Please be patient as the page loads
|
MT2_HUMAN
|
||||||
| SwissProt Accessions | P02795, Q14823, Q2HXR9, Q53XT9 | Gene names | MT2A, CES1, MT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-2 (MT-2) (Metallothionein-II) (MT-II) (Metallothionein-2A). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
MT2_HUMAN
|
||||||
| θ value | 7.09661e-13 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P02795, Q14823, Q2HXR9, Q53XT9 | Gene names | MT2A, CES1, MT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-2 (MT-2) (Metallothionein-II) (MT-II) (Metallothionein-2A). | |||||
|
MT1I_HUMAN
|
||||||
| θ value | 2.28291e-11 (rank : 2) | NC score | 0.986700 (rank : 2) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | P80295 | Gene names | MT1I | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1I (MT-1I) (Metallothionein-II). | |||||
|
MT1E_HUMAN
|
||||||
| θ value | 5.08577e-11 (rank : 3) | NC score | 0.981287 (rank : 4) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P04732, Q86YX4 | Gene names | MT1E | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1E (MT-1E) (Metallothionein-IE) (MT-IE). | |||||
|
MT1F_HUMAN
|
||||||
| θ value | 8.67504e-11 (rank : 4) | NC score | 0.984021 (rank : 3) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P04733, Q9UI97 | Gene names | MT1F | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1F (MT-1F) (Metallothionein-IF) (MT-IF). | |||||
|
MT1M_HUMAN
|
||||||
| θ value | 8.67504e-11 (rank : 5) | NC score | 0.967633 (rank : 5) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8N339, Q8TDN3 | Gene names | MT1M | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1M (MT-1M) (Metallothionein-IM) (MT-IM). | |||||
|
MT1A_HUMAN
|
||||||
| θ value | 1.133e-10 (rank : 6) | NC score | 0.893290 (rank : 10) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P04731 | Gene names | MT1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1A (MT-1A) (Metallothionein-IA) (MT-IA). | |||||
|
MT1H_HUMAN
|
||||||
| θ value | 1.9326e-10 (rank : 7) | NC score | 0.902145 (rank : 9) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | P80294 | Gene names | MT1H | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1H (MT-1H) (Metallothionein-IH) (MT-IH) (Metallothionein-0) (MT-0). | |||||
|
MT1L_HUMAN
|
||||||
| θ value | 7.34386e-10 (rank : 8) | NC score | 0.917024 (rank : 8) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q93083 | Gene names | MT1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1L (MT-1L) (Metallothionein-IL) (MT-IL). | |||||
|
MT1G_HUMAN
|
||||||
| θ value | 2.79066e-09 (rank : 9) | NC score | 0.928633 (rank : 7) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P13640, P80296 | Gene names | MT1G, MT1K, MT1M | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1G (MT-1G) (Metallothionein-IG) (MT-IG) (Metallothionein-1K) (MT-1K). | |||||
|
MT1_MOUSE
|
||||||
| θ value | 2.79066e-09 (rank : 10) | NC score | 0.882860 (rank : 11) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P02802, Q64485 | Gene names | Mt1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1 (MT-1) (Metallothionein-I) (MT-I). | |||||
|
MT2_MOUSE
|
||||||
| θ value | 3.64472e-09 (rank : 11) | NC score | 0.846436 (rank : 13) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P02798 | Gene names | Mt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-2 (MT-2) (Metallothionein-II) (MT-II). | |||||
|
MT1B_HUMAN
|
||||||
| θ value | 4.76016e-09 (rank : 12) | NC score | 0.866267 (rank : 12) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | P07438 | Gene names | MT1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1B (MT-1B) (Metallothionein-IB) (MT-IB). | |||||
|
MT1X_HUMAN
|
||||||
| θ value | 4.76016e-09 (rank : 13) | NC score | 0.941448 (rank : 6) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P80297 | Gene names | MT1X | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1X (MT-1X) (Metallothionein-IX) (MT-IX). | |||||
|
CT127_HUMAN
|
||||||
| θ value | 3.41135e-07 (rank : 14) | NC score | 0.734564 (rank : 16) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9BQN2 | Gene names | C20orf127 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative metallothionein C20orf127. | |||||
|
MT4_MOUSE
|
||||||
| θ value | 0.000121331 (rank : 15) | NC score | 0.754040 (rank : 15) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P47945 | Gene names | Mt4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-4 (MT-4) (Metallothionein-IV) (MT-IV). | |||||
|
MT4_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 16) | NC score | 0.758874 (rank : 14) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P47944 | Gene names | MT4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-4 (MT-4) (Metallothionein-IV) (MT-IV). | |||||
|
MT3_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 17) | NC score | 0.564255 (rank : 17) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P25713, Q2V574 | Gene names | MT3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-3 (MT-3) (Metallothionein-III) (MT-III) (Growth inhibitory factor) (GIF) (GIFB). | |||||
|
MT3_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 18) | NC score | 0.472518 (rank : 18) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P28184 | Gene names | Mt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-3 (MT-3) (Metallothionein-III) (MT-III) (Growth inhibitory factor) (GIF). | |||||
|
PCSK5_MOUSE
|
||||||
| θ value | 0.125558 (rank : 19) | NC score | 0.138728 (rank : 32) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 613 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q04592, Q62040 | Gene names | Pcsk5 | |||
|
Domain Architecture |

|
|||||
| Description | Proprotein convertase subtilisin/kexin type 5 precursor (EC 3.4.21.-) (Proprotein convertase PC5) (Subtilisin/kexin-like protease PC5) (PC6) (Subtilisin-like proprotein convertase 6) (SPC6). | |||||
|
KIF4A_HUMAN
|
||||||
| θ value | 0.279714 (rank : 20) | NC score | 0.055837 (rank : 142) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O95239, Q86TN3, Q86XX7, Q9NNY6, Q9NY24, Q9UMW3 | Gene names | KIF4A, KIF4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
LAMA5_MOUSE
|
||||||
| θ value | 0.279714 (rank : 21) | NC score | 0.110482 (rank : 38) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q61001, Q9JHQ6 | Gene names | Lama5 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-5 chain precursor. | |||||
|
ADA28_HUMAN
|
||||||
| θ value | 0.365318 (rank : 22) | NC score | 0.112133 (rank : 37) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9UKQ2, Q9Y339, Q9Y3S0 | Gene names | ADAM28, MDCL | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 28 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 28) (Metalloproteinase-like, disintegrin-like, and cysteine- rich protein-L) (MDC-L) (eMDC II) (ADAM23). | |||||
|
ITB5_MOUSE
|
||||||
| θ value | 0.365318 (rank : 23) | NC score | 0.196900 (rank : 19) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O70309, O70308, O88347 | Gene names | Itgb5 | |||
|
Domain Architecture |
|
|||||
| Description | Integrin beta-5 precursor. | |||||
|
KIF4A_MOUSE
|
||||||
| θ value | 0.365318 (rank : 24) | NC score | 0.055013 (rank : 144) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1033 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P33174 | Gene names | Kif4a, Kif4, Kns4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
ADA28_MOUSE
|
||||||
| θ value | 0.47712 (rank : 25) | NC score | 0.109924 (rank : 39) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9JLN6, Q8K5D2, Q8K5D3 | Gene names | Adam28 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 28 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 28) (Thymic epithelial cell-ADAM) (TECADAM). | |||||
|
ITB1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 26) | NC score | 0.191509 (rank : 20) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P05556, P78466, P78467, Q13089, Q13090, Q13091, Q13212, Q14622, Q14647 | Gene names | ITGB1, FNRB | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-1 precursor (Fibronectin receptor subunit beta) (Integrin VLA-4 subunit beta) (CD29 antigen). | |||||
|
ITB1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 27) | NC score | 0.176058 (rank : 23) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P09055 | Gene names | Itgb1 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-1 precursor (Fibronectin receptor subunit beta) (Integrin VLA-4 subunit beta) (CD29 antigen). | |||||
|
ITB3_HUMAN
|
||||||
| θ value | 0.62314 (rank : 28) | NC score | 0.175411 (rank : 24) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P05106, O15495, Q13413, Q14648, Q16499 | Gene names | ITGB3, GP3A | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-3 precursor (Platelet membrane glycoprotein IIIa) (GPIIIa) (CD61 antigen). | |||||
|
ITB5_HUMAN
|
||||||
| θ value | 0.813845 (rank : 29) | NC score | 0.184611 (rank : 21) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P18084 | Gene names | ITGB5 | |||
|
Domain Architecture |
|
|||||
| Description | Integrin beta-5 precursor. | |||||
|
TNR9_HUMAN
|
||||||
| θ value | 0.813845 (rank : 30) | NC score | 0.132551 (rank : 33) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q07011 | Gene names | TNFRSF9, CD137, ILA | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 9 precursor (4-1BB ligand receptor) (T-cell antigen 4-1BB homolog) (T-cell antigen ILA) (CD137 antigen). | |||||
|
ITB8_HUMAN
|
||||||
| θ value | 1.06291 (rank : 31) | NC score | 0.166499 (rank : 25) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P26012 | Gene names | ITGB8 | |||
|
Domain Architecture |
|
|||||
| Description | Integrin beta-8 precursor. | |||||
|
CRIM1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 32) | NC score | 0.121723 (rank : 36) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9NZV1, Q59GH0, Q7LCQ5, Q9H318 | Gene names | CRIM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich motor neuron 1 protein precursor (CRIM-1) (Cysteine-rich repeat-containing protein S52). | |||||
|
MEGF6_HUMAN
|
||||||
| θ value | 1.38821 (rank : 33) | NC score | 0.097404 (rank : 52) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | O75095, Q5VV39 | Gene names | MEGF6, EGFL3, KIAA0815 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multiple epidermal growth factor-like domains 6 precursor (EGF-like domain-containing protein 3) (Multiple EGF-like domain protein 3). | |||||
|
ADA11_HUMAN
|
||||||
| θ value | 1.81305 (rank : 34) | NC score | 0.104859 (rank : 44) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O75078, Q14808, Q14809, Q14810 | Gene names | ADAM11, MDC | |||
|
Domain Architecture |
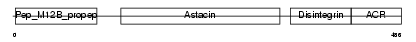
|
|||||
| Description | ADAM 11 precursor (A disintegrin and metalloproteinase domain 11) (Metalloproteinase-like, disintegrin-like, and cysteine-rich protein) (MDC). | |||||
|
ADA11_MOUSE
|
||||||
| θ value | 1.81305 (rank : 35) | NC score | 0.104136 (rank : 45) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9R1V4 | Gene names | Adam11, Mdc | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 11 precursor (A disintegrin and metalloproteinase domain 11) (Metalloproteinase-like, disintegrin-like, and cysteine-rich protein) (MDC). | |||||
|
ADA22_HUMAN
|
||||||
| θ value | 1.81305 (rank : 36) | NC score | 0.123373 (rank : 35) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9P0K1, O75075, O75076, Q9P0K2, Q9UIA1, Q9UKK2 | Gene names | ADAM22, MDC2 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 22 precursor (A disintegrin and metalloproteinase domain 22) (Metalloproteinase-like, disintegrin-like, and cysteine-rich protein 2) (Metalloproteinase-disintegrin ADAM22-3). | |||||
|
ADAM9_MOUSE
|
||||||
| θ value | 1.81305 (rank : 37) | NC score | 0.105464 (rank : 43) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q61072, Q60618, Q61853, Q80U94 | Gene names | Adam9, Kiaa0021, Mdc9, Mltng | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAM 9 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 9) (Metalloprotease/disintegrin/cysteine-rich protein 9) (Myeloma cell metalloproteinase) (Meltrin gamma). | |||||
|
ITB6_HUMAN
|
||||||
| θ value | 1.81305 (rank : 38) | NC score | 0.165321 (rank : 26) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P18564, Q16500 | Gene names | ITGB6 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-6 precursor. | |||||
|
LAMA1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 39) | NC score | 0.095162 (rank : 56) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 804 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P25391 | Gene names | LAMA1, LAMA | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-1 chain precursor (Laminin A chain). | |||||
|
LAMA2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 40) | NC score | 0.087296 (rank : 65) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1198 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q60675, Q05003, Q64061 | Gene names | Lama2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
LAMB2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 41) | NC score | 0.077951 (rank : 85) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q61292, Q62182 | Gene names | Lamb2, Lams | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (S-LAM). | |||||
|
ADA29_HUMAN
|
||||||
| θ value | 2.36792 (rank : 42) | NC score | 0.079274 (rank : 78) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9UKF5, Q9UHP1, Q9UKF3, Q9UKF4 | Gene names | ADAM29 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 29 precursor (A disintegrin and metalloproteinase domain 29). | |||||
|
ITB3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 43) | NC score | 0.177114 (rank : 22) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O54890 | Gene names | Itgb3 | |||
|
Domain Architecture |
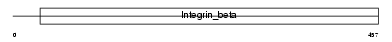
|
|||||
| Description | Integrin beta-3 precursor (Platelet membrane glycoprotein IIIa) (GPIIIa) (CD61 antigen). | |||||
|
USH2A_MOUSE
|
||||||
| θ value | 2.36792 (rank : 44) | NC score | 0.099100 (rank : 49) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 626 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q2QI47, Q9JLP3 | Gene names | Ush2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Usherin precursor (Usher syndrome type-2A protein homolog) (Usher syndrome type IIa protein homolog). | |||||
|
ADA22_MOUSE
|
||||||
| θ value | 3.0926 (rank : 45) | NC score | 0.129936 (rank : 34) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9R1V6, Q5TLI8, Q5TLI9, Q5TLJ0, Q5TLJ1, Q5TLJ2, Q5TLJ3, Q5TLJ4, Q5TLJ5, Q5TLJ6, Q5TLJ7, Q5TLJ8, Q5TLJ9, Q5TLK0, Q5TLK1, Q5TLK2, Q5TLK3, Q5TLK4, Q5TLK5, Q5TLK6, Q5TLK7, Q5TLK8, Q8BSF2, Q9R1V5 | Gene names | Adam22 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 22 precursor (A disintegrin and metalloproteinase domain 22). | |||||
|
ADAM9_HUMAN
|
||||||
| θ value | 3.0926 (rank : 46) | NC score | 0.098533 (rank : 50) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q13443, Q10718, Q8NFM6 | Gene names | ADAM9, KIAA0021, MCMP, MDC9, MLTNG | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 9 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 9) (Metalloprotease/disintegrin/cysteine-rich protein 9) (Myeloma cell metalloproteinase) (Meltrin gamma) (Cellular disintegrin-related protein). | |||||
|
FRAS1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 47) | NC score | 0.084022 (rank : 69) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q80T14, Q80TC7, Q811H8, Q8BPZ4 | Gene names | Fras1, Kiaa1500 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Extracellular matrix protein FRAS1 precursor. | |||||
|
JAG2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 48) | NC score | 0.056630 (rank : 140) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9QYE5, O55139, O70219 | Gene names | Jag2 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-2 precursor (Jagged2). | |||||
|
TNR7_MOUSE
|
||||||
| θ value | 3.0926 (rank : 49) | NC score | 0.109078 (rank : 40) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P41272 | Gene names | Tnfrsf7, Cd27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor necrosis factor receptor superfamily member 7 precursor (CD27L receptor) (T-cell activation antigen CD27). | |||||
|
ADA24_MOUSE
|
||||||
| θ value | 4.03905 (rank : 50) | NC score | 0.081369 (rank : 72) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9R160 | Gene names | Adam24 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 24 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 24) (Testase 1). | |||||
|
FRAS1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 51) | NC score | 0.083529 (rank : 70) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 759 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q86XX4, Q86UZ4, Q8N3U9, Q8NAU7, Q96JW7, Q9H6N9, Q9P228 | Gene names | FRAS1, KIAA1500 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Extracellular matrix protein FRAS1 precursor. | |||||
|
ITB7_MOUSE
|
||||||
| θ value | 4.03905 (rank : 52) | NC score | 0.163651 (rank : 27) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P26011, Q3U2M1, Q64656 | Gene names | Itgb7 | |||
|
Domain Architecture |
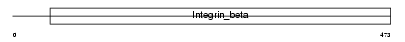
|
|||||
| Description | Integrin beta-7 precursor (Integrin beta-P) (M290 IEL antigen). | |||||
|
PCSK5_HUMAN
|
||||||
| θ value | 4.03905 (rank : 53) | NC score | 0.085827 (rank : 68) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q92824, Q13527 | Gene names | PCSK5, PC5, PC6 | |||
|
Domain Architecture |

