Please be patient as the page loads
|
FA13C_MOUSE
|
||||||
| SwissProt Accessions | Q9DBR2 | Gene names | Fam13c1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13C1. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
FA13C_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.964783 (rank : 2) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8NE31, Q99787 | Gene names | FAM13C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13C1. | |||||
|
FA13C_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q9DBR2 | Gene names | Fam13c1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13C1. | |||||
|
FA13A_HUMAN
|
||||||
| θ value | 6.34462e-54 (rank : 3) | NC score | 0.840403 (rank : 4) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O94988, Q8NBA3 | Gene names | FAM13A1, KIAA0914 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13A1. | |||||
|
FA13A_MOUSE
|
||||||
| θ value | 2.66562e-52 (rank : 4) | NC score | 0.845183 (rank : 3) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BGI4, Q8BZ91 | Gene names | Fam13a1, Precm1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13A1 (Precm1 protein). | |||||
|
CE005_MOUSE
|
||||||
| θ value | 5.95217e-44 (rank : 5) | NC score | 0.617555 (rank : 5) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8K2H3 | Gene names | ||||
|
Domain Architecture |
|
|||||
| Description | Protein C5orf5 homolog. | |||||
|
CE005_HUMAN
|
||||||
| θ value | 2.76489e-41 (rank : 6) | NC score | 0.613744 (rank : 6) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NYF5, Q6PGQ2, Q9P0I7 | Gene names | C5orf5 | |||
|
Domain Architecture |
|
|||||
| Description | Protein C5orf5 (GAP-like protein N61). | |||||
|
MCM3A_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 7) | NC score | 0.070213 (rank : 7) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60318, Q9UMT4 | Gene names | MCM3AP, GANP, KIAA0572, MAP80 | |||
|
Domain Architecture |

|
|||||
| Description | 80 kDa MCM3-associated protein (GANP protein). | |||||
|
TXND2_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 8) | NC score | 0.059232 (rank : 12) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
CLIC6_HUMAN
|
||||||
| θ value | 0.125558 (rank : 9) | NC score | 0.030804 (rank : 44) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96NY7, Q8IX31 | Gene names | CLIC6, CLIC1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel 6. | |||||
|
EP15R_MOUSE
|
||||||
| θ value | 0.125558 (rank : 10) | NC score | 0.025956 (rank : 51) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q60902, Q8CB60, Q8CB70, Q91WH8 | Gene names | Eps15l1, Eps15-rs, Eps15R | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal growth factor receptor substrate 15-like 1 (Eps15-related protein) (Eps15R) (Epidermal growth factor receptor pathway substrate 15-related sequence) (Eps15-rs). | |||||
|
ITB3_HUMAN
|
||||||
| θ value | 0.125558 (rank : 11) | NC score | 0.023310 (rank : 59) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P05106, O15495, Q13413, Q14648, Q16499 | Gene names | ITGB3, GP3A | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-3 precursor (Platelet membrane glycoprotein IIIa) (GPIIIa) (CD61 antigen). | |||||
|
BRPF3_HUMAN
|
||||||
| θ value | 0.21417 (rank : 12) | NC score | 0.027714 (rank : 47) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9ULD4, Q5R3K8 | Gene names | BRPF3, KIAA1286 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and PHD finger-containing protein 3. | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 0.279714 (rank : 13) | NC score | 0.038219 (rank : 35) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
G3BP_MOUSE
|
||||||
| θ value | 0.365318 (rank : 14) | NC score | 0.033891 (rank : 39) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P97855 | Gene names | G3bp, G3bp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras-GTPase-activating protein-binding protein 1 (EC 3.6.1.-) (ATP- dependent DNA helicase VIII) (GAP SH3-domain-binding protein 1) (G3BP- 1) (HDH-VIII). | |||||
|
TITIN_HUMAN
|
||||||
| θ value | 0.365318 (rank : 15) | NC score | 0.011574 (rank : 84) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 0.62314 (rank : 16) | NC score | 0.033799 (rank : 40) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
NCOR2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 17) | NC score | 0.041355 (rank : 32) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
SETBP_HUMAN
|
||||||
| θ value | 0.62314 (rank : 18) | NC score | 0.052510 (rank : 19) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Y6X0, Q9UEF3 | Gene names | SETBP1, KIAA0437 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SET-binding protein (SEB). | |||||
|
X3CL1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 19) | NC score | 0.037509 (rank : 36) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O35188, O35933 | Gene names | Cx3cl1, Cx3c, Fkn, Scyd1 | |||
|
Domain Architecture |
|
|||||
| Description | Fractalkine precursor (CX3CL1) (Neurotactin) (CX3C membrane-anchored chemokine) (Small inducible cytokine D1). | |||||
|
ZF106_HUMAN
|
||||||
| θ value | 0.813845 (rank : 20) | NC score | 0.022290 (rank : 61) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9H2Y7, Q6NSD9, Q6PEK1, Q86T43, Q86T45, Q86T50, Q86T58, Q86TA9, Q96M37, Q9H7B8 | Gene names | ZFP106, ZNF474 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 106 homolog (Zfp-106) (Zinc finger protein 474). | |||||
|
APLP1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 21) | NC score | 0.026362 (rank : 50) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P51693, O00113, Q96A92 | Gene names | APLP1 | |||
|
Domain Architecture |
|
|||||
| Description | Amyloid-like protein 1 precursor (APLP) (APLP-1) [Contains: C30]. | |||||
|
DIDO1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 22) | NC score | 0.039109 (rank : 34) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
DLEC1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 23) | NC score | 0.031535 (rank : 43) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y238, Q9NSW0, Q9NTG5 | Gene names | DLEC1, DLC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Deleted in lung and esophageal cancer protein 1 (Deleted in lung cancer protein 1) (DLC-1). | |||||
|
PKHA4_MOUSE
|
||||||
| θ value | 1.38821 (rank : 24) | NC score | 0.025585 (rank : 55) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8VC98, Q8R3N3 | Gene names | Plekha4, Pepp1 | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin homology domain-containing family A member 4 (Phosphoinositol 3-phosphate-binding protein 1) (PEPP-1). | |||||
|
PRGC2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 25) | NC score | 0.036061 (rank : 37) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8VHJ7, Q8C1C0 | Gene names | Ppargc1b, Errl1, Pgc1, Pgc1b, Ppargc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta protein) (PGC-1beta) (ERR ligand 1) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
WDR60_HUMAN
|
||||||
| θ value | 1.38821 (rank : 26) | NC score | 0.026982 (rank : 48) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8WVS4, Q9NW58 | Gene names | WDR60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 60. | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 1.81305 (rank : 27) | NC score | 0.034821 (rank : 38) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
SALL2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 28) | NC score | 0.004817 (rank : 92) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9QX96 | Gene names | Sall2, Sal2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 2 (Spalt-like protein 2) (MSal-2). | |||||
|
SETBP_MOUSE
|
||||||
| θ value | 1.81305 (rank : 29) | NC score | 0.047098 (rank : 26) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Z180, Q66JL8 | Gene names | Setbp1, Kiaa0437 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SET-binding protein (SEB). | |||||
|
SYN1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 30) | NC score | 0.025662 (rank : 54) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P17600, O75825, Q5H9A9 | Gene names | SYN1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I) (Brain protein 4.1). | |||||
|
BASP_HUMAN
|
||||||
| θ value | 2.36792 (rank : 31) | NC score | 0.041594 (rank : 31) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
IF39_HUMAN
|
||||||
| θ value | 2.36792 (rank : 32) | NC score | 0.039176 (rank : 33) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P55884, Q9UMF9 | Gene names | EIF3S9 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 9 (eIF-3 eta) (eIF3 p116) (eIF3 p110) (eIF3b) (Prt1 homolog) (hPrt1). | |||||
|
K0256_HUMAN
|
||||||
| θ value | 2.36792 (rank : 33) | NC score | 0.022527 (rank : 60) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q93073, Q8N767 | Gene names | KIAA0256 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0256. | |||||
|
PRG4_MOUSE
|
||||||
| θ value | 2.36792 (rank : 34) | NC score | 0.044453 (rank : 27) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
PRP1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.029939 (rank : 45) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
S30BP_HUMAN
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.029624 (rank : 46) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UHR5, Q8N1R5, Q8TDR8, Q96D27 | Gene names | SAP30BP, HCNGP, HTRG, HTRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAP30-binding protein (Transcriptional regulator protein HCNGP). | |||||
|
TCAL5_HUMAN
|
||||||
| θ value | 2.36792 (rank : 37) | NC score | 0.024718 (rank : 57) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
AFF1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 38) | NC score | 0.041899 (rank : 29) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
CSMD3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 39) | NC score | 0.007491 (rank : 87) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z407, Q96PZ3 | Gene names | CSMD3, KIAA1894 | |||
|
Domain Architecture |

