Please be patient as the page loads
|
DSRAD_MOUSE
|
||||||
| SwissProt Accessions | Q99MU3, O70375, Q80UZ6, Q8C222, Q99MU2, Q99MU4, Q99MU7 | Gene names | Adar | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific adenosine deaminase (EC 3.5.4.-) (DRADA) (RNA adenosine deaminase 1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
DSRAD_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.982060 (rank : 2) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P55265, O15223, O43859, O43860, Q9BYM3, Q9BYM4 | Gene names | ADAR, ADAR1, DSRAD, IFI4 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific adenosine deaminase (EC 3.5.4.-) (DRADA) (136 kDa double-stranded RNA-binding protein) (P136) (K88DSRBP) (Interferon-inducible protein 4) (IFI-4 protein). | |||||
|
DSRAD_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q99MU3, O70375, Q80UZ6, Q8C222, Q99MU2, Q99MU4, Q99MU7 | Gene names | Adar | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific adenosine deaminase (EC 3.5.4.-) (DRADA) (RNA adenosine deaminase 1). | |||||
|
RED1_MOUSE
|
||||||
| θ value | 1.04338e-64 (rank : 3) | NC score | 0.835251 (rank : 4) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q91ZS8, Q8K3X1, Q91ZS6, Q91ZS7, Q91ZS9, Q99MU8 | Gene names | Adarb1, Adar2, Red1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-specific editase 1 (EC 3.5.-.-) (dsRNA adenosine deaminase) (RNA-editing deaminase 1) (RNA-editing enzyme 1). | |||||
|
RED2_HUMAN
|
||||||
| θ value | 1.15359e-63 (rank : 4) | NC score | 0.838665 (rank : 3) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NS39 | Gene names | ADARB2, ADAR3, RED2 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase B2 (EC 3.5.-.-) (dsRNA adenosine deaminase B2) (RNA-dependent adenosine deaminase 3) (RNA-editing deaminase 2) (RNA-editing enzyme 2). | |||||
|
RED1_HUMAN
|
||||||
| θ value | 4.38363e-63 (rank : 5) | NC score | 0.831313 (rank : 6) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P78563, O00395, O00465, O00691, O00692, P78555, Q8NFD1 | Gene names | ADARB1, ADAR2, DRADA2, RED1 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase 1 (EC 3.5.-.-) (dsRNA adenosine deaminase) (RNA-editing deaminase 1) (RNA-editing enzyme 1). | |||||
|
RED2_MOUSE
|
||||||
| θ value | 6.99855e-61 (rank : 6) | NC score | 0.833387 (rank : 5) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9JI20, Q3UTP1, Q8CC51, Q9JIE5 | Gene names | Adarb2, Adar3, Red2 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase B2 (EC 3.5.-.-) (dsRNA adenosine deaminase B2) (RNA-dependent adenosine deaminase 3) (RNA-editing deaminase 2) (RNA-editing enzyme 2). | |||||
|
STAU2_MOUSE
|
||||||
| θ value | 1.38499e-08 (rank : 7) | NC score | 0.400085 (rank : 9) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8CJ67, Q8BSY8, Q8CJ66, Q8R175, Q91Z19, Q9D5N7 | Gene names | Stau2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 2. | |||||
|
STAU2_HUMAN
|
||||||
| θ value | 1.53129e-07 (rank : 8) | NC score | 0.375865 (rank : 10) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9NUL3, Q6AHY7, Q96HM0, Q96HM1, Q9NVI5, Q9UGG6 | Gene names | STAU2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 2. | |||||
|
PRKRA_MOUSE
|
||||||
| θ value | 9.92553e-07 (rank : 9) | NC score | 0.335662 (rank : 11) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9WTX2, Q9CZB7 | Gene names | Prkra, Rax | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interferon-inducible double stranded RNA-dependent protein kinase activator A (Protein kinase, interferon-inducible double stranded RNA- dependent activator) (Protein activator of the interferon-induced protein kinase) (PKR-associated protein X) (PKR-associating protein X) (RAX). | |||||
|
PRKRA_HUMAN
|
||||||
| θ value | 1.29631e-06 (rank : 10) | NC score | 0.334566 (rank : 12) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O75569, Q53G24, Q6X7T5, Q8NDK4 | Gene names | PRKRA, PACT, RAX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interferon-inducible double stranded RNA-dependent protein kinase activator A (Protein kinase, interferon-inducible double stranded RNA- dependent activator) (Protein activator of the interferon-induced protein kinase) (PKR-associated protein X) (PKR-associating protein X). | |||||