|
|||||
| Description | Proprotein convertase subtilisin/kexin type 5 precursor (EC 3.4.21.-) (Proprotein convertase PC5) (Subtilisin/kexin-like protease PC5) (PC6) (hPC6). | |||||
|
CREL1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | 0.076199 (rank : 91) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 524 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q91XD7, Q8BGJ8 | Gene names | Creld1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich with EGF-like domain protein 1 precursor. | |||||
|
TNR7_HUMAN
|
||||||
| θ value | 5.27518 (rank : 55) | NC score | 0.102999 (rank : 47) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P26842 | Gene names | TNFRSF7, CD27 | |||
|
Domain Architecture |
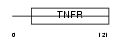
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 7 precursor (CD27L receptor) (T-cell activation antigen CD27) (T14). | |||||
|
ADAM7_HUMAN
|
||||||
| θ value | 6.88961 (rank : 56) | NC score | 0.077701 (rank : 86) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9H2U9, O75959 | Gene names | ADAM7, GP83 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 7 precursor (A disintegrin and metalloproteinase domain 7) (Sperm maturation-related glycoprotein GP-83). | |||||
|
ADAM8_MOUSE
|
||||||
| θ value | 6.88961 (rank : 57) | NC score | 0.080545 (rank : 74) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q05910 | Gene names | Adam8, Ms2 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 8 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 8) (Cell surface antigen MS2) (Macrophage cysteine-rich glycoprotein) (CD156 antigen). | |||||
|
FBLN2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | 0.031466 (rank : 159) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P37889, Q9WUI2 | Gene names | Fbln2 | |||
|
Domain Architecture |

|
|||||
| Description | Fibulin-2 precursor. | |||||
|
ITB7_HUMAN
|
||||||
| θ value | 6.88961 (rank : 59) | NC score | 0.161665 (rank : 28) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P26010 | Gene names | ITGB7 | |||
|
Domain Architecture |
|
|||||
| Description | Integrin beta-7 precursor. | |||||
|
PGBM_HUMAN
|
||||||
| θ value | 6.88961 (rank : 60) | NC score | 0.048024 (rank : 158) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1113 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P98160, Q16287, Q9H3V5 | Gene names | HSPG2 | |||
|
Domain Architecture |

|
|||||
| Description | Basement membrane-specific heparan sulfate proteoglycan core protein precursor (HSPG) (Perlecan) (PLC). | |||||
|
ST14_HUMAN
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | 0.013574 (rank : 160) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y5Y6, Q9BS01, Q9H3S0, Q9HB36, Q9HCA3 | Gene names | ST14, PRSS14, SNC19, TADG15 | |||
|
Domain Architecture |

