Please be patient as the page loads
|
GGA2_MOUSE
|
||||||
| SwissProt Accessions | Q6P5E6, Q3U374, Q6PFX3, Q6ZPY8, Q8BM76, Q9DC15 | Gene names | Gga2, Kiaa1080 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADP-ribosylation factor-binding protein GGA2 (Golgi-localized, gamma ear-containing, ARF-binding protein 2) (Gamma-adaptin-related protein 2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
GGA2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.988972 (rank : 2) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UJY4, O14564, Q9NYN2, Q9UPS2 | Gene names | GGA2, KIAA1080 | |||
|
Domain Architecture |

|
|||||
| Description | ADP-ribosylation factor-binding protein GGA2 (Golgi-localized, gamma ear-containing, ARF-binding protein 2) (Gamma-adaptin-related protein 2) (VHS domain and ear domain of gamma-adaptin) (Vear). | |||||
|
GGA2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q6P5E6, Q3U374, Q6PFX3, Q6ZPY8, Q8BM76, Q9DC15 | Gene names | Gga2, Kiaa1080 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADP-ribosylation factor-binding protein GGA2 (Golgi-localized, gamma ear-containing, ARF-binding protein 2) (Gamma-adaptin-related protein 2). | |||||
|
GGA1_MOUSE
|
||||||
| θ value | 3.4095e-124 (rank : 3) | NC score | 0.952732 (rank : 3) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8R0H9, Q3U2N1 | Gene names | Gga1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADP-ribosylation factor-binding protein GGA1 (Golgi-localized, gamma ear-containing, ARF-binding protein 1) (Gamma-adaptin-related protein 1). | |||||
|
GGA1_HUMAN
|
||||||
| θ value | 2.06847e-121 (rank : 4) | NC score | 0.952646 (rank : 4) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UJY5, Q5TG07, Q9BW94, Q9UG00, Q9UGW0, Q9UGW1 | Gene names | GGA1 | |||
|
Domain Architecture |

|
|||||
| Description | ADP-ribosylation factor-binding protein GGA1 (Golgi-localized, gamma ear-containing, ARF-binding protein 1) (Gamma-adaptin-related protein 1). | |||||
|
GGA3_HUMAN
|
||||||
| θ value | 2.02215e-84 (rank : 5) | NC score | 0.936838 (rank : 5) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9NZ52, Q15017, Q6IS16, Q9UJY3 | Gene names | GGA3, KIAA0154 | |||
|
Domain Architecture |

|
|||||
| Description | ADP-ribosylation factor-binding protein GGA3 (Golgi-localized, gamma ear-containing, ARF-binding protein 3). | |||||
|
GGA3_MOUSE
|
||||||
| θ value | 1.00358e-83 (rank : 6) | NC score | 0.925488 (rank : 6) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BMI3, Q80U71 | Gene names | Gga3, Kiaa0154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADP-ribosylation factor-binding protein GGA3 (Golgi-localized, gamma ear-containing, ARF-binding protein 3). | |||||
|
TM1L1_MOUSE
|
||||||
| θ value | 3.40345e-15 (rank : 7) | NC score | 0.645011 (rank : 9) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q923U0, Q99KE0, Q9D6Y5 | Gene names | Tom1l1, Srcasm | |||
|
Domain Architecture |
|
|||||
| Description | TOM1-like 1 protein (Target of myb-like 1 protein) (Src-activating and signaling molecule protein). | |||||
|
AP1G1_HUMAN
|
||||||
| θ value | 7.58209e-15 (rank : 8) | NC score | 0.396207 (rank : 11) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43747, O75709, O75842, Q9UG09, Q9Y3U4 | Gene names | AP1G1, ADTG, CLAPG1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit gamma-1 (Adapter-related protein complex 1 gamma- 1 subunit) (Gamma-adaptin) (Adaptor protein complex AP-1 gamma-1 subunit) (Golgi adaptor HA1/AP1 adaptin subunit gamma-1) (Clathrin assembly protein complex 1 gamma-1 large chain). | |||||
|
TOM1_MOUSE
|
||||||
| θ value | 1.68911e-14 (rank : 9) | NC score | 0.662699 (rank : 7) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O88746, Q3V4C6, Q9D120 | Gene names | Tom1 | |||
|
Domain Architecture |
|
|||||
| Description | Target of Myb protein 1. | |||||
|
AP1G1_MOUSE
|
||||||
| θ value | 3.76295e-14 (rank : 10) | NC score | 0.388645 (rank : 15) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P22892 | Gene names | Ap1g1, Adtg, Clapg1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit gamma-1 (Adapter-related protein complex 1 gamma- 1 subunit) (Gamma-adaptin) (Adaptor protein complex AP-1 gamma-1 subunit) (Golgi adaptor HA1/AP1 adaptin subunit gamma-1) (Clathrin assembly protein complex 1 gamma-1 large chain). | |||||
|
TOM1_HUMAN
|
||||||
| θ value | 4.91457e-14 (rank : 11) | NC score | 0.657628 (rank : 8) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O60784, Q5TIJ6 | Gene names | TOM1 | |||
|
Domain Architecture |
|
|||||
| Description | Target of Myb protein 1. | |||||
|
AP1G2_MOUSE
|
||||||
| θ value | 1.86753e-13 (rank : 12) | NC score | 0.381169 (rank : 17) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O88512 | Gene names | Ap1g2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit gamma-2 (Adapter-related protein complex 1 gamma- 2 subunit) (Gamma2-adaptin) (Adaptor protein complex AP-1 gamma-2 subunit) (G2ad). | |||||
|
TM1L1_HUMAN
|
||||||
| θ value | 1.86753e-13 (rank : 13) | NC score | 0.631511 (rank : 10) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O75674 | Gene names | TOM1L1, SRCASM | |||
|
Domain Architecture |
|
|||||
| Description | TOM1-like 1 protein (Target of myb-like 1 protein) (Src-activating and signaling molecule protein). | |||||
|
AP1G2_HUMAN
|
||||||
| θ value | 1.47974e-10 (rank : 14) | NC score | 0.371195 (rank : 20) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O75843, O75504 | Gene names | AP1G2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit gamma-2 (Adapter-related protein complex 1 gamma- 2 subunit) (Gamma2-adaptin) (Adaptor protein complex AP-1 gamma-2 subunit) (G2ad). | |||||
|
HGS_MOUSE
|
||||||
| θ value | 1.9326e-10 (rank : 15) | NC score | 0.391938 (rank : 14) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q99LI8, Q61691, Q8BQW3 | Gene names | Hgs, Hrs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate. | |||||
|
HGS_HUMAN
|
||||||
| θ value | 3.29651e-10 (rank : 16) | NC score | 0.392918 (rank : 12) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O14964, Q9NR36 | Gene names | HGS, HRS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate (Protein pp110) (Hrs). | |||||
|
STAM2_HUMAN
|
||||||
| θ value | 1.63604e-09 (rank : 17) | NC score | 0.392372 (rank : 13) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O75886, Q7LDQ0, Q9UF58 | Gene names | STAM2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal transducing adapter molecule 2 (STAM-2). | |||||
|
STAM2_MOUSE
|
||||||
| θ value | 3.64472e-09 (rank : 18) | NC score | 0.377008 (rank : 19) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O88811, Q8C8Y4 | Gene names | Stam2, Hbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal transducing adapter molecule 2 (STAM-2) (Hrs-binding protein). | |||||
|
STAM1_HUMAN
|
||||||
| θ value | 1.06045e-08 (rank : 19) | NC score | 0.381971 (rank : 16) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q92783, Q8N6D9 | Gene names | STAM, STAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal transducing adapter molecule 1 (STAM-1). | |||||
|
STAM1_MOUSE
|
||||||
| θ value | 1.38499e-08 (rank : 20) | NC score | 0.377046 (rank : 18) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P70297 | Gene names | Stam, Stam1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal transducing adapter molecule 1 (STAM-1). | |||||
|
SRBP1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 21) | NC score | 0.048823 (rank : 32) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
K0555_HUMAN
|
||||||
| θ value | 0.125558 (rank : 22) | NC score | 0.022184 (rank : 44) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 880 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96AA8, O60302 | Gene names | KIAA0555 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0555. | |||||
|
HNF1A_HUMAN
|
||||||
| θ value | 0.163984 (rank : 23) | NC score | 0.051917 (rank : 29) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P20823, Q99861 | Gene names | TCF1, HNF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-alpha (HNF-1A) (Liver-specific transcription factor LF-B1) (LFB1) (Transcription factor 1) (TCF-1). | |||||
|
HNF1A_MOUSE
|
||||||
| θ value | 0.21417 (rank : 24) | NC score | 0.045457 (rank : 34) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P22361 | Gene names | Tcf1, Hnf-1, Hnf-1a, Hnf1a | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-alpha (HNF-1A) (Liver-specific transcription factor LF-B1) (LFB1). | |||||
|
SMCA4_HUMAN
|
||||||
| θ value | 0.365318 (rank : 25) | NC score | 0.022361 (rank : 43) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P51532, O95052, Q9HBD3 | Gene names | SMARCA4, BRG1, SNF2B, SNF2L4 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L4 (EC 3.6.1.-) (ATP- dependent helicase SMARCA4) (SNF2-beta) (BRG-1 protein) (Mitotic growth and transcription activator) (Brahma protein homolog 1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 4). | |||||
|
IRX4_MOUSE
|
||||||
| θ value | 0.47712 (rank : 26) | NC score | 0.023125 (rank : 41) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QY61 | Gene names | Irx4, Irxa3 | |||
|
Domain Architecture |

