Please be patient as the page loads
|
S30BP_MOUSE
|
||||||
| SwissProt Accessions | Q02614, Q8VDJ5 | Gene names | Sap30bp, Hcngp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAP30-binding protein (Transcriptional regulator protein HCNGP). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
S30BP_MOUSE
|
||||||
| θ value | 9.92011e-124 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q02614, Q8VDJ5 | Gene names | Sap30bp, Hcngp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAP30-binding protein (Transcriptional regulator protein HCNGP). | |||||
|
S30BP_HUMAN
|
||||||
| θ value | 1.02657e-120 (rank : 2) | NC score | 0.992571 (rank : 2) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UHR5, Q8N1R5, Q8TDR8, Q96D27 | Gene names | SAP30BP, HCNGP, HTRG, HTRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAP30-binding protein (Transcriptional regulator protein HCNGP). | |||||
|
LEO1_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 3) | NC score | 0.098723 (rank : 5) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
VPS72_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 4) | NC score | 0.160876 (rank : 4) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15906, Q53GJ2, Q5U0R4 | Gene names | VPS72, TCFL1, YL1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
VPS72_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 5) | NC score | 0.162224 (rank : 3) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q62481, Q3U2N4, Q810A9, Q99K81 | Gene names | Vps72, Tcfl1, Yl1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
SEC63_MOUSE
|
||||||
| θ value | 0.125558 (rank : 6) | NC score | 0.097943 (rank : 6) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
DHX16_HUMAN
|
||||||
| θ value | 0.365318 (rank : 7) | NC score | 0.040978 (rank : 27) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O60231, O60322, Q969X7, Q96QC1 | Gene names | DHX16, DBP2, DDX16, KIAA0577 | |||
|
Domain Architecture |

|
|||||
| Description | Putative pre-mRNA-splicing factor ATP-dependent RNA helicase DHX16 (EC 3.6.1.-) (DEAH-box protein 16) (ATP-dependent RNA helicase #3). | |||||
|
SEC63_HUMAN
|
||||||
| θ value | 0.365318 (rank : 8) | NC score | 0.095057 (rank : 7) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
M3K13_HUMAN
|
||||||
| θ value | 0.47712 (rank : 9) | NC score | 0.010971 (rank : 40) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 911 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43283 | Gene names | MAP3K13, LZK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 13 (EC 2.7.11.25) (Mixed lineage kinase) (MLK) (Leucine zipper-bearing kinase). | |||||
|
TXND2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 10) | NC score | 0.047340 (rank : 24) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
CC47_MOUSE
|
||||||
| θ value | 0.813845 (rank : 11) | NC score | 0.073589 (rank : 15) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9D024, Q3TSB5, Q3V1S7, Q8C5D9, Q920S6 | Gene names | Ccdc47, Asp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 47 precursor (Adipocyte-specific protein 4). | |||||
|
NF2L3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 12) | NC score | 0.060123 (rank : 18) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y4A8, Q6NUS0, Q7Z498, Q86UJ4, Q86VR5, Q9UQA4 | Gene names | NFE2L3, NRF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear factor erythroid 2-related factor 3 (NF-E2-related factor 3) (NFE2-related factor 3) (Nuclear factor, erythroid derived 2, like 3). | |||||
|
DSPP_HUMAN
|
||||||
| θ value | 1.38821 (rank : 13) | NC score | 0.075043 (rank : 14) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 14) | NC score | 0.085851 (rank : 8) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
NSBP1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 15) | NC score | 0.076081 (rank : 13) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
PPIG_HUMAN
|
||||||
| θ value | 1.81305 (rank : 16) | NC score | 0.036003 (rank : 29) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q13427, O00706, Q96DG9 | Gene names | PPIG | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase G (EC 5.2.1.8) (Peptidyl-prolyl isomerase G) (PPIase G) (Rotamase G) (Cyclophilin G) (Clk-associating RS-cyclophilin) (CARS-cyclophilin) (CARS-Cyp) (SR-cyclophilin) (SRcyp) (SR-cyp) (CASP10). | |||||
