Please be patient as the page loads
|
EST1A_MOUSE
|
||||||
| SwissProt Accessions | P61406, Q5NC64 | Gene names | Est1a, Kiaa0732 | |||
|
Domain Architecture |

|
|||||
| Description | Telomerase-binding protein EST1A (Ever shorter telomeres 1A) (Telomerase subunit EST1A) (EST1-like protein A). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
EST1A_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.966858 (rank : 2) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q86US8, O94837, Q86VH6, Q9UF60 | Gene names | EST1A, C17orf31, KIAA0732 | |||
|
Domain Architecture |
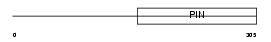
|
|||||
| Description | Telomerase-binding protein EST1A (Ever shorter telomeres 1A) (Telomerase subunit EST1A) (EST1-like protein A) (hSmg5/7a). | |||||
|
EST1A_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 138 | |
| SwissProt Accessions | P61406, Q5NC64 | Gene names | Est1a, Kiaa0732 | |||
|
Domain Architecture |

|
|||||
| Description | Telomerase-binding protein EST1A (Ever shorter telomeres 1A) (Telomerase subunit EST1A) (EST1-like protein A). | |||||
|
SMG7_HUMAN
|
||||||
| θ value | 7.0802e-21 (rank : 3) | NC score | 0.521121 (rank : 3) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92540, Q5T1Q0, Q6PIE0, Q7Z7H9, Q8IXC1, Q8IXC2 | Gene names | SMG7, C1orf16, EST1C, KIAA0250 | |||
|
Domain Architecture |

|
|||||
| Description | Protein SMG7 (SMG-7 homolog) (EST1-like protein C). | |||||
|
SMG7_MOUSE
|
||||||
| θ value | 3.51386e-20 (rank : 4) | NC score | 0.519478 (rank : 4) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5RJH6, Q63ZW5, Q6ZQF3 | Gene names | Smg7, Est1c, Kiaa0250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein SMG7 (SMG-7 homolog) (EST1-like protein C). | |||||
|
SMG5_MOUSE
|
||||||
| θ value | 8.38298e-14 (rank : 5) | NC score | 0.450613 (rank : 5) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6ZPY2, Q497S2, Q8R3S4 | Gene names | Smg5, Est1b, Kiaa1089 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein SMG5 (SMG-5 homolog) (EST1-like protein B). | |||||
|
SMG5_HUMAN
|
||||||
| θ value | 1.2105e-12 (rank : 6) | NC score | 0.425382 (rank : 6) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UPR3, Q5QJE7, Q8IXC0, Q8IY09, Q96IJ7 | Gene names | SMG5, EST1B, KIAA1089 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein SMG5 (SMG-5 homolog) (EST1-like protein B) (LPTS-interacting protein) (LPTS-RP1). | |||||
|
CA026_MOUSE
|
||||||
| θ value | 5.62301e-10 (rank : 7) | NC score | 0.335907 (rank : 7) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9DBQ9, Q6PAN2, Q7TMQ6 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C1orf26 homolog. | |||||
|
CA026_HUMAN
|
||||||
| θ value | 6.87365e-08 (rank : 8) | NC score | 0.310077 (rank : 8) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5T5J6, Q8NEK9, Q9BZQ7, Q9NXQ0 | Gene names | C1orf26 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C1orf26. | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 8.40245e-06 (rank : 9) | NC score | 0.091053 (rank : 12) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
SPT5H_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 10) | NC score | 0.141586 (rank : 9) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O00267, O43279, Q59G52, Q99639 | Gene names | SUPT5H, SPT5, SPT5H | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT5 (hSPT5) (DRB sensitivity-inducing factor large subunit) (DSIF large subunit) (DSIF p160) (Tat- cotransactivator 1 protein) (Tat-CT1 protein). | |||||
|
SPT5H_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 11) | NC score | 0.139204 (rank : 10) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 437 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O55201, Q3UJ77, Q3UJH1, Q3UKD7, Q3UM54, Q6PB73, Q6PDP0, Q6PFR4 | Gene names | Supt5h | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT5 (DRB sensitivity-inducing factor large subunit) (DSIF large subunit). | |||||
|
REN3B_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 12) | NC score | 0.105090 (rank : 11) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9BZI7, Q9H1J0 | Gene names | UPF3B, RENT3B, UPF3X | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of nonsense transcripts 3B (Nonsense mRNA reducing factor 3B) (Up-frameshift suppressor 3 homolog B) (hUpf3B) (hUpf3p-X). | |||||
|
MYH11_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 13) | NC score | 0.070582 (rank : 20) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
ROCK2_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 14) | NC score | 0.036606 (rank : 114) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 2072 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P70336, Q8BR64, Q8CBR0 | Gene names | Rock2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2). | |||||
|
MYH8_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 15) | NC score | 0.067876 (rank : 24) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1600 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P13542, Q5SX36 | Gene names | Myh8, Myhsp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
ZNFX1_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 16) | NC score | 0.071100 (rank : 18) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 407 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8R151, Q3U016, Q3UD71, Q3UEZ0 | Gene names | ZNFX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
MAST2_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 17) | NC score | 0.019337 (rank : 148) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1110 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q60592, Q6P9M1, Q8CHD1 | Gene names | Mast2, Mast205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 2 (EC 2.7.11.1). | |||||
|
KLC1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 18) | NC score | 0.074828 (rank : 14) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q07866, Q7RTM2, Q7RTM3, Q7RTM5, Q7RTP9, Q7RTQ5, Q86SF5, Q86TF5, Q86V74, Q86V75, Q86V76, Q86V77, Q86V78, Q86V79, Q96H62 | Gene names | KNS2, KLC, KLC1 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin light chain 1 (KLC 1). | |||||
|
MYH8_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 19) | NC score | 0.067313 (rank : 29) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1610 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | P13535, Q14910 | Gene names | MYH8 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
CAR11_HUMAN
|
||||||
| θ value | 0.125558 (rank : 20) | NC score | 0.073743 (rank : 15) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9BXL7, Q548H3 | Gene names | CARD11, CARMA1 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 11 (CARD-containing MAGUK protein 3) (Carma 1). | |||||
|
MYH11_MOUSE
|
||||||
| θ value | 0.125558 (rank : 21) | NC score | 0.067840 (rank : 25) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
MYH13_HUMAN
|
||||||
| θ value | 0.125558 (rank : 22) | NC score | 0.068560 (rank : 22) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
EHMT2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 23) | NC score | 0.035954 (rank : 116) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q96KQ7, Q14349, Q6PK06, Q96MH5, Q96QD0, Q9UQL8, Q9Y331 | Gene names | EHMT2, BAT8, G9A, NG36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
EHMT2_MOUSE
|
||||||
| θ value | 0.163984 (rank : 24) | NC score | 0.035439 (rank : 120) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Z148, Q6PE08, Q8K4R6, Q8K4R7, Q9Z149 | Gene names | Ehmt2, Bat8, G9a, Ng36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
MYH1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 25) | NC score | 0.067735 (rank : 26) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1478 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | P12882, Q9Y622 | Gene names | MYH1 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-1 (Myosin heavy chain, skeletal muscle, adult 1) (Myosin heavy chain IIx/d) (MyHC-IIx/d). | |||||
|
CALD1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 26) | NC score | 0.071196 (rank : 17) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q05682, Q13978, Q13979, Q14741, Q14742 | Gene names | CALD1, CAD, CDM | |||
|
Domain Architecture |

|
|||||
| Description | Caldesmon (CDM). | |||||
|
ATRX_HUMAN
|
||||||
| θ value | 0.279714 (rank : 27) | NC score | 0.057969 (rank : 53) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P46100, P51068, Q15886, Q59FB5, Q59H31, Q5H9A2, Q5JWI4, Q7Z2J1, Q9H0Z1, Q9NTS3 | Gene names | ATRX, RAD54L, XH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked helicase II) (X-linked nuclear protein) (XNP) (Znf- HX). | |||||
|
MORC4_HUMAN
|
||||||
| θ value | 0.279714 (rank : 28) | NC score | 0.056070 (rank : 61) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8TE76, Q5JUK7, Q96MZ2, Q9HAI7 | Gene names | MORC4, ZCWCC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 4 (Zinc finger CW-type coiled- coil domain protein 2). | |||||
|
MYH2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 29) | NC score | 0.066916 (rank : 30) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1491 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9UKX2, Q14322, Q16229 | Gene names | MYH2, MYHSA2 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, skeletal muscle, adult 2 (Myosin heavy chain IIa) (MyHC-IIa). | |||||
|
TOP1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 30) | NC score | 0.051677 (rank : 87) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P11387, O43256, Q12855, Q12856, Q15610, Q5TFY3, Q9UJN0 | Gene names | TOP1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 1 (EC 5.99.1.2) (DNA topoisomerase I). | |||||
|
CHD1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 31) | NC score | 0.035413 (rank : 121) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
CP135_HUMAN
|
||||||
| θ value | 0.365318 (rank : 32) | NC score | 0.070870 (rank : 19) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1290 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q66GS9, O75130, Q9H8H7 | Gene names | CEP135, CEP4, KIAA0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
HTSF1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 33) | NC score | 0.054781 (rank : 66) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BGC0, Q1WWK0, Q9CT41, Q9DAU3 | Gene names | Htatsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 homolog. | |||||
|
BRWD1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 34) | NC score | 0.027161 (rank : 134) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NSI6, O43721, Q5R2V0, Q5R2V1, Q8TCV3, Q96QG9, Q96QH0, Q9NUK1 | Gene names | BRWD1, C21orf107, WDR9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain and WD repeat domain-containing protein 1 (WD repeat protein 9). | |||||
|
CING_MOUSE
|
||||||
| θ value | 0.47712 (rank : 35) | NC score | 0.068312 (rank : 23) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1432 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | P59242 | Gene names | Cgn | |||
|
Domain Architecture |

|
|||||
| Description | Cingulin. | |||||
|
MAST2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 36) | NC score | 0.014221 (rank : 163) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1189 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6P0Q8, O94899, Q5VT07, Q5VT08, Q7LGC4, Q8NDG1, Q96B94, Q9BYE8 | Gene names | MAST2, KIAA0807, MAST205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 2 (EC 2.7.11.1). | |||||
|
MICA1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 37) | NC score | 0.048759 (rank : 99) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8VDP3 | Gene names | Mical1, Mical, Nical | |||
|
Domain Architecture |

|
|||||
| Description | NEDD9-interacting protein with calponin homology and LIM domains (Molecule interacting with CasL protein 1). | |||||
|
MYH9_MOUSE
|
||||||
| θ value | 0.47712 (rank : 38) | NC score | 0.069824 (rank : 21) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
BPAEA_MOUSE
|
||||||
| θ value | 0.62314 (rank : 39) | NC score | 0.058180 (rank : 50) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1480 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q91ZU8, Q8K5D4 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoform 5 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
CHD1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 40) | NC score | 0.032704 (rank : 129) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
IF4G1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 41) | NC score | 0.039677 (rank : 112) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6NZJ6, Q6NZN8, Q8BW99 | Gene names | Eif4g1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1). | |||||
|
MDN1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 42) | NC score | 0.054918 (rank : 65) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9NU22, O15019 | Gene names | MDN1, KIAA0301 | |||
|
Domain Architecture |