|
|||||
| Description | Iroquois-class homeodomain protein IRX-4 (Iroquois homeobox protein 4) (Homeodomain protein IRXA3). | |||||
|
RAPH1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 27) | NC score | 0.024654 (rank : 37) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
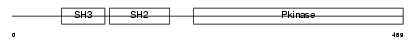
ABL1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 28) | NC score | 0.006093 (rank : 59) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1178 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P00520, P97896, Q61252, Q61253, Q61254, Q61255, Q61256, Q61257, Q61258, Q61259, Q61260, Q61261, Q6PCM5, Q8C1X4 | Gene names | Abl1, Abl | |||
|
Domain Architecture |
|
|||||
| Description | Proto-oncogene tyrosine-protein kinase ABL1 (EC 2.7.10.2) (p150) (c- ABL) (Abelson murine leukemia viral oncogene homolog 1). | |||||
|
SWP70_MOUSE
|
||||||
| θ value | 0.813845 (rank : 29) | NC score | 0.017765 (rank : 48) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1097 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6A028, O88443, Q3TQR6, Q3UCA3, Q6P1D0 | Gene names | Swap70, Kiaa0640 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Switch-associated protein 70 (SWAP-70). | |||||
|
CCD96_MOUSE
|
||||||
| θ value | 1.38821 (rank : 30) | NC score | 0.022615 (rank : 42) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9CR92 | Gene names | Ccdc96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
LDB3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 31) | NC score | 0.018824 (rank : 47) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75112, Q5K6N9, Q5K6P0, Q5K6P1, Q96FH2, Q9Y4Z3, Q9Y4Z4, Q9Y4Z5 | Gene names | LDB3, KIAA0613, ZASP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LIM domain-binding protein 3 (Z-band alternatively spliced PDZ-motif protein) (Protein cypher). | |||||
|
SYNE1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 32) | NC score | 0.011434 (rank : 55) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
COBA2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 33) | NC score | 0.005837 (rank : 62) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P13942, Q07751, Q13271, Q13272, Q13273, Q99866, Q9UIP9 | Gene names | COL11A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
PASK_HUMAN
|
||||||
| θ value | 1.81305 (rank : 34) | NC score | 0.000872 (rank : 68) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 857 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96RG2, Q86XH6, Q99763, Q9UFR7 | Gene names | PASK, KIAA0135 | |||
|
Domain Architecture |