|
|||||
| Description | CUB and sushi domain-containing protein 3 precursor (CUB and sushi multiple domains protein 3). | |||||
|
ESCO1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 40) | NC score | 0.023897 (rank : 58) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5FWF5, Q69YG4, Q69YS3, Q6IMD7, Q8N3Z5, Q8NBG2, Q96PX7 | Gene names | ESCO1, EFO1, KIAA1911 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyltransferase ESCO1 (EC 2.3.1.-) (Establishment of cohesion 1 homolog 1) (ECO1 homolog 1) (ESO1 homolog 1) (Establishment factor- like protein 1) (EFO1p) (hEFO1) (CTF7 homolog 1). | |||||
|
NUCL_MOUSE
|
||||||
| θ value | 3.0926 (rank : 41) | NC score | 0.019843 (rank : 68) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P09405, Q548M9, Q61991, Q99K50 | Gene names | Ncl, Nuc | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolin (Protein C23). | |||||
|
PEX14_HUMAN
|
||||||
| θ value | 3.0926 (rank : 42) | NC score | 0.025887 (rank : 53) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75381, Q8WX51 | Gene names | PEX14 | |||
|
Domain Architecture |
|
|||||
| Description | Peroxisomal membrane protein PEX14 (Peroxin-14) (Peroxisomal membrane anchor protein PEX14) (PTS1 receptor docking protein). | |||||
|
TRPM7_MOUSE
|
||||||
| θ value | 3.0926 (rank : 43) | NC score | 0.011879 (rank : 83) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q923J1, Q8C7S7, Q8CE54, Q921Y5, Q925B2, Q9CUT2, Q9JLQ1 | Gene names | Trpm7, Chak, Ltrpc7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 7 (EC 2.7.11.1) (Long transient receptor potential channel 7) (LTrpC7) (Channel-kinase 1) (Transient receptor potential-phospholipase C- interacting kinase) (TRP-PLIK). | |||||
|
TXND2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 44) | NC score | 0.042997 (rank : 28) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
ARHG1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 45) | NC score | 0.017354 (rank : 73) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q61210, O89074, Q80YE8, Q80YE9, Q91VL3 | Gene names | Arhgef1, Lbcl2, Lsc | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 1 (Lymphoid blast crisis-like 2) (Lbc's second cousin). | |||||
|
BAT2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.032562 (rank : 41) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
CP053_HUMAN
|
||||||
| θ value | 4.03905 (rank : 47) | NC score | 0.041690 (rank : 30) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BTK6 | Gene names | C16orf53 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf53. | |||||
|
IF4G1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 48) | NC score | 0.017593 (rank : 72) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6NZJ6, Q6NZN8, Q8BW99 | Gene names | Eif4g1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1). | |||||
|
ITB3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 49) | NC score | 0.017342 (rank : 74) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O54890 | Gene names | Itgb3 | |||
|
Domain Architecture |
|
|||||
| Description | Integrin beta-3 precursor (Platelet membrane glycoprotein IIIa) (GPIIIa) (CD61 antigen). | |||||
|
MYBB_HUMAN
|
||||||
| θ value | 4.03905 (rank : 50) | NC score | 0.024922 (rank : 56) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P10244 | Gene names | MYBL2, BMYB | |||
|
Domain Architecture |

