Please be patient as the page loads
|
MCM3A_MOUSE
|
||||||
| SwissProt Accessions | Q9WUU9 | Gene names | Mcm3ap, Ganp, Map80 | |||
|
Domain Architecture |

|
|||||
| Description | 80 kDa MCM3-associated protein (GANP protein). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
MCM3A_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.977005 (rank : 2) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O60318, Q9UMT4 | Gene names | MCM3AP, GANP, KIAA0572, MAP80 | |||
|
Domain Architecture |

|
|||||
| Description | 80 kDa MCM3-associated protein (GANP protein). | |||||
|
MCM3A_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q9WUU9 | Gene names | Mcm3ap, Ganp, Map80 | |||
|
Domain Architecture |

|
|||||
| Description | 80 kDa MCM3-associated protein (GANP protein). | |||||
|
EYA4_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 3) | NC score | 0.101839 (rank : 3) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O95677, O95464, O95679, Q8IW39, Q9NTR7 | Gene names | EYA4 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 4 (EC 3.1.3.48). | |||||
|
NU153_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 4) | NC score | 0.092062 (rank : 4) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P49790 | Gene names | NUP153 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup153 (Nucleoporin Nup153) (153 kDa nucleoporin). | |||||
|
CCD21_HUMAN
|
||||||
| θ value | 0.365318 (rank : 5) | NC score | 0.019018 (rank : 43) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1228 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6P2H3, Q5VY68, Q5VY70, Q9H6Q1, Q9H828, Q9UF52 | Gene names | CCDC21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
TROP_HUMAN
|
||||||
| θ value | 0.365318 (rank : 6) | NC score | 0.027531 (rank : 25) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q12816, Q9NU89, Q9UPN8 | Gene names | TRO, KIAA1114, MAGED3 | |||
|
Domain Architecture |

|
|||||
| Description | Trophinin (MAGE-D3 antigen). | |||||
|
BSN_HUMAN
|
||||||
| θ value | 0.62314 (rank : 7) | NC score | 0.041999 (rank : 16) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 0.62314 (rank : 8) | NC score | 0.042132 (rank : 15) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
ENAM_MOUSE
|
||||||
| θ value | 0.62314 (rank : 9) | NC score | 0.040984 (rank : 17) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O55196 | Gene names | Enam | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Enamelin precursor. | |||||
|
EYA1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 10) | NC score | 0.059528 (rank : 8) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P97767, O08818 | Gene names | Eya1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
|
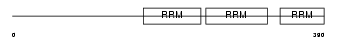
HNRPR_HUMAN
|
||||||
| θ value | 0.62314 (rank : 11) | NC score | 0.030247 (rank : 21) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O43390 | Gene names | HNRPR | |||
|
Domain Architecture |
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein R (hnRNP R). | |||||
|
NUP98_HUMAN
|
||||||
| θ value | 0.813845 (rank : 12) | NC score | 0.058919 (rank : 9) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P52948, Q8IUT2, Q8WYB0, Q96E54, Q9H3Q4, Q9NT02, Q9UF57, Q9UHX0, Q9Y6J4, Q9Y6J5 | Gene names | NUP98 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup98-Nup96 precursor [Contains: Nuclear pore complex protein Nup98 (Nucleoporin Nup98) (98 kDa nucleoporin); Nuclear pore complex protein Nup96 (Nucleoporin Nup96) (96 kDa nucleoporin)]. | |||||
|
ZN646_HUMAN
|
||||||
| θ value | 0.813845 (rank : 13) | NC score | 0.006089 (rank : 68) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 813 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O15015 | Gene names | ZNF646, KIAA0296 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 646. | |||||
|
BUB1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 14) | NC score | 0.024676 (rank : 32) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O43683, O43430, O43643, O60626 | Gene names | BUB1, BUB1L | |||
|
Domain Architecture |

|
|||||
| Description | Mitotic checkpoint serine/threonine-protein kinase BUB1 (EC 2.7.11.1) (hBUB1) (BUB1A). | |||||
|
HORN_MOUSE
|
||||||
| θ value | 1.06291 (rank : 15) | NC score | 0.026665 (rank : 29) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8VHD8 | Gene names | Hrnr | |||
|
Domain Architecture |