|
|||||
| Description | PAS domain-containing serine/threonine-protein kinase (EC 2.7.11.1) (PAS-kinase) (PASKIN) (hPASK). | |||||
|
SREC2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 35) | NC score | 0.004932 (rank : 64) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 605 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P59222 | Gene names | Scarf2, Srec2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor class F member 2 precursor (Scavenger receptor expressed by endothelial cells 2 protein) (SREC-II). | |||||
|
ALEX_MOUSE
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.047946 (rank : 33) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6R0H6, Q8BIR3 | Gene names | Gnas, Gnas1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ALEX (Alternative gene product encoded by XL-exon). | |||||
|
CCDC7_HUMAN
|
||||||
| θ value | 2.36792 (rank : 37) | NC score | 0.022133 (rank : 45) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 253 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96M83, Q8IVQ0, Q8NEQ0 | Gene names | CCDC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 7. | |||||
|
SPTN2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 38) | NC score | 0.011889 (rank : 54) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 535 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O15020, O14872, O14873 | Gene names | SPTBN2, KIAA0302 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 2 (Spectrin, non-erythroid beta chain 2) (Beta-III spectrin). | |||||
|
T240L_HUMAN
|
||||||
| θ value | 2.36792 (rank : 39) | NC score | 0.024448 (rank : 39) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q71F56, Q68DN4, Q9H8C0, Q9NSY9, Q9UFD8, Q9UPX5 | Gene names | THRAP2, KIAA1025, PROSIT240, TRAP240L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 2 (Thyroid hormone receptor-associated protein complex 240 kDa component-like). | |||||
|
T240L_MOUSE
|
||||||
| θ value | 2.36792 (rank : 40) | NC score | 0.024647 (rank : 38) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6JPI3, Q3TRF4, Q3UQI8, Q80TM3, Q80WQ0 | Gene names | Thrap2, Kiaa1025, Trap240l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 2 (Thyroid hormone receptor-associated protein complex 240 kDa component-like). | |||||
|
GNDS_HUMAN
|
||||||
| θ value | 3.0926 (rank : 41) | NC score | 0.010598 (rank : 56) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q12967, Q9HAX7, Q9HAY1, Q9HCT1 | Gene names | RALGDS, RGF | |||
|
Domain Architecture |

|
|||||
| Description | Ral guanine nucleotide dissociation stimulator (RalGEF) (RalGDS). | |||||
|
LRC8C_MOUSE
|
||||||
| θ value | 3.0926 (rank : 42) | NC score | 0.007860 (rank : 58) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R502, Q3TEP9, Q8C296, Q8R0N7, Q8R3G5 | Gene names | Lrrc8c, Ad158 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 8C (Protein AD158). | |||||
|
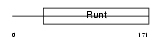
RUNX2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 43) | NC score | 0.031620 (rank : 36) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13950, O14614, O14615, O95181 | Gene names | RUNX2, AML3, CBFA1, OSF2, PEBP2A | |||
|
Domain Architecture |
|
|||||
| Description | Runt-related transcription factor 2 (Core-binding factor, alpha 1 subunit) (CBF-alpha 1) (Acute myeloid leukemia 3 protein) (Oncogene AML-3) (Polyomavirus enhancer-binding protein 2 alpha A subunit) (PEBP2-alpha A) (PEA2-alpha A) (SL3-3 enhancer factor 1 alpha A subunit) (SL3/AKV core-binding factor alpha A subunit) (Osteoblast- specific transcription factor 2) (OSF-2). | |||||
|
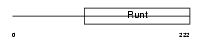
RUNX2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 44) | NC score | 0.031732 (rank : 35) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q08775, O35183, Q08776, Q9JLN0, Q9QUQ6, Q9QY29, Q9R0U4, Q9Z2J7 | Gene names | Runx2, Aml3, Cbfa1, Osf2, Pebp2a | |||
|
Domain Architecture |
|
|||||
| Description | Runt-related transcription factor 2 (Core-binding factor, alpha 1 subunit) (CBF-alpha 1) (Acute myeloid leukemia 3 protein) (Oncogene AML-3) (Polyomavirus enhancer-binding protein 2 alpha A subunit) (PEBP2-alpha A) (PEA2-alpha A) (SL3-3 enhancer factor 1 alpha A subunit) (SL3/AKV core-binding factor alpha A subunit) (Osteoblast- specific transcription factor 2) (OSF-2). | |||||
|
ITSN2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 45) | NC score | 0.020473 (rank : 46) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1173 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NZM3, O95062, Q15812, Q9HAK4, Q9NXE6, Q9NYG0, Q9NZM2, Q9ULG4 | Gene names | ITSN2, KIAA1256, SH3D1B, SWAP | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-2 (SH3 domain-containing protein 1B) (SH3P18) (SH3P18-like WASP-associated protein). | |||||
|
LDB3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.012600 (rank : 53) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 510 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9JKS4, Q6A038, Q811P2, Q811P3, Q811P4, Q811P5, Q9D130, Q9JKS3, Q9R0Z1, Q9WVH1, Q9WVH2 | Gene names | Ldb3, Kiaa0613 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LIM domain-binding protein 3 (Z-band alternatively spliced PDZ-motif protein) (Protein cypher) (Protein oracle). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 47) | NC score | 0.014554 (rank : 50) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
AP2B1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 48) | NC score | 0.049631 (rank : 31) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P63010, P21851, Q96J19 | Gene names | AP2B1, CLAPB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-2 complex subunit beta-1 (Adapter-related protein complex 2 beta-1 subunit) (Beta-adaptin) (Plasma membrane adaptor HA2/AP2 adaptin beta subunit) (Clathrin assembly protein complex 2 beta large chain) (AP105B). | |||||
|
AP2B1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | 0.049634 (rank : 30) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9DBG3, Q80XJ4 | Gene names | Ap2b1, Clapb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-2 complex subunit beta-1 (Adapter-related protein complex 2 beta-1 subunit) (Beta-adaptin) (Plasma membrane adaptor HA2/AP2 adaptin beta subunit) (Clathrin assembly protein complex 2 beta large chain) (AP105B). | |||||
|
CEP68_MOUSE
|
||||||
| θ value | 5.27518 (rank : 50) | NC score | 0.014816 (rank : 49) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C0D9, Q3US49, Q5SW85, Q8R2V0 | Gene names | Cep68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 68 kDa (Cep68 protein). | |||||
|
CRBB1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 51) | NC score | 0.009532 (rank : 57) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P53674 | Gene names | CRYBB1 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin B1. | |||||
|
MRP2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 52) | NC score | 0.005946 (rank : 60) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VI47 | Gene names | Abcc2 | |||
|
Domain Architecture |