|
|||||
| Description | Midasin (MIDAS-containing protein). | |||||
|
E41L2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 43) | NC score | 0.026436 (rank : 136) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O43491 | Gene names | EPB41L2 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 2 (Generally expressed protein 4.1) (4.1G). | |||||
|
KLC1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 44) | NC score | 0.061288 (rank : 40) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O88447 | Gene names | Kns2, Klc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin light chain 1 (KLC 1). | |||||
|
LUZP1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 45) | NC score | 0.059506 (rank : 48) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 940 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q8R4U7, Q3UV18, Q7TS71, Q8BQW1, Q99NG3, Q9CSL6 | Gene names | Luzp1, Luzp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper protein 1 (Leucine zipper motif-containing protein). | |||||
|
TCF8_HUMAN
|
||||||
| θ value | 0.813845 (rank : 46) | NC score | 0.006071 (rank : 175) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1057 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P37275, Q12924, Q13800 | Gene names | TCF8, AREB6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (NIL-2-A zinc finger protein) (Negative regulator of IL2). | |||||
|
UBF1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 47) | NC score | 0.056057 (rank : 62) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P17480 | Gene names | UBTF, UBF, UBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1) (Autoantigen NOR-90). | |||||
|
UBF1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 48) | NC score | 0.055718 (rank : 63) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P25976 | Gene names | Ubtf, Tcfubf, Ubf-1, Ubf1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1). | |||||
|
AKAP9_HUMAN
|
||||||
| θ value | 1.06291 (rank : 49) | NC score | 0.058098 (rank : 51) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1710 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q99996, O14869, O43355, O94895, Q9UQH3, Q9UQQ4, Q9Y6B8, Q9Y6Y2 | Gene names | AKAP9, AKAP350, AKAP450, KIAA0803 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 9 (Protein kinase A-anchoring protein 9) (PRKA9) (A-kinase anchor protein 450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP 350) (hgAKAP 350) (AKAP 120-like protein) (Protein hyperion) (Protein yotiao) (Centrosome- and Golgi-localized PKN-associated protein) (CG-NAP). | |||||
|
CRSP2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 50) | NC score | 0.055636 (rank : 64) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O60244, Q9UNB3 | Gene names | CRSP2, ARC150, DRIP150, EXLM1, TRAP170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CRSP complex subunit 2 (Cofactor required for Sp1 transcriptional activation subunit 2) (Transcriptional coactivator CRSP150) (Vitamin D3 receptor-interacting protein complex 150 kDa component) (DRIP150) (Thyroid hormone receptor-associated protein complex 170 kDa component) (Trap170) (Activator-recruited cofactor 150 kDa component) (ARC150). | |||||
|
PF21B_HUMAN
|
||||||
| θ value | 1.06291 (rank : 51) | NC score | 0.025845 (rank : 137) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96EK2, Q5TFL2, Q6ICC0 | Gene names | PHF21B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
RBM25_HUMAN
|
||||||
| θ value | 1.06291 (rank : 52) | NC score | 0.067363 (rank : 27) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P49756, Q9UEQ5, Q9UIE9 | Gene names | RBM25, RNPC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable RNA-binding protein 25 (RNA-binding motif protein 25) (RNA- binding region-containing protein 7) (Protein S164). | |||||
|
SETD2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 53) | NC score | 0.047678 (rank : 101) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
SOX9_HUMAN
|
||||||
| θ value | 1.06291 (rank : 54) | NC score | 0.030877 (rank : 130) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P48436 | Gene names | SOX9 | |||
|
Domain Architecture |
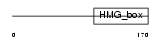
|
|||||
| Description | Transcription factor SOX-9. | |||||
|
BSN_HUMAN
|
||||||
| θ value | 1.38821 (rank : 55) | NC score | 0.035469 (rank : 118) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
CDC37_HUMAN
|
||||||
| θ value | 1.38821 (rank : 56) | NC score | 0.057974 (rank : 52) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q16543 | Gene names | CDC37 | |||
|
Domain Architecture |
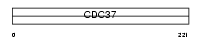
|
|||||
| Description | Hsp90 co-chaperone Cdc37 (Hsp90 chaperone protein kinase-targeting subunit) (p50Cdc37). | |||||
|
CP135_MOUSE
|
||||||
| θ value | 1.38821 (rank : 57) | NC score | 0.067335 (rank : 28) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1278 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q6P5D4, Q6A030 | Gene names | Cep135, Cep4, Kiaa0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
FOXN4_HUMAN
|
||||||
| θ value | 1.38821 (rank : 58) | NC score | 0.013677 (rank : 164) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96NZ1, Q6ZMR4, Q96NZ0 | Gene names | FOXN4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein N4. | |||||
|
IF4G1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 59) | NC score | 0.033411 (rank : 127) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q04637, O43177, O95066, Q5HYG0, Q6ZN21, Q8N102 | Gene names | EIF4G1, EIF4G, EIF4GI | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1) (p220). | |||||
|
MYH7_HUMAN
|
||||||
| θ value | 1.38821 (rank : 60) | NC score | 0.066456 (rank : 31) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
PLEC1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 61) | NC score | 0.048164 (rank : 100) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1326 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q15149, Q15148, Q16640 | Gene names | PLEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Plectin-1 (PLTN) (PCN) (Hemidesmosomal protein 1) (HD1). | |||||
|
SDCG8_MOUSE
|
||||||
| θ value | 1.38821 (rank : 62) | NC score | 0.060273 (rank : 45) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q80UF4 | Gene names | Sdccag8, Cccap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serologically defined colon cancer antigen 8 (Centrosomal colon cancer autoantigen protein) (mCCCAP). | |||||
|
TRHY_HUMAN
|
||||||
| θ value | 1.38821 (rank : 63) | NC score | 0.057595 (rank : 56) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
CCD41_MOUSE
|
||||||
| θ value | 1.81305 (rank : 64) | NC score | 0.056920 (rank : 59) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9D5R3, Q3U7X7, Q3UX57, Q80VF0 | Gene names | Ccdc41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41. | |||||
|
CUTL1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | 0.046953 (rank : 103) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
LATS1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 66) | NC score | 0.014494 (rank : 162) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1027 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BYR2, Q9Z0W4 | Gene names | Lats1, Warts | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase LATS1 (EC 2.7.11.1) (Large tumor suppressor homolog 1) (WARTS protein kinase). | |||||
|
TS101_HUMAN
|
||||||
| θ value | 1.81305 (rank : 67) | NC score | 0.051857 (rank : 85) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q99816, Q9BUM5 | Gene names | TSG101 | |||
|
Domain Architecture |
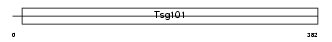
|
|||||
| Description | Tumor susceptibility gene 101 protein. | |||||
|
BRWD1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.022335 (rank : 142) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 478 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q921C3, Q921C2 | Gene names | Brwd1, Wdr9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and WD repeat domain-containing protein 1 (WD repeat protein 9). | |||||
|
CBX4_HUMAN
|
||||||
| θ value | 2.36792 (rank : 69) | NC score | 0.023458 (rank : 140) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O00257, Q96C04 | Gene names | CBX4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 4 (Polycomb 2 homolog) (Pc2) (hPc2). | |||||
|
CING_HUMAN
|
||||||
| θ value | 2.36792 (rank : 70) | NC score | 0.065231 (rank : 32) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1275 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9P2M7, Q5T386, Q9NR25 | Gene names | CGN, KIAA1319 | |||
|
Domain Architecture |

|
|||||
| Description | Cingulin. | |||||
|
DDX11_MOUSE
|
||||||
| θ value | 2.36792 (rank : 71) | NC score | 0.019595 (rank : 147) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6AXC6 | Gene names | Ddx11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX11 (EC 3.6.1.-) (DEAD/H box protein 11). | |||||
|
NFM_MOUSE
|
||||||
| θ value | 2.36792 (rank : 72) | NC score | 0.035274 (rank : 123) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P08553, Q61961 | Gene names | Nef3, Nefm, Nfm | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 2.36792 (rank : 73) | NC score | 0.016882 (rank : 154) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
PC11X_HUMAN
|
||||||
| θ value | 2.36792 (rank : 74) | NC score | 0.007517 (rank : 171) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BZA7, Q2TJH0, Q2TJH1, Q2TJH3, Q5JVZ0, Q70LR8, Q70LS7, Q70LS8, Q70LS9, Q70LT7, Q70LT8, Q70LT9, Q70LU0, Q70LU1, Q96RV4, Q96RW0, Q9BZA6, Q9H4E0, Q9P2M0, Q9P2X5 | Gene names | PCDH11X, PCDH11, PCDHX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protocadherin-11 X-linked precursor (Protocadherin-11) (Protocadherin on the X chromosome) (PCDH-X) (Protocadherin-S). | |||||
|
TRI35_MOUSE
|
||||||
| θ value | 2.36792 (rank : 75) | NC score | 0.018841 (rank : 151) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 465 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8C006, Q6ZPY0, Q7TQL7, Q810V7, Q8BVY9, Q8VID4, Q9CW74, Q9D208 | Gene names | Trim35, Hls5, Kiaa1098, Mair | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 35 (Macrophage-derived apoptosis- inducing RBCC protein) (Protein MAIR) (Protein Nc8) (Hemopoietic lineage switch protein 5). | |||||
|
DIAP2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 76) | NC score | 0.020831 (rank : 145) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O60879, O60878, Q8WX06, Q8WX48, Q9UJL2 | Gene names | DIAPH2, DIA | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 2 (Diaphanous-related formin-2) (DRF2). | |||||
|
GOGA4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 77) | NC score | 0.054352 (rank : 69) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
KI67_HUMAN
|
||||||
| θ value | 3.0926 (rank : 78) | NC score | 0.036518 (rank : 115) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
MYH1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 79) | NC score | 0.064402 (rank : 33) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1461 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q5SX40 | Gene names | Myh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-1 (Myosin heavy chain, skeletal muscle, adult 1). | |||||
|
MYST3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 80) | NC score | 0.022305 (rank : 143) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
SH24A_MOUSE
|
||||||
| θ value | 3.0926 (rank : 81) | NC score | 0.034328 (rank : 126) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9D7V1 | Gene names | Sh2d4a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH2 domain-containing protein 4A. | |||||
|
SMCA2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 82) | NC score | 0.034381 (rank : 125) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P51531 | Gene names | SMARCA2, BRM, SNF2A, SNF2L2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L2 (EC 3.6.1.-) (ATP- dependent helicase SMARCA2) (SNF2-alpha) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin subfamily A member 2) (hBRM). | |||||
|
SPRL1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 83) | NC score | 0.028363 (rank : 131) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q14515, Q14800 | Gene names | SPARCL1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (High endothelial venule protein) (Hevin) (MAST 9). | |||||
|
THOC2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 84) | NC score | 0.072273 (rank : 16) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8NI27, Q9H8I6 | Gene names | THOC2, CXorf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | THO complex subunit 2 (Tho2). | |||||
|
WDR60_MOUSE
|
||||||
| θ value | 3.0926 (rank : 85) | NC score | 0.044424 (rank : 104) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8C761, Q3UNM4 | Gene names | Wdr60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 60. | |||||
|
ZFP38_HUMAN
|
||||||
| θ value | 3.0926 (rank : 86) | NC score | -0.001291 (rank : 177) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 766 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y5A6, Q9H0B5 | Gene names | ZNF38, ZFP38, ZSCAN21 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 38 homolog (Zfp-38) (Zinc finger and SCAN domain- containing protein 21) (NY-REN-21 antigen). | |||||
|
DDX41_MOUSE
|
||||||
| θ value | 4.03905 (rank : 87) | NC score | 0.007364 (rank : 172) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q91VN6 | Gene names | Ddx41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX41 (EC 3.6.1.-) (DEAD box protein 41). | |||||
|
EDF1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 88) | NC score | 0.035757 (rank : 117) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JMG1, Q99NE6 | Gene names | Edf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelial differentiation-related factor 1 (EDF-1) (Multiprotein- bridging factor 1) (MBF1). | |||||
|
EYA2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 89) | NC score | 0.011964 (rank : 167) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O08575, P97925 | Gene names | Eya2, Eab1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 2 (EC 3.1.3.48). | |||||
|
GCYB2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 90) | NC score | 0.016472 (rank : 155) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75343, Q9NZ64 | Gene names | GUCY1B2 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylate cyclase soluble subunit beta-2 (EC 4.6.1.2) (GCS-beta-2). | |||||
|
LIPA3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 91) | NC score | 0.039894 (rank : 111) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O75145, Q9H8B5, Q9UEW4 | Gene names | PPFIA3, KIAA0654 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-3 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-3) (PTPRF-interacting protein alpha-3). | |||||
|
MBP_MOUSE
|
||||||
| θ value | 4.03905 (rank : 92) | NC score | 0.024629 (rank : 138) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P04370, Q01585, Q03139, Q03176, Q61836, Q61837, Q99KE4, Q9QWP1 | Gene names | Mbp, Shi | |||
|
Domain Architecture |