|
|||||
| Description | Suppressor of tumorigenicity protein 14 (EC 3.4.21.-) (Serine protease 14) (Matriptase) (Membrane-type serine protease 1) (MT-SP1) (Prostamin) (Serine protease TADG-15) (Tumor-associated differentially-expressed gene 15 protein). | |||||
|
ZAN_MOUSE
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.079561 (rank : 75) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | O88799, O08647 | Gene names | Zan | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
ADA33_MOUSE
|
||||||
| θ value | 8.99809 (rank : 63) | NC score | 0.077398 (rank : 88) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q923W9 | Gene names | Adam33 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 33 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 33). | |||||
|
ADM1B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 64) | NC score | 0.078543 (rank : 82) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8R534, Q9R156 | Gene names | Adam1b | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 1b precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 1b) (Fertilin alpha 1b subunit). | |||||
|
ITB6_MOUSE
|
||||||
| θ value | 8.99809 (rank : 65) | NC score | 0.153381 (rank : 29) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Z0T9 | Gene names | Itgb6 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-6 precursor. | |||||
|
LAMB2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 66) | NC score | 0.073060 (rank : 105) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 692 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P55268, Q16321 | Gene names | LAMB2, LAMS | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (Laminin B1s chain). | |||||
|
SREC2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 67) | NC score | 0.091791 (rank : 60) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q96GP6, Q8IXF3, Q9BW74 | Gene names | SCARF2, SREC2, SREPCR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor class F member 2 precursor (Scavenger receptor expressed by endothelial cells 2 protein) (SREC-II) (SRECRP-1). | |||||
|
SREC2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 68) | NC score | 0.094085 (rank : 59) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 605 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P59222 | Gene names | Scarf2, Srec2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor class F member 2 precursor (Scavenger receptor expressed by endothelial cells 2 protein) (SREC-II). | |||||
|
USH2A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 69) | NC score | 0.078008 (rank : 84) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 600 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O75445, Q5VVM9, Q6S362, Q9NS27 | Gene names | USH2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Usherin precursor (Usher syndrome type-2A protein) (Usher syndrome type IIa protein). | |||||
|
AD26A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.073801 (rank : 99) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9R158 | Gene names | Adam26a, Adam26 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 26A precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 26A) (Testase-3). | |||||
|
ADA10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.070917 (rank : 117) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O14672, Q10742, Q92650 | Gene names | ADAM10, KUZ, MADM | |||
|
Domain Architecture |
|
|||||
| Description | ADAM 10 precursor (EC 3.4.24.81) (A disintegrin and metalloproteinase domain 10) (Mammalian disintegrin-metalloprotease) (Kuzbanian protein homolog) (CDw156c antigen). | |||||
|
ADA10_MOUSE
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.070793 (rank : 118) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O35598 | Gene names | Adam10, Kuz, Madm | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 10 precursor (EC 3.4.24.81) (A disintegrin and metalloproteinase domain 10) (Mammalian disintegrin-metalloprotease) (Kuzbanian protein homolog). | |||||
|
ADA12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 73) | NC score | 0.071391 (rank : 113) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 224 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O43184, O60470 | Gene names | ADAM12, MLTN | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 12 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 12) (Meltrin alpha). | |||||
|
ADA12_MOUSE
|
||||||
| θ value | θ > 10 (rank : 74) | NC score | 0.071473 (rank : 112) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q61824 | Gene names | Adam12, Mltna | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 12 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 12) (Meltrin alpha). | |||||
|
ADA15_HUMAN
|
||||||
| θ value | θ > 10 (rank : 75) | NC score | 0.079083 (rank : 79) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q13444, Q13493, Q96C78 | Gene names | ADAM15, MDC15 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 15 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 15) (Metalloproteinase-like, disintegrin-like, and cysteine- rich protein 15) (MDC-15) (Metalloprotease RGD disintegrin protein) (Metargidin). | |||||
|
ADA15_MOUSE
|
||||||
| θ value | θ > 10 (rank : 76) | NC score | 0.078674 (rank : 81) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O88839, Q3U7C2, Q8C7Z0, Q91VS9, Q9QYL2 | Gene names | Adam15, Mdc15 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 15 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 15) (Metalloproteinase-like, disintegrin-like, and cysteine- rich protein 15) (MDC-15) (Metalloprotease RGD disintegrin protein) (Metargidin) (AD56). | |||||
|
ADA17_HUMAN
|
||||||
| θ value | θ > 10 (rank : 77) | NC score | 0.073900 (rank : 98) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P78536, O60226 | Gene names | ADAM17, CSVP, TACE | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 17 precursor (EC 3.4.24.86) (A disintegrin and metalloproteinase domain 17) (TNF-alpha-converting enzyme) (TNF-alpha convertase) (Snake venom-like protease) (CD156b antigen). | |||||
|
ADA17_MOUSE
|
||||||
| θ value | θ > 10 (rank : 78) | NC score | 0.073323 (rank : 102) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9Z0F8, O88726, Q9R1U4, Q9Z0K3 | Gene names | Adam17, Tace | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 17 precursor (EC 3.4.24.86) (A disintegrin and metalloproteinase domain 17) (TNF-alpha-converting enzyme) (TNF-alpha convertase) (CD156b antigen). | |||||
|
ADA18_HUMAN
|
||||||
| θ value | θ > 10 (rank : 79) | NC score | 0.078237 (rank : 83) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y3Q7, Q6UXJ9 | Gene names | ADAM18, TMDC3 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 18 precursor (A disintegrin and metalloproteinase domain 18) (Transmembrane metalloproteinase-like, disintegrin-like, and cysteine- rich protein III) (tMDC III). | |||||
|
ADA18_MOUSE
|
||||||
| θ value | θ > 10 (rank : 80) | NC score | 0.077512 (rank : 87) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9R157, Q60621 | Gene names | Adam18, Adam27, Dtgn3, Tmdc3 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 18 precursor (A disintegrin and metalloproteinase domain 18) (Transmembrane metalloproteinase-like, disintegrin-like, and cysteine- rich protein III) (tMDC III) (ADAM 27). | |||||
|
ADA19_HUMAN
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.073358 (rank : 101) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9H013, Q9BZL5, Q9UHP2 | Gene names | ADAM19, MLTNB | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta) (Metalloprotease and disintegrin dentritic antigen marker) (MADDAM). | |||||
|
ADA19_MOUSE
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.075720 (rank : 92) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O35674 | Gene names | Adam19, Mltnb | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta). | |||||
|
ADA20_HUMAN
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.074155 (rank : 96) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O43506, Q9UKJ9 | Gene names | ADAM20 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 20 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 20). | |||||
|
ADA21_HUMAN
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.071657 (rank : 111) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UKJ8, O43507 | Gene names | ADAM21 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 21 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 21). | |||||
|
ADA21_MOUSE
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.072013 (rank : 110) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9JI76 | Gene names | Adam21, Adam31 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 21 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 21) (ADAM 31). | |||||
|
ADA23_HUMAN
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.079495 (rank : 77) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O75077 | Gene names | ADAM23, MDC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAM 23 precursor (A disintegrin and metalloproteinase domain 23) (Metalloproteinase-like, disintegrin-like, and cysteine-rich protein 3) (MDC-3). | |||||
|
ADA23_MOUSE
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.079514 (rank : 76) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9R1V7, Q6QDY9, Q6QDZ0, Q8CC33 | Gene names | Adam23, Mdc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAM 23 precursor (A disintegrin and metalloproteinase domain 23) (Metalloproteinase-like, disintegrin-like, and cysteine-rich protein 3) (MDC-3). | |||||
|
ADA25_MOUSE
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.073222 (rank : 103) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9R159, Q9D4E4 | Gene names | Adam25 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 25 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 25) (Testase 2). | |||||
|
ADA30_HUMAN
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.073206 (rank : 104) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UKF2, Q9UKF1 | Gene names | ADAM30 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 30 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 30). | |||||
|
ADA32_HUMAN
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.076214 (rank : 90) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8TC27, Q8TC42 | Gene names | ADAM32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAM 32 precursor (A disintegrin and metalloproteinase domain 32). | |||||
|
ADA32_MOUSE
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.071231 (rank : 114) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8K410, Q6P901, Q8BJ80 | Gene names | Adam32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAM 32 precursor (A disintegrin and metalloproteinase domain 32). | |||||
|
ADA33_HUMAN
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.074216 (rank : 95) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9BZ11, Q5JT75, Q5JT76, Q8N0W6 | Gene names | ADAM33 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 33 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 33). | |||||
|
ADAM2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.072133 (rank : 109) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q99965, P78326, Q9UQQ8 | Gene names | ADAM2, FTNB | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 2 precursor (A disintegrin and metalloproteinase domain 2) (Fertilin subunit beta) (PH-30) (PH30). | |||||
|
ADAM2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.073746 (rank : 100) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q60718, Q60814, Q9D4G3, Q9QWJ0 | Gene names | Adam2, Ftnb | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 2 precursor (A disintegrin and metalloproteinase domain 2) (Fertilin subunit beta) (PH-30) (PH30) (PH30-beta). | |||||
|
ADAM7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.075578 (rank : 93) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O35227 | Gene names | Adam7 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 7 precursor (A disintegrin and metalloproteinase domain 7). | |||||
|
ADAM8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.076528 (rank : 89) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P78325 | Gene names | ADAM8, MS2 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 8 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 8) (Cell surface antigen MS2) (CD156a antigen) (CD156). | |||||
|
ADEC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.069563 (rank : 119) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O15204 | Gene names | ADAMDEC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAM DEC1 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain-like protein decysin 1) (ADAM-like protein decysin 1). | |||||
|
ADEC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.070929 (rank : 116) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9R0X2 | Gene names | Adamdec1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAM DEC1 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain-like protein decysin 1) (ADAM-like protein decysin 1). | |||||
|
ADM1A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.072956 (rank : 106) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q60813, Q60617, Q80WR7, Q8R533 | Gene names | Adam1a, Adam1, Ftna | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 1a precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 1a) (Fertilin alpha 1a subunit). | |||||
|
AGRIN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.060308 (rank : 137) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 582 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O00468, Q5SVA2, Q60FE1, Q7KYS8, Q8N4J5, Q96IC1, Q9BTD4 | Gene names | AGRIN, AGRN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Agrin precursor. | |||||
|
ATRN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.065160 (rank : 129) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 525 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O75882, O60295, O95414, Q3MIT3, Q5TDA2, Q5TDA4, Q5VYW3, Q9NTQ3, Q9NTQ4, Q9NU01, Q9NZ57, Q9NZ58 | Gene names | ATRN, KIAA0548, MGCA | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany homolog) (DPPT-L). | |||||
|
ATRN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.067407 (rank : 125) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9WU60, Q9R263, Q9WU77 | Gene names | Atrn, mg, Mgca | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany protein). | |||||
|
CREL1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.062495 (rank : 133) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q96HD1, Q6I9X5, Q8NFT4, Q9Y409 | Gene names | CRELD1, CIRRIN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich with EGF-like domain protein 1 precursor. | |||||
|
ITB2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.139340 (rank : 31) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P05107, Q16418 | Gene names | ITGB2, CD18 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-2 precursor (Cell surface adhesion glycoproteins LFA- 1/CR3/p150,95 subunit beta) (Complement receptor C3 subunit beta) (CD18 antigen). | |||||
|
ITB2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.139794 (rank : 30) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P11835, Q64482 | Gene names | Itgb2 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-2 precursor (Cell surface adhesion glycoproteins LFA- 1/CR3/P150,95 subunit beta) (Complement receptor C3 subunit beta) (CD18 antigen). | |||||
|
ITB4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.106666 (rank : 42) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P16144, O14690, O14691, O15339, O15340, O15341, Q9UIQ4 | Gene names | ITGB4 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-4 precursor (GP150) (CD104 antigen). | |||||
|
JAG1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.051923 (rank : 150) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 514 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P78504, O14902, O15122, Q15816 | Gene names | JAG1, JAGL1 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-1 precursor (Jagged1) (hJ1) (CD339 antigen). | |||||
|
JAG1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.052519 (rank : 148) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9QXX0 | Gene names | Jag1 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-1 precursor (Jagged1) (CD339 antigen). | |||||
|
JAG2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.052217 (rank : 149) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9Y219, Q9UE17, Q9UE99, Q9UNK8, Q9Y6P9, Q9Y6Q0 | Gene names | JAG2 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-2 precursor (Jagged2) (HJ2). | |||||
|
KR104_HUMAN
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.055702 (rank : 143) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P60372 | Gene names | KRTAP10-4, KAP10.4, KAP18-4, KRTAP10.4, KRTAP18-4, KRTAP18.4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 10-4 (Keratin-associated protein 10.4) (High sulfur keratin-associated protein 10.4) (Keratin-associated protein 18-4) (Keratin-associated protein 18.4). | |||||
|
KR106_HUMAN
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.050067 (rank : 157) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P60371 | Gene names | KRTAP10-6, KAP10.6, KAP18-6, KRTAP10.6, KRTAP18-6, KRTAP18.6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 10-6 (Keratin-associated protein 10.6) (High sulfur keratin-associated protein 10.6) (Keratin-associated protein 18-6) (Keratin-associated protein 18.6). | |||||
|
KR171_HUMAN
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.058472 (rank : 139) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9BYP8 | Gene names | KRTAP17-1, KAP17.1, KRTAP16.1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 17-1 (Keratin-associated protein 16.1). | |||||
|
KR510_HUMAN
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.107589 (rank : 41) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6L8G5 | Gene names | KRTAP5-10, KAP5.10, KRTAP5.10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-10 (Keratin-associated protein 5.10) (Ultrahigh sulfur keratin-associated protein 5.10). | |||||
|
KR511_HUMAN
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.073905 (rank : 97) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q6L8G4 | Gene names | KRTAP5-11, KAP5.11, KRTAP5.11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-11 (Keratin-associated protein 5.11) (Ultrahigh sulfur keratin-associated protein 5.11). | |||||
|
KRA51_HUMAN
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.103113 (rank : 46) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q6L8H4 | Gene names | KRTAP5-1, KAP5.1, KRTAP5.1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-1 (Keratin-associated protein 5.1) (Ultrahigh sulfur keratin-associated protein 5.1). | |||||
|
KRA52_HUMAN
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.088969 (rank : 64) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q701N4 | Gene names | KRTAP5-2, KAP5-8, KAP5.2, KRTAP5-8, KRTAP5.2, KRTAP5.8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-2 (Keratin-associated protein 5.2) (Ultrahigh sulfur keratin-associated protein 5.2) (Keratin-associated protein 5-8) (Keratin-associated protein 5.8). | |||||
|
KRA53_HUMAN
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.098157 (rank : 51) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q6L8H2, Q6PL44, Q701N3 | Gene names | KRTAP5-3, KAP5-9, KAP5.3, KRTAP5-9, KRTAP5.3, KRTAP5.9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-3 (Keratin-associated protein 5.3) (Ultrahigh sulfur keratin-associated protein 5.3) (Keratin-associated protein 5-9) (Keratin-associated protein 5.9) (UHS KerB-like). | |||||
|
KRA54_HUMAN
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.096888 (rank : 53) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q6L8H1 | Gene names | KRTAP5-4, KAP5.4, KRTAP5.4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-4 (Keratin-associated protein 5.4) (Ultrahigh sulfur keratin-associated protein 5.4). | |||||
|
KRA55_HUMAN
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.090507 (rank : 62) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q701N2 | Gene names | KRTAP5-5, KAP5-11, KAP5.5, KRTAP5-11, KRTAP5.11, KRTAP5.5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-5 (Keratin-associated protein 5.5) (Ultrahigh sulfur keratin-associated protein 5.5) (Keratin-associated protein 5-11) (Keratin-associated protein 5.11). | |||||
|
KRA56_HUMAN
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.095808 (rank : 54) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6L8G9 | Gene names | KRTAP5-6, KAP5.6, KRTAP5.6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-6 (Keratin-associated protein 5.6) (Ultrahigh sulfur keratin-associated protein 5.6). | |||||
|
KRA57_HUMAN
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.091665 (rank : 61) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q6L8G8, Q701N5 | Gene names | KRTAP5-7, KAP5-7, KAP5.3, KRTAP5.3, KRTAP5.7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-7 (Keratin-associated protein 5.7) (Ultrahigh sulfur keratin-associated protein 5.7) (Keratin-associated protein 5-3) (Keratin-associated protein 5.3). | |||||
|
KRA58_HUMAN
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.094671 (rank : 58) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O75690, Q6L8G7, Q6UTX6 | Gene names | KRTAP5-8, KAP5.8, KRTAP5.8, UHSKB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-8 (Keratin-associated protein 5.8) (Ultrahigh sulfur keratin-associated protein 5.8) (Keratin, ultra high-sulfur matrix protein B) (UHS keratin B) (UHS KerB). | |||||
|
KRA59_HUMAN
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.101741 (rank : 48) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P26371, Q14564 | Gene names | KRTAP5-9, KAP5.9, KRN1, KRTAP5.9, UHSK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-9 (Keratin-associated protein 5.9) (Ultrahigh sulfur keratin-associated protein 5.9) (Keratin, cuticle, ultrahigh sulfur 1) (Keratin, ultra high-sulfur matrix protein A) (UHS keratin A) (UHS KerA). | |||||
|
LAMA1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.085903 (rank : 67) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 838 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P19137 | Gene names | Lama1, Lama, Lama-1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-1 chain precursor (Laminin A chain). | |||||
|
LAMA2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.082896 (rank : 71) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1092 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P24043, Q14736, Q93022 | Gene names | LAMA2, LAMM | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
LAMA3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.062041 (rank : 134) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q16787, Q13679, Q13680, Q96TG0 | Gene names | LAMA3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-3 chain precursor (Epiligrin 170 kDa subunit) (E170) (Nicein subunit alpha). | |||||
|
LAMA3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.067592 (rank : 123) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q61789, O08751, Q61788, Q61966, Q9JHQ7 | Gene names | Lama3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-3 chain precursor (Nicein subunit alpha). | |||||
|
LAMA4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.056178 (rank : 141) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P97927, O88785, P70409 | Gene names | Lama4 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-4 chain precursor. | |||||
|
LAMA5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.087282 (rank : 66) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 976 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O15230, Q8WZA7, Q9H1P1 | Gene names | LAMA5, KIAA0533, KIAA1907 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-5 chain precursor. | |||||
|
LAMB1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.067283 (rank : 126) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P07942 | Gene names | LAMB1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
LAMB1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.068930 (rank : 121) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P02469 | Gene names | Lamb1-1, Lamb-1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
LAMB3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.065221 (rank : 128) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q13751, O14947, Q14733, Q9UJK4, Q9UJL1 | Gene names | LAMB3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-3 chain precursor (Laminin 5 beta 3) (Laminin B1k chain) (Kalinin B1 chain). | |||||
|
LAMB3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.064523 (rank : 130) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 444 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q61087 | Gene names | Lamb3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-3 chain precursor (Laminin 5 beta 3) (Kalinin B1 chain). | |||||
|
LAMC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.067577 (rank : 124) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P11047 | Gene names | LAMC1, LAMB2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-1 chain precursor (Laminin B2 chain). | |||||
|
LAMC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.070991 (rank : 115) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P02468 | Gene names | Lamc1, Lamb-2, Lamc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-1 chain precursor (Laminin B2 chain). | |||||
|
LAMC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.063524 (rank : 131) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 715 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q13753, Q02536, Q02537, Q13752, Q14941, Q5VYE8 | Gene names | LAMC2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-2 chain precursor (Laminin 5 gamma 2 subunit) (Kalinin/nicein/epiligrin 100 kDa subunit) (Laminin B2t chain) (Cell- scattering factor 140 kDa subunit) (CSF 140 kDa subunit) (Large adhesive scatter factor 140 kDa subunit) (Ladsin 140 kDa subunit). | |||||
|
LAMC2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.067838 (rank : 122) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q61092 | Gene names | Lamc2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-2 chain precursor (Laminin 5 gamma 2 subunit) (Kalinin/nicein/epiligrin 100 kDa subunit) (Laminin B2t chain). | |||||
|
LAMC3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.069252 (rank : 120) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 588 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9Y6N6 | Gene names | LAMC3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-3 chain precursor (Laminin 12 gamma 3 subunit). | |||||
|
LAMC3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.065372 (rank : 127) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9R0B6, Q9WTW6 | Gene names | Lamc3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-3 chain precursor (Laminin 12 gamma 3 subunit). | |||||
|
MEGF8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.053623 (rank : 146) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q7Z7M0, O75097 | Gene names | MEGF8, EGFL4, KIAA0817 | |||
|
Domain Architecture |
|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
MEGF8_MOUSE
|
||||||
| θ value | θ > 10 (rank : 141) | NC score | 0.050525 (rank : 156) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 520 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P60882, Q80TR3, Q80V41, Q8BMN9, Q8JZW7, Q8K0J3 | Gene names | Megf8, Egfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
MEGF9_HUMAN
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.078739 (rank : 80) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9H1U4, O75098 | Gene names | MEGF9, EGFL5, KIAA0818 | |||
|
Domain Architecture |
|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
MEGF9_MOUSE
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.075053 (rank : 94) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8BH27, Q8BWI4 | Gene names | Megf9, Egfl5, Kiaa0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
NAGPA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.072530 (rank : 107) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UK23, Q96EJ8 | Gene names | NAGPA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase precursor (EC 3.1.4.45) (Phosphodiester alpha-GlcNAcase) (Mannose 6- phosphate-uncovering enzyme). | |||||
|
NAGPA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.081318 (rank : 73) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8BJ48, Q3UUT5, Q8CHQ8, Q9QZE6 | Gene names | Nagpa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase precursor (EC 3.1.4.45) (Phosphodiester alpha-GlcNAcase) (Mannose 6- phosphate-uncovering enzyme). | |||||
|
NET4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 146) | NC score | 0.050746 (rank : 155) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9HB63, Q658K9, Q7L3F1, Q7L9D6, Q7Z5B6, Q9BZP1, Q9NT44, Q9P133 | Gene names | NTN4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin-4 precursor (Beta-netrin) (Hepar-derived netrin-like protein). | |||||
|
NET4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.051004 (rank : 153) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9JI33 | Gene names | Ntn4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin-4 precursor (Beta-netrin). | |||||
|
NTNG2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.050883 (rank : 154) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q96CW9, Q6UXY0, Q96JH0 | Gene names | NTNG2, KIAA1857, LMNT2 | |||
|
Domain Architecture |