|
|||||
| Description | Canalicular multispecific organic anion transporter 1 (ATP-binding cassette sub-family C member 2). | |||||
|
PEX1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.005526 (rank : 63) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O43933, Q96S71, Q96S72, Q96S73, Q99994 | Gene names | PEX1 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisome biogenesis factor 1 (Peroxin-1) (Peroxisome biogenesis disorder protein 1). | |||||
|
CD248_MOUSE
|
||||||
| θ value | 6.88961 (rank : 54) | NC score | 0.004769 (rank : 65) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 578 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q91V98, Q3UMV6, Q91ZV1 | Gene names | Cd248, Tem1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endosialin precursor (Tumor endothelial marker 1) (CD248 antigen). | |||||
|
CN092_MOUSE
|
||||||
| θ value | 6.88961 (rank : 55) | NC score | 0.023309 (rank : 40) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BU11, Q80UI2, Q99PN9, Q9CS16 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Epidermal Langerhans cell protein LCP1. | |||||
|
SORC3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 56) | NC score | 0.005899 (rank : 61) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UPU3, Q5VXF9, Q9NQJ2 | Gene names | SORCS3, KIAA1059 | |||
|
Domain Architecture |

|
|||||
| Description | VPS10 domain-containing receptor SorCS3 precursor. | |||||
|
CE290_HUMAN
|
||||||
| θ value | 8.99809 (rank : 57) | NC score | 0.014044 (rank : 51) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
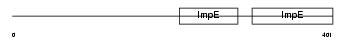
KLC4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 58) | NC score | 0.003923 (rank : 66) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 336 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9DBS5, Q3TCC5 | Gene names | Klc4, Knsl8 | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin light chain 4 (KLC 4) (Kinesin-like protein 8). | |||||
|
RFIP3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 59) | NC score | 0.013471 (rank : 52) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1055 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O75154, Q4VXV7, Q9H155, Q9H1G0, Q9NUI0 | Gene names | RAB11FIP3, KIAA0665 | |||
|
Domain Architecture |

|
|||||
| Description | Rab11 family-interacting protein 3 (Rab11-FIP3) (EF hands-containing Rab-interacting protein) (Eferin). | |||||
|
SPB6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 60) | NC score | 0.001366 (rank : 67) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P35237, Q96J44 | Gene names | SERPINB6, PI6, PTI | |||
|
Domain Architecture |

|
|||||
| Description | Serpin B6 (Placental thrombin inhibitor) (Cytoplasmic antiproteinase) (CAP) (Protease inhibitor 6) (PI-6). | |||||
|
AP2A1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.110720 (rank : 22) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O95782, Q96CI7, Q96PP6, Q96PP7, Q9H070 | Gene names | AP2A1, ADTAA | |||
|
Domain Architecture |

|
|||||
| Description | AP-2 complex subunit alpha-1 (Adapter-related protein complex 2 alpha- 1 subunit) (Alpha-adaptin A) (Adaptor protein complex AP-2 alpha-1 subunit) (Clathrin assembly protein complex 2 alpha-A large chain) (100 kDa coated vesicle protein A) (Plasma membrane adaptor HA2/AP2 adaptin alpha A subunit). | |||||
|
AP2A1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.108116 (rank : 24) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P17426 | Gene names | Ap2a1, Adtaa, Clapa1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-2 complex subunit alpha-1 (Adapter-related protein complex 2 alpha- 1 subunit) (Alpha-adaptin A) (Adaptor protein complex AP-2 alpha-1 subunit) (Clathrin assembly protein complex 2 alpha-A large chain) (100 kDa coated vesicle protein A) (Plasma membrane adaptor HA2/AP2 adaptin alpha A subunit). | |||||
|
AP2A2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.107777 (rank : 26) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O94973, O75403, Q96SI8 | Gene names | AP2A2, ADTAB, KIAA0899 | |||
|
Domain Architecture |

|
|||||
| Description | AP-2 complex subunit alpha-2 (Adapter-related protein complex 2 alpha- 2 subunit) (Alpha-adaptin C) (Adaptor protein complex AP-2 alpha-2 subunit) (Clathrin assembly protein complex 2 alpha-C large chain) (100 kDa coated vesicle protein C) (Plasma membrane adaptor HA2/AP2 adaptin alpha C subunit) (Huntingtin-interacting protein HYPJ). | |||||
|
AP2A2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.107941 (rank : 25) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P17427, Q921V0 | Gene names | Ap2a2, Adtab | |||
|
Domain Architecture |

|
|||||
| Description | AP-2 complex subunit alpha-2 (Adapter-related protein complex 2 alpha- 2 subunit) (Alpha-adaptin C) (Adaptor protein complex AP-2 alpha-2 subunit) (Clathrin assembly protein complex 2 alpha-C large chain) (100 kDa coated vesicle protein C) (Plasma membrane adaptor HA2/AP2 adaptin alpha C subunit). | |||||
|
AP3D1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.095340 (rank : 28) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O14617, O00202, O75262, Q59HF5, Q96G11, Q9H3C6 | Gene names | AP3D1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit delta-1 (Adapter-related protein complex 3 subunit delta-1) (Delta-adaptin 3) (AP-3 complex subunit delta) (Delta-adaptin). | |||||
|
AP3D1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.100562 (rank : 27) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O54774 | Gene names | Ap3d1, Ap3d | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-3 complex subunit delta-1 (Adapter-related protein complex 3 subunit delta-1) (Delta-adaptin 3) (AP-3 complex subunit delta) (Delta-adaptin) (mBLVR1). | |||||
|
AP4E1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 67) | NC score | 0.111394 (rank : 21) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UPM8, Q9Y588 | Gene names | AP4E1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-4 complex subunit epsilon-1 (Adapter-related protein complex 4 epsilon-1 subunit) (Epsilon subunit of AP-4) (AP-4 adapter complex epsilon subunit). | |||||
|
AP4E1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 68) | NC score | 0.110644 (rank : 23) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80V94 | Gene names | Ap4e1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-4 complex subunit epsilon-1 (Adapter-related protein complex 4 epsilon-1 subunit) (Epsilon subunit of AP-4) (AP-4 adapter complex epsilon subunit). | |||||
|
GGA2_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q6P5E6, Q3U374, Q6PFX3, Q6ZPY8, Q8BM76, Q9DC15 | Gene names | Gga2, Kiaa1080 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADP-ribosylation factor-binding protein GGA2 (Golgi-localized, gamma ear-containing, ARF-binding protein 2) (Gamma-adaptin-related protein 2). | |||||
|
GGA2_HUMAN
|
||||||
| NC score | 0.988972 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UJY4, O14564, Q9NYN2, Q9UPS2 | Gene names | GGA2, KIAA1080 | |||
|
Domain Architecture |