|
|||||
| Description | Hornerin. | |||||
|
NU214_HUMAN
|
||||||
| θ value | 1.06291 (rank : 16) | NC score | 0.061595 (rank : 7) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P35658 | Gene names | NUP214, CAIN, CAN | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup214 (Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein). | |||||
|
PO121_MOUSE
|
||||||
| θ value | 1.06291 (rank : 17) | NC score | 0.066144 (rank : 6) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8K3Z9, Q7TSH5 | Gene names | Pom121, Nup121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa). | |||||
|
EYA4_MOUSE
|
||||||
| θ value | 1.38821 (rank : 18) | NC score | 0.068516 (rank : 5) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Z191 | Gene names | Eya4 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 4 (EC 3.1.3.48). | |||||
|
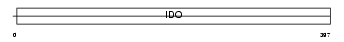
I23O_HUMAN
|
||||||
| θ value | 1.38821 (rank : 19) | NC score | 0.051910 (rank : 11) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P14902 | Gene names | INDO, IDO | |||
|
Domain Architecture |
|
|||||
| Description | Indoleamine 2,3-dioxygenase (EC 1.13.11.52) (IDO) (Indoleamine-pyrrole 2,3-dioxygenase). | |||||
|
K0256_MOUSE
|
||||||
| θ value | 1.38821 (rank : 20) | NC score | 0.030035 (rank : 22) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6A098 | Gene names | Kiaa0256 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0256. | |||||
|
CEP57_MOUSE
|
||||||
| θ value | 1.81305 (rank : 21) | NC score | 0.026283 (rank : 30) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 580 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8CEE0, Q6ZQJ3, Q7TN18, Q80X65, Q810F2, Q9D4J4, Q9D5S4, Q9D5W5 | Gene names | Cep57, Kiaa0092, Tsp57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 57 kDa (Cep57 protein) (Testis-specific protein 57) (Translokin). | |||||
|
EWS_HUMAN
|
||||||
| θ value | 1.81305 (rank : 22) | NC score | 0.039282 (rank : 19) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q01844, Q92635 | Gene names | EWSR1, EWS | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein EWS (EWS oncogene) (Ewing sarcoma breakpoint region 1 protein). | |||||
|
EWS_MOUSE
|
||||||
| θ value | 1.81305 (rank : 23) | NC score | 0.039627 (rank : 18) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61545 | Gene names | Ewsr1, Ews, Ewsh | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein EWS. | |||||
|
MLTK_HUMAN
|
||||||
| θ value | 1.81305 (rank : 24) | NC score | 0.005098 (rank : 71) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 911 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NYL2, Q53SX1, Q580W8, Q59GY5, Q86YW8, Q9HCC4, Q9HCC5, Q9HDD2, Q9NYE9 | Gene names | MLTK, ZAK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase MLT (EC 2.7.11.25) (MLK-like mitogen-activated protein triple kinase) (Leucine zipper- and sterile alpha motif-containing kinase) (Sterile alpha motif- and leucine zipper-containing kinase AZK) (Mixed lineage kinase-related kinase) (MLK-related kinase) (MRK) (Cervical cancer suppressor gene 4 protein). | |||||
|
NPAS4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 25) | NC score | 0.026797 (rank : 28) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
PF21B_MOUSE
|
||||||
| θ value | 1.81305 (rank : 26) | NC score | 0.023466 (rank : 35) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8C966 | Gene names | Phf21b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
SHAN1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 27) | NC score | 0.013456 (rank : 52) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Y566, Q9NYW9 | Gene names | SHANK1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 1 (Shank1) (Somatostatin receptor-interacting protein) (SSTR-interacting protein) (SSTRIP). | |||||
|
CMGA_MOUSE
|
||||||
| θ value | 2.36792 (rank : 28) | NC score | 0.029970 (rank : 23) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P26339 | Gene names | Chga | |||
|
Domain Architecture |

|
|||||
| Description | Chromogranin A precursor (CgA) [Contains: Pancreastatin; Beta-granin; WE-14]. | |||||
|
VPS54_MOUSE
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.027247 (rank : 26) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5SPW0, Q8BPB3, Q8CFZ7, Q8CHL5, Q8R3R4, Q8R3X1 | Gene names | Vps54 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vacuolar protein sorting-associated protein 54 (Tumor antigen SLP-8p homolog). | |||||
|
ZN592_HUMAN
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.006979 (rank : 66) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 762 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92610 | Gene names | ZNF592, KIAA0211 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 592. | |||||
|
APEH_MOUSE
|
||||||
| θ value | 3.0926 (rank : 31) | NC score | 0.022949 (rank : 38) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R146, Q8K029, Q8R0M9 | Gene names | Apeh | |||
|
Domain Architecture |

|
|||||
| Description | Acylamino-acid-releasing enzyme (EC 3.4.19.1) (AARE) (Acyl-peptide hydrolase) (APH) (Acylaminoacyl-peptidase). | |||||
|
CA026_HUMAN
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.024321 (rank : 33) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5T5J6, Q8NEK9, Q9BZQ7, Q9NXQ0 | Gene names | C1orf26 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C1orf26. | |||||
|
LIPA1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.009692 (rank : 61) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1294 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q13136, Q13135, Q14567, Q8N4I2 | Gene names | PPFIA1, LIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-1) (PTPRF-interacting protein alpha-1) (LAR-interacting protein 1) (LIP.1). | |||||
|
SYN1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 34) | NC score | 0.017115 (rank : 45) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P17600, O75825, Q5H9A9 | Gene names | SYN1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I) (Brain protein 4.1). | |||||
|
ZBT20_HUMAN
|
||||||
| θ value | 3.0926 (rank : 35) | NC score | 0.005319 (rank : 70) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 835 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9HC78 | Gene names | ZBTB20, DPZF, ZNF288 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 20 (Zinc finger protein 288) (Dendritic-derived BTB/POZ zinc finger protein). | |||||
|
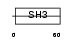
CASL_MOUSE
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.014917 (rank : 47) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O35177, Q8BJL8, Q8BK90, Q8BL52, Q8BM94, Q8BMI9, Q99KE7 | Gene names | Nedd9, Casl | |||
|
Domain Architecture |
|
|||||
| Description | Enhancer of filamentation 1 (MEF1) (CRK-associated substrate-related protein) (CAS-L) (p105) (Protein NEDD9) (Neural precursor cell expressed developmentally down-regulated protein 9). | |||||
|
CF152_HUMAN
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.025011 (rank : 31) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q86VQ0, Q9BWX7 | Gene names | C6orf152 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf152. | |||||
|
CLC4F_MOUSE
|
||||||
| θ value | 4.03905 (rank : 38) | NC score | 0.009806 (rank : 60) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 572 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P70194, Q8BLZ8 | Gene names | Clec4f, Clecsf13, Kclr | |||
|
Domain Architecture |