|
ZBP1_MOUSE
|
||||||
| θ value | 2.21117e-06 (rank : 11) | NC score | 0.202321 (rank : 17) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9QY24, Q8VE02, Q9D8B9 | Gene names | Zbp1, Dlm1 | |||
|
Domain Architecture |
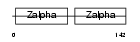
|
|||||
| Description | Z-DNA-binding protein 1 (Tumor stroma and activated macrophage protein DLM-1). | |||||
|
E2AK2_MOUSE
|
||||||
| θ value | 1.43324e-05 (rank : 12) | NC score | 0.039524 (rank : 24) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q03963, Q61742, Q62026 | Gene names | Eif2ak2, Pkr, Prkr, Tik | |||
|
Domain Architecture |
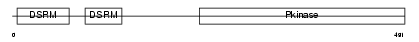
|
|||||
| Description | Interferon-induced, double-stranded RNA-activated protein kinase (EC 2.7.11.1) (Interferon-inducible RNA-dependent protein kinase) (Protein kinase RNA-activated) (PKR) (p68 kinase) (P1/eIF-2A protein kinase) (Serine/threonine-protein kinase TIK). | |||||
|
ILF3_MOUSE
|
||||||
| θ value | 1.87187e-05 (rank : 13) | NC score | 0.452207 (rank : 7) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Z1X4, Q80VD5, Q812A1, Q8BP80, Q8C2H8, Q8K588 | Gene names | Ilf3 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin enhancer-binding factor 3. | |||||
|
ILF3_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 14) | NC score | 0.451859 (rank : 8) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q12906, O43409, Q6P1X1, Q86XY7, Q99544, Q99545, Q9BZH4, Q9BZH5, Q9NQ95, Q9NQ96, Q9NQ97, Q9NQ98, Q9NQ99, Q9NQA0, Q9NQA1, Q9NQA2, Q9NRN2, Q9NRN3, Q9NRN4, Q9UMZ9, Q9UN00, Q9UN84, Q9UNA2 | Gene names | ILF3, DRBF, MPHOSPH4, NF90 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin enhancer-binding factor 3 (Nuclear factor of activated T- cells 90 kDa) (NF-AT-90) (Double-stranded RNA-binding protein 76) (DRBP76) (Translational control protein 80) (TCP80) (Nuclear factor associated with dsRNA) (NFAR) (M-phase phosphoprotein 4) (MPP4). | |||||
|
E2AK2_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 15) | NC score | 0.053059 (rank : 21) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P19525 | Gene names | EIF2AK2, PKR, PRKR | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced, double-stranded RNA-activated protein kinase (EC 2.7.11.1) (Interferon-inducible RNA-dependent protein kinase) (Protein kinase RNA-activated) (PKR) (p68 kinase) (P1/eIF-2A protein kinase). | |||||
|
TRBP2_MOUSE
|
||||||
| θ value | 0.00102713 (rank : 16) | NC score | 0.324699 (rank : 13) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P97473 | Gene names | Tarbp2, Prbp | |||
|
Domain Architecture |
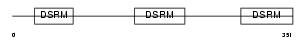
|
|||||
| Description | TAR RNA-binding protein 2 (Protamine-1 RNA-binding protein) (PRM-1 RNA-binding protein). | |||||
|
ZBP1_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 17) | NC score | 0.143385 (rank : 18) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H171, Q5JY39, Q9BYW4 | Gene names | ZBP1, C20orf183, DLM1 | |||
|
Domain Architecture |
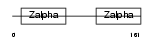
|
|||||
| Description | Z-DNA-binding protein 1 (Tumor stroma and activated macrophage protein DLM-1). | |||||
|
PIWL2_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 18) | NC score | 0.034428 (rank : 26) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TC59, Q96SW6, Q9NW28 | Gene names | PIWIL2, HILI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 2. | |||||
|
MYO15_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 19) | NC score | 0.019268 (rank : 39) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9QZZ4, O70395, Q9QWL6 | Gene names | Myo15a, Myo15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
NKRF_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 20) | NC score | 0.074797 (rank : 19) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O15226, Q9UJ91 | Gene names | NKRF, ITBA4, NRF | |||
|
Domain Architecture |
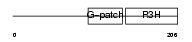
|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF) (ITBA4 protein). | |||||
|
TRBP2_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 21) | NC score | 0.306785 (rank : 14) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q15633, Q12878 | Gene names | TARBP2, TRBP | |||