|
|||||
| Description | Myb-related protein B (B-Myb). | |||||
|
RECQ5_HUMAN
|
||||||
| θ value | 4.03905 (rank : 51) | NC score | 0.020511 (rank : 65) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O94762, Q9P1W7, Q9UNC8 | Gene names | RECQL5, RECQ5 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase Q5 (EC 3.6.1.-) (RecQ protein-like 5) (RecQ5). | |||||
|
ADCY4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 52) | NC score | 0.006465 (rank : 89) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NFM4, Q96ML7 | Gene names | ADCY4 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 4 (EC 4.6.1.1) (Adenylate cyclase type IV) (ATP pyrophosphate-lyase 4) (Adenylyl cyclase 4). | |||||
|
ARHG1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.015740 (rank : 77) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92888, O00513, Q8N4J4, Q96BF4, Q96F17, Q9BSB1 | Gene names | ARHGEF1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 1 (p115-RhoGEF) (p115RhoGEF) (115 kDa guanine nucleotide exchange factor) (Sub1.5). | |||||
|
CHD4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | 0.009660 (rank : 86) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6PDQ2 | Gene names | Chd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (CHD-4). | |||||
|
MEN1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 55) | NC score | 0.018622 (rank : 70) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O88559 | Gene names | Men1 | |||
|
Domain Architecture |

|
|||||
| Description | Menin. | |||||
|
NRK_MOUSE
|
||||||
| θ value | 5.27518 (rank : 56) | NC score | 0.001380 (rank : 94) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 899 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9R0G8, Q6NV55, Q8C9S9, Q9R0S4 | Gene names | Nrk, Nesk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nik-related protein kinase (EC 2.7.11.1) (Nck-interacting kinase-like embryo specific kinase) (NIK-like embryo-specific kinase) (NESK). | |||||
|
PTN1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 57) | NC score | 0.004990 (rank : 91) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P18031, Q5TGD8, Q9BQV9, Q9NQQ4 | Gene names | PTPN1, PTP1B | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 1 (EC 3.1.3.48) (Protein-tyrosine phosphatase 1B) (PTP-1B). | |||||
|
SYN3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 58) | NC score | 0.021923 (rank : 62) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O14994 | Gene names | SYN3 | |||
|
Domain Architecture |
|
|||||
| Description | Synapsin-3 (Synapsin III). | |||||
|
TAIP2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 59) | NC score | 0.013417 (rank : 81) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P59055 | Gene names | Taip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TGF-beta-induced apoptosis protein 2 (TAIP-2). | |||||
|
CD2L7_HUMAN
|
||||||
| θ value | 6.88961 (rank : 60) | NC score | 0.001896 (rank : 93) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1275 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NYV4, O94978 | Gene names | CRKRS, CRK7, KIAA0904 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle 2-related protein kinase 7 (EC 2.7.11.22) (CDC2- related protein kinase 7) (Cdc2-related kinase, arginine/serine-rich) (CrkRS). | |||||
|
CMGA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | 0.019911 (rank : 67) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P10645, Q96E84, Q96GL7, Q9BQB5 | Gene names | CHGA | |||
|
Domain Architecture |

|
|||||
| Description | Chromogranin A precursor (CgA) (Pituitary secretory protein I) (SP-I) [Contains: Vasostatin-1 (Vasostatin I); Vasostatin-2 (Vasostatin II); EA-92; ES-43; Pancreastatin; SS-18; WA-8; WE-14; LF-19; AL-11; GV-19; GR-44; ER-37]. | |||||
|
IFT74_HUMAN
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.014311 (rank : 80) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96LB3, Q6PGQ8, Q9H643, Q9H8G7 | Gene names | IFT74, CCDC2, CMG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 74 homolog (Coiled-coil domain-containing protein 2) (Capillary morphogenesis protein 1) (CMG-1). | |||||
|
IRF3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 63) | NC score | 0.009852 (rank : 85) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q14653 | Gene names | IRF3 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon regulatory factor 3 (IRF-3). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 64) | NC score | 0.026853 (rank : 49) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 65) | NC score | 0.032309 (rank : 42) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
K1045_HUMAN
|
||||||
| θ value | 6.88961 (rank : 66) | NC score | 0.016236 (rank : 75) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UPV7, Q58FE9, Q5T662 | Gene names | KIAA1045 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1045. | |||||
|
MCM3A_MOUSE
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.048199 (rank : 25) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9WUU9 | Gene names | Mcm3ap, Ganp, Map80 | |||
|
Domain Architecture |

|
|||||
| Description | 80 kDa MCM3-associated protein (GANP protein). | |||||
|
MKL1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.012132 (rank : 82) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8K4J6 | Gene names | Mkl1, Bsac | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 1 (Myocardin-related transcription factor A) (MRTF-A) (Megakaryoblastic leukemia 1 protein homolog) (Basic SAP coiled-coil transcription activator). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.025932 (rank : 52) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
TCAL3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | 0.020065 (rank : 66) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q969E4 | Gene names | TCEAL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 3 (TCEA-like protein 3) (Transcription elongation factor S-II protein-like 3). | |||||
|
UTRO_HUMAN
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.005083 (rank : 90) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P46939 | Gene names | UTRN, DMDL | |||
|
Domain Architecture |

