Please be patient as the page loads
|
PWWP2_HUMAN
|
||||||
| SwissProt Accessions | Q6NUJ5, Q5SZI0, Q6ZQX5, Q96F43 | Gene names | PWWP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PWWP domain-containing protein 2. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PWWP2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q6NUJ5, Q5SZI0, Q6ZQX5, Q96F43 | Gene names | PWWP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PWWP domain-containing protein 2. | |||||
|
DNM3A_HUMAN
|
||||||
| θ value | 1.133e-10 (rank : 2) | NC score | 0.467719 (rank : 2) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y6K1, Q8WXU9 | Gene names | DNMT3A | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3A (EC 2.1.1.37) (Dnmt3a) (DNA methyltransferase HsaIIIA) (DNA MTase HsaIIIA) (M.HsaIIIA). | |||||
|
DNM3A_MOUSE
|
||||||
| θ value | 1.133e-10 (rank : 3) | NC score | 0.466598 (rank : 3) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O88508, Q3UH24, Q8CJ60, Q922J0, Q9CSE1 | Gene names | Dnmt3a | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3A (EC 2.1.1.37) (Dnmt3a) (DNA methyltransferase MmuIIIA) (DNA MTase MmuIIIA) (M.MmuIIIA). | |||||
|
DNM3B_HUMAN
|
||||||
| θ value | 2.13673e-09 (rank : 4) | NC score | 0.463424 (rank : 4) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UBC3, Q9UBD4, Q9UJQ5, Q9UKA6, Q9UNE5, Q9Y5R9, Q9Y5S0 | Gene names | DNMT3B | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3B (EC 2.1.1.37) (Dnmt3b) (DNA methyltransferase HsaIIIB) (DNA MTase HsaIIIB) (M.HsaIIIB). | |||||
|
DNM3B_MOUSE
|
||||||
| θ value | 3.64472e-09 (rank : 5) | NC score | 0.461210 (rank : 5) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O88509, O88510, O88511 | Gene names | Dnmt3b | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3B (EC 2.1.1.37) (Dnmt3b) (DNA methyltransferase MmuIIIB) (DNA MTase MmuIIIB) (M.MmuIIIB). | |||||
|
PSIP1_HUMAN
|
||||||
| θ value | 0.000121331 (rank : 6) | NC score | 0.319844 (rank : 11) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O75475, O00256, O95368, Q6P391, Q86YB9, Q9NZI3, Q9UER6 | Gene names | PSIP1, DFS70, LEDGF, PSIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PC4 and SFRS1-interacting protein (Lens epithelium-derived growth factor) (Transcriptional coactivator p75/p52) (Dense fine speckles 70 kDa protein) (DFS 70) (CLL-associated antigen KW-7). | |||||
|
HDGR3_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 7) | NC score | 0.374703 (rank : 7) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y3E1 | Gene names | HDGFRP3, HDGF2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatoma-derived growth factor-related protein 3 (HRP-3) (Hepatoma- derived growth factor 2). | |||||
|
HDGR3_MOUSE
|
||||||
| θ value | 0.000158464 (rank : 8) | NC score | 0.375835 (rank : 6) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9JMG7, Q3TRX2, Q8BQ69, Q8BR62, Q9D2M7 | Gene names | Hdgfrp3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatoma-derived growth factor-related protein 3 (HRP-3). | |||||
|
PSIP1_MOUSE
|
||||||
| θ value | 0.000158464 (rank : 9) | NC score | 0.328047 (rank : 10) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q99JF8, Q3TEJ7, Q3TTD7, Q3UTA1, Q80WQ7, Q99JF7, Q99LR4, Q9CT03 | Gene names | Psip1, Ledgf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PC4 and SFRS1-interacting protein (Lens epithelium-derived growth factor) (mLEDGF). | |||||
|
MSH6_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 10) | NC score | 0.203315 (rank : 16) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P54276, O54710 | Gene names | Msh6, Gtmbp | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein MSH6 (MutS-alpha 160 kDa subunit) (G/T mismatch-binding protein) (GTBP) (GTMBP) (p160). | |||||
|
TARA_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 11) | NC score | 0.073832 (rank : 26) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
HDGF_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 12) | NC score | 0.347747 (rank : 8) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P51858, Q5SZ09 | Gene names | HDGF, HMG1L2 | |||
|
Domain Architecture |
|
|||||
| Description | Hepatoma-derived growth factor (HDGF) (High-mobility group protein 1- like 2) (HMG-1L2). | |||||
|
HDGF_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 13) | NC score | 0.335397 (rank : 9) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P51859, Q8BPG7, Q9CYA4, Q9JK87 | Gene names | Hdgf, Tdrm1 | |||
|
Domain Architecture |
|
|||||
| Description | Hepatoma-derived growth factor (HDGF). | |||||
|
MSH6_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 14) | NC score | 0.195746 (rank : 17) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P52701, O43706, O43917, Q8TCX4, Q9BTB5 | Gene names | MSH6, GTBP | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein MSH6 (MutS-alpha 160 kDa subunit) (G/T mismatch-binding protein) (GTBP) (GTMBP) (p160). | |||||
|
ZCPW1_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 15) | NC score | 0.205481 (rank : 15) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6IR42 | Gene names | Zcwpw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CW-type PWWP domain protein 1 homolog. | |||||
|
JPH2_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 16) | NC score | 0.063463 (rank : 28) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9BR39, O95913, Q5JY74, Q9UJN4 | Gene names | JPH2, JP2 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 17) | NC score | 0.054524 (rank : 32) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 18) | NC score | 0.047610 (rank : 41) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
NSD1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 19) | NC score | 0.127338 (rank : 19) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
GLI2_HUMAN
|
||||||
| θ value | 0.125558 (rank : 20) | NC score | 0.012922 (rank : 76) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 971 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P10070, O60252, O60253, O60254, O60255, Q15590, Q15591 | Gene names | GLI2, THP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLI2 (Tax helper protein). | |||||
|
TAF1C_MOUSE
|
||||||
| θ value | 0.125558 (rank : 21) | NC score | 0.080909 (rank : 24) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PDZ2, P97359, Q3U3B6, Q8BN54 | Gene names | Taf1c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA box-binding protein-associated factor, RNA polymerase I, subunit C (TATA box-binding protein-associated factor 1C) (TBP-associated factor 1C) (TBP-associated factor RNA polymerase I 95 kDa) (TAFI95). | |||||
|
ZCPW1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 22) | NC score | 0.210265 (rank : 14) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9H0M4, Q8NA98, Q9BUD0, Q9NWF7 | Gene names | ZCWPW1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CW-type PWWP domain protein 1. | |||||
|
MBD5_HUMAN
|
||||||
| θ value | 0.279714 (rank : 23) | NC score | 0.138878 (rank : 18) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9P267, Q9NUV6 | Gene names | MBD5, KIAA1461 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 5 (Methyl-CpG-binding protein MBD5). | |||||
|
PKCB1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 24) | NC score | 0.123967 (rank : 22) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
SC24C_HUMAN
|
||||||
| θ value | 0.279714 (rank : 25) | NC score | 0.051240 (rank : 34) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P53992, Q8WV25 | Gene names | SEC24C, KIAA0079 | |||
|
Domain Architecture |