|
|||||
| Description | Netrin G2 precursor (Laminet-2). | |||||
|
PCSK6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.052732 (rank : 147) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P29122, Q15099, Q15100, Q9UEG7, Q9UEJ1, Q9UEJ2, Q9UEJ7, Q9UEJ8, Q9UEJ9, Q9Y4G9, Q9Y4H0, Q9Y4H1 | Gene names | PCSK6, PACE4 | |||
|
Domain Architecture |

|
|||||
| Description | Proprotein convertase subtilisin/kexin type 6 precursor (EC 3.4.21.-) (Paired basic amino acid cleaving enzyme 4) (Subtilisin/kexin-like protease PACE4) (Subtilisin-like proprotein convertase 4) (SPC4). | |||||
|
SIVA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.095534 (rank : 55) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O15304, Q96P98, Q9UPD6 | Gene names | SIVA | |||
|
Domain Architecture |
|
|||||
| Description | Apoptosis regulatory protein Siva (CD27-binding protein) (CD27BP). | |||||
|
SREC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.089508 (rank : 63) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q14162, O43701 | Gene names | SCARF1, KIAA0149, SREC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelial cells scavenger receptor precursor (Acetyl LDL receptor) (Scavenger receptor class F member 1). | |||||
|
STAB1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.051850 (rank : 151) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 566 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8R4Y4, Q8K0K6, Q8VC09 | Gene names | Stab1, Feel1 | |||
|
Domain Architecture |

|
|||||
| Description | Stabilin-1 precursor (FEEL-1 protein). | |||||
|
STAB2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.051171 (rank : 152) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8WWQ8, Q6ZMK2, Q7Z5N9, Q86UR4, Q8IUG9, Q8TES1, Q9H7H7, Q9NRY3 | Gene names | STAB2, FEEL2, FELL, FEX2, HARE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stabilin-2 precursor (FEEL-2 protein) (Fasciclin EGF-like laminin-type EGF-like and link domain-containing scavenger receptor 1) (FAS1 EGF- like and X-link domain-containing adhesion molecule 2) (Hyaluronan receptor for endocytosis) [Contains: 190 kDa form stabilin-2 (190 kDa hyaluronan receptor for endocytosis)]. | |||||
|
STAB2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.053681 (rank : 145) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8R4U0, Q8BM87 | Gene names | Stab2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stabilin-2 precursor [Contains: Short form stabilin-2]. | |||||
|
TENA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.059192 (rank : 138) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 631 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P24821, Q14583, Q15567 | Gene names | TNC, HXB | |||
|
Domain Architecture |

|
|||||
| Description | Tenascin precursor (TN) (Tenascin-C) (TN-C) (Hexabrachion) (Cytotactin) (Neuronectin) (GMEM) (JI) (Myotendinous antigen) (Glioma- associated-extracellular matrix antigen) (GP 150-225). | |||||
|
TENA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.061841 (rank : 135) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q80YX1, Q64706, Q80YX2 | Gene names | Tnc, Hxb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tenascin precursor (TN) (Tenascin-C) (TN-C) (Hexabrachion). | |||||
|
TENX_HUMAN
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.072229 (rank : 108) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 789 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P22105, P78530, P78531, Q08424, Q9UMG7 | Gene names | TNXB, HXBL, TNX, XB | |||
|
Domain Architecture |