|
|||||
| Description | Myelin basic protein (MBP) (Myelin A1 protein). | |||||
|
WDHD1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 93) | NC score | 0.016125 (rank : 158) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P59328, Q6P408 | Gene names | Wdhd1, And1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and HMG-box DNA-binding protein 1 (Acidic nucleoplasmic DNA- binding protein 1) (And-1). | |||||
|
ZCHC8_MOUSE
|
||||||
| θ value | 4.03905 (rank : 94) | NC score | 0.027618 (rank : 133) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9CYA6, Q3TK71, Q8CCB7, Q91WQ1 | Gene names | Zcchc8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 8. | |||||
|
ACINU_MOUSE
|
||||||
| θ value | 5.27518 (rank : 95) | NC score | 0.041789 (rank : 109) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9JIX8, Q9CSN7, Q9CSR9, Q9CSX7, Q9R046, Q9R047 | Gene names | Acin1, Acinus | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
BCN1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 96) | NC score | 0.044281 (rank : 105) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 367 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O88597 | Gene names | Becn1 | |||
|
Domain Architecture |

|
|||||
| Description | Beclin-1 (Coiled-coil myosin-like BCL2-interacting protein). | |||||
|
CIR_HUMAN
|
||||||
| θ value | 5.27518 (rank : 97) | NC score | 0.038546 (rank : 113) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q86X95, O95367, Q12804, Q4G1B9, Q6PJI4, Q8IWI2 | Gene names | CIR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor (Recepin). | |||||
|
DDX46_HUMAN
|
||||||
| θ value | 5.27518 (rank : 98) | NC score | 0.015136 (rank : 159) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q7L014, O94894, Q96EI0, Q9Y658 | Gene names | DDX46, KIAA0801 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX46 (EC 3.6.1.-) (DEAD box protein 46) (PRP5 homolog). | |||||
|
EDF1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 99) | NC score | 0.033332 (rank : 128) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60869, Q5T5T2, Q9UIM1 | Gene names | EDF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelial differentiation-related factor 1 (EDF-1) (Multiprotein- bridging factor 1) (MBF1). | |||||
|
ESCO1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 100) | NC score | 0.035403 (rank : 122) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q5FWF5, Q69YG4, Q69YS3, Q6IMD7, Q8N3Z5, Q8NBG2, Q96PX7 | Gene names | ESCO1, EFO1, KIAA1911 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyltransferase ESCO1 (EC 2.3.1.-) (Establishment of cohesion 1 homolog 1) (ECO1 homolog 1) (ESO1 homolog 1) (Establishment factor- like protein 1) (EFO1p) (hEFO1) (CTF7 homolog 1). | |||||
|
KLC2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 101) | NC score | 0.043652 (rank : 106) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9H0B6, Q9H9C8, Q9HA20 | Gene names | KLC2 | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin light chain 2 (KLC 2). | |||||
|
LATS1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 102) | NC score | 0.011391 (rank : 168) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1057 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O95835, Q6PKD0 | Gene names | LATS1, WARTS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase LATS1 (EC 2.7.11.1) (Large tumor suppressor homolog 1) (WARTS protein kinase) (h-warts). | |||||
|
NO40_MOUSE
|
||||||
| θ value | 5.27518 (rank : 103) | NC score | 0.024296 (rank : 139) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9ESX4 | Gene names | Zcchc17, Ps1d | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar protein of 40 kDa (pNO40) (Zinc finger CCHC domain- containing protein 17) (Putative S1 RNA-binding domain protein) (PS1D protein). | |||||
|
PARC_HUMAN
|
||||||
| θ value | 5.27518 (rank : 104) | NC score | 0.012267 (rank : 166) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IWT3, O75188, Q68CP2, Q68D92, Q8N3W9, Q9BU56 | Gene names | PARC, H7AP1, KIAA0708 | |||
|
Domain Architecture |

|
|||||
| Description | p53-associated parkin-like cytoplasmic protein (UbcH7-associated protein 1). | |||||
|
PCNT_HUMAN
|
||||||
| θ value | 5.27518 (rank : 105) | NC score | 0.051625 (rank : 88) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1551 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | O95613, O43152, Q7Z7C9 | Gene names | PCNT, KIAA0402, PCNT2 | |||
|
Domain Architecture |

|
|||||
| Description | Pericentrin (Pericentrin B) (Kendrin). | |||||
|
PHF20_HUMAN
|
||||||
| θ value | 5.27518 (rank : 106) | NC score | 0.026768 (rank : 135) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9BVI0, Q5JWY9, Q9BWV4, Q9BXA3, Q9BZW3, Q9H421, Q9H4J6, Q9NZ22 | Gene names | PHF20, C20orf104, GLEA2, HCA58, NZF, TZP | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58) (Glioma-expressed antigen 2) (Transcription factor TZP) (Novel zinc finger protein). | |||||
|
RNF31_MOUSE
|
||||||
| θ value | 5.27518 (rank : 107) | NC score | 0.014598 (rank : 161) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q924T7 | Gene names | Rnf31 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 31. | |||||
|
SOX9_MOUSE
|
||||||
| θ value | 5.27518 (rank : 108) | NC score | 0.017024 (rank : 153) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q04887, Q8C7L2, Q91ZK2, Q99KQ0 | Gene names | Sox9, Sox-9 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-9. | |||||
|
CD209_HUMAN
|
||||||
| θ value | 6.88961 (rank : 109) | NC score | 0.015028 (rank : 160) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NNX6, Q96QP7, Q96QP8, Q96QP9, Q96QQ0, Q96QQ1, Q96QQ2, Q96QQ3, Q96QQ4, Q96QQ5, Q96QQ6, Q96QQ7, Q96QQ8 | Gene names | CD209, CLEC4L | |||
|
Domain Architecture |

|
|||||
| Description | CD209 antigen (Dendritic cell-specific ICAM-3-grabbing nonintegrin 1) (DC-SIGN1) (DC-SIGN) (C-type lectin domain family 4 member L). | |||||
|
CENPF_HUMAN
|
||||||
| θ value | 6.88961 (rank : 110) | NC score | 0.052777 (rank : 79) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
FGD6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 111) | NC score | 0.006107 (rank : 174) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q69ZL1, Q8C8W5, Q8K3B0, Q9D3Y7 | Gene names | Fgd6, Kiaa1362 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 6. | |||||
|
HPPD_HUMAN
|
||||||
| θ value | 6.88961 (rank : 112) | NC score | 0.016227 (rank : 157) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P32754, Q13234 | Gene names | HPD, PPD | |||
|
Domain Architecture |

|
|||||
| Description | 4-hydroxyphenylpyruvate dioxygenase (EC 1.13.11.27) (4HPPD) (HPD) (HPPDase) (4-hydroxyphenylpyruvic acid oxidase). | |||||
|
IG2AS_HUMAN
|
||||||
| θ value | 6.88961 (rank : 113) | NC score | 0.035201 (rank : 124) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6U949, Q7Z712, Q9P2W0 | Gene names | IGF2AS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative insulin-like growth factor 2 antisense gene protein (IGF2-AS) (PEG8/IGF2AS protein). | |||||
|
MICA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 114) | NC score | 0.021780 (rank : 144) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8TDZ2, Q7Z633, Q8IVS9, Q96G47, Q9H6X6, Q9H7I0, Q9HAA1, Q9UFF7 | Gene names | MICAL1, MICAL, NICAL | |||
|
Domain Architecture |

|
|||||
| Description | NEDD9-interacting protein with calponin homology and LIM domains (Molecule interacting with CasL protein 1). | |||||
|
PHLB3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 115) | NC score | 0.043093 (rank : 107) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 304 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6NSJ2, Q8N7Z4 | Gene names | PHLDB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 3. | |||||
|
PMS2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 116) | NC score | 0.012762 (rank : 165) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P54279 | Gene names | Pms2 | |||
|
Domain Architecture |
|
|||||
| Description | PMS1 protein homolog 2 (DNA mismatch repair protein PMS2). | |||||
|
TTC14_HUMAN
|
||||||
| θ value | 6.88961 (rank : 117) | NC score | 0.042089 (rank : 108) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q96N46, Q6UWJ7, Q8TF22 | Gene names | TTC14, KIAA1980 | |||
|
Domain Architecture |
|
|||||
| Description | Tetratricopeptide repeat protein 14 (TPR repeat protein 14). | |||||
|
ZNFX1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 118) | NC score | 0.035452 (rank : 119) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9P2E3, Q9BQM7, Q9BQM8, Q9H8C1, Q9H9S2, Q9NUM1, Q9NWW1 | Gene names | ZNFX1, KIAA1404 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
CA065_HUMAN
|
||||||
| θ value | 8.99809 (rank : 119) | NC score | 0.040600 (rank : 110) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 469 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8N715, Q8N746, Q8NA93 | Gene names | C1orf65 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C1orf65. | |||||
|
CSKI1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 120) | NC score | 0.008589 (rank : 170) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
DEK_HUMAN
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | 0.023342 (rank : 141) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P35659, Q5TGV4, Q5TGV5 | Gene names | DEK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
EEA1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 122) | NC score | 0.050515 (rank : 92) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1796 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q15075, Q14221 | Gene names | EEA1, ZFYVE2 | |||
|
Domain Architecture |

