Please be patient as the page loads
|
FA35A_MOUSE
|
||||||
| SwissProt Accessions | Q3UEN2 | Gene names | Fam35a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM35A. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
FA35A_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.946480 (rank : 2) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86V20, O95885, Q9H991 | Gene names | FAM35A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM35A. | |||||
|
FA35A_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q3UEN2 | Gene names | Fam35a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM35A. | |||||
|
CHK2_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 3) | NC score | 0.018481 (rank : 33) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 860 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z265 | Gene names | Chek2, Chk2, Rad53 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Chk2 (EC 2.7.11.1). | |||||
|
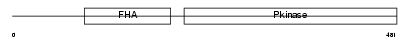
CHK2_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 4) | NC score | 0.017346 (rank : 36) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O96017, Q6QA03, Q6QA04, Q6QA05, Q6QA06, Q6QA07, Q6QA08, Q6QA10, Q6QA11, Q6QA12, Q6QA13, Q9HCQ8, Q9UGF0, Q9UGF1 | Gene names | CHEK2, CHK2 | |||
|
Domain Architecture |
|
|||||
| Description | Serine/threonine-protein kinase Chk2 (EC 2.7.11.1) (Cds1). | |||||
|
DMP1_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 5) | NC score | 0.095749 (rank : 4) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
RSRC1_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 6) | NC score | 0.118595 (rank : 3) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9DBU6, Q3TF06 | Gene names | Rsrc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine/serine-rich coiled coil protein 1. | |||||
|
DMP1_MOUSE
|
||||||
| θ value | 0.163984 (rank : 7) | NC score | 0.080763 (rank : 5) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 8) | NC score | 0.050712 (rank : 12) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
ATBF1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 9) | NC score | 0.022856 (rank : 27) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1841 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q61329 | Gene names | Atbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
NBN_MOUSE
|
||||||
| θ value | 0.279714 (rank : 10) | NC score | 0.061460 (rank : 10) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9R207, O88981, Q3UY57, Q811I6, Q8CCY0, Q9R1X1 | Gene names | Nbn, Nbs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nibrin (Nijmegen breakage syndrome protein 1 homolog) (Cell cycle regulatory protein p95). | |||||
|
TGON1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 11) | NC score | 0.080008 (rank : 6) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q62313 | Gene names | Tgoln1, Ttgn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 1 precursor (TGN38A). | |||||
|
C8AP2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 12) | NC score | 0.043651 (rank : 17) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UKL3, Q6PH76, Q7LCQ7, Q86YD9, Q9NUQ4, Q9NZV9, Q9P2N1, Q9Y563 | Gene names | CASP8AP2, FLASH, KIAA1315, RIP25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
TGON2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 13) | NC score | 0.072285 (rank : 8) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q62314 | Gene names | Tgoln2, Ttgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (TGN38B). | |||||
|
BANK1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 14) | NC score | 0.045221 (rank : 16) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NDB2, Q8N5K8, Q8NB56, Q8WYN5, Q9NWP2 | Gene names | BANK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell scaffold protein with ankyrin repeats. | |||||
|
ENL_HUMAN
|
||||||
| θ value | 1.06291 (rank : 15) | NC score | 0.062052 (rank : 9) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
