) (Glioblastoma-expressed RING finger protein). | |||||
|
BLNK_MOUSE
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.017154 (rank : 78) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9QUN3, O88504 | Gene names | Blnk, Bash, Ly57, Slp65 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell linker protein (Cytoplasmic adapter protein) (B-cell adapter containing SH2 domain protein) (B-cell adapter containing Src homology 2 domain protein) (Src homology 2 domain-containing leukocyte protein of 65 kDa) (Slp-65) (Lymphocyte antigen 57). | |||||
|
CAC1H_MOUSE
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.009006 (rank : 94) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O88427, Q80TJ2, Q9JKU5 | Gene names | Cacna1h, Kiaa1120 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1H (Voltage- gated calcium channel subunit alpha Cav3.2). | |||||
|
FOXP4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.007001 (rank : 100) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9DBY0, Q80V92, Q8CG10, Q8CIS1 | Gene names | Foxp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein P4 (Fork head-related protein like A) (mFKHLA). | |||||
|
GON4L_MOUSE
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.013997 (rank : 86) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DB00, Q80TB4, Q91YI9 | Gene names | Gon4l, Gon4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
HRX_HUMAN
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.018450 (rank : 75) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
MUCDL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.031827 (rank : 55) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9HBB8, Q9H746, Q9HAU3, Q9HBB5, Q9HBB6, Q9HBB7, Q9NX86, Q9NXI9 | Gene names | MUCDHL | |||
|
Domain Architecture |

|
|||||
| Description | Mucin and cadherin-like protein precursor (Mu-protocadherin). | |||||
|
PCDH7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.002108 (rank : 105) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60245, O60246, O60247 | Gene names | PCDH7, BHPCDH | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin-7 precursor (Brain-heart protocadherin) (BH-Pcdh). | |||||
|
RBMS2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 93) | NC score | 0.007212 (rank : 99) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VC70 | Gene names | Rbms2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding motif, single-stranded-interacting protein 2. | |||||
|
SASH1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.016957 (rank : 79) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O94885, Q8TAI0, Q9H7R7 | Gene names | SASH1, KIAA0790, PEPE1 | |||
|
Domain Architecture |

|
|||||
| Description | SAM and SH3 domain-containing protein 1 (Proline-glutamate repeat- containing protein). | |||||
|
SDK1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 95) | NC score | 0.007359 (rank : 98) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 511 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q7Z5N4, Q8TEN9, Q8TEP5, Q96N44 | Gene names | SDK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein sidekick-1 precursor. | |||||
|
SN1L2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | 0.001680 (rank : 106) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CFH6 | Gene names | Snf1lk2, Sik2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SNF1-like kinase 2 (EC 2.7.11.1) (Salt-inducible kinase 2). | |||||
|
MUC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.051878 (rank : 30) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
PRB3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.092117 (rank : 10) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
PRB4S_HUMAN
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.082354 (rank : 12) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
PRP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.056881 (rank : 23) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
PRP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.053013 (rank : 26) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
PRP5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.074133 (rank : 14) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P04281 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic proline-rich peptide IB-1. | |||||
|
S18L1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.053526 (rank : 25) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O75177, Q5JXJ3, Q8NE69, Q9BR55, Q9H4K6 | Gene names | SS18L1, KIAA0693 | |||
|
Domain Architecture |
|
|||||
| Description | SS18-like protein 1 (SYT homolog 1). | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.057821 (rank : 20) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.052349 (rank : 28) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
WASIP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.057529 (rank : 22) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
YLPM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.050583 (rank : 31) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
BAG4_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 4.24358e-175 (rank : 1) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 96 | |
| SwissProt Accessions | O95429, O95818 | Gene names | BAG4, SODD | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 4 (BCL2-associated athanogene 4) (BAG-4) (Silencer of death domains). | |||||
|
BAG4_MOUSE
|
||||||
| NC score | 0.956571 (rank : 2) | θ value | 2.12569e-142 (rank : 2) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8CI61, Q3TRL9, Q91VT5, Q9CWG2 | Gene names | Bag4, Sodd | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 4 (BCL2-associated athanogene 4) (BAG-4) (Silencer of death domains). | |||||
|
BAG3_HUMAN
|
||||||
| NC score | 0.547145 (rank : 3) | θ value | 6.62687e-19 (rank : 3) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O95817, Q9NT20, Q9P120 | Gene names | BAG3, BIS | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis) (Docking protein CAIR-1). | |||||
|
BAG5_HUMAN
|
||||||
| NC score | 0.508522 (rank : 4) | θ value | 1.9326e-10 (rank : 5) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UL15, O94950, Q86W59 | Gene names | BAG5, KIAA0873 | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 5 (BCL2-associated athanogene 5) (BAG-5). | |||||
|
BAG5_MOUSE
|
||||||
| NC score | 0.505999 (rank : 5) | θ value | 7.34386e-10 (rank : 6) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CI32, Q3TVA8, Q8K175, Q9DAU0 | Gene names | Bag5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BAG family molecular chaperone regulator 5 (BCL2-associated athanogene 5) (BAG-5). | |||||
|
BAG3_MOUSE
|
||||||
| NC score | 0.505563 (rank : 6) | θ value | 6.62687e-19 (rank : 4) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9JLV1, Q9JJC7 | Gene names | Bag3, Bis | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis). | |||||
|
PRRT3_HUMAN
|
||||||
| NC score | 0.106771 (rank : 7) | θ value | 0.0148317 (rank : 7) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
SSXT_HUMAN
|
||||||
| NC score | 0.103460 (rank : 8) | θ value | 0.163984 (rank : 12) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15532, Q16404, Q9BXC6 | Gene names | SS18, SSXT, SYT | |||
|
Domain Architecture |