|
CAD16_MOUSE
|
||||||
| θ value | 2.36792 (rank : 17) | NC score | 0.015106 (rank : 39) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88338, Q9JLZ5 | Gene names | Cdh16 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-16 precursor (Kidney-specific cadherin) (Ksp-cadherin). | |||||
|
CLSPN_HUMAN
|
||||||
| θ value | 2.36792 (rank : 18) | NC score | 0.057011 (rank : 21) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9HAW4, Q5VYG0, Q6P6H5, Q8IWI1 | Gene names | CLSPN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Claspin (hClaspin) (Hu-Claspin). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 19) | NC score | 0.049335 (rank : 22) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
TCF8_HUMAN
|
||||||
| θ value | 2.36792 (rank : 20) | NC score | 0.010943 (rank : 41) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 1057 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P37275, Q12924, Q13800 | Gene names | TCF8, AREB6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (NIL-2-A zinc finger protein) (Negative regulator of IL2). | |||||
|
TGON2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 21) | NC score | 0.041806 (rank : 26) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q62314 | Gene names | Tgoln2, Ttgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (TGN38B). | |||||
|
ZCH11_HUMAN
|
||||||
| θ value | 2.36792 (rank : 22) | NC score | 0.043616 (rank : 25) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5TAX3, Q12764, Q5TAX4, Q86XZ3 | Gene names | ZCCHC11, KIAA0191 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 11. | |||||
|
TCAL5_HUMAN
|
||||||
| θ value | 3.0926 (rank : 23) | NC score | 0.039578 (rank : 28) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
ZN292_HUMAN
|
||||||
| θ value | 3.0926 (rank : 24) | NC score | 0.033141 (rank : 32) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 918 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60281, Q9H8G3, Q9H8J4 | Gene names | ZNF292, KIAA0530 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 292. | |||||
|
ERIC1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 25) | NC score | 0.080797 (rank : 9) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
I10R2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 26) | NC score | 0.033484 (rank : 31) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61190 | Gene names | Il10rb, Crfb4 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-10 receptor beta chain precursor (IL-10R-B) (IL-10R2) (Cytokine receptor family 2 member 4) (Cytokine receptor class-II member 4) (CRF2-4) (CDw210b antigen). | |||||
|
K0494_MOUSE
|
||||||
| θ value | 4.03905 (rank : 27) | NC score | 0.033550 (rank : 30) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BGQ6, Q3TQN6, Q8BJV7, Q8C0R1 | Gene names | Kiaa0494 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized calcium-binding protein KIAA0494. | |||||
|
DCDC2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 28) | NC score | 0.031180 (rank : 33) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5DU00, Q5SZU0, Q80Y99 | Gene names | Dcdc2, Kiaa1154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublecortin domain-containing protein 2. | |||||
|
F10A4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 29) | NC score | 0.018415 (rank : 38) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IZP2 | Gene names | FAM10A4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM10A4. | |||||
|
TF2AY_MOUSE
|
||||||
| θ value | 5.27518 (rank : 30) | NC score | 0.030234 (rank : 34) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8R4I4, Q99PM2, Q9DAF4 | Gene names | Gtf2a1lf, Alf | |||
|
Domain Architecture |

|
|||||
| Description | TFIIA-alpha and beta-like factor (General transcription factor II A, 1-like factor). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 31) | NC score | 0.048530 (rank : 23) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
PMS2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 32) | NC score | 0.019520 (rank : 36) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P54278 | Gene names | PMS2, PMSL2 | |||
|
Domain Architecture |
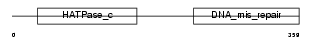
|
|||||