|
|||||
| Description | Early endosome antigen 1 (Endosome-associated protein p162) (Zinc finger FYVE domain-containing protein 2). | |||||
|
GCP60_MOUSE
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | 0.028173 (rank : 132) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8BMP6, O35371 | Gene names | Acbd3, Gcp60, Pap7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi resident protein GCP60 (Acyl-CoA-binding domain-containing protein 3) (Golgi phosphoprotein 1) (GOLPH1) (Golgi complex-associated protein 1) (GOCAP1) (PBR- and PKA-associated protein 7) (Peripheral benzodiazepine receptor-associated protein PAP7). | |||||
|
GOGA2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.054780 (rank : 67) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q921M4 | Gene names | Golga2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgin subfamily A member 2 (Cis-Golgi matrix protein GM130). | |||||
|
IQEC2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 125) | NC score | 0.017622 (rank : 152) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5DU25 | Gene names | Iqsec2, Kiaa0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
LDB3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 126) | NC score | 0.006673 (rank : 173) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 510 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9JKS4, Q6A038, Q811P2, Q811P3, Q811P4, Q811P5, Q9D130, Q9JKS3, Q9R0Z1, Q9WVH1, Q9WVH2 | Gene names | Ldb3, Kiaa0613 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LIM domain-binding protein 3 (Z-band alternatively spliced PDZ-motif protein) (Protein cypher) (Protein oracle). | |||||
|
MPP8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 127) | NC score | 0.053443 (rank : 73) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q99549, Q86TK3, Q9BTP1 | Gene names | MPHOSPH8, MPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | M-phase phosphoprotein 8. | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 128) | NC score | 0.084087 (rank : 13) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
PGCA_MOUSE
|
||||||
| θ value | 8.99809 (rank : 129) | NC score | 0.004354 (rank : 176) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 643 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q61282, Q64021 | Gene names | Agc1, Agc | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP). | |||||
|
PRP18_HUMAN
|
||||||
| θ value | 8.99809 (rank : 130) | NC score | 0.019192 (rank : 149) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99633, Q9BUI9 | Gene names | PRPF18, HPRP18 | |||
|
Domain Architecture |
|
|||||
| Description | Pre-mRNA-splicing factor 18 (PRP18 homolog) (hPRP18). | |||||
|
PRP18_MOUSE
|
||||||
| θ value | 8.99809 (rank : 131) | NC score | 0.020050 (rank : 146) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BM39, Q3UX46, Q8BZ54 | Gene names | Prpf18, Prp18 | |||
|
Domain Architecture |
|
|||||
| Description | Pre-mRNA-splicing factor 18 (PRP18 homolog). | |||||
|
RBBP6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 132) | NC score | 0.047548 (rank : 102) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
RU17_MOUSE
|
||||||
| θ value | 8.99809 (rank : 133) | NC score | 0.011171 (rank : 169) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q62376, Q3UIW4 | Gene names | Snrp70 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U1 small nuclear ribonucleoprotein 70 kDa (U1 SNRNP 70 kDa) (snRNP70). | |||||
|
SEC62_MOUSE
|
||||||
| θ value | 8.99809 (rank : 134) | NC score | 0.019047 (rank : 150) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BU14, Q6NX81 | Gene names | Tloc1, Sec62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Translocation protein SEC62 (Translocation protein 1) (TP-1). | |||||
|
SENP7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 135) | NC score | 0.016240 (rank : 156) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BQF6, Q96PS5, Q9C0F6, Q9HBT5 | Gene names | SENP7, KIAA1707, SSP2, SUSP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 7 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP7) (SUMO-1-specific protease 2). | |||||
|
SMC2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 136) | NC score | 0.050812 (rank : 91) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | O95347, Q9P1P2 | Gene names | SMC2L1, CAPE, SMC2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 2-like 1 protein (Chromosome- associated protein E) (hCAP-E) (XCAP-E homolog). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 8.99809 (rank : 137) | NC score | 0.052837 (rank : 77) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
UACA_MOUSE
|
||||||
| θ value | 8.99809 (rank : 138) | NC score | 0.049059 (rank : 98) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q8CGB3, Q69ZG3, Q7TN77, Q8BJC8 | Gene names | Uaca, Kiaa1561 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uveal autoantigen with coiled-coil domains and ankyrin repeats protein (Nucling) (Nuclear membrane-binding protein). | |||||
|
C102B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.053695 (rank : 71) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q68D86, Q7Z467, Q8NDK7, Q9H5C1 | Gene names | CCDC102B, C18orf14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 102B. | |||||
|
CCD39_MOUSE
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.054419 (rank : 68) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1026 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9D5Y1, Q8CDG6 | Gene names | Ccdc39, D3Ertd789e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 39. | |||||
|
CCD41_HUMAN
|
||||||
| θ value | θ > 10 (rank : 141) | NC score | 0.052196 (rank : 82) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1139 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9Y592 | Gene names | CCDC41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41 (NY-REN-58 antigen). | |||||
|
CCD46_HUMAN
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.050371 (rank : 96) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1430 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q8N8E3, Q6PIB5, Q8NCR4, Q8NFR4 | Gene names | CCDC46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 46. | |||||
|
CE152_HUMAN
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.052813 (rank : 78) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 976 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O94986, Q6NTA0 | Gene names | CEP152, KIAA0912 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 152 kDa (Cep152 protein). | |||||
|
CI039_HUMAN
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.051228 (rank : 90) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1171 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9NXG0, Q5VYJ0, Q9HAJ5 | Gene names | C9orf39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C9orf39. | |||||
|
CI093_HUMAN
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.051501 (rank : 89) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1088 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q6TFL3, Q5SU58, Q6P1W1, Q6ZR13, Q8N1Z4, Q8N8L3, Q8TBR2 | Gene names | C9orf93 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C9orf93. | |||||
|
DESP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 146) | NC score | 0.053096 (rank : 76) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P15924, O75993, Q14189, Q9UHN4 | Gene names | DSP | |||
|
Domain Architecture |

|
|||||
| Description | Desmoplakin (DP) (250/210 kDa paraneoplastic pemphigus antigen). | |||||
|
GOGA4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.050434 (rank : 94) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
GOGB1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.052091 (rank : 83) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
HOOK3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.050480 (rank : 93) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8BUK6, Q8BV48, Q8BW57, Q8BY41 | Gene names | Hook3 | |||
|
Domain Architecture |
|
|||||
| Description | Hook homolog 3. | |||||
|
K0555_HUMAN
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.050182 (rank : 97) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 880 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q96AA8, O60302 | Gene names | KIAA0555 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0555. | |||||
|
MY18A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.057633 (rank : 55) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1929 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q92614, Q8IXP8 | Gene names | MYO18A, KIAA0216, MYSPDZ | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
MY18A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.060329 (rank : 44) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1989 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q9JMH9, Q811D7 | Gene names | Myo18a, Myspdz | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
MY18B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.057764 (rank : 54) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1299 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8IUG5, Q8NDI8, Q8TE65, Q8WWS0, Q96KH2, Q96KR8, Q96KR9 | Gene names | MYO18B | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18B (Myosin XVIIIb). | |||||
|
MYH10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.060429 (rank : 43) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P35580, Q16087 | Gene names | MYH10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MYH10_MOUSE
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.060846 (rank : 42) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1612 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q61879, Q5SV63 | Gene names | Myh10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MYH14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.059563 (rank : 47) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 995 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q7Z406, Q5CZ75, Q6XYE4, Q76B62, Q8WV23, Q96I22, Q9BT27, Q9BW35, Q9H882 | Gene names | MYH14, KIAA2034 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-14 (Myosin heavy chain, nonmuscle IIc) (Nonmuscle myosin heavy chain IIc) (NMHC II-C). | |||||
|
MYH14_MOUSE
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.061137 (rank : 41) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1225 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q6URW6, Q80V64, Q80ZE6 | Gene names | Myh14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-14 (Myosin heavy chain, nonmuscle IIc) (Nonmuscle myosin heavy chain IIc) (NMHC II-C). | |||||
|
MYH3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.063953 (rank : 34) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1660 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P11055, Q15492 | Gene names | MYH3 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, fast skeletal muscle, embryonic (Muscle embryonic myosin heavy chain) (SMHCE). | |||||
|
MYH4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.062294 (rank : 37) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1377 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9Y623 | Gene names | MYH4 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-4 (Myosin heavy chain, skeletal muscle, fetal) (Myosin heavy chain IIb) (MyHC-IIb). | |||||
|
MYH4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.062219 (rank : 39) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1476 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q5SX39 | Gene names | Myh4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-4 (Myosin heavy chain, skeletal muscle, fetal). | |||||
|
MYH6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.062275 (rank : 38) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.063151 (rank : 35) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH9_HUMAN
|
||||||
| θ value | θ > 10 (rank : 163) | NC score | 0.063128 (rank : 36) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
MYO5A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 164) | NC score | 0.056922 (rank : 58) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1092 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9Y4I1, O60653, Q07902, Q16249, Q9UE30, Q9UE31 | Gene names | MYO5A, MYH12 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5A (Myosin Va) (Dilute myosin heavy chain, non-muscle) (Myosin heavy chain 12) (Myoxin). | |||||
|
MYO5A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 165) | NC score | 0.057542 (rank : 57) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1117 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q99104 | Gene names | Myo5a, Dilute | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5A (Myosin Va) (Dilute myosin heavy chain, non-muscle). | |||||
|
MYO5B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 166) | NC score | 0.056302 (rank : 60) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 911 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9ULV0 | Gene names | MYO5B, KIAA1119 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5B (Myosin Vb). | |||||
|
MYO5C_HUMAN
|
||||||
| θ value | θ > 10 (rank : 167) | NC score | 0.053950 (rank : 70) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1206 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9NQX4 | Gene names | MYO5C | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5C (Myosin Vc). | |||||
|
MYO6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 168) | NC score | 0.052751 (rank : 80) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 727 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9UM54, Q9BZZ7, Q9UEG2 | Gene names | MYO6, KIAA0389 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-6 (Myosin VI). | |||||
|
MYO6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 169) | NC score | 0.051858 (rank : 84) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q64331 | Gene names | Myo6, Sv | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-6 (Myosin VI). | |||||
|
NASP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 170) | NC score | 0.059918 (rank : 46) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P49321, Q96A69, Q9BTW2 | Gene names | NASP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigenic sperm protein (NASP). | |||||
|
NSBP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 171) | NC score | 0.059292 (rank : 49) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
OPTN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 172) | NC score | 0.050385 (rank : 95) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1120 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q8K3K8 | Gene names | Optn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Optineurin. | |||||
|
PIBF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 173) | NC score | 0.051833 (rank : 86) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8WXW3, O95664, Q6U9V2, Q6UG50, Q86V07, Q96SF4 | Gene names | PIBF1, C13orf24, PIBF | |||
|
Domain Architecture |

|
|||||
| Description | Progesterone-induced-blocking factor 1. | |||||
|
RAD50_HUMAN
|
||||||
| θ value | θ > 10 (rank : 174) | NC score | 0.053285 (rank : 75) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q92878, O43254, Q6GMT7, Q6P5X3, Q9UP86 | Gene names | RAD50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (hRAD50). | |||||
|
RAD50_MOUSE
|
||||||
| θ value | θ > 10 (rank : 175) | NC score | 0.052231 (rank : 81) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1366 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P70388, Q6PEB0, Q8C2T7, Q9CU59 | Gene names | Rad50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (mRad50). | |||||
|
RRBP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 176) | NC score | 0.053388 (rank : 74) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
SDCG8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 177) | NC score | 0.053672 (rank : 72) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1002 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q86SQ7, O60527, Q3ZCR6, Q8N5F2, Q9P0F1 | Gene names | SDCCAG8, CCCAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serologically defined colon cancer antigen 8 (Centrosomal colon cancer autoantigen protein) (hCCCAP) (Antigen NY-CO-8). | |||||
|
EST1A_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 138 | |
| SwissProt Accessions | P61406, Q5NC64 | Gene names | Est1a, Kiaa0732 | |||
|
Domain Architecture |

|
|||||
| Description | Telomerase-binding protein EST1A (Ever shorter telomeres 1A) (Telomerase subunit EST1A) (EST1-like protein A). | |||||
|
EST1A_HUMAN
|
||||||
| NC score | 0.966858 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q86US8, O94837, Q86VH6, Q9UF60 | Gene names | EST1A, C17orf31, KIAA0732 | |||
|
Domain Architecture |

|
|||||
| Description | Telomerase-binding protein EST1A (Ever shorter telomeres 1A) (Telomerase subunit EST1A) (EST1-like protein A) (hSmg5/7a). | |||||
|
SMG7_HUMAN
|
||||||
| NC score | 0.521121 (rank : 3) | θ value | 7.0802e-21 (rank : 3) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92540, Q5T1Q0, Q6PIE0, Q7Z7H9, Q8IXC1, Q8IXC2 | Gene names | SMG7, C1orf16, EST1C, KIAA0250 | |||
|
Domain Architecture |