|
|||||
| Description | SSXT protein (Synovial sarcoma, translocated to X chromosome) (SYT protein). | |||||
|
SSXT_MOUSE
|
||||||
| NC score | 0.102532 (rank : 9) | θ value | 0.21417 (rank : 14) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q62280 | Gene names | Ss18, Ssxt, Syt | |||
|
Domain Architecture |

|
|||||
| Description | SSXT protein (SYT protein) (Synovial sarcoma-associated Ss18-alpha). | |||||
|
PRB3_HUMAN
|
||||||
| NC score | 0.092117 (rank : 10) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
ARI1B_HUMAN
|
||||||
| NC score | 0.088056 (rank : 11) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 567 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8NFD5, Q5JRD1, Q5VYC4, Q8IZY8, Q8TEV0, Q8TF02, Q99491, Q9ULI5 | Gene names | ARID1B, DAN15, KIAA1235, OSA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-rich interactive domain-containing protein 1B (ARID domain- containing protein 1B) (Osa homolog 2) (hOsa2) (p250R) (BRG1-binding protein hELD/OSA1) (BRG1-associated factor 250b) (BAF250B). | |||||
|
PRB4S_HUMAN
|
||||||
| NC score | 0.082354 (rank : 12) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
GGNB1_HUMAN
|
||||||
| NC score | 0.076058 (rank : 13) | θ value | 1.81305 (rank : 31) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5YKI7, Q5YKI8 | Gene names | GGNBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin-binding protein 1. | |||||
|
PRP5_HUMAN
|
||||||
| NC score | 0.074133 (rank : 14) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P04281 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic proline-rich peptide IB-1. | |||||
|
ARI1A_HUMAN
|
||||||
| NC score | 0.070488 (rank : 15) | θ value | 0.47712 (rank : 18) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
PRRT3_MOUSE
|
||||||
| NC score | 0.068641 (rank : 16) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PE13 | Gene names | Prrt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
EP300_HUMAN
|
||||||
| NC score | 0.065057 (rank : 17) | θ value | 4.03905 (rank : 53) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
MAGE1_HUMAN
|
||||||
| NC score | 0.060300 (rank : 18) | θ value | 0.0961366 (rank : 9) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
FMN2_HUMAN
|
||||||
| NC score | 0.058236 (rank : 19) | θ value | 0.0961366 (rank : 8) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.057821 (rank : 20) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
YLPM1_HUMAN
|
||||||
| NC score | 0.057536 (rank : 21) | θ value | 1.06291 (rank : 26) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
WASIP_HUMAN
|
||||||
| NC score | 0.057529 (rank : 22) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
PRP1_HUMAN
|
||||||
| NC score | 0.056881 (rank : 23) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
UBP2L_MOUSE
|
||||||
| NC score | 0.053577 (rank : 24) | θ value | 3.0926 (rank : 48) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q80X50, Q8BIT6, Q8BIW4, Q8BJ01, Q8CIG7, Q8K102, Q9CRT6 | Gene names | Ubap2l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated protein 2-like. | |||||
|
S18L1_HUMAN
|
||||||
| NC score | 0.053526 (rank : 25) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O75177, Q5JXJ3, Q8NE69, Q9BR55, Q9H4K6 | Gene names | SS18L1, KIAA0693 | |||
|
Domain Architecture |