| Description | PMS1 protein homolog 2 (DNA mismatch repair protein PMS2). | |||||
|
FA35A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 33) | NC score | 0.020388 (rank : 35) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3UEN2 | Gene names | Fam35a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM35A. | |||||
|
NBEA_MOUSE
|
||||||
| θ value | 8.99809 (rank : 34) | NC score | 0.009768 (rank : 42) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9EPN1, Q8C931, Q9EPM9, Q9EPN0, Q9WVM9 | Gene names | Nbea, Lyst2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein neurobeachin (Lysosomal-trafficking regulator 2). | |||||
|
THUM1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 35) | NC score | 0.019290 (rank : 37) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99J36, Q3U6Y3 | Gene names | Thumpd1 | |||
|
Domain Architecture |

|
|||||
| Description | THUMP domain-containing protein 1. | |||||
|
NSBP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 36) | NC score | 0.060295 (rank : 17) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
NUCKS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 37) | NC score | 0.058507 (rank : 20) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
NUCKS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 38) | NC score | 0.059000 (rank : 19) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q80XU3, Q8BVD8 | Gene names | Nucks1, Nucks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (JC7). | |||||
|
PTMA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 39) | NC score | 0.076193 (rank : 12) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P06454, Q15249, Q15592 | Gene names | PTMA, TMSA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha-1]. | |||||
|
PTMA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 40) | NC score | 0.079468 (rank : 10) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P26350, Q3UQV6 | Gene names | Ptma | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha]. | |||||
|
PTMS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 41) | NC score | 0.060755 (rank : 16) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P20962 | Gene names | PTMS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
PTMS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 42) | NC score | 0.076845 (rank : 11) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9D0J8 | Gene names | Ptms | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
S30BP_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 9.92011e-124 (rank : 1) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q02614, Q8VDJ5 | Gene names | Sap30bp, Hcngp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAP30-binding protein (Transcriptional regulator protein HCNGP). | |||||
|
S30BP_HUMAN
|
||||||
| NC score | 0.992571 (rank : 2) | θ value | 1.02657e-120 (rank : 2) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UHR5, Q8N1R5, Q8TDR8, Q96D27 | Gene names | SAP30BP, HCNGP, HTRG, HTRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAP30-binding protein (Transcriptional regulator protein HCNGP). | |||||
|
VPS72_MOUSE
|
||||||
| NC score | 0.162224 (rank : 3) | θ value | 0.0563607 (rank : 5) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q62481, Q3U2N4, Q810A9, Q99K81 | Gene names | Vps72, Tcfl1, Yl1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
VPS72_HUMAN
|
||||||
| NC score | 0.160876 (rank : 4) | θ value | 0.0563607 (rank : 4) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15906, Q53GJ2, Q5U0R4 | Gene names | VPS72, TCFL1, YL1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.098723 (rank : 5) | θ value | 0.0330416 (rank : 3) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
SEC63_MOUSE
|
||||||
| NC score | 0.097943 (rank : 6) | θ value | 0.125558 (rank : 6) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
SEC63_HUMAN
|
||||||
| NC score | 0.095057 (rank : 7) | θ value | 0.365318 (rank : 8) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.085851 (rank : 8) | θ value | 1.38821 (rank : 14) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
ERIC1_HUMAN
|
||||||
| NC score | 0.080797 (rank : 9) | θ value | 4.03905 (rank : 25) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
PTMA_MOUSE
|
||||||
