Please be patient as the page loads
|
HS90B_HUMAN
|
||||||
| SwissProt Accessions | P08238, Q5T9W7, Q9NQW0 | Gene names | HSP90AB1, HSP90B, HSPC2, HSPCB | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock protein HSP 90-beta (HSP 84) (HSP 90). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
HS90A_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.977168 (rank : 4) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P07900, Q2PP14, Q9BVQ5 | Gene names | HSP90AA1, HSP90A, HSPC1, HSPCA | |||
|
Domain Architecture |
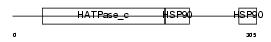
|
|||||
| Description | Heat shock protein HSP 90-alpha (HSP 86) (NY-REN-38 antigen). | |||||
|
HS90A_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.979917 (rank : 3) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P07901 | Gene names | Hsp90aa1, Hsp86, Hsp86-1, Hspca | |||
|
Domain Architecture |
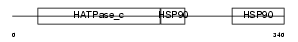
|
|||||
| Description | Heat shock protein HSP 90-alpha (HSP 86) (Tumor-specific transplantation 86 kDa antigen) (TSTA). | |||||
|
HS90B_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 98 | |
| SwissProt Accessions | P08238, Q5T9W7, Q9NQW0 | Gene names | HSP90AB1, HSP90B, HSPC2, HSPCB | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock protein HSP 90-beta (HSP 84) (HSP 90). | |||||
|
HS90B_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.998135 (rank : 2) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P11499, O89078 | Gene names | Hsp90ab1, Hsp84, Hsp84-1, Hspcb | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock protein HSP 90-beta (HSP 84) (Tumor-specific transplantation 84 kDa antigen) (TSTA). | |||||
|
ENPL_HUMAN
|
||||||
| θ value | 2.6227e-108 (rank : 5) | NC score | 0.933458 (rank : 5) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P14625, Q96A97 | Gene names | HSP90B1, TRA1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmin precursor (Heat shock protein 90 kDa beta member 1) (94 kDa glucose-regulated protein) (GRP94) (gp96 homolog) (Tumor rejection antigen 1). | |||||
|
ENPL_MOUSE
|
||||||
| θ value | 2.89973e-107 (rank : 6) | NC score | 0.931922 (rank : 6) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P08113, P11427 | Gene names | Hsp90b1, Tra-1, Tra1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmin precursor (Heat shock protein 90 kDa beta member 1) (94 kDa glucose-regulated protein) (GRP94) (ERP99) (Polymorphic tumor rejection antigen 1) (Tumor rejection antigen gp96). | |||||
|
TRAP1_HUMAN
|
||||||
| θ value | 5.55816e-50 (rank : 7) | NC score | 0.856840 (rank : 7) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q12931, O43642, O75235, Q9UHL5 | Gene names | TRAP1, HSP75 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock protein 75 kDa, mitochondrial precursor (HSP 75) (Tumor necrosis factor type 1 receptor-associated protein) (TRAP-1) (TNFR- associated protein 1). | |||||
|
TRAP1_MOUSE
|
||||||
| θ value | 2.1121e-49 (rank : 8) | NC score | 0.855764 (rank : 8) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9CQN1, Q542I4 | Gene names | Trap1 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock protein 75 kDa, mitochondrial precursor (HSP 75) (Tumor necrosis factor type 1 receptor-associated protein) (TRAP-1) (TNFR- associated protein 1). | |||||
|
SEC63_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 9) | NC score | 0.108499 (rank : 10) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
TRIPB_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 10) | NC score | 0.038416 (rank : 17) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1587 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q15643, O14689, O15154, O95949 | Gene names | TRIP11, CEV14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid receptor-interacting protein 11 (TRIP-11) (Golgi-associated microtubule-binding protein 210) (GMAP-210) (Trip230) (Clonal evolution-related gene on chromosome 14). | |||||
|
KI67_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 11) | NC score | 0.060042 (rank : 14) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
SEC63_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 12) | NC score | 0.101651 (rank : 11) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
CND3_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 13) | NC score | 0.080253 (rank : 13) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BPX3, Q96SV9, Q9BUR3, Q9BVY1, Q9H914, Q9H9Z6, Q9HBI9 | Gene names | NCAPG, CAPG, NYMEL3 | |||
|
Domain Architecture |

|
|||||
| Description | Condensin complex subunit 3 (Non-SMC condensin I complex subunit G) (Chromosome-associated protein G) (Condensin subunit CAP-G) (hCAP-G) (XCAP-G homolog) (Antigen NY-MEL-3). | |||||
|
NUF2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 14) | NC score | 0.036815 (rank : 20) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 661 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9BZD4, Q8WU69, Q96HJ4, Q96Q78 | Gene names | CDCA1, NUF2, NUF2R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinetochore protein Nuf2 (hNuf2) (hNuf2R) (hsNuf2) (Cell division cycle-associated protein 1). | |||||
|
PRI1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 15) | NC score | 0.091735 (rank : 12) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P20664 | Gene names | Prim1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA primase small subunit (EC 2.7.7.-) (DNA primase 49 kDa subunit) (p49). | |||||
|
SMC2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 16) | NC score | 0.035334 (rank : 24) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O95347, Q9P1P2 | Gene names | SMC2L1, CAPE, SMC2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 2-like 1 protein (Chromosome- associated protein E) (hCAP-E) (XCAP-E homolog). | |||||
|
HSF2B_HUMAN
|
||||||
| θ value | 0.279714 (rank : 17) | NC score | 0.144641 (rank : 9) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O75031 | Gene names | HSF2BP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heat shock factor 2-binding protein. | |||||
|
LAMA2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 18) | NC score | 0.016235 (rank : 67) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1198 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q60675, Q05003, Q64061 | Gene names | Lama2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
OXSR1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 19) | NC score | 0.012383 (rank : 80) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 865 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6P9R2, Q8BVZ9, Q8C0B9 | Gene names | Oxsr1, Osr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase OSR1 (EC 2.7.11.1) (Oxidative stress- responsive 1 protein). | |||||
|
SYNE1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 20) | NC score | 0.028237 (rank : 32) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
AT10A_HUMAN
|
||||||
| θ value | 0.813845 (rank : 21) | NC score | 0.022481 (rank : 51) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60312, Q969I4 | Gene names | ATP10A, ATP10C, ATPVC, KIAA0566 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (ATPVA) (Aminophospholipid translocase VA). | |||||
|
RELN_HUMAN
|
||||||
| θ value | 0.813845 (rank : 22) | NC score | 0.036019 (rank : 22) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P78509, Q86UJ0, Q86UJ8, Q8NDV0, Q9UDQ2 | Gene names | RELN | |||
|
Domain Architecture |
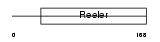
|
|||||
| Description | Reelin precursor (EC 3.4.21.-). | |||||
|
EVI1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 23) | NC score | 0.002959 (rank : 96) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 924 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q03112, Q16122, Q99917 | Gene names | EVI1 | |||
|
Domain Architecture |