|
Domain Architecture |

|
|||||
| Description | TAR RNA-binding protein 2 (Trans-activation-responsive RNA-binding protein). | |||||
|
SON_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 22) | NC score | 0.067643 (rank : 20) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
APBA2_HUMAN
|
||||||
| θ value | 0.125558 (rank : 23) | NC score | 0.030669 (rank : 28) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99767, O60571 | Gene names | APBA2, MINT2 | |||
|
Domain Architecture |

|
|||||
| Description | Amyloid beta A4 precursor protein-binding family A member 2 (Neuron- specific X11L protein) (Neuronal Munc18-1-interacting protein 2) (Mint-2) (Adapter protein X11beta). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 0.163984 (rank : 24) | NC score | 0.015424 (rank : 47) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
VMD2_MOUSE
|
||||||
| θ value | 0.163984 (rank : 25) | NC score | 0.030150 (rank : 29) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O88870, Q6H1V0 | Gene names | Vmd2, Bmd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bestrophin-1 (Vitelliform macular dystrophy protein 2 homolog). | |||||
|
DLG5_HUMAN
|
||||||
| θ value | 0.279714 (rank : 26) | NC score | 0.018735 (rank : 40) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 1088 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8TDM6, Q5T1H7, Q5T1H8, Q6DKG3, Q86WC0, Q8TDM7, Q9UE73, Q9Y4E3 | Gene names | DLG5, KIAA0583, PDLG | |||
|
Domain Architecture |

|
|||||
| Description | Discs large homolog 5 (Placenta and prostate DLG) (Discs large protein P-dlg). | |||||
|
SON_HUMAN
|
||||||
| θ value | 0.47712 (rank : 27) | NC score | 0.043697 (rank : 23) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
ULK2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 28) | NC score | 0.007990 (rank : 56) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 979 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IYT8, O75119 | Gene names | ULK2, KIAA0623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ULK2 (EC 2.7.11.1) (Unc-51-like kinase 2). | |||||
|
BRD4_MOUSE
|
||||||
| θ value | 0.62314 (rank : 29) | NC score | 0.024810 (rank : 32) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
HGS_HUMAN
|
||||||
| θ value | 0.62314 (rank : 30) | NC score | 0.028256 (rank : 30) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O14964, Q9NR36 | Gene names | HGS, HRS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate (Protein pp110) (Hrs). | |||||
|
MAGD1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 31) | NC score | 0.015902 (rank : 46) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9QYH6, Q543L6, Q99PB5, Q9CYX1 | Gene names | Maged1, Nrage | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen D1 (MAGE-D1 antigen) (Neurotrophin receptor-interacting MAGE homolog) (Dlxin-1). | |||||
|
PDCD7_MOUSE
|
||||||
| θ value | 0.62314 (rank : 32) | NC score | 0.038266 (rank : 25) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9WTY1, Q8R5D9 | Gene names | Pdcd7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Programmed cell death protein 7 (ES18). | |||||
|
FOXM1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 33) | NC score | 0.015291 (rank : 48) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q08050, O43258, O43259, O43260, Q4ZGG7, Q9BRL2 | Gene names | FOXM1, FKHL16, HFH11, MPP2, WIN | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein M1 (Forkhead-related protein FKHL16) (Hepatocyte nuclear factor 3 forkhead homolog 11) (HNF-3/fork-head homolog 11) (HFH-11) (Winged helix factor from INS-1 cells) (M-phase phosphoprotein 2) (MPM-2 reactive phosphoprotein 2) (Transcription factor Trident). | |||||
|
CTGE6_HUMAN
|
||||||
| θ value | 1.38821 (rank : 34) | NC score | 0.012620 (rank : 51) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q86UF2 | Gene names | CTAGE6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein cTAGE-6 (cTAGE family member 6). | |||||
|
HGS_MOUSE
|
||||||
| θ value | 1.38821 (rank : 35) | NC score | 0.026565 (rank : 31) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99LI8, Q61691, Q8BQW3 | Gene names | Hgs, Hrs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate. | |||||
