Please be patient as the page loads
|
PR285_HUMAN
|
||||||
| SwissProt Accessions | Q9BYK8, Q3C2G2, Q4VXQ1, Q8TEF3, Q96ND3, Q9C094 | Gene names | PRIC285, KIAA1769 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal proliferator-activated receptor A-interacting complex 285 kDa protein (EC 3.6.1.-) (ATP-dependent helicase PRIC285) (PPAR-alpha- interacting complex protein 285) (PPAR-gamma DBD-interacting protein 1) (PDIP1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PR285_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9BYK8, Q3C2G2, Q4VXQ1, Q8TEF3, Q96ND3, Q9C094 | Gene names | PRIC285, KIAA1769 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal proliferator-activated receptor A-interacting complex 285 kDa protein (EC 3.6.1.-) (ATP-dependent helicase PRIC285) (PPAR-alpha- interacting complex protein 285) (PPAR-gamma DBD-interacting protein 1) (PDIP1). | |||||
|
HELZ_HUMAN
|
||||||
| θ value | 2.20485e-131 (rank : 2) | NC score | 0.851997 (rank : 2) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P42694 | Gene names | HELZ, KIAA0054 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable helicase with zinc-finger domain (EC 3.6.1.-). | |||||
|
RENT1_HUMAN
|
||||||
| θ value | 2.18568e-46 (rank : 3) | NC score | 0.783493 (rank : 3) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q92900, O00239, O43343, Q86Z25, Q92842 | Gene names | UPF1, KIAA0221, RENT1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of nonsense transcripts 1 (EC 3.6.1.-) (ATP-dependent helicase RENT1) (Nonsense mRNA reducing factor 1) (NORF1) (Up- frameshift suppressor 1 homolog) (hUpf1). | |||||
|
RENT1_MOUSE
|
||||||
| θ value | 2.85459e-46 (rank : 4) | NC score | 0.783328 (rank : 4) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9EPU0, Q99PR4 | Gene names | Upf1, Rent1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of nonsense transcripts 1 (EC 3.6.1.-) (ATP-dependent helicase RENT1) (Nonsense mRNA reducing factor 1) (NORF1) (Up- frameshift suppressor 1 homolog) (mUpf1). | |||||
|
M10L1_HUMAN
|
||||||
| θ value | 1.37539e-32 (rank : 5) | NC score | 0.729842 (rank : 5) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9BXT6, Q5TGD5, Q8NBD4, Q9NXW3, Q9UFB3, Q9UGX9 | Gene names | MOV10L1 | |||
|
Domain Architecture |

|
|||||
| Description | Putative helicase Mov10l1 (EC 3.6.1.-) (Moloney leukemia virus 10-like protein 1) (MOV10-like 1). | |||||
|
M10L1_MOUSE
|
||||||
| θ value | 9.8567e-31 (rank : 6) | NC score | 0.724986 (rank : 8) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q99MV5, Q7TPA9, Q8C3W0, Q924C2 | Gene names | Mov10l1, Champ | |||
|
Domain Architecture |

|
|||||
| Description | Putative helicase Mov10l1 (EC 3.6.1.-) (Moloney leukemia virus 10-like protein 1) (MOV10-like 1) (Cardiac helicase activated by MEF2 protein) (Cardiac-specific RNA helicase). | |||||
|
SMBP2_MOUSE
|
||||||
| θ value | 8.3442e-30 (rank : 7) | NC score | 0.714336 (rank : 9) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P40694 | Gene names | Ighmbp2, Smbp-2, Smbp2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein SMUBP-2 (EC 3.6.1.-) (ATP-dependent helicase IGHMBP2) (Immunoglobulin mu-binding protein 2) (SMUBP-2) (Cardiac transcription factor 1) (CATF1). | |||||
|
MOV10_MOUSE
|
||||||
| θ value | 1.12775e-26 (rank : 8) | NC score | 0.726550 (rank : 6) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P23249, Q9DC64 | Gene names | Mov10, Gb110 | |||
|
Domain Architecture |

|
|||||
| Description | Putative helicase MOV-10 (EC 3.6.1.-) (Moloney leukemia virus 10 protein). | |||||
|
MOV10_HUMAN
|
||||||
| θ value | 2.51237e-26 (rank : 9) | NC score | 0.725220 (rank : 7) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9HCE1, Q8TEF0, Q9BSY3, Q9BUJ9 | Gene names | MOV10, KIAA1631 | |||
|
Domain Architecture |

|
|||||
| Description | Putative helicase MOV-10 (EC 3.6.1.-) (Moloney leukemia virus 10 protein). | |||||
|
SMBP2_HUMAN
|
||||||
| θ value | 3.62785e-25 (rank : 10) | NC score | 0.708850 (rank : 10) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P38935, Q00443, Q14177 | Gene names | IGHMBP2, SMBP2, SMUBP2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein SMUBP-2 (EC 3.6.1.-) (ATP-dependent helicase IGHMBP2) (Immunoglobulin mu-binding protein 2) (SMUBP-2) (Glial factor 1) (GF-1). | |||||
|
RRP44_HUMAN
|
||||||
| θ value | 3.07116e-24 (rank : 11) | NC score | 0.362803 (rank : 16) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y2L1, Q5W0P7, Q5W0P8, Q658Z7, Q7Z481, Q8WWI2, Q9UG36 | Gene names | DIS3, KIAA1008, RRP44 | |||
|
Domain Architecture |