|
|||||
| Description | Ecotropic virus integration 1 site protein. | |||||
|
OXSR1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 24) | NC score | 0.011040 (rank : 82) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O95747, Q3LR53, Q7Z501, Q9UPQ1 | Gene names | OXSR1, KIAA1101, OSR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase OSR1 (EC 2.7.11.1) (Oxidative stress- responsive 1 protein). | |||||
|
BAZ1A_HUMAN
|
||||||
| θ value | 1.38821 (rank : 25) | NC score | 0.023659 (rank : 47) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9NRL2, Q9NZ15, Q9P065, Q9UIG1, Q9Y3V3 | Gene names | BAZ1A, ACF1, WCRF180 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1A (ATP-utilizing chromatin assembly and remodeling factor 1) (hACF1) (ATP-dependent chromatin remodelling protein) (Williams syndrome transcription factor-related chromatin remodeling factor 180) (WCRF180) (hWALp1) (CHRAC subunit ACF1). | |||||
|
CCD46_MOUSE
|
||||||
| θ value | 1.38821 (rank : 26) | NC score | 0.023349 (rank : 48) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q5PR68, Q80ZN6, Q99JS4, Q9D9T1 | Gene names | Ccdc46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled coil domain-containing protein 46. | |||||
|
KIF5A_MOUSE
|
||||||
| θ value | 1.38821 (rank : 27) | NC score | 0.017530 (rank : 63) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P33175, Q5DTP1, Q6PDY7, Q9Z2F9 | Gene names | Kif5a, Kiaa4086, Kif5, Nkhc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin heavy chain isoform 5A (Neuronal kinesin heavy chain) (NKHC) (Kinesin heavy chain neuron-specific 1). | |||||
|
DYH11_HUMAN
|
||||||
| θ value | 1.81305 (rank : 28) | NC score | 0.023981 (rank : 45) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q96DT5, Q9UJ82 | Gene names | DNAH11 | |||
|
Domain Architecture |

|
|||||
| Description | Ciliary dynein heavy chain 11 (Axonemal beta dynein heavy chain 11). | |||||
|
BPAEA_HUMAN
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.022284 (rank : 52) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1369 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O94833, Q5TBT0, Q8N1T8, Q8N8J3, Q8WXK9, Q96AK9, Q96DQ5, Q9H555 | Gene names | DST, BPAG1, DMH, DT, KIAA0728 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 6/9/10 (Trabeculin-beta) (Bullous pemphigoid antigen) (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
CEP55_MOUSE
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.025713 (rank : 38) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8BT07, Q8C2J0, Q8K2I8, Q8R2Y4, Q9DBZ8 | Gene names | Cep55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome protein 55. | |||||
|
CLC6A_HUMAN
|
||||||
| θ value | 2.36792 (rank : 31) | NC score | 0.013102 (rank : 77) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6EIG7 | Gene names | CLEC6A, CLECSF10, DECTIN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-type lectin domain family 6 member A (C-type lectin superfamily member 10) (Dendritic cell-associated lectin 2) (DC-associated C-type lectin-2) (DECTIN-2). | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 32) | NC score | 0.045057 (rank : 16) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
TCAL5_HUMAN
|
||||||
| θ value | 2.36792 (rank : 33) | NC score | 0.033505 (rank : 28) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
ANLN_HUMAN
|
||||||
| θ value | 3.0926 (rank : 34) | NC score | 0.034749 (rank : 27) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NQW6, Q5CZ78, Q6NSK5, Q9H8Y4, Q9NVN9, Q9NVP0 | Gene names | ANLN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
BPA1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 35) | NC score | 0.018562 (rank : 60) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
CCD46_HUMAN
|
||||||
| θ value | 3.0926 (rank : 36) | NC score | 0.021550 (rank : 53) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1430 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8N8E3, Q6PIB5, Q8NCR4, Q8NFR4 | Gene names | CCDC46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 46. | |||||
|
CK5P2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 37) | NC score | 0.016948 (rank : 65) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1433 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96SN8, Q7Z3L4, Q7Z3U1, Q7Z7I6, Q9BSW0, Q9H6J6, Q9HCD9, Q9NV90, Q9UIW9 | Gene names | CDK5RAP2, CEP215, KIAA1633 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48) (Centrosome-associated protein 215). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 38) | NC score | 0.017189 (rank : 64) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
LEO1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 39) | NC score | 0.048149 (rank : 15) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
NAF1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 40) | NC score | 0.024844 (rank : 43) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q15025, O76008, Q96EL9, Q99833, Q9H1J3 | Gene names | TNIP1, KIAA0113, NAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Nef-associated factor 1 (Naf1) (TNFAIP3-interacting protein 1) (HIV-1 Nef-interacting protein) (Virion-associated nuclear shuttling protein) (VAN) (hVAN) (Nip40-1). | |||||
|
RGS6_HUMAN
|
||||||
| θ value | 3.0926 (rank : 41) | NC score | 0.012383 (rank : 79) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P49758, O75576, O75577, Q8TE13, Q8TE14, Q8TE15, Q8TE16, Q8TE17, Q8TE18, Q8TE19, Q8TE20, Q8TE21, Q8TE22, Q9Y245, Q9Y647 | Gene names | RGS6 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 6 (RGS6) (S914). | |||||
|
SMC2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 42) | NC score | 0.035887 (rank : 23) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1131 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8CG48, Q61076, Q9CS17, Q9CSD8 | Gene names | Smc2l1, Cape, Fin16, Smc2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 2-like 1 protein (Chromosome- associated protein E) (XCAP-E homolog) (FGF-inducible protein 16). | |||||
|
SMRC2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 43) | NC score | 0.028817 (rank : 31) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8TAQ2, Q92923, Q96E12, Q96GY4 | Gene names | SMARCC2, BAF170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
TXND2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 44) | NC score | 0.034898 (rank : 26) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
ZA2G_MOUSE
|
||||||
| θ value | 3.0926 (rank : 45) | NC score | 0.007738 (rank : 84) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q64726 | Gene names | Azgp1 | |||
|
Domain Architecture |
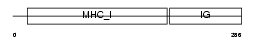
|
|||||
| Description | Zinc-alpha-2-glycoprotein precursor (Zn-alpha-2-glycoprotein) (Zn- alpha-2-GP). | |||||
|
ZBT24_HUMAN
|
||||||
| θ value | 3.0926 (rank : 46) | NC score | -0.000210 (rank : 98) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43167, Q5TED5, Q8N455 | Gene names | ZBTB24, KIAA0441, ZNF450 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and BTB domain-containing protein 24 (Zinc finger protein 450). | |||||
|
CJ092_HUMAN
|
||||||
| θ value | 4.03905 (rank : 47) | NC score | 0.031205 (rank : 29) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IYW2, Q5JSF7, Q9NTQ5 | Gene names | C10orf92 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf92. | |||||
|
NOC2L_HUMAN
|
||||||
| θ value | 4.03905 (rank : 48) | NC score | 0.025225 (rank : 41) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y3T9, Q5SVA3, Q9BTN6 | Gene names | NOC2L | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar complex protein 2 homolog (Protein NOC2 homolog) (NOC2- like). | |||||
|
NSBP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 49) | NC score | 0.037316 (rank : 19) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
ST1E1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 50) | NC score | 0.022705 (rank : 50) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P49888, Q8N6X5 | Gene names | SULT1E1, STE | |||
|
Domain Architecture |