LIMA1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 16) | NC score | 0.033053 (rank : 22) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 309 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UHB6, Q2TAN7, Q9BVF2, Q9H8J1, Q9HBN5, Q9NX96, Q9NXC3, Q9NXU6, Q9P0H8, Q9UHB5 | Gene names | LIMA1, EPLIN, SREBP3 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain and actin-binding protein 1 (Epithelial protein lost in neoplasm). | |||||
|
CLSPN_MOUSE
|
||||||
| θ value | 1.81305 (rank : 17) | NC score | 0.038287 (rank : 19) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80YR7, Q69GM2 | Gene names | Clspn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Claspin. | |||||
|
DEK_MOUSE
|
||||||
| θ value | 1.81305 (rank : 18) | NC score | 0.047693 (rank : 14) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q7TNV0, Q80VC5, Q8BZV6 | Gene names | Dek | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
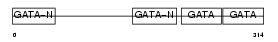
GATA4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 19) | NC score | 0.021533 (rank : 29) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P43694 | Gene names | GATA4 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor GATA-4 (GATA-binding factor 4). | |||||
|
MYCPP_HUMAN
|
||||||
| θ value | 1.81305 (rank : 20) | NC score | 0.029002 (rank : 26) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z401, Q14655, Q86T77, Q8IVX2, Q8NB93 | Gene names | MYCPBP, IRLB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-myc promoter-binding protein. | |||||
|
RSRC1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 21) | NC score | 0.079733 (rank : 7) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96IZ7, Q96QK2, Q9NZE5 | Gene names | RSRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine/serine-rich coiled coil protein 1. | |||||
|
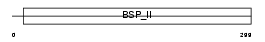
SIAL_MOUSE
|
||||||
| θ value | 1.81305 (rank : 22) | NC score | 0.050851 (rank : 11) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61711, Q61363 | Gene names | Ibsp | |||
|
Domain Architecture |
|
|||||
| Description | Bone sialoprotein-2 precursor (Bone sialoprotein II) (BSP II) (Cell- binding sialoprotein) (Integrin-binding sialoprotein). | |||||
|
SLC2B_HUMAN
|
||||||
| θ value | 1.81305 (rank : 23) | NC score | 0.046269 (rank : 15) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8NEV8, Q9Y4D6 | Gene names | EXPH5, KIAA0624, SLAC2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Slp homolog lacking C2 domains b (Exophilin-5). | |||||
|
CR025_MOUSE
|
||||||
| θ value | 2.36792 (rank : 24) | NC score | 0.048310 (rank : 13) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BH50, Q3TQ72, Q8BU43 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C18orf25 homolog. | |||||
|
PRGC1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 25) | NC score | 0.037109 (rank : 20) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UBK2, Q9UN32 | Gene names | PPARGC1A, LEM6, PGC1, PGC1A, PPARGC1 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PPAR gamma coactivator 1-alpha) (PPARGC-1-alpha) (PGC-1-alpha) (Ligand effect modulator 6). | |||||
|
RIN2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 26) | NC score | 0.021382 (rank : 30) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D684, Q99K06 | Gene names | Rin2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 2 (Ras interaction/interference protein 2). | |||||
|
ZN406_MOUSE
|
||||||
| θ value | 3.0926 (rank : 27) | NC score | 0.003984 (rank : 53) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 800 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7TS63 | Gene names | Znf406, Zfp406 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 406. | |||||
|
AT1A1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 28) | NC score | 0.008141 (rank : 50) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VDN2, Q91Z09 | Gene names | Atp1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-1 chain precursor (EC 3.6.3.9) (Sodium pump 1) (Na+/K+ ATPase 1). | |||||
|