|
|||||
| Description | Protein transport protein Sec24C (SEC24-related protein C). | |||||
|
DDX51_HUMAN
|
||||||
| θ value | 0.365318 (rank : 26) | NC score | 0.029501 (rank : 54) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
SH3K1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 27) | NC score | 0.050009 (rank : 35) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96B97, Q8IWX6, Q8IX98, Q96RN4, Q9NYR0 | Gene names | SH3KBP1, CIN85 | |||
|
Domain Architecture |

|
|||||
| Description | SH3-domain kinase-binding protein 1 (Cbl-interacting protein of 85 kDa) (Human Src-family kinase-binding protein 1) (HSB-1) (CD2-binding protein 3) (CD2BP3). | |||||
|
NSD1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 28) | NC score | 0.104737 (rank : 23) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
SET1A_HUMAN
|
||||||
| θ value | 0.62314 (rank : 29) | NC score | 0.075227 (rank : 25) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
TXND2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 30) | NC score | 0.067233 (rank : 27) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
IF2B2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 31) | NC score | 0.037389 (rank : 45) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y6M1 | Gene names | IGF2BP2, IMP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin-like growth factor 2 mRNA-binding protein 2 (IGF-II mRNA- binding protein 2) (IMP-2) (Hepatocellular carcinoma autoantigen p62). | |||||
|
IF4G3_MOUSE
|
||||||
| θ value | 0.813845 (rank : 32) | NC score | 0.049154 (rank : 37) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q80XI3, Q80Y69, Q8BJ68, Q8BQF3, Q8C6Q3, Q8CIH0 | Gene names | Eif4g3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 3 (eIF-4-gamma 3) (eIF-4G 3) (eIF4G 3) (eIF-4-gamma II) (eIF4GII). | |||||
|
PGCA_MOUSE
|
||||||
| θ value | 0.813845 (rank : 33) | NC score | 0.027066 (rank : 56) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 643 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61282, Q64021 | Gene names | Agc1, Agc | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP). | |||||
|
RABX5_HUMAN
|
||||||
| θ value | 0.813845 (rank : 34) | NC score | 0.048177 (rank : 39) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UJ41 | Gene names | RABGEF1, RABEX5 | |||
|
Domain Architecture |

|
|||||
| Description | Rab5 GDP/GTP exchange factor (Rabex-5) (Rabaptin-5-associated exchange factor for Rab5). | |||||
|
XRCC1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 35) | NC score | 0.060504 (rank : 30) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q60596, Q3THC5, Q5U435, Q7TNQ5 | Gene names | Xrcc1, Xrcc-1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein XRCC1 (X-ray repair cross-complementing protein 1). | |||||
|
BLNK_MOUSE
|
||||||
| θ value | 1.06291 (rank : 36) | NC score | 0.049575 (rank : 36) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9QUN3, O88504 | Gene names | Blnk, Bash, Ly57, Slp65 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell linker protein (Cytoplasmic adapter protein) (B-cell adapter containing SH2 domain protein) (B-cell adapter containing Src homology 2 domain protein) (Src homology 2 domain-containing leukocyte protein of 65 kDa) (Slp-65) (Lymphocyte antigen 57). | |||||
|
SH3K1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 37) | NC score | 0.045859 (rank : 42) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8R550, Q8CAL8, Q8CEF6, Q8R545, Q8R546, Q8R547, Q8R548, Q8R549, Q8R551, Q9CTQ9, Q9DC14, Q9JKC3 | Gene names | Sh3kbp1, Ruk, Seta | |||
|
Domain Architecture |