|
|||||
| Description | Exosome complex exonuclease RRP44 (EC 3.1.13.-) (Ribosomal RNA- processing protein 44) (DIS3 protein homolog). | |||||
|
SETX_HUMAN
|
||||||
| θ value | 6.84181e-24 (rank : 12) | NC score | 0.636749 (rank : 11) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q7Z333, O75120, Q3KQX4, Q68DW5, Q6AZD7, Q7Z3J6, Q8WX33, Q9H9D1, Q9NVP9 | Gene names | SETX, ALS4, KIAA0625, SCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable helicase senataxin (EC 3.6.1.-) (SEN1 homolog). | |||||
|
ZNFX1_MOUSE
|
||||||
| θ value | 3.07829e-16 (rank : 13) | NC score | 0.463239 (rank : 12) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 407 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8R151, Q3U016, Q3UD71, Q3UEZ0 | Gene names | ZNFX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
ZNFX1_HUMAN
|
||||||
| θ value | 1.29331e-14 (rank : 14) | NC score | 0.460263 (rank : 13) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9P2E3, Q9BQM7, Q9BQM8, Q9H8C1, Q9H9S2, Q9NUM1, Q9NWW1 | Gene names | ZNFX1, KIAA1404 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
AQR_HUMAN
|
||||||
| θ value | 1.06045e-08 (rank : 15) | NC score | 0.441107 (rank : 15) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O60306, Q2YDX9, Q6IRU8, Q6PIC8 | Gene names | AQR, KIAA0560 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intron-binding protein aquarius (Intron-binding protein of 160 kDa) (IBP160). | |||||
|
AQR_MOUSE
|
||||||
| θ value | 1.06045e-08 (rank : 16) | NC score | 0.443960 (rank : 14) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CFQ3, P97871, Q3U9N1, Q3ULE8, Q80TX8 | Gene names | Aqr, Kiaa0560 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intron-binding protein aquarius. | |||||
|
Z3H7A_HUMAN
|
||||||
| θ value | 5.81887e-07 (rank : 17) | NC score | 0.170864 (rank : 18) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IWR0, Q9NPE9 | Gene names | ZC3H7A, ZC3H7, ZC3HDC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH-type domain-containing protein 7A. | |||||
|
Z3H7B_HUMAN
|
||||||
| θ value | 9.29e-05 (rank : 18) | NC score | 0.147465 (rank : 19) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UGR2, Q5TFX9, Q8TBT9, Q9H8B6, Q9UGQ9, Q9UGR0, Q9UGR1, Q9UK03, Q9UPW9 | Gene names | ZC3H7B, KIAA1031 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger CCCH-type domain-containing protein 7B (Rotavirus 'X'- associated non-structural protein) (RoXaN). | |||||
|
CLIC6_HUMAN
|
||||||
| θ value | 0.125558 (rank : 19) | NC score | 0.028652 (rank : 35) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96NY7, Q8IX31 | Gene names | CLIC6, CLIC1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel 6. | |||||
|
NEBU_HUMAN
|
||||||
| θ value | 0.365318 (rank : 20) | NC score | 0.025551 (rank : 37) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P20929, Q15346 | Gene names | NEB | |||
|
Domain Architecture |
|
|||||
| Description | Nebulin. | |||||
|
STRC_HUMAN
|
||||||
| θ value | 0.47712 (rank : 21) | NC score | 0.040794 (rank : 30) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7RTU9 | Gene names | STRC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stereocilin precursor. | |||||
|
NAGAB_MOUSE
|
||||||
| θ value | 0.813845 (rank : 22) | NC score | 0.034472 (rank : 33) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QWR8, O88620, Q8R437, Q8VDK2 | Gene names | Naga | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-N-acetylgalactosaminidase precursor (EC 3.2.1.49) (Alpha- galactosidase B). | |||||
|
RFC4_HUMAN
|
||||||
| θ value | 0.813845 (rank : 23) | NC score | 0.073827 (rank : 21) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P35249, Q6FHX7 | Gene names | RFC4 | |||
|
Domain Architecture |
|
|||||
| Description | Replication factor C subunit 4 (Replication factor C 37 kDa subunit) (RF-C 37 kDa subunit) (RFC37) (Activator 1 37 kDa subunit) (A1 37 kDa subunit). | |||||
|
RFC4_MOUSE
|
||||||
| θ value | 0.813845 (rank : 24) | NC score | 0.075108 (rank : 20) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99J62 | Gene names | Rfc4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Replication factor C subunit 4. | |||||
|
ZN473_MOUSE
|
||||||
| θ value | 1.06291 (rank : 25) | NC score | 0.000746 (rank : 69) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 824 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BI67, Q8BI98, Q8BIB7 | Gene names | Znf473, Zfp100, Zfp473 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 473 homolog (Zinc finger protein 100) (Zfp-100). | |||||
|
ZN638_HUMAN
|
||||||
| θ value | 1.06291 (rank : 26) | NC score | 0.040046 (rank : 31) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q14966, Q53R34, Q5XJ05, Q68DP3, Q6P2H2, Q7Z3T7, Q8NF92, Q8TCA1, Q9H2G1, Q9NP37 | Gene names | ZNF638, NP220, ZFML | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein) (Cutaneous T-cell lymphoma-associated antigen se33-1) (CTCL tumor antigen se33-1). | |||||
|
ZN638_MOUSE
|
||||||
| θ value | 1.06291 (rank : 27) | NC score | 0.038436 (rank : 32) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61464, Q6DFV9, Q8C941 | Gene names | Znf638, Np220, Zfml | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein). | |||||
|
ITA2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 28) | NC score | 0.010912 (rank : 57) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q62469, Q62163 | Gene names | Itga2 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-2 precursor (Platelet membrane glycoprotein Ia) (GPIa) (Collagen receptor) (VLA-2 alpha chain) (CD49b antigen). | |||||
|
RFC5_HUMAN
|
||||||
| θ value | 1.38821 (rank : 29) | NC score | 0.069707 (rank : 23) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P40937 | Gene names | RFC5 | |||
|
Domain Architecture |
|
|||||
| Description | Replication factor C subunit 5 (Replication factor C 36 kDa subunit) (RF-C 36 kDa subunit) (RFC36) (Activator 1 36 kDa subunit) (A1 36 kDa subunit). | |||||
|
RFC5_MOUSE
|
||||||
| θ value | 1.38821 (rank : 30) | NC score | 0.072529 (rank : 22) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9D0F6 | Gene names | Rfc5 | |||
|
Domain Architecture |
|
|||||
| Description | Replication factor C subunit 5 (Replication factor C 36 kDa subunit) (RF-C 36 kDa subunit) (RFC36) (Activator 1 36 kDa subunit) (A1 36 kDa subunit). | |||||
|
ZN142_HUMAN
|
||||||
| θ value | 1.38821 (rank : 31) | NC score | 0.002522 (rank : 64) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P52746, Q92510 | Gene names | ZNF142, KIAA0236 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 142 (HA4654). | |||||
|
AKAP2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 32) | NC score | 0.025505 (rank : 38) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O54931, O54932, O54933 | Gene names | Akap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2) (AKAP expressed in kidney and lung) (AKAP-KL). | |||||
|
BRPF3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 33) | NC score | 0.013120 (rank : 52) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ULD4, Q5R3K8 | Gene names | BRPF3, KIAA1286 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and PHD finger-containing protein 3. | |||||
|
CIZ1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 34) | NC score | 0.031403 (rank : 34) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9ULV3, Q9NYM8, Q9UHK4, Q9Y3F9, Q9Y3G0 | Gene names | CIZ1, LSFR1, NP94 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cip1-interacting zinc finger protein (Nuclear protein NP94). | |||||
|
TIF1B_MOUSE
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.016697 (rank : 44) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q62318, P70391, Q8C283, Q99PN4 | Gene names | Trim28, Krip1, Tif1b | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-beta (TIF1-beta) (Tripartite motif-containing protein 28) (KRAB-A-interacting protein) (KRIP-1). | |||||
|
ARMX5_MOUSE
|
||||||
| θ value | 3.0926 (rank : 36) | NC score | 0.016307 (rank : 46) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3UZB0 | Gene names | Armcx5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 5. | |||||
|
EPIPL_MOUSE
|
||||||
| θ value | 3.0926 (rank : 37) | NC score | 0.014312 (rank : 48) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R0W0 | Gene names | Eppk1 | |||
|
Domain Architecture |