|
|||||
| Description | Protein SMG7 (SMG-7 homolog) (EST1-like protein C). | |||||
|
SMG7_MOUSE
|
||||||
| NC score | 0.519478 (rank : 4) | θ value | 3.51386e-20 (rank : 4) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5RJH6, Q63ZW5, Q6ZQF3 | Gene names | Smg7, Est1c, Kiaa0250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein SMG7 (SMG-7 homolog) (EST1-like protein C). | |||||
|
SMG5_MOUSE
|
||||||
| NC score | 0.450613 (rank : 5) | θ value | 8.38298e-14 (rank : 5) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6ZPY2, Q497S2, Q8R3S4 | Gene names | Smg5, Est1b, Kiaa1089 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein SMG5 (SMG-5 homolog) (EST1-like protein B). | |||||
|
SMG5_HUMAN
|
||||||
| NC score | 0.425382 (rank : 6) | θ value | 1.2105e-12 (rank : 6) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UPR3, Q5QJE7, Q8IXC0, Q8IY09, Q96IJ7 | Gene names | SMG5, EST1B, KIAA1089 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein SMG5 (SMG-5 homolog) (EST1-like protein B) (LPTS-interacting protein) (LPTS-RP1). | |||||
|
CA026_MOUSE
|
||||||
| NC score | 0.335907 (rank : 7) | θ value | 5.62301e-10 (rank : 7) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9DBQ9, Q6PAN2, Q7TMQ6 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C1orf26 homolog. | |||||
|
CA026_HUMAN
|
||||||
| NC score | 0.310077 (rank : 8) | θ value | 6.87365e-08 (rank : 8) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5T5J6, Q8NEK9, Q9BZQ7, Q9NXQ0 | Gene names | C1orf26 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C1orf26. | |||||
|
SPT5H_HUMAN
|
||||||
| NC score | 0.141586 (rank : 9) | θ value | 0.00175202 (rank : 10) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O00267, O43279, Q59G52, Q99639 | Gene names | SUPT5H, SPT5, SPT5H | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT5 (hSPT5) (DRB sensitivity-inducing factor large subunit) (DSIF large subunit) (DSIF p160) (Tat- cotransactivator 1 protein) (Tat-CT1 protein). | |||||
|
SPT5H_MOUSE
|
||||||
| NC score | 0.139204 (rank : 10) | θ value | 0.00175202 (rank : 11) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 437 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O55201, Q3UJ77, Q3UJH1, Q3UKD7, Q3UM54, Q6PB73, Q6PDP0, Q6PFR4 | Gene names | Supt5h | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT5 (DRB sensitivity-inducing factor large subunit) (DSIF large subunit). | |||||
|
REN3B_HUMAN
|
||||||
| NC score | 0.105090 (rank : 11) | θ value | 0.00228821 (rank : 12) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9BZI7, Q9H1J0 | Gene names | UPF3B, RENT3B, UPF3X | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of nonsense transcripts 3B (Nonsense mRNA reducing factor 3B) (Up-frameshift suppressor 3 homolog B) (hUpf3B) (hUpf3p-X). | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.091053 (rank : 12) | θ value | 8.40245e-06 (rank : 9) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.084087 (rank : 13) | θ value | 8.99809 (rank : 128) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
KLC1_HUMAN
|
||||||
| NC score | 0.074828 (rank : 14) | θ value | 0.0961366 (rank : 18) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q07866, Q7RTM2, Q7RTM3, Q7RTM5, Q7RTP9, Q7RTQ5, Q86SF5, Q86TF5, Q86V74, Q86V75, Q86V76, Q86V77, Q86V78, Q86V79, Q96H62 | Gene names | KNS2, KLC, KLC1 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin light chain 1 (KLC 1). | |||||
|
CAR11_HUMAN
|
||||||
| NC score | 0.073743 (rank : 15) | θ value | 0.125558 (rank : 20) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9BXL7, Q548H3 | Gene names | CARD11, CARMA1 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 11 (CARD-containing MAGUK protein 3) (Carma 1). | |||||
|
THOC2_HUMAN
|
||||||
| NC score | 0.072273 (rank : 16) | θ value | 3.0926 (rank : 84) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8NI27, Q9H8I6 | Gene names | THOC2, CXorf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | THO complex subunit 2 (Tho2). | |||||
|
CALD1_HUMAN
|
||||||
| NC score | 0.071196 (rank : 17) | θ value | 0.21417 (rank : 26) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q05682, Q13978, Q13979, Q14741, Q14742 | Gene names | CALD1, CAD, CDM | |||
|
Domain Architecture |

|
|||||
| Description | Caldesmon (CDM). | |||||
|
ZNFX1_MOUSE
|
||||||
| NC score | 0.071100 (rank : 18) | θ value | 0.0563607 (rank : 16) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 407 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8R151, Q3U016, Q3UD71, Q3UEZ0 | Gene names | ZNFX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
CP135_HUMAN
|
||||||
| NC score | 0.070870 (rank : 19) | θ value | 0.365318 (rank : 32) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1290 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q66GS9, O75130, Q9H8H7 | Gene names | CEP135, CEP4, KIAA0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
MYH11_HUMAN
|
||||||
| NC score | 0.070582 (rank : 20) | θ value | 0.00390308 (rank : 13) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
MYH9_MOUSE
|
||||||
| NC score | 0.069824 (rank : 21) | θ value | 0.47712 (rank : 38) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
MYH13_HUMAN
|
||||||
| NC score | 0.068560 (rank : 22) | θ value | 0.125558 (rank : 22) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
CING_MOUSE
|
||||||
| NC score | 0.068312 (rank : 23) | θ value | 0.47712 (rank : 35) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1432 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | P59242 | Gene names | Cgn | |||
|
Domain Architecture |

|
|||||
| Description | Cingulin. | |||||
|
MYH8_MOUSE
|
||||||
| NC score | 0.067876 (rank : 24) | θ value | 0.0330416 (rank : 15) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1600 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P13542, Q5SX36 | Gene names | Myh8, Myhsp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
MYH11_MOUSE
|
||||||
| NC score | 0.067840 (rank : 25) | θ value | 0.125558 (rank : 21) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
MYH1_HUMAN
|
||||||
| NC score | 0.067735 (rank : 26) | θ value | 0.163984 (rank : 25) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1478 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | P12882, Q9Y622 | Gene names | MYH1 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-1 (Myosin heavy chain, skeletal muscle, adult 1) (Myosin heavy chain IIx/d) (MyHC-IIx/d). | |||||
|
RBM25_HUMAN
|
||||||
| NC score | 0.067363 (rank : 27) | θ value | 1.06291 (rank : 52) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P49756, Q9UEQ5, Q9UIE9 | Gene names | RBM25, RNPC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable RNA-binding protein 25 (RNA-binding motif protein 25) (RNA- binding region-containing protein 7) (Protein S164). | |||||
|
CP135_MOUSE
|
||||||
| NC score | 0.067335 (rank : 28) | θ value | 1.38821 (rank : 57) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1278 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q6P5D4, Q6A030 | Gene names | Cep135, Cep4, Kiaa0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
MYH8_HUMAN
|
||||||
| NC score | 0.067313 (rank : 29) | θ value | 0.0961366 (rank : 19) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1610 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | P13535, Q14910 | Gene names | MYH8 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
MYH2_HUMAN
|
||||||
| NC score | 0.066916 (rank : 30) | θ value | 0.279714 (rank : 29) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1491 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9UKX2, Q14322, Q16229 | Gene names | MYH2, MYHSA2 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, skeletal muscle, adult 2 (Myosin heavy chain IIa) (MyHC-IIa). | |||||
|
MYH7_HUMAN
|
||||||
| NC score | 0.066456 (rank : 31) | θ value | 1.38821 (rank : 60) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
CING_HUMAN
|
||||||
| NC score | 0.065231 (rank : 32) | θ value | 2.36792 (rank : 70) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1275 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9P2M7, Q5T386, Q9NR25 | Gene names | CGN, KIAA1319 | |||
|
Domain Architecture |

|
|||||
| Description | Cingulin. | |||||
|
MYH1_MOUSE
|
||||||
| NC score | 0.064402 (rank : 33) | θ value | 3.0926 (rank : 79) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1461 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q5SX40 | Gene names | Myh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-1 (Myosin heavy chain, skeletal muscle, adult 1). | |||||
|
MYH3_HUMAN
|
||||||
| NC score | 0.063953 (rank : 34) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1660 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P11055, Q15492 | Gene names | MYH3 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, fast skeletal muscle, embryonic (Muscle embryonic myosin heavy chain) (SMHCE). | |||||
|
MYH6_MOUSE
|
||||||
| NC score | 0.063151 (rank : 35) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH9_HUMAN
|
||||||
| NC score | 0.063128 (rank : 36) | θ value | θ > 10 (rank : 163) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
MYH4_HUMAN
|
||||||
| NC score | 0.062294 (rank : 37) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1377 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9Y623 | Gene names | MYH4 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-4 (Myosin heavy chain, skeletal muscle, fetal) (Myosin heavy chain IIb) (MyHC-IIb). | |||||
|
MYH6_HUMAN
|
||||||
| NC score | 0.062275 (rank : 38) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH4_MOUSE
|
||||||
| NC score | 0.062219 (rank : 39) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1476 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q5SX39 | Gene names | Myh4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-4 (Myosin heavy chain, skeletal muscle, fetal). | |||||
|
KLC1_MOUSE
|
||||||
| NC score | 0.061288 (rank : 40) | θ value | 0.813845 (rank : 44) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O88447 | Gene names | Kns2, Klc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin light chain 1 (KLC 1). | |||||
|
MYH14_MOUSE
|
||||||
| NC score | 0.061137 (rank : 41) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1225 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q6URW6, Q80V64, Q80ZE6 | Gene names | Myh14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-14 (Myosin heavy chain, nonmuscle IIc) (Nonmuscle myosin heavy chain IIc) (NMHC II-C). | |||||
|
MYH10_MOUSE
|
||||||
| NC score | 0.060846 (rank : 42) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1612 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q61879, Q5SV63 | Gene names | Myh10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MYH10_HUMAN
|
||||||
| NC score | 0.060429 (rank : 43) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P35580, Q16087 | Gene names | MYH10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MY18A_MOUSE
|
||||||
| NC score | 0.060329 (rank : 44) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1989 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q9JMH9, Q811D7 | Gene names | Myo18a, Myspdz | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
SDCG8_MOUSE
|
||||||
| NC score | 0.060273 (rank : 45) | θ value | 1.38821 (rank : 62) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q80UF4 | Gene names | Sdccag8, Cccap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serologically defined colon cancer antigen 8 (Centrosomal colon cancer autoantigen protein) (mCCCAP). | |||||
|
NASP_HUMAN
|
||||||
| NC score | 0.059918 (rank : 46) | θ value | θ > 10 (rank : 170) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P49321, Q96A69, Q9BTW2 | Gene names | NASP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigenic sperm protein (NASP). | |||||
|
MYH14_HUMAN
|
||||||
| NC score | 0.059563 (rank : 47) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 995 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q7Z406, Q5CZ75, Q6XYE4, Q76B62, Q8WV23, Q96I22, Q9BT27, Q9BW35, Q9H882 | Gene names | MYH14, KIAA2034 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-14 (Myosin heavy chain, nonmuscle IIc) (Nonmuscle myosin heavy chain IIc) (NMHC II-C). | |||||
|
LUZP1_MOUSE
|
||||||
| NC score | 0.059506 (rank : 48) | θ value | 0.813845 (rank : 45) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 940 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q8R4U7, Q3UV18, Q7TS71, Q8BQW1, Q99NG3, Q9CSL6 | Gene names | Luzp1, Luzp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper protein 1 (Leucine zipper motif-containing protein). | |||||
|
NSBP1_HUMAN
|
||||||
| NC score | 0.059292 (rank : 49) | θ value | θ > 10 (rank : 171) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
BPAEA_MOUSE
|
||||||
| NC score | 0.058180 (rank : 50) | θ value | 0.62314 (rank : 39) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1480 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q91ZU8, Q8K5D4 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoform 5 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
AKAP9_HUMAN
|
||||||
| NC score | 0.058098 (rank : 51) | θ value | 1.06291 (rank : 49) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1710 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q99996, O14869, O43355, O94895, Q9UQH3, Q9UQQ4, Q9Y6B8, Q9Y6Y2 | Gene names | AKAP9, AKAP350, AKAP450, KIAA0803 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 9 (Protein kinase A-anchoring protein 9) (PRKA9) (A-kinase anchor protein 450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP 350) (hgAKAP 350) (AKAP 120-like protein) (Protein hyperion) (Protein yotiao) (Centrosome- and Golgi-localized PKN-associated protein) (CG-NAP). | |||||
|
CDC37_HUMAN
|
||||||
| NC score | 0.057974 (rank : 52) | θ value | 1.38821 (rank : 56) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q16543 | Gene names | CDC37 | |||
|
Domain Architecture |