|
|||||
| Description | SH3-domain kinase-binding protein 1 (SH3-containing, expressed in tumorigenic astrocytes) (Regulator of ubiquitous kinase) (Ruk). | |||||
|
TITIN_HUMAN
|
||||||
| θ value | 1.06291 (rank : 38) | NC score | 0.017444 (rank : 70) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
ADA19_HUMAN
|
||||||
| θ value | 1.38821 (rank : 39) | NC score | 0.014104 (rank : 74) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H013, Q9BZL5, Q9UHP2 | Gene names | ADAM19, MLTNB | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta) (Metalloprotease and disintegrin dentritic antigen marker) (MADDAM). | |||||
|
ANR11_HUMAN
|
||||||
| θ value | 1.38821 (rank : 40) | NC score | 0.030298 (rank : 51) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
SIVA_HUMAN
|
||||||
| θ value | 1.38821 (rank : 41) | NC score | 0.057558 (rank : 31) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15304, Q96P98, Q9UPD6 | Gene names | SIVA | |||
|
Domain Architecture |
|
|||||
| Description | Apoptosis regulatory protein Siva (CD27-binding protein) (CD27BP). | |||||
|
SOX13_MOUSE
|
||||||
| θ value | 1.38821 (rank : 42) | NC score | 0.026123 (rank : 60) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 495 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q04891, Q922L3 | Gene names | Sox13, Sox-13 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-13 protein. | |||||
|
ZMY11_HUMAN
|
||||||
| θ value | 1.38821 (rank : 43) | NC score | 0.124808 (rank : 21) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q15326 | Gene names | ZMYND11, BS69 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11 (Adenovirus 5 E1A- binding protein) (BS69 protein). | |||||
|
ZMY11_MOUSE
|
||||||
| θ value | 1.38821 (rank : 44) | NC score | 0.125490 (rank : 20) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8R5C8, Q8C155 | Gene names | Zmynd11 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11. | |||||
|
CS007_MOUSE
|
||||||
| θ value | 1.81305 (rank : 45) | NC score | 0.048878 (rank : 38) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 464 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6ZPZ3 | Gene names | Kiaa1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7 homolog. | |||||
|
JHD2B_HUMAN
|
||||||
| θ value | 1.81305 (rank : 46) | NC score | 0.026361 (rank : 58) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7LBC6, Q9BVH6, Q9BW93, Q9BZ52, Q9NYF4, Q9UPS0 | Gene names | JMJD1B, C5orf7, JHDM2B, KIAA1082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 2B (EC 1.14.11.-) (Jumonji domain-containing protein 1B) (Nuclear protein 5qNCA). | |||||
|
PUNC_MOUSE
|
||||||
| θ value | 1.81305 (rank : 47) | NC score | 0.017622 (rank : 69) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 501 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BQC3, O70246, Q792T2, Q9Z2S6 | Gene names | Punc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative neuronal cell adhesion molecule precursor. | |||||
|
ROBO4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 48) | NC score | 0.026605 (rank : 57) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8C310, Q9DBW1 | Gene names | Robo4 | |||
|
Domain Architecture |
|
|||||
| Description | Roundabout homolog 4 precursor. | |||||
|
UBP51_HUMAN
|
||||||
| θ value | 1.81305 (rank : 49) | NC score | 0.019895 (rank : 63) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q70EK9, Q8IWJ8 | Gene names | USP51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 51 (EC 3.1.2.15) (Ubiquitin thioesterase 51) (Ubiquitin-specific-processing protease 51) (Deubiquitinating enzyme 51). | |||||
|
GP176_MOUSE
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.011793 (rank : 77) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80WT4, Q80UC4 | Gene names | Gpr176, Agr9, Gm1012 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 176 (G-protein coupled receptor AGR9). | |||||
|
RFX3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.023822 (rank : 61) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P48380, Q5JTL7, Q5JTL8, Q6NW13, Q8WTU4, Q95HL5, Q95HL6 | Gene names | RFX3 | |||
|
Domain Architecture |
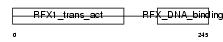
|
|||||
| Description | Transcription factor RFX3. | |||||
|
TF65_HUMAN
|
||||||
| θ value | 2.36792 (rank : 52) | NC score | 0.026133 (rank : 59) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q04206, Q6SLK1 | Gene names | RELA, NFKB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor p65 (Nuclear factor NF-kappa-B p65 subunit). | |||||
|
ATN1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 53) | NC score | 0.048001 (rank : 40) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
NOP56_HUMAN
|
||||||
| θ value | 3.0926 (rank : 54) | NC score | 0.033480 (rank : 47) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O00567, Q9NQ05 | Gene names | NOL5A, NOP56 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar protein Nop56 (Nucleolar protein 5A). | |||||
|
REPS2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.033994 (rank : 46) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8NFH8, O43428, Q8NFI5 | Gene names | REPS2, POB1 | |||
|
Domain Architecture |

|
|||||
| Description | RalBP1-associated Eps domain-containing protein 2 (RalBP1-interacting protein 2) (Partner of RalBP1). | |||||
|
BRE1B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 56) | NC score | 0.010531 (rank : 79) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1040 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O75150, Q6AHZ6, Q6N005, Q7L3T6, Q8N615, Q96T18, Q9BSV9, Q9HC82 | Gene names | RNF40, BRE1B, KIAA0661 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase BRE1B (EC 6.3.2.-) (BRE1-B) (RING finger protein 40) (95 kDa retinoblastoma-associated protein) (RBP95). | |||||
|
JIP3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 57) | NC score | 0.017753 (rank : 68) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1039 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UPT6, Q96RY4, Q9H4I4, Q9H7P1, Q9NUG0 | Gene names | MAPK8IP3, JIP3, KIAA1066 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 3 (JNK-interacting protein 3) (JIP-3) (JNK MAP kinase scaffold protein 3) (Mitogen- activated protein kinase 8-interacting protein 3). | |||||
|
TENA_MOUSE
|
||||||
| θ value | 4.03905 (rank : 58) | NC score | 0.010214 (rank : 80) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80YX1, Q64706, Q80YX2 | Gene names | Tnc, Hxb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tenascin precursor (TN) (Tenascin-C) (TN-C) (Hexabrachion). | |||||
|
ULK1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 59) | NC score | 0.003429 (rank : 83) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1214 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O70405, Q6PGB2 | Gene names | Ulk1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ULK1 (EC 2.7.11.1) (Unc-51-like kinase 1) (Serine/threonine-protein kinase Unc51.1). | |||||
|
DCX_HUMAN
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.018429 (rank : 65) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O43602, O43911 | Gene names | DCX, DBCN, LISX | |||
|
Domain Architecture |
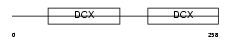
|
|||||
| Description | Neuronal migration protein doublecortin (Lissencephalin-X) (Lis-X) (Doublin). | |||||
|
PDLI4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.015143 (rank : 73) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P50479, Q96AT8, Q9BTW8, Q9Y292 | Gene names | PDLIM4, RIL | |||
|
Domain Architecture |

