Please be patient as the page loads
|
FA35A_HUMAN
|
||||||
| SwissProt Accessions | Q86V20, O95885, Q9H991 | Gene names | FAM35A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM35A. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
FA35A_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q86V20, O95885, Q9H991 | Gene names | FAM35A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM35A. | |||||
|
FA35A_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.946480 (rank : 2) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q3UEN2 | Gene names | Fam35a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM35A. | |||||
|
NEDD1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 3) | NC score | 0.043211 (rank : 5) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 294 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NHV4, Q8NA30 | Gene names | NEDD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein NEDD1. | |||||
|
CADH2_MOUSE
|
||||||
| θ value | 0.21417 (rank : 4) | NC score | 0.014148 (rank : 13) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P15116, Q64260 | Gene names | Cdh2 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-2 precursor (Neural-cadherin) (N-cadherin) (CDw325 antigen). | |||||
|
IFT74_MOUSE
|
||||||
| θ value | 0.21417 (rank : 5) | NC score | 0.026744 (rank : 10) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 939 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BKE9, Q80ZI3, Q9CSY1, Q9CUS0, Q9D9T5 | Gene names | Ift74, Ccdc2, Cmg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 74 homolog (Coiled-coil domain-containing protein 2) (Capillary morphogenesis protein 1) (CMG-1). | |||||
|
BPAEA_MOUSE
|
||||||
| θ value | 0.47712 (rank : 6) | NC score | 0.015864 (rank : 11) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 1480 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q91ZU8, Q8K5D4 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoform 5 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 0.47712 (rank : 7) | NC score | 0.028048 (rank : 8) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
OLIG3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 8) | NC score | 0.027513 (rank : 9) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7RTU3, Q8N8Q0 | Gene names | OLIG3, BHLHB7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oligodendrocyte transcription factor 3 (Oligo3) (Basic helix-loop- helix domain-containing class B protein 7). | |||||
|
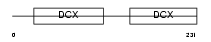
RP1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 9) | NC score | 0.031250 (rank : 7) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P56716 | Gene names | Rp1, Orp1, Rp1h | |||
|
Domain Architecture |
|
|||||
| Description | Oxygen-regulated protein 1 (Retinitis pigmentosa RP1 protein homolog). | |||||
|
SPT16_HUMAN
|
||||||
| θ value | 1.38821 (rank : 10) | NC score | 0.036946 (rank : 6) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5B9, Q6GMT8, Q6P2F1, Q6PJM1, Q9NRX0 | Gene names | SUPT16H, FACT140, FACTP140 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FACT complex subunit SPT16 (Facilitates chromatin transcription complex subunit SPT16) (hSPT16) (FACT 140 kDa subunit) (FACTp140) (Chromatin-specific transcription elongation factor 140 kDa subunit). | |||||
|
ENL_HUMAN
|
||||||
| θ value | 2.36792 (rank : 11) | NC score | 0.051153 (rank : 4) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
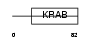
ZN175_HUMAN
|
||||||
| θ value | 3.0926 (rank : 12) | NC score | 0.001579 (rank : 20) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 774 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y473 | Gene names | ZNF175 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 175 (Zinc finger protein OTK18). | |||||
|
SYNE1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 13) | NC score | 0.009741 (rank : 16) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
GOGA5_MOUSE
|
||||||
| θ value | 6.88961 (rank : 14) | NC score | 0.007388 (rank : 17) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 1434 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9QYE6, O88317, Q3TGE7, Q3U6S5, Q3UUF9 | Gene names | Golga5, Retii | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (Sumiko protein) (Ret-II protein). | |||||
|
MLTK_MOUSE
|
||||||