|
|||||
| Description | Epiplakin. | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 38) | NC score | 0.019805 (rank : 40) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
MAST4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 39) | NC score | 0.002017 (rank : 66) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 4.03905 (rank : 40) | NC score | 0.007277 (rank : 60) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
TIF1A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 41) | NC score | 0.012636 (rank : 53) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q64127, Q64126 | Gene names | Trim24, Tif1, Tif1a | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24). | |||||
|
TNF10_MOUSE
|
||||||
| θ value | 4.03905 (rank : 42) | NC score | 0.019494 (rank : 41) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P50592 | Gene names | Tnfsf10, Trail | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor ligand superfamily member 10 (TNF-related apoptosis-inducing ligand) (Protein TRAIL) (CD253 antigen). | |||||
|
ASAHL_HUMAN
|
||||||
| θ value | 5.27518 (rank : 43) | NC score | 0.016374 (rank : 45) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q02083, Q5KTF2, Q96EY2, Q9BRA8 | Gene names | ASAHL, NAAA, PLT | |||
|
Domain Architecture |
|
|||||
| Description | N-acylethanolamine-hydrolyzing acid amidase precursor (EC 3.5.1.-) (N- acylsphingosine amidohydrolase-like) (ASAH-like protein) (Acid ceramidase-like protein). | |||||
|
EDD1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 44) | NC score | 0.014261 (rank : 49) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80TP3, Q698K9, Q6PEQ8, Q6PFQ9, Q80VL4, Q810V6, Q9CXE9 | Gene names | Edd1, Edd, Kiaa0896 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog). | |||||
|
FBN1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 45) | NC score | 0.001926 (rank : 67) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P35555, Q15972, Q75N87 | Gene names | FBN1, FBN | |||
|
Domain Architecture |