CBPB2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 29) | NC score | 0.017568 (rank : 35) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96IY4, Q15114, Q5T9K1, Q5T9K2, Q9P2Y6 | Gene names | CPB2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase B2 precursor (EC 3.4.17.20) (Carboxypeptidase U) (CPU) (Thrombin-activable fibrinolysis inhibitor) (TAFI) (Plasma carboxypeptidase B) (pCPB). | |||||
|
HTSF1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 30) | NC score | 0.030875 (rank : 24) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O43719, Q59G06, Q99730 | Gene names | HTATSF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 (Tat-SF1). | |||||
|
PK3CD_HUMAN
|
||||||
| θ value | 4.03905 (rank : 31) | NC score | 0.011802 (rank : 47) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O00329, O15445 | Gene names | PIK3CD | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit delta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit delta) (PtdIns-3- kinase p110) (PI3K) (p110delta). | |||||
|
PRGC1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 32) | NC score | 0.034802 (rank : 21) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O70343 | Gene names | Ppargc1a, Pgc1, Pgc1a, Ppargc1 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PPAR gamma coactivator 1-alpha) (PPARGC-1-alpha) (PGC-1-alpha). | |||||
|
CF010_HUMAN
|
||||||
| θ value | 5.27518 (rank : 33) | NC score | 0.029666 (rank : 25) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5SRN2, Q5SPI9, Q5SPJ0, Q5SPK9, Q5SPL0, Q5SRN3, Q5TG25, Q5TG26, Q8N4B6 | Gene names | C6orf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf10. | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 5.27518 (rank : 34) | NC score | 0.010763 (rank : 48) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
PITM1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 35) | NC score | 0.017867 (rank : 34) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O00562, Q6T7X3, Q8TBN3, Q9BZ73 | Gene names | PITPNM1, DRES9, NIR2, PITPNM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 1 (Phosphatidylinositol transfer protein, membrane-associated 1) (Pyk2 N-terminal domain-interacting receptor 2) (NIR-2) (Drosophila retinal degeneration B homolog). | |||||
|
SCFD2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 36) | NC score | 0.022842 (rank : 28) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BTY8, Q3USN5, Q8BFV9, Q8BKU5, Q8BKZ0, Q8BTP6, Q8BTQ9, Q8BXI1, Q8BXS0, Q8BZ58 | Gene names | Scfd2, Stxbp1l1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sec1 family domain-containing protein 2 (Syntaxin-binding protein 1- like 1) (Neuronal Sec1). | |||||
|
CBPB2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 37) | NC score | 0.015195 (rank : 42) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JHH6, Q5EBI3, Q9QZF0 | Gene names | Cpb2, Tafi | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase B2 precursor (EC 3.4.17.20) (Carboxypeptidase U) (CPU) (Thrombin-activable fibrinolysis inhibitor) (TAFI) (Carboxypeptidase R) (CPR). | |||||
|
CERK1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 38) | NC score | 0.016691 (rank : 38) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TCT0, Q9BYB3, Q9UGE5 | Gene names | CERK, KIAA1646 | |||
|
Domain Architecture |

|
|||||
| Description | Ceramide kinase (EC 2.7.1.138) (Acylsphingosine kinase) (hCERK) (Lipid kinase 4) (LK4). | |||||
|
KCNH8_HUMAN
|
||||||
| θ value | 6.88961 (rank : 39) | NC score | 0.007902 (rank : 51) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96L42 | Gene names | KCNH8 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 8 (Voltage-gated potassium channel subunit Kv12.1) (Ether-a-go-go-like potassium channel 3) (ELK channel 3) (ELK3) (ELK1) (hElk1). | |||||
|
MYOC_HUMAN
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | 0.013912 (rank : 44) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 489 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99972, O00620, Q7Z6Q9 | Gene names | MYOC, GLC1A, TIGR | |||
|
Domain Architecture |

|
|||||
| Description | Myocilin precursor (Trabecular meshwork-induced glucocorticoid response protein). | |||||
|
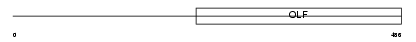
MYOC_MOUSE