|
|||||
| Description | PDZ and LIM domain protein 4 (LIM protein RIL) (Reversion-induced LIM protein). | |||||
|
RT35_MOUSE
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.029781 (rank : 53) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BJZ4, Q8VCG7 | Gene names | Mrps35 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 28S ribosomal protein S35, mitochondrial precursor (S35mt) (MRP-S35). | |||||
|
SASH1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 63) | NC score | 0.030352 (rank : 50) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P59808 | Gene names | Sash1 | |||
|
Domain Architecture |

|
|||||
| Description | SAM and SH3 domain-containing protein 1. | |||||
|
T22D4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.028948 (rank : 55) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y3Q8 | Gene names | TSC22D4, TILZ2 | |||
|
Domain Architecture |

|
|||||
| Description | TSC22 domain family protein 4 (TSC22-related-inducible leucine zipper protein 2) (Tsc-22-like protein THG-1). | |||||
|
ZN318_HUMAN
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.030856 (rank : 49) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5VUA4, O94796, Q4G0E4, Q8NEM6, Q9UNU8, Q9Y2W9 | Gene names | ZNF318 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 318 (Endocrine regulatory protein). | |||||
|
ADCY9_HUMAN
|
||||||
| θ value | 6.88961 (rank : 66) | NC score | 0.010851 (rank : 78) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O60503, O60273, Q4ZHT9, Q4ZIR5, Q9BWT4, Q9UGP2 | Gene names | ADCY9, KIAA0520 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 9 (EC 4.6.1.1) (Adenylate cyclase type IX) (ATP pyrophosphate-lyase 9) (Adenylyl cyclase 9). | |||||
|
ETV3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.018313 (rank : 66) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
FZD6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.007955 (rank : 82) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61089 | Gene names | Fzd6 | |||
|
Domain Architecture |

|
|||||
| Description | Frizzled-6 precursor (Fz-6) (mFz6). | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.041027 (rank : 44) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
VGFR3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | 0.002890 (rank : 84) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1056 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P35917 | Gene names | Flt4, Flt-4 | |||
|
Domain Architecture |

|
|||||
| Description | Vascular endothelial growth factor receptor 3 precursor (EC 2.7.10.1) (VEGFR-3) (Tyrosine-protein kinase receptor FLT4). | |||||
|
CBP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 71) | NC score | 0.029976 (rank : 52) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CEL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.014091 (rank : 75) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
CS029_MOUSE
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.020348 (rank : 62) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9CS00 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf29 homolog. | |||||
|
ELL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.016789 (rank : 71) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P55199 | Gene names | ELL, C19orf17 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II elongation factor ELL (Eleven-nineteen lysine-rich leukemia protein). | |||||
|
IF4G1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.041709 (rank : 43) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q04637, O43177, O95066, Q5HYG0, Q6ZN21, Q8N102 | Gene names | EIF4G1, EIF4G, EIF4GI | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1) (p220). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.031500 (rank : 48) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
PTN23_HUMAN
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.018520 (rank : 64) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 646 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9H3S7, Q7KZF8, Q8N6Z5, Q9BSR5, Q9P257, Q9UG03, Q9UMZ4 | Gene names | PTPN23, KIAA1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48) (His- domain-containing protein tyrosine phosphatase) (HD-PTP) (Protein tyrosine phosphatase TD14) (PTP-TD14). | |||||
|
RIMB1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.017838 (rank : 67) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
TENX_HUMAN
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.009399 (rank : 81) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 789 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P22105, P78530, P78531, Q08424, Q9UMG7 | Gene names | TNXB, HXBL, TNX, XB | |||
|
Domain Architecture |

|
|||||
| Description | Tenascin-X precursor (TN-X) (Hexabrachion-like protein). | |||||
|
ZN533_MOUSE
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.016478 (rank : 72) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BXJ8, Q8BWQ7 | Gene names | Znf533, Zfp533 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 533. | |||||
|
ATRX_MOUSE
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.054093 (rank : 33) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61687 | Gene names | Atrx, Hp1bp2, Xnp | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked nuclear protein) (Heterochromatin protein 2) (HP1 alpha-interacting protein) (HP1-BP38 protein). | |||||
|
DNM3L_HUMAN
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.281135 (rank : 12) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UJW3, Q9BUJ4 | Gene names | DNMT3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA (cytosine-5)-methyltransferase 3-like. | |||||
|
DNM3L_MOUSE
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.277762 (rank : 13) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CWR8 | Gene names | Dnmt3l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA (cytosine-5)-methyltransferase 3-like. | |||||
|
TMAP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.062030 (rank : 29) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P84157 | Gene names | TMAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane anchor protein 1. | |||||
|
PWWP2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q6NUJ5, Q5SZI0, Q6ZQX5, Q96F43 | Gene names | PWWP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PWWP domain-containing protein 2. | |||||
|
DNM3A_HUMAN
|
||||||
| NC score | 0.467719 (rank : 2) | θ value | 1.133e-10 (rank : 2) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y6K1, Q8WXU9 | Gene names | DNMT3A | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3A (EC 2.1.1.37) (Dnmt3a) (DNA methyltransferase HsaIIIA) (DNA MTase HsaIIIA) (M.HsaIIIA). | |||||
|
DNM3A_MOUSE
|
||||||
| NC score | 0.466598 (rank : 3) | θ value | 1.133e-10 (rank : 3) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O88508, Q3UH24, Q8CJ60, Q922J0, Q9CSE1 | Gene names | Dnmt3a | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3A (EC 2.1.1.37) (Dnmt3a) (DNA methyltransferase MmuIIIA) (DNA MTase MmuIIIA) (M.MmuIIIA). | |||||
|
DNM3B_HUMAN
|
||||||
| NC score | 0.463424 (rank : 4) | θ value | 2.13673e-09 (rank : 4) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UBC3, Q9UBD4, Q9UJQ5, Q9UKA6, Q9UNE5, Q9Y5R9, Q9Y5S0 | Gene names | DNMT3B | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3B (EC 2.1.1.37) (Dnmt3b) (DNA methyltransferase HsaIIIB) (DNA MTase HsaIIIB) (M.HsaIIIB). | |||||
|
DNM3B_MOUSE
|
||||||
| NC score | 0.461210 (rank : 5) | θ value | 3.64472e-09 (rank : 5) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O88509, O88510, O88511 | Gene names | Dnmt3b | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3B (EC 2.1.1.37) (Dnmt3b) (DNA methyltransferase MmuIIIB) (DNA MTase MmuIIIB) (M.MmuIIIB). | |||||
|
HDGR3_MOUSE
|
||||||
| NC score | 0.375835 (rank : 6) | θ value | 0.000158464 (rank : 8) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9JMG7, Q3TRX2, Q8BQ69, Q8BR62, Q9D2M7 | Gene names | Hdgfrp3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatoma-derived growth factor-related protein 3 (HRP-3). | |||||
|
HDGR3_HUMAN
|
||||||
| NC score | 0.374703 (rank : 7) | θ value | 0.000158464 (rank : 7) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y3E1 | Gene names | HDGFRP3, HDGF2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatoma-derived growth factor-related protein 3 (HRP-3) (Hepatoma- derived growth factor 2). | |||||
|
HDGF_HUMAN
|
||||||
| NC score | 0.347747 (rank : 8) | θ value | 0.00869519 (rank : 12) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P51858, Q5SZ09 | Gene names | HDGF, HMG1L2 | |||
|
Domain Architecture |