|
|||||
| Description | ADP-ribosylation factor-binding protein GGA2 (Golgi-localized, gamma ear-containing, ARF-binding protein 2) (Gamma-adaptin-related protein 2) (VHS domain and ear domain of gamma-adaptin) (Vear). | |||||
|
GGA1_MOUSE
|
||||||
| NC score | 0.952732 (rank : 3) | θ value | 3.4095e-124 (rank : 3) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8R0H9, Q3U2N1 | Gene names | Gga1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADP-ribosylation factor-binding protein GGA1 (Golgi-localized, gamma ear-containing, ARF-binding protein 1) (Gamma-adaptin-related protein 1). | |||||
|
GGA1_HUMAN
|
||||||
| NC score | 0.952646 (rank : 4) | θ value | 2.06847e-121 (rank : 4) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UJY5, Q5TG07, Q9BW94, Q9UG00, Q9UGW0, Q9UGW1 | Gene names | GGA1 | |||
|
Domain Architecture |

|
|||||
| Description | ADP-ribosylation factor-binding protein GGA1 (Golgi-localized, gamma ear-containing, ARF-binding protein 1) (Gamma-adaptin-related protein 1). | |||||
|
GGA3_HUMAN
|
||||||
| NC score | 0.936838 (rank : 5) | θ value | 2.02215e-84 (rank : 5) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9NZ52, Q15017, Q6IS16, Q9UJY3 | Gene names | GGA3, KIAA0154 | |||
|
Domain Architecture |

|
|||||
| Description | ADP-ribosylation factor-binding protein GGA3 (Golgi-localized, gamma ear-containing, ARF-binding protein 3). | |||||
|
GGA3_MOUSE
|
||||||
| NC score | 0.925488 (rank : 6) | θ value | 1.00358e-83 (rank : 6) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BMI3, Q80U71 | Gene names | Gga3, Kiaa0154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADP-ribosylation factor-binding protein GGA3 (Golgi-localized, gamma ear-containing, ARF-binding protein 3). | |||||
|
TOM1_MOUSE
|
||||||
| NC score | 0.662699 (rank : 7) | θ value | 1.68911e-14 (rank : 9) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O88746, Q3V4C6, Q9D120 | Gene names | Tom1 | |||
|
Domain Architecture |

|
|||||
| Description | Target of Myb protein 1. | |||||
|
TOM1_HUMAN
|
||||||
| NC score | 0.657628 (rank : 8) | θ value | 4.91457e-14 (rank : 11) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O60784, Q5TIJ6 | Gene names | TOM1 | |||
|
Domain Architecture |

|
|||||
| Description | Target of Myb protein 1. | |||||
|
TM1L1_MOUSE
|
||||||
| NC score | 0.645011 (rank : 9) | θ value | 3.40345e-15 (rank : 7) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q923U0, Q99KE0, Q9D6Y5 | Gene names | Tom1l1, Srcasm | |||
|
Domain Architecture |

|
|||||
| Description | TOM1-like 1 protein (Target of myb-like 1 protein) (Src-activating and signaling molecule protein). | |||||
|
TM1L1_HUMAN
|
||||||
| NC score | 0.631511 (rank : 10) | θ value | 1.86753e-13 (rank : 13) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O75674 | Gene names | TOM1L1, SRCASM | |||
|
Domain Architecture |

|
|||||
| Description | TOM1-like 1 protein (Target of myb-like 1 protein) (Src-activating and signaling molecule protein). | |||||
|
AP1G1_HUMAN
|
||||||
| NC score | 0.396207 (rank : 11) | θ value | 7.58209e-15 (rank : 8) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43747, O75709, O75842, Q9UG09, Q9Y3U4 | Gene names | AP1G1, ADTG, CLAPG1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit gamma-1 (Adapter-related protein complex 1 gamma- 1 subunit) (Gamma-adaptin) (Adaptor protein complex AP-1 gamma-1 subunit) (Golgi adaptor HA1/AP1 adaptin subunit gamma-1) (Clathrin assembly protein complex 1 gamma-1 large chain). | |||||
|
HGS_HUMAN
|
||||||
| NC score | 0.392918 (rank : 12) | θ value | 3.29651e-10 (rank : 16) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O14964, Q9NR36 | Gene names | HGS, HRS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate (Protein pp110) (Hrs). | |||||
|
STAM2_HUMAN
|
||||||
| NC score | 0.392372 (rank : 13) | θ value | 1.63604e-09 (rank : 17) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O75886, Q7LDQ0, Q9UF58 | Gene names | STAM2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal transducing adapter molecule 2 (STAM-2). | |||||
|
HGS_MOUSE
|
||||||
| NC score | 0.391938 (rank : 14) | θ value | 1.9326e-10 (rank : 15) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q99LI8, Q61691, Q8BQW3 | Gene names | Hgs, Hrs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate. | |||||
|
AP1G1_MOUSE
|
||||||
| NC score | 0.388645 (rank : 15) | θ value | 3.76295e-14 (rank : 10) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P22892 | Gene names | Ap1g1, Adtg, Clapg1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit gamma-1 (Adapter-related protein complex 1 gamma- 1 subunit) (Gamma-adaptin) (Adaptor protein complex AP-1 gamma-1 subunit) (Golgi adaptor HA1/AP1 adaptin subunit gamma-1) (Clathrin assembly protein complex 1 gamma-1 large chain). | |||||
|
STAM1_HUMAN
|
||||||
| NC score | 0.381971 (rank : 16) | θ value | 1.06045e-08 (rank : 19) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q92783, Q8N6D9 | Gene names | STAM, STAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal transducing adapter molecule 1 (STAM-1). | |||||
|
AP1G2_MOUSE
|
||||||
| NC score | 0.381169 (rank : 17) | θ value | 1.86753e-13 (rank : 12) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O88512 | Gene names | Ap1g2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit gamma-2 (Adapter-related protein complex 1 gamma- 2 subunit) (Gamma2-adaptin) (Adaptor protein complex AP-1 gamma-2 subunit) (G2ad). | |||||
|
STAM1_MOUSE
|
||||||
| NC score | 0.377046 (rank : 18) | θ value | 1.38499e-08 (rank : 20) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P70297 | Gene names | Stam, Stam1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal transducing adapter molecule 1 (STAM-1). | |||||
|
STAM2_MOUSE
|
||||||
| NC score | 0.377008 (rank : 19) | θ value | 3.64472e-09 (rank : 18) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O88811, Q8C8Y4 | Gene names | Stam2, Hbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal transducing adapter molecule 2 (STAM-2) (Hrs-binding protein). | |||||
|
AP1G2_HUMAN
|
||||||
| NC score | 0.371195 (rank : 20) | θ value | 1.47974e-10 (rank : 14) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O75843, O75504 | Gene names | AP1G2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit gamma-2 (Adapter-related protein complex 1 gamma- 2 subunit) (Gamma2-adaptin) (Adaptor protein complex AP-1 gamma-2 subunit) (G2ad). | |||||
|
AP4E1_HUMAN
|
||||||
| NC score | 0.111394 (rank : 21) | θ value | θ > 10 (rank : 67) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UPM8, Q9Y588 | Gene names | AP4E1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-4 complex subunit epsilon-1 (Adapter-related protein complex 4 epsilon-1 subunit) (Epsilon subunit of AP-4) (AP-4 adapter complex epsilon subunit). | |||||
|
AP2A1_HUMAN
|
||||||
| NC score | 0.110720 (rank : 22) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O95782, Q96CI7, Q96PP6, Q96PP7, Q9H070 | Gene names | AP2A1, ADTAA | |||
|
Domain Architecture |