|
|||||
| Description | Fibrillin-1 precursor. | |||||
|
FUT4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 46) | NC score | 0.011280 (rank : 56) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P22083 | Gene names | FUT4, ELFT | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-(1,3)-fucosyltransferase (EC 2.4.1.-) (Galactoside 3-L- fucosyltransferase) (Fucosyltransferase 4) (FUCT-IV) (Fuc-TIV) (ELAM-1 ligand fucosyltransferase). | |||||
|
YETS2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 47) | NC score | 0.013623 (rank : 50) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ULM3, Q641P6, Q9NW96 | Gene names | YEATS2, KIAA1197 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YEATS domain-containing protein 2. | |||||
|
ZN622_MOUSE
|
||||||
| θ value | 5.27518 (rank : 48) | NC score | 0.018234 (rank : 43) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91VY9 | Gene names | Znf622, D15Ertd806e, Zfp622 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 622. | |||||
|
AKAP2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 49) | NC score | 0.019095 (rank : 42) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y2D5, Q9UG26 | Gene names | AKAP2, KIAA0920 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2). | |||||
|
DDX51_HUMAN
|
||||||
| θ value | 6.88961 (rank : 50) | NC score | 0.005515 (rank : 61) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
EDD1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 51) | NC score | 0.013431 (rank : 51) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95071, O94970, Q9NPL3 | Gene names | EDD1, EDD, HYD, KIAA0896 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog) (hHYD) (Progestin-induced protein). | |||||
|
FBX32_HUMAN
|
||||||
| θ value | 6.88961 (rank : 52) | NC score | 0.011781 (rank : 55) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q969P5 | Gene names | FBXO32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 32 (Muscle atrophy F-box protein) (MAFbx) (Atrogin- 1). | |||||
|
GCP5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 53) | NC score | 0.016280 (rank : 47) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96RT8, Q96PY8 | Gene names | TUBGCP5, GCP5, KIAA1899 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-tubulin complex component 5 (GCP-5). | |||||
|
NAGAB_HUMAN
|
||||||
| θ value | 6.88961 (rank : 54) | NC score | 0.023577 (rank : 39) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P17050 | Gene names | NAGA | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-N-acetylgalactosaminidase precursor (EC 3.2.1.49) (Alpha- galactosidase B). | |||||
|
SPTN2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 55) | NC score | 0.003795 (rank : 62) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 535 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15020, O14872, O14873 | Gene names | SPTBN2, KIAA0302 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 2 (Spectrin, non-erythroid beta chain 2) (Beta-III spectrin). | |||||
|
STRC_MOUSE
|
||||||
| θ value | 6.88961 (rank : 56) | NC score | 0.028143 (rank : 36) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VIM6 | Gene names | Strc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stereocilin precursor. | |||||
|
AAKG3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 57) | NC score | 0.008106 (rank : 59) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BGM7, Q80WK8, Q8CJ41 | Gene names | Prkag3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 5'-AMP-activated protein kinase subunit gamma-3 (AMPK gamma-3 chain) (AMPK gamma3). | |||||
|
ATX2L_MOUSE
|
||||||
| θ value | 8.99809 (rank : 58) | NC score | 0.010493 (rank : 58) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7TQH0, Q80XN9, Q8K059 | Gene names | Atxn2l, A2lp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2-like protein. | |||||
|
GSHB_HUMAN
|
||||||
| θ value | 8.99809 (rank : 59) | NC score | 0.012414 (rank : 54) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P48637, Q4TTD9 | Gene names | GSS | |||
|
Domain Architecture |
|
|||||
| Description | Glutathione synthetase (EC 6.3.2.3) (Glutathione synthase) (GSH synthetase) (GSH-S). | |||||
|
SHAN1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 60) | NC score | 0.002282 (rank : 65) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y566, Q9NYW9 | Gene names | SHANK1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 1 (Shank1) (Somatostatin receptor-interacting protein) (SSTR-interacting protein) (SSTRIP). | |||||
|
WD51A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 61) | NC score | 0.000945 (rank : 68) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8JZX3, Q9CY09 | Gene names | Wdr51a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 51A. | |||||
|
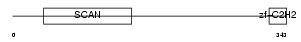
ZN174_HUMAN
|
||||||
| θ value | 8.99809 (rank : 62) | NC score | -0.000752 (rank : 70) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q15697, Q9BQ34 | Gene names | ZNF174, ZSCAN8 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 174 (AW-1) (Zinc finger and SCAN domain-containing protein 8). | |||||
|
ZN451_MOUSE
|
||||||
| θ value | 8.99809 (rank : 63) | NC score | 0.003312 (rank : 63) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 643 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8C0P7 | Gene names | Znf451, Zfp451 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 451. | |||||
|
CX048_HUMAN
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.197890 (rank : 17) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8WUE5, Q9NWY8 | Gene names | CXorf48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CXorf48 (Tumor antigen BJ-HCC-20). | |||||
|
SPR2B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.060461 (rank : 26) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O70554, Q7TSB2 | Gene names | Sprr2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small proline-rich protein 2B. | |||||
|
SPR2D_MOUSE
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.061973 (rank : 24) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O70555 | Gene names | Sprr2d | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small proline-rich protein 2D. | |||||
|
SPR2E_HUMAN
|
||||||
| θ value | θ > 10 (rank : 67) | NC score | 0.056560 (rank : 28) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P22531, Q96RM2 | Gene names | SPRR2E | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small proline-rich protein 2E (SPR-2E) (Small proline-rich protein II) (SPR-II). | |||||
|
SPR2F_HUMAN
|
||||||
| θ value | θ > 10 (rank : 68) | NC score | 0.060637 (rank : 25) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96RM1 | Gene names | SPRR2F | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small proline-rich protein 2F (SPR-2F). | |||||
|
SPR2G_MOUSE
|
||||||
| θ value | θ > 10 (rank : 69) | NC score | 0.060150 (rank : 27) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O70558 | Gene names | Sprr2g | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small proline-rich protein 2G. | |||||
|
SPR2H_MOUSE
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.053769 (rank : 29) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70559 | Gene names | Sprr2h | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small proline-rich protein 2H. | |||||
|
PR285_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9BYK8, Q3C2G2, Q4VXQ1, Q8TEF3, Q96ND3, Q9C094 | Gene names | PRIC285, KIAA1769 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal proliferator-activated receptor A-interacting complex 285 kDa protein (EC 3.6.1.-) (ATP-dependent helicase PRIC285) (PPAR-alpha- interacting complex protein 285) (PPAR-gamma DBD-interacting protein 1) (PDIP1). | |||||
|
HELZ_HUMAN
|
||||||
| NC score | 0.851997 (rank : 2) | θ value | 2.20485e-131 (rank : 2) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P42694 | Gene names | HELZ, KIAA0054 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable helicase with zinc-finger domain (EC 3.6.1.-). | |||||
|
RENT1_HUMAN
|
||||||
| NC score | 0.783493 (rank : 3) | θ value | 2.18568e-46 (rank : 3) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q92900, O00239, O43343, Q86Z25, Q92842 | Gene names | UPF1, KIAA0221, RENT1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of nonsense transcripts 1 (EC 3.6.1.-) (ATP-dependent helicase RENT1) (Nonsense mRNA reducing factor 1) (NORF1) (Up- frameshift suppressor 1 homolog) (hUpf1). | |||||
|
RENT1_MOUSE
|
||||||
| NC score | 0.783328 (rank : 4) | θ value | 2.85459e-46 (rank : 4) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9EPU0, Q99PR4 | Gene names | Upf1, Rent1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of nonsense transcripts 1 (EC 3.6.1.-) (ATP-dependent helicase RENT1) (Nonsense mRNA reducing factor 1) (NORF1) (Up- frameshift suppressor 1 homolog) (mUpf1). | |||||
|
M10L1_HUMAN
|
||||||
| NC score | 0.729842 (rank : 5) | θ value | 1.37539e-32 (rank : 5) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9BXT6, Q5TGD5, Q8NBD4, Q9NXW3, Q9UFB3, Q9UGX9 | Gene names | MOV10L1 | |||
|
Domain Architecture |