|
SEPT9_MOUSE
|
||||||
| θ value | 1.38821 (rank : 36) | NC score | 0.017409 (rank : 42) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80UG5, Q3URP2, Q80TM7, Q9QYX9 | Gene names | Sept9, Kiaa0991, Sint1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Septin-9 (SL3-3 integration site 1 protein). | |||||
|
STAU1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 37) | NC score | 0.248059 (rank : 16) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Z108 | Gene names | Stau1, Stau | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 1. | |||||
|
BRD4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 38) | NC score | 0.022929 (rank : 36) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
CIC_HUMAN
|
||||||
| θ value | 1.81305 (rank : 39) | NC score | 0.017166 (rank : 44) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
SHB_MOUSE
|
||||||
| θ value | 1.81305 (rank : 40) | NC score | 0.017234 (rank : 43) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PD21, Q3ULM3 | Gene names | Shb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH2 domain-containing adapter protein B. | |||||
|
STAU1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 41) | NC score | 0.251661 (rank : 15) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O95793, Q6GTM4, Q9H5B4, Q9H5B5, Q9Y3Q2 | Gene names | STAU1, STAU | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 1. | |||||
|
ULK2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 42) | NC score | 0.006846 (rank : 59) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 943 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9QY01, Q80TV7, Q9WTP4 | Gene names | Ulk2, Kiaa0623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ULK2 (EC 2.7.11.1) (Unc-51-like kinase 2) (Serine/threonine-protein kinase Unc51.2). | |||||
|
CPSF6_HUMAN
|
||||||
| θ value | 3.0926 (rank : 43) | NC score | 0.023787 (rank : 34) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q16630, Q53ES1, Q9BSJ7, Q9BW18 | Gene names | CPSF6, CFIM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6 (Cleavage and polyadenylation specificity factor 68 kDa subunit) (CPSF 68 kDa subunit) (Pre-mRNA cleavage factor Im 68 kDa subunit) (Protein HPBRII- 4/7). | |||||
|
CPSF6_MOUSE
|
||||||
| θ value | 3.0926 (rank : 44) | NC score | 0.024076 (rank : 33) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6NVF9, Q8BX86, Q8BXI8 | Gene names | Cpsf6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6. | |||||
|
LMO7_HUMAN
|
||||||
| θ value | 3.0926 (rank : 45) | NC score | 0.017580 (rank : 41) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8WWI1, O15462, O95346, Q9UKC1, Q9UQM5, Q9Y6A7 | Gene names | LMO7, FBX20, FBXO20, KIAA0858 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain only protein 7 (LOMP) (F-box only protein 20). | |||||
|
TXD11_HUMAN
|
||||||
| θ value | 3.0926 (rank : 46) | NC score | 0.014698 (rank : 49) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6PKC3, O95887, Q6PJA6, Q8N2Q4, Q96K45, Q96K53 | Gene names | TXNDC11, EFP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 11 (EF-hand-binding protein 1). | |||||
|
WASF4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 47) | NC score | 0.017139 (rank : 45) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8IV90 | Gene names | WASF4, SCAR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 4 (WASP-family protein member 4). | |||||
|
BAT2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 48) | NC score | 0.020618 (rank : 38) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
CABIN_HUMAN
|
||||||
| θ value | 4.03905 (rank : 49) | NC score | 0.022309 (rank : 37) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y6J0, Q9Y460 | Gene names | CABIN1, KIAA0330 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin-binding protein Cabin 1 (Calcineurin inhibitor) (CAIN). | |||||
|
FBSH_MOUSE
|
||||||
| θ value | 4.03905 (rank : 50) | NC score | 0.023346 (rank : 35) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8R089 | Gene names | Fbs1, Fbs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable fibrosin-1 long transcript protein. | |||||
|