|
|||||
| Description | AP-2 complex subunit alpha-1 (Adapter-related protein complex 2 alpha- 1 subunit) (Alpha-adaptin A) (Adaptor protein complex AP-2 alpha-1 subunit) (Clathrin assembly protein complex 2 alpha-A large chain) (100 kDa coated vesicle protein A) (Plasma membrane adaptor HA2/AP2 adaptin alpha A subunit). | |||||
|
AP4E1_MOUSE
|
||||||
| NC score | 0.110644 (rank : 23) | θ value | θ > 10 (rank : 68) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80V94 | Gene names | Ap4e1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-4 complex subunit epsilon-1 (Adapter-related protein complex 4 epsilon-1 subunit) (Epsilon subunit of AP-4) (AP-4 adapter complex epsilon subunit). | |||||
|
AP2A1_MOUSE
|
||||||
| NC score | 0.108116 (rank : 24) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P17426 | Gene names | Ap2a1, Adtaa, Clapa1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-2 complex subunit alpha-1 (Adapter-related protein complex 2 alpha- 1 subunit) (Alpha-adaptin A) (Adaptor protein complex AP-2 alpha-1 subunit) (Clathrin assembly protein complex 2 alpha-A large chain) (100 kDa coated vesicle protein A) (Plasma membrane adaptor HA2/AP2 adaptin alpha A subunit). | |||||
|
AP2A2_MOUSE
|
||||||
| NC score | 0.107941 (rank : 25) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P17427, Q921V0 | Gene names | Ap2a2, Adtab | |||
|
Domain Architecture |

|
|||||
| Description | AP-2 complex subunit alpha-2 (Adapter-related protein complex 2 alpha- 2 subunit) (Alpha-adaptin C) (Adaptor protein complex AP-2 alpha-2 subunit) (Clathrin assembly protein complex 2 alpha-C large chain) (100 kDa coated vesicle protein C) (Plasma membrane adaptor HA2/AP2 adaptin alpha C subunit). | |||||
|
AP2A2_HUMAN
|
||||||
| NC score | 0.107777 (rank : 26) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O94973, O75403, Q96SI8 | Gene names | AP2A2, ADTAB, KIAA0899 | |||
|
Domain Architecture |

|
|||||
| Description | AP-2 complex subunit alpha-2 (Adapter-related protein complex 2 alpha- 2 subunit) (Alpha-adaptin C) (Adaptor protein complex AP-2 alpha-2 subunit) (Clathrin assembly protein complex 2 alpha-C large chain) (100 kDa coated vesicle protein C) (Plasma membrane adaptor HA2/AP2 adaptin alpha C subunit) (Huntingtin-interacting protein HYPJ). | |||||
|
AP3D1_MOUSE
|
||||||
| NC score | 0.100562 (rank : 27) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O54774 | Gene names | Ap3d1, Ap3d | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-3 complex subunit delta-1 (Adapter-related protein complex 3 subunit delta-1) (Delta-adaptin 3) (AP-3 complex subunit delta) (Delta-adaptin) (mBLVR1). | |||||
|
AP3D1_HUMAN
|
||||||
| NC score | 0.095340 (rank : 28) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O14617, O00202, O75262, Q59HF5, Q96G11, Q9H3C6 | Gene names | AP3D1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit delta-1 (Adapter-related protein complex 3 subunit delta-1) (Delta-adaptin 3) (AP-3 complex subunit delta) (Delta-adaptin). | |||||
|
HNF1A_HUMAN
|
||||||
| NC score | 0.051917 (rank : 29) | θ value | 0.163984 (rank : 23) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P20823, Q99861 | Gene names | TCF1, HNF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-alpha (HNF-1A) (Liver-specific transcription factor LF-B1) (LFB1) (Transcription factor 1) (TCF-1). | |||||
|
AP2B1_MOUSE
|
||||||
| NC score | 0.049634 (rank : 30) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9DBG3, Q80XJ4 | Gene names | Ap2b1, Clapb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-2 complex subunit beta-1 (Adapter-related protein complex 2 beta-1 subunit) (Beta-adaptin) (Plasma membrane adaptor HA2/AP2 adaptin beta subunit) (Clathrin assembly protein complex 2 beta large chain) (AP105B). | |||||
|
AP2B1_HUMAN
|
||||||
| NC score | 0.049631 (rank : 31) | θ value | 5.27518 (rank : 48) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P63010, P21851, Q96J19 | Gene names | AP2B1, CLAPB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-2 complex subunit beta-1 (Adapter-related protein complex 2 beta-1 subunit) (Beta-adaptin) (Plasma membrane adaptor HA2/AP2 adaptin beta subunit) (Clathrin assembly protein complex 2 beta large chain) (AP105B). | |||||
|
SRBP1_HUMAN
|
||||||
| NC score | 0.048823 (rank : 32) | θ value | 0.0961366 (rank : 21) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
ALEX_MOUSE
|
||||||
| NC score | 0.047946 (rank : 33) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6R0H6, Q8BIR3 | Gene names | Gnas, Gnas1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ALEX (Alternative gene product encoded by XL-exon). | |||||
|
HNF1A_MOUSE
|
||||||
| NC score | 0.045457 (rank : 34) | θ value | 0.21417 (rank : 24) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P22361 | Gene names | Tcf1, Hnf-1, Hnf-1a, Hnf1a | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-alpha (HNF-1A) (Liver-specific transcription factor LF-B1) (LFB1). | |||||
|
RUNX2_MOUSE
|
||||||
| NC score | 0.031732 (rank : 35) | θ value | 3.0926 (rank : 44) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q08775, O35183, Q08776, Q9JLN0, Q9QUQ6, Q9QY29, Q9R0U4, Q9Z2J7 | Gene names | Runx2, Aml3, Cbfa1, Osf2, Pebp2a | |||
|
Domain Architecture |