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.015901 (rank : 40) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70624, O70289 | Gene names | Myoc, Tigr | |||
|
Domain Architecture |
|
|||||
| Description | Myocilin precursor (Trabecular meshwork-induced glucocorticoid response protein). | |||||
|
NCKX1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.020643 (rank : 31) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60721, O43485, O75184 | Gene names | SLC24A1, KIAA0702, NCKX1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium/calcium exchanger 1 (Na(+)/K(+)/Ca(2+)-exchange protein 1) (Retinal rod Na-Ca+K exchanger). | |||||
|
PGCB_MOUSE
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.007378 (rank : 52) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61361 | Gene names | Bcan | |||
|
Domain Architecture |

|
|||||
| Description | Brevican core protein precursor. | |||||
|
PITM1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.016571 (rank : 39) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35954 | Gene names | Pitpnm1, Dres9, Mpt1, Nir2, Pitpnm | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 1 (Phosphatidylinositol transfer protein, membrane-associated 1) (PITPnm 1) (Mpt-1) (Pyk2 N-terminal domain-interacting receptor 2) (NIR-2) (Drosophila retinal degeneration B homolog 1) (RdgB1). | |||||
|
SYTL2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.012967 (rank : 46) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9HCH5, Q2YDA7, Q8ND34, Q96BJ2, Q9H768, Q9NXM1 | Gene names | SYTL2, KIAA1597, SLP2 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 2 (Exophilin-4). | |||||
|
AF9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 46) | NC score | 0.038495 (rank : 18) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P42568 | Gene names | MLLT3, AF9, YEATS3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein AF-9 (ALL1 fused gene from chromosome 9 protein) (Myeloid/lymphoid or mixed-lineage leukemia translocated to chromosome 3 protein) (YEATS domain-containing protein 3). | |||||
|
AHTF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 47) | NC score | 0.015357 (rank : 41) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WYP5, Q7Z4E3, Q8IZA4, Q96EH9, Q9Y4Q6 | Gene names | AHCTF1, ELYS, TMBS62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
AP3B2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 48) | NC score | 0.015039 (rank : 43) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q13367, O14808 | Gene names | AP3B2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (AP-3 complex beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain) (Neuron-specific vesicle coat protein beta-NAP). | |||||
|
CR025_HUMAN
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.032137 (rank : 23) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96B23, Q5XG78, Q6N058, Q86TB5, Q8TCQ5 | Gene names | C18orf25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C18orf25. | |||||
|
ESCO1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 50) | NC score | 0.017232 (rank : 37) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5FWF5, Q69YG4, Q69YS3, Q6IMD7, Q8N3Z5, Q8NBG2, Q96PX7 | Gene names | ESCO1, EFO1, KIAA1911 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyltransferase ESCO1 (EC 2.3.1.-) (Establishment of cohesion 1 homolog 1) (ECO1 homolog 1) (ESO1 homolog 1) (Establishment factor- like protein 1) (EFO1p) (hEFO1) (CTF7 homolog 1). | |||||
|
MCM3A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.013876 (rank : 45) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9WUU9 | Gene names | Mcm3ap, Ganp, Map80 | |||
|
Domain Architecture |

|
|||||
| Description | 80 kDa MCM3-associated protein (GANP protein). | |||||
|
NPAS3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.008680 (rank : 49) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
S30BP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.020388 (rank : 32) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q02614, Q8VDJ5 | Gene names | Sap30bp, Hcngp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAP30-binding protein (Transcriptional regulator protein HCNGP). | |||||