|
|||||
| Description | Estrogen sulfotransferase (EC 2.8.2.4) (Sulfotransferase, estrogen- preferring) (EST-1). | |||||
|
STK39_HUMAN
|
||||||
| θ value | 4.03905 (rank : 51) | NC score | 0.007045 (rank : 89) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 882 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UEW8, O14774, Q9UER4 | Gene names | STK39, SPAK | |||
|
Domain Architecture |
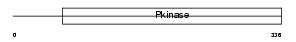
|
|||||
| Description | STE20/SPS1-related proline-alanine-rich protein kinase (EC 2.7.11.1) (Ste-20-related kinase) (Serine/threonine-protein kinase 39) (DCHT). | |||||
|
STK39_MOUSE
|
||||||
| θ value | 4.03905 (rank : 52) | NC score | 0.007667 (rank : 85) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 915 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Z1W9, Q80W13 | Gene names | Stk39, Spak | |||
|
Domain Architecture |
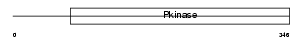
|
|||||
| Description | STE20/SPS1-related proline-alanine-rich protein kinase (EC 2.7.11.1) (Ste-20-related kinase) (Serine/threonine-protein kinase 39). | |||||
|
ADA30_HUMAN
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.014589 (rank : 72) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UKF2, Q9UKF1 | Gene names | ADAM30 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 30 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 30). | |||||
|
AT10A_MOUSE
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | 0.013063 (rank : 78) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O54827, Q8R3B8 | Gene names | Atp10a, Atpc5, Pfatp | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (P-locus fat-associated ATPase). | |||||
|
MAP1A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 55) | NC score | 0.027969 (rank : 35) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
MPIP3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 56) | NC score | 0.029898 (rank : 30) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P30307, Q96PL3, Q9H168, Q9H2E8, Q9H2E9, Q9H2F1 | Gene names | CDC25C | |||
|
Domain Architecture |

|
|||||
| Description | M-phase inducer phosphatase 3 (EC 3.1.3.48) (Dual specificity phosphatase Cdc25C). | |||||
|
MRIP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 57) | NC score | 0.026513 (rank : 37) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 578 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6WCQ1, Q3KQZ5, Q5FB94, Q6DHW2, Q7Z5Y2, Q8N390, Q96G40 | Gene names | MRIP, KIAA0864, RHOIP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin phosphatase Rho-interacting protein (Rho-interacting protein 3) (M-RIP) (RIP3) (p116Rip). | |||||
|
MRIP_MOUSE
|
||||||
| θ value | 5.27518 (rank : 58) | NC score | 0.026903 (rank : 36) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 569 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P97434, Q3TBR5, Q3TC85, Q3TRS0, Q3U1Y8, Q3U3E4, Q5SWZ3, Q5SWZ4, Q5SWZ7, Q6P559, Q80YC8 | Gene names | Mrip, Kiaa0864, Rhoip3 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin phosphatase Rho-interacting protein (Rho-interacting protein 3) (p116Rip) (RIP3). | |||||
|
PHF3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 59) | NC score | 0.035032 (rank : 25) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q92576, Q5T1T6, Q9NQ16, Q9UI45 | Gene names | PHF3, KIAA0244 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 3. | |||||
|
PITX2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.010583 (rank : 83) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99697, O60578, O60579, O60580, Q9BY17 | Gene names | PITX2, ARP1, RGS, RIEG, RIEG1 | |||
|
Domain Architecture |
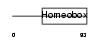
|
|||||
| Description | Pituitary homeobox 2 (RIEG bicoid-related homeobox transcription factor) (Solurshin) (ALL1-responsive protein ARP1). | |||||
|
PMS2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.025166 (rank : 42) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P54278 | Gene names | PMS2, PMSL2 | |||
|
Domain Architecture |
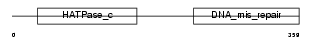
|
|||||
| Description | PMS1 protein homolog 2 (DNA mismatch repair protein PMS2). | |||||
|
S12A6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.013577 (rank : 75) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UHW9, Q8TDD4, Q9UFR2, Q9Y642, Q9Y665 | Gene names | SLC12A6, KCC3 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 12 member 6 (Electroneutral potassium-chloride cotransporter 3) (K-Cl cotransporter 3). | |||||
|
S12A6_MOUSE
|
||||||
| θ value | 5.27518 (rank : 63) | NC score | 0.013552 (rank : 76) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q924N4, Q924N3 | Gene names | Slc12a6, Kcc3 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 12 member 6 (Electroneutral potassium-chloride cotransporter 3) (K-Cl cotransporter 3). | |||||
|
TARA_MOUSE
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.025319 (rank : 40) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
TGON1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.023082 (rank : 49) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q62313 | Gene names | Tgoln1, Ttgn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 1 precursor (TGN38A). | |||||
|
TRI36_HUMAN
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.004983 (rank : 91) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NQ86 | Gene names | TRIM36, RBCC728, RNF98 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 36 (Zinc-binding protein Rbcc728) (RING finger protein 98). | |||||
|
ZN318_MOUSE
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.037682 (rank : 18) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 642 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q99PP2, Q3TZL5, Q9JJ01 | Gene names | Znf318, Tzf, Zfp318 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 318 (Testicular zinc finger protein). | |||||
|
AFF4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.013666 (rank : 74) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
ANLN_MOUSE
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.036384 (rank : 21) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8K298, Q8BL79, Q8BLB3, Q8K2N0, Q9CUY0 | Gene names | Anln | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
ANR11_HUMAN
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | 0.018412 (rank : 61) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
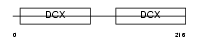
DCDC2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.019915 (rank : 57) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UHG0, Q5VTR8, Q5VTR9, Q86W35, Q9UFD1, Q9ULR6 | Gene names | DCDC2, KIAA1154 | |||
|
Domain Architecture |
|
|||||
| Description | Doublecortin domain-containing protein 2 (RU2S protein). | |||||
|
DESP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.018338 (rank : 62) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P15924, O75993, Q14189, Q9UHN4 | Gene names | DSP | |||
|
Domain Architecture |

|
|||||
| Description | Desmoplakin (DP) (250/210 kDa paraneoplastic pemphigus antigen). | |||||
|
DYH9_HUMAN
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | 0.023809 (rank : 46) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9NYC9, O95494, Q9NQ28 | Gene names | DNAH9, DNAH17L, DNAL1 | |||
|
Domain Architecture |