|
|||||
| Description | Runt-related transcription factor 2 (Core-binding factor, alpha 1 subunit) (CBF-alpha 1) (Acute myeloid leukemia 3 protein) (Oncogene AML-3) (Polyomavirus enhancer-binding protein 2 alpha A subunit) (PEBP2-alpha A) (PEA2-alpha A) (SL3-3 enhancer factor 1 alpha A subunit) (SL3/AKV core-binding factor alpha A subunit) (Osteoblast- specific transcription factor 2) (OSF-2). | |||||
|
RUNX2_HUMAN
|
||||||
| NC score | 0.031620 (rank : 36) | θ value | 3.0926 (rank : 43) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13950, O14614, O14615, O95181 | Gene names | RUNX2, AML3, CBFA1, OSF2, PEBP2A | |||
|
Domain Architecture |

|
|||||
| Description | Runt-related transcription factor 2 (Core-binding factor, alpha 1 subunit) (CBF-alpha 1) (Acute myeloid leukemia 3 protein) (Oncogene AML-3) (Polyomavirus enhancer-binding protein 2 alpha A subunit) (PEBP2-alpha A) (PEA2-alpha A) (SL3-3 enhancer factor 1 alpha A subunit) (SL3/AKV core-binding factor alpha A subunit) (Osteoblast- specific transcription factor 2) (OSF-2). | |||||
|
RAPH1_HUMAN
|
||||||
| NC score | 0.024654 (rank : 37) | θ value | 0.62314 (rank : 27) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
T240L_MOUSE
|
||||||
| NC score | 0.024647 (rank : 38) | θ value | 2.36792 (rank : 40) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6JPI3, Q3TRF4, Q3UQI8, Q80TM3, Q80WQ0 | Gene names | Thrap2, Kiaa1025, Trap240l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 2 (Thyroid hormone receptor-associated protein complex 240 kDa component-like). | |||||
|
T240L_HUMAN
|
||||||
| NC score | 0.024448 (rank : 39) | θ value | 2.36792 (rank : 39) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q71F56, Q68DN4, Q9H8C0, Q9NSY9, Q9UFD8, Q9UPX5 | Gene names | THRAP2, KIAA1025, PROSIT240, TRAP240L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 2 (Thyroid hormone receptor-associated protein complex 240 kDa component-like). | |||||
|
CN092_MOUSE
|
||||||
| NC score | 0.023309 (rank : 40) | θ value | 6.88961 (rank : 55) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BU11, Q80UI2, Q99PN9, Q9CS16 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Epidermal Langerhans cell protein LCP1. | |||||
|
IRX4_MOUSE
|
||||||
| NC score | 0.023125 (rank : 41) | θ value | 0.47712 (rank : 26) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QY61 | Gene names | Irx4, Irxa3 | |||
|
Domain Architecture |

|
|||||
| Description | Iroquois-class homeodomain protein IRX-4 (Iroquois homeobox protein 4) (Homeodomain protein IRXA3). | |||||
|
CCD96_MOUSE
|
||||||
| NC score | 0.022615 (rank : 42) | θ value | 1.38821 (rank : 30) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9CR92 | Gene names | Ccdc96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
SMCA4_HUMAN
|
||||||
| NC score | 0.022361 (rank : 43) | θ value | 0.365318 (rank : 25) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P51532, O95052, Q9HBD3 | Gene names | SMARCA4, BRG1, SNF2B, SNF2L4 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L4 (EC 3.6.1.-) (ATP- dependent helicase SMARCA4) (SNF2-beta) (BRG-1 protein) (Mitotic growth and transcription activator) (Brahma protein homolog 1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 4). | |||||
|
K0555_HUMAN
|
||||||
| NC score | 0.022184 (rank : 44) | θ value | 0.125558 (rank : 22) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 880 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96AA8, O60302 | Gene names | KIAA0555 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0555. | |||||
|
CCDC7_HUMAN
|
||||||
| NC score | 0.022133 (rank : 45) | θ value | 2.36792 (rank : 37) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 253 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96M83, Q8IVQ0, Q8NEQ0 | Gene names | CCDC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 7. | |||||
|
ITSN2_HUMAN
|
||||||
| NC score | 0.020473 (rank : 46) | θ value | 4.03905 (rank : 45) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1173 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NZM3, O95062, Q15812, Q9HAK4, Q9NXE6, Q9NYG0, Q9NZM2, Q9ULG4 | Gene names | ITSN2, KIAA1256, SH3D1B, SWAP | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-2 (SH3 domain-containing protein 1B) (SH3P18) (SH3P18-like WASP-associated protein). | |||||
|
LDB3_HUMAN
|
||||||
| NC score | 0.018824 (rank : 47) | θ value | 1.38821 (rank : 31) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75112, Q5K6N9, Q5K6P0, Q5K6P1, Q96FH2, Q9Y4Z3, Q9Y4Z4, Q9Y4Z5 | Gene names | LDB3, KIAA0613, ZASP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LIM domain-binding protein 3 (Z-band alternatively spliced PDZ-motif protein) (Protein cypher). | |||||
|
SWP70_MOUSE
|
||||||
| NC score | 0.017765 (rank : 48) | θ value | 0.813845 (rank : 29) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1097 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6A028, O88443, Q3TQR6, Q3UCA3, Q6P1D0 | Gene names | Swap70, Kiaa0640 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Switch-associated protein 70 (SWAP-70). | |||||
|
CEP68_MOUSE
|
||||||
| NC score | 0.014816 (rank : 49) | θ value | 5.27518 (rank : 50) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C0D9, Q3US49, Q5SW85, Q8R2V0 | Gene names | Cep68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 68 kDa (Cep68 protein). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.014554 (rank : 50) | θ value | 4.03905 (rank : 47) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
CE290_HUMAN
|
||||||
| NC score | 0.014044 (rank : 51) | θ value | 8.99809 (rank : 57) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
RFIP3_HUMAN
|
||||||
| NC score | 0.013471 (rank : 52) | θ value | 8.99809 (rank : 59) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1055 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O75154, Q4VXV7, Q9H155, Q9H1G0, Q9NUI0 | Gene names | RAB11FIP3, KIAA0665 | |||
|
Domain Architecture |

