Please be patient as the page loads
|
DNM3A_MOUSE
|
||||||
| SwissProt Accessions | O88508, Q3UH24, Q8CJ60, Q922J0, Q9CSE1 | Gene names | Dnmt3a | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3A (EC 2.1.1.37) (Dnmt3a) (DNA methyltransferase MmuIIIA) (DNA MTase MmuIIIA) (M.MmuIIIA). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
DNM3A_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.987655 (rank : 2) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Y6K1, Q8WXU9 | Gene names | DNMT3A | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3A (EC 2.1.1.37) (Dnmt3a) (DNA methyltransferase HsaIIIA) (DNA MTase HsaIIIA) (M.HsaIIIA). | |||||
|
DNM3A_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | O88508, Q3UH24, Q8CJ60, Q922J0, Q9CSE1 | Gene names | Dnmt3a | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3A (EC 2.1.1.37) (Dnmt3a) (DNA methyltransferase MmuIIIA) (DNA MTase MmuIIIA) (M.MmuIIIA). | |||||
|
DNM3B_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.976314 (rank : 4) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UBC3, Q9UBD4, Q9UJQ5, Q9UKA6, Q9UNE5, Q9Y5R9, Q9Y5S0 | Gene names | DNMT3B | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3B (EC 2.1.1.37) (Dnmt3b) (DNA methyltransferase HsaIIIB) (DNA MTase HsaIIIB) (M.HsaIIIB). | |||||
|
DNM3B_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.976615 (rank : 3) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O88509, O88510, O88511 | Gene names | Dnmt3b | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3B (EC 2.1.1.37) (Dnmt3b) (DNA methyltransferase MmuIIIB) (DNA MTase MmuIIIB) (M.MmuIIIB). | |||||
|
DNM3L_HUMAN
|
||||||
| θ value | 6.53529e-67 (rank : 5) | NC score | 0.906775 (rank : 5) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UJW3, Q9BUJ4 | Gene names | DNMT3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA (cytosine-5)-methyltransferase 3-like. | |||||
|
DNM3L_MOUSE
|
||||||
| θ value | 5.7252e-63 (rank : 6) | NC score | 0.898085 (rank : 6) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9CWR8 | Gene names | Dnmt3l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA (cytosine-5)-methyltransferase 3-like. | |||||
|
PWWP2_HUMAN
|
||||||
| θ value | 1.133e-10 (rank : 7) | NC score | 0.466598 (rank : 7) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6NUJ5, Q5SZI0, Q6ZQX5, Q96F43 | Gene names | PWWP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PWWP domain-containing protein 2. | |||||
|
ATRX_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 8) | NC score | 0.166204 (rank : 9) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P46100, P51068, Q15886, Q59FB5, Q59H31, Q5H9A2, Q5JWI4, Q7Z2J1, Q9H0Z1, Q9NTS3 | Gene names | ATRX, RAD54L, XH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked helicase II) (X-linked nuclear protein) (XNP) (Znf- HX). | |||||
|
ATRX_MOUSE
|
||||||
| θ value | 0.000461057 (rank : 9) | NC score | 0.169216 (rank : 8) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q61687 | Gene names | Atrx, Hp1bp2, Xnp | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked nuclear protein) (Heterochromatin protein 2) (HP1 alpha-interacting protein) (HP1-BP38 protein). | |||||
|
RRBP1_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 10) | NC score | 0.029924 (rank : 41) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1306 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9P2E9, O75300, O75301, Q5W165, Q96SB2, Q9BWP1, Q9H476 | Gene names | RRBP1, KIAA1398 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (180 kDa ribosome receptor homolog) (ES/130-related protein). | |||||
|
MBD5_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 11) | NC score | 0.145812 (rank : 10) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9P267, Q9NUV6 | Gene names | MBD5, KIAA1461 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 5 (Methyl-CpG-binding protein MBD5). | |||||
|
CHD5_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 12) | NC score | 0.087813 (rank : 14) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8TDI0, O75032, Q5TG89, Q9UFR9 | Gene names | CHD5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 5 (EC 3.6.1.-) (ATP- dependent helicase CHD5) (CHD-5). | |||||
|
MSH6_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 13) | NC score | 0.103051 (rank : 11) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P54276, O54710 | Gene names | Msh6, Gtmbp | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein MSH6 (MutS-alpha 160 kDa subunit) (G/T mismatch-binding protein) (GTBP) (GTMBP) (p160). | |||||
|
CHD3_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 14) | NC score | 0.083943 (rank : 17) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q12873, Q9Y4I0 | Gene names | CHD3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 3 (EC 3.6.1.-) (ATP- dependent helicase CHD3) (CHD-3) (Mi-2 autoantigen 240 kDa protein) (Mi2-alpha) (Zinc-finger helicase) (hZFH). | |||||
|
CHD4_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 15) | NC score | 0.083525 (rank : 19) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q14839, Q8IXZ5 | Gene names | CHD4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (EC 3.6.1.-) (ATP- dependent helicase CHD4) (CHD-4) (Mi-2 autoantigen 218 kDa protein) (Mi2-beta). | |||||
|
CHD4_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 16) | NC score | 0.083625 (rank : 18) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q6PDQ2 | Gene names | Chd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (CHD-4). | |||||
|
MEFV_HUMAN
|
||||||
| θ value | 0.279714 (rank : 17) | NC score | 0.014823 (rank : 67) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O15553, Q96PN4, Q96PN5 | Gene names | MEFV, MEF | |||
|
Domain Architecture |