|
|||||
| Description | Hepatoma-derived growth factor (HDGF) (High-mobility group protein 1- like 2) (HMG-1L2). | |||||
|
HDGF_MOUSE
|
||||||
| NC score | 0.335397 (rank : 9) | θ value | 0.00869519 (rank : 13) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P51859, Q8BPG7, Q9CYA4, Q9JK87 | Gene names | Hdgf, Tdrm1 | |||
|
Domain Architecture |

|
|||||
| Description | Hepatoma-derived growth factor (HDGF). | |||||
|
PSIP1_MOUSE
|
||||||
| NC score | 0.328047 (rank : 10) | θ value | 0.000158464 (rank : 9) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q99JF8, Q3TEJ7, Q3TTD7, Q3UTA1, Q80WQ7, Q99JF7, Q99LR4, Q9CT03 | Gene names | Psip1, Ledgf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PC4 and SFRS1-interacting protein (Lens epithelium-derived growth factor) (mLEDGF). | |||||
|
PSIP1_HUMAN
|
||||||
| NC score | 0.319844 (rank : 11) | θ value | 0.000121331 (rank : 6) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O75475, O00256, O95368, Q6P391, Q86YB9, Q9NZI3, Q9UER6 | Gene names | PSIP1, DFS70, LEDGF, PSIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PC4 and SFRS1-interacting protein (Lens epithelium-derived growth factor) (Transcriptional coactivator p75/p52) (Dense fine speckles 70 kDa protein) (DFS 70) (CLL-associated antigen KW-7). | |||||
|
DNM3L_HUMAN
|
||||||
| NC score | 0.281135 (rank : 12) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UJW3, Q9BUJ4 | Gene names | DNMT3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA (cytosine-5)-methyltransferase 3-like. | |||||
|
DNM3L_MOUSE
|
||||||
| NC score | 0.277762 (rank : 13) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CWR8 | Gene names | Dnmt3l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA (cytosine-5)-methyltransferase 3-like. | |||||
|
ZCPW1_HUMAN
|
||||||
| NC score | 0.210265 (rank : 14) | θ value | 0.163984 (rank : 22) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9H0M4, Q8NA98, Q9BUD0, Q9NWF7 | Gene names | ZCWPW1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CW-type PWWP domain protein 1. | |||||
|
ZCPW1_MOUSE
|
||||||
| NC score | 0.205481 (rank : 15) | θ value | 0.0193708 (rank : 15) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6IR42 | Gene names | Zcwpw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CW-type PWWP domain protein 1 homolog. | |||||
|
MSH6_MOUSE
|
||||||
| NC score | 0.203315 (rank : 16) | θ value | 0.00228821 (rank : 10) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P54276, O54710 | Gene names | Msh6, Gtmbp | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein MSH6 (MutS-alpha 160 kDa subunit) (G/T mismatch-binding protein) (GTBP) (GTMBP) (p160). | |||||
|
MSH6_HUMAN
|
||||||
| NC score | 0.195746 (rank : 17) | θ value | 0.0113563 (rank : 14) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P52701, O43706, O43917, Q8TCX4, Q9BTB5 | Gene names | MSH6, GTBP | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein MSH6 (MutS-alpha 160 kDa subunit) (G/T mismatch-binding protein) (GTBP) (GTMBP) (p160). | |||||
|
MBD5_HUMAN
|
||||||
| NC score | 0.138878 (rank : 18) | θ value | 0.279714 (rank : 23) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9P267, Q9NUV6 | Gene names | MBD5, KIAA1461 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 5 (Methyl-CpG-binding protein MBD5). | |||||
|
NSD1_HUMAN
|
||||||
| NC score | 0.127338 (rank : 19) | θ value | 0.0961366 (rank : 19) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
ZMY11_MOUSE
|
||||||
| NC score | 0.125490 (rank : 20) | θ value | 1.38821 (rank : 44) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8R5C8, Q8C155 | Gene names | Zmynd11 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11. | |||||
|
ZMY11_HUMAN
|
||||||
| NC score | 0.124808 (rank : 21) | θ value | 1.38821 (rank : 43) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q15326 | Gene names | ZMYND11, BS69 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11 (Adenovirus 5 E1A- binding protein) (BS69 protein). | |||||
|
PKCB1_HUMAN
|
||||||
| NC score | 0.123967 (rank : 22) | θ value | 0.279714 (rank : 24) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
NSD1_MOUSE
|
||||||
| NC score | 0.104737 (rank : 23) | θ value | 0.47712 (rank : 28) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
TAF1C_MOUSE
|
||||||
| NC score | 0.080909 (rank : 24) | θ value | 0.125558 (rank : 21) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PDZ2, P97359, Q3U3B6, Q8BN54 | Gene names | Taf1c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA box-binding protein-associated factor, RNA polymerase I, subunit C (TATA box-binding protein-associated factor 1C) (TBP-associated factor 1C) (TBP-associated factor RNA polymerase I 95 kDa) (TAFI95). | |||||
|
SET1A_HUMAN
|
||||||
| NC score | 0.075227 (rank : 25) | θ value | 0.62314 (rank : 29) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
TARA_MOUSE
|
||||||
| NC score | 0.073832 (rank : 26) | θ value | 0.00509761 (rank : 11) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
TXND2_HUMAN
|
||||||
| NC score | 0.067233 (rank : 27) | θ value | 0.62314 (rank : 30) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
JPH2_HUMAN
|
||||||
| NC score | 0.063463 (rank : 28) | θ value | 0.0252991 (rank : 16) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9BR39, O95913, Q5JY74, Q9UJN4 | Gene names | JPH2, JP2 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
TMAP1_HUMAN
|
||||||
| NC score | 0.062030 (rank : 29) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P84157 | Gene names | TMAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane anchor protein 1. | |||||
|
XRCC1_MOUSE
|
||||||
| NC score | 0.060504 (rank : 30) | θ value | 0.813845 (rank : 35) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q60596, Q3THC5, Q5U435, Q7TNQ5 | Gene names | Xrcc1, Xrcc-1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein XRCC1 (X-ray repair cross-complementing protein 1). | |||||
|
SIVA_HUMAN
|
||||||
| NC score | 0.057558 (rank : 31) | θ value | 1.38821 (rank : 41) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15304, Q96P98, Q9UPD6 | Gene names | SIVA | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis regulatory protein Siva (CD27-binding protein) (CD27BP). | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.054524 (rank : 32) | θ value | 0.0736092 (rank : 17) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
ATRX_MOUSE
|
||||||
| NC score | 0.054093 (rank : 33) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61687 | Gene names | Atrx, Hp1bp2, Xnp | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked nuclear protein) (Heterochromatin protein 2) (HP1 alpha-interacting protein) (HP1-BP38 protein). | |||||
|
SC24C_HUMAN
|
||||||
| NC score | 0.051240 (rank : 34) | θ value | 0.279714 (rank : 25) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P53992, Q8WV25 | Gene names | SEC24C, KIAA0079 | |||
|
Domain Architecture |