|
|||||
| Description | SS18-like protein 1 (SYT homolog 1). | |||||
|
PRP2_MOUSE
|
||||||
| NC score | 0.053013 (rank : 26) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
UBP2L_HUMAN
|
||||||
| NC score | 0.052434 (rank : 27) | θ value | 3.0926 (rank : 47) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14157, Q5VU75, Q5VU76, Q9BTU3, Q9UGL5 | Gene names | UBAP2L, KIAA0144, NICE4 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-associated protein 2-like (Protein NICE-4). | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.052349 (rank : 28) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.052007 (rank : 29) | θ value | 1.06291 (rank : 25) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.051878 (rank : 30) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
YLPM1_MOUSE
|
||||||
| NC score | 0.050583 (rank : 31) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
REXO1_HUMAN
|
||||||
| NC score | 0.050401 (rank : 32) | θ value | 0.365318 (rank : 17) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8N1G1, Q9ULT2 | Gene names | REXO1, ELOABP1, KIAA1138, TCEB3BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA exonuclease 1 homolog (EC 3.1.-.-) (Elongin A-binding protein 1) (EloA-BP1) (Transcription elongation factor B polypeptide 3-binding protein 1). | |||||
|
EYA1_MOUSE
|
||||||
| NC score | 0.049833 (rank : 33) | θ value | 1.81305 (rank : 30) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P97767, O08818 | Gene names | Eya1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
|
CN032_HUMAN
|
||||||
| NC score | 0.048077 (rank : 34) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8NDC0, Q96BG5 | Gene names | C14orf32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32. | |||||
|
IL3RB_HUMAN
|
||||||
| NC score | 0.048055 (rank : 35) | θ value | 1.06291 (rank : 24) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P32927 | Gene names | CSF2RB, IL3RB, IL5RB | |||
|
Domain Architecture |

|
|||||
| Description | Cytokine receptor common beta chain precursor (GM-CSF/IL-3/IL-5 receptor common beta-chain) (CD131 antigen) (CDw131). | |||||
|
DAB1_HUMAN
|
||||||
| NC score | 0.047192 (rank : 36) | θ value | 0.279714 (rank : 15) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75553, Q9NYA8 | Gene names | DAB1 | |||
|
Domain Architecture |