| NC score | 0.079468 (rank : 10) | θ value | θ > 10 (rank : 40) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P26350, Q3UQV6 | Gene names | Ptma | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha]. | |||||
|
PTMS_MOUSE
|
||||||
| NC score | 0.076845 (rank : 11) | θ value | θ > 10 (rank : 42) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9D0J8 | Gene names | Ptms | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
PTMA_HUMAN
|
||||||
| NC score | 0.076193 (rank : 12) | θ value | θ > 10 (rank : 39) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P06454, Q15249, Q15592 | Gene names | PTMA, TMSA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha-1]. | |||||
|
NSBP1_MOUSE
|
||||||
| NC score | 0.076081 (rank : 13) | θ value | 1.38821 (rank : 15) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.075043 (rank : 14) | θ value | 1.38821 (rank : 13) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
CC47_MOUSE
|
||||||
| NC score | 0.073589 (rank : 15) | θ value | 0.813845 (rank : 11) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9D024, Q3TSB5, Q3V1S7, Q8C5D9, Q920S6 | Gene names | Ccdc47, Asp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 47 precursor (Adipocyte-specific protein 4). | |||||
|
PTMS_HUMAN
|
||||||
| NC score | 0.060755 (rank : 16) | θ value | θ > 10 (rank : 41) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P20962 | Gene names | PTMS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
NSBP1_HUMAN
|
||||||
| NC score | 0.060295 (rank : 17) | θ value | θ > 10 (rank : 36) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
NF2L3_HUMAN
|
||||||
| NC score | 0.060123 (rank : 18) | θ value | 0.813845 (rank : 12) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y4A8, Q6NUS0, Q7Z498, Q86UJ4, Q86VR5, Q9UQA4 | Gene names | NFE2L3, NRF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear factor erythroid 2-related factor 3 (NF-E2-related factor 3) (NFE2-related factor 3) (Nuclear factor, erythroid derived 2, like 3). | |||||
|
NUCKS_MOUSE
|
||||||
| NC score | 0.059000 (rank : 19) | θ value | θ > 10 (rank : 38) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q80XU3, Q8BVD8 | Gene names | Nucks1, Nucks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (JC7). | |||||
|
NUCKS_HUMAN
|
||||||
| NC score | 0.058507 (rank : 20) | θ value | θ > 10 (rank : 37) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
CLSPN_HUMAN
|
||||||
| NC score | 0.057011 (rank : 21) | θ value | 2.36792 (rank : 18) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9HAW4, Q5VYG0, Q6P6H5, Q8IWI1 | Gene names | CLSPN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Claspin (hClaspin) (Hu-Claspin). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.049335 (rank : 22) | θ value | 2.36792 (rank : 19) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.048530 (rank : 23) | θ value | 6.88961 (rank : 31) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.047340 (rank : 24) | θ value | 0.62314 (rank : 10) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
ZCH11_HUMAN
|
||||||
| NC score | 0.043616 (rank : 25) | θ value | 2.36792 (rank : 22) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5TAX3, Q12764, Q5TAX4, Q86XZ3 | Gene names | ZCCHC11, KIAA0191 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 11. | |||||
|
TGON2_MOUSE
|
||||||
| NC score | 0.041806 (rank : 26) | θ value | 2.36792 (rank : 21) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q62314 | Gene names | Tgoln2, Ttgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (TGN38B). | |||||
|
DHX16_HUMAN
|
||||||
| NC score | 0.040978 (rank : 27) | θ value | 0.365318 (rank : 7) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O60231, O60322, Q969X7, Q96QC1 | Gene names | DHX16, DBP2, DDX16, KIAA0577 | |||
|
Domain Architecture |

|
|||||
| Description | Putative pre-mRNA-splicing factor ATP-dependent RNA helicase DHX16 (EC 3.6.1.-) (DEAH-box protein 16) (ATP-dependent RNA helicase #3). | |||||
|
TCAL5_HUMAN
|
||||||
| NC score | 0.039578 (rank : 28) | θ value | 3.0926 (rank : 23) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
PPIG_HUMAN
|
||||||
| NC score | 0.036003 (rank : 29) | θ value | 1.81305 (rank : 16) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q13427, O00706, Q96DG9 | Gene names | PPIG | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase G (EC 5.2.1.8) (Peptidyl-prolyl isomerase G) (PPIase G) (Rotamase G) (Cyclophilin G) (Clk-associating RS-cyclophilin) (CARS-cyclophilin) (CARS-Cyp) (SR-cyclophilin) (SRcyp) (SR-cyp) (CASP10). | |||||