|
|||||
| Description | Ciliary dynein heavy chain 9 (Axonemal beta dynein heavy chain 9). | |||||
|
IF4G3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.015343 (rank : 70) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O43432, Q15597, Q8NEN1 | Gene names | EIF4G3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 3 (eIF-4-gamma 3) (eIF-4G 3) (eIF4G 3) (eIF-4-gamma II) (eIF4GII). | |||||
|
IF4G3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.015480 (rank : 69) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80XI3, Q80Y69, Q8BJ68, Q8BQF3, Q8C6Q3, Q8CIH0 | Gene names | Eif4g3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 3 (eIF-4-gamma 3) (eIF-4G 3) (eIF4G 3) (eIF-4-gamma II) (eIF4GII). | |||||
|
KIF5C_HUMAN
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.014426 (rank : 73) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 974 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O60282, O95079 | Gene names | KIF5C, KIAA0531, NKHC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin heavy chain isoform 5C (Kinesin heavy chain neuron-specific 2). | |||||
|
LMO7_HUMAN
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.019868 (rank : 58) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8WWI1, O15462, O95346, Q9UKC1, Q9UQM5, Q9Y6A7 | Gene names | LMO7, FBX20, FBXO20, KIAA0858 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain only protein 7 (LOMP) (F-box only protein 20). | |||||
|
MPP8_HUMAN
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.024690 (rank : 44) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q99549, Q86TK3, Q9BTP1 | Gene names | MPHOSPH8, MPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | M-phase phosphoprotein 8. | |||||
|
NKX61_MOUSE
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.004050 (rank : 92) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99MA9, Q9ERQ7 | Gene names | Nkx6-1, Nkx6.1, Nkx6a | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Nkx-6.1. | |||||
|
P52K_HUMAN
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.021524 (rank : 54) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43422, Q8WTW1, Q9Y3Z4 | Gene names | PRKRIR, DAP4, P52RIPK, THAP0 | |||
|
Domain Architecture |

|
|||||
| Description | 52 kDa repressor of the inhibitor of the protein kinase (p58IPK- interacting protein) (58 kDa interferon-induced protein kinase- interacting protein) (P52rIPK) (Death-associated protein 4) (THAP domain-containing protein 0). | |||||
|
SALL1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.001178 (rank : 97) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 826 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9ER74, Q920R5 | Gene names | Sall1, Sal3 | |||
|
Domain Architecture |

|
|||||
| Description | Sal-like protein 1 (Zinc finger protein Spalt-3) (Sal-3) (MSal-3). | |||||
|
SC10A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.003020 (rank : 95) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5Y9 | Gene names | SCN10A, PN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 10 subunit alpha (Sodium channel protein type X subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.8) (Peripheral nerve sodium channel 3) (hPN3). | |||||
|
SMRC2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.027975 (rank : 34) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6PDG5, Q6P6P3 | Gene names | Smarcc2, Baf170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
SYCP1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.028161 (rank : 33) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1456 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q62209, O09205, P70192, Q62329 | Gene names | Sycp1, Scp1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
TEX14_MOUSE
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.007599 (rank : 87) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1333 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q7M6U3, Q3UT36, Q5NC10, Q5NC11, Q8CGK1, Q8CGK2, Q99MV8 | Gene names | Tex14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed protein 14 (Testis-expressed sequence 14). | |||||
|
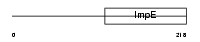
TTC1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.015123 (rank : 71) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q99614, Q9BVT3 | Gene names | TTC1, TPR1 | |||
|
Domain Architecture |
|
|||||
| Description | Tetratricopeptide repeat protein 1 (TPR repeat protein 1). | |||||
|
ABCG2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.003495 (rank : 94) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7TMS5, Q9R004, Q9Z1T0 | Gene names | Abcg2, Abcp, Bcrp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-binding cassette sub-family G member 2 (Breast cancer resistance protein 1 homolog). | |||||
|
ANR26_MOUSE
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.011855 (rank : 81) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1554 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q811D2, Q69ZS2, Q9CS61 | Gene names | Ankrd26, Kiaa1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
CP135_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.016702 (rank : 66) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1290 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q66GS9, O75130, Q9H8H7 | Gene names | CEP135, CEP4, KIAA0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
DMP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.019997 (rank : 56) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
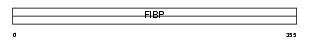
FIBP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.020119 (rank : 55) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JI19 | Gene names | Fibp | |||
|
Domain Architecture |
|
|||||
| Description | Acidic fibroblast growth factor intracellular-binding protein (aFGF intracellular-binding protein) (FGF-1 intracellular-binding protein). | |||||
|
NKX61_HUMAN
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.003674 (rank : 93) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P78426 | Gene names | NKX6-1, NKX6A | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Nkx-6.1. | |||||
|
PEG3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 93) | NC score | 0.007640 (rank : 86) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
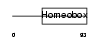
PITX2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.007368 (rank : 88) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P97474, O08646, O70336, P97933, Q9JLA0, Q9QXB8, Q9R1V9, Q9Z141 | Gene names | Pitx2, Arp1, Brx1, Otlx2, Ptx2, Rgs | |||
|
Domain Architecture |
|
|||||
| Description | Pituitary homeobox 2 (Orthodenticle-like homeobox 2) (Solurshin) (ALL1-responsive protein ARP1) (BRX1 homeoprotein) (Paired-like homeodomain transcription factor Munc 30). | |||||
|
RELN_MOUSE
|
||||||
| θ value | 8.99809 (rank : 95) | NC score | 0.025495 (rank : 39) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q60841, Q9CUA6 | Gene names | Reln, Rl | |||
|
Domain Architecture |
|
|||||
| Description | Reelin precursor (EC 3.4.21.-) (Reeler protein). | |||||
|
RXINP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | 0.019831 (rank : 59) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96RL1, Q5XKQ1, Q7Z3W7, Q8N5B9, Q9BZR1, Q9BZR5, Q9UHX7 | Gene names | RXRIP110, RAP80 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoid X receptor-interacting protein 110 (Receptor-associated protein 80) (Nuclear zinc finger protein RAP80). | |||||
|
SHOX_HUMAN
|
||||||
| θ value | 8.99809 (rank : 97) | NC score | 0.006950 (rank : 90) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O15266, O00412, O00413, O15267 | Gene names | SHOX, PHOG | |||
|
Domain Architecture |