|
|||||
| Description | C-type lectin domain family 4 member F (C-type lectin superfamily member 13) (C-type lectin 13) (Kupffer cell receptor). | |||||
|
DOT1L_HUMAN
|
||||||
| θ value | 4.03905 (rank : 39) | NC score | 0.044843 (rank : 14) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8TEK3, O60379, Q96JL1 | Gene names | DOT1L, KIAA1814 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-79 specific (EC 2.1.1.43) (Histone H3-K79 methyltransferase) (H3-K79-HMTase) (DOT1-like protein). | |||||
|
EYA1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 40) | NC score | 0.052743 (rank : 10) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99502 | Gene names | EYA1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
|
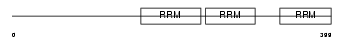
HNRPQ_MOUSE
|
||||||
| θ value | 4.03905 (rank : 41) | NC score | 0.024042 (rank : 34) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7TMK9, O88991, Q80YV6, Q8BGP1, Q8C5K6, Q8CGC2, Q91ZR0, Q99KU9, Q9QYF4 | Gene names | Syncrip, Hnrpq, Nsap1, Nsap1l | |||
|
Domain Architecture |
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein Q (hnRNP Q) (hnRNP-Q) (Synaptotagmin-binding, cytoplasmic RNA-interacting protein) (Glycine- and tyrosine-rich RNA-binding protein) (GRY-RBP) (NS1-associated protein 1) (pp68). | |||||
|
KLC2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 42) | NC score | 0.012149 (rank : 53) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H0B6, Q9H9C8, Q9HA20 | Gene names | KLC2 | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin light chain 2 (KLC 2). | |||||
|
NUPL2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 43) | NC score | 0.029155 (rank : 24) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CIC2, Q8BJX3, Q8BVP7, Q924S0 | Gene names | Nupl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin-like 2 (NLP-1). | |||||
|
RFA1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 44) | NC score | 0.023046 (rank : 36) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P27694, Q59ES9 | Gene names | RPA1, REPA1, RPA70 | |||
|
Domain Architecture |
|
|||||
| Description | Replication protein A 70 kDa DNA-binding subunit (RP-A) (RF-A) (Replication factor-A protein 1) (Single-stranded DNA-binding protein) (p70). | |||||
|
CEP57_HUMAN
|
||||||
| θ value | 5.27518 (rank : 45) | NC score | 0.021522 (rank : 40) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 576 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q86XR8, Q14704, Q8IXP0, Q9BVF9 | Gene names | CEP57, KIAA0092, TSP57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 57 kDa (Cep57 protein) (Testis-specific protein 57) (Translokin) (FGF2-interacting protein). | |||||
|
GOGB1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 46) | NC score | 0.006687 (rank : 67) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
HNRPQ_HUMAN
|
||||||
| θ value | 5.27518 (rank : 47) | NC score | 0.023043 (rank : 37) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60506, Q53H05, Q5TCG2, Q8IW78, Q8N599, Q96LC1, Q96LC2, Q9Y583 | Gene names | SYNCRIP, HNRPQ, NSAP1 | |||
|
Domain Architecture |
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein Q (hnRNP Q) (hnRNP-Q) (Synaptotagmin-binding, cytoplasmic RNA-interacting protein) (Glycine- and tyrosine-rich RNA-binding protein) (GRY-RBP) (NS1-associated protein 1). | |||||
|
KI13A_MOUSE
|
||||||
| θ value | 5.27518 (rank : 48) | NC score | 0.008949 (rank : 62) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9EQW7, O35062 | Gene names | Kif13a | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13A. | |||||
|
MAST2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | 0.003570 (rank : 74) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1110 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q60592, Q6P9M1, Q8CHD1 | Gene names | Mast2, Mast205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 2 (EC 2.7.11.1). | |||||
|
PDPK1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 50) | NC score | 0.003182 (rank : 77) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O15530 | Gene names | PDPK1, PDK1 | |||
|
Domain Architecture |
|
|||||
| Description | 3-phosphoinositide-dependent protein kinase 1 (EC 2.7.11.1) (hPDK1). | |||||
|
PDPK1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 51) | NC score | 0.003171 (rank : 78) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 852 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Z2A0, Q9R1D8, Q9R215 | Gene names | Pdpk1, Pdk1 | |||
|
Domain Architecture |
|
|||||
| Description | 3-phosphoinositide-dependent protein kinase 1 (EC 2.7.11.1) (mPDK1). | |||||
|
PO121_HUMAN
|
||||||
| θ value | 5.27518 (rank : 52) | NC score | 0.047605 (rank : 13) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y2N3, O75115, Q9Y4S7 | Gene names | POM121, KIAA0618, NUP121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa) (P145). | |||||
|
PSMF1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.022216 (rank : 39) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BHL8, Q8C0G9 | Gene names | Psmf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteasome inhibitor PI31 subunit. | |||||
|
CD34_MOUSE
|
||||||
| θ value | 6.88961 (rank : 54) | NC score | 0.020176 (rank : 41) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64314, Q62550, Q62551 | Gene names | Cd34 | |||
|
Domain Architecture |
|
|||||
| Description | Hematopoietic progenitor cell antigen CD34 precursor. | |||||
|
DGKI_HUMAN
|
||||||
| θ value | 6.88961 (rank : 55) | NC score | 0.007643 (rank : 64) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O75912, Q9NZ49 | Gene names | DGKI | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase iota (EC 2.7.1.107) (Diglyceride kinase iota) (DGK-iota) (DAG kinase iota). | |||||
|
FA13C_MOUSE
|
||||||
| θ value | 6.88961 (rank : 56) | NC score | 0.048199 (rank : 12) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9DBR2 | Gene names | Fam13c1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13C1. | |||||
|
FAK2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 57) | NC score | 0.003458 (rank : 75) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9QVP9 | Gene names | Ptk2b, Fak2, Pyk2, Raftk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein tyrosine kinase 2 beta (EC 2.7.10.2) (Focal adhesion kinase 2) (FADK 2) (Proline-rich tyrosine kinase 2) (Cell adhesion kinase beta) (CAK beta) (Calcium-dependent tyrosine kinase) (CADTK) (Related adhesion focal tyrosine kinase) (RAFTK). | |||||
|
FXL18_HUMAN
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | 0.035276 (rank : 20) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96ME1, Q9BR90, Q9BTC7, Q9HAK7 | Gene names | FBXL18, FBL18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 18 (F-box and leucine-rich repeat protein 18). | |||||
|
ILDR1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 59) | NC score | 0.016524 (rank : 46) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CBR1, Q6PFB3, Q8CB39, Q91VS0 | Gene names | Ildr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immunoglobulin-like domain-containing receptor 1 precursor. | |||||
|
SNG4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 60) | NC score | 0.014371 (rank : 50) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95473 | Gene names | SYNGR4 | |||
|
Domain Architecture |
|
|||||
| Description | Synaptogyrin-4. | |||||
|
VP13B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | 0.018314 (rank : 44) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7Z7G8, Q709C6, Q709C7, Q7Z7G4, Q7Z7G5, Q7Z7G6, Q7Z7G7, Q8NB77, Q9NWV1, Q9Y4E7 | Gene names | VPS13B, CHS1, COH1, KIAA0532 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vacuolar protein sorting-associated protein 13B (Cohen syndrome protein 1). | |||||
|
ADDB_MOUSE
|
||||||
| θ value | 8.99809 (rank : 62) | NC score | 0.007796 (rank : 63) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 574 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9QYB8, Q3U0E1, Q80VH9, Q8C1C4, Q9CXE3, Q9JLE5, Q9QYB7 | Gene names | Add2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adducin (Erythrocyte adducin subunit beta) (Add97). | |||||
|
ARNT_MOUSE
|
||||||
| θ value | 8.99809 (rank : 63) | NC score | 0.010396 (rank : 58) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
CAC1A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 64) | NC score | 0.011465 (rank : 56) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P97445 | Gene names | Cacna1a, Cach4, Cacn3, Cacnl1a4, Ccha1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
CHD9_MOUSE
|
||||||
| θ value | 8.99809 (rank : 65) | NC score | 0.005027 (rank : 72) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BYH8, Q7TMV5, Q8BJG8, Q8BZJ2, Q8CHG8 | Gene names | Chd9, Kiaa0308, Pric320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha- interacting complex protein 320 kDa). | |||||
|
FA35A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 66) | NC score | 0.013876 (rank : 51) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3UEN2 | Gene names | Fam35a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM35A. | |||||
|
FAK2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 67) | NC score | 0.003210 (rank : 76) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 847 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q14289, Q13475, Q14290, Q16709 | Gene names | PTK2B, FAK2, PYK2, RAFTK | |||
|
Domain Architecture |