|
|||||
| Description | Putative helicase Mov10l1 (EC 3.6.1.-) (Moloney leukemia virus 10-like protein 1) (MOV10-like 1). | |||||
|
MOV10_MOUSE
|
||||||
| NC score | 0.726550 (rank : 6) | θ value | 1.12775e-26 (rank : 8) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P23249, Q9DC64 | Gene names | Mov10, Gb110 | |||
|
Domain Architecture |

|
|||||
| Description | Putative helicase MOV-10 (EC 3.6.1.-) (Moloney leukemia virus 10 protein). | |||||
|
MOV10_HUMAN
|
||||||
| NC score | 0.725220 (rank : 7) | θ value | 2.51237e-26 (rank : 9) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9HCE1, Q8TEF0, Q9BSY3, Q9BUJ9 | Gene names | MOV10, KIAA1631 | |||
|
Domain Architecture |

|
|||||
| Description | Putative helicase MOV-10 (EC 3.6.1.-) (Moloney leukemia virus 10 protein). | |||||
|
M10L1_MOUSE
|
||||||
| NC score | 0.724986 (rank : 8) | θ value | 9.8567e-31 (rank : 6) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q99MV5, Q7TPA9, Q8C3W0, Q924C2 | Gene names | Mov10l1, Champ | |||
|
Domain Architecture |

|
|||||
| Description | Putative helicase Mov10l1 (EC 3.6.1.-) (Moloney leukemia virus 10-like protein 1) (MOV10-like 1) (Cardiac helicase activated by MEF2 protein) (Cardiac-specific RNA helicase). | |||||
|
SMBP2_MOUSE
|
||||||
| NC score | 0.714336 (rank : 9) | θ value | 8.3442e-30 (rank : 7) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P40694 | Gene names | Ighmbp2, Smbp-2, Smbp2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein SMUBP-2 (EC 3.6.1.-) (ATP-dependent helicase IGHMBP2) (Immunoglobulin mu-binding protein 2) (SMUBP-2) (Cardiac transcription factor 1) (CATF1). | |||||
|
SMBP2_HUMAN
|
||||||
| NC score | 0.708850 (rank : 10) | θ value | 3.62785e-25 (rank : 10) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P38935, Q00443, Q14177 | Gene names | IGHMBP2, SMBP2, SMUBP2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein SMUBP-2 (EC 3.6.1.-) (ATP-dependent helicase IGHMBP2) (Immunoglobulin mu-binding protein 2) (SMUBP-2) (Glial factor 1) (GF-1). | |||||
|
SETX_HUMAN
|
||||||
| NC score | 0.636749 (rank : 11) | θ value | 6.84181e-24 (rank : 12) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q7Z333, O75120, Q3KQX4, Q68DW5, Q6AZD7, Q7Z3J6, Q8WX33, Q9H9D1, Q9NVP9 | Gene names | SETX, ALS4, KIAA0625, SCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable helicase senataxin (EC 3.6.1.-) (SEN1 homolog). | |||||
|
ZNFX1_MOUSE
|
||||||
| NC score | 0.463239 (rank : 12) | θ value | 3.07829e-16 (rank : 13) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 407 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8R151, Q3U016, Q3UD71, Q3UEZ0 | Gene names | ZNFX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
ZNFX1_HUMAN
|
||||||
| NC score | 0.460263 (rank : 13) | θ value | 1.29331e-14 (rank : 14) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9P2E3, Q9BQM7, Q9BQM8, Q9H8C1, Q9H9S2, Q9NUM1, Q9NWW1 | Gene names | ZNFX1, KIAA1404 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
AQR_MOUSE
|
||||||
| NC score | 0.443960 (rank : 14) | θ value | 1.06045e-08 (rank : 16) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CFQ3, P97871, Q3U9N1, Q3ULE8, Q80TX8 | Gene names | Aqr, Kiaa0560 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intron-binding protein aquarius. | |||||
|
AQR_HUMAN
|
||||||
| NC score | 0.441107 (rank : 15) | θ value | 1.06045e-08 (rank : 15) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O60306, Q2YDX9, Q6IRU8, Q6PIC8 | Gene names | AQR, KIAA0560 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intron-binding protein aquarius (Intron-binding protein of 160 kDa) (IBP160). | |||||
|
RRP44_HUMAN
|
||||||
| NC score | 0.362803 (rank : 16) | θ value | 3.07116e-24 (rank : 11) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y2L1, Q5W0P7, Q5W0P8, Q658Z7, Q7Z481, Q8WWI2, Q9UG36 | Gene names | DIS3, KIAA1008, RRP44 | |||
|
Domain Architecture |