|
|||||
| Description | Utrophin (Dystrophin-related protein 1) (DRP1) (DRP). | |||||
|
AFF2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.019123 (rank : 69) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O55112 | Gene names | Aff2, Fmr2, Ox19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation protein 2 homolog) (Protein FMR-2) (FMR2P) (Protein Ox19). | |||||
|
BCAS1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.015470 (rank : 79) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75363, Q68CZ3 | Gene names | BCAS1, AIBC1, NABC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Breast carcinoma amplified sequence 1 (Novel amplified in breast cancer 1) (Amplified and overexpressed in breast cancer). | |||||
|
JHD1B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.006554 (rank : 88) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6P1G2, Q3V396, Q6PFD0, Q6ZPE8, Q9CSF7, Q9QZN6 | Gene names | Fbxl10, Fbl10, Jhdm1b, Kiaa3014 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10). | |||||
|
LAS1L_HUMAN
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.018010 (rank : 71) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y4W2, Q5JXQ0, Q8TEN5, Q9H9V5 | Gene names | LAS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LAS1-like protein. | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.021214 (rank : 63) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
ORC2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.016160 (rank : 76) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q60862 | Gene names | Orc2l, Orc2 | |||
|
Domain Architecture |

|
|||||
| Description | Origin recognition complex subunit 2. | |||||
|
RIMS1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.015681 (rank : 78) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 437 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86UR5, O15048, Q8TDY9, Q8TDZ5, Q9HBA1, Q9HBA2, Q9HBA3, Q9HBA4, Q9HBA5, Q9HBA6 | Gene names | RIMS1, KIAA0340, RIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 1 (Rab3-interacting molecule 1) (RIM 1). | |||||
|
SRCA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.020563 (rank : 64) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q86TD4 | Gene names | SRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
GMIP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 80) | NC score | 0.057106 (rank : 15) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9P107 | Gene names | GMIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GEM-interacting protein (GMIP). | |||||
|
GMIP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.058926 (rank : 13) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6PGG2, Q6P9S3 | Gene names | Gmip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GEM-interacting protein (GMIP). | |||||
|
K1688_HUMAN
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.052077 (rank : 20) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9C0H5 | Gene names | KIAA1688 | |||
|
Domain Architecture |

|
|||||
| Description | Protein KIAA1688. | |||||
|
K1688_MOUSE
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.052050 (rank : 21) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P59281, Q69ZD4, Q6P9R6 | Gene names | Kiaa1688, D15Wsu169e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1688. | |||||
|
RGAP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.056753 (rank : 17) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H0H5, Q6PJ26, Q9NWN2, Q9P250, Q9P2W2 | Gene names | RACGAP1, KIAA1478, MGCRACGAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rac GTPase-activating protein 1 (MgcRacGAP). | |||||
|
RHG01_HUMAN
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.068441 (rank : 8) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q07960 | Gene names | ARHGAP1, CDC42GAP, RHOGAP1 | |||
|
Domain Architecture |
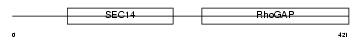
|
|||||
| Description | Rho-GTPase-activating protein 1 (GTPase-activating protein rhoOGAP) (Rho-related small GTPase protein activator) (CDC42 GTPase-activating protein) (p50-RhoGAP). | |||||
|
RHG05_HUMAN
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.051751 (rank : 24) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13017 | Gene names | ARHGAP5, RHOGAP5 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 5 (p190-B). | |||||
|
RHG05_MOUSE
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.051917 (rank : 22) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P97393 | Gene names | Arhgap5, Rhogap5 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 5 (p190-B). | |||||
|
RHG06_HUMAN
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.057807 (rank : 14) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43182, O43437, Q9P1B3, Q9UK81, Q9UK82 | Gene names | ARHGAP6, RHOGAP6 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 6 (Rho-type GTPase-activating protein RhoGAPX-1). | |||||
|
RHG06_MOUSE
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.057043 (rank : 16) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O54834, Q9QZL8 | Gene names | Arhgap6 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 6 (Rho-type GTPase-activating protein RhoGAPX-1). | |||||
|
RHG08_HUMAN
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.051884 (rank : 23) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NSG0, O75983, O95695, Q96RW1, Q96RW2, Q9HA49, Q9HC46, Q9NVX8, Q9NXL1, Q9UH20 | Gene names | ARHGAP8 | |||
|
Domain Architecture |
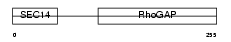
|
|||||
| Description | Rho-GTPase-activating protein 8. | |||||
|
RHG08_MOUSE
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.054408 (rank : 18) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CXP4, Q99JY7 | Gene names | Arhgap8 | |||
|
Domain Architecture |
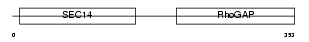
|
|||||
| Description | Rho-GTPase-activating protein 8. | |||||
|
RHG18_MOUSE
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.059688 (rank : 11) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K0Q5, Q8BP03, Q8R196 | Gene names | Arhgap18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho-GTPase-activating protein 18. | |||||
|
RHG25_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.063363 (rank : 10) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 550 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P42331, Q8IXQ2 | Gene names | ARHGAP25, KIAA0053 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
RHG25_MOUSE
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.066741 (rank : 9) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BYW1, Q2VPR2, Q3TE04, Q3TVX0, Q3UVB7, Q6A0E0, Q8BX98 | Gene names | Arhgap25, Kiaa0053 | |||
|
Domain Architecture |
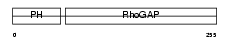
|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
FA13C_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q9DBR2 | Gene names | Fam13c1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13C1. | |||||
|
FA13C_HUMAN
|
||||||
| NC score | 0.964783 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8NE31, Q99787 | Gene names | FAM13C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13C1. | |||||
|
FA13A_MOUSE
|
||||||
| NC score | 0.845183 (rank : 3) | θ value | 2.66562e-52 (rank : 4) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BGI4, Q8BZ91 | Gene names | Fam13a1, Precm1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13A1 (Precm1 protein). | |||||
|
FA13A_HUMAN
|
||||||
| NC score | 0.840403 (rank : 4) | θ value | 6.34462e-54 (rank : 3) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O94988, Q8NBA3 | Gene names | FAM13A1, KIAA0914 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM13A1. | |||||
|
CE005_MOUSE
|
||||||
| NC score | 0.617555 (rank : 5) | θ value | 5.95217e-44 (rank : 5) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8K2H3 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Protein C5orf5 homolog. | |||||
|
CE005_HUMAN
|
||||||
| NC score | 0.613744 (rank : 6) | θ value | 2.76489e-41 (rank : 6) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NYF5, Q6PGQ2, Q9P0I7 | Gene names | C5orf5 | |||
|
Domain Architecture |