|
|||||
| Description | Disabled homolog 1. | |||||
|
EYA1_HUMAN
|
||||||
| NC score | 0.046936 (rank : 37) | θ value | 0.813845 (rank : 23) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99502 | Gene names | EYA1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
|
STRAD_MOUSE
|
||||||
| NC score | 0.045300 (rank : 38) | θ value | 3.0926 (rank : 45) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q60924 | Gene names | Stra13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-inducible E3 protein (Hematopoietic-specific protein E3). | |||||
|
CEL_HUMAN
|
||||||
| NC score | 0.043540 (rank : 39) | θ value | 0.813845 (rank : 22) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
TULP4_HUMAN
|
||||||
| NC score | 0.042833 (rank : 40) | θ value | 0.62314 (rank : 21) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9NRJ4, Q9HD22, Q9P2F0 | Gene names | TULP4, KIAA1397, TUSP | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-like protein 4 (Tubby superfamily protein). | |||||
|
LEF1_HUMAN
|
||||||
| NC score | 0.041027 (rank : 41) | θ value | 0.62314 (rank : 19) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UJU2, Q9HAZ0 | Gene names | LEF1 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphoid enhancer-binding factor 1 (LEF-1) (T cell-specific transcription factor 1-alpha) (TCF1-alpha). | |||||
|
LEF1_MOUSE
|
||||||
| NC score | 0.040942 (rank : 42) | θ value | 0.62314 (rank : 20) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P27782 | Gene names | Lef1, Lef-1 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphoid enhancer-binding factor 1 (LEF-1). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.040216 (rank : 43) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
R3HD2_MOUSE
|
||||||
| NC score | 0.039724 (rank : 44) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
CX020_HUMAN
|
||||||
| NC score | 0.038767 (rank : 45) | θ value | 3.0926 (rank : 37) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NDZ0, Q5JXE5 | Gene names | CXorf20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein CXorf20. | |||||
|
SFPQ_MOUSE
|
||||||
| NC score | 0.038603 (rank : 46) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8VIJ6, Q9ERW2 | Gene names | Sfpq | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit). | |||||
|
IASPP_MOUSE
|
||||||
| NC score | 0.037957 (rank : 47) | θ value | 0.279714 (rank : 16) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1024 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q5I1X5 | Gene names | Ppp1r13l, Nkip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
ZN750_HUMAN
|
||||||
| NC score | 0.037954 (rank : 48) | θ value | 1.38821 (rank : 29) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q32MQ0, Q9H899 | Gene names | ZNF750 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ZNF750. | |||||
|
T22D4_HUMAN
|
||||||
| NC score | 0.034235 (rank : 49) | θ value | 4.03905 (rank : 58) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y3Q8 | Gene names | TSC22D4, TILZ2 | |||
|
Domain Architecture |

|
|||||
| Description | TSC22 domain family protein 4 (TSC22-related-inducible leucine zipper protein 2) (Tsc-22-like protein THG-1). | |||||
|
CTND2_HUMAN
|
||||||
| NC score | 0.033838 (rank : 50) | θ value | 0.163984 (rank : 11) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UQB3, O00379, O15390, O43206, O43840, Q13589, Q9UM66, Q9UPM3 | Gene names | CTNND2, NPRAP | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Delta-catenin) (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin) (GT24). | |||||
|
RUNX2_MOUSE
|
||||||
| NC score | 0.033397 (rank : 51) | θ value | 3.0926 (rank : 43) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q08775, O35183, Q08776, Q9JLN0, Q9QUQ6, Q9QY29, Q9R0U4, Q9Z2J7 | Gene names | Runx2, Aml3, Cbfa1, Osf2, Pebp2a | |||
|
Domain Architecture |

|
|||||
| Description | Runt-related transcription factor 2 (Core-binding factor, alpha 1 subunit) (CBF-alpha 1) (Acute myeloid leukemia 3 protein) (Oncogene AML-3) (Polyomavirus enhancer-binding protein 2 alpha A subunit) (PEBP2-alpha A) (PEA2-alpha A) (SL3-3 enhancer factor 1 alpha A subunit) (SL3/AKV core-binding factor alpha A subunit) (Osteoblast- specific transcription factor 2) (OSF-2). | |||||
|
DYN1_HUMAN
|
||||||
| NC score | 0.032887 (rank : 52) | θ value | 2.36792 (rank : 34) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q05193 | Gene names | DNM1, DNM | |||
|
Domain Architecture |

|
|||||
| Description | Dynamin-1 (EC 3.6.5.5). | |||||
|
DYN1_MOUSE
|
||||||
| NC score | 0.032841 (rank : 53) | θ value | 1.38821 (rank : 27) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P39053, Q61358, Q61359, Q61360, Q8JZZ4, Q9CSY7, Q9QXX1 | Gene names | Dnm1, Dnm | |||
|
Domain Architecture |