|
|||||
| Description | Rab11 family-interacting protein 3 (Rab11-FIP3) (EF hands-containing Rab-interacting protein) (Eferin). | |||||
|
LDB3_MOUSE
|
||||||
| NC score | 0.012600 (rank : 53) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 510 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9JKS4, Q6A038, Q811P2, Q811P3, Q811P4, Q811P5, Q9D130, Q9JKS3, Q9R0Z1, Q9WVH1, Q9WVH2 | Gene names | Ldb3, Kiaa0613 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LIM domain-binding protein 3 (Z-band alternatively spliced PDZ-motif protein) (Protein cypher) (Protein oracle). | |||||
|
SPTN2_HUMAN
|
||||||
| NC score | 0.011889 (rank : 54) | θ value | 2.36792 (rank : 38) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 535 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O15020, O14872, O14873 | Gene names | SPTBN2, KIAA0302 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 2 (Spectrin, non-erythroid beta chain 2) (Beta-III spectrin). | |||||
|
SYNE1_HUMAN
|
||||||
| NC score | 0.011434 (rank : 55) | θ value | 1.38821 (rank : 32) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
GNDS_HUMAN
|
||||||
| NC score | 0.010598 (rank : 56) | θ value | 3.0926 (rank : 41) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q12967, Q9HAX7, Q9HAY1, Q9HCT1 | Gene names | RALGDS, RGF | |||
|
Domain Architecture |

|
|||||
| Description | Ral guanine nucleotide dissociation stimulator (RalGEF) (RalGDS). | |||||
|
CRBB1_HUMAN
|
||||||
| NC score | 0.009532 (rank : 57) | θ value | 5.27518 (rank : 51) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P53674 | Gene names | CRYBB1 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin B1. | |||||
|
LRC8C_MOUSE
|
||||||
| NC score | 0.007860 (rank : 58) | θ value | 3.0926 (rank : 42) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R502, Q3TEP9, Q8C296, Q8R0N7, Q8R3G5 | Gene names | Lrrc8c, Ad158 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 8C (Protein AD158). | |||||
|
ABL1_MOUSE
|
||||||
| NC score | 0.006093 (rank : 59) | θ value | 0.813845 (rank : 28) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1178 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P00520, P97896, Q61252, Q61253, Q61254, Q61255, Q61256, Q61257, Q61258, Q61259, Q61260, Q61261, Q6PCM5, Q8C1X4 | Gene names | Abl1, Abl | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene tyrosine-protein kinase ABL1 (EC 2.7.10.2) (p150) (c- ABL) (Abelson murine leukemia viral oncogene homolog 1). | |||||
|
MRP2_MOUSE
|
||||||
| NC score | 0.005946 (rank : 60) | θ value | 5.27518 (rank : 52) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VI47 | Gene names | Abcc2 | |||
|
Domain Architecture |

|
|||||
| Description | Canalicular multispecific organic anion transporter 1 (ATP-binding cassette sub-family C member 2). | |||||
|
SORC3_HUMAN
|
||||||
| NC score | 0.005899 (rank : 61) | θ value | 6.88961 (rank : 56) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UPU3, Q5VXF9, Q9NQJ2 | Gene names | SORCS3, KIAA1059 | |||
|
Domain Architecture |

|
|||||
| Description | VPS10 domain-containing receptor SorCS3 precursor. | |||||
|
COBA2_HUMAN
|
||||||
| NC score | 0.005837 (rank : 62) | θ value | 1.81305 (rank : 33) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P13942, Q07751, Q13271, Q13272, Q13273, Q99866, Q9UIP9 | Gene names | COL11A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
PEX1_HUMAN
|
||||||
| NC score | 0.005526 (rank : 63) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O43933, Q96S71, Q96S72, Q96S73, Q99994 | Gene names | PEX1 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisome biogenesis factor 1 (Peroxin-1) (Peroxisome biogenesis disorder protein 1). | |||||
|
SREC2_MOUSE
|
||||||
| NC score | 0.004932 (rank : 64) | θ value | 1.81305 (rank : 35) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 605 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P59222 | Gene names | Scarf2, Srec2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor class F member 2 precursor (Scavenger receptor expressed by endothelial cells 2 protein) (SREC-II). | |||||
|
CD248_MOUSE
|
||||||
| NC score | 0.004769 (rank : 65) | θ value | 6.88961 (rank : 54) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 578 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q91V98, Q3UMV6, Q91ZV1 | Gene names | Cd248, Tem1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endosialin precursor (Tumor endothelial marker 1) (CD248 antigen). | |||||
|
KLC4_MOUSE
|
||||||
| NC score | 0.003923 (rank : 66) | θ value | 8.99809 (rank : 58) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 336 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9DBS5, Q3TCC5 | Gene names | Klc4, Knsl8 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin light chain 4 (KLC 4) (Kinesin-like protein 8). | |||||
|
SPB6_HUMAN
|
||||||
| NC score | 0.001366 (rank : 67) | θ value | 8.99809 (rank : 60) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P35237, Q96J44 | Gene names | SERPINB6, PI6, PTI | |||
|
Domain Architecture |

|
|||||
| Description | Serpin B6 (Placental thrombin inhibitor) (Cytoplasmic antiproteinase) (CAP) (Protease inhibitor 6) (PI-6). | |||||
|
PASK_HUMAN
|
||||||
| NC score | 0.000872 (rank : 68) | θ value | 1.81305 (rank : 34) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 857 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96RG2, Q86XH6, Q99763, Q9UFR7 | Gene names | PASK, KIAA0135 | |||
|
Domain Architecture |

|
|||||
| Description | PAS domain-containing serine/threonine-protein kinase (EC 2.7.11.1) (PAS-kinase) (PASKIN) (hPASK). | |||||