|
ZN639_HUMAN
|
||||||
| θ value | 8.99809 (rank : 54) | NC score | 0.002773 (rank : 54) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 723 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UID6 | Gene names | ZNF639, ZASC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 639 (Zinc finger protein ZASC1) (Zinc finger protein ANC_2H01). | |||||
|
FA35A_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q3UEN2 | Gene names | Fam35a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM35A. | |||||
|
FA35A_HUMAN
|
||||||
| NC score | 0.946480 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86V20, O95885, Q9H991 | Gene names | FAM35A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM35A. | |||||
|
RSRC1_MOUSE
|
||||||
| NC score | 0.118595 (rank : 3) | θ value | 0.0330416 (rank : 6) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9DBU6, Q3TF06 | Gene names | Rsrc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine/serine-rich coiled coil protein 1. | |||||
|
DMP1_HUMAN
|
||||||
| NC score | 0.095749 (rank : 4) | θ value | 0.0330416 (rank : 5) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
DMP1_MOUSE
|
||||||
| NC score | 0.080763 (rank : 5) | θ value | 0.163984 (rank : 7) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
TGON1_MOUSE
|
||||||
| NC score | 0.080008 (rank : 6) | θ value | 0.279714 (rank : 11) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q62313 | Gene names | Tgoln1, Ttgn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 1 precursor (TGN38A). | |||||
|
RSRC1_HUMAN
|
||||||
| NC score | 0.079733 (rank : 7) | θ value | 1.81305 (rank : 21) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96IZ7, Q96QK2, Q9NZE5 | Gene names | RSRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine/serine-rich coiled coil protein 1. | |||||
|
TGON2_MOUSE
|
||||||
| NC score | 0.072285 (rank : 8) | θ value | 0.62314 (rank : 13) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q62314 | Gene names | Tgoln2, Ttgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (TGN38B). | |||||
|
ENL_HUMAN
|
||||||
| NC score | 0.062052 (rank : 9) | θ value | 1.06291 (rank : 15) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
NBN_MOUSE
|
||||||
| NC score | 0.061460 (rank : 10) | θ value | 0.279714 (rank : 10) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9R207, O88981, Q3UY57, Q811I6, Q8CCY0, Q9R1X1 | Gene names | Nbn, Nbs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nibrin (Nijmegen breakage syndrome protein 1 homolog) (Cell cycle regulatory protein p95). | |||||
|
SIAL_MOUSE
|
||||||
| NC score | 0.050851 (rank : 11) | θ value | 1.81305 (rank : 22) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61711, Q61363 | Gene names | Ibsp | |||
|
Domain Architecture |

|
|||||
| Description | Bone sialoprotein-2 precursor (Bone sialoprotein II) (BSP II) (Cell- binding sialoprotein) (Integrin-binding sialoprotein). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.050712 (rank : 12) | θ value | 0.163984 (rank : 8) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
CR025_MOUSE
|
||||||
| NC score | 0.048310 (rank : 13) | θ value | 2.36792 (rank : 24) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BH50, Q3TQ72, Q8BU43 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C18orf25 homolog. | |||||
|
DEK_MOUSE
|
||||||
| NC score | 0.047693 (rank : 14) | θ value | 1.81305 (rank : 18) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q7TNV0, Q80VC5, Q8BZV6 | Gene names | Dek | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
SLC2B_HUMAN
|
||||||
| NC score | 0.046269 (rank : 15) | θ value | 1.81305 (rank : 23) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8NEV8, Q9Y4D6 | Gene names | EXPH5, KIAA0624, SLAC2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Slp homolog lacking C2 domains b (Exophilin-5). | |||||
|
BANK1_HUMAN
|
||||||