| θ value | 6.88961 (rank : 15) | NC score | 0.001547 (rank : 21) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ESL4, Q3V1X8, Q8BR73, Q9ESL3 | Gene names | Mltk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase MLT (EC 2.7.11.25) (MLK-like mitogen-activated protein triple kinase) (Leucine zipper- and sterile alpha motif kinase ZAK) (Sterile alpha motif- and leucine zipper-containing kinase AZK) (Mixed lineage kinase-related kinase) (MLK-related kinase) (MRK). | |||||
|
SENP1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 16) | NC score | 0.014555 (rank : 12) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9P0U3, Q86XC8 | Gene names | SENP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 1 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP1). | |||||
|
UBE1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 17) | NC score | 0.013693 (rank : 14) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P22314, Q96E13 | Gene names | UBE1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-activating enzyme E1 (A1S9 protein). | |||||
|
CAC1A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 18) | NC score | 0.006449 (rank : 18) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P97445 | Gene names | Cacna1a, Cach4, Cacn3, Cacnl1a4, Ccha1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
MKL2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 19) | NC score | 0.011568 (rank : 15) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P59759 | Gene names | Mkl2, Mrtfb | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 2 (Myocardin-related transcription factor B) (MRTF-B). | |||||
|
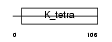
ZN238_HUMAN
|
||||||
| θ value | 8.99809 (rank : 20) | NC score | 0.001911 (rank : 19) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99592, Q13397, Q5VU40, Q8N463 | Gene names | ZNF238, RP58, TAZ1, ZBTB18 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 238 (Transcriptional repressor RP58) (58 kDa repressor protein) (Zinc finger protein C2H2-171) (Translin-associated zinc finger protein 1) (TAZ-1) (Zinc finger and BTB domain-containing protein 18). | |||||
|
RSRC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 21) | NC score | 0.053378 (rank : 3) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9DBU6, Q3TF06 | Gene names | Rsrc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine/serine-rich coiled coil protein 1. | |||||
|
FA35A_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q86V20, O95885, Q9H991 | Gene names | FAM35A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM35A. | |||||
|
FA35A_MOUSE
|
||||||
| NC score | 0.946480 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q3UEN2 | Gene names | Fam35a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM35A. | |||||
|
RSRC1_MOUSE
|
||||||
| NC score | 0.053378 (rank : 3) | θ value | θ > 10 (rank : 21) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9DBU6, Q3TF06 | Gene names | Rsrc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine/serine-rich coiled coil protein 1. | |||||
|
ENL_HUMAN
|
||||||
| NC score | 0.051153 (rank : 4) | θ value | 2.36792 (rank : 11) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
NEDD1_HUMAN
|
||||||
| NC score | 0.043211 (rank : 5) | θ value | 0.125558 (rank : 3) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 294 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NHV4, Q8NA30 | Gene names | NEDD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein NEDD1. | |||||
|
SPT16_HUMAN
|
||||||
| NC score | 0.036946 (rank : 6) | θ value | 1.38821 (rank : 10) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5B9, Q6GMT8, Q6P2F1, Q6PJM1, Q9NRX0 | Gene names | SUPT16H, FACT140, FACTP140 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FACT complex subunit SPT16 (Facilitates chromatin transcription complex subunit SPT16) (hSPT16) (FACT 140 kDa subunit) (FACTp140) (Chromatin-specific transcription elongation factor 140 kDa subunit). | |||||
|
RP1_MOUSE
|
||||||
| NC score | 0.031250 (rank : 7) | θ value | 1.38821 (rank : 9) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P56716 | Gene names | Rp1, Orp1, Rp1h | |||
|
Domain Architecture |

|
|||||
| Description | Oxygen-regulated protein 1 (Retinitis pigmentosa RP1 protein homolog). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.028048 (rank : 8) | θ value | 0.47712 (rank : 7) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