|
|||||
| Description | Dynamin-1 (EC 3.6.5.5). | |||||
|
RUNX2_HUMAN
|
||||||
| NC score | 0.031913 (rank : 54) | θ value | 3.0926 (rank : 42) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q13950, O14614, O14615, O95181 | Gene names | RUNX2, AML3, CBFA1, OSF2, PEBP2A | |||
|
Domain Architecture |

|
|||||
| Description | Runt-related transcription factor 2 (Core-binding factor, alpha 1 subunit) (CBF-alpha 1) (Acute myeloid leukemia 3 protein) (Oncogene AML-3) (Polyomavirus enhancer-binding protein 2 alpha A subunit) (PEBP2-alpha A) (PEA2-alpha A) (SL3-3 enhancer factor 1 alpha A subunit) (SL3/AKV core-binding factor alpha A subunit) (Osteoblast- specific transcription factor 2) (OSF-2). | |||||
|
MUCDL_HUMAN
|
||||||
| NC score | 0.031827 (rank : 55) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9HBB8, Q9H746, Q9HAU3, Q9HBB5, Q9HBB6, Q9HBB7, Q9NX86, Q9NXI9 | Gene names | MUCDHL | |||
|
Domain Architecture |

|
|||||
| Description | Mucin and cadherin-like protein precursor (Mu-protocadherin). | |||||
|
CECR2_HUMAN
|
||||||
| NC score | 0.031028 (rank : 56) | θ value | 4.03905 (rank : 51) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9BXF3, Q658Z4, Q96P58, Q9C0C3 | Gene names | CECR2, KIAA1740 | |||
|
Domain Architecture |

|
|||||
| Description | Cat eye syndrome critical region protein 2. | |||||
|
RT30_HUMAN
|
||||||
| NC score | 0.030638 (rank : 57) | θ value | 3.0926 (rank : 41) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NP92, Q96I91, Q96Q19, Q9H0P8, Q9NSF9, Q9NZ76 | Gene names | MRPS30, PDCD9 | |||
|
Domain Architecture |

|
|||||
| Description | Mitochondrial 28S ribosomal protein S30 (S30mt) (MRP-S30) (Programmed cell death protein 9). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.029387 (rank : 58) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
PHLB1_HUMAN
|
||||||
| NC score | 0.027474 (rank : 59) | θ value | 3.0926 (rank : 40) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1038 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q86UU1, O75133, Q4KMF8, Q8TEQ2 | Gene names | PHLDB1, DLNB07, KIAA0638, LL5A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 1 (Protein LL5-alpha). | |||||
|
DX26B_MOUSE
|
||||||
| NC score | 0.027221 (rank : 60) | θ value | 3.0926 (rank : 38) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BND4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DDX26B. | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.026978 (rank : 61) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
TCGAP_HUMAN
|
||||||
| NC score | 0.026171 (rank : 62) | θ value | 1.81305 (rank : 33) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 872 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O14559, O14552, O14560, Q6ZSP6, Q96CP3, Q9NT23 | Gene names | SNX26, TCGAP | |||
|
Domain Architecture |

|
|||||
| Description | TC10/CDC42 GTPase-activating protein (Sorting nexin-26). | |||||
|
UBQL3_HUMAN
|
||||||
| NC score | 0.025356 (rank : 63) | θ value | 3.0926 (rank : 49) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9H347, Q9NRE0 | Gene names | UBQLN3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquilin-3. | |||||
|
MAML1_HUMAN
|
||||||
| NC score | 0.024775 (rank : 64) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q92585, Q9NZ12 | Gene names | MAML1, KIAA0200 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 1 (Mam-1). | |||||
|
ANTR1_MOUSE
|
||||||
| NC score | 0.024741 (rank : 65) | θ value | 4.03905 (rank : 50) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9CZ52 | Gene names | Antxr1, Atr, Tem8 | |||
|
Domain Architecture |