|
|||||
| Description | Tenascin-X precursor (TN-X) (Hexabrachion-like protein). | |||||
|
TNR9_MOUSE
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.095047 (rank : 57) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P20334 | Gene names | Tnfrsf9, Cd137, Cd157, Ila, Ly63 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 9 precursor (4-1BB ligand receptor) (T-cell antigen 4-1BB) (CD137 antigen). | |||||
|
WIF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.060833 (rank : 136) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9Y5W5, Q6UXI1, Q8WVG4 | Gene names | WIF1 | |||
|
Domain Architecture |
|
|||||
| Description | Wnt inhibitory factor 1 precursor (WIF-1). | |||||
|
WIF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.063054 (rank : 132) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9WUA1 | Gene names | Wif1 | |||
|
Domain Architecture |
|
|||||
| Description | Wnt inhibitory factor 1 precursor (WIF-1). | |||||
|
MT2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 7.09661e-13 (rank : 1) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P02795, Q14823, Q2HXR9, Q53XT9 | Gene names | MT2A, CES1, MT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-2 (MT-2) (Metallothionein-II) (MT-II) (Metallothionein-2A). | |||||
|
MT1I_HUMAN
|
||||||
| NC score | 0.986700 (rank : 2) | θ value | 2.28291e-11 (rank : 2) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | P80295 | Gene names | MT1I | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1I (MT-1I) (Metallothionein-II). | |||||
|
MT1F_HUMAN
|
||||||
| NC score | 0.984021 (rank : 3) | θ value | 8.67504e-11 (rank : 4) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P04733, Q9UI97 | Gene names | MT1F | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1F (MT-1F) (Metallothionein-IF) (MT-IF). | |||||
|
MT1E_HUMAN
|
||||||
| NC score | 0.981287 (rank : 4) | θ value | 5.08577e-11 (rank : 3) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P04732, Q86YX4 | Gene names | MT1E | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1E (MT-1E) (Metallothionein-IE) (MT-IE). | |||||
|
MT1M_HUMAN
|
||||||
| NC score | 0.967633 (rank : 5) | θ value | 8.67504e-11 (rank : 5) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8N339, Q8TDN3 | Gene names | MT1M | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1M (MT-1M) (Metallothionein-IM) (MT-IM). | |||||
|
MT1X_HUMAN
|
||||||
| NC score | 0.941448 (rank : 6) | θ value | 4.76016e-09 (rank : 13) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P80297 | Gene names | MT1X | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1X (MT-1X) (Metallothionein-IX) (MT-IX). | |||||
|
MT1G_HUMAN
|
||||||
| NC score | 0.928633 (rank : 7) | θ value | 2.79066e-09 (rank : 9) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P13640, P80296 | Gene names | MT1G, MT1K, MT1M | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1G (MT-1G) (Metallothionein-IG) (MT-IG) (Metallothionein-1K) (MT-1K). | |||||
|
MT1L_HUMAN
|
||||||
| NC score | 0.917024 (rank : 8) | θ value | 7.34386e-10 (rank : 8) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q93083 | Gene names | MT1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1L (MT-1L) (Metallothionein-IL) (MT-IL). | |||||
|
MT1H_HUMAN
|
||||||
| NC score | 0.902145 (rank : 9) | θ value | 1.9326e-10 (rank : 7) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | P80294 | Gene names | MT1H | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1H (MT-1H) (Metallothionein-IH) (MT-IH) (Metallothionein-0) (MT-0). | |||||
|
MT1A_HUMAN
|
||||||
| NC score | 0.893290 (rank : 10) | θ value | 1.133e-10 (rank : 6) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P04731 | Gene names | MT1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1A (MT-1A) (Metallothionein-IA) (MT-IA). | |||||
|
MT1_MOUSE
|
||||||
| NC score | 0.882860 (rank : 11) | θ value | 2.79066e-09 (rank : 10) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P02802, Q64485 | Gene names | Mt1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1 (MT-1) (Metallothionein-I) (MT-I). | |||||
|
MT1B_HUMAN
|
||||||
| NC score | 0.866267 (rank : 12) | θ value | 4.76016e-09 (rank : 12) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | P07438 | Gene names | MT1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1B (MT-1B) (Metallothionein-IB) (MT-IB). | |||||
|
MT2_MOUSE
|
||||||
| NC score | 0.846436 (rank : 13) | θ value | 3.64472e-09 (rank : 11) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P02798 | Gene names | Mt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-2 (MT-2) (Metallothionein-II) (MT-II). | |||||
|
MT4_HUMAN
|
||||||
| NC score | 0.758874 (rank : 14) | θ value | 0.000270298 (rank : 16) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P47944 | Gene names | MT4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-4 (MT-4) (Metallothionein-IV) (MT-IV). | |||||
|
MT4_MOUSE
|
||||||
| NC score | 0.754040 (rank : 15) | θ value | 0.000121331 (rank : 15) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P47945 | Gene names | Mt4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-4 (MT-4) (Metallothionein-IV) (MT-IV). | |||||
|
CT127_HUMAN
|
||||||
| NC score | 0.734564 (rank : 16) | θ value | 3.41135e-07 (rank : 14) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9BQN2 | Gene names | C20orf127 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative metallothionein C20orf127. | |||||
|
MT3_HUMAN
|
||||||
| NC score | 0.564255 (rank : 17) | θ value | 0.0148317 (rank : 17) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P25713, Q2V574 | Gene names | MT3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-3 (MT-3) (Metallothionein-III) (MT-III) (Growth inhibitory factor) (GIF) (GIFB). | |||||
|
MT3_MOUSE
|
||||||
| NC score | 0.472518 (rank : 18) | θ value | 0.0563607 (rank : 18) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P28184 | Gene names | Mt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-3 (MT-3) (Metallothionein-III) (MT-III) (Growth inhibitory factor) (GIF). | |||||
|
ITB5_MOUSE
|
||||||
| NC score | 0.196900 (rank : 19) | θ value | 0.365318 (rank : 23) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O70309, O70308, O88347 | Gene names | Itgb5 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-5 precursor. | |||||
|
ITB1_HUMAN
|
||||||
| NC score | 0.191509 (rank : 20) | θ value | 0.47712 (rank : 26) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P05556, P78466, P78467, Q13089, Q13090, Q13091, Q13212, Q14622, Q14647 | Gene names | ITGB1, FNRB | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-1 precursor (Fibronectin receptor subunit beta) (Integrin VLA-4 subunit beta) (CD29 antigen). | |||||
|
ITB5_HUMAN
|
||||||
| NC score | 0.184611 (rank : 21) | θ value | 0.813845 (rank : 29) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P18084 | Gene names | ITGB5 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-5 precursor. | |||||
|
ITB3_MOUSE
|
||||||
| NC score | 0.177114 (rank : 22) | θ value | 2.36792 (rank : 43) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O54890 | Gene names | Itgb3 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-3 precursor (Platelet membrane glycoprotein IIIa) (GPIIIa) (CD61 antigen). | |||||
|
ITB1_MOUSE
|
||||||
| NC score | 0.176058 (rank : 23) | θ value | 0.62314 (rank : 27) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P09055 | Gene names | Itgb1 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-1 precursor (Fibronectin receptor subunit beta) (Integrin VLA-4 subunit beta) (CD29 antigen). | |||||
|
ITB3_HUMAN
|
||||||
| NC score | 0.175411 (rank : 24) | θ value | 0.62314 (rank : 28) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P05106, O15495, Q13413, Q14648, Q16499 | Gene names | ITGB3, GP3A | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-3 precursor (Platelet membrane glycoprotein IIIa) (GPIIIa) (CD61 antigen). | |||||
|
ITB8_HUMAN
|
||||||
| NC score | 0.166499 (rank : 25) | θ value | 1.06291 (rank : 31) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P26012 | Gene names | ITGB8 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-8 precursor. | |||||
|
ITB6_HUMAN
|
||||||
| NC score | 0.165321 (rank : 26) | θ value | 1.81305 (rank : 38) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P18564, Q16500 | Gene names | ITGB6 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-6 precursor. | |||||
|
ITB7_MOUSE
|
||||||
| NC score | 0.163651 (rank : 27) | θ value | 4.03905 (rank : 52) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P26011, Q3U2M1, Q64656 | Gene names | Itgb7 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-7 precursor (Integrin beta-P) (M290 IEL antigen). | |||||
|
ITB7_HUMAN
|
||||||
| NC score | 0.161665 (rank : 28) | θ value | 6.88961 (rank : 59) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P26010 | Gene names | ITGB7 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-7 precursor. | |||||
|
ITB6_MOUSE
|
||||||
| NC score | 0.153381 (rank : 29) | θ value | 8.99809 (rank : 65) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Z0T9 | Gene names | Itgb6 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-6 precursor. | |||||
|
ITB2_MOUSE
|
||||||
| NC score | 0.139794 (rank : 30) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P11835, Q64482 | Gene names | Itgb2 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-2 precursor (Cell surface adhesion glycoproteins LFA- 1/CR3/P150,95 subunit beta) (Complement receptor C3 subunit beta) (CD18 antigen). | |||||
|
ITB2_HUMAN
|
||||||
| NC score | 0.139340 (rank : 31) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P05107, Q16418 | Gene names | ITGB2, CD18 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-2 precursor (Cell surface adhesion glycoproteins LFA- 1/CR3/p150,95 subunit beta) (Complement receptor C3 subunit beta) (CD18 antigen). | |||||
|
PCSK5_MOUSE
|
||||||
| NC score | 0.138728 (rank : 32) | θ value | 0.125558 (rank : 19) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 613 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q04592, Q62040 | Gene names | Pcsk5 | |||
|
Domain Architecture |