|
|||||
| Description | Protein C5orf5 (GAP-like protein N61). | |||||
|
MCM3A_HUMAN
|
||||||
| NC score | 0.070213 (rank : 7) | θ value | 0.0193708 (rank : 7) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60318, Q9UMT4 | Gene names | MCM3AP, GANP, KIAA0572, MAP80 | |||
|
Domain Architecture |

|
|||||
| Description | 80 kDa MCM3-associated protein (GANP protein). | |||||
|
RHG01_HUMAN
|
||||||
| NC score | 0.068441 (rank : 8) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q07960 | Gene names | ARHGAP1, CDC42GAP, RHOGAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 1 (GTPase-activating protein rhoOGAP) (Rho-related small GTPase protein activator) (CDC42 GTPase-activating protein) (p50-RhoGAP). | |||||
|
RHG25_MOUSE
|
||||||
| NC score | 0.066741 (rank : 9) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BYW1, Q2VPR2, Q3TE04, Q3TVX0, Q3UVB7, Q6A0E0, Q8BX98 | Gene names | Arhgap25, Kiaa0053 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
RHG25_HUMAN
|
||||||
| NC score | 0.063363 (rank : 10) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 550 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P42331, Q8IXQ2 | Gene names | ARHGAP25, KIAA0053 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
RHG18_MOUSE
|
||||||
| NC score | 0.059688 (rank : 11) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K0Q5, Q8BP03, Q8R196 | Gene names | Arhgap18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho-GTPase-activating protein 18. | |||||
|
TXND2_HUMAN
|
||||||
| NC score | 0.059232 (rank : 12) | θ value | 0.0961366 (rank : 8) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
GMIP_MOUSE
|
||||||
| NC score | 0.058926 (rank : 13) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6PGG2, Q6P9S3 | Gene names | Gmip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GEM-interacting protein (GMIP). | |||||
|
RHG06_HUMAN
|
||||||
| NC score | 0.057807 (rank : 14) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43182, O43437, Q9P1B3, Q9UK81, Q9UK82 | Gene names | ARHGAP6, RHOGAP6 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 6 (Rho-type GTPase-activating protein RhoGAPX-1). | |||||
|
GMIP_HUMAN
|
||||||
| NC score | 0.057106 (rank : 15) | θ value | θ > 10 (rank : 80) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9P107 | Gene names | GMIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GEM-interacting protein (GMIP). | |||||
|
RHG06_MOUSE
|
||||||
| NC score | 0.057043 (rank : 16) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O54834, Q9QZL8 | Gene names | Arhgap6 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 6 (Rho-type GTPase-activating protein RhoGAPX-1). | |||||
|
RGAP1_HUMAN
|
||||||
| NC score | 0.056753 (rank : 17) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H0H5, Q6PJ26, Q9NWN2, Q9P250, Q9P2W2 | Gene names | RACGAP1, KIAA1478, MGCRACGAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rac GTPase-activating protein 1 (MgcRacGAP). | |||||
|
RHG08_MOUSE
|
||||||
| NC score | 0.054408 (rank : 18) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CXP4, Q99JY7 | Gene names | Arhgap8 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 8. | |||||
|
SETBP_HUMAN
|
||||||
| NC score | 0.052510 (rank : 19) | θ value | 0.62314 (rank : 18) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Y6X0, Q9UEF3 | Gene names | SETBP1, KIAA0437 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SET-binding protein (SEB). | |||||
|
K1688_HUMAN
|
||||||
| NC score | 0.052077 (rank : 20) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9C0H5 | Gene names | KIAA1688 | |||
|
Domain Architecture |

|
|||||
| Description | Protein KIAA1688. | |||||
|
K1688_MOUSE
|
||||||
| NC score | 0.052050 (rank : 21) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P59281, Q69ZD4, Q6P9R6 | Gene names | Kiaa1688, D15Wsu169e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1688. | |||||
|
RHG05_MOUSE
|
||||||
| NC score | 0.051917 (rank : 22) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P97393 | Gene names | Arhgap5, Rhogap5 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 5 (p190-B). | |||||
|
RHG08_HUMAN
|
||||||
| NC score | 0.051884 (rank : 23) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NSG0, O75983, O95695, Q96RW1, Q96RW2, Q9HA49, Q9HC46, Q9NVX8, Q9NXL1, Q9UH20 | Gene names | ARHGAP8 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 8. | |||||
|
RHG05_HUMAN
|
||||||
| NC score | 0.051751 (rank : 24) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13017 | Gene names | ARHGAP5, RHOGAP5 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 5 (p190-B). | |||||
|
MCM3A_MOUSE
|
||||||
| NC score | 0.048199 (rank : 25) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9WUU9 | Gene names | Mcm3ap, Ganp, Map80 | |||
|
Domain Architecture |