|
|||||
| Description | Hsp90 co-chaperone Cdc37 (Hsp90 chaperone protein kinase-targeting subunit) (p50Cdc37). | |||||
|
ATRX_HUMAN
|
||||||
| NC score | 0.057969 (rank : 53) | θ value | 0.279714 (rank : 27) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P46100, P51068, Q15886, Q59FB5, Q59H31, Q5H9A2, Q5JWI4, Q7Z2J1, Q9H0Z1, Q9NTS3 | Gene names | ATRX, RAD54L, XH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked helicase II) (X-linked nuclear protein) (XNP) (Znf- HX). | |||||
|
MY18B_HUMAN
|
||||||
| NC score | 0.057764 (rank : 54) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1299 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8IUG5, Q8NDI8, Q8TE65, Q8WWS0, Q96KH2, Q96KR8, Q96KR9 | Gene names | MYO18B | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18B (Myosin XVIIIb). | |||||
|
MY18A_HUMAN
|
||||||
| NC score | 0.057633 (rank : 55) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1929 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q92614, Q8IXP8 | Gene names | MYO18A, KIAA0216, MYSPDZ | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
TRHY_HUMAN
|
||||||
| NC score | 0.057595 (rank : 56) | θ value | 1.38821 (rank : 63) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
MYO5A_MOUSE
|
||||||
| NC score | 0.057542 (rank : 57) | θ value | θ > 10 (rank : 165) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1117 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q99104 | Gene names | Myo5a, Dilute | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5A (Myosin Va) (Dilute myosin heavy chain, non-muscle). | |||||
|
MYO5A_HUMAN
|
||||||
| NC score | 0.056922 (rank : 58) | θ value | θ > 10 (rank : 164) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1092 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9Y4I1, O60653, Q07902, Q16249, Q9UE30, Q9UE31 | Gene names | MYO5A, MYH12 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5A (Myosin Va) (Dilute myosin heavy chain, non-muscle) (Myosin heavy chain 12) (Myoxin). | |||||
|
CCD41_MOUSE
|
||||||
| NC score | 0.056920 (rank : 59) | θ value | 1.81305 (rank : 64) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9D5R3, Q3U7X7, Q3UX57, Q80VF0 | Gene names | Ccdc41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41. | |||||
|
MYO5B_HUMAN
|
||||||
| NC score | 0.056302 (rank : 60) | θ value | θ > 10 (rank : 166) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 911 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9ULV0 | Gene names | MYO5B, KIAA1119 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5B (Myosin Vb). | |||||
|
MORC4_HUMAN
|
||||||
| NC score | 0.056070 (rank : 61) | θ value | 0.279714 (rank : 28) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8TE76, Q5JUK7, Q96MZ2, Q9HAI7 | Gene names | MORC4, ZCWCC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 4 (Zinc finger CW-type coiled- coil domain protein 2). | |||||
|
UBF1_HUMAN
|
||||||
| NC score | 0.056057 (rank : 62) | θ value | 0.813845 (rank : 47) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P17480 | Gene names | UBTF, UBF, UBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1) (Autoantigen NOR-90). | |||||
|
UBF1_MOUSE
|
||||||
| NC score | 0.055718 (rank : 63) | θ value | 0.813845 (rank : 48) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P25976 | Gene names | Ubtf, Tcfubf, Ubf-1, Ubf1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1). | |||||
|
CRSP2_HUMAN
|
||||||
| NC score | 0.055636 (rank : 64) | θ value | 1.06291 (rank : 50) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O60244, Q9UNB3 | Gene names | CRSP2, ARC150, DRIP150, EXLM1, TRAP170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CRSP complex subunit 2 (Cofactor required for Sp1 transcriptional activation subunit 2) (Transcriptional coactivator CRSP150) (Vitamin D3 receptor-interacting protein complex 150 kDa component) (DRIP150) (Thyroid hormone receptor-associated protein complex 170 kDa component) (Trap170) (Activator-recruited cofactor 150 kDa component) (ARC150). | |||||
|
MDN1_HUMAN
|
||||||
| NC score | 0.054918 (rank : 65) | θ value | 0.62314 (rank : 42) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9NU22, O15019 | Gene names | MDN1, KIAA0301 | |||
|
Domain Architecture |

|
|||||
| Description | Midasin (MIDAS-containing protein). | |||||
|
HTSF1_MOUSE
|
||||||
| NC score | 0.054781 (rank : 66) | θ value | 0.365318 (rank : 33) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BGC0, Q1WWK0, Q9CT41, Q9DAU3 | Gene names | Htatsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 homolog. | |||||
|
GOGA2_MOUSE
|
||||||
| NC score | 0.054780 (rank : 67) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q921M4 | Gene names | Golga2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgin subfamily A member 2 (Cis-Golgi matrix protein GM130). | |||||
|
CCD39_MOUSE
|
||||||
| NC score | 0.054419 (rank : 68) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1026 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9D5Y1, Q8CDG6 | Gene names | Ccdc39, D3Ertd789e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 39. | |||||
|
GOGA4_HUMAN
|
||||||
| NC score | 0.054352 (rank : 69) | θ value | 3.0926 (rank : 77) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
MYO5C_HUMAN
|
||||||
| NC score | 0.053950 (rank : 70) | θ value | θ > 10 (rank : 167) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1206 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9NQX4 | Gene names | MYO5C | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5C (Myosin Vc). | |||||
|
C102B_HUMAN
|
||||||
| NC score | 0.053695 (rank : 71) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q68D86, Q7Z467, Q8NDK7, Q9H5C1 | Gene names | CCDC102B, C18orf14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 102B. | |||||
|
SDCG8_HUMAN
|
||||||
| NC score | 0.053672 (rank : 72) | θ value | θ > 10 (rank : 177) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1002 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q86SQ7, O60527, Q3ZCR6, Q8N5F2, Q9P0F1 | Gene names | SDCCAG8, CCCAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serologically defined colon cancer antigen 8 (Centrosomal colon cancer autoantigen protein) (hCCCAP) (Antigen NY-CO-8). | |||||
|
MPP8_HUMAN
|
||||||
| NC score | 0.053443 (rank : 73) | θ value | 8.99809 (rank : 127) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q99549, Q86TK3, Q9BTP1 | Gene names | MPHOSPH8, MPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | M-phase phosphoprotein 8. | |||||
|
RRBP1_MOUSE
|
||||||
| NC score | 0.053388 (rank : 74) | θ value | θ > 10 (rank : 176) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
RAD50_HUMAN
|
||||||
| NC score | 0.053285 (rank : 75) | θ value | θ > 10 (rank : 174) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q92878, O43254, Q6GMT7, Q6P5X3, Q9UP86 | Gene names | RAD50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (hRAD50). | |||||
|
DESP_HUMAN
|
||||||
| NC score | 0.053096 (rank : 76) | θ value | θ > 10 (rank : 146) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P15924, O75993, Q14189, Q9UHN4 | Gene names | DSP | |||
|
Domain Architecture |

|
|||||
| Description | Desmoplakin (DP) (250/210 kDa paraneoplastic pemphigus antigen). | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.052837 (rank : 77) | θ value | 8.99809 (rank : 137) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
CE152_HUMAN
|
||||||
| NC score | 0.052813 (rank : 78) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 976 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O94986, Q6NTA0 | Gene names | CEP152, KIAA0912 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 152 kDa (Cep152 protein). | |||||
|
CENPF_HUMAN
|
||||||
| NC score | 0.052777 (rank : 79) | θ value | 6.88961 (rank : 110) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
MYO6_HUMAN
|
||||||
| NC score | 0.052751 (rank : 80) | θ value | θ > 10 (rank : 168) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 727 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9UM54, Q9BZZ7, Q9UEG2 | Gene names | MYO6, KIAA0389 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-6 (Myosin VI). | |||||
|
RAD50_MOUSE
|
||||||
| NC score | 0.052231 (rank : 81) | θ value | θ > 10 (rank : 175) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1366 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P70388, Q6PEB0, Q8C2T7, Q9CU59 | Gene names | Rad50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (mRad50). | |||||
|
CCD41_HUMAN
|
||||||
| NC score | 0.052196 (rank : 82) | θ value | θ > 10 (rank : 141) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1139 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9Y592 | Gene names | CCDC41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41 (NY-REN-58 antigen). | |||||
|
GOGB1_HUMAN
|
||||||
| NC score | 0.052091 (rank : 83) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
MYO6_MOUSE
|
||||||
| NC score | 0.051858 (rank : 84) | θ value | θ > 10 (rank : 169) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q64331 | Gene names | Myo6, Sv | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-6 (Myosin VI). | |||||
|
TS101_HUMAN
|
||||||
| NC score | 0.051857 (rank : 85) | θ value | 1.81305 (rank : 67) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q99816, Q9BUM5 | Gene names | TSG101 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor susceptibility gene 101 protein. | |||||
|
PIBF1_HUMAN
|
||||||
| NC score | 0.051833 (rank : 86) | θ value | θ > 10 (rank : 173) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8WXW3, O95664, Q6U9V2, Q6UG50, Q86V07, Q96SF4 | Gene names | PIBF1, C13orf24, PIBF | |||
|
Domain Architecture |

|
|||||
| Description | Progesterone-induced-blocking factor 1. | |||||
|
TOP1_HUMAN
|
||||||
| NC score | 0.051677 (rank : 87) | θ value | 0.279714 (rank : 30) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P11387, O43256, Q12855, Q12856, Q15610, Q5TFY3, Q9UJN0 | Gene names | TOP1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 1 (EC 5.99.1.2) (DNA topoisomerase I). | |||||
|
PCNT_HUMAN
|
||||||
| NC score | 0.051625 (rank : 88) | θ value | 5.27518 (rank : 105) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1551 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | O95613, O43152, Q7Z7C9 | Gene names | PCNT, KIAA0402, PCNT2 | |||
|
Domain Architecture |

|
|||||
| Description | Pericentrin (Pericentrin B) (Kendrin). | |||||
|
CI093_HUMAN
|
||||||
| NC score | 0.051501 (rank : 89) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1088 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q6TFL3, Q5SU58, Q6P1W1, Q6ZR13, Q8N1Z4, Q8N8L3, Q8TBR2 | Gene names | C9orf93 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C9orf93. | |||||
|
CI039_HUMAN
|
||||||
| NC score | 0.051228 (rank : 90) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1171 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9NXG0, Q5VYJ0, Q9HAJ5 | Gene names | C9orf39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C9orf39. | |||||
|
SMC2_HUMAN
|
||||||
| NC score | 0.050812 (rank : 91) | θ value | 8.99809 (rank : 136) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | O95347, Q9P1P2 | Gene names | SMC2L1, CAPE, SMC2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 2-like 1 protein (Chromosome- associated protein E) (hCAP-E) (XCAP-E homolog). | |||||
|
EEA1_HUMAN
|
||||||
| NC score | 0.050515 (rank : 92) | θ value | 8.99809 (rank : 122) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1796 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q15075, Q14221 | Gene names | EEA1, ZFYVE2 | |||
|
Domain Architecture |

|
|||||
| Description | Early endosome antigen 1 (Endosome-associated protein p162) (Zinc finger FYVE domain-containing protein 2). | |||||
|
HOOK3_MOUSE
|
||||||
| NC score | 0.050480 (rank : 93) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8BUK6, Q8BV48, Q8BW57, Q8BY41 | Gene names | Hook3 | |||
|
Domain Architecture |

|
|||||
| Description | Hook homolog 3. | |||||
|
GOGA4_MOUSE
|
||||||
| NC score | 0.050434 (rank : 94) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
OPTN_MOUSE
|
||||||
| NC score | 0.050385 (rank : 95) | θ value | θ > 10 (rank : 172) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1120 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q8K3K8 | Gene names | Optn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Optineurin. | |||||
|
CCD46_HUMAN
|
||||||
| NC score | 0.050371 (rank : 96) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1430 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q8N8E3, Q6PIB5, Q8NCR4, Q8NFR4 | Gene names | CCDC46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 46. | |||||
|
K0555_HUMAN
|
||||||
| NC score | 0.050182 (rank : 97) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 880 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q96AA8, O60302 | Gene names | KIAA0555 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0555. | |||||
|
UACA_MOUSE
|
||||||
| NC score | 0.049059 (rank : 98) | θ value | 8.99809 (rank : 138) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q8CGB3, Q69ZG3, Q7TN77, Q8BJC8 | Gene names | Uaca, Kiaa1561 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uveal autoantigen with coiled-coil domains and ankyrin repeats protein (Nucling) (Nuclear membrane-binding protein). | |||||
|
MICA1_MOUSE
|
||||||
| NC score | 0.048759 (rank : 99) | θ value | 0.47712 (rank : 37) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8VDP3 | Gene names | Mical1, Mical, Nical | |||
|
Domain Architecture |