|
|||||
| Description | Proprotein convertase subtilisin/kexin type 5 precursor (EC 3.4.21.-) (Proprotein convertase PC5) (Subtilisin/kexin-like protease PC5) (PC6) (Subtilisin-like proprotein convertase 6) (SPC6). | |||||
|
TNR9_HUMAN
|
||||||
| NC score | 0.132551 (rank : 33) | θ value | 0.813845 (rank : 30) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q07011 | Gene names | TNFRSF9, CD137, ILA | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 9 precursor (4-1BB ligand receptor) (T-cell antigen 4-1BB homolog) (T-cell antigen ILA) (CD137 antigen). | |||||
|
ADA22_MOUSE
|
||||||
| NC score | 0.129936 (rank : 34) | θ value | 3.0926 (rank : 45) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9R1V6, Q5TLI8, Q5TLI9, Q5TLJ0, Q5TLJ1, Q5TLJ2, Q5TLJ3, Q5TLJ4, Q5TLJ5, Q5TLJ6, Q5TLJ7, Q5TLJ8, Q5TLJ9, Q5TLK0, Q5TLK1, Q5TLK2, Q5TLK3, Q5TLK4, Q5TLK5, Q5TLK6, Q5TLK7, Q5TLK8, Q8BSF2, Q9R1V5 | Gene names | Adam22 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 22 precursor (A disintegrin and metalloproteinase domain 22). | |||||
|
ADA22_HUMAN
|
||||||
| NC score | 0.123373 (rank : 35) | θ value | 1.81305 (rank : 36) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9P0K1, O75075, O75076, Q9P0K2, Q9UIA1, Q9UKK2 | Gene names | ADAM22, MDC2 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 22 precursor (A disintegrin and metalloproteinase domain 22) (Metalloproteinase-like, disintegrin-like, and cysteine-rich protein 2) (Metalloproteinase-disintegrin ADAM22-3). | |||||
|
CRIM1_HUMAN
|
||||||
| NC score | 0.121723 (rank : 36) | θ value | 1.38821 (rank : 32) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9NZV1, Q59GH0, Q7LCQ5, Q9H318 | Gene names | CRIM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich motor neuron 1 protein precursor (CRIM-1) (Cysteine-rich repeat-containing protein S52). | |||||
|
ADA28_HUMAN
|
||||||
| NC score | 0.112133 (rank : 37) | θ value | 0.365318 (rank : 22) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9UKQ2, Q9Y339, Q9Y3S0 | Gene names | ADAM28, MDCL | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 28 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 28) (Metalloproteinase-like, disintegrin-like, and cysteine- rich protein-L) (MDC-L) (eMDC II) (ADAM23). | |||||
|
LAMA5_MOUSE
|
||||||
| NC score | 0.110482 (rank : 38) | θ value | 0.279714 (rank : 21) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q61001, Q9JHQ6 | Gene names | Lama5 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-5 chain precursor. | |||||
|
ADA28_MOUSE
|
||||||
| NC score | 0.109924 (rank : 39) | θ value | 0.47712 (rank : 25) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9JLN6, Q8K5D2, Q8K5D3 | Gene names | Adam28 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 28 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 28) (Thymic epithelial cell-ADAM) (TECADAM). | |||||
|
TNR7_MOUSE
|
||||||
| NC score | 0.109078 (rank : 40) | θ value | 3.0926 (rank : 49) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P41272 | Gene names | Tnfrsf7, Cd27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor necrosis factor receptor superfamily member 7 precursor (CD27L receptor) (T-cell activation antigen CD27). | |||||
|
KR510_HUMAN
|
||||||
| NC score | 0.107589 (rank : 41) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6L8G5 | Gene names | KRTAP5-10, KAP5.10, KRTAP5.10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-10 (Keratin-associated protein 5.10) (Ultrahigh sulfur keratin-associated protein 5.10). | |||||
|
ITB4_HUMAN
|
||||||
| NC score | 0.106666 (rank : 42) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P16144, O14690, O14691, O15339, O15340, O15341, Q9UIQ4 | Gene names | ITGB4 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-4 precursor (GP150) (CD104 antigen). | |||||
|
ADAM9_MOUSE
|
||||||
| NC score | 0.105464 (rank : 43) | θ value | 1.81305 (rank : 37) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q61072, Q60618, Q61853, Q80U94 | Gene names | Adam9, Kiaa0021, Mdc9, Mltng | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAM 9 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 9) (Metalloprotease/disintegrin/cysteine-rich protein 9) (Myeloma cell metalloproteinase) (Meltrin gamma). | |||||
|
ADA11_HUMAN
|
||||||
| NC score | 0.104859 (rank : 44) | θ value | 1.81305 (rank : 34) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O75078, Q14808, Q14809, Q14810 | Gene names | ADAM11, MDC | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 11 precursor (A disintegrin and metalloproteinase domain 11) (Metalloproteinase-like, disintegrin-like, and cysteine-rich protein) (MDC). | |||||
|
ADA11_MOUSE
|
||||||
| NC score | 0.104136 (rank : 45) | θ value | 1.81305 (rank : 35) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9R1V4 | Gene names | Adam11, Mdc | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 11 precursor (A disintegrin and metalloproteinase domain 11) (Metalloproteinase-like, disintegrin-like, and cysteine-rich protein) (MDC). | |||||
|
KRA51_HUMAN
|
||||||
| NC score | 0.103113 (rank : 46) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q6L8H4 | Gene names | KRTAP5-1, KAP5.1, KRTAP5.1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-1 (Keratin-associated protein 5.1) (Ultrahigh sulfur keratin-associated protein 5.1). | |||||
|
TNR7_HUMAN
|
||||||
| NC score | 0.102999 (rank : 47) | θ value | 5.27518 (rank : 55) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P26842 | Gene names | TNFRSF7, CD27 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 7 precursor (CD27L receptor) (T-cell activation antigen CD27) (T14). | |||||
|
KRA59_HUMAN
|
||||||
| NC score | 0.101741 (rank : 48) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P26371, Q14564 | Gene names | KRTAP5-9, KAP5.9, KRN1, KRTAP5.9, UHSK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-9 (Keratin-associated protein 5.9) (Ultrahigh sulfur keratin-associated protein 5.9) (Keratin, cuticle, ultrahigh sulfur 1) (Keratin, ultra high-sulfur matrix protein A) (UHS keratin A) (UHS KerA). | |||||
|
USH2A_MOUSE
|
||||||
| NC score | 0.099100 (rank : 49) | θ value | 2.36792 (rank : 44) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 626 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q2QI47, Q9JLP3 | Gene names | Ush2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Usherin precursor (Usher syndrome type-2A protein homolog) (Usher syndrome type IIa protein homolog). | |||||
|
ADAM9_HUMAN
|
||||||
| NC score | 0.098533 (rank : 50) | θ value | 3.0926 (rank : 46) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q13443, Q10718, Q8NFM6 | Gene names | ADAM9, KIAA0021, MCMP, MDC9, MLTNG | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 9 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 9) (Metalloprotease/disintegrin/cysteine-rich protein 9) (Myeloma cell metalloproteinase) (Meltrin gamma) (Cellular disintegrin-related protein). | |||||
|
KRA53_HUMAN
|
||||||
| NC score | 0.098157 (rank : 51) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q6L8H2, Q6PL44, Q701N3 | Gene names | KRTAP5-3, KAP5-9, KAP5.3, KRTAP5-9, KRTAP5.3, KRTAP5.9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-3 (Keratin-associated protein 5.3) (Ultrahigh sulfur keratin-associated protein 5.3) (Keratin-associated protein 5-9) (Keratin-associated protein 5.9) (UHS KerB-like). | |||||
|
MEGF6_HUMAN
|
||||||
| NC score | 0.097404 (rank : 52) | θ value | 1.38821 (rank : 33) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | O75095, Q5VV39 | Gene names | MEGF6, EGFL3, KIAA0815 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multiple epidermal growth factor-like domains 6 precursor (EGF-like domain-containing protein 3) (Multiple EGF-like domain protein 3). | |||||
|
KRA54_HUMAN
|
||||||
| NC score | 0.096888 (rank : 53) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q6L8H1 | Gene names | KRTAP5-4, KAP5.4, KRTAP5.4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-4 (Keratin-associated protein 5.4) (Ultrahigh sulfur keratin-associated protein 5.4). | |||||
|
KRA56_HUMAN
|
||||||
| NC score | 0.095808 (rank : 54) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6L8G9 | Gene names | KRTAP5-6, KAP5.6, KRTAP5.6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-6 (Keratin-associated protein 5.6) (Ultrahigh sulfur keratin-associated protein 5.6). | |||||
|
SIVA_HUMAN
|
||||||
| NC score | 0.095534 (rank : 55) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O15304, Q96P98, Q9UPD6 | Gene names | SIVA | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis regulatory protein Siva (CD27-binding protein) (CD27BP). | |||||
|
LAMA1_HUMAN
|
||||||
| NC score | 0.095162 (rank : 56) | θ value | 1.81305 (rank : 39) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 804 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P25391 | Gene names | LAMA1, LAMA | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-1 chain precursor (Laminin A chain). | |||||
|
TNR9_MOUSE
|
||||||
| NC score | 0.095047 (rank : 57) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P20334 | Gene names | Tnfrsf9, Cd137, Cd157, Ila, Ly63 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 9 precursor (4-1BB ligand receptor) (T-cell antigen 4-1BB) (CD137 antigen). | |||||
|
KRA58_HUMAN
|
||||||
| NC score | 0.094671 (rank : 58) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O75690, Q6L8G7, Q6UTX6 | Gene names | KRTAP5-8, KAP5.8, KRTAP5.8, UHSKB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-8 (Keratin-associated protein 5.8) (Ultrahigh sulfur keratin-associated protein 5.8) (Keratin, ultra high-sulfur matrix protein B) (UHS keratin B) (UHS KerB). | |||||
|
SREC2_MOUSE
|
||||||
| NC score | 0.094085 (rank : 59) | θ value | 8.99809 (rank : 68) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 605 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P59222 | Gene names | Scarf2, Srec2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor class F member 2 precursor (Scavenger receptor expressed by endothelial cells 2 protein) (SREC-II). | |||||
|
SREC2_HUMAN
|
||||||
| NC score | 0.091791 (rank : 60) | θ value | 8.99809 (rank : 67) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q96GP6, Q8IXF3, Q9BW74 | Gene names | SCARF2, SREC2, SREPCR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor class F member 2 precursor (Scavenger receptor expressed by endothelial cells 2 protein) (SREC-II) (SRECRP-1). | |||||
|
KRA57_HUMAN
|
||||||
| NC score | 0.091665 (rank : 61) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q6L8G8, Q701N5 | Gene names | KRTAP5-7, KAP5-7, KAP5.3, KRTAP5.3, KRTAP5.7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-7 (Keratin-associated protein 5.7) (Ultrahigh sulfur keratin-associated protein 5.7) (Keratin-associated protein 5-3) (Keratin-associated protein 5.3). | |||||
|
KRA55_HUMAN
|
||||||
| NC score | 0.090507 (rank : 62) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q701N2 | Gene names | KRTAP5-5, KAP5-11, KAP5.5, KRTAP5-11, KRTAP5.11, KRTAP5.5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-5 (Keratin-associated protein 5.5) (Ultrahigh sulfur keratin-associated protein 5.5) (Keratin-associated protein 5-11) (Keratin-associated protein 5.11). | |||||
|
SREC_HUMAN
|
||||||
| NC score | 0.089508 (rank : 63) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q14162, O43701 | Gene names | SCARF1, KIAA0149, SREC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelial cells scavenger receptor precursor (Acetyl LDL receptor) (Scavenger receptor class F member 1). | |||||
|
KRA52_HUMAN
|
||||||
| NC score | 0.088969 (rank : 64) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q701N4 | Gene names | KRTAP5-2, KAP5-8, KAP5.2, KRTAP5-8, KRTAP5.2, KRTAP5.8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-2 (Keratin-associated protein 5.2) (Ultrahigh sulfur keratin-associated protein 5.2) (Keratin-associated protein 5-8) (Keratin-associated protein 5.8). | |||||
|
LAMA2_MOUSE
|
||||||
| NC score | 0.087296 (rank : 65) | θ value | 1.81305 (rank : 40) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1198 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q60675, Q05003, Q64061 | Gene names | Lama2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
LAMA5_HUMAN
|
||||||
| NC score | 0.087282 (rank : 66) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 976 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O15230, Q8WZA7, Q9H1P1 | Gene names | LAMA5, KIAA0533, KIAA1907 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-5 chain precursor. | |||||
|
LAMA1_MOUSE
|
||||||
| NC score | 0.085903 (rank : 67) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 838 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P19137 | Gene names | Lama1, Lama, Lama-1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-1 chain precursor (Laminin A chain). | |||||
|
PCSK5_HUMAN
|
||||||
| NC score | 0.085827 (rank : 68) | θ value | 4.03905 (rank : 53) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q92824, Q13527 | Gene names | PCSK5, PC5, PC6 | |||
|
Domain Architecture |