| NC score | 0.045221 (rank : 16) | θ value | 1.06291 (rank : 14) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NDB2, Q8N5K8, Q8NB56, Q8WYN5, Q9NWP2 | Gene names | BANK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell scaffold protein with ankyrin repeats. | |||||
|
C8AP2_HUMAN
|
||||||
| NC score | 0.043651 (rank : 17) | θ value | 0.62314 (rank : 12) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UKL3, Q6PH76, Q7LCQ7, Q86YD9, Q9NUQ4, Q9NZV9, Q9P2N1, Q9Y563 | Gene names | CASP8AP2, FLASH, KIAA1315, RIP25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
AF9_HUMAN
|
||||||
| NC score | 0.038495 (rank : 18) | θ value | 8.99809 (rank : 46) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P42568 | Gene names | MLLT3, AF9, YEATS3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein AF-9 (ALL1 fused gene from chromosome 9 protein) (Myeloid/lymphoid or mixed-lineage leukemia translocated to chromosome 3 protein) (YEATS domain-containing protein 3). | |||||
|
CLSPN_MOUSE
|
||||||
| NC score | 0.038287 (rank : 19) | θ value | 1.81305 (rank : 17) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80YR7, Q69GM2 | Gene names | Clspn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Claspin. | |||||
|
PRGC1_HUMAN
|
||||||
| NC score | 0.037109 (rank : 20) | θ value | 3.0926 (rank : 25) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UBK2, Q9UN32 | Gene names | PPARGC1A, LEM6, PGC1, PGC1A, PPARGC1 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PPAR gamma coactivator 1-alpha) (PPARGC-1-alpha) (PGC-1-alpha) (Ligand effect modulator 6). | |||||
|
PRGC1_MOUSE
|
||||||
| NC score | 0.034802 (rank : 21) | θ value | 4.03905 (rank : 32) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O70343 | Gene names | Ppargc1a, Pgc1, Pgc1a, Ppargc1 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PPAR gamma coactivator 1-alpha) (PPARGC-1-alpha) (PGC-1-alpha). | |||||
|
LIMA1_HUMAN
|
||||||
| NC score | 0.033053 (rank : 22) | θ value | 1.06291 (rank : 16) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 309 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UHB6, Q2TAN7, Q9BVF2, Q9H8J1, Q9HBN5, Q9NX96, Q9NXC3, Q9NXU6, Q9P0H8, Q9UHB5 | Gene names | LIMA1, EPLIN, SREBP3 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain and actin-binding protein 1 (Epithelial protein lost in neoplasm). | |||||
|
CR025_HUMAN
|
||||||
| NC score | 0.032137 (rank : 23) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96B23, Q5XG78, Q6N058, Q86TB5, Q8TCQ5 | Gene names | C18orf25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C18orf25. | |||||
|
HTSF1_HUMAN
|
||||||
| NC score | 0.030875 (rank : 24) | θ value | 4.03905 (rank : 30) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O43719, Q59G06, Q99730 | Gene names | HTATSF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 (Tat-SF1). | |||||
|
CF010_HUMAN
|
||||||
| NC score | 0.029666 (rank : 25) | θ value | 5.27518 (rank : 33) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5SRN2, Q5SPI9, Q5SPJ0, Q5SPK9, Q5SPL0, Q5SRN3, Q5TG25, Q5TG26, Q8N4B6 | Gene names | C6orf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf10. | |||||
|
MYCPP_HUMAN
|
||||||
| NC score | 0.029002 (rank : 26) | θ value | 1.81305 (rank : 20) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z401, Q14655, Q86T77, Q8IVX2, Q8NB93 | Gene names | MYCPBP, IRLB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-myc promoter-binding protein. | |||||
|
ATBF1_MOUSE
|
||||||
| NC score | 0.022856 (rank : 27) | θ value | 0.279714 (rank : 9) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1841 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q61329 | Gene names | Atbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
SCFD2_MOUSE
|
||||||
| NC score | 0.022842 (rank : 28) | θ value | 5.27518 (rank : 36) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BTY8, Q3USN5, Q8BFV9, Q8BKU5, Q8BKZ0, Q8BTP6, Q8BTQ9, Q8BXI1, Q8BXS0, Q8BZ58 | Gene names | Scfd2, Stxbp1l1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sec1 family domain-containing protein 2 (Syntaxin-binding protein 1- like 1) (Neuronal Sec1). | |||||
|
GATA4_HUMAN
|
||||||