|
|||||
| Description | NEDD9-interacting protein with calponin homology and LIM domains (Molecule interacting with CasL protein 1). | |||||
|
PLEC1_HUMAN
|
||||||
| NC score | 0.048164 (rank : 100) | θ value | 1.38821 (rank : 61) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1326 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q15149, Q15148, Q16640 | Gene names | PLEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Plectin-1 (PLTN) (PCN) (Hemidesmosomal protein 1) (HD1). | |||||
|
SETD2_HUMAN
|
||||||
| NC score | 0.047678 (rank : 101) | θ value | 1.06291 (rank : 53) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
RBBP6_HUMAN
|
||||||
| NC score | 0.047548 (rank : 102) | θ value | 8.99809 (rank : 132) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
CUTL1_MOUSE
|
||||||
| NC score | 0.046953 (rank : 103) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
WDR60_MOUSE
|
||||||
| NC score | 0.044424 (rank : 104) | θ value | 3.0926 (rank : 85) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8C761, Q3UNM4 | Gene names | Wdr60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 60. | |||||
|
BCN1_MOUSE
|
||||||
| NC score | 0.044281 (rank : 105) | θ value | 5.27518 (rank : 96) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 367 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O88597 | Gene names | Becn1 | |||
|
Domain Architecture |

|
|||||
| Description | Beclin-1 (Coiled-coil myosin-like BCL2-interacting protein). | |||||
|
KLC2_HUMAN
|
||||||
| NC score | 0.043652 (rank : 106) | θ value | 5.27518 (rank : 101) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9H0B6, Q9H9C8, Q9HA20 | Gene names | KLC2 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin light chain 2 (KLC 2). | |||||
|
PHLB3_HUMAN
|
||||||
| NC score | 0.043093 (rank : 107) | θ value | 6.88961 (rank : 115) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 304 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6NSJ2, Q8N7Z4 | Gene names | PHLDB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 3. | |||||
|
TTC14_HUMAN
|
||||||
| NC score | 0.042089 (rank : 108) | θ value | 6.88961 (rank : 117) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q96N46, Q6UWJ7, Q8TF22 | Gene names | TTC14, KIAA1980 | |||
|
Domain Architecture |

|
|||||
| Description | Tetratricopeptide repeat protein 14 (TPR repeat protein 14). | |||||
|
ACINU_MOUSE
|
||||||
| NC score | 0.041789 (rank : 109) | θ value | 5.27518 (rank : 95) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9JIX8, Q9CSN7, Q9CSR9, Q9CSX7, Q9R046, Q9R047 | Gene names | Acin1, Acinus | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
CA065_HUMAN
|
||||||
| NC score | 0.040600 (rank : 110) | θ value | 8.99809 (rank : 119) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 469 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8N715, Q8N746, Q8NA93 | Gene names | C1orf65 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C1orf65. | |||||
|
LIPA3_HUMAN
|
||||||
| NC score | 0.039894 (rank : 111) | θ value | 4.03905 (rank : 91) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O75145, Q9H8B5, Q9UEW4 | Gene names | PPFIA3, KIAA0654 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-3 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-3) (PTPRF-interacting protein alpha-3). | |||||
|
IF4G1_MOUSE
|
||||||
| NC score | 0.039677 (rank : 112) | θ value | 0.62314 (rank : 41) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6NZJ6, Q6NZN8, Q8BW99 | Gene names | Eif4g1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1). | |||||
|
CIR_HUMAN
|
||||||
| NC score | 0.038546 (rank : 113) | θ value | 5.27518 (rank : 97) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q86X95, O95367, Q12804, Q4G1B9, Q6PJI4, Q8IWI2 | Gene names | CIR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor (Recepin). | |||||
|
ROCK2_MOUSE
|
||||||
| NC score | 0.036606 (rank : 114) | θ value | 0.0148317 (rank : 14) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 2072 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P70336, Q8BR64, Q8CBR0 | Gene names | Rock2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2). | |||||
|
KI67_HUMAN
|
||||||
| NC score | 0.036518 (rank : 115) | θ value | 3.0926 (rank : 78) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
EHMT2_HUMAN
|
||||||
| NC score | 0.035954 (rank : 116) | θ value | 0.163984 (rank : 23) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q96KQ7, Q14349, Q6PK06, Q96MH5, Q96QD0, Q9UQL8, Q9Y331 | Gene names | EHMT2, BAT8, G9A, NG36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
EDF1_MOUSE
|
||||||
| NC score | 0.035757 (rank : 117) | θ value | 4.03905 (rank : 88) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JMG1, Q99NE6 | Gene names | Edf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelial differentiation-related factor 1 (EDF-1) (Multiprotein- bridging factor 1) (MBF1). | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.035469 (rank : 118) | θ value | 1.38821 (rank : 55) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
ZNFX1_HUMAN
|
||||||
| NC score | 0.035452 (rank : 119) | θ value | 6.88961 (rank : 118) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9P2E3, Q9BQM7, Q9BQM8, Q9H8C1, Q9H9S2, Q9NUM1, Q9NWW1 | Gene names | ZNFX1, KIAA1404 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
EHMT2_MOUSE
|
||||||
| NC score | 0.035439 (rank : 120) | θ value | 0.163984 (rank : 24) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Z148, Q6PE08, Q8K4R6, Q8K4R7, Q9Z149 | Gene names | Ehmt2, Bat8, G9a, Ng36 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 3) (H3-K9-HMTase 3) (Euchromatic histone-lysine N-methyltransferase 2) (HLA-B-associated transcript 8) (Protein G9a). | |||||
|
CHD1_HUMAN
|
||||||
| NC score | 0.035413 (rank : 121) | θ value | 0.365318 (rank : 31) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
ESCO1_HUMAN
|
||||||
| NC score | 0.035403 (rank : 122) | θ value | 5.27518 (rank : 100) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q5FWF5, Q69YG4, Q69YS3, Q6IMD7, Q8N3Z5, Q8NBG2, Q96PX7 | Gene names | ESCO1, EFO1, KIAA1911 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyltransferase ESCO1 (EC 2.3.1.-) (Establishment of cohesion 1 homolog 1) (ECO1 homolog 1) (ESO1 homolog 1) (Establishment factor- like protein 1) (EFO1p) (hEFO1) (CTF7 homolog 1). | |||||
|
NFM_MOUSE
|
||||||
| NC score | 0.035274 (rank : 123) | θ value | 2.36792 (rank : 72) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P08553, Q61961 | Gene names | Nef3, Nefm, Nfm | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M). | |||||
|
IG2AS_HUMAN
|
||||||
| NC score | 0.035201 (rank : 124) | θ value | 6.88961 (rank : 113) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6U949, Q7Z712, Q9P2W0 | Gene names | IGF2AS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative insulin-like growth factor 2 antisense gene protein (IGF2-AS) (PEG8/IGF2AS protein). | |||||
|
SMCA2_HUMAN
|
||||||
| NC score | 0.034381 (rank : 125) | θ value | 3.0926 (rank : 82) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P51531 | Gene names | SMARCA2, BRM, SNF2A, SNF2L2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L2 (EC 3.6.1.-) (ATP- dependent helicase SMARCA2) (SNF2-alpha) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin subfamily A member 2) (hBRM). | |||||
|
SH24A_MOUSE
|
||||||
| NC score | 0.034328 (rank : 126) | θ value | 3.0926 (rank : 81) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9D7V1 | Gene names | Sh2d4a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH2 domain-containing protein 4A. | |||||
|
IF4G1_HUMAN
|
||||||
| NC score | 0.033411 (rank : 127) | θ value | 1.38821 (rank : 59) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q04637, O43177, O95066, Q5HYG0, Q6ZN21, Q8N102 | Gene names | EIF4G1, EIF4G, EIF4GI | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1) (p220). | |||||
|
EDF1_HUMAN
|
||||||
| NC score | 0.033332 (rank : 128) | θ value | 5.27518 (rank : 99) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60869, Q5T5T2, Q9UIM1 | Gene names | EDF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelial differentiation-related factor 1 (EDF-1) (Multiprotein- bridging factor 1) (MBF1). | |||||
|
CHD1_MOUSE
|
||||||
| NC score | 0.032704 (rank : 129) | θ value | 0.62314 (rank : 40) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
SOX9_HUMAN
|
||||||
| NC score | 0.030877 (rank : 130) | θ value | 1.06291 (rank : 54) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P48436 | Gene names | SOX9 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-9. | |||||
|
SPRL1_HUMAN
|
||||||
| NC score | 0.028363 (rank : 131) | θ value | 3.0926 (rank : 83) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q14515, Q14800 | Gene names | SPARCL1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (High endothelial venule protein) (Hevin) (MAST 9). | |||||
|
GCP60_MOUSE
|
||||||
| NC score | 0.028173 (rank : 132) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8BMP6, O35371 | Gene names | Acbd3, Gcp60, Pap7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi resident protein GCP60 (Acyl-CoA-binding domain-containing protein 3) (Golgi phosphoprotein 1) (GOLPH1) (Golgi complex-associated protein 1) (GOCAP1) (PBR- and PKA-associated protein 7) (Peripheral benzodiazepine receptor-associated protein PAP7). | |||||
|
ZCHC8_MOUSE
|
||||||
| NC score | 0.027618 (rank : 133) | θ value | 4.03905 (rank : 94) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9CYA6, Q3TK71, Q8CCB7, Q91WQ1 | Gene names | Zcchc8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 8. | |||||
|
BRWD1_HUMAN
|
||||||
| NC score | 0.027161 (rank : 134) | θ value | 0.47712 (rank : 34) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NSI6, O43721, Q5R2V0, Q5R2V1, Q8TCV3, Q96QG9, Q96QH0, Q9NUK1 | Gene names | BRWD1, C21orf107, WDR9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain and WD repeat domain-containing protein 1 (WD repeat protein 9). | |||||
|
PHF20_HUMAN
|
||||||
| NC score | 0.026768 (rank : 135) | θ value | 5.27518 (rank : 106) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9BVI0, Q5JWY9, Q9BWV4, Q9BXA3, Q9BZW3, Q9H421, Q9H4J6, Q9NZ22 | Gene names | PHF20, C20orf104, GLEA2, HCA58, NZF, TZP | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58) (Glioma-expressed antigen 2) (Transcription factor TZP) (Novel zinc finger protein). | |||||
|
E41L2_HUMAN
|
||||||
| NC score | 0.026436 (rank : 136) | θ value | 0.813845 (rank : 43) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O43491 | Gene names | EPB41L2 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 2 (Generally expressed protein 4.1) (4.1G). | |||||
|
PF21B_HUMAN
|
||||||
| NC score | 0.025845 (rank : 137) | θ value | 1.06291 (rank : 51) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96EK2, Q5TFL2, Q6ICC0 | Gene names | PHF21B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
MBP_MOUSE
|
||||||
| NC score | 0.024629 (rank : 138) | θ value | 4.03905 (rank : 92) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P04370, Q01585, Q03139, Q03176, Q61836, Q61837, Q99KE4, Q9QWP1 | Gene names | Mbp, Shi | |||
|
Domain Architecture |