|
|||||
| Description | Proprotein convertase subtilisin/kexin type 5 precursor (EC 3.4.21.-) (Proprotein convertase PC5) (Subtilisin/kexin-like protease PC5) (PC6) (hPC6). | |||||
|
FRAS1_MOUSE
|
||||||
| NC score | 0.084022 (rank : 69) | θ value | 3.0926 (rank : 47) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q80T14, Q80TC7, Q811H8, Q8BPZ4 | Gene names | Fras1, Kiaa1500 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Extracellular matrix protein FRAS1 precursor. | |||||
|
FRAS1_HUMAN
|
||||||
| NC score | 0.083529 (rank : 70) | θ value | 4.03905 (rank : 51) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 759 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q86XX4, Q86UZ4, Q8N3U9, Q8NAU7, Q96JW7, Q9H6N9, Q9P228 | Gene names | FRAS1, KIAA1500 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Extracellular matrix protein FRAS1 precursor. | |||||
|
LAMA2_HUMAN
|
||||||
| NC score | 0.082896 (rank : 71) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1092 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P24043, Q14736, Q93022 | Gene names | LAMA2, LAMM | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
ADA24_MOUSE
|
||||||
| NC score | 0.081369 (rank : 72) | θ value | 4.03905 (rank : 50) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9R160 | Gene names | Adam24 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 24 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 24) (Testase 1). | |||||
|
NAGPA_MOUSE
|
||||||
| NC score | 0.081318 (rank : 73) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8BJ48, Q3UUT5, Q8CHQ8, Q9QZE6 | Gene names | Nagpa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase precursor (EC 3.1.4.45) (Phosphodiester alpha-GlcNAcase) (Mannose 6- phosphate-uncovering enzyme). | |||||
|
ADAM8_MOUSE
|
||||||
| NC score | 0.080545 (rank : 74) | θ value | 6.88961 (rank : 57) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q05910 | Gene names | Adam8, Ms2 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 8 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 8) (Cell surface antigen MS2) (Macrophage cysteine-rich glycoprotein) (CD156 antigen). | |||||
|
ZAN_MOUSE
|
||||||
| NC score | 0.079561 (rank : 75) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | O88799, O08647 | Gene names | Zan | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
ADA23_MOUSE
|
||||||
| NC score | 0.079514 (rank : 76) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9R1V7, Q6QDY9, Q6QDZ0, Q8CC33 | Gene names | Adam23, Mdc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAM 23 precursor (A disintegrin and metalloproteinase domain 23) (Metalloproteinase-like, disintegrin-like, and cysteine-rich protein 3) (MDC-3). | |||||
|
ADA23_HUMAN
|
||||||
| NC score | 0.079495 (rank : 77) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O75077 | Gene names | ADAM23, MDC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAM 23 precursor (A disintegrin and metalloproteinase domain 23) (Metalloproteinase-like, disintegrin-like, and cysteine-rich protein 3) (MDC-3). | |||||
|
ADA29_HUMAN
|
||||||
| NC score | 0.079274 (rank : 78) | θ value | 2.36792 (rank : 42) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9UKF5, Q9UHP1, Q9UKF3, Q9UKF4 | Gene names | ADAM29 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 29 precursor (A disintegrin and metalloproteinase domain 29). | |||||
|
ADA15_HUMAN
|
||||||
| NC score | 0.079083 (rank : 79) | θ value | θ > 10 (rank : 75) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q13444, Q13493, Q96C78 | Gene names | ADAM15, MDC15 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 15 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 15) (Metalloproteinase-like, disintegrin-like, and cysteine- rich protein 15) (MDC-15) (Metalloprotease RGD disintegrin protein) (Metargidin). | |||||
|
MEGF9_HUMAN
|
||||||
| NC score | 0.078739 (rank : 80) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9H1U4, O75098 | Gene names | MEGF9, EGFL5, KIAA0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
ADA15_MOUSE
|
||||||
| NC score | 0.078674 (rank : 81) | θ value | θ > 10 (rank : 76) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O88839, Q3U7C2, Q8C7Z0, Q91VS9, Q9QYL2 | Gene names | Adam15, Mdc15 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 15 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 15) (Metalloproteinase-like, disintegrin-like, and cysteine- rich protein 15) (MDC-15) (Metalloprotease RGD disintegrin protein) (Metargidin) (AD56). | |||||
|
ADM1B_MOUSE
|
||||||
| NC score | 0.078543 (rank : 82) | θ value | 8.99809 (rank : 64) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8R534, Q9R156 | Gene names | Adam1b | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 1b precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 1b) (Fertilin alpha 1b subunit). | |||||
|
ADA18_HUMAN
|
||||||
| NC score | 0.078237 (rank : 83) | θ value | θ > 10 (rank : 79) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y3Q7, Q6UXJ9 | Gene names | ADAM18, TMDC3 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 18 precursor (A disintegrin and metalloproteinase domain 18) (Transmembrane metalloproteinase-like, disintegrin-like, and cysteine- rich protein III) (tMDC III). | |||||
|
USH2A_HUMAN
|
||||||
| NC score | 0.078008 (rank : 84) | θ value | 8.99809 (rank : 69) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 600 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O75445, Q5VVM9, Q6S362, Q9NS27 | Gene names | USH2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Usherin precursor (Usher syndrome type-2A protein) (Usher syndrome type IIa protein). | |||||
|
LAMB2_MOUSE
|
||||||
| NC score | 0.077951 (rank : 85) | θ value | 1.81305 (rank : 41) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q61292, Q62182 | Gene names | Lamb2, Lams | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (S-LAM). | |||||
|
ADAM7_HUMAN
|
||||||
| NC score | 0.077701 (rank : 86) | θ value | 6.88961 (rank : 56) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9H2U9, O75959 | Gene names | ADAM7, GP83 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 7 precursor (A disintegrin and metalloproteinase domain 7) (Sperm maturation-related glycoprotein GP-83). | |||||
|
ADA18_MOUSE
|
||||||
| NC score | 0.077512 (rank : 87) | θ value | θ > 10 (rank : 80) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9R157, Q60621 | Gene names | Adam18, Adam27, Dtgn3, Tmdc3 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 18 precursor (A disintegrin and metalloproteinase domain 18) (Transmembrane metalloproteinase-like, disintegrin-like, and cysteine- rich protein III) (tMDC III) (ADAM 27). | |||||
|
ADA33_MOUSE
|
||||||
| NC score | 0.077398 (rank : 88) | θ value | 8.99809 (rank : 63) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q923W9 | Gene names | Adam33 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 33 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 33). | |||||
|
ADAM8_HUMAN
|
||||||
| NC score | 0.076528 (rank : 89) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P78325 | Gene names | ADAM8, MS2 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 8 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 8) (Cell surface antigen MS2) (CD156a antigen) (CD156). | |||||
|
ADA32_HUMAN
|
||||||
| NC score | 0.076214 (rank : 90) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8TC27, Q8TC42 | Gene names | ADAM32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAM 32 precursor (A disintegrin and metalloproteinase domain 32). | |||||
|
CREL1_MOUSE
|
||||||
| NC score | 0.076199 (rank : 91) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 524 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q91XD7, Q8BGJ8 | Gene names | Creld1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich with EGF-like domain protein 1 precursor. | |||||
|
ADA19_MOUSE
|
||||||
| NC score | 0.075720 (rank : 92) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O35674 | Gene names | Adam19, Mltnb | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta). | |||||
|
ADAM7_MOUSE
|
||||||
| NC score | 0.075578 (rank : 93) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O35227 | Gene names | Adam7 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 7 precursor (A disintegrin and metalloproteinase domain 7). | |||||
|
MEGF9_MOUSE
|
||||||
| NC score | 0.075053 (rank : 94) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8BH27, Q8BWI4 | Gene names | Megf9, Egfl5, Kiaa0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
ADA33_HUMAN
|
||||||
| NC score | 0.074216 (rank : 95) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9BZ11, Q5JT75, Q5JT76, Q8N0W6 | Gene names | ADAM33 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 33 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 33). | |||||
|
ADA20_HUMAN
|
||||||
| NC score | 0.074155 (rank : 96) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O43506, Q9UKJ9 | Gene names | ADAM20 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 20 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 20). | |||||
|
KR511_HUMAN
|
||||||
| NC score | 0.073905 (rank : 97) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q6L8G4 | Gene names | KRTAP5-11, KAP5.11, KRTAP5.11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-11 (Keratin-associated protein 5.11) (Ultrahigh sulfur keratin-associated protein 5.11). | |||||
|
ADA17_HUMAN
|
||||||
| NC score | 0.073900 (rank : 98) | θ value | θ > 10 (rank : 77) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P78536, O60226 | Gene names | ADAM17, CSVP, TACE | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 17 precursor (EC 3.4.24.86) (A disintegrin and metalloproteinase domain 17) (TNF-alpha-converting enzyme) (TNF-alpha convertase) (Snake venom-like protease) (CD156b antigen). | |||||
|
AD26A_MOUSE
|
||||||
| NC score | 0.073801 (rank : 99) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9R158 | Gene names | Adam26a, Adam26 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 26A precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 26A) (Testase-3). | |||||
|
ADAM2_MOUSE
|
||||||
| NC score | 0.073746 (rank : 100) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q60718, Q60814, Q9D4G3, Q9QWJ0 | Gene names | Adam2, Ftnb | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 2 precursor (A disintegrin and metalloproteinase domain 2) (Fertilin subunit beta) (PH-30) (PH30) (PH30-beta). | |||||
|
ADA19_HUMAN
|
||||||
| NC score | 0.073358 (rank : 101) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9H013, Q9BZL5, Q9UHP2 | Gene names | ADAM19, MLTNB | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta) (Metalloprotease and disintegrin dentritic antigen marker) (MADDAM). | |||||
|
ADA17_MOUSE
|
||||||
| NC score | 0.073323 (rank : 102) | θ value | θ > 10 (rank : 78) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9Z0F8, O88726, Q9R1U4, Q9Z0K3 | Gene names | Adam17, Tace | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 17 precursor (EC 3.4.24.86) (A disintegrin and metalloproteinase domain 17) (TNF-alpha-converting enzyme) (TNF-alpha convertase) (CD156b antigen). | |||||
|
ADA25_MOUSE
|
||||||
| NC score | 0.073222 (rank : 103) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9R159, Q9D4E4 | Gene names | Adam25 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 25 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 25) (Testase 2). | |||||
|
ADA30_HUMAN
|
||||||
| NC score | 0.073206 (rank : 104) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UKF2, Q9UKF1 | Gene names | ADAM30 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 30 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 30). | |||||
|
LAMB2_HUMAN
|
||||||
| NC score | 0.073060 (rank : 105) | θ value | 8.99809 (rank : 66) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 692 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P55268, Q16321 | Gene names | LAMB2, LAMS | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (Laminin B1s chain). | |||||
|
ADM1A_MOUSE
|
||||||
| NC score | 0.072956 (rank : 106) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q60813, Q60617, Q80WR7, Q8R533 | Gene names | Adam1a, Adam1, Ftna | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 1a precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 1a) (Fertilin alpha 1a subunit). | |||||
|
NAGPA_HUMAN
|
||||||
| NC score | 0.072530 (rank : 107) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UK23, Q96EJ8 | Gene names | NAGPA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase precursor (EC 3.1.4.45) (Phosphodiester alpha-GlcNAcase) (Mannose 6- phosphate-uncovering enzyme). | |||||
|
TENX_HUMAN
|
||||||
| NC score | 0.072229 (rank : 108) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 789 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P22105, P78530, P78531, Q08424, Q9UMG7 | Gene names | TNXB, HXBL, TNX, XB | |||
|
Domain Architecture |

|
|||||
| Description | Tenascin-X precursor (TN-X) (Hexabrachion-like protein). | |||||
|
ADAM2_HUMAN
|
||||||
| NC score | 0.072133 (rank : 109) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q99965, P78326, Q9UQQ8 | Gene names | ADAM2, FTNB | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 2 precursor (A disintegrin and metalloproteinase domain 2) (Fertilin subunit beta) (PH-30) (PH30). | |||||
|
ADA21_MOUSE
|
||||||
| NC score | 0.072013 (rank : 110) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9JI76 | Gene names | Adam21, Adam31 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 21 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 21) (ADAM 31). | |||||
|
ADA21_HUMAN
|
||||||
| NC score | 0.071657 (rank : 111) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UKJ8, O43507 | Gene names | ADAM21 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 21 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 21). | |||||
|
ADA12_MOUSE
|
||||||
| NC score | 0.071473 (rank : 112) | θ value | θ > 10 (rank : 74) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q61824 | Gene names | Adam12, Mltna | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 12 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 12) (Meltrin alpha). | |||||
|
ADA12_HUMAN
|
||||||
| NC score | 0.071391 (rank : 113) | θ value | θ > 10 (rank : 73) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 224 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O43184, O60470 | Gene names | ADAM12, MLTN | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 12 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 12) (Meltrin alpha). | |||||
|
ADA32_MOUSE
|
||||||
| NC score | 0.071231 (rank : 114) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8K410, Q6P901, Q8BJ80 | Gene names | Adam32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAM 32 precursor (A disintegrin and metalloproteinase domain 32). | |||||
|
LAMC1_MOUSE
|
||||||
| NC score | 0.070991 (rank : 115) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P02468 | Gene names | Lamc1, Lamb-2, Lamc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-1 chain precursor (Laminin B2 chain). | |||||
|
ADEC1_MOUSE
|
||||||
| NC score | 0.070929 (rank : 116) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9R0X2 | Gene names | Adamdec1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAM DEC1 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain-like protein decysin 1) (ADAM-like protein decysin 1). | |||||
|
ADA10_HUMAN
|
||||||
| NC score | 0.070917 (rank : 117) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O14672, Q10742, Q92650 | Gene names | ADAM10, KUZ, MADM | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 10 precursor (EC 3.4.24.81) (A disintegrin and metalloproteinase domain 10) (Mammalian disintegrin-metalloprotease) (Kuzbanian protein homolog) (CDw156c antigen). | |||||
|
ADA10_MOUSE
|
||||||
| NC score | 0.070793 (rank : 118) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O35598 | Gene names | Adam10, Kuz, Madm | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 10 precursor (EC 3.4.24.81) (A disintegrin and metalloproteinase domain 10) (Mammalian disintegrin-metalloprotease) (Kuzbanian protein homolog). | |||||
|
ADEC1_HUMAN
|
||||||
| NC score | 0.069563 (rank : 119) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O15204 | Gene names | ADAMDEC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAM DEC1 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain-like protein decysin 1) (ADAM-like protein decysin 1). | |||||
|
LAMC3_HUMAN
|
||||||
| NC score | 0.069252 (rank : 120) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 588 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9Y6N6 | Gene names | LAMC3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-3 chain precursor (Laminin 12 gamma 3 subunit). | |||||
|
LAMB1_MOUSE
|
||||||
| NC score | 0.068930 (rank : 121) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P02469 | Gene names | Lamb1-1, Lamb-1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
LAMC2_MOUSE
|
||||||
| NC score | 0.067838 (rank : 122) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q61092 | Gene names | Lamc2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-2 chain precursor (Laminin 5 gamma 2 subunit) (Kalinin/nicein/epiligrin 100 kDa subunit) (Laminin B2t chain). | |||||
|
LAMA3_MOUSE
|
||||||
| NC score | 0.067592 (rank : 123) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q61789, O08751, Q61788, Q61966, Q9JHQ7 | Gene names | Lama3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-3 chain precursor (Nicein subunit alpha). | |||||
|
LAMC1_HUMAN
|
||||||
| NC score | 0.067577 (rank : 124) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P11047 | Gene names | LAMC1, LAMB2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-1 chain precursor (Laminin B2 chain). | |||||
|
ATRN_MOUSE
|
||||||
| NC score | 0.067407 (rank : 125) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9WU60, Q9R263, Q9WU77 | Gene names | Atrn, mg, Mgca | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany protein). | |||||
|
LAMB1_HUMAN
|
||||||
| NC score | 0.067283 (rank : 126) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P07942 | Gene names | LAMB1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
LAMC3_MOUSE
|
||||||
| NC score | 0.065372 (rank : 127) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9R0B6, Q9WTW6 | Gene names | Lamc3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-3 chain precursor (Laminin 12 gamma 3 subunit). | |||||
|
LAMB3_HUMAN
|
||||||
| NC score | 0.065221 (rank : 128) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q13751, O14947, Q14733, Q9UJK4, Q9UJL1 | Gene names | LAMB3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-3 chain precursor (Laminin 5 beta 3) (Laminin B1k chain) (Kalinin B1 chain). | |||||
|
ATRN_HUMAN
|
||||||
| NC score | 0.065160 (rank : 129) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 525 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O75882, O60295, O95414, Q3MIT3, Q5TDA2, Q5TDA4, Q5VYW3, Q9NTQ3, Q9NTQ4, Q9NU01, Q9NZ57, Q9NZ58 | Gene names | ATRN, KIAA0548, MGCA | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany homolog) (DPPT-L). | |||||
|
LAMB3_MOUSE
|
||||||
| NC score | 0.064523 (rank : 130) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 444 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q61087 | Gene names | Lamb3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-3 chain precursor (Laminin 5 beta 3) (Kalinin B1 chain). | |||||
|
LAMC2_HUMAN
|
||||||
| NC score | 0.063524 (rank : 131) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 715 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q13753, Q02536, Q02537, Q13752, Q14941, Q5VYE8 | Gene names | LAMC2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-2 chain precursor (Laminin 5 gamma 2 subunit) (Kalinin/nicein/epiligrin 100 kDa subunit) (Laminin B2t chain) (Cell- scattering factor 140 kDa subunit) (CSF 140 kDa subunit) (Large adhesive scatter factor 140 kDa subunit) (Ladsin 140 kDa subunit). | |||||
|
WIF1_MOUSE
|
||||||
| NC score | 0.063054 (rank : 132) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9WUA1 | Gene names | Wif1 | |||
|
Domain Architecture |