|
|||||
| Description | Short stature homeobox protein (Short stature homeobox-containing protein) (Pseudoautosomal homeobox-containing osteogenic protein). | |||||
|
TGON2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 98) | NC score | 0.015611 (rank : 68) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q62314 | Gene names | Tgoln2, Ttgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (TGN38B). | |||||
|
HS90B_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 98 | |
| SwissProt Accessions | P08238, Q5T9W7, Q9NQW0 | Gene names | HSP90AB1, HSP90B, HSPC2, HSPCB | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock protein HSP 90-beta (HSP 84) (HSP 90). | |||||
|
HS90B_MOUSE
|
||||||
| NC score | 0.998135 (rank : 2) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P11499, O89078 | Gene names | Hsp90ab1, Hsp84, Hsp84-1, Hspcb | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock protein HSP 90-beta (HSP 84) (Tumor-specific transplantation 84 kDa antigen) (TSTA). | |||||
|
HS90A_MOUSE
|
||||||
| NC score | 0.979917 (rank : 3) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P07901 | Gene names | Hsp90aa1, Hsp86, Hsp86-1, Hspca | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock protein HSP 90-alpha (HSP 86) (Tumor-specific transplantation 86 kDa antigen) (TSTA). | |||||
|
HS90A_HUMAN
|
||||||
| NC score | 0.977168 (rank : 4) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P07900, Q2PP14, Q9BVQ5 | Gene names | HSP90AA1, HSP90A, HSPC1, HSPCA | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock protein HSP 90-alpha (HSP 86) (NY-REN-38 antigen). | |||||
|
ENPL_HUMAN
|
||||||
| NC score | 0.933458 (rank : 5) | θ value | 2.6227e-108 (rank : 5) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P14625, Q96A97 | Gene names | HSP90B1, TRA1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmin precursor (Heat shock protein 90 kDa beta member 1) (94 kDa glucose-regulated protein) (GRP94) (gp96 homolog) (Tumor rejection antigen 1). | |||||
|
ENPL_MOUSE
|
||||||
| NC score | 0.931922 (rank : 6) | θ value | 2.89973e-107 (rank : 6) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P08113, P11427 | Gene names | Hsp90b1, Tra-1, Tra1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmin precursor (Heat shock protein 90 kDa beta member 1) (94 kDa glucose-regulated protein) (GRP94) (ERP99) (Polymorphic tumor rejection antigen 1) (Tumor rejection antigen gp96). | |||||
|
TRAP1_HUMAN
|
||||||
| NC score | 0.856840 (rank : 7) | θ value | 5.55816e-50 (rank : 7) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q12931, O43642, O75235, Q9UHL5 | Gene names | TRAP1, HSP75 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock protein 75 kDa, mitochondrial precursor (HSP 75) (Tumor necrosis factor type 1 receptor-associated protein) (TRAP-1) (TNFR- associated protein 1). | |||||
|
TRAP1_MOUSE
|
||||||
| NC score | 0.855764 (rank : 8) | θ value | 2.1121e-49 (rank : 8) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9CQN1, Q542I4 | Gene names | Trap1 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock protein 75 kDa, mitochondrial precursor (HSP 75) (Tumor necrosis factor type 1 receptor-associated protein) (TRAP-1) (TNFR- associated protein 1). | |||||
|
HSF2B_HUMAN
|
||||||
| NC score | 0.144641 (rank : 9) | θ value | 0.279714 (rank : 17) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O75031 | Gene names | HSF2BP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heat shock factor 2-binding protein. | |||||
|
SEC63_HUMAN
|
||||||
| NC score | 0.108499 (rank : 10) | θ value | 0.0148317 (rank : 9) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
SEC63_MOUSE
|
||||||
| NC score | 0.101651 (rank : 11) | θ value | 0.0330416 (rank : 12) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
PRI1_MOUSE
|
||||||
| NC score | 0.091735 (rank : 12) | θ value | 0.21417 (rank : 15) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P20664 | Gene names | Prim1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA primase small subunit (EC 2.7.7.-) (DNA primase 49 kDa subunit) (p49). | |||||
|
CND3_HUMAN
|
||||||
| NC score | 0.080253 (rank : 13) | θ value | 0.0961366 (rank : 13) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BPX3, Q96SV9, Q9BUR3, Q9BVY1, Q9H914, Q9H9Z6, Q9HBI9 | Gene names | NCAPG, CAPG, NYMEL3 | |||
|
Domain Architecture |

|
|||||
| Description | Condensin complex subunit 3 (Non-SMC condensin I complex subunit G) (Chromosome-associated protein G) (Condensin subunit CAP-G) (hCAP-G) (XCAP-G homolog) (Antigen NY-MEL-3). | |||||
|
KI67_HUMAN
|
||||||
| NC score | 0.060042 (rank : 14) | θ value | 0.0330416 (rank : 11) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.048149 (rank : 15) | θ value | 3.0926 (rank : 39) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.045057 (rank : 16) | θ value | 2.36792 (rank : 32) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
TRIPB_HUMAN
|
||||||
| NC score | 0.038416 (rank : 17) | θ value | 0.0148317 (rank : 10) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1587 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q15643, O14689, O15154, O95949 | Gene names | TRIP11, CEV14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid receptor-interacting protein 11 (TRIP-11) (Golgi-associated microtubule-binding protein 210) (GMAP-210) (Trip230) (Clonal evolution-related gene on chromosome 14). | |||||
|
ZN318_MOUSE
|
||||||
| NC score | 0.037682 (rank : 18) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 642 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q99PP2, Q3TZL5, Q9JJ01 | Gene names | Znf318, Tzf, Zfp318 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 318 (Testicular zinc finger protein). | |||||
|
NSBP1_MOUSE
|
||||||
| NC score | 0.037316 (rank : 19) | θ value | 4.03905 (rank : 49) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
NUF2_HUMAN
|
||||||
| NC score | 0.036815 (rank : 20) | θ value | 0.163984 (rank : 14) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 661 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9BZD4, Q8WU69, Q96HJ4, Q96Q78 | Gene names | CDCA1, NUF2, NUF2R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinetochore protein Nuf2 (hNuf2) (hNuf2R) (hsNuf2) (Cell division cycle-associated protein 1). | |||||
|
ANLN_MOUSE
|
||||||
| NC score | 0.036384 (rank : 21) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8K298, Q8BL79, Q8BLB3, Q8K2N0, Q9CUY0 | Gene names | Anln | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
RELN_HUMAN
|
||||||
| NC score | 0.036019 (rank : 22) | θ value | 0.813845 (rank : 22) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P78509, Q86UJ0, Q86UJ8, Q8NDV0, Q9UDQ2 | Gene names | RELN | |||
|
Domain Architecture |

|
|||||
| Description | Reelin precursor (EC 3.4.21.-). | |||||
|
SMC2_MOUSE
|
||||||
| NC score | 0.035887 (rank : 23) | θ value | 3.0926 (rank : 42) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1131 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8CG48, Q61076, Q9CS17, Q9CSD8 | Gene names | Smc2l1, Cape, Fin16, Smc2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 2-like 1 protein (Chromosome- associated protein E) (XCAP-E homolog) (FGF-inducible protein 16). | |||||
|
SMC2_HUMAN
|
||||||
| NC score | 0.035334 (rank : 24) | θ value | 0.21417 (rank : 16) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O95347, Q9P1P2 | Gene names | SMC2L1, CAPE, SMC2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 2-like 1 protein (Chromosome- associated protein E) (hCAP-E) (XCAP-E homolog). | |||||
|
PHF3_HUMAN
|
||||||
| NC score | 0.035032 (rank : 25) | θ value | 5.27518 (rank : 59) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q92576, Q5T1T6, Q9NQ16, Q9UI45 | Gene names | PHF3, KIAA0244 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 3. | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.034898 (rank : 26) | θ value | 3.0926 (rank : 44) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
ANLN_HUMAN
|
||||||
| NC score | 0.034749 (rank : 27) | θ value | 3.0926 (rank : 34) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NQW6, Q5CZ78, Q6NSK5, Q9H8Y4, Q9NVN9, Q9NVP0 | Gene names | ANLN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
TCAL5_HUMAN
|
||||||
| NC score | 0.033505 (rank : 28) | θ value | 2.36792 (rank : 33) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
CJ092_HUMAN
|
||||||
| NC score | 0.031205 (rank : 29) | θ value | 4.03905 (rank : 47) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IYW2, Q5JSF7, Q9NTQ5 | Gene names | C10orf92 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf92. | |||||
|
MPIP3_HUMAN
|
||||||
| NC score | 0.029898 (rank : 30) | θ value | 5.27518 (rank : 56) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P30307, Q96PL3, Q9H168, Q9H2E8, Q9H2E9, Q9H2F1 | Gene names | CDC25C | |||
|
Domain Architecture |