|
|||||
| Description | Protein transport protein Sec24C (SEC24-related protein C). | |||||
|
SH3K1_HUMAN
|
||||||
| NC score | 0.050009 (rank : 35) | θ value | 0.365318 (rank : 27) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96B97, Q8IWX6, Q8IX98, Q96RN4, Q9NYR0 | Gene names | SH3KBP1, CIN85 | |||
|
Domain Architecture |

|
|||||
| Description | SH3-domain kinase-binding protein 1 (Cbl-interacting protein of 85 kDa) (Human Src-family kinase-binding protein 1) (HSB-1) (CD2-binding protein 3) (CD2BP3). | |||||
|
BLNK_MOUSE
|
||||||
| NC score | 0.049575 (rank : 36) | θ value | 1.06291 (rank : 36) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9QUN3, O88504 | Gene names | Blnk, Bash, Ly57, Slp65 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell linker protein (Cytoplasmic adapter protein) (B-cell adapter containing SH2 domain protein) (B-cell adapter containing Src homology 2 domain protein) (Src homology 2 domain-containing leukocyte protein of 65 kDa) (Slp-65) (Lymphocyte antigen 57). | |||||
|
IF4G3_MOUSE
|
||||||
| NC score | 0.049154 (rank : 37) | θ value | 0.813845 (rank : 32) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q80XI3, Q80Y69, Q8BJ68, Q8BQF3, Q8C6Q3, Q8CIH0 | Gene names | Eif4g3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 3 (eIF-4-gamma 3) (eIF-4G 3) (eIF4G 3) (eIF-4-gamma II) (eIF4GII). | |||||
|
CS007_MOUSE
|
||||||
| NC score | 0.048878 (rank : 38) | θ value | 1.81305 (rank : 45) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 464 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6ZPZ3 | Gene names | Kiaa1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7 homolog. | |||||
|
RABX5_HUMAN
|
||||||
| NC score | 0.048177 (rank : 39) | θ value | 0.813845 (rank : 34) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UJ41 | Gene names | RABGEF1, RABEX5 | |||
|
Domain Architecture |

|
|||||
| Description | Rab5 GDP/GTP exchange factor (Rabex-5) (Rabaptin-5-associated exchange factor for Rab5). | |||||
|
ATN1_HUMAN
|
||||||
| NC score | 0.048001 (rank : 40) | θ value | 3.0926 (rank : 53) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.047610 (rank : 41) | θ value | 0.0961366 (rank : 18) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
SH3K1_MOUSE
|
||||||
| NC score | 0.045859 (rank : 42) | θ value | 1.06291 (rank : 37) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8R550, Q8CAL8, Q8CEF6, Q8R545, Q8R546, Q8R547, Q8R548, Q8R549, Q8R551, Q9CTQ9, Q9DC14, Q9JKC3 | Gene names | Sh3kbp1, Ruk, Seta | |||
|
Domain Architecture |