|
|||||
| Description | 80 kDa MCM3-associated protein (GANP protein). | |||||
|
SETBP_MOUSE
|
||||||
| NC score | 0.047098 (rank : 26) | θ value | 1.81305 (rank : 29) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Z180, Q66JL8 | Gene names | Setbp1, Kiaa0437 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SET-binding protein (SEB). | |||||
|
PRG4_MOUSE
|
||||||
| NC score | 0.044453 (rank : 27) | θ value | 2.36792 (rank : 34) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.042997 (rank : 28) | θ value | 3.0926 (rank : 44) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
AFF1_HUMAN
|
||||||
| NC score | 0.041899 (rank : 29) | θ value | 3.0926 (rank : 38) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
CP053_HUMAN
|
||||||
| NC score | 0.041690 (rank : 30) | θ value | 4.03905 (rank : 47) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BTK6 | Gene names | C16orf53 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf53. | |||||
|
BASP_HUMAN
|
||||||
| NC score | 0.041594 (rank : 31) | θ value | 2.36792 (rank : 31) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
NCOR2_MOUSE
|
||||||
| NC score | 0.041355 (rank : 32) | θ value | 0.62314 (rank : 17) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
IF39_HUMAN
|
||||||
| NC score | 0.039176 (rank : 33) | θ value | 2.36792 (rank : 32) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P55884, Q9UMF9 | Gene names | EIF3S9 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 9 (eIF-3 eta) (eIF3 p116) (eIF3 p110) (eIF3b) (Prt1 homolog) (hPrt1). | |||||
|
DIDO1_HUMAN
|
||||||
| NC score | 0.039109 (rank : 34) | θ value | 1.06291 (rank : 22) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.038219 (rank : 35) | θ value | 0.279714 (rank : 13) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
X3CL1_MOUSE
|
||||||
| NC score | 0.037509 (rank : 36) | θ value | 0.813845 (rank : 19) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O35188, O35933 | Gene names | Cx3cl1, Cx3c, Fkn, Scyd1 | |||
|
Domain Architecture |

|
|||||
| Description | Fractalkine precursor (CX3CL1) (Neurotactin) (CX3C membrane-anchored chemokine) (Small inducible cytokine D1). | |||||
|
PRGC2_MOUSE
|
||||||
| NC score | 0.036061 (rank : 37) | θ value | 1.38821 (rank : 25) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8VHJ7, Q8C1C0 | Gene names | Ppargc1b, Errl1, Pgc1, Pgc1b, Ppargc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta protein) (PGC-1beta) (ERR ligand 1) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.034821 (rank : 38) | θ value | 1.81305 (rank : 27) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
G3BP_MOUSE
|
||||||
| NC score | 0.033891 (rank : 39) | θ value | 0.365318 (rank : 14) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P97855 | Gene names | G3bp, G3bp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras-GTPase-activating protein-binding protein 1 (EC 3.6.1.-) (ATP- dependent DNA helicase VIII) (GAP SH3-domain-binding protein 1) (G3BP- 1) (HDH-VIII). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.033799 (rank : 40) | θ value | 0.62314 (rank : 16) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
BAT2_MOUSE
|
||||||
| NC score | 0.032562 (rank : 41) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.032309 (rank : 42) | θ value | 6.88961 (rank : 65) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
DLEC1_HUMAN
|
||||||
| NC score | 0.031535 (rank : 43) | θ value | 1.38821 (rank : 23) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y238, Q9NSW0, Q9NTG5 | Gene names | DLEC1, DLC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Deleted in lung and esophageal cancer protein 1 (Deleted in lung cancer protein 1) (DLC-1). | |||||
|
CLIC6_HUMAN
|
||||||
| NC score | 0.030804 (rank : 44) | θ value | 0.125558 (rank : 9) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96NY7, Q8IX31 | Gene names | CLIC6, CLIC1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel 6. | |||||
|
PRP1_HUMAN
|
||||||
| NC score | 0.029939 (rank : 45) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
S30BP_HUMAN
|
||||||
| NC score | 0.029624 (rank : 46) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UHR5, Q8N1R5, Q8TDR8, Q96D27 | Gene names | SAP30BP, HCNGP, HTRG, HTRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAP30-binding protein (Transcriptional regulator protein HCNGP). | |||||
|
BRPF3_HUMAN
|
||||||
| NC score | 0.027714 (rank : 47) | θ value | 0.21417 (rank : 12) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9ULD4, Q5R3K8 | Gene names | BRPF3, KIAA1286 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and PHD finger-containing protein 3. | |||||
|
WDR60_HUMAN
|
||||||
| NC score | 0.026982 (rank : 48) | θ value | 1.38821 (rank : 26) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8WVS4, Q9NW58 | Gene names | WDR60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 60. | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.026853 (rank : 49) | θ value | 6.88961 (rank : 64) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
APLP1_HUMAN
|
||||||
| NC score | 0.026362 (rank : 50) | θ value | 1.06291 (rank : 21) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P51693, O00113, Q96A92 | Gene names | APLP1 | |||
|
Domain Architecture |

|
|||||
| Description | Amyloid-like protein 1 precursor (APLP) (APLP-1) [Contains: C30]. | |||||
|
EP15R_MOUSE
|
||||||
| NC score | 0.025956 (rank : 51) | θ value | 0.125558 (rank : 10) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q60902, Q8CB60, Q8CB70, Q91WH8 | Gene names | Eps15l1, Eps15-rs, Eps15R | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal growth factor receptor substrate 15-like 1 (Eps15-related protein) (Eps15R) (Epidermal growth factor receptor pathway substrate 15-related sequence) (Eps15-rs). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.025932 (rank : 52) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
PEX14_HUMAN
|
||||||
| NC score | 0.025887 (rank : 53) | θ value | 3.0926 (rank : 42) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75381, Q8WX51 | Gene names | PEX14 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal membrane protein PEX14 (Peroxin-14) (Peroxisomal membrane anchor protein PEX14) (PTS1 receptor docking protein). | |||||
|
SYN1_HUMAN
|
||||||
| NC score | 0.025662 (rank : 54) | θ value | 1.81305 (rank : 30) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P17600, O75825, Q5H9A9 | Gene names | SYN1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I) (Brain protein 4.1). | |||||
|
PKHA4_MOUSE
|
||||||
| NC score | 0.025585 (rank : 55) | θ value | 1.38821 (rank : 24) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8VC98, Q8R3N3 | Gene names | Plekha4, Pepp1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 4 (Phosphoinositol 3-phosphate-binding protein 1) (PEPP-1). | |||||
|
MYBB_HUMAN
|
||||||
| NC score | 0.024922 (rank : 56) | θ value | 4.03905 (rank : 50) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P10244 | Gene names | MYBL2, BMYB | |||
|
Domain Architecture |