|
|||||
| Description | M-phase inducer phosphatase 3 (EC 3.1.3.48) (Dual specificity phosphatase Cdc25C). | |||||
|
SMRC2_HUMAN
|
||||||
| NC score | 0.028817 (rank : 31) | θ value | 3.0926 (rank : 43) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8TAQ2, Q92923, Q96E12, Q96GY4 | Gene names | SMARCC2, BAF170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
SYNE1_HUMAN
|
||||||
| NC score | 0.028237 (rank : 32) | θ value | 0.365318 (rank : 20) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
SYCP1_MOUSE
|
||||||
| NC score | 0.028161 (rank : 33) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1456 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q62209, O09205, P70192, Q62329 | Gene names | Sycp1, Scp1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
SMRC2_MOUSE
|
||||||
| NC score | 0.027975 (rank : 34) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6PDG5, Q6P6P3 | Gene names | Smarcc2, Baf170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
MAP1A_HUMAN
|
||||||
| NC score | 0.027969 (rank : 35) | θ value | 5.27518 (rank : 55) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
MRIP_MOUSE
|
||||||
| NC score | 0.026903 (rank : 36) | θ value | 5.27518 (rank : 58) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 569 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P97434, Q3TBR5, Q3TC85, Q3TRS0, Q3U1Y8, Q3U3E4, Q5SWZ3, Q5SWZ4, Q5SWZ7, Q6P559, Q80YC8 | Gene names | Mrip, Kiaa0864, Rhoip3 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin phosphatase Rho-interacting protein (Rho-interacting protein 3) (p116Rip) (RIP3). | |||||
|
MRIP_HUMAN
|
||||||
| NC score | 0.026513 (rank : 37) | θ value | 5.27518 (rank : 57) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 578 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6WCQ1, Q3KQZ5, Q5FB94, Q6DHW2, Q7Z5Y2, Q8N390, Q96G40 | Gene names | MRIP, KIAA0864, RHOIP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin phosphatase Rho-interacting protein (Rho-interacting protein 3) (M-RIP) (RIP3) (p116Rip). | |||||
|
CEP55_MOUSE
|
||||||
| NC score | 0.025713 (rank : 38) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8BT07, Q8C2J0, Q8K2I8, Q8R2Y4, Q9DBZ8 | Gene names | Cep55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome protein 55. | |||||
|
RELN_MOUSE
|
||||||
| NC score | 0.025495 (rank : 39) | θ value | 8.99809 (rank : 95) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q60841, Q9CUA6 | Gene names | Reln, Rl | |||
|
Domain Architecture |

|
|||||
| Description | Reelin precursor (EC 3.4.21.-) (Reeler protein). | |||||
|
TARA_MOUSE
|
||||||
| NC score | 0.025319 (rank : 40) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
NOC2L_HUMAN
|
||||||
| NC score | 0.025225 (rank : 41) | θ value | 4.03905 (rank : 48) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y3T9, Q5SVA3, Q9BTN6 | Gene names | NOC2L | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar complex protein 2 homolog (Protein NOC2 homolog) (NOC2- like). | |||||
|
PMS2_HUMAN
|
||||||
| NC score | 0.025166 (rank : 42) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P54278 | Gene names | PMS2, PMSL2 | |||
|
Domain Architecture |

|
|||||
| Description | PMS1 protein homolog 2 (DNA mismatch repair protein PMS2). | |||||
|
NAF1_HUMAN
|
||||||
| NC score | 0.024844 (rank : 43) | θ value | 3.0926 (rank : 40) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q15025, O76008, Q96EL9, Q99833, Q9H1J3 | Gene names | TNIP1, KIAA0113, NAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Nef-associated factor 1 (Naf1) (TNFAIP3-interacting protein 1) (HIV-1 Nef-interacting protein) (Virion-associated nuclear shuttling protein) (VAN) (hVAN) (Nip40-1). | |||||
|
MPP8_HUMAN
|
||||||
| NC score | 0.024690 (rank : 44) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q99549, Q86TK3, Q9BTP1 | Gene names | MPHOSPH8, MPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | M-phase phosphoprotein 8. | |||||
|
DYH11_HUMAN
|
||||||
| NC score | 0.023981 (rank : 45) | θ value | 1.81305 (rank : 28) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q96DT5, Q9UJ82 | Gene names | DNAH11 | |||
|
Domain Architecture |

|
|||||
| Description | Ciliary dynein heavy chain 11 (Axonemal beta dynein heavy chain 11). | |||||
|
DYH9_HUMAN
|
||||||
| NC score | 0.023809 (rank : 46) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9NYC9, O95494, Q9NQ28 | Gene names | DNAH9, DNAH17L, DNAL1 | |||
|
Domain Architecture |

|
|||||
| Description | Ciliary dynein heavy chain 9 (Axonemal beta dynein heavy chain 9). | |||||
|
BAZ1A_HUMAN
|
||||||
| NC score | 0.023659 (rank : 47) | θ value | 1.38821 (rank : 25) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9NRL2, Q9NZ15, Q9P065, Q9UIG1, Q9Y3V3 | Gene names | BAZ1A, ACF1, WCRF180 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1A (ATP-utilizing chromatin assembly and remodeling factor 1) (hACF1) (ATP-dependent chromatin remodelling protein) (Williams syndrome transcription factor-related chromatin remodeling factor 180) (WCRF180) (hWALp1) (CHRAC subunit ACF1). | |||||
|
CCD46_MOUSE
|
||||||
| NC score | 0.023349 (rank : 48) | θ value | 1.38821 (rank : 26) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q5PR68, Q80ZN6, Q99JS4, Q9D9T1 | Gene names | Ccdc46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled coil domain-containing protein 46. | |||||
|
TGON1_MOUSE
|
||||||
| NC score | 0.023082 (rank : 49) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q62313 | Gene names | Tgoln1, Ttgn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 1 precursor (TGN38A). | |||||
|
ST1E1_HUMAN
|
||||||
| NC score | 0.022705 (rank : 50) | θ value | 4.03905 (rank : 50) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P49888, Q8N6X5 | Gene names | SULT1E1, STE | |||
|
Domain Architecture |

|
|||||
| Description | Estrogen sulfotransferase (EC 2.8.2.4) (Sulfotransferase, estrogen- preferring) (EST-1). | |||||
|
AT10A_HUMAN
|
||||||
| NC score | 0.022481 (rank : 51) | θ value | 0.813845 (rank : 21) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60312, Q969I4 | Gene names | ATP10A, ATP10C, ATPVC, KIAA0566 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (ATPVA) (Aminophospholipid translocase VA). | |||||
|
BPAEA_HUMAN
|
||||||
| NC score | 0.022284 (rank : 52) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1369 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O94833, Q5TBT0, Q8N1T8, Q8N8J3, Q8WXK9, Q96AK9, Q96DQ5, Q9H555 | Gene names | DST, BPAG1, DMH, DT, KIAA0728 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 6/9/10 (Trabeculin-beta) (Bullous pemphigoid antigen) (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
CCD46_HUMAN
|
||||||
| NC score | 0.021550 (rank : 53) | θ value | 3.0926 (rank : 36) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1430 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8N8E3, Q6PIB5, Q8NCR4, Q8NFR4 | Gene names | CCDC46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 46. | |||||
|
P52K_HUMAN
|
||||||
| NC score | 0.021524 (rank : 54) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43422, Q8WTW1, Q9Y3Z4 | Gene names | PRKRIR, DAP4, P52RIPK, THAP0 | |||
|
Domain Architecture |