|
|||||
| Description | Anthrax toxin receptor 1 precursor (Tumor endothelial marker 8). | |||||
|
SPEG_MOUSE
|
||||||
| NC score | 0.024309 (rank : 66) | θ value | 0.0961366 (rank : 10) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
CNOT4_MOUSE
|
||||||
| NC score | 0.023462 (rank : 67) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BT14, Q8CCR4, Q9CV74, Q9Z1D0 | Gene names | Cnot4, Not4 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
CO4A3_HUMAN
|
||||||
| NC score | 0.023156 (rank : 68) | θ value | 0.21417 (rank : 13) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 421 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q01955, Q9BQT2 | Gene names | COL4A3 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-3(IV) chain precursor (Goodpasture antigen). | |||||
|
JHD2A_MOUSE
|
||||||
| NC score | 0.022691 (rank : 69) | θ value | 4.03905 (rank : 55) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6PCM1, Q2MJQ6, Q3TKW8, Q3UML3, Q6ZQ57, Q8K2J6, Q8K2K4, Q8R350 | Gene names | Jmjd1a, Jhdm2a, Kiaa0742 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 2A (EC 1.14.11.-) (Jumonji domain-containing protein 1A). | |||||
|
SMCA4_HUMAN
|
||||||
| NC score | 0.021904 (rank : 70) | θ value | 3.0926 (rank : 44) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P51532, O95052, Q9HBD3 | Gene names | SMARCA4, BRG1, SNF2B, SNF2L4 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L4 (EC 3.6.1.-) (ATP- dependent helicase SMARCA4) (SNF2-beta) (BRG-1 protein) (Mitotic growth and transcription activator) (Brahma protein homolog 1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 4). | |||||
|
EPN2_MOUSE
|
||||||
| NC score | 0.020369 (rank : 71) | θ value | 4.03905 (rank : 54) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8CHU3, O70447, Q8BZ85 | Gene names | Epn2 | |||
|
Domain Architecture |

|
|||||
| Description | Epsin-2 (EPS-15-interacting protein 2) (Intersectin-EH-binding protein 2) (Ibp2). | |||||
|
SPEG_HUMAN
|
||||||
| NC score | 0.019586 (rank : 72) | θ value | 1.38821 (rank : 28) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
GCM1_MOUSE
|
||||||
| NC score | 0.019220 (rank : 73) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P70348, O09103 | Gene names | Gcm1, Gcma | |||
|
Domain Architecture |

|
|||||
| Description | Chorion-specific transcription factor GCMa (Glial cells missing homolog 1) (GCM motif protein 1) (mGCMa) (mGCM1). | |||||
|
UBIP1_HUMAN
|
||||||
| NC score | 0.018752 (rank : 74) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NZI7, Q68CT0, Q86Y57, Q9H8V0, Q9UD76, Q9UD78 | Gene names | UBP1, LBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Upstream-binding protein 1 (LBP-1). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.018450 (rank : 75) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
OTU7B_HUMAN
|
||||||
| NC score | 0.017960 (rank : 76) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6GQQ9, Q8WWA7, Q9NQ53, Q9UFF4 | Gene names | OTUD7B, ZA20D1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | OTU domain-containing protein 7B (EC 3.-.-.-) (Zinc finger protein Cezanne) (Zinc finger A20 domain-containing protein 1) (Cellular zinc finger anti-NF-kappa B protein). | |||||
|
EST2_HUMAN
|
||||||
| NC score | 0.017699 (rank : 77) | θ value | 3.0926 (rank : 39) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O00748, Q16859, Q5MAB8, Q7Z366, Q8IUP4, Q8TCP8 | Gene names | CES2, ICE | |||
|
Domain Architecture |

|
|||||
| Description | Carboxylesterase 2 precursor (EC 3.1.1.1) (CE-2) (hCE-2). | |||||
|
BLNK_MOUSE
|
||||||
| NC score | 0.017154 (rank : 78) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9QUN3, O88504 | Gene names | Blnk, Bash, Ly57, Slp65 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell linker protein (Cytoplasmic adapter protein) (B-cell adapter containing SH2 domain protein) (B-cell adapter containing Src homology 2 domain protein) (Src homology 2 domain-containing leukocyte protein of 65 kDa) (Slp-65) (Lymphocyte antigen 57). | |||||
|
SASH1_HUMAN
|
||||||
| NC score | 0.016957 (rank : 79) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O94885, Q8TAI0, Q9H7R7 | Gene names | SASH1, KIAA0790, PEPE1 | |||
|
Domain Architecture |