|
|||||
| Description | Protein tyrosine kinase 2 beta (EC 2.7.10.2) (Focal adhesion kinase 2) (FADK 2) (Proline-rich tyrosine kinase 2) (Cell adhesion kinase beta) (CAK beta) (Calcium-dependent tyrosine kinase) (CADTK) (Related adhesion focal tyrosine kinase) (RAFTK). | |||||
|
IF4G1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 68) | NC score | 0.010213 (rank : 59) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6NZJ6, Q6NZN8, Q8BW99 | Gene names | Eif4g1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1). | |||||
|
K1543_HUMAN
|
||||||
| θ value | 8.99809 (rank : 69) | NC score | 0.020175 (rank : 42) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9P1Y5, Q8NDF1 | Gene names | KIAA1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
KI13A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 70) | NC score | 0.007395 (rank : 65) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9H1H9, Q9H1H8 | Gene names | KIF13A, RBKIN | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13A (Kinesin-like protein RBKIN). | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 71) | NC score | 0.014732 (rank : 49) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
P85A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.011677 (rank : 55) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P27986 | Gene names | PIK3R1, GRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit alpha (PI3-kinase p85-subunit alpha) (PtdIns-3-kinase p85-alpha) (PI3K). | |||||
|
P85A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.011894 (rank : 54) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P26450 | Gene names | Pik3r1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit alpha (PI3-kinase p85-subunit alpha) (PtdIns-3-kinase p85-alpha) (PI3K). | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.027170 (rank : 27) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
RFIP3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.010661 (rank : 57) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1055 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O75154, Q4VXV7, Q9H155, Q9H1G0, Q9NUI0 | Gene names | RAB11FIP3, KIAA0665 | |||
|
Domain Architecture |

|
|||||
| Description | Rab11 family-interacting protein 3 (Rab11-FIP3) (EF hands-containing Rab-interacting protein) (Eferin). | |||||
|
SNIP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.014821 (rank : 48) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9QWI6, O70298 | Gene names | P140, Kiaa1684 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | p130Cas-associated protein (p140Cap) (SNAP-25-interacting protein) (SNIP). | |||||
|
SOX14_HUMAN
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.005544 (rank : 69) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95416, Q3KPH7 | Gene names | SOX14, SOX28 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-14. | |||||
|
ZN445_HUMAN
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.000708 (rank : 79) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P59923 | Gene names | ZNF445, ZNF168 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 445. | |||||
|
ZN518_HUMAN
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.003742 (rank : 73) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6AHZ1, O15044, Q32MP4 | Gene names | ZNF518, KIAA0335 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 518. | |||||
|
MCM3A_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q9WUU9 | Gene names | Mcm3ap, Ganp, Map80 | |||
|
Domain Architecture |

|
|||||
| Description | 80 kDa MCM3-associated protein (GANP protein). | |||||
|
MCM3A_HUMAN
|
||||||
| NC score | 0.977005 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O60318, Q9UMT4 | Gene names | MCM3AP, GANP, KIAA0572, MAP80 | |||
|
Domain Architecture |