|
|||||
| Description | Pyrin (Marenostrin). | |||||
|
TOIP2_MOUSE
|
||||||
| θ value | 0.279714 (rank : 18) | NC score | 0.052529 (rank : 32) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BYU6, Q6NY10, Q8BJP0 | Gene names | Tor1aip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Torsin-1A-interacting protein 2. | |||||
|
SGOL2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 19) | NC score | 0.036883 (rank : 36) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7TSY8, Q811I4, Q8C9C4, Q8CGJ4, Q9CS55 | Gene names | Sgol2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 2. | |||||
|
PDZD2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 20) | NC score | 0.022960 (rank : 51) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O15018, Q9BXD4 | Gene names | PDZD2, AIPC, KIAA0300, PDZK3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 2 (PDZ domain-containing protein 3) (Activated in prostate cancer protein). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 0.47712 (rank : 21) | NC score | 0.024428 (rank : 47) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
C8AP2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 22) | NC score | 0.029160 (rank : 42) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9WUF3, Q9CSW2 | Gene names | Casp8ap2, Flash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
CS007_HUMAN
|
||||||
| θ value | 0.62314 (rank : 23) | NC score | 0.040260 (rank : 33) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UPT8, Q9Y420 | Gene names | C19orf7, KIAA1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7. | |||||
|
MRCKG_HUMAN
|
||||||
| θ value | 0.62314 (rank : 24) | NC score | 0.004033 (rank : 86) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1519 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6DT37, O00565 | Gene names | CDC42BPG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK gamma (EC 2.7.11.1) (CDC42- binding protein kinase gamma) (Myotonic dystrophy kinase-related CDC42-binding kinase gamma) (Myotonic dystrophy protein kinase-like alpha) (MRCK gamma) (DMPK-like gamma). | |||||
|
PF21A_HUMAN
|
||||||
| θ value | 0.62314 (rank : 25) | NC score | 0.089004 (rank : 13) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96BD5, Q6AWA2, Q9C0G7, Q9H8V9, Q9HAK6, Q9NZE9 | Gene names | PHF21A, BHC80, KIAA1696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (BHC80a). | |||||
|
MSH6_HUMAN
|
||||||
| θ value | 0.813845 (rank : 26) | NC score | 0.094091 (rank : 12) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P52701, O43706, O43917, Q8TCX4, Q9BTB5 | Gene names | MSH6, GTBP | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein MSH6 (MutS-alpha 160 kDa subunit) (G/T mismatch-binding protein) (GTBP) (GTMBP) (p160). | |||||
|
RFX1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 27) | NC score | 0.021360 (rank : 53) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P48377 | Gene names | Rfx1 | |||
|
Domain Architecture |
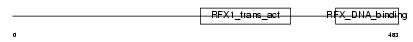
|
|||||
| Description | DNA-binding protein RFX1. | |||||
|
TACC1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 28) | NC score | 0.016418 (rank : 60) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6Y685, Q6Y686 | Gene names | Tacc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transforming acidic coiled-coil-containing protein 1. | |||||
|
INVS_HUMAN
|
||||||
| θ value | 1.38821 (rank : 29) | NC score | 0.011242 (rank : 77) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Y283, Q5W0T6, Q8IVX8, Q9BRB9, Q9Y488, Q9Y498 | Gene names | INVS, INV, NPHP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning homolog) (Nephrocystin-2). | |||||
|
BAT2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 30) | NC score | 0.034529 (rank : 39) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
HDGR3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 31) | NC score | 0.079467 (rank : 21) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y3E1 | Gene names | HDGFRP3, HDGF2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatoma-derived growth factor-related protein 3 (HRP-3) (Hepatoma- derived growth factor 2). | |||||
|
HDGR3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 32) | NC score | 0.080292 (rank : 20) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9JMG7, Q3TRX2, Q8BQ69, Q8BR62, Q9D2M7 | Gene names | Hdgfrp3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatoma-derived growth factor-related protein 3 (HRP-3). | |||||
|
K1949_MOUSE
|
||||||
| θ value | 1.81305 (rank : 33) | NC score | 0.027382 (rank : 44) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BQ30, Q3UDS6, Q8CEY9, Q8R3D8 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1949 homolog. | |||||
|
PF21A_MOUSE
|
||||||
| θ value | 1.81305 (rank : 34) | NC score | 0.086763 (rank : 15) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6ZPK0, Q6XVG0, Q80Z33, Q8CAZ4, Q8VEC8 | Gene names | Phf21a, Bhc80, Kiaa1696, Pftf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (mBHC80) (BHC80a). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 35) | NC score | 0.026482 (rank : 45) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
ST65G_HUMAN
|
||||||
| θ value | 1.81305 (rank : 36) | NC score | 0.036360 (rank : 37) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O94864, Q6IB21, Q9H2T6 | Gene names | SUPT7L, KIAA0764 | |||
|
Domain Architecture |

|
|||||
| Description | STAGA complex 65 subunit gamma (STAF65gamma) (SPTF-associated factor 65 gamma) (Suppressor of Ty 7-like) (Adenocarcinoma antigen ART1). | |||||
|
ST65G_MOUSE
|
||||||
| θ value | 1.81305 (rank : 37) | NC score | 0.036331 (rank : 38) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CZV5 | Gene names | Supt7l | |||
|
Domain Architecture |

|
|||||
| Description | STAGA complex 65 subunit gamma (STAF65gamma) (SPTF-associated factor 65 gamma) (Suppressor of Ty 7-like). | |||||
|
TP53B_HUMAN
|
||||||
| θ value | 1.81305 (rank : 38) | NC score | 0.037617 (rank : 35) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 413 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q12888, Q2M1Z7, Q4LE46, Q5FWZ3, Q7Z3U4 | Gene names | TP53BP1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
WNK2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 39) | NC score | 0.007107 (rank : 83) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
ZBT45_HUMAN
|
||||||
| θ value | 1.81305 (rank : 40) | NC score | 0.004887 (rank : 85) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 901 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96K62 | Gene names | ZBTB45, ZNF499 | |||
|
Domain Architecture |
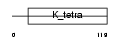
|
|||||
| Description | Zinc finger and BTB domain-containing protein 45 (Zinc finger protein 499). | |||||
|
ZCPW1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 41) | NC score | 0.085168 (rank : 16) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9H0M4, Q8NA98, Q9BUD0, Q9NWF7 | Gene names | ZCWPW1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CW-type PWWP domain protein 1. | |||||
|
GP124_MOUSE
|
||||||
| θ value | 2.36792 (rank : 42) | NC score | 0.012096 (rank : 76) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q91ZV8 | Gene names | Gpr124, Tem5 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 124 precursor (Tumor endothelial marker 5). | |||||
|
HGS_HUMAN
|
||||||
| θ value | 2.36792 (rank : 43) | NC score | 0.015448 (rank : 63) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O14964, Q9NR36 | Gene names | HGS, HRS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate (Protein pp110) (Hrs). | |||||
|
LAD1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 44) | NC score | 0.033844 (rank : 40) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P57016 | Gene names | Lad1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (Linear IgA disease autoantigen). | |||||
|
PSL1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 45) | NC score | 0.016896 (rank : 59) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8TCT7, O60365, Q567S3, Q8IUH9, Q9BUY6, Q9H3M4, Q9NPN2, Q9P1Z6 | Gene names | SPPL2B, KIAA1532, PSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal peptide peptidase-like 2B (EC 3.4.23.-) (Protein SPP-like 2B) (Protein SPPL2b) (Intramembrane protease 4) (IMP4) (Presenilin-like protein 1). | |||||
|
SP110_HUMAN
|
||||||
| θ value | 2.36792 (rank : 46) | NC score | 0.068632 (rank : 25) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9HB58, Q14976, Q14977, Q8WUZ6, Q9HCT8 | Gene names | SP110 | |||
|
Domain Architecture |