|
|||||
| Description | SAM and SH3 domain-containing protein 1 (Proline-glutamate repeat- containing protein). | |||||
|
CBL_HUMAN
|
||||||
| NC score | 0.016815 (rank : 80) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 353 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P22681 | Gene names | CBL, CBL2, RNF55 | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase CBL (EC 6.3.2.-) (Signal transduction protein CBL) (Proto-oncogene c-CBL) (Casitas B-lineage lymphoma proto- oncogene) (RING finger protein 55). | |||||
|
TMC2_HUMAN
|
||||||
| NC score | 0.016809 (rank : 81) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8TDI7, Q5JXT0, Q5JXT1, Q6UWW4, Q6ZS41, Q8N9F3, Q9BYN2, Q9BYN3, Q9BYN4, Q9BYN5 | Gene names | TMC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane cochlear-expressed protein 2. | |||||
|
PDLI7_HUMAN
|
||||||
| NC score | 0.016604 (rank : 82) | θ value | 5.27518 (rank : 63) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 547 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9NR12, Q14250, Q5XG82, Q6NVZ5, Q96C91, Q9BXB8, Q9BXB9 | Gene names | PDLIM7, ENIGMA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ and LIM domain protein 7 (LIM mineralization protein) (LMP) (Protein enigma). | |||||
|
TRIP6_HUMAN
|
||||||
| NC score | 0.016458 (rank : 83) | θ value | 3.0926 (rank : 46) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q15654, O15170, O15275, Q9BTB2, Q9UNT4 | Gene names | TRIP6, OIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 6 (TRIP6) (OPA-interacting protein 1) (Zyxin-related protein 1) (ZRP-1). | |||||
|
ARRD1_HUMAN
|
||||||
| NC score | 0.015292 (rank : 84) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N5I2 | Gene names | ARRDC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arrestin domain-containing protein 1. | |||||
|
NFAC3_MOUSE
|
||||||
| NC score | 0.014022 (rank : 85) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P97305, Q60896 | Gene names | Nfatc3, Nfat4 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells, cytoplasmic 3 (NF-ATc3) (NFATc3) (T cell transcription factor NFAT4) (NF-AT4) (NFATx). | |||||
|
GON4L_MOUSE
|
||||||
| NC score | 0.013997 (rank : 86) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DB00, Q80TB4, Q91YI9 | Gene names | Gon4l, Gon4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
PER2_HUMAN
|
||||||
| NC score | 0.013778 (rank : 87) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
SOX15_MOUSE
|
||||||
| NC score | 0.013214 (rank : 88) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P43267, O70204, P70418, Q62246, Q91V00, Q91V43, Q920T1, Q9JLG2 | Gene names | Sox15, Sox-15 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-15 protein. | |||||
|
PTN4_HUMAN
|
||||||
| NC score | 0.013212 (rank : 89) | θ value | 1.81305 (rank : 32) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P29074 | Gene names | PTPN4 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 4 (EC 3.1.3.48) (Protein-tyrosine phosphatase MEG1) (PTPase-MEG1) (MEG). | |||||
|
SEM6C_MOUSE
|
||||||
| NC score | 0.012817 (rank : 90) | θ value | 4.03905 (rank : 56) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9WTM3 | Gene names | Sema6c, Semay | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6C precursor (Semaphorin Y) (Sema Y). | |||||
|
COCA1_HUMAN
|
||||||
| NC score | 0.012708 (rank : 91) | θ value | 5.27518 (rank : 59) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99715, Q99716 | Gene names | COL12A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XII) chain precursor. | |||||
|
AMOL1_MOUSE
|
||||||
| NC score | 0.012605 (rank : 92) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9D4H4, Q571F1 | Gene names | Amotl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
DHX30_MOUSE
|
||||||
| NC score | 0.011401 (rank : 93) | θ value | 4.03905 (rank : 52) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99PU8, Q3U4Z4, Q3UFA0, Q3UJS4, Q91WA7, Q99KN7 | Gene names | Dhx30, Helg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX30 (EC 3.6.1.-) (DEAH box protein 30). | |||||
|