| NC score | 0.021533 (rank : 29) | θ value | 1.81305 (rank : 19) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P43694 | Gene names | GATA4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor GATA-4 (GATA-binding factor 4). | |||||
|
RIN2_MOUSE
|
||||||
| NC score | 0.021382 (rank : 30) | θ value | 3.0926 (rank : 26) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D684, Q99K06 | Gene names | Rin2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 2 (Ras interaction/interference protein 2). | |||||
|
NCKX1_HUMAN
|
||||||
| NC score | 0.020643 (rank : 31) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60721, O43485, O75184 | Gene names | SLC24A1, KIAA0702, NCKX1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium/calcium exchanger 1 (Na(+)/K(+)/Ca(2+)-exchange protein 1) (Retinal rod Na-Ca+K exchanger). | |||||
|
S30BP_MOUSE
|
||||||
| NC score | 0.020388 (rank : 32) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q02614, Q8VDJ5 | Gene names | Sap30bp, Hcngp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAP30-binding protein (Transcriptional regulator protein HCNGP). | |||||
|
CHK2_MOUSE
|
||||||
| NC score | 0.018481 (rank : 33) | θ value | 0.00509761 (rank : 3) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 860 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z265 | Gene names | Chek2, Chk2, Rad53 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Chk2 (EC 2.7.11.1). | |||||
|
PITM1_HUMAN
|
||||||
| NC score | 0.017867 (rank : 34) | θ value | 5.27518 (rank : 35) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O00562, Q6T7X3, Q8TBN3, Q9BZ73 | Gene names | PITPNM1, DRES9, NIR2, PITPNM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 1 (Phosphatidylinositol transfer protein, membrane-associated 1) (Pyk2 N-terminal domain-interacting receptor 2) (NIR-2) (Drosophila retinal degeneration B homolog). | |||||
|
CBPB2_HUMAN
|
||||||
| NC score | 0.017568 (rank : 35) | θ value | 4.03905 (rank : 29) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96IY4, Q15114, Q5T9K1, Q5T9K2, Q9P2Y6 | Gene names | CPB2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase B2 precursor (EC 3.4.17.20) (Carboxypeptidase U) (CPU) (Thrombin-activable fibrinolysis inhibitor) (TAFI) (Plasma carboxypeptidase B) (pCPB). | |||||
|
CHK2_HUMAN
|
||||||
| NC score | 0.017346 (rank : 36) | θ value | 0.0252991 (rank : 4) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O96017, Q6QA03, Q6QA04, Q6QA05, Q6QA06, Q6QA07, Q6QA08, Q6QA10, Q6QA11, Q6QA12, Q6QA13, Q9HCQ8, Q9UGF0, Q9UGF1 | Gene names | CHEK2, CHK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Chk2 (EC 2.7.11.1) (Cds1). | |||||
|
ESCO1_HUMAN
|
||||||
| NC score | 0.017232 (rank : 37) | θ value | 8.99809 (rank : 50) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5FWF5, Q69YG4, Q69YS3, Q6IMD7, Q8N3Z5, Q8NBG2, Q96PX7 | Gene names | ESCO1, EFO1, KIAA1911 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyltransferase ESCO1 (EC 2.3.1.-) (Establishment of cohesion 1 homolog 1) (ECO1 homolog 1) (ESO1 homolog 1) (Establishment factor- like protein 1) (EFO1p) (hEFO1) (CTF7 homolog 1). | |||||
|
CERK1_HUMAN
|
||||||
| NC score | 0.016691 (rank : 38) | θ value | 6.88961 (rank : 38) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TCT0, Q9BYB3, Q9UGE5 | Gene names | CERK, KIAA1646 | |||
|
Domain Architecture |

|
|||||
| Description | Ceramide kinase (EC 2.7.1.138) (Acylsphingosine kinase) (hCERK) (Lipid kinase 4) (LK4). | |||||
|
PITM1_MOUSE
|
||||||
| NC score | 0.016571 (rank : 39) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35954 | Gene names | Pitpnm1, Dres9, Mpt1, Nir2, Pitpnm | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 1 (Phosphatidylinositol transfer protein, membrane-associated 1) (PITPnm 1) (Mpt-1) (Pyk2 N-terminal domain-interacting receptor 2) (NIR-2) (Drosophila retinal degeneration B homolog 1) (RdgB1). | |||||
|
MYOC_MOUSE
|
||||||
| NC score | 0.015901 (rank : 40) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70624, O70289 | Gene names | Myoc, Tigr | |||
|
Domain Architecture |

|
|||||
| Description | Myocilin precursor (Trabecular meshwork-induced glucocorticoid response protein). | |||||