|
|||||
| Description | Sp110 nuclear body protein (Speckled 110 kDa) (Transcriptional coactivator Sp110) (Interferon-induced protein 41/75). | |||||
|
TXND2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 47) | NC score | 0.021051 (rank : 54) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
ZFHX2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 48) | NC score | 0.016247 (rank : 61) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1178 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9C0A1 | Gene names | ZFHX2, KIAA1762 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger homeobox protein 2. | |||||
|
ZMY11_HUMAN
|
||||||
| θ value | 2.36792 (rank : 49) | NC score | 0.077480 (rank : 22) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q15326 | Gene names | ZMYND11, BS69 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11 (Adenovirus 5 E1A- binding protein) (BS69 protein). | |||||
|
ZMY11_MOUSE
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.076640 (rank : 23) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8R5C8, Q8C155 | Gene names | Zmynd11 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11. | |||||
|
ZN335_HUMAN
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.000689 (rank : 92) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 973 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H4Z2, Q9H684 | Gene names | ZNF335 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 335. | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 52) | NC score | 0.015037 (rank : 65) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
CASC3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 53) | NC score | 0.021552 (rank : 52) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K3W3, Q8K219, Q99NF0 | Gene names | Casc3, Mln51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASC3 (Cancer susceptibility candidate gene 3 protein homolog) (Metastatic lymph node protein 51 homolog) (Protein MLN 51 homolog) (Protein barentsz) (Btz). | |||||
|
KSR1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 54) | NC score | 0.003437 (rank : 89) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q61097, Q61648, Q78DX8 | Gene names | Ksr1, Ksr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinase suppressor of ras-1 (Kinase suppressor of ras) (mKSR1) (Hb protein). | |||||
|
LRC50_MOUSE
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.014743 (rank : 68) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D2H9, Q9CVS9 | Gene names | Lrrc50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 50. | |||||
|
NSD1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.062965 (rank : 29) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 3.0926 (rank : 57) | NC score | 0.016114 (rank : 62) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
PSIP1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.065893 (rank : 26) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q99JF8, Q3TEJ7, Q3TTD7, Q3UTA1, Q80WQ7, Q99JF7, Q99LR4, Q9CT03 | Gene names | Psip1, Ledgf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PC4 and SFRS1-interacting protein (Lens epithelium-derived growth factor) (mLEDGF). | |||||
|
SYTL2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | 0.010916 (rank : 78) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9HCH5, Q2YDA7, Q8ND34, Q96BJ2, Q9H768, Q9NXM1 | Gene names | SYTL2, KIAA1597, SLP2 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 2 (Exophilin-4). | |||||
|
TCAL5_HUMAN
|
||||||
| θ value | 3.0926 (rank : 60) | NC score | 0.024123 (rank : 48) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
TMAP1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 61) | NC score | 0.065241 (rank : 27) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P84157 | Gene names | TMAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane anchor protein 1. | |||||
|
ZCPW1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 62) | NC score | 0.065228 (rank : 28) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6IR42 | Gene names | Zcwpw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CW-type PWWP domain protein 1 homolog. | |||||
|
CSKI2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.012234 (rank : 74) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8WXE0, Q7LG69, Q9ULT1 | Gene names | CASKIN2, KIAA1139 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-2. | |||||
|
LAD1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.027806 (rank : 43) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O00515, O95614 | Gene names | LAD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (120 kDa linear IgA bullous dermatosis antigen) (97 kDa linear IgA bullous dermatosis antigen) (Linear IgA disease antigen homolog) (LadA). | |||||
|
LSD1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 65) | NC score | 0.012101 (rank : 75) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O60341, Q5TH94, Q5TH95, Q86VT7, Q8IXK4, Q8NDP6, Q8TAZ3, Q96AW4 | Gene names | AOF2, KIAA0601, LSD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lysine-specific histone demethylase 1 (EC 1.-.-.-) (Flavin-containing amine oxidase domain-containing protein 2) (BRAF35-HDAC complex protein BHC110). | |||||
|
PAK4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 66) | NC score | 0.001722 (rank : 90) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1098 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BTW9, Q6ZPX0, Q80Z97, Q9CS71 | Gene names | Pak4, Kiaa1142 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PAK 4 (EC 2.7.11.1) (p21-activated kinase 4) (PAK-4). | |||||
|
PI51C_HUMAN
|
||||||
| θ value | 4.03905 (rank : 67) | NC score | 0.013826 (rank : 69) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60331, Q7LE07 | Gene names | PIP5K1C, KIAA0589 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-4-phosphate 5-kinase type-1 gamma (EC 2.7.1.68) (Phosphatidylinositol-4-phosphate 5-kinase type I gamma) (PtdIns(4)P- 5-kinase gamma) (PtdInsPKIgamma) (PIP5KIgamma). | |||||
|
PRGR_HUMAN
|
||||||
| θ value | 4.03905 (rank : 68) | NC score | 0.012591 (rank : 73) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P06401, Q9UPF7 | Gene names | PGR, NR3C3 | |||
|
Domain Architecture |

|
|||||
| Description | Progesterone receptor (PR). | |||||
|
REN3A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 69) | NC score | 0.020181 (rank : 55) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H1J1, Q5T8C3, Q5T8C9, Q7Z6N3, Q86YK1, Q9BZI8 | Gene names | UPF3A, RENT3A, UPF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of nonsense transcripts 3A (Nonsense mRNA reducing factor 3A) (Up-frameshift suppressor 3 homolog A) (hUpf3). | |||||
|
WDR46_MOUSE
|
||||||
| θ value | 4.03905 (rank : 70) | NC score | 0.023242 (rank : 50) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z0H1 | Gene names | Wdr46, Bing4 | |||
|
Domain Architecture |
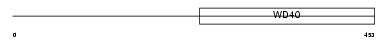
|
|||||
| Description | WD repeat protein 46 (WD repeat protein BING4). | |||||
|
AT1B4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 71) | NC score | 0.014837 (rank : 66) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99ME6, Q99ME5 | Gene names | Atp1b4 | |||
|
Domain Architecture |
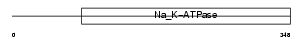
|
|||||
| Description | X/potassium-transporting ATPase subunit beta-m (X,K-ATPase beta-m subunit). | |||||
|
CD008_HUMAN
|
||||||
| θ value | 5.27518 (rank : 72) | NC score | 0.019751 (rank : 57) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P78312, O43607, P78311, P78313, Q9UEG8 | Gene names | C4orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C4orf8 (Protein IT14). | |||||
|
EHMT2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 73) | NC score | 0.008422 (rank : 81) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Z148, Q6PE08, Q8K4R6, Q8K4R7, Q9Z149 | Gene names | Ehmt2, Bat8, G9a, Ng36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
LY10_HUMAN
|
||||||
| θ value | 5.27518 (rank : 74) | NC score | 0.040254 (rank : 34) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q13342, Q13341, Q92881, Q96TG3 | Gene names | SP140, LYSP100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear body protein SP140 (Nuclear autoantigen Sp-140) (Speckled 140 kDa) (LYSp100 protein) (Lymphoid-restricted homolog of Sp100). | |||||
|
MDC1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 75) | NC score | 0.019327 (rank : 58) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
PSIP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 76) | NC score | 0.055049 (rank : 31) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O75475, O00256, O95368, Q6P391, Q86YB9, Q9NZI3, Q9UER6 | Gene names | PSIP1, DFS70, LEDGF, PSIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PC4 and SFRS1-interacting protein (Lens epithelium-derived growth factor) (Transcriptional coactivator p75/p52) (Dense fine speckles 70 kDa protein) (DFS 70) (CLL-associated antigen KW-7). | |||||
|
BAT2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.024102 (rank : 49) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
BCL7C_MOUSE
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.025030 (rank : 46) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O08664, Q99LS2 | Gene names | Bcl7c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell CLL/lymphoma 7 protein family member C (B-cell chronic lymphocytic leukemia/lymphoma 7C protein). | |||||
|
BRSK1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.001173 (rank : 91) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 942 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8TDC3, Q5J5B5, Q8NDD0, Q8NDR4, Q8TDC2, Q96AV4, Q96JL4 | Gene names | BRSK1, KIAA1811, SAD1 | |||
|
Domain Architecture |