|
|||||
| Description | Myelin basic protein (MBP) (Myelin A1 protein). | |||||
|
NO40_MOUSE
|
||||||
| NC score | 0.024296 (rank : 139) | θ value | 5.27518 (rank : 103) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9ESX4 | Gene names | Zcchc17, Ps1d | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar protein of 40 kDa (pNO40) (Zinc finger CCHC domain- containing protein 17) (Putative S1 RNA-binding domain protein) (PS1D protein). | |||||
|
CBX4_HUMAN
|
||||||
| NC score | 0.023458 (rank : 140) | θ value | 2.36792 (rank : 69) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O00257, Q96C04 | Gene names | CBX4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 4 (Polycomb 2 homolog) (Pc2) (hPc2). | |||||
|
DEK_HUMAN
|
||||||
| NC score | 0.023342 (rank : 141) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P35659, Q5TGV4, Q5TGV5 | Gene names | DEK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
BRWD1_MOUSE
|
||||||
| NC score | 0.022335 (rank : 142) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 478 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q921C3, Q921C2 | Gene names | Brwd1, Wdr9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and WD repeat domain-containing protein 1 (WD repeat protein 9). | |||||
|
MYST3_MOUSE
|
||||||
| NC score | 0.022305 (rank : 143) | θ value | 3.0926 (rank : 80) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
MICA1_HUMAN
|
||||||
| NC score | 0.021780 (rank : 144) | θ value | 6.88961 (rank : 114) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8TDZ2, Q7Z633, Q8IVS9, Q96G47, Q9H6X6, Q9H7I0, Q9HAA1, Q9UFF7 | Gene names | MICAL1, MICAL, NICAL | |||
|
Domain Architecture |

|
|||||
| Description | NEDD9-interacting protein with calponin homology and LIM domains (Molecule interacting with CasL protein 1). | |||||
|
DIAP2_HUMAN
|
||||||
| NC score | 0.020831 (rank : 145) | θ value | 3.0926 (rank : 76) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O60879, O60878, Q8WX06, Q8WX48, Q9UJL2 | Gene names | DIAPH2, DIA | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 2 (Diaphanous-related formin-2) (DRF2). | |||||
|
PRP18_MOUSE
|
||||||
| NC score | 0.020050 (rank : 146) | θ value | 8.99809 (rank : 131) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BM39, Q3UX46, Q8BZ54 | Gene names | Prpf18, Prp18 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-splicing factor 18 (PRP18 homolog). | |||||
|
DDX11_MOUSE
|
||||||
| NC score | 0.019595 (rank : 147) | θ value | 2.36792 (rank : 71) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6AXC6 | Gene names | Ddx11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX11 (EC 3.6.1.-) (DEAD/H box protein 11). | |||||
|
MAST2_MOUSE
|
||||||
| NC score | 0.019337 (rank : 148) | θ value | 0.0736092 (rank : 17) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1110 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q60592, Q6P9M1, Q8CHD1 | Gene names | Mast2, Mast205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 2 (EC 2.7.11.1). | |||||
|
PRP18_HUMAN
|
||||||
| NC score | 0.019192 (rank : 149) | θ value | 8.99809 (rank : 130) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99633, Q9BUI9 | Gene names | PRPF18, HPRP18 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-splicing factor 18 (PRP18 homolog) (hPRP18). | |||||
|
SEC62_MOUSE
|
||||||
| NC score | 0.019047 (rank : 150) | θ value | 8.99809 (rank : 134) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BU14, Q6NX81 | Gene names | Tloc1, Sec62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Translocation protein SEC62 (Translocation protein 1) (TP-1). | |||||
|
TRI35_MOUSE
|
||||||
| NC score | 0.018841 (rank : 151) | θ value | 2.36792 (rank : 75) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 465 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8C006, Q6ZPY0, Q7TQL7, Q810V7, Q8BVY9, Q8VID4, Q9CW74, Q9D208 | Gene names | Trim35, Hls5, Kiaa1098, Mair | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 35 (Macrophage-derived apoptosis- inducing RBCC protein) (Protein MAIR) (Protein Nc8) (Hemopoietic lineage switch protein 5). | |||||
|
IQEC2_MOUSE
|
||||||
| NC score | 0.017622 (rank : 152) | θ value | 8.99809 (rank : 125) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5DU25 | Gene names | Iqsec2, Kiaa0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
SOX9_MOUSE
|
||||||
| NC score | 0.017024 (rank : 153) | θ value | 5.27518 (rank : 108) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q04887, Q8C7L2, Q91ZK2, Q99KQ0 | Gene names | Sox9, Sox-9 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-9. | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.016882 (rank : 154) | θ value | 2.36792 (rank : 73) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
GCYB2_HUMAN
|
||||||
| NC score | 0.016472 (rank : 155) | θ value | 4.03905 (rank : 90) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75343, Q9NZ64 | Gene names | GUCY1B2 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylate cyclase soluble subunit beta-2 (EC 4.6.1.2) (GCS-beta-2). | |||||
|
SENP7_HUMAN
|
||||||
| NC score | 0.016240 (rank : 156) | θ value | 8.99809 (rank : 135) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BQF6, Q96PS5, Q9C0F6, Q9HBT5 | Gene names | SENP7, KIAA1707, SSP2, SUSP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 7 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP7) (SUMO-1-specific protease 2). | |||||
|
HPPD_HUMAN
|
||||||
| NC score | 0.016227 (rank : 157) | θ value | 6.88961 (rank : 112) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P32754, Q13234 | Gene names | HPD, PPD | |||
|
Domain Architecture |

|
|||||
| Description | 4-hydroxyphenylpyruvate dioxygenase (EC 1.13.11.27) (4HPPD) (HPD) (HPPDase) (4-hydroxyphenylpyruvic acid oxidase). | |||||
|
WDHD1_MOUSE
|
||||||
| NC score | 0.016125 (rank : 158) | θ value | 4.03905 (rank : 93) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P59328, Q6P408 | Gene names | Wdhd1, And1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and HMG-box DNA-binding protein 1 (Acidic nucleoplasmic DNA- binding protein 1) (And-1). | |||||
|
DDX46_HUMAN
|
||||||
| NC score | 0.015136 (rank : 159) | θ value | 5.27518 (rank : 98) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q7L014, O94894, Q96EI0, Q9Y658 | Gene names | DDX46, KIAA0801 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX46 (EC 3.6.1.-) (DEAD box protein 46) (PRP5 homolog). | |||||
|
CD209_HUMAN
|
||||||
| NC score | 0.015028 (rank : 160) | θ value | 6.88961 (rank : 109) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NNX6, Q96QP7, Q96QP8, Q96QP9, Q96QQ0, Q96QQ1, Q96QQ2, Q96QQ3, Q96QQ4, Q96QQ5, Q96QQ6, Q96QQ7, Q96QQ8 | Gene names | CD209, CLEC4L | |||
|
Domain Architecture |

|
|||||
| Description | CD209 antigen (Dendritic cell-specific ICAM-3-grabbing nonintegrin 1) (DC-SIGN1) (DC-SIGN) (C-type lectin domain family 4 member L). | |||||
|
RNF31_MOUSE
|
||||||
| NC score | 0.014598 (rank : 161) | θ value | 5.27518 (rank : 107) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q924T7 | Gene names | Rnf31 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 31. | |||||
|
LATS1_MOUSE
|
||||||
| NC score | 0.014494 (rank : 162) | θ value | 1.81305 (rank : 66) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1027 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BYR2, Q9Z0W4 | Gene names | Lats1, Warts | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase LATS1 (EC 2.7.11.1) (Large tumor suppressor homolog 1) (WARTS protein kinase). | |||||
|
MAST2_HUMAN
|
||||||
| NC score | 0.014221 (rank : 163) | θ value | 0.47712 (rank : 36) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1189 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6P0Q8, O94899, Q5VT07, Q5VT08, Q7LGC4, Q8NDG1, Q96B94, Q9BYE8 | Gene names | MAST2, KIAA0807, MAST205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 2 (EC 2.7.11.1). | |||||
|
FOXN4_HUMAN
|
||||||
| NC score | 0.013677 (rank : 164) | θ value | 1.38821 (rank : 58) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96NZ1, Q6ZMR4, Q96NZ0 | Gene names | FOXN4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein N4. | |||||
|
PMS2_MOUSE
|
||||||
| NC score | 0.012762 (rank : 165) | θ value | 6.88961 (rank : 116) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P54279 | Gene names | Pms2 | |||
|
Domain Architecture |

|
|||||
| Description | PMS1 protein homolog 2 (DNA mismatch repair protein PMS2). | |||||
|
PARC_HUMAN
|
||||||
| NC score | 0.012267 (rank : 166) | θ value | 5.27518 (rank : 104) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IWT3, O75188, Q68CP2, Q68D92, Q8N3W9, Q9BU56 | Gene names | PARC, H7AP1, KIAA0708 | |||
|
Domain Architecture |

|
|||||
| Description | p53-associated parkin-like cytoplasmic protein (UbcH7-associated protein 1). | |||||
|
EYA2_MOUSE
|
||||||
| NC score | 0.011964 (rank : 167) | θ value | 4.03905 (rank : 89) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O08575, P97925 | Gene names | Eya2, Eab1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 2 (EC 3.1.3.48). | |||||
|
LATS1_HUMAN
|
||||||
| NC score | 0.011391 (rank : 168) | θ value | 5.27518 (rank : 102) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1057 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O95835, Q6PKD0 | Gene names | LATS1, WARTS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase LATS1 (EC 2.7.11.1) (Large tumor suppressor homolog 1) (WARTS protein kinase) (h-warts). | |||||
|
RU17_MOUSE
|
||||||
| NC score | 0.011171 (rank : 169) | θ value | 8.99809 (rank : 133) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q62376, Q3UIW4 | Gene names | Snrp70 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U1 small nuclear ribonucleoprotein 70 kDa (U1 SNRNP 70 kDa) (snRNP70). | |||||
|
CSKI1_MOUSE
|
||||||
| NC score | 0.008589 (rank : 170) | θ value | 8.99809 (rank : 120) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
PC11X_HUMAN
|
||||||
| NC score | 0.007517 (rank : 171) | θ value | 2.36792 (rank : 74) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BZA7, Q2TJH0, Q2TJH1, Q2TJH3, Q5JVZ0, Q70LR8, Q70LS7, Q70LS8, Q70LS9, Q70LT7, Q70LT8, Q70LT9, Q70LU0, Q70LU1, Q96RV4, Q96RW0, Q9BZA6, Q9H4E0, Q9P2M0, Q9P2X5 | Gene names | PCDH11X, PCDH11, PCDHX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protocadherin-11 X-linked precursor (Protocadherin-11) (Protocadherin on the X chromosome) (PCDH-X) (Protocadherin-S). | |||||
|
DDX41_MOUSE
|
||||||
| NC score | 0.007364 (rank : 172) | θ value | 4.03905 (rank : 87) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q91VN6 | Gene names | Ddx41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX41 (EC 3.6.1.-) (DEAD box protein 41). | |||||
|
LDB3_MOUSE
|
||||||
| NC score | 0.006673 (rank : 173) | θ value | 8.99809 (rank : 126) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 510 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9JKS4, Q6A038, Q811P2, Q811P3, Q811P4, Q811P5, Q9D130, Q9JKS3, Q9R0Z1, Q9WVH1, Q9WVH2 | Gene names | Ldb3, Kiaa0613 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LIM domain-binding protein 3 (Z-band alternatively spliced PDZ-motif protein) (Protein cypher) (Protein oracle). | |||||
|
FGD6_MOUSE
|
||||||
| NC score | 0.006107 (rank : 174) | θ value | 6.88961 (rank : 111) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q69ZL1, Q8C8W5, Q8K3B0, Q9D3Y7 | Gene names | Fgd6, Kiaa1362 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 6. | |||||
|
TCF8_HUMAN
|
||||||
| NC score | 0.006071 (rank : 175) | θ value | 0.813845 (rank : 46) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1057 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P37275, Q12924, Q13800 | Gene names | TCF8, AREB6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (NIL-2-A zinc finger protein) (Negative regulator of IL2). | |||||
|
PGCA_MOUSE
|
||||||
| NC score | 0.004354 (rank : 176) | θ value | 8.99809 (rank : 129) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 643 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q61282, Q64021 | Gene names | Agc1, Agc | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP). | |||||
|
ZFP38_HUMAN
|
||||||
| NC score | -0.001291 (rank : 177) | θ value | 3.0926 (rank : 86) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 766 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y5A6, Q9H0B5 | Gene names | ZNF38, ZFP38, ZSCAN21 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 38 homolog (Zfp-38) (Zinc finger and SCAN domain- containing protein 21) (NY-REN-21 antigen). | |||||