|
K0494_MOUSE
|
||||||
| NC score | 0.033550 (rank : 30) | θ value | 4.03905 (rank : 27) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BGQ6, Q3TQN6, Q8BJV7, Q8C0R1 | Gene names | Kiaa0494 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized calcium-binding protein KIAA0494. | |||||
|
I10R2_MOUSE
|
||||||
| NC score | 0.033484 (rank : 31) | θ value | 4.03905 (rank : 26) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61190 | Gene names | Il10rb, Crfb4 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-10 receptor beta chain precursor (IL-10R-B) (IL-10R2) (Cytokine receptor family 2 member 4) (Cytokine receptor class-II member 4) (CRF2-4) (CDw210b antigen). | |||||
|
ZN292_HUMAN
|
||||||
| NC score | 0.033141 (rank : 32) | θ value | 3.0926 (rank : 24) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 918 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60281, Q9H8G3, Q9H8J4 | Gene names | ZNF292, KIAA0530 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 292. | |||||
|
DCDC2_MOUSE
|
||||||
| NC score | 0.031180 (rank : 33) | θ value | 5.27518 (rank : 28) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5DU00, Q5SZU0, Q80Y99 | Gene names | Dcdc2, Kiaa1154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublecortin domain-containing protein 2. | |||||
|
TF2AY_MOUSE
|
||||||
| NC score | 0.030234 (rank : 34) | θ value | 5.27518 (rank : 30) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8R4I4, Q99PM2, Q9DAF4 | Gene names | Gtf2a1lf, Alf | |||
|
Domain Architecture |

|
|||||
| Description | TFIIA-alpha and beta-like factor (General transcription factor II A, 1-like factor). | |||||
|
FA35A_MOUSE
|
||||||
| NC score | 0.020388 (rank : 35) | θ value | 8.99809 (rank : 33) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3UEN2 | Gene names | Fam35a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM35A. | |||||
|
PMS2_HUMAN
|
||||||
| NC score | 0.019520 (rank : 36) | θ value | 6.88961 (rank : 32) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P54278 | Gene names | PMS2, PMSL2 | |||
|
Domain Architecture |

|
|||||
| Description | PMS1 protein homolog 2 (DNA mismatch repair protein PMS2). | |||||
|
THUM1_MOUSE
|
||||||
| NC score | 0.019290 (rank : 37) | θ value | 8.99809 (rank : 35) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99J36, Q3U6Y3 | Gene names | Thumpd1 | |||
|
Domain Architecture |

|
|||||
| Description | THUMP domain-containing protein 1. | |||||
|
F10A4_HUMAN
|
||||||
| NC score | 0.018415 (rank : 38) | θ value | 5.27518 (rank : 29) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IZP2 | Gene names | FAM10A4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM10A4. | |||||
|
CAD16_MOUSE
|
||||||
| NC score | 0.015106 (rank : 39) | θ value | 2.36792 (rank : 17) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88338, Q9JLZ5 | Gene names | Cdh16 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-16 precursor (Kidney-specific cadherin) (Ksp-cadherin). | |||||
|
M3K13_HUMAN
|
||||||
| NC score | 0.010971 (rank : 40) | θ value | 0.47712 (rank : 9) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 911 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43283 | Gene names | MAP3K13, LZK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 13 (EC 2.7.11.25) (Mixed lineage kinase) (MLK) (Leucine zipper-bearing kinase). | |||||
|
TCF8_HUMAN
|
||||||
| NC score | 0.010943 (rank : 41) | θ value | 2.36792 (rank : 20) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 1057 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P37275, Q12924, Q13800 | Gene names | TCF8, AREB6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (NIL-2-A zinc finger protein) (Negative regulator of IL2). | |||||
|
NBEA_MOUSE
|
||||||
| NC score | 0.009768 (rank : 42) | θ value | 8.99809 (rank : 34) | |||
| Query Neighborhood Hits | 35 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9EPN1, Q8C931, Q9EPM9, Q9EPN0, Q9WVM9 | Gene names | Nbea, Lyst2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein neurobeachin (Lysosomal-trafficking regulator 2). | |||||