|
|||||
| Description | BR serine/threonine-protein kinase 1 (EC 2.7.11.1) (SAD1 kinase) (SAD1A). | |||||
|
CSKI2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.009648 (rank : 79) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 812 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8VHK1, Q6ZPX1, Q7TS77 | Gene names | Caskin2, Kiaa1139 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-2. | |||||
|
ELOA3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.012996 (rank : 71) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NG57 | Gene names | TCEB3C, TCEB3L2 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II transcription factor SIII subunit A3 (Elongin-A3) (EloA3) (Transcription elongation factor B polypeptide 3C). | |||||
|
KC1D_HUMAN
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.003647 (rank : 87) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 737 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P48730, Q96KZ6, Q9BTN5 | Gene names | CSNK1D | |||
|
Domain Architecture |

|
|||||
| Description | Casein kinase I isoform delta (EC 2.7.11.1) (CKI-delta) (CKId). | |||||
|
KC1D_MOUSE
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.003647 (rank : 88) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 737 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DC28, Q3TZK2, Q99KK4 | Gene names | Csnk1d | |||
|
Domain Architecture |

|
|||||
| Description | Casein kinase I isoform delta (EC 2.7.11.1) (CKI-delta) (CKId). | |||||
|
RP3A_MOUSE
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.007922 (rank : 82) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 510 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P47708 | Gene names | Rph3a | |||
|
Domain Architecture |

|
|||||
| Description | Rabphilin-3A (Exophilin-1). | |||||
|
WDR81_MOUSE
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.012816 (rank : 72) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5ND34, Q3TA90, Q3TUS9, Q6KAR1 | Gene names | Wdr81 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 81. | |||||
|
ES8L1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.005533 (rank : 84) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R5F8, Q8R0D6, Q9D2M6 | Gene names | Eps8l1, Eps8r1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Epidermal growth factor receptor kinase substrate 8-like protein 1 (Epidermal growth factor receptor pathway substrate 8-related protein 1) (EPS8-like protein 1). | |||||
|
HGS_MOUSE
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.013322 (rank : 70) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q99LI8, Q61691, Q8BQW3 | Gene names | Hgs, Hrs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate. | |||||
|
MYLK_HUMAN
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.000184 (rank : 93) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q15746, O95796, O95797, O95798, O95799, Q14844, Q16794, Q5MY99, Q5MYA0, Q7Z4J0, Q9C0L5, Q9UBG5, Q9UIT9 | Gene names | MYLK, MLCK | |||
|
Domain Architecture |

|
|||||
| Description | Myosin light chain kinase, smooth muscle (EC 2.7.11.18) (MLCK) (Telokin) (Kinase-related protein) (KRP). | |||||
|
RRBP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.019937 (rank : 56) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
SASH1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.008666 (rank : 80) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O94885, Q8TAI0, Q9H7R7 | Gene names | SASH1, KIAA0790, PEPE1 | |||
|
Domain Architecture |