|
|||||
| Description | Myb-related protein B (B-Myb). | |||||
|
TCAL5_HUMAN
|
||||||
| NC score | 0.024718 (rank : 57) | θ value | 2.36792 (rank : 37) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
ESCO1_HUMAN
|
||||||
| NC score | 0.023897 (rank : 58) | θ value | 3.0926 (rank : 40) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5FWF5, Q69YG4, Q69YS3, Q6IMD7, Q8N3Z5, Q8NBG2, Q96PX7 | Gene names | ESCO1, EFO1, KIAA1911 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyltransferase ESCO1 (EC 2.3.1.-) (Establishment of cohesion 1 homolog 1) (ECO1 homolog 1) (ESO1 homolog 1) (Establishment factor- like protein 1) (EFO1p) (hEFO1) (CTF7 homolog 1). | |||||
|
ITB3_HUMAN
|
||||||
| NC score | 0.023310 (rank : 59) | θ value | 0.125558 (rank : 11) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P05106, O15495, Q13413, Q14648, Q16499 | Gene names | ITGB3, GP3A | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-3 precursor (Platelet membrane glycoprotein IIIa) (GPIIIa) (CD61 antigen). | |||||
|
K0256_HUMAN
|
||||||
| NC score | 0.022527 (rank : 60) | θ value | 2.36792 (rank : 33) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q93073, Q8N767 | Gene names | KIAA0256 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0256. | |||||
|
ZF106_HUMAN
|
||||||
| NC score | 0.022290 (rank : 61) | θ value | 0.813845 (rank : 20) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9H2Y7, Q6NSD9, Q6PEK1, Q86T43, Q86T45, Q86T50, Q86T58, Q86TA9, Q96M37, Q9H7B8 | Gene names | ZFP106, ZNF474 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 106 homolog (Zfp-106) (Zinc finger protein 474). | |||||
|
SYN3_HUMAN
|
||||||
| NC score | 0.021923 (rank : 62) | θ value | 5.27518 (rank : 58) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O14994 | Gene names | SYN3 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-3 (Synapsin III). | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.021214 (rank : 63) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
SRCA_HUMAN
|
||||||
| NC score | 0.020563 (rank : 64) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q86TD4 | Gene names | SRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
RECQ5_HUMAN
|
||||||
| NC score | 0.020511 (rank : 65) | θ value | 4.03905 (rank : 51) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O94762, Q9P1W7, Q9UNC8 | Gene names | RECQL5, RECQ5 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase Q5 (EC 3.6.1.-) (RecQ protein-like 5) (RecQ5). | |||||
|
TCAL3_HUMAN
|
||||||
| NC score | 0.020065 (rank : 66) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q969E4 | Gene names | TCEAL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 3 (TCEA-like protein 3) (Transcription elongation factor S-II protein-like 3). | |||||
|
CMGA_HUMAN
|
||||||
| NC score | 0.019911 (rank : 67) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P10645, Q96E84, Q96GL7, Q9BQB5 | Gene names | CHGA | |||
|
Domain Architecture |

|
|||||
| Description | Chromogranin A precursor (CgA) (Pituitary secretory protein I) (SP-I) [Contains: Vasostatin-1 (Vasostatin I); Vasostatin-2 (Vasostatin II); EA-92; ES-43; Pancreastatin; SS-18; WA-8; WE-14; LF-19; AL-11; GV-19; GR-44; ER-37]. | |||||
|
NUCL_MOUSE
|
||||||
| NC score | 0.019843 (rank : 68) | θ value | 3.0926 (rank : 41) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P09405, Q548M9, Q61991, Q99K50 | Gene names | Ncl, Nuc | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolin (Protein C23). | |||||
|
AFF2_MOUSE
|
||||||
| NC score | 0.019123 (rank : 69) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O55112 | Gene names | Aff2, Fmr2, Ox19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation protein 2 homolog) (Protein FMR-2) (FMR2P) (Protein Ox19). | |||||
|
MEN1_MOUSE
|
||||||
| NC score | 0.018622 (rank : 70) | θ value | 5.27518 (rank : 55) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O88559 | Gene names | Men1 | |||
|
Domain Architecture |

|
|||||
| Description | Menin. | |||||
|
LAS1L_HUMAN
|
||||||
| NC score | 0.018010 (rank : 71) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y4W2, Q5JXQ0, Q8TEN5, Q9H9V5 | Gene names | LAS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LAS1-like protein. | |||||
|
IF4G1_MOUSE
|
||||||
| NC score | 0.017593 (rank : 72) | θ value | 4.03905 (rank : 48) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6NZJ6, Q6NZN8, Q8BW99 | Gene names | Eif4g1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1). | |||||
|
ARHG1_MOUSE
|
||||||
| NC score | 0.017354 (rank : 73) | θ value | 4.03905 (rank : 45) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q61210, O89074, Q80YE8, Q80YE9, Q91VL3 | Gene names | Arhgef1, Lbcl2, Lsc | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 1 (Lymphoid blast crisis-like 2) (Lbc's second cousin). | |||||
|
ITB3_MOUSE
|
||||||
| NC score | 0.017342 (rank : 74) | θ value | 4.03905 (rank : 49) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O54890 | Gene names | Itgb3 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-3 precursor (Platelet membrane glycoprotein IIIa) (GPIIIa) (CD61 antigen). | |||||
|
K1045_HUMAN
|
||||||
| NC score | 0.016236 (rank : 75) | θ value | 6.88961 (rank : 66) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UPV7, Q58FE9, Q5T662 | Gene names | KIAA1045 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1045. | |||||
|
ORC2_MOUSE
|
||||||
| NC score | 0.016160 (rank : 76) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q60862 | Gene names | Orc2l, Orc2 | |||
|
Domain Architecture |