|
|||||
| Description | 80 kDa MCM3-associated protein (GANP protein). | |||||
|
EYA4_HUMAN
|
||||||
| NC score | 0.101839 (rank : 3) | θ value | 0.00869519 (rank : 3) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O95677, O95464, O95679, Q8IW39, Q9NTR7 | Gene names | EYA4 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 4 (EC 3.1.3.48). | |||||
|
NU153_HUMAN
|
||||||
| NC score | 0.092062 (rank : 4) | θ value | 0.0431538 (rank : 4) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P49790 | Gene names | NUP153 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup153 (Nucleoporin Nup153) (153 kDa nucleoporin). | |||||
|
EYA4_MOUSE
|
||||||
| NC score | 0.068516 (rank : 5) | θ value | 1.38821 (rank : 18) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Z191 | Gene names | Eya4 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 4 (EC 3.1.3.48). | |||||
|
PO121_MOUSE
|
||||||
| NC score | 0.066144 (rank : 6) | θ value | 1.06291 (rank : 17) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8K3Z9, Q7TSH5 | Gene names | Pom121, Nup121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa). | |||||
|
NU214_HUMAN
|
||||||
| NC score | 0.061595 (rank : 7) | θ value | 1.06291 (rank : 16) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P35658 | Gene names | NUP214, CAIN, CAN | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup214 (Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein). | |||||
|
EYA1_MOUSE
|
||||||
| NC score | 0.059528 (rank : 8) | θ value | 0.62314 (rank : 10) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P97767, O08818 | Gene names | Eya1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
|
NUP98_HUMAN
|
||||||
| NC score | 0.058919 (rank : 9) | θ value | 0.813845 (rank : 12) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P52948, Q8IUT2, Q8WYB0, Q96E54, Q9H3Q4, Q9NT02, Q9UF57, Q9UHX0, Q9Y6J4, Q9Y6J5 | Gene names | NUP98 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup98-Nup96 precursor [Contains: Nuclear pore complex protein Nup98 (Nucleoporin Nup98) (98 kDa nucleoporin); Nuclear pore complex protein Nup96 (Nucleoporin Nup96) (96 kDa nucleoporin)]. | |||||
|
EYA1_HUMAN
|
||||||
| NC score | 0.052743 (rank : 10) | θ value | 4.03905 (rank : 40) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99502 | Gene names | EYA1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 1 (EC 3.1.3.48). | |||||
|
I23O_HUMAN
|
||||||
| NC score | 0.051910 (rank : 11) | θ value | 1.38821 (rank : 19) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P14902 | Gene names | INDO, IDO | |||
|
Domain Architecture |

|
|||||
| Description | Indoleamine 2,3-dioxygenase (EC 1.13.11.52) (IDO) (Indoleamine-pyrrole 2,3-dioxygenase). | |||||
|
FA13C_MOUSE
|
||||||
| NC score | 0.048199 (rank : 12) | θ value | 6.88961 (rank : 56) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9DBR2 | Gene names | Fam13c1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13C1. | |||||
|
PO121_HUMAN
|
||||||
| NC score | 0.047605 (rank : 13) | θ value | 5.27518 (rank : 52) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y2N3, O75115, Q9Y4S7 | Gene names | POM121, KIAA0618, NUP121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa) (P145). | |||||
|
DOT1L_HUMAN
|
||||||
| NC score | 0.044843 (rank : 14) | θ value | 4.03905 (rank : 39) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8TEK3, O60379, Q96JL1 | Gene names | DOT1L, KIAA1814 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-79 specific (EC 2.1.1.43) (Histone H3-K79 methyltransferase) (H3-K79-HMTase) (DOT1-like protein). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.042132 (rank : 15) | θ value | 0.62314 (rank : 8) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.041999 (rank : 16) | θ value | 0.62314 (rank : 7) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
ENAM_MOUSE
|
||||||
| NC score | 0.040984 (rank : 17) | θ value | 0.62314 (rank : 9) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O55196 | Gene names | Enam | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Enamelin precursor. | |||||
|
EWS_MOUSE
|
||||||
| NC score | 0.039627 (rank : 18) | θ value | 1.81305 (rank : 23) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61545 | Gene names | Ewsr1, Ews, Ewsh | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein EWS. | |||||
|
EWS_HUMAN
|
||||||
| NC score | 0.039282 (rank : 19) | θ value | 1.81305 (rank : 22) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q01844, Q92635 | Gene names | EWSR1, EWS | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein EWS (EWS oncogene) (Ewing sarcoma breakpoint region 1 protein). | |||||
|
FXL18_HUMAN
|
||||||
| NC score | 0.035276 (rank : 20) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96ME1, Q9BR90, Q9BTC7, Q9HAK7 | Gene names | FBXL18, FBL18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 18 (F-box and leucine-rich repeat protein 18). | |||||
|
HNRPR_HUMAN
|
||||||
| NC score | 0.030247 (rank : 21) | θ value | 0.62314 (rank : 11) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O43390 | Gene names | HNRPR | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein R (hnRNP R). | |||||
|
K0256_MOUSE
|
||||||
| NC score | 0.030035 (rank : 22) | θ value | 1.38821 (rank : 20) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6A098 | Gene names | Kiaa0256 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0256. | |||||
|
CMGA_MOUSE
|
||||||
| NC score | 0.029970 (rank : 23) | θ value | 2.36792 (rank : 28) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P26339 | Gene names | Chga | |||
|
Domain Architecture |

|
|||||
| Description | Chromogranin A precursor (CgA) [Contains: Pancreastatin; Beta-granin; WE-14]. | |||||
|
NUPL2_MOUSE
|
||||||
| NC score | 0.029155 (rank : 24) | θ value | 4.03905 (rank : 43) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CIC2, Q8BJX3, Q8BVP7, Q924S0 | Gene names | Nupl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin-like 2 (NLP-1). | |||||
|
TROP_HUMAN
|
||||||
| NC score | 0.027531 (rank : 25) | θ value | 0.365318 (rank : 6) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q12816, Q9NU89, Q9UPN8 | Gene names | TRO, KIAA1114, MAGED3 | |||
|
Domain Architecture |