|
|||||
| Description | Exosome complex exonuclease RRP44 (EC 3.1.13.-) (Ribosomal RNA- processing protein 44) (DIS3 protein homolog). | |||||
|
CX048_HUMAN
|
||||||
| NC score | 0.197890 (rank : 17) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8WUE5, Q9NWY8 | Gene names | CXorf48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CXorf48 (Tumor antigen BJ-HCC-20). | |||||
|
Z3H7A_HUMAN
|
||||||
| NC score | 0.170864 (rank : 18) | θ value | 5.81887e-07 (rank : 17) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IWR0, Q9NPE9 | Gene names | ZC3H7A, ZC3H7, ZC3HDC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH-type domain-containing protein 7A. | |||||
|
Z3H7B_HUMAN
|
||||||
| NC score | 0.147465 (rank : 19) | θ value | 9.29e-05 (rank : 18) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UGR2, Q5TFX9, Q8TBT9, Q9H8B6, Q9UGQ9, Q9UGR0, Q9UGR1, Q9UK03, Q9UPW9 | Gene names | ZC3H7B, KIAA1031 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger CCCH-type domain-containing protein 7B (Rotavirus 'X'- associated non-structural protein) (RoXaN). | |||||
|
RFC4_MOUSE
|
||||||
| NC score | 0.075108 (rank : 20) | θ value | 0.813845 (rank : 24) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99J62 | Gene names | Rfc4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Replication factor C subunit 4. | |||||
|
RFC4_HUMAN
|
||||||
| NC score | 0.073827 (rank : 21) | θ value | 0.813845 (rank : 23) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P35249, Q6FHX7 | Gene names | RFC4 | |||
|
Domain Architecture |

|
|||||
| Description | Replication factor C subunit 4 (Replication factor C 37 kDa subunit) (RF-C 37 kDa subunit) (RFC37) (Activator 1 37 kDa subunit) (A1 37 kDa subunit). | |||||
|
RFC5_MOUSE
|
||||||
| NC score | 0.072529 (rank : 22) | θ value | 1.38821 (rank : 30) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9D0F6 | Gene names | Rfc5 | |||
|
Domain Architecture |

|
|||||
| Description | Replication factor C subunit 5 (Replication factor C 36 kDa subunit) (RF-C 36 kDa subunit) (RFC36) (Activator 1 36 kDa subunit) (A1 36 kDa subunit). | |||||
|
RFC5_HUMAN
|
||||||
| NC score | 0.069707 (rank : 23) | θ value | 1.38821 (rank : 29) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P40937 | Gene names | RFC5 | |||
|
Domain Architecture |

|
|||||
| Description | Replication factor C subunit 5 (Replication factor C 36 kDa subunit) (RF-C 36 kDa subunit) (RFC36) (Activator 1 36 kDa subunit) (A1 36 kDa subunit). | |||||
|
SPR2D_MOUSE
|
||||||
| NC score | 0.061973 (rank : 24) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O70555 | Gene names | Sprr2d | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small proline-rich protein 2D. | |||||
|
SPR2F_HUMAN
|
||||||
| NC score | 0.060637 (rank : 25) | θ value | θ > 10 (rank : 68) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96RM1 | Gene names | SPRR2F | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small proline-rich protein 2F (SPR-2F). | |||||
|
SPR2B_MOUSE
|
||||||
| NC score | 0.060461 (rank : 26) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O70554, Q7TSB2 | Gene names | Sprr2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small proline-rich protein 2B. | |||||
|
SPR2G_MOUSE
|
||||||
| NC score | 0.060150 (rank : 27) | θ value | θ > 10 (rank : 69) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O70558 | Gene names | Sprr2g | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small proline-rich protein 2G. | |||||
|
SPR2E_HUMAN
|
||||||
| NC score | 0.056560 (rank : 28) | θ value | θ > 10 (rank : 67) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P22531, Q96RM2 | Gene names | SPRR2E | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small proline-rich protein 2E (SPR-2E) (Small proline-rich protein II) (SPR-II). | |||||
|
SPR2H_MOUSE
|
||||||
| NC score | 0.053769 (rank : 29) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70559 | Gene names | Sprr2h | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small proline-rich protein 2H. | |||||
|
STRC_HUMAN
|
||||||
| NC score | 0.040794 (rank : 30) | θ value | 0.47712 (rank : 21) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7RTU9 | Gene names | STRC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stereocilin precursor. | |||||
|
ZN638_HUMAN
|
||||||
| NC score | 0.040046 (rank : 31) | θ value | 1.06291 (rank : 26) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q14966, Q53R34, Q5XJ05, Q68DP3, Q6P2H2, Q7Z3T7, Q8NF92, Q8TCA1, Q9H2G1, Q9NP37 | Gene names | ZNF638, NP220, ZFML | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein) (Cutaneous T-cell lymphoma-associated antigen se33-1) (CTCL tumor antigen se33-1). | |||||
|
ZN638_MOUSE
|
||||||
| NC score | 0.038436 (rank : 32) | θ value | 1.06291 (rank : 27) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61464, Q6DFV9, Q8C941 | Gene names | Znf638, Np220, Zfml | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein). | |||||
|
NAGAB_MOUSE
|
||||||
| NC score | 0.034472 (rank : 33) | θ value | 0.813845 (rank : 22) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QWR8, O88620, Q8R437, Q8VDK2 | Gene names | Naga | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-N-acetylgalactosaminidase precursor (EC 3.2.1.49) (Alpha- galactosidase B). | |||||
|
CIZ1_HUMAN
|
||||||
| NC score | 0.031403 (rank : 34) | θ value | 2.36792 (rank : 34) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9ULV3, Q9NYM8, Q9UHK4, Q9Y3F9, Q9Y3G0 | Gene names | CIZ1, LSFR1, NP94 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cip1-interacting zinc finger protein (Nuclear protein NP94). | |||||
|
CLIC6_HUMAN
|
||||||
| NC score | 0.028652 (rank : 35) | θ value | 0.125558 (rank : 19) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96NY7, Q8IX31 | Gene names | CLIC6, CLIC1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel 6. | |||||
|
STRC_MOUSE
|
||||||
| NC score | 0.028143 (rank : 36) | θ value | 6.88961 (rank : 56) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VIM6 | Gene names | Strc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stereocilin precursor. | |||||
|
NEBU_HUMAN
|
||||||
| NC score | 0.025551 (rank : 37) | θ value | 0.365318 (rank : 20) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P20929, Q15346 | Gene names | NEB | |||
|
Domain Architecture |