|
|||||
| Description | SH3-domain kinase-binding protein 1 (SH3-containing, expressed in tumorigenic astrocytes) (Regulator of ubiquitous kinase) (Ruk). | |||||
|
IF4G1_HUMAN
|
||||||
| NC score | 0.041709 (rank : 43) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q04637, O43177, O95066, Q5HYG0, Q6ZN21, Q8N102 | Gene names | EIF4G1, EIF4G, EIF4GI | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1) (p220). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.041027 (rank : 44) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
IF2B2_HUMAN
|
||||||
| NC score | 0.037389 (rank : 45) | θ value | 0.813845 (rank : 31) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y6M1 | Gene names | IGF2BP2, IMP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin-like growth factor 2 mRNA-binding protein 2 (IGF-II mRNA- binding protein 2) (IMP-2) (Hepatocellular carcinoma autoantigen p62). | |||||
|
REPS2_HUMAN
|
||||||
| NC score | 0.033994 (rank : 46) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8NFH8, O43428, Q8NFI5 | Gene names | REPS2, POB1 | |||
|
Domain Architecture |

|
|||||
| Description | RalBP1-associated Eps domain-containing protein 2 (RalBP1-interacting protein 2) (Partner of RalBP1). | |||||
|
NOP56_HUMAN
|
||||||
| NC score | 0.033480 (rank : 47) | θ value | 3.0926 (rank : 54) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O00567, Q9NQ05 | Gene names | NOL5A, NOP56 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar protein Nop56 (Nucleolar protein 5A). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.031500 (rank : 48) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
ZN318_HUMAN
|
||||||
| NC score | 0.030856 (rank : 49) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5VUA4, O94796, Q4G0E4, Q8NEM6, Q9UNU8, Q9Y2W9 | Gene names | ZNF318 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 318 (Endocrine regulatory protein). | |||||
|
SASH1_MOUSE
|
||||||
| NC score | 0.030352 (rank : 50) | θ value | 5.27518 (rank : 63) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P59808 | Gene names | Sash1 | |||
|
Domain Architecture |

|
|||||
| Description | SAM and SH3 domain-containing protein 1. | |||||
|
ANR11_HUMAN
|
||||||
| NC score | 0.030298 (rank : 51) | θ value | 1.38821 (rank : 40) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
CBP_HUMAN
|
||||||
| NC score | 0.029976 (rank : 52) | θ value | 8.99809 (rank : 71) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
RT35_MOUSE
|
||||||
| NC score | 0.029781 (rank : 53) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BJZ4, Q8VCG7 | Gene names | Mrps35 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 28S ribosomal protein S35, mitochondrial precursor (S35mt) (MRP-S35). | |||||
|
DDX51_HUMAN
|
||||||
| NC score | 0.029501 (rank : 54) | θ value | 0.365318 (rank : 26) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
T22D4_HUMAN
|
||||||
| NC score | 0.028948 (rank : 55) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y3Q8 | Gene names | TSC22D4, TILZ2 | |||
|
Domain Architecture |

|
|||||
| Description | TSC22 domain family protein 4 (TSC22-related-inducible leucine zipper protein 2) (Tsc-22-like protein THG-1). | |||||
|
PGCA_MOUSE
|
||||||
| NC score | 0.027066 (rank : 56) | θ value | 0.813845 (rank : 33) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 643 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61282, Q64021 | Gene names | Agc1, Agc | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP). | |||||
|
ROBO4_MOUSE
|
||||||
| NC score | 0.026605 (rank : 57) | θ value | 1.81305 (rank : 48) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8C310, Q9DBW1 | Gene names | Robo4 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 4 precursor. | |||||
|
JHD2B_HUMAN
|
||||||
| NC score | 0.026361 (rank : 58) | θ value | 1.81305 (rank : 46) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7LBC6, Q9BVH6, Q9BW93, Q9BZ52, Q9NYF4, Q9UPS0 | Gene names | JMJD1B, C5orf7, JHDM2B, KIAA1082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 2B (EC 1.14.11.-) (Jumonji domain-containing protein 1B) (Nuclear protein 5qNCA). | |||||
|
TF65_HUMAN
|
||||||
| NC score | 0.026133 (rank : 59) | θ value | 2.36792 (rank : 52) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q04206, Q6SLK1 | Gene names | RELA, NFKB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor p65 (Nuclear factor NF-kappa-B p65 subunit). | |||||
|
SOX13_MOUSE
|
||||||
| NC score | 0.026123 (rank : 60) | θ value | 1.38821 (rank : 42) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 495 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q04891, Q922L3 | Gene names | Sox13, Sox-13 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-13 protein. | |||||
|
RFX3_HUMAN
|
||||||
| NC score | 0.023822 (rank : 61) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P48380, Q5JTL7, Q5JTL8, Q6NW13, Q8WTU4, Q95HL5, Q95HL6 | Gene names | RFX3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor RFX3. | |||||
|
CS029_MOUSE
|
||||||
| NC score | 0.020348 (rank : 62) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9CS00 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf29 homolog. | |||||
|
UBP51_HUMAN
|
||||||
| NC score | 0.019895 (rank : 63) | θ value | 1.81305 (rank : 49) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q70EK9, Q8IWJ8 | Gene names | USP51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 51 (EC 3.1.2.15) (Ubiquitin thioesterase 51) (Ubiquitin-specific-processing protease 51) (Deubiquitinating enzyme 51). | |||||
|
PTN23_HUMAN
|
||||||
| NC score | 0.018520 (rank : 64) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 646 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9H3S7, Q7KZF8, Q8N6Z5, Q9BSR5, Q9P257, Q9UG03, Q9UMZ4 | Gene names | PTPN23, KIAA1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48) (His- domain-containing protein tyrosine phosphatase) (HD-PTP) (Protein tyrosine phosphatase TD14) (PTP-TD14). | |||||
|
DCX_HUMAN
|
||||||
| NC score | 0.018429 (rank : 65) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O43602, O43911 | Gene names | DCX, DBCN, LISX | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal migration protein doublecortin (Lissencephalin-X) (Lis-X) (Doublin). | |||||
|
ETV3_HUMAN
|
||||||
| NC score | 0.018313 (rank : 66) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
RIMB1_MOUSE
|
||||||
| NC score | 0.017838 (rank : 67) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
JIP3_HUMAN
|
||||||
| NC score | 0.017753 (rank : 68) | θ value | 4.03905 (rank : 57) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1039 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UPT6, Q96RY4, Q9H4I4, Q9H7P1, Q9NUG0 | Gene names | MAPK8IP3, JIP3, KIAA1066 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 3 (JNK-interacting protein 3) (JIP-3) (JNK MAP kinase scaffold protein 3) (Mitogen- activated protein kinase 8-interacting protein 3). | |||||
|
PUNC_MOUSE
|
||||||
| NC score | 0.017622 (rank : 69) | θ value | 1.81305 (rank : 47) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 501 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BQC3, O70246, Q792T2, Q9Z2S6 | Gene names | Punc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative neuronal cell adhesion molecule precursor. | |||||
|
TITIN_HUMAN
|
||||||
| NC score | 0.017444 (rank : 70) | θ value | 1.06291 (rank : 38) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
ELL_HUMAN
|
||||||
| NC score | 0.016789 (rank : 71) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P55199 | Gene names | ELL, C19orf17 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II elongation factor ELL (Eleven-nineteen lysine-rich leukemia protein). | |||||
|
ZN533_MOUSE
|
||||||
| NC score | 0.016478 (rank : 72) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BXJ8, Q8BWQ7 | Gene names | Znf533, Zfp533 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 533. | |||||
|
PDLI4_HUMAN
|
||||||
| NC score | 0.015143 (rank : 73) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P50479, Q96AT8, Q9BTW8, Q9Y292 | Gene names | PDLIM4, RIL | |||
|
Domain Architecture |