|
|||||
| Description | Trophinin (MAGE-D3 antigen). | |||||
|
VPS54_MOUSE
|
||||||
| NC score | 0.027247 (rank : 26) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5SPW0, Q8BPB3, Q8CFZ7, Q8CHL5, Q8R3R4, Q8R3X1 | Gene names | Vps54 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vacuolar protein sorting-associated protein 54 (Tumor antigen SLP-8p homolog). | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.027170 (rank : 27) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
NPAS4_MOUSE
|
||||||
| NC score | 0.026797 (rank : 28) | θ value | 1.81305 (rank : 25) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
HORN_MOUSE
|
||||||
| NC score | 0.026665 (rank : 29) | θ value | 1.06291 (rank : 15) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8VHD8 | Gene names | Hrnr | |||
|
Domain Architecture |

|
|||||
| Description | Hornerin. | |||||
|
CEP57_MOUSE
|
||||||
| NC score | 0.026283 (rank : 30) | θ value | 1.81305 (rank : 21) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 580 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8CEE0, Q6ZQJ3, Q7TN18, Q80X65, Q810F2, Q9D4J4, Q9D5S4, Q9D5W5 | Gene names | Cep57, Kiaa0092, Tsp57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 57 kDa (Cep57 protein) (Testis-specific protein 57) (Translokin). | |||||
|
CF152_HUMAN
|
||||||
| NC score | 0.025011 (rank : 31) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q86VQ0, Q9BWX7 | Gene names | C6orf152 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf152. | |||||
|
BUB1_HUMAN
|
||||||
| NC score | 0.024676 (rank : 32) | θ value | 1.06291 (rank : 14) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O43683, O43430, O43643, O60626 | Gene names | BUB1, BUB1L | |||
|
Domain Architecture |

|
|||||
| Description | Mitotic checkpoint serine/threonine-protein kinase BUB1 (EC 2.7.11.1) (hBUB1) (BUB1A). | |||||
|
CA026_HUMAN
|
||||||
| NC score | 0.024321 (rank : 33) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5T5J6, Q8NEK9, Q9BZQ7, Q9NXQ0 | Gene names | C1orf26 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C1orf26. | |||||
|
HNRPQ_MOUSE
|
||||||
| NC score | 0.024042 (rank : 34) | θ value | 4.03905 (rank : 41) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7TMK9, O88991, Q80YV6, Q8BGP1, Q8C5K6, Q8CGC2, Q91ZR0, Q99KU9, Q9QYF4 | Gene names | Syncrip, Hnrpq, Nsap1, Nsap1l | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein Q (hnRNP Q) (hnRNP-Q) (Synaptotagmin-binding, cytoplasmic RNA-interacting protein) (Glycine- and tyrosine-rich RNA-binding protein) (GRY-RBP) (NS1-associated protein 1) (pp68). | |||||
|
PF21B_MOUSE
|
||||||
| NC score | 0.023466 (rank : 35) | θ value | 1.81305 (rank : 26) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8C966 | Gene names | Phf21b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
RFA1_HUMAN
|
||||||
| NC score | 0.023046 (rank : 36) | θ value | 4.03905 (rank : 44) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P27694, Q59ES9 | Gene names | RPA1, REPA1, RPA70 | |||
|
Domain Architecture |

|
|||||
| Description | Replication protein A 70 kDa DNA-binding subunit (RP-A) (RF-A) (Replication factor-A protein 1) (Single-stranded DNA-binding protein) (p70). | |||||
|
HNRPQ_HUMAN
|
||||||
| NC score | 0.023043 (rank : 37) | θ value | 5.27518 (rank : 47) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60506, Q53H05, Q5TCG2, Q8IW78, Q8N599, Q96LC1, Q96LC2, Q9Y583 | Gene names | SYNCRIP, HNRPQ, NSAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein Q (hnRNP Q) (hnRNP-Q) (Synaptotagmin-binding, cytoplasmic RNA-interacting protein) (Glycine- and tyrosine-rich RNA-binding protein) (GRY-RBP) (NS1-associated protein 1). | |||||
|
APEH_MOUSE
|
||||||
| NC score | 0.022949 (rank : 38) | θ value | 3.0926 (rank : 31) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R146, Q8K029, Q8R0M9 | Gene names | Apeh | |||
|
Domain Architecture |