LAP4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 51) | NC score | 0.006539 (rank : 60) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 608 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14160, Q6P496, Q7Z5D1, Q8WWV8, Q96C69, Q96GG1 | Gene names | SCRIB, CRIB1, KIAA0147, LAP4, SCRB1, VARTUL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP4 (Protein scribble homolog) (hScrib). | |||||
|
SMR3A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 52) | NC score | 0.031439 (rank : 27) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99954 | Gene names | SMR3A, PBI, PROL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Submaxillary gland androgen-regulated protein 3 homolog A precursor (Proline-rich protein 5) (Proline-rich protein PBI). | |||||
|
DNL3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.012827 (rank : 50) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P49916, Q16714 | Gene names | LIG3 | |||
|
Domain Architecture |
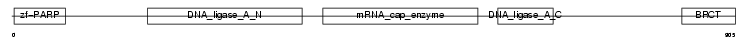
|
|||||
| Description | DNA ligase 3 (EC 6.5.1.1) (DNA ligase III) (Polydeoxyribonucleotide synthase [ATP] 3). | |||||
|
ETV3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | 0.009766 (rank : 55) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
CAC1B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 55) | NC score | 0.004073 (rank : 64) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q00975 | Gene names | CACNA1B, CACH5, CACNL1A5 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent N-type calcium channel subunit alpha-1B (Voltage- gated calcium channel subunit alpha Cav2.2) (Calcium channel, L type, alpha-1 polypeptide isoform 5) (Brain calcium channel III) (BIII). | |||||
|
DUX4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 56) | NC score | 0.004653 (rank : 63) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UBX2 | Gene names | DUX4, DUX10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double homeobox protein 4 (Double homeobox protein 10) (Double homeobox protein 4/10). | |||||
|
E41L2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 57) | NC score | 0.005642 (rank : 62) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O70318 | Gene names | Epb41l2, Epb4.1l2 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 2 (Generally expressed protein 4.1) (4.1G). | |||||
|
NKRF_MOUSE
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | 0.045712 (rank : 22) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BY02, Q8BJU5, Q9CRL2 | Gene names | Nkrf, Nrf | |||
|
Domain Architecture |

|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF). | |||||
|
RD23B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 59) | NC score | 0.012617 (rank : 52) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P54727, Q8WUB0 | Gene names | RAD23B | |||
|
Domain Architecture |
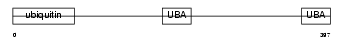
|
|||||
| Description | UV excision repair protein RAD23 homolog B (hHR23B) (XP-C repair- complementing complex 58 kDa protein) (p58). | |||||
|
SLIK3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 60) | NC score | 0.003278 (rank : 65) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O94933 | Gene names | SLITRK3, KIAA0848 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SLIT and NTRK-like protein 3 precursor. | |||||
|
TAIP2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | 0.009892 (rank : 54) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8WYN3 | Gene names | TAIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TGF-beta-induced apoptosis protein 2 (TAIP-2). | |||||
|
ZEP2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.001666 (rank : 67) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 964 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q3UHF7, O55140, Q3UVD4, Q3UVH5 | Gene names | Hivep2, Mibp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Human immunodeficiency virus type I enhancer-binding protein 2 homolog (Myc intron-binding protein 1) (MIBP-1). | |||||
|
CO8A2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 63) | NC score | 0.007175 (rank : 58) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 299 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P25067