|
|||||
| Description | Nebulin. | |||||
|
AKAP2_MOUSE
|
||||||
| NC score | 0.025505 (rank : 38) | θ value | 2.36792 (rank : 32) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O54931, O54932, O54933 | Gene names | Akap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2) (AKAP expressed in kidney and lung) (AKAP-KL). | |||||
|
NAGAB_HUMAN
|
||||||
| NC score | 0.023577 (rank : 39) | θ value | 6.88961 (rank : 54) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P17050 | Gene names | NAGA | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-N-acetylgalactosaminidase precursor (EC 3.2.1.49) (Alpha- galactosidase B). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.019805 (rank : 40) | θ value | 3.0926 (rank : 38) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
TNF10_MOUSE
|
||||||
| NC score | 0.019494 (rank : 41) | θ value | 4.03905 (rank : 42) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P50592 | Gene names | Tnfsf10, Trail | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor ligand superfamily member 10 (TNF-related apoptosis-inducing ligand) (Protein TRAIL) (CD253 antigen). | |||||
|
AKAP2_HUMAN
|
||||||
| NC score | 0.019095 (rank : 42) | θ value | 6.88961 (rank : 49) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y2D5, Q9UG26 | Gene names | AKAP2, KIAA0920 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2). | |||||
|
ZN622_MOUSE
|
||||||
| NC score | 0.018234 (rank : 43) | θ value | 5.27518 (rank : 48) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91VY9 | Gene names | Znf622, D15Ertd806e, Zfp622 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 622. | |||||
|
TIF1B_MOUSE
|
||||||
| NC score | 0.016697 (rank : 44) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q62318, P70391, Q8C283, Q99PN4 | Gene names | Trim28, Krip1, Tif1b | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-beta (TIF1-beta) (Tripartite motif-containing protein 28) (KRAB-A-interacting protein) (KRIP-1). | |||||
|
ASAHL_HUMAN
|
||||||
| NC score | 0.016374 (rank : 45) | θ value | 5.27518 (rank : 43) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q02083, Q5KTF2, Q96EY2, Q9BRA8 | Gene names | ASAHL, NAAA, PLT | |||
|
Domain Architecture |

|
|||||
| Description | N-acylethanolamine-hydrolyzing acid amidase precursor (EC 3.5.1.-) (N- acylsphingosine amidohydrolase-like) (ASAH-like protein) (Acid ceramidase-like protein). | |||||
|
ARMX5_MOUSE
|
||||||
| NC score | 0.016307 (rank : 46) | θ value | 3.0926 (rank : 36) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3UZB0 | Gene names | Armcx5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 5. | |||||
|
GCP5_HUMAN
|
||||||
| NC score | 0.016280 (rank : 47) | θ value | 6.88961 (rank : 53) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96RT8, Q96PY8 | Gene names | TUBGCP5, GCP5, KIAA1899 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-tubulin complex component 5 (GCP-5). | |||||
|
EPIPL_MOUSE
|
||||||
| NC score | 0.014312 (rank : 48) | θ value | 3.0926 (rank : 37) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R0W0 | Gene names | Eppk1 | |||
|
Domain Architecture |