OLIG3_HUMAN
|
||||||
| NC score | 0.027513 (rank : 9) | θ value | 1.38821 (rank : 8) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7RTU3, Q8N8Q0 | Gene names | OLIG3, BHLHB7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oligodendrocyte transcription factor 3 (Oligo3) (Basic helix-loop- helix domain-containing class B protein 7). | |||||
|
IFT74_MOUSE
|
||||||
| NC score | 0.026744 (rank : 10) | θ value | 0.21417 (rank : 5) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 939 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BKE9, Q80ZI3, Q9CSY1, Q9CUS0, Q9D9T5 | Gene names | Ift74, Ccdc2, Cmg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 74 homolog (Coiled-coil domain-containing protein 2) (Capillary morphogenesis protein 1) (CMG-1). | |||||
|
BPAEA_MOUSE
|
||||||
| NC score | 0.015864 (rank : 11) | θ value | 0.47712 (rank : 6) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 1480 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q91ZU8, Q8K5D4 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoform 5 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
SENP1_HUMAN
|
||||||
| NC score | 0.014555 (rank : 12) | θ value | 6.88961 (rank : 16) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9P0U3, Q86XC8 | Gene names | SENP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 1 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP1). | |||||
|
CADH2_MOUSE
|
||||||
| NC score | 0.014148 (rank : 13) | θ value | 0.21417 (rank : 4) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P15116, Q64260 | Gene names | Cdh2 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-2 precursor (Neural-cadherin) (N-cadherin) (CDw325 antigen). | |||||
|
UBE1_HUMAN
|
||||||
| NC score | 0.013693 (rank : 14) | θ value | 6.88961 (rank : 17) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P22314, Q96E13 | Gene names | UBE1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-activating enzyme E1 (A1S9 protein). | |||||
|
MKL2_MOUSE
|
||||||
| NC score | 0.011568 (rank : 15) | θ value | 8.99809 (rank : 19) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P59759 | Gene names | Mkl2, Mrtfb | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 2 (Myocardin-related transcription factor B) (MRTF-B). | |||||
|
SYNE1_HUMAN
|
||||||
| NC score | 0.009741 (rank : 16) | θ value | 5.27518 (rank : 13) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
GOGA5_MOUSE
|
||||||
| NC score | 0.007388 (rank : 17) | θ value | 6.88961 (rank : 14) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 1434 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9QYE6, O88317, Q3TGE7, Q3U6S5, Q3UUF9 | Gene names | Golga5, Retii | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (Sumiko protein) (Ret-II protein). | |||||
|
CAC1A_MOUSE
|
||||||
| NC score | 0.006449 (rank : 18) | θ value | 8.99809 (rank : 18) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P97445 | Gene names | Cacna1a, Cach4, Cacn3, Cacnl1a4, Ccha1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
ZN238_HUMAN
|
||||||
| NC score | 0.001911 (rank : 19) | θ value | 8.99809 (rank : 20) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99592, Q13397, Q5VU40, Q8N463 | Gene names | ZNF238, RP58, TAZ1, ZBTB18 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 238 (Transcriptional repressor RP58) (58 kDa repressor protein) (Zinc finger protein C2H2-171) (Translin-associated zinc finger protein 1) (TAZ-1) (Zinc finger and BTB domain-containing protein 18). | |||||
|
ZN175_HUMAN
|
||||||
| NC score | 0.001579 (rank : 20) | θ value | 3.0926 (rank : 12) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 774 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y473 | Gene names | ZNF175 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 175 (Zinc finger protein OTK18). | |||||
|
MLTK_MOUSE
|
||||||
| NC score | 0.001547 (rank : 21) | θ value | 6.88961 (rank : 15) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ESL4, Q3V1X8, Q8BR73, Q9ESL3 | Gene names | Mltk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase MLT (EC 2.7.11.25) (MLK-like mitogen-activated protein triple kinase) (Leucine zipper- and sterile alpha motif kinase ZAK) (Sterile alpha motif- and leucine zipper-containing kinase AZK) (Mixed lineage kinase-related kinase) (MLK-related kinase) (MRK). | |||||