|
|||||
| Description | Wnt inhibitory factor 1 precursor (WIF-1). | |||||
|
CREL1_HUMAN
|
||||||
| NC score | 0.062495 (rank : 133) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q96HD1, Q6I9X5, Q8NFT4, Q9Y409 | Gene names | CRELD1, CIRRIN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich with EGF-like domain protein 1 precursor. | |||||
|
LAMA3_HUMAN
|
||||||
| NC score | 0.062041 (rank : 134) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q16787, Q13679, Q13680, Q96TG0 | Gene names | LAMA3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-3 chain precursor (Epiligrin 170 kDa subunit) (E170) (Nicein subunit alpha). | |||||
|
TENA_MOUSE
|
||||||
| NC score | 0.061841 (rank : 135) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q80YX1, Q64706, Q80YX2 | Gene names | Tnc, Hxb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tenascin precursor (TN) (Tenascin-C) (TN-C) (Hexabrachion). | |||||
|
WIF1_HUMAN
|
||||||
| NC score | 0.060833 (rank : 136) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9Y5W5, Q6UXI1, Q8WVG4 | Gene names | WIF1 | |||
|
Domain Architecture |

|
|||||
| Description | Wnt inhibitory factor 1 precursor (WIF-1). | |||||
|
AGRIN_HUMAN
|
||||||
| NC score | 0.060308 (rank : 137) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 582 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O00468, Q5SVA2, Q60FE1, Q7KYS8, Q8N4J5, Q96IC1, Q9BTD4 | Gene names | AGRIN, AGRN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Agrin precursor. | |||||
|
TENA_HUMAN
|
||||||
| NC score | 0.059192 (rank : 138) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 631 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P24821, Q14583, Q15567 | Gene names | TNC, HXB | |||
|
Domain Architecture |

|
|||||
| Description | Tenascin precursor (TN) (Tenascin-C) (TN-C) (Hexabrachion) (Cytotactin) (Neuronectin) (GMEM) (JI) (Myotendinous antigen) (Glioma- associated-extracellular matrix antigen) (GP 150-225). | |||||
|
KR171_HUMAN
|
||||||
| NC score | 0.058472 (rank : 139) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9BYP8 | Gene names | KRTAP17-1, KAP17.1, KRTAP16.1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 17-1 (Keratin-associated protein 16.1). | |||||
|
JAG2_MOUSE
|
||||||
| NC score | 0.056630 (rank : 140) | θ value | 3.0926 (rank : 48) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9QYE5, O55139, O70219 | Gene names | Jag2 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-2 precursor (Jagged2). | |||||
|
LAMA4_MOUSE
|
||||||
| NC score | 0.056178 (rank : 141) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P97927, O88785, P70409 | Gene names | Lama4 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-4 chain precursor. | |||||
|
KIF4A_HUMAN
|
||||||
| NC score | 0.055837 (rank : 142) | θ value | 0.279714 (rank : 20) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O95239, Q86TN3, Q86XX7, Q9NNY6, Q9NY24, Q9UMW3 | Gene names | KIF4A, KIF4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
KR104_HUMAN
|
||||||
| NC score | 0.055702 (rank : 143) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P60372 | Gene names | KRTAP10-4, KAP10.4, KAP18-4, KRTAP10.4, KRTAP18-4, KRTAP18.4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 10-4 (Keratin-associated protein 10.4) (High sulfur keratin-associated protein 10.4) (Keratin-associated protein 18-4) (Keratin-associated protein 18.4). | |||||
|
KIF4A_MOUSE
|
||||||
| NC score | 0.055013 (rank : 144) | θ value | 0.365318 (rank : 24) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1033 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P33174 | Gene names | Kif4a, Kif4, Kns4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
STAB2_MOUSE
|
||||||
| NC score | 0.053681 (rank : 145) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8R4U0, Q8BM87 | Gene names | Stab2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stabilin-2 precursor [Contains: Short form stabilin-2]. | |||||
|
MEGF8_HUMAN
|
||||||
| NC score | 0.053623 (rank : 146) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q7Z7M0, O75097 | Gene names | MEGF8, EGFL4, KIAA0817 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
PCSK6_HUMAN
|
||||||
| NC score | 0.052732 (rank : 147) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P29122, Q15099, Q15100, Q9UEG7, Q9UEJ1, Q9UEJ2, Q9UEJ7, Q9UEJ8, Q9UEJ9, Q9Y4G9, Q9Y4H0, Q9Y4H1 | Gene names | PCSK6, PACE4 | |||
|
Domain Architecture |

|
|||||
| Description | Proprotein convertase subtilisin/kexin type 6 precursor (EC 3.4.21.-) (Paired basic amino acid cleaving enzyme 4) (Subtilisin/kexin-like protease PACE4) (Subtilisin-like proprotein convertase 4) (SPC4). | |||||
|
JAG1_MOUSE
|
||||||
| NC score | 0.052519 (rank : 148) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9QXX0 | Gene names | Jag1 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-1 precursor (Jagged1) (CD339 antigen). | |||||
|
JAG2_HUMAN
|
||||||
| NC score | 0.052217 (rank : 149) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9Y219, Q9UE17, Q9UE99, Q9UNK8, Q9Y6P9, Q9Y6Q0 | Gene names | JAG2 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-2 precursor (Jagged2) (HJ2). | |||||
|
JAG1_HUMAN
|
||||||
| NC score | 0.051923 (rank : 150) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 514 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P78504, O14902, O15122, Q15816 | Gene names | JAG1, JAGL1 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-1 precursor (Jagged1) (hJ1) (CD339 antigen). | |||||
|
STAB1_MOUSE
|
||||||
| NC score | 0.051850 (rank : 151) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 566 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8R4Y4, Q8K0K6, Q8VC09 | Gene names | Stab1, Feel1 | |||
|
Domain Architecture |

|
|||||
| Description | Stabilin-1 precursor (FEEL-1 protein). | |||||
|
STAB2_HUMAN
|
||||||
| NC score | 0.051171 (rank : 152) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8WWQ8, Q6ZMK2, Q7Z5N9, Q86UR4, Q8IUG9, Q8TES1, Q9H7H7, Q9NRY3 | Gene names | STAB2, FEEL2, FELL, FEX2, HARE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stabilin-2 precursor (FEEL-2 protein) (Fasciclin EGF-like laminin-type EGF-like and link domain-containing scavenger receptor 1) (FAS1 EGF- like and X-link domain-containing adhesion molecule 2) (Hyaluronan receptor for endocytosis) [Contains: 190 kDa form stabilin-2 (190 kDa hyaluronan receptor for endocytosis)]. | |||||
|
NET4_MOUSE
|
||||||
| NC score | 0.051004 (rank : 153) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9JI33 | Gene names | Ntn4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin-4 precursor (Beta-netrin). | |||||
|
NTNG2_HUMAN
|
||||||
| NC score | 0.050883 (rank : 154) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q96CW9, Q6UXY0, Q96JH0 | Gene names | NTNG2, KIAA1857, LMNT2 | |||
|
Domain Architecture |

|
|||||
| Description | Netrin G2 precursor (Laminet-2). | |||||
|
NET4_HUMAN
|
||||||
| NC score | 0.050746 (rank : 155) | θ value | θ > 10 (rank : 146) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9HB63, Q658K9, Q7L3F1, Q7L9D6, Q7Z5B6, Q9BZP1, Q9NT44, Q9P133 | Gene names | NTN4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin-4 precursor (Beta-netrin) (Hepar-derived netrin-like protein). | |||||
|
MEGF8_MOUSE
|
||||||
| NC score | 0.050525 (rank : 156) | θ value | θ > 10 (rank : 141) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 520 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P60882, Q80TR3, Q80V41, Q8BMN9, Q8JZW7, Q8K0J3 | Gene names | Megf8, Egfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
KR106_HUMAN
|
||||||
| NC score | 0.050067 (rank : 157) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P60371 | Gene names | KRTAP10-6, KAP10.6, KAP18-6, KRTAP10.6, KRTAP18-6, KRTAP18.6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 10-6 (Keratin-associated protein 10.6) (High sulfur keratin-associated protein 10.6) (Keratin-associated protein 18-6) (Keratin-associated protein 18.6). | |||||
|
PGBM_HUMAN
|
||||||
| NC score | 0.048024 (rank : 158) | θ value | 6.88961 (rank : 60) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1113 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P98160, Q16287, Q9H3V5 | Gene names | HSPG2 | |||
|
Domain Architecture |

|
|||||
| Description | Basement membrane-specific heparan sulfate proteoglycan core protein precursor (HSPG) (Perlecan) (PLC). | |||||
|
FBLN2_MOUSE
|
||||||
| NC score | 0.031466 (rank : 159) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P37889, Q9WUI2 | Gene names | Fbln2 | |||
|
Domain Architecture |

|
|||||
| Description | Fibulin-2 precursor. | |||||
|
ST14_HUMAN
|
||||||
| NC score | 0.013574 (rank : 160) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y5Y6, Q9BS01, Q9H3S0, Q9HB36, Q9HCA3 | Gene names | ST14, PRSS14, SNC19, TADG15 | |||
|
Domain Architecture |

|
|||||
| Description | Suppressor of tumorigenicity protein 14 (EC 3.4.21.-) (Serine protease 14) (Matriptase) (Membrane-type serine protease 1) (MT-SP1) (Prostamin) (Serine protease TADG-15) (Tumor-associated differentially-expressed gene 15 protein). | |||||