|
|||||
| Description | Acylamino-acid-releasing enzyme (EC 3.4.19.1) (AARE) (Acyl-peptide hydrolase) (APH) (Acylaminoacyl-peptidase). | |||||
|
PSMF1_MOUSE
|
||||||
| NC score | 0.022216 (rank : 39) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BHL8, Q8C0G9 | Gene names | Psmf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteasome inhibitor PI31 subunit. | |||||
|
CEP57_HUMAN
|
||||||
| NC score | 0.021522 (rank : 40) | θ value | 5.27518 (rank : 45) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 576 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q86XR8, Q14704, Q8IXP0, Q9BVF9 | Gene names | CEP57, KIAA0092, TSP57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 57 kDa (Cep57 protein) (Testis-specific protein 57) (Translokin) (FGF2-interacting protein). | |||||
|
CD34_MOUSE
|
||||||
| NC score | 0.020176 (rank : 41) | θ value | 6.88961 (rank : 54) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64314, Q62550, Q62551 | Gene names | Cd34 | |||
|
Domain Architecture |
|
|||||
| Description | Hematopoietic progenitor cell antigen CD34 precursor. | |||||
|
K1543_HUMAN
|
||||||
| NC score | 0.020175 (rank : 42) | θ value | 8.99809 (rank : 69) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9P1Y5, Q8NDF1 | Gene names | KIAA1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
CCD21_HUMAN
|
||||||
| NC score | 0.019018 (rank : 43) | θ value | 0.365318 (rank : 5) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1228 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6P2H3, Q5VY68, Q5VY70, Q9H6Q1, Q9H828, Q9UF52 | Gene names | CCDC21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
VP13B_HUMAN
|
||||||
| NC score | 0.018314 (rank : 44) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7Z7G8, Q709C6, Q709C7, Q7Z7G4, Q7Z7G5, Q7Z7G6, Q7Z7G7, Q8NB77, Q9NWV1, Q9Y4E7 | Gene names | VPS13B, CHS1, COH1, KIAA0532 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vacuolar protein sorting-associated protein 13B (Cohen syndrome protein 1). | |||||
|
SYN1_HUMAN
|
||||||
| NC score | 0.017115 (rank : 45) | θ value | 3.0926 (rank : 34) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P17600, O75825, Q5H9A9 | Gene names | SYN1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I) (Brain protein 4.1). | |||||
|
ILDR1_MOUSE
|
||||||
| NC score | 0.016524 (rank : 46) | θ value | 6.88961 (rank : 59) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CBR1, Q6PFB3, Q8CB39, Q91VS0 | Gene names | Ildr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immunoglobulin-like domain-containing receptor 1 precursor. | |||||
|
CASL_MOUSE
|
||||||
| NC score | 0.014917 (rank : 47) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O35177, Q8BJL8, Q8BK90, Q8BL52, Q8BM94, Q8BMI9, Q99KE7 | Gene names | Nedd9, Casl | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of filamentation 1 (MEF1) (CRK-associated substrate-related protein) (CAS-L) (p105) (Protein NEDD9) (Neural precursor cell expressed developmentally down-regulated protein 9). | |||||
|
SNIP_MOUSE
|
||||||
| NC score | 0.014821 (rank : 48) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9QWI6, O70298 | Gene names | P140, Kiaa1684 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | p130Cas-associated protein (p140Cap) (SNAP-25-interacting protein) (SNIP). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.014732 (rank : 49) | θ value | 8.99809 (rank : 71) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
SNG4_HUMAN
|
||||||
| NC score | 0.014371 (rank : 50) | θ value | 6.88961 (rank : 60) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95473 | Gene names | SYNGR4 | |||
|
Domain Architecture |
|
|||||
| Description | Synaptogyrin-4. | |||||
|
FA35A_MOUSE
|
||||||
| NC score | 0.013876 (rank : 51) | θ value | 8.99809 (rank : 66) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3UEN2 | Gene names | Fam35a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM35A. | |||||
|
SHAN1_HUMAN
|
||||||
| NC score | 0.013456 (rank : 52) | θ value | 1.81305 (rank : 27) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Y566, Q9NYW9 | Gene names | SHANK1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 1 (Shank1) (Somatostatin receptor-interacting protein) (SSTR-interacting protein) (SSTRIP). | |||||
|
KLC2_HUMAN
|
||||||
| NC score | 0.012149 (rank : 53) | θ value | 4.03905 (rank : 42) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H0B6, Q9H9C8, Q9HA20 | Gene names | KLC2 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin light chain 2 (KLC 2). | |||||
|
P85A_MOUSE
|
||||||
| NC score | 0.011894 (rank : 54) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P26450 | Gene names | Pik3r1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit alpha (PI3-kinase p85-subunit alpha) (PtdIns-3-kinase p85-alpha) (PI3K). | |||||
|
P85A_HUMAN
|
||||||
| NC score | 0.011677 (rank : 55) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P27986 | Gene names | PIK3R1, GRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit alpha (PI3-kinase p85-subunit alpha) (PtdIns-3-kinase p85-alpha) (PI3K). | |||||
|
CAC1A_MOUSE
|
||||||
| NC score | 0.011465 (rank : 56) | θ value | 8.99809 (rank : 64) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P97445 | Gene names | Cacna1a, Cach4, Cacn3, Cacnl1a4, Ccha1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
RFIP3_HUMAN
|
||||||
| NC score | 0.010661 (rank : 57) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1055 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O75154, Q4VXV7, Q9H155, Q9H1G0, Q9NUI0 | Gene names | RAB11FIP3, KIAA0665 | |||
|
Domain Architecture |

|
|||||
| Description | Rab11 family-interacting protein 3 (Rab11-FIP3) (EF hands-containing Rab-interacting protein) (Eferin). | |||||
|
ARNT_MOUSE
|
||||||
| NC score | 0.010396 (rank : 58) | θ value | 8.99809 (rank : 63) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
IF4G1_MOUSE
|
||||||
| NC score | 0.010213 (rank : 59) | θ value | 8.99809 (rank : 68) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6NZJ6, Q6NZN8, Q8BW99 | Gene names | Eif4g1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1). | |||||
|
CLC4F_MOUSE
|
||||||
| NC score | 0.009806 (rank : 60) | θ value | 4.03905 (rank : 38) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 572 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P70194, Q8BLZ8 | Gene names | Clec4f, Clecsf13, Kclr | |||
|
Domain Architecture |