|
|||||
| Description | 52 kDa repressor of the inhibitor of the protein kinase (p58IPK- interacting protein) (58 kDa interferon-induced protein kinase- interacting protein) (P52rIPK) (Death-associated protein 4) (THAP domain-containing protein 0). | |||||
|
FIBP_MOUSE
|
||||||
| NC score | 0.020119 (rank : 55) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JI19 | Gene names | Fibp | |||
|
Domain Architecture |

|
|||||
| Description | Acidic fibroblast growth factor intracellular-binding protein (aFGF intracellular-binding protein) (FGF-1 intracellular-binding protein). | |||||
|
DMP1_HUMAN
|
||||||
| NC score | 0.019997 (rank : 56) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
DCDC2_HUMAN
|
||||||
| NC score | 0.019915 (rank : 57) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UHG0, Q5VTR8, Q5VTR9, Q86W35, Q9UFD1, Q9ULR6 | Gene names | DCDC2, KIAA1154 | |||
|
Domain Architecture |

|
|||||
| Description | Doublecortin domain-containing protein 2 (RU2S protein). | |||||
|
LMO7_HUMAN
|
||||||
| NC score | 0.019868 (rank : 58) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8WWI1, O15462, O95346, Q9UKC1, Q9UQM5, Q9Y6A7 | Gene names | LMO7, FBX20, FBXO20, KIAA0858 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain only protein 7 (LOMP) (F-box only protein 20). | |||||
|
RXINP_HUMAN
|
||||||
| NC score | 0.019831 (rank : 59) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96RL1, Q5XKQ1, Q7Z3W7, Q8N5B9, Q9BZR1, Q9BZR5, Q9UHX7 | Gene names | RXRIP110, RAP80 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoid X receptor-interacting protein 110 (Receptor-associated protein 80) (Nuclear zinc finger protein RAP80). | |||||
|
BPA1_MOUSE
|
||||||
| NC score | 0.018562 (rank : 60) | θ value | 3.0926 (rank : 35) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
ANR11_HUMAN
|
||||||
| NC score | 0.018412 (rank : 61) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
DESP_HUMAN
|
||||||
| NC score | 0.018338 (rank : 62) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P15924, O75993, Q14189, Q9UHN4 | Gene names | DSP | |||
|
Domain Architecture |

|
|||||
| Description | Desmoplakin (DP) (250/210 kDa paraneoplastic pemphigus antigen). | |||||
|
KIF5A_MOUSE
|
||||||
| NC score | 0.017530 (rank : 63) | θ value | 1.38821 (rank : 27) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P33175, Q5DTP1, Q6PDY7, Q9Z2F9 | Gene names | Kif5a, Kiaa4086, Kif5, Nkhc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin heavy chain isoform 5A (Neuronal kinesin heavy chain) (NKHC) (Kinesin heavy chain neuron-specific 1). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.017189 (rank : 64) | θ value | 3.0926 (rank : 38) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
CK5P2_HUMAN
|
||||||
| NC score | 0.016948 (rank : 65) | θ value | 3.0926 (rank : 37) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1433 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96SN8, Q7Z3L4, Q7Z3U1, Q7Z7I6, Q9BSW0, Q9H6J6, Q9HCD9, Q9NV90, Q9UIW9 | Gene names | CDK5RAP2, CEP215, KIAA1633 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48) (Centrosome-associated protein 215). | |||||
|
CP135_HUMAN
|
||||||
| NC score | 0.016702 (rank : 66) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1290 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q66GS9, O75130, Q9H8H7 | Gene names | CEP135, CEP4, KIAA0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
LAMA2_MOUSE
|
||||||
| NC score | 0.016235 (rank : 67) | θ value | 0.365318 (rank : 18) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1198 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q60675, Q05003, Q64061 | Gene names | Lama2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
TGON2_MOUSE
|
||||||
| NC score | 0.015611 (rank : 68) | θ value | 8.99809 (rank : 98) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q62314 | Gene names | Tgoln2, Ttgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (TGN38B). | |||||
|
IF4G3_MOUSE
|
||||||
| NC score | 0.015480 (rank : 69) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80XI3, Q80Y69, Q8BJ68, Q8BQF3, Q8C6Q3, Q8CIH0 | Gene names | Eif4g3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 3 (eIF-4-gamma 3) (eIF-4G 3) (eIF4G 3) (eIF-4-gamma II) (eIF4GII). | |||||
|
IF4G3_HUMAN
|
||||||
| NC score | 0.015343 (rank : 70) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O43432, Q15597, Q8NEN1 | Gene names | EIF4G3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 3 (eIF-4-gamma 3) (eIF-4G 3) (eIF4G 3) (eIF-4-gamma II) (eIF4GII). | |||||
|
TTC1_HUMAN
|
||||||
| NC score | 0.015123 (rank : 71) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q99614, Q9BVT3 | Gene names | TTC1, TPR1 | |||
|
Domain Architecture |