|
|||||
| Description | SAM and SH3 domain-containing protein 1 (Proline-glutamate repeat- containing protein). | |||||
|
ZN638_HUMAN
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.015191 (rank : 64) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14966, Q53R34, Q5XJ05, Q68DP3, Q6P2H2, Q7Z3T7, Q8NF92, Q8TCA1, Q9H2G1, Q9NP37 | Gene names | ZNF638, NP220, ZFML | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein) (Cutaneous T-cell lymphoma-associated antigen se33-1) (CTCL tumor antigen se33-1). | |||||
|
HDGF_HUMAN
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.075210 (rank : 24) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P51858, Q5SZ09 | Gene names | HDGF, HMG1L2 | |||
|
Domain Architecture |
|
|||||
| Description | Hepatoma-derived growth factor (HDGF) (High-mobility group protein 1- like 2) (HMG-1L2). | |||||
|
HDGF_MOUSE
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.059220 (rank : 30) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P51859, Q8BPG7, Q9CYA4, Q9JK87 | Gene names | Hdgf, Tdrm1 | |||
|
Domain Architecture |
|
|||||
| Description | Hepatoma-derived growth factor (HDGF). | |||||
|
DNM3A_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | O88508, Q3UH24, Q8CJ60, Q922J0, Q9CSE1 | Gene names | Dnmt3a | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3A (EC 2.1.1.37) (Dnmt3a) (DNA methyltransferase MmuIIIA) (DNA MTase MmuIIIA) (M.MmuIIIA). | |||||
|
DNM3A_HUMAN
|
||||||
| NC score | 0.987655 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Y6K1, Q8WXU9 | Gene names | DNMT3A | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3A (EC 2.1.1.37) (Dnmt3a) (DNA methyltransferase HsaIIIA) (DNA MTase HsaIIIA) (M.HsaIIIA). | |||||
|
DNM3B_MOUSE
|
||||||
| NC score | 0.976615 (rank : 3) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O88509, O88510, O88511 | Gene names | Dnmt3b | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3B (EC 2.1.1.37) (Dnmt3b) (DNA methyltransferase MmuIIIB) (DNA MTase MmuIIIB) (M.MmuIIIB). | |||||
|
DNM3B_HUMAN
|
||||||
| NC score | 0.976314 (rank : 4) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UBC3, Q9UBD4, Q9UJQ5, Q9UKA6, Q9UNE5, Q9Y5R9, Q9Y5S0 | Gene names | DNMT3B | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3B (EC 2.1.1.37) (Dnmt3b) (DNA methyltransferase HsaIIIB) (DNA MTase HsaIIIB) (M.HsaIIIB). | |||||
|
DNM3L_HUMAN
|
||||||
| NC score | 0.906775 (rank : 5) | θ value | 6.53529e-67 (rank : 5) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UJW3, Q9BUJ4 | Gene names | DNMT3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA (cytosine-5)-methyltransferase 3-like. | |||||
|
DNM3L_MOUSE
|
||||||
| NC score | 0.898085 (rank : 6) | θ value | 5.7252e-63 (rank : 6) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9CWR8 | Gene names | Dnmt3l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA (cytosine-5)-methyltransferase 3-like. | |||||
|
PWWP2_HUMAN
|
||||||
| NC score | 0.466598 (rank : 7) | θ value | 1.133e-10 (rank : 7) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6NUJ5, Q5SZI0, Q6ZQX5, Q96F43 | Gene names | PWWP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PWWP domain-containing protein 2. | |||||
|
ATRX_MOUSE
|
||||||
| NC score | 0.169216 (rank : 8) | θ value | 0.000461057 (rank : 9) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q61687 | Gene names | Atrx, Hp1bp2, Xnp | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked nuclear protein) (Heterochromatin protein 2) (HP1 alpha-interacting protein) (HP1-BP38 protein). | |||||
|
ATRX_HUMAN
|
||||||
| NC score | 0.166204 (rank : 9) | θ value | 0.000461057 (rank : 8) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P46100, P51068, Q15886, Q59FB5, Q59H31, Q5H9A2, Q5JWI4, Q7Z2J1, Q9H0Z1, Q9NTS3 | Gene names | ATRX, RAD54L, XH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked helicase II) (X-linked nuclear protein) (XNP) (Znf- HX). | |||||
|
MBD5_HUMAN
|
||||||
| NC score | 0.145812 (rank : 10) | θ value | 0.0193708 (rank : 11) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9P267, Q9NUV6 | Gene names | MBD5, KIAA1461 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 5 (Methyl-CpG-binding protein MBD5). | |||||
|
MSH6_MOUSE
|
||||||
| NC score | 0.103051 (rank : 11) | θ value | 0.0431538 (rank : 13) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P54276, O54710 | Gene names | Msh6, Gtmbp | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein MSH6 (MutS-alpha 160 kDa subunit) (G/T mismatch-binding protein) (GTBP) (GTMBP) (p160). | |||||
|
MSH6_HUMAN
|
||||||
| NC score | 0.094091 (rank : 12) | θ value | 0.813845 (rank : 26) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P52701, O43706, O43917, Q8TCX4, Q9BTB5 | Gene names | MSH6, GTBP | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein MSH6 (MutS-alpha 160 kDa subunit) (G/T mismatch-binding protein) (GTBP) (GTMBP) (p160). | |||||
|
PF21A_HUMAN
|
||||||
| NC score | 0.089004 (rank : 13) | θ value | 0.62314 (rank : 25) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96BD5, Q6AWA2, Q9C0G7, Q9H8V9, Q9HAK6, Q9NZE9 | Gene names | PHF21A, BHC80, KIAA1696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (BHC80a). | |||||
|
CHD5_HUMAN
|
||||||
| NC score | 0.087813 (rank : 14) | θ value | 0.0330416 (rank : 12) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8TDI0, O75032, Q5TG89, Q9UFR9 | Gene names | CHD5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 5 (EC 3.6.1.-) (ATP- dependent helicase CHD5) (CHD-5). | |||||
|
PF21A_MOUSE
|
||||||
| NC score | 0.086763 (rank : 15) | θ value | 1.81305 (rank : 34) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6ZPK0, Q6XVG0, Q80Z33, Q8CAZ4, Q8VEC8 | Gene names | Phf21a, Bhc80, Kiaa1696, Pftf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (mBHC80) (BHC80a). | |||||
|
ZCPW1_HUMAN
|
||||||
| NC score | 0.085168 (rank : 16) | θ value | 1.81305 (rank : 41) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9H0M4, Q8NA98, Q9BUD0, Q9NWF7 | Gene names | ZCWPW1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CW-type PWWP domain protein 1. | |||||
|
CHD3_HUMAN
|
||||||
| NC score | 0.083943 (rank : 17) | θ value | 0.0961366 (rank : 14) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q12873, Q9Y4I0 | Gene names | CHD3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 3 (EC 3.6.1.-) (ATP- dependent helicase CHD3) (CHD-3) (Mi-2 autoantigen 240 kDa protein) (Mi2-alpha) (Zinc-finger helicase) (hZFH). | |||||
|
CHD4_MOUSE
|
||||||
| NC score | 0.083625 (rank : 18) | θ value | 0.0961366 (rank : 16) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q6PDQ2 | Gene names | Chd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (CHD-4). | |||||
|
CHD4_HUMAN
|
||||||
| NC score | 0.083525 (rank : 19) | θ value | 0.0961366 (rank : 15) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q14839, Q8IXZ5 | Gene names | CHD4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (EC 3.6.1.-) (ATP- dependent helicase CHD4) (CHD-4) (Mi-2 autoantigen 218 kDa protein) (Mi2-beta). | |||||
|
HDGR3_MOUSE
|
||||||
| NC score | 0.080292 (rank : 20) | θ value | 1.81305 (rank : 32) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9JMG7, Q3TRX2, Q8BQ69, Q8BR62, Q9D2M7 | Gene names | Hdgfrp3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatoma-derived growth factor-related protein 3 (HRP-3). | |||||
|
HDGR3_HUMAN
|
||||||
| NC score | 0.079467 (rank : 21) | θ value | 1.81305 (rank : 31) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y3E1 | Gene names | HDGFRP3, HDGF2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatoma-derived growth factor-related protein 3 (HRP-3) (Hepatoma- derived growth factor 2). | |||||
|
ZMY11_HUMAN
|
||||||
| NC score | 0.077480 (rank : 22) | θ value | 2.36792 (rank : 49) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q15326 | Gene names | ZMYND11, BS69 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11 (Adenovirus 5 E1A- binding protein) (BS69 protein). | |||||
|
ZMY11_MOUSE
|
||||||
| NC score | 0.076640 (rank : 23) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8R5C8, Q8C155 | Gene names | Zmynd11 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11. | |||||
|
HDGF_HUMAN
|
||||||
| NC score | 0.075210 (rank : 24) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P51858, Q5SZ09 | Gene names | HDGF, HMG1L2 | |||
|
Domain Architecture |