|
AHTF1_HUMAN
|
||||||
| NC score | 0.015357 (rank : 41) | θ value | 8.99809 (rank : 47) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WYP5, Q7Z4E3, Q8IZA4, Q96EH9, Q9Y4Q6 | Gene names | AHCTF1, ELYS, TMBS62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
CBPB2_MOUSE
|
||||||
| NC score | 0.015195 (rank : 42) | θ value | 6.88961 (rank : 37) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JHH6, Q5EBI3, Q9QZF0 | Gene names | Cpb2, Tafi | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase B2 precursor (EC 3.4.17.20) (Carboxypeptidase U) (CPU) (Thrombin-activable fibrinolysis inhibitor) (TAFI) (Carboxypeptidase R) (CPR). | |||||
|
AP3B2_HUMAN
|
||||||
| NC score | 0.015039 (rank : 43) | θ value | 8.99809 (rank : 48) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q13367, O14808 | Gene names | AP3B2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (AP-3 complex beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain) (Neuron-specific vesicle coat protein beta-NAP). | |||||
|
MYOC_HUMAN
|
||||||
| NC score | 0.013912 (rank : 44) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 489 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99972, O00620, Q7Z6Q9 | Gene names | MYOC, GLC1A, TIGR | |||
|
Domain Architecture |

|
|||||
| Description | Myocilin precursor (Trabecular meshwork-induced glucocorticoid response protein). | |||||
|
MCM3A_MOUSE
|
||||||
| NC score | 0.013876 (rank : 45) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9WUU9 | Gene names | Mcm3ap, Ganp, Map80 | |||
|
Domain Architecture |

|
|||||
| Description | 80 kDa MCM3-associated protein (GANP protein). | |||||
|
SYTL2_HUMAN
|
||||||
| NC score | 0.012967 (rank : 46) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9HCH5, Q2YDA7, Q8ND34, Q96BJ2, Q9H768, Q9NXM1 | Gene names | SYTL2, KIAA1597, SLP2 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 2 (Exophilin-4). | |||||
|
PK3CD_HUMAN
|
||||||
| NC score | 0.011802 (rank : 47) | θ value | 4.03905 (rank : 31) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O00329, O15445 | Gene names | PIK3CD | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit delta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit delta) (PtdIns-3- kinase p110) (PI3K) (p110delta). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.010763 (rank : 48) | θ value | 5.27518 (rank : 34) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
NPAS3_HUMAN
|
||||||
| NC score | 0.008680 (rank : 49) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
AT1A1_MOUSE
|
||||||
| NC score | 0.008141 (rank : 50) | θ value | 4.03905 (rank : 28) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VDN2, Q91Z09 | Gene names | Atp1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-1 chain precursor (EC 3.6.3.9) (Sodium pump 1) (Na+/K+ ATPase 1). | |||||
|
KCNH8_HUMAN
|
||||||
| NC score | 0.007902 (rank : 51) | θ value | 6.88961 (rank : 39) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96L42 | Gene names | KCNH8 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 8 (Voltage-gated potassium channel subunit Kv12.1) (Ether-a-go-go-like potassium channel 3) (ELK channel 3) (ELK3) (ELK1) (hElk1). | |||||
|
PGCB_MOUSE
|
||||||
| NC score | 0.007378 (rank : 52) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61361 | Gene names | Bcan | |||
|
Domain Architecture |

|
|||||
| Description | Brevican core protein precursor. | |||||
|
ZN406_MOUSE
|
||||||
| NC score | 0.003984 (rank : 53) | θ value | 3.0926 (rank : 27) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 800 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7TS63 | Gene names | Znf406, Zfp406 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 406. | |||||
|
ZN639_HUMAN
|
||||||
| NC score | 0.002773 (rank : 54) | θ value | 8.99809 (rank : 54) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 723 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UID6 | Gene names | ZNF639, ZASC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 639 (Zinc finger protein ZASC1) (Zinc finger protein ANC_2H01). | |||||