|
|||||
| Description | C-type lectin domain family 4 member F (C-type lectin superfamily member 13) (C-type lectin 13) (Kupffer cell receptor). | |||||
|
LIPA1_HUMAN
|
||||||
| NC score | 0.009692 (rank : 61) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1294 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q13136, Q13135, Q14567, Q8N4I2 | Gene names | PPFIA1, LIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-1) (PTPRF-interacting protein alpha-1) (LAR-interacting protein 1) (LIP.1). | |||||
|
KI13A_MOUSE
|
||||||
| NC score | 0.008949 (rank : 62) | θ value | 5.27518 (rank : 48) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9EQW7, O35062 | Gene names | Kif13a | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13A. | |||||
|
ADDB_MOUSE
|
||||||
| NC score | 0.007796 (rank : 63) | θ value | 8.99809 (rank : 62) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 574 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9QYB8, Q3U0E1, Q80VH9, Q8C1C4, Q9CXE3, Q9JLE5, Q9QYB7 | Gene names | Add2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adducin (Erythrocyte adducin subunit beta) (Add97). | |||||
|
DGKI_HUMAN
|
||||||
| NC score | 0.007643 (rank : 64) | θ value | 6.88961 (rank : 55) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O75912, Q9NZ49 | Gene names | DGKI | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase iota (EC 2.7.1.107) (Diglyceride kinase iota) (DGK-iota) (DAG kinase iota). | |||||
|
KI13A_HUMAN
|
||||||
| NC score | 0.007395 (rank : 65) | θ value | 8.99809 (rank : 70) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9H1H9, Q9H1H8 | Gene names | KIF13A, RBKIN | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13A (Kinesin-like protein RBKIN). | |||||
|
ZN592_HUMAN
|
||||||
| NC score | 0.006979 (rank : 66) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 762 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92610 | Gene names | ZNF592, KIAA0211 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 592. | |||||
|
GOGB1_HUMAN
|
||||||
| NC score | 0.006687 (rank : 67) | θ value | 5.27518 (rank : 46) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
ZN646_HUMAN
|
||||||
| NC score | 0.006089 (rank : 68) | θ value | 0.813845 (rank : 13) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 813 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O15015 | Gene names | ZNF646, KIAA0296 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 646. | |||||
|
SOX14_HUMAN
|
||||||
| NC score | 0.005544 (rank : 69) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95416, Q3KPH7 | Gene names | SOX14, SOX28 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-14. | |||||
|
ZBT20_HUMAN
|
||||||
| NC score | 0.005319 (rank : 70) | θ value | 3.0926 (rank : 35) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 835 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9HC78 | Gene names | ZBTB20, DPZF, ZNF288 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 20 (Zinc finger protein 288) (Dendritic-derived BTB/POZ zinc finger protein). | |||||
|
MLTK_HUMAN
|
||||||
| NC score | 0.005098 (rank : 71) | θ value | 1.81305 (rank : 24) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 911 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NYL2, Q53SX1, Q580W8, Q59GY5, Q86YW8, Q9HCC4, Q9HCC5, Q9HDD2, Q9NYE9 | Gene names | MLTK, ZAK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase MLT (EC 2.7.11.25) (MLK-like mitogen-activated protein triple kinase) (Leucine zipper- and sterile alpha motif-containing kinase) (Sterile alpha motif- and leucine zipper-containing kinase AZK) (Mixed lineage kinase-related kinase) (MLK-related kinase) (MRK) (Cervical cancer suppressor gene 4 protein). | |||||
|
CHD9_MOUSE
|
||||||
| NC score | 0.005027 (rank : 72) | θ value | 8.99809 (rank : 65) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BYH8, Q7TMV5, Q8BJG8, Q8BZJ2, Q8CHG8 | Gene names | Chd9, Kiaa0308, Pric320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha- interacting complex protein 320 kDa). | |||||
|
ZN518_HUMAN
|
||||||
| NC score | 0.003742 (rank : 73) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6AHZ1, O15044, Q32MP4 | Gene names | ZNF518, KIAA0335 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 518. | |||||
|
MAST2_MOUSE
|
||||||
| NC score | 0.003570 (rank : 74) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1110 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q60592, Q6P9M1, Q8CHD1 | Gene names | Mast2, Mast205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 2 (EC 2.7.11.1). | |||||
|
FAK2_MOUSE
|
||||||
| NC score | 0.003458 (rank : 75) | θ value | 6.88961 (rank : 57) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9QVP9 | Gene names | Ptk2b, Fak2, Pyk2, Raftk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein tyrosine kinase 2 beta (EC 2.7.10.2) (Focal adhesion kinase 2) (FADK 2) (Proline-rich tyrosine kinase 2) (Cell adhesion kinase beta) (CAK beta) (Calcium-dependent tyrosine kinase) (CADTK) (Related adhesion focal tyrosine kinase) (RAFTK). | |||||
|
FAK2_HUMAN
|
||||||
| NC score | 0.003210 (rank : 76) | θ value | 8.99809 (rank : 67) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 847 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q14289, Q13475, Q14290, Q16709 | Gene names | PTK2B, FAK2, PYK2, RAFTK | |||
|
Domain Architecture |

|
|||||
| Description | Protein tyrosine kinase 2 beta (EC 2.7.10.2) (Focal adhesion kinase 2) (FADK 2) (Proline-rich tyrosine kinase 2) (Cell adhesion kinase beta) (CAK beta) (Calcium-dependent tyrosine kinase) (CADTK) (Related adhesion focal tyrosine kinase) (RAFTK). | |||||
|
PDPK1_HUMAN
|
||||||
| NC score | 0.003182 (rank : 77) | θ value | 5.27518 (rank : 50) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O15530 | Gene names | PDPK1, PDK1 | |||
|
Domain Architecture |

|
|||||
| Description | 3-phosphoinositide-dependent protein kinase 1 (EC 2.7.11.1) (hPDK1). | |||||
|
PDPK1_MOUSE
|
||||||
| NC score | 0.003171 (rank : 78) | θ value | 5.27518 (rank : 51) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 852 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Z2A0, Q9R1D8, Q9R215 | Gene names | Pdpk1, Pdk1 | |||
|
Domain Architecture |

|
|||||
| Description | 3-phosphoinositide-dependent protein kinase 1 (EC 2.7.11.1) (mPDK1). | |||||
|
ZN445_HUMAN
|
||||||
| NC score | 0.000708 (rank : 79) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P59923 | Gene names | ZNF445, ZNF168 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 445. | |||||