|
|||||
| Description | PDZ and LIM domain protein 4 (LIM protein RIL) (Reversion-induced LIM protein). | |||||
|
ADA19_HUMAN
|
||||||
| NC score | 0.014104 (rank : 74) | θ value | 1.38821 (rank : 39) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H013, Q9BZL5, Q9UHP2 | Gene names | ADAM19, MLTNB | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta) (Metalloprotease and disintegrin dentritic antigen marker) (MADDAM). | |||||
|
CEL_HUMAN
|
||||||
| NC score | 0.014091 (rank : 75) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
GLI2_HUMAN
|
||||||
| NC score | 0.012922 (rank : 76) | θ value | 0.125558 (rank : 20) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 971 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P10070, O60252, O60253, O60254, O60255, Q15590, Q15591 | Gene names | GLI2, THP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLI2 (Tax helper protein). | |||||
|
GP176_MOUSE
|
||||||
| NC score | 0.011793 (rank : 77) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80WT4, Q80UC4 | Gene names | Gpr176, Agr9, Gm1012 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 176 (G-protein coupled receptor AGR9). | |||||
|
ADCY9_HUMAN
|
||||||
| NC score | 0.010851 (rank : 78) | θ value | 6.88961 (rank : 66) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O60503, O60273, Q4ZHT9, Q4ZIR5, Q9BWT4, Q9UGP2 | Gene names | ADCY9, KIAA0520 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 9 (EC 4.6.1.1) (Adenylate cyclase type IX) (ATP pyrophosphate-lyase 9) (Adenylyl cyclase 9). | |||||
|
BRE1B_HUMAN
|
||||||
| NC score | 0.010531 (rank : 79) | θ value | 4.03905 (rank : 56) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1040 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O75150, Q6AHZ6, Q6N005, Q7L3T6, Q8N615, Q96T18, Q9BSV9, Q9HC82 | Gene names | RNF40, BRE1B, KIAA0661 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase BRE1B (EC 6.3.2.-) (BRE1-B) (RING finger protein 40) (95 kDa retinoblastoma-associated protein) (RBP95). | |||||
|
TENA_MOUSE
|
||||||
| NC score | 0.010214 (rank : 80) | θ value | 4.03905 (rank : 58) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80YX1, Q64706, Q80YX2 | Gene names | Tnc, Hxb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tenascin precursor (TN) (Tenascin-C) (TN-C) (Hexabrachion). | |||||
|
TENX_HUMAN
|
||||||
| NC score | 0.009399 (rank : 81) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 789 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P22105, P78530, P78531, Q08424, Q9UMG7 | Gene names | TNXB, HXBL, TNX, XB | |||
|
Domain Architecture |

|
|||||
| Description | Tenascin-X precursor (TN-X) (Hexabrachion-like protein). | |||||
|
FZD6_MOUSE
|
||||||
| NC score | 0.007955 (rank : 82) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61089 | Gene names | Fzd6 | |||
|
Domain Architecture |

|
|||||
| Description | Frizzled-6 precursor (Fz-6) (mFz6). | |||||
|
ULK1_MOUSE
|
||||||
| NC score | 0.003429 (rank : 83) | θ value | 4.03905 (rank : 59) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1214 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O70405, Q6PGB2 | Gene names | Ulk1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ULK1 (EC 2.7.11.1) (Unc-51-like kinase 1) (Serine/threonine-protein kinase Unc51.1). | |||||
|
VGFR3_MOUSE
|
||||||
| NC score | 0.002890 (rank : 84) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1056 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P35917 | Gene names | Flt4, Flt-4 | |||
|
Domain Architecture |

|
|||||
| Description | Vascular endothelial growth factor receptor 3 precursor (EC 2.7.10.1) (VEGFR-3) (Tyrosine-protein kinase receptor FLT4). | |||||