|
|||||
| Description | Hepatoma-derived growth factor (HDGF) (High-mobility group protein 1- like 2) (HMG-1L2). | |||||
|
SP110_HUMAN
|
||||||
| NC score | 0.068632 (rank : 25) | θ value | 2.36792 (rank : 46) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9HB58, Q14976, Q14977, Q8WUZ6, Q9HCT8 | Gene names | SP110 | |||
|
Domain Architecture |

|
|||||
| Description | Sp110 nuclear body protein (Speckled 110 kDa) (Transcriptional coactivator Sp110) (Interferon-induced protein 41/75). | |||||
|
PSIP1_MOUSE
|
||||||
| NC score | 0.065893 (rank : 26) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q99JF8, Q3TEJ7, Q3TTD7, Q3UTA1, Q80WQ7, Q99JF7, Q99LR4, Q9CT03 | Gene names | Psip1, Ledgf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PC4 and SFRS1-interacting protein (Lens epithelium-derived growth factor) (mLEDGF). | |||||
|
TMAP1_HUMAN
|
||||||
| NC score | 0.065241 (rank : 27) | θ value | 3.0926 (rank : 61) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P84157 | Gene names | TMAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane anchor protein 1. | |||||
|
ZCPW1_MOUSE
|
||||||
| NC score | 0.065228 (rank : 28) | θ value | 3.0926 (rank : 62) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6IR42 | Gene names | Zcwpw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CW-type PWWP domain protein 1 homolog. | |||||
|
NSD1_HUMAN
|
||||||
| NC score | 0.062965 (rank : 29) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
HDGF_MOUSE
|
||||||
| NC score | 0.059220 (rank : 30) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P51859, Q8BPG7, Q9CYA4, Q9JK87 | Gene names | Hdgf, Tdrm1 | |||
|
Domain Architecture |

|
|||||
| Description | Hepatoma-derived growth factor (HDGF). | |||||
|
PSIP1_HUMAN
|
||||||
| NC score | 0.055049 (rank : 31) | θ value | 5.27518 (rank : 76) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O75475, O00256, O95368, Q6P391, Q86YB9, Q9NZI3, Q9UER6 | Gene names | PSIP1, DFS70, LEDGF, PSIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PC4 and SFRS1-interacting protein (Lens epithelium-derived growth factor) (Transcriptional coactivator p75/p52) (Dense fine speckles 70 kDa protein) (DFS 70) (CLL-associated antigen KW-7). | |||||
|
TOIP2_MOUSE
|
||||||
| NC score | 0.052529 (rank : 32) | θ value | 0.279714 (rank : 18) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BYU6, Q6NY10, Q8BJP0 | Gene names | Tor1aip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Torsin-1A-interacting protein 2. | |||||
|
CS007_HUMAN
|
||||||
| NC score | 0.040260 (rank : 33) | θ value | 0.62314 (rank : 23) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UPT8, Q9Y420 | Gene names | C19orf7, KIAA1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7. | |||||
|
LY10_HUMAN
|
||||||
| NC score | 0.040254 (rank : 34) | θ value | 5.27518 (rank : 74) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q13342, Q13341, Q92881, Q96TG3 | Gene names | SP140, LYSP100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear body protein SP140 (Nuclear autoantigen Sp-140) (Speckled 140 kDa) (LYSp100 protein) (Lymphoid-restricted homolog of Sp100). | |||||
|
TP53B_HUMAN
|
||||||
| NC score | 0.037617 (rank : 35) | θ value | 1.81305 (rank : 38) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 413 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q12888, Q2M1Z7, Q4LE46, Q5FWZ3, Q7Z3U4 | Gene names | TP53BP1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
SGOL2_MOUSE
|
||||||
| NC score | 0.036883 (rank : 36) | θ value | 0.365318 (rank : 19) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7TSY8, Q811I4, Q8C9C4, Q8CGJ4, Q9CS55 | Gene names | Sgol2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 2. | |||||
|
ST65G_HUMAN
|
||||||
| NC score | 0.036360 (rank : 37) | θ value | 1.81305 (rank : 36) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O94864, Q6IB21, Q9H2T6 | Gene names | SUPT7L, KIAA0764 | |||
|
Domain Architecture |

|
|||||
| Description | STAGA complex 65 subunit gamma (STAF65gamma) (SPTF-associated factor 65 gamma) (Suppressor of Ty 7-like) (Adenocarcinoma antigen ART1). | |||||
|
ST65G_MOUSE
|
||||||
| NC score | 0.036331 (rank : 38) | θ value | 1.81305 (rank : 37) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CZV5 | Gene names | Supt7l | |||
|
Domain Architecture |