|
|||||
| Description | Tetratricopeptide repeat protein 1 (TPR repeat protein 1). | |||||
|
ADA30_HUMAN
|
||||||
| NC score | 0.014589 (rank : 72) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UKF2, Q9UKF1 | Gene names | ADAM30 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 30 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 30). | |||||
|
KIF5C_HUMAN
|
||||||
| NC score | 0.014426 (rank : 73) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 974 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O60282, O95079 | Gene names | KIF5C, KIAA0531, NKHC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin heavy chain isoform 5C (Kinesin heavy chain neuron-specific 2). | |||||
|
AFF4_HUMAN
|
||||||
| NC score | 0.013666 (rank : 74) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
S12A6_HUMAN
|
||||||
| NC score | 0.013577 (rank : 75) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UHW9, Q8TDD4, Q9UFR2, Q9Y642, Q9Y665 | Gene names | SLC12A6, KCC3 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 12 member 6 (Electroneutral potassium-chloride cotransporter 3) (K-Cl cotransporter 3). | |||||
|
S12A6_MOUSE
|
||||||
| NC score | 0.013552 (rank : 76) | θ value | 5.27518 (rank : 63) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q924N4, Q924N3 | Gene names | Slc12a6, Kcc3 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 12 member 6 (Electroneutral potassium-chloride cotransporter 3) (K-Cl cotransporter 3). | |||||
|
CLC6A_HUMAN
|
||||||
| NC score | 0.013102 (rank : 77) | θ value | 2.36792 (rank : 31) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6EIG7 | Gene names | CLEC6A, CLECSF10, DECTIN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-type lectin domain family 6 member A (C-type lectin superfamily member 10) (Dendritic cell-associated lectin 2) (DC-associated C-type lectin-2) (DECTIN-2). | |||||
|
AT10A_MOUSE
|
||||||
| NC score | 0.013063 (rank : 78) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O54827, Q8R3B8 | Gene names | Atp10a, Atpc5, Pfatp | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (P-locus fat-associated ATPase). | |||||
|
RGS6_HUMAN
|
||||||
| NC score | 0.012383 (rank : 79) | θ value | 3.0926 (rank : 41) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P49758, O75576, O75577, Q8TE13, Q8TE14, Q8TE15, Q8TE16, Q8TE17, Q8TE18, Q8TE19, Q8TE20, Q8TE21, Q8TE22, Q9Y245, Q9Y647 | Gene names | RGS6 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 6 (RGS6) (S914). | |||||
|
OXSR1_MOUSE
|
||||||
| NC score | 0.012383 (rank : 80) | θ value | 0.365318 (rank : 19) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 865 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6P9R2, Q8BVZ9, Q8C0B9 | Gene names | Oxsr1, Osr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase OSR1 (EC 2.7.11.1) (Oxidative stress- responsive 1 protein). | |||||
|
ANR26_MOUSE
|
||||||
| NC score | 0.011855 (rank : 81) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1554 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q811D2, Q69ZS2, Q9CS61 | Gene names | Ankrd26, Kiaa1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
OXSR1_HUMAN
|
||||||
| NC score | 0.011040 (rank : 82) | θ value | 1.06291 (rank : 24) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O95747, Q3LR53, Q7Z501, Q9UPQ1 | Gene names | OXSR1, KIAA1101, OSR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase OSR1 (EC 2.7.11.1) (Oxidative stress- responsive 1 protein). | |||||
|
PITX2_HUMAN
|
||||||
| NC score | 0.010583 (rank : 83) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99697, O60578, O60579, O60580, Q9BY17 | Gene names | PITX2, ARP1, RGS, RIEG, RIEG1 | |||
|
Domain Architecture |

|
|||||
| Description | Pituitary homeobox 2 (RIEG bicoid-related homeobox transcription factor) (Solurshin) (ALL1-responsive protein ARP1). | |||||
|
ZA2G_MOUSE
|
||||||
| NC score | 0.007738 (rank : 84) | θ value | 3.0926 (rank : 45) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q64726 | Gene names | Azgp1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-alpha-2-glycoprotein precursor (Zn-alpha-2-glycoprotein) (Zn- alpha-2-GP). | |||||
|
STK39_MOUSE
|
||||||
| NC score | 0.007667 (rank : 85) | θ value | 4.03905 (rank : 52) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 915 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Z1W9, Q80W13 | Gene names | Stk39, Spak | |||
|
Domain Architecture |

|
|||||
| Description | STE20/SPS1-related proline-alanine-rich protein kinase (EC 2.7.11.1) (Ste-20-related kinase) (Serine/threonine-protein kinase 39). | |||||
|
PEG3_MOUSE
|
||||||
| NC score | 0.007640 (rank : 86) | θ value | 8.99809 (rank : 93) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
TEX14_MOUSE
|
||||||
| NC score | 0.007599 (rank : 87) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1333 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q7M6U3, Q3UT36, Q5NC10, Q5NC11, Q8CGK1, Q8CGK2, Q99MV8 | Gene names | Tex14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed protein 14 (Testis-expressed sequence 14). | |||||
|
PITX2_MOUSE
|
||||||
| NC score | 0.007368 (rank : 88) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P97474, O08646, O70336, P97933, Q9JLA0, Q9QXB8, Q9R1V9, Q9Z141 | Gene names | Pitx2, Arp1, Brx1, Otlx2, Ptx2, Rgs | |||
|
Domain Architecture |

|
|||||
| Description | Pituitary homeobox 2 (Orthodenticle-like homeobox 2) (Solurshin) (ALL1-responsive protein ARP1) (BRX1 homeoprotein) (Paired-like homeodomain transcription factor Munc 30). | |||||
|
STK39_HUMAN
|
||||||
| NC score | 0.007045 (rank : 89) | θ value | 4.03905 (rank : 51) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 882 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UEW8, O14774, Q9UER4 | Gene names | STK39, SPAK | |||
|
Domain Architecture |

|
|||||
| Description | STE20/SPS1-related proline-alanine-rich protein kinase (EC 2.7.11.1) (Ste-20-related kinase) (Serine/threonine-protein kinase 39) (DCHT). | |||||
|
SHOX_HUMAN
|
||||||
| NC score | 0.006950 (rank : 90) | θ value | 8.99809 (rank : 97) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O15266, O00412, O00413, O15267 | Gene names | SHOX, PHOG | |||
|
Domain Architecture |

|
|||||
| Description | Short stature homeobox protein (Short stature homeobox-containing protein) (Pseudoautosomal homeobox-containing osteogenic protein). | |||||
|
TRI36_HUMAN
|
||||||
| NC score | 0.004983 (rank : 91) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NQ86 | Gene names | TRIM36, RBCC728, RNF98 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 36 (Zinc-binding protein Rbcc728) (RING finger protein 98). | |||||
|
NKX61_MOUSE
|
||||||
| NC score | 0.004050 (rank : 92) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99MA9, Q9ERQ7 | Gene names | Nkx6-1, Nkx6.1, Nkx6a | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Nkx-6.1. | |||||
|
NKX61_HUMAN
|
||||||
| NC score | 0.003674 (rank : 93) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P78426 | Gene names | NKX6-1, NKX6A | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Nkx-6.1. | |||||
|
ABCG2_MOUSE
|
||||||
| NC score | 0.003495 (rank : 94) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7TMS5, Q9R004, Q9Z1T0 | Gene names | Abcg2, Abcp, Bcrp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-binding cassette sub-family G member 2 (Breast cancer resistance protein 1 homolog). | |||||
|
SC10A_HUMAN
|
||||||
| NC score | 0.003020 (rank : 95) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5Y9 | Gene names | SCN10A, PN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 10 subunit alpha (Sodium channel protein type X subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.8) (Peripheral nerve sodium channel 3) (hPN3). | |||||
|
EVI1_HUMAN
|
||||||
| NC score | 0.002959 (rank : 96) | θ value | 1.06291 (rank : 23) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 924 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q03112, Q16122, Q99917 | Gene names | EVI1 | |||
|
Domain Architecture |

|
|||||
| Description | Ecotropic virus integration 1 site protein. | |||||
|
SALL1_MOUSE
|
||||||
| NC score | 0.001178 (rank : 97) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 826 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9ER74, Q920R5 | Gene names | Sall1, Sal3 | |||
|
Domain Architecture |

|
|||||
| Description | Sal-like protein 1 (Zinc finger protein Spalt-3) (Sal-3) (MSal-3). | |||||
|
ZBT24_HUMAN
|
||||||
| NC score | -0.000210 (rank : 98) | θ value | 3.0926 (rank : 46) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43167, Q5TED5, Q8N455 | Gene names | ZBTB24, KIAA0441, ZNF450 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and BTB domain-containing protein 24 (Zinc finger protein 450). | |||||