|
|||||
| Description | Epiplakin. | |||||
|
EDD1_MOUSE
|
||||||
| NC score | 0.014261 (rank : 49) | θ value | 5.27518 (rank : 44) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80TP3, Q698K9, Q6PEQ8, Q6PFQ9, Q80VL4, Q810V6, Q9CXE9 | Gene names | Edd1, Edd, Kiaa0896 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog). | |||||
|
YETS2_HUMAN
|
||||||
| NC score | 0.013623 (rank : 50) | θ value | 5.27518 (rank : 47) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ULM3, Q641P6, Q9NW96 | Gene names | YEATS2, KIAA1197 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YEATS domain-containing protein 2. | |||||
|
EDD1_HUMAN
|
||||||
| NC score | 0.013431 (rank : 51) | θ value | 6.88961 (rank : 51) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95071, O94970, Q9NPL3 | Gene names | EDD1, EDD, HYD, KIAA0896 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog) (hHYD) (Progestin-induced protein). | |||||
|
BRPF3_HUMAN
|
||||||
| NC score | 0.013120 (rank : 52) | θ value | 2.36792 (rank : 33) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ULD4, Q5R3K8 | Gene names | BRPF3, KIAA1286 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and PHD finger-containing protein 3. | |||||
|
TIF1A_MOUSE
|
||||||
| NC score | 0.012636 (rank : 53) | θ value | 4.03905 (rank : 41) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q64127, Q64126 | Gene names | Trim24, Tif1, Tif1a | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24). | |||||
|
GSHB_HUMAN
|
||||||
| NC score | 0.012414 (rank : 54) | θ value | 8.99809 (rank : 59) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P48637, Q4TTD9 | Gene names | GSS | |||
|
Domain Architecture |
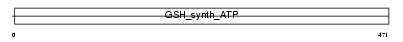
|
|||||
| Description | Glutathione synthetase (EC 6.3.2.3) (Glutathione synthase) (GSH synthetase) (GSH-S). | |||||
|
FBX32_HUMAN
|
||||||
| NC score | 0.011781 (rank : 55) | θ value | 6.88961 (rank : 52) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q969P5 | Gene names | FBXO32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 32 (Muscle atrophy F-box protein) (MAFbx) (Atrogin- 1). | |||||
|
FUT4_HUMAN
|
||||||
| NC score | 0.011280 (rank : 56) | θ value | 5.27518 (rank : 46) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P22083 | Gene names | FUT4, ELFT | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-(1,3)-fucosyltransferase (EC 2.4.1.-) (Galactoside 3-L- fucosyltransferase) (Fucosyltransferase 4) (FUCT-IV) (Fuc-TIV) (ELAM-1 ligand fucosyltransferase). | |||||
|
ITA2_MOUSE
|
||||||
| NC score | 0.010912 (rank : 57) | θ value | 1.38821 (rank : 28) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q62469, Q62163 | Gene names | Itga2 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-2 precursor (Platelet membrane glycoprotein Ia) (GPIa) (Collagen receptor) (VLA-2 alpha chain) (CD49b antigen). | |||||
|
ATX2L_MOUSE
|
||||||
| NC score | 0.010493 (rank : 58) | θ value | 8.99809 (rank : 58) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7TQH0, Q80XN9, Q8K059 | Gene names | Atxn2l, A2lp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2-like protein. | |||||
|
AAKG3_MOUSE
|
||||||
| NC score | 0.008106 (rank : 59) | θ value | 8.99809 (rank : 57) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BGM7, Q80WK8, Q8CJ41 | Gene names | Prkag3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 5'-AMP-activated protein kinase subunit gamma-3 (AMPK gamma-3 chain) (AMPK gamma3). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.007277 (rank : 60) | θ value | 4.03905 (rank : 40) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
DDX51_HUMAN
|
||||||
| NC score | 0.005515 (rank : 61) | θ value | 6.88961 (rank : 50) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
SPTN2_HUMAN
|
||||||
| NC score | 0.003795 (rank : 62) | θ value | 6.88961 (rank : 55) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 535 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15020, O14872, O14873 | Gene names | SPTBN2, KIAA0302 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 2 (Spectrin, non-erythroid beta chain 2) (Beta-III spectrin). | |||||
|
ZN451_MOUSE
|
||||||
| NC score | 0.003312 (rank : 63) | θ value | 8.99809 (rank : 63) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 643 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8C0P7 | Gene names | Znf451, Zfp451 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 451. | |||||
|
ZN142_HUMAN
|
||||||
| NC score | 0.002522 (rank : 64) | θ value | 1.38821 (rank : 31) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P52746, Q92510 | Gene names | ZNF142, KIAA0236 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 142 (HA4654). | |||||
|
SHAN1_HUMAN
|
||||||
| NC score | 0.002282 (rank : 65) | θ value | 8.99809 (rank : 60) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y566, Q9NYW9 | Gene names | SHANK1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 1 (Shank1) (Somatostatin receptor-interacting protein) (SSTR-interacting protein) (SSTRIP). | |||||
|
MAST4_HUMAN
|
||||||
| NC score | 0.002017 (rank : 66) | θ value | 4.03905 (rank : 39) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||
|
FBN1_HUMAN
|
||||||
| NC score | 0.001926 (rank : 67) | θ value | 5.27518 (rank : 45) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P35555, Q15972, Q75N87 | Gene names | FBN1, FBN | |||
|
Domain Architecture |

|
|||||
| Description | Fibrillin-1 precursor. | |||||
|
WD51A_MOUSE
|
||||||
| NC score | 0.000945 (rank : 68) | θ value | 8.99809 (rank : 61) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8JZX3, Q9CY09 | Gene names | Wdr51a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 51A. | |||||
|
ZN473_MOUSE
|
||||||
| NC score | 0.000746 (rank : 69) | θ value | 1.06291 (rank : 25) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 824 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BI67, Q8BI98, Q8BIB7 | Gene names | Znf473, Zfp100, Zfp473 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 473 homolog (Zinc finger protein 100) (Zfp-100). | |||||
|
ZN174_HUMAN
|
||||||
| NC score | -0.000752 (rank : 70) | θ value | 8.99809 (rank : 62) | |||
| Query Neighborhood Hits | 63 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q15697, Q9BQ34 | Gene names | ZNF174, ZSCAN8 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 174 (AW-1) (Zinc finger and SCAN domain-containing protein 8). | |||||