|
|||||
| Description | STAGA complex 65 subunit gamma (STAF65gamma) (SPTF-associated factor 65 gamma) (Suppressor of Ty 7-like). | |||||
|
BAT2_MOUSE
|
||||||
| NC score | 0.034529 (rank : 39) | θ value | 1.81305 (rank : 30) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
LAD1_MOUSE
|
||||||
| NC score | 0.033844 (rank : 40) | θ value | 2.36792 (rank : 44) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P57016 | Gene names | Lad1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (Linear IgA disease autoantigen). | |||||
|
RRBP1_HUMAN
|
||||||
| NC score | 0.029924 (rank : 41) | θ value | 0.00509761 (rank : 10) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1306 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9P2E9, O75300, O75301, Q5W165, Q96SB2, Q9BWP1, Q9H476 | Gene names | RRBP1, KIAA1398 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (180 kDa ribosome receptor homolog) (ES/130-related protein). | |||||
|
C8AP2_MOUSE
|
||||||
| NC score | 0.029160 (rank : 42) | θ value | 0.62314 (rank : 22) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9WUF3, Q9CSW2 | Gene names | Casp8ap2, Flash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
LAD1_HUMAN
|
||||||
| NC score | 0.027806 (rank : 43) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O00515, O95614 | Gene names | LAD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (120 kDa linear IgA bullous dermatosis antigen) (97 kDa linear IgA bullous dermatosis antigen) (Linear IgA disease antigen homolog) (LadA). | |||||
|
K1949_MOUSE
|
||||||
| NC score | 0.027382 (rank : 44) | θ value | 1.81305 (rank : 33) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BQ30, Q3UDS6, Q8CEY9, Q8R3D8 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1949 homolog. | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.026482 (rank : 45) | θ value | 1.81305 (rank : 35) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
BCL7C_MOUSE
|
||||||
| NC score | 0.025030 (rank : 46) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O08664, Q99LS2 | Gene names | Bcl7c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell CLL/lymphoma 7 protein family member C (B-cell chronic lymphocytic leukemia/lymphoma 7C protein). | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.024428 (rank : 47) | θ value | 0.47712 (rank : 21) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
TCAL5_HUMAN
|
||||||
| NC score | 0.024123 (rank : 48) | θ value | 3.0926 (rank : 60) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
BAT2_HUMAN
|
||||||
| NC score | 0.024102 (rank : 49) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
WDR46_MOUSE
|
||||||
| NC score | 0.023242 (rank : 50) | θ value | 4.03905 (rank : 70) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z0H1 | Gene names | Wdr46, Bing4 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 46 (WD repeat protein BING4). | |||||
|
PDZD2_HUMAN
|
||||||
| NC score | 0.022960 (rank : 51) | θ value | 0.47712 (rank : 20) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O15018, Q9BXD4 | Gene names | PDZD2, AIPC, KIAA0300, PDZK3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 2 (PDZ domain-containing protein 3) (Activated in prostate cancer protein). | |||||
|
CASC3_MOUSE
|
||||||
| NC score | 0.021552 (rank : 52) | θ value | 3.0926 (rank : 53) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K3W3, Q8K219, Q99NF0 | Gene names | Casc3, Mln51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASC3 (Cancer susceptibility candidate gene 3 protein homolog) (Metastatic lymph node protein 51 homolog) (Protein MLN 51 homolog) (Protein barentsz) (Btz). | |||||
|
RFX1_MOUSE
|
||||||
| NC score | 0.021360 (rank : 53) | θ value | 1.06291 (rank : 27) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P48377 | Gene names | Rfx1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein RFX1. | |||||
|
TXND2_HUMAN
|
||||||
| NC score | 0.021051 (rank : 54) | θ value | 2.36792 (rank : 47) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
REN3A_HUMAN
|
||||||
| NC score | 0.020181 (rank : 55) | θ value | 4.03905 (rank : 69) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H1J1, Q5T8C3, Q5T8C9, Q7Z6N3, Q86YK1, Q9BZI8 | Gene names | UPF3A, RENT3A, UPF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of nonsense transcripts 3A (Nonsense mRNA reducing factor 3A) (Up-frameshift suppressor 3 homolog A) (hUpf3). | |||||
|
RRBP1_MOUSE
|
||||||
| NC score | 0.019937 (rank : 56) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
CD008_HUMAN
|
||||||
| NC score | 0.019751 (rank : 57) | θ value | 5.27518 (rank : 72) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P78312, O43607, P78311, P78313, Q9UEG8 | Gene names | C4orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C4orf8 (Protein IT14). | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.019327 (rank : 58) | θ value | 5.27518 (rank : 75) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
PSL1_HUMAN
|
||||||
| NC score | 0.016896 (rank : 59) | θ value | 2.36792 (rank : 45) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8TCT7, O60365, Q567S3, Q8IUH9, Q9BUY6, Q9H3M4, Q9NPN2, Q9P1Z6 | Gene names | SPPL2B, KIAA1532, PSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal peptide peptidase-like 2B (EC 3.4.23.-) (Protein SPP-like 2B) (Protein SPPL2b) (Intramembrane protease 4) (IMP4) (Presenilin-like protein 1). | |||||
|
TACC1_MOUSE
|
||||||
| NC score | 0.016418 (rank : 60) | θ value | 1.06291 (rank : 28) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6Y685, Q6Y686 | Gene names | Tacc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transforming acidic coiled-coil-containing protein 1. | |||||
|
ZFHX2_HUMAN
|
||||||
| NC score | 0.016247 (rank : 61) | θ value | 2.36792 (rank : 48) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1178 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9C0A1 | Gene names | ZFHX2, KIAA1762 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger homeobox protein 2. | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.016114 (rank : 62) | θ value | 3.0926 (rank : 57) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
HGS_HUMAN
|
||||||
| NC score | 0.015448 (rank : 63) | θ value | 2.36792 (rank : 43) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O14964, Q9NR36 | Gene names | HGS, HRS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate (Protein pp110) (Hrs). | |||||
|
ZN638_HUMAN
|
||||||
| NC score | 0.015191 (rank : 64) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14966, Q53R34, Q5XJ05, Q68DP3, Q6P2H2, Q7Z3T7, Q8NF92, Q8TCA1, Q9H2G1, Q9NP37 | Gene names | ZNF638, NP220, ZFML | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein) (Cutaneous T-cell lymphoma-associated antigen se33-1) (CTCL tumor antigen se33-1). | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.015037 (rank : 65) | θ value | 3.0926 (rank : 52) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
AT1B4_MOUSE
|
||||||
| NC score | 0.014837 (rank : 66) | θ value | 5.27518 (rank : 71) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99ME6, Q99ME5 | Gene names | Atp1b4 | |||
|
Domain Architecture |

|
|||||
| Description | X/potassium-transporting ATPase subunit beta-m (X,K-ATPase beta-m subunit). | |||||
|
MEFV_HUMAN
|
||||||
| NC score | 0.014823 (rank : 67) | θ value | 0.279714 (rank : 17) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O15553, Q96PN4, Q96PN5 | Gene names | MEFV, MEF | |||
|
Domain Architecture |

|
|||||
| Description | Pyrin (Marenostrin). | |||||
|
LRC50_MOUSE
|
||||||
| NC score | 0.014743 (rank : 68) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D2H9, Q9CVS9 | Gene names | Lrrc50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 50. | |||||
|
PI51C_HUMAN
|
||||||
| NC score | 0.013826 (rank : 69) | θ value | 4.03905 (rank : 67) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60331, Q7LE07 | Gene names | PIP5K1C, KIAA0589 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-4-phosphate 5-kinase type-1 gamma (EC 2.7.1.68) (Phosphatidylinositol-4-phosphate 5-kinase type I gamma) (PtdIns(4)P- 5-kinase gamma) (PtdInsPKIgamma) (PIP5KIgamma). | |||||
|
HGS_MOUSE
|
||||||
| NC score | 0.013322 (rank : 70) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q99LI8, Q61691, Q8BQW3 | Gene names | Hgs, Hrs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate. | |||||
|
ELOA3_HUMAN
|
||||||
| NC score | 0.012996 (rank : 71) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NG57 | Gene names | TCEB3C, TCEB3L2 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II transcription factor SIII subunit A3 (Elongin-A3) (EloA3) (Transcription elongation factor B polypeptide 3C). | |||||
|
WDR81_MOUSE
|
||||||
| NC score | 0.012816 (rank : 72) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5ND34, Q3TA90, Q3TUS9, Q6KAR1 | Gene names | Wdr81 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 81. | |||||
|
PRGR_HUMAN
|
||||||
| NC score | 0.012591 (rank : 73) | θ value | 4.03905 (rank : 68) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P06401, Q9UPF7 | Gene names | PGR, NR3C3 | |||
|
Domain Architecture |