|
|||||
| Description | Origin recognition complex subunit 2. | |||||
|
ARHG1_HUMAN
|
||||||
| NC score | 0.015740 (rank : 77) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92888, O00513, Q8N4J4, Q96BF4, Q96F17, Q9BSB1 | Gene names | ARHGEF1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 1 (p115-RhoGEF) (p115RhoGEF) (115 kDa guanine nucleotide exchange factor) (Sub1.5). | |||||
|
RIMS1_HUMAN
|
||||||
| NC score | 0.015681 (rank : 78) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 437 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86UR5, O15048, Q8TDY9, Q8TDZ5, Q9HBA1, Q9HBA2, Q9HBA3, Q9HBA4, Q9HBA5, Q9HBA6 | Gene names | RIMS1, KIAA0340, RIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 1 (Rab3-interacting molecule 1) (RIM 1). | |||||
|
BCAS1_HUMAN
|
||||||
| NC score | 0.015470 (rank : 79) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75363, Q68CZ3 | Gene names | BCAS1, AIBC1, NABC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Breast carcinoma amplified sequence 1 (Novel amplified in breast cancer 1) (Amplified and overexpressed in breast cancer). | |||||
|
IFT74_HUMAN
|
||||||
| NC score | 0.014311 (rank : 80) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96LB3, Q6PGQ8, Q9H643, Q9H8G7 | Gene names | IFT74, CCDC2, CMG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 74 homolog (Coiled-coil domain-containing protein 2) (Capillary morphogenesis protein 1) (CMG-1). | |||||
|
TAIP2_MOUSE
|
||||||
| NC score | 0.013417 (rank : 81) | θ value | 5.27518 (rank : 59) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P59055 | Gene names | Taip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TGF-beta-induced apoptosis protein 2 (TAIP-2). | |||||
|
MKL1_MOUSE
|
||||||
| NC score | 0.012132 (rank : 82) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8K4J6 | Gene names | Mkl1, Bsac | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 1 (Myocardin-related transcription factor A) (MRTF-A) (Megakaryoblastic leukemia 1 protein homolog) (Basic SAP coiled-coil transcription activator). | |||||
|
TRPM7_MOUSE
|
||||||
| NC score | 0.011879 (rank : 83) | θ value | 3.0926 (rank : 43) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q923J1, Q8C7S7, Q8CE54, Q921Y5, Q925B2, Q9CUT2, Q9JLQ1 | Gene names | Trpm7, Chak, Ltrpc7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 7 (EC 2.7.11.1) (Long transient receptor potential channel 7) (LTrpC7) (Channel-kinase 1) (Transient receptor potential-phospholipase C- interacting kinase) (TRP-PLIK). | |||||
|
TITIN_HUMAN
|
||||||
| NC score | 0.011574 (rank : 84) | θ value | 0.365318 (rank : 15) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
IRF3_HUMAN
|
||||||
| NC score | 0.009852 (rank : 85) | θ value | 6.88961 (rank : 63) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q14653 | Gene names | IRF3 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon regulatory factor 3 (IRF-3). | |||||
|
CHD4_MOUSE
|
||||||
| NC score | 0.009660 (rank : 86) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6PDQ2 | Gene names | Chd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (CHD-4). | |||||
|
CSMD3_HUMAN
|
||||||
| NC score | 0.007491 (rank : 87) | θ value | 3.0926 (rank : 39) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z407, Q96PZ3 | Gene names | CSMD3, KIAA1894 | |||
|
Domain Architecture |

|
|||||
| Description | CUB and sushi domain-containing protein 3 precursor (CUB and sushi multiple domains protein 3). | |||||
|
JHD1B_MOUSE
|
||||||
| NC score | 0.006554 (rank : 88) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6P1G2, Q3V396, Q6PFD0, Q6ZPE8, Q9CSF7, Q9QZN6 | Gene names | Fbxl10, Fbl10, Jhdm1b, Kiaa3014 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10). | |||||
|
ADCY4_HUMAN
|
||||||
| NC score | 0.006465 (rank : 89) | θ value | 5.27518 (rank : 52) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NFM4, Q96ML7 | Gene names | ADCY4 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 4 (EC 4.6.1.1) (Adenylate cyclase type IV) (ATP pyrophosphate-lyase 4) (Adenylyl cyclase 4). | |||||
|
UTRO_HUMAN
|
||||||
| NC score | 0.005083 (rank : 90) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P46939 | Gene names | UTRN, DMDL | |||
|
Domain Architecture |

|
|||||
| Description | Utrophin (Dystrophin-related protein 1) (DRP1) (DRP). | |||||
|
PTN1_HUMAN
|
||||||
| NC score | 0.004990 (rank : 91) | θ value | 5.27518 (rank : 57) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P18031, Q5TGD8, Q9BQV9, Q9NQQ4 | Gene names | PTPN1, PTP1B | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 1 (EC 3.1.3.48) (Protein-tyrosine phosphatase 1B) (PTP-1B). | |||||
|
SALL2_MOUSE
|
||||||
| NC score | 0.004817 (rank : 92) | θ value | 1.81305 (rank : 28) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9QX96 | Gene names | Sall2, Sal2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 2 (Spalt-like protein 2) (MSal-2). | |||||
|
CD2L7_HUMAN
|
||||||
| NC score | 0.001896 (rank : 93) | θ value | 6.88961 (rank : 60) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1275 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NYV4, O94978 | Gene names | CRKRS, CRK7, KIAA0904 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle 2-related protein kinase 7 (EC 2.7.11.22) (CDC2- related protein kinase 7) (Cdc2-related kinase, arginine/serine-rich) (CrkRS). | |||||
|
NRK_MOUSE
|
||||||
| NC score | 0.001380 (rank : 94) | θ value | 5.27518 (rank : 56) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 899 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9R0G8, Q6NV55, Q8C9S9, Q9R0S4 | Gene names | Nrk, Nesk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nik-related protein kinase (EC 2.7.11.1) (Nck-interacting kinase-like embryo specific kinase) (NIK-like embryo-specific kinase) (NESK). | |||||