CAC1H_MOUSE
|
||||||
| NC score | 0.009006 (rank : 94) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O88427, Q80TJ2, Q9JKU5 | Gene names | Cacna1h, Kiaa1120 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1H (Voltage- gated calcium channel subunit alpha Cav3.2). | |||||
|
SYCP1_HUMAN
|
||||||
| NC score | 0.007972 (rank : 95) | θ value | 4.03905 (rank : 57) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1313 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q15431, O14963 | Gene names | SYCP1, SCP1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
TRIM8_MOUSE
|
||||||
| NC score | 0.007851 (rank : 96) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99PJ2, Q8C508, Q8C700, Q8CGI2, Q99PQ4 | Gene names | Trim8, Gerp, Rnf27 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 8 (RING finger protein 27) (Glioblastoma-expressed RING finger protein). | |||||
|
TRAF7_HUMAN
|
||||||
| NC score | 0.007645 (rank : 97) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6Q0C0, Q9H073 | Gene names | TRAF7, RFWD1, RNF119 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin protein ligase TRAF7 (EC 6.3.2.-) (TNF receptor- associated factor 7) (RING finger and WD repeat domain protein 1) (RING finger protein 119). | |||||
|
SDK1_HUMAN
|
||||||
| NC score | 0.007359 (rank : 98) | θ value | 8.99809 (rank : 95) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 511 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q7Z5N4, Q8TEN9, Q8TEP5, Q96N44 | Gene names | SDK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein sidekick-1 precursor. | |||||
|
RBMS2_MOUSE
|
||||||
| NC score | 0.007212 (rank : 99) | θ value | 8.99809 (rank : 93) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VC70 | Gene names | Rbms2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding motif, single-stranded-interacting protein 2. | |||||
|
FOXP4_MOUSE
|
||||||
| NC score | 0.007001 (rank : 100) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9DBY0, Q80V92, Q8CG10, Q8CIS1 | Gene names | Foxp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein P4 (Fork head-related protein like A) (mFKHLA). | |||||
|
GLI2_HUMAN
|
||||||
| NC score | 0.006462 (rank : 101) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 971 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P10070, O60252, O60253, O60254, O60255, Q15590, Q15591 | Gene names | GLI2, THP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLI2 (Tax helper protein). | |||||
|
REST_MOUSE
|
||||||
| NC score | 0.003834 (rank : 102) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1506 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q922J3, Q571L7 | Gene names | Rsn, Kiaa4046 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Restin. | |||||
|
SP5_MOUSE
|
||||||
| NC score | 0.003702 (rank : 103) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9JHX2 | Gene names | Sp5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor Sp5. | |||||
|
SP5_HUMAN
|
||||||
| NC score | 0.003555 (rank : 104) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 832 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6BEB4 | Gene names | SP5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Sp5. | |||||
|
PCDH7_HUMAN
|
||||||
| NC score | 0.002108 (rank : 105) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60245, O60246, O60247 | Gene names | PCDH7, BHPCDH | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin-7 precursor (Brain-heart protocadherin) (BH-Pcdh). | |||||
|
SN1L2_MOUSE
|
||||||
| NC score | 0.001680 (rank : 106) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CFH6 | Gene names | Snf1lk2, Sik2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SNF1-like kinase 2 (EC 2.7.11.1) (Salt-inducible kinase 2). | |||||
|
ZNF8_HUMAN
|
||||||
| NC score | -0.000370 (rank : 107) | θ value | 5.27518 (rank : 70) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 748 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P17098, Q6PI99 | Gene names | ZNF8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 8 (Zinc finger protein HF.18). | |||||