|
|||||
| Description | Progesterone receptor (PR). | |||||
|
CSKI2_HUMAN
|
||||||
| NC score | 0.012234 (rank : 74) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8WXE0, Q7LG69, Q9ULT1 | Gene names | CASKIN2, KIAA1139 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-2. | |||||
|
LSD1_HUMAN
|
||||||
| NC score | 0.012101 (rank : 75) | θ value | 4.03905 (rank : 65) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O60341, Q5TH94, Q5TH95, Q86VT7, Q8IXK4, Q8NDP6, Q8TAZ3, Q96AW4 | Gene names | AOF2, KIAA0601, LSD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lysine-specific histone demethylase 1 (EC 1.-.-.-) (Flavin-containing amine oxidase domain-containing protein 2) (BRAF35-HDAC complex protein BHC110). | |||||
|
GP124_MOUSE
|
||||||
| NC score | 0.012096 (rank : 76) | θ value | 2.36792 (rank : 42) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q91ZV8 | Gene names | Gpr124, Tem5 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 124 precursor (Tumor endothelial marker 5). | |||||
|
INVS_HUMAN
|
||||||
| NC score | 0.011242 (rank : 77) | θ value | 1.38821 (rank : 29) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Y283, Q5W0T6, Q8IVX8, Q9BRB9, Q9Y488, Q9Y498 | Gene names | INVS, INV, NPHP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning homolog) (Nephrocystin-2). | |||||
|
SYTL2_HUMAN
|
||||||
| NC score | 0.010916 (rank : 78) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9HCH5, Q2YDA7, Q8ND34, Q96BJ2, Q9H768, Q9NXM1 | Gene names | SYTL2, KIAA1597, SLP2 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 2 (Exophilin-4). | |||||
|
CSKI2_MOUSE
|
||||||
| NC score | 0.009648 (rank : 79) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 812 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8VHK1, Q6ZPX1, Q7TS77 | Gene names | Caskin2, Kiaa1139 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-2. | |||||
|
SASH1_HUMAN
|
||||||
| NC score | 0.008666 (rank : 80) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O94885, Q8TAI0, Q9H7R7 | Gene names | SASH1, KIAA0790, PEPE1 | |||
|
Domain Architecture |

|
|||||
| Description | SAM and SH3 domain-containing protein 1 (Proline-glutamate repeat- containing protein). | |||||
|
EHMT2_MOUSE
|
||||||
| NC score | 0.008422 (rank : 81) | θ value | 5.27518 (rank : 73) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Z148, Q6PE08, Q8K4R6, Q8K4R7, Q9Z149 | Gene names | Ehmt2, Bat8, G9a, Ng36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
RP3A_MOUSE
|
||||||
| NC score | 0.007922 (rank : 82) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 510 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P47708 | Gene names | Rph3a | |||
|
Domain Architecture |

|
|||||
| Description | Rabphilin-3A (Exophilin-1). | |||||
|
WNK2_HUMAN
|
||||||
| NC score | 0.007107 (rank : 83) | θ value | 1.81305 (rank : 39) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
ES8L1_MOUSE
|
||||||
| NC score | 0.005533 (rank : 84) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R5F8, Q8R0D6, Q9D2M6 | Gene names | Eps8l1, Eps8r1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Epidermal growth factor receptor kinase substrate 8-like protein 1 (Epidermal growth factor receptor pathway substrate 8-related protein 1) (EPS8-like protein 1). | |||||
|
ZBT45_HUMAN
|
||||||
| NC score | 0.004887 (rank : 85) | θ value | 1.81305 (rank : 40) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 901 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96K62 | Gene names | ZBTB45, ZNF499 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 45 (Zinc finger protein 499). | |||||
|
MRCKG_HUMAN
|
||||||
| NC score | 0.004033 (rank : 86) | θ value | 0.62314 (rank : 24) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1519 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6DT37, O00565 | Gene names | CDC42BPG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK gamma (EC 2.7.11.1) (CDC42- binding protein kinase gamma) (Myotonic dystrophy kinase-related CDC42-binding kinase gamma) (Myotonic dystrophy protein kinase-like alpha) (MRCK gamma) (DMPK-like gamma). | |||||
|
KC1D_HUMAN
|
||||||
| NC score | 0.003647 (rank : 87) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 737 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P48730, Q96KZ6, Q9BTN5 | Gene names | CSNK1D | |||
|
Domain Architecture |

|
|||||
| Description | Casein kinase I isoform delta (EC 2.7.11.1) (CKI-delta) (CKId). | |||||
|
KC1D_MOUSE
|
||||||
| NC score | 0.003647 (rank : 88) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 737 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DC28, Q3TZK2, Q99KK4 | Gene names | Csnk1d | |||
|
Domain Architecture |

|
|||||
| Description | Casein kinase I isoform delta (EC 2.7.11.1) (CKI-delta) (CKId). | |||||
|
KSR1_MOUSE
|
||||||
| NC score | 0.003437 (rank : 89) | θ value | 3.0926 (rank : 54) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q61097, Q61648, Q78DX8 | Gene names | Ksr1, Ksr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinase suppressor of ras-1 (Kinase suppressor of ras) (mKSR1) (Hb protein). | |||||
|
PAK4_MOUSE
|
||||||
| NC score | 0.001722 (rank : 90) | θ value | 4.03905 (rank : 66) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1098 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BTW9, Q6ZPX0, Q80Z97, Q9CS71 | Gene names | Pak4, Kiaa1142 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PAK 4 (EC 2.7.11.1) (p21-activated kinase 4) (PAK-4). | |||||
|
BRSK1_HUMAN
|
||||||
| NC score | 0.001173 (rank : 91) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 942 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8TDC3, Q5J5B5, Q8NDD0, Q8NDR4, Q8TDC2, Q96AV4, Q96JL4 | Gene names | BRSK1, KIAA1811, SAD1 | |||
|
Domain Architecture |

|
|||||
| Description | BR serine/threonine-protein kinase 1 (EC 2.7.11.1) (SAD1 kinase) (SAD1A). | |||||
|
ZN335_HUMAN
|
||||||
| NC score | 0.000689 (rank : 92) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 973 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H4Z2, Q9H684 | Gene names | ZNF335 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 335. | |||||
|
MYLK_HUMAN
|
||||||
| NC score | 0.000184 (rank : 93) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 91 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q15746, O95796, O95797, O95798, O95799, Q14844, Q16794, Q5MY99, Q5MYA0, Q7Z4J0, Q9C0L5, Q9UBG5, Q9UIT9 | Gene names | MYLK, MLCK | |||
|
Domain Architecture |

|
|||||
| Description | Myosin light chain kinase, smooth muscle (EC 2.7.11.18) (MLCK) (Telokin) (Kinase-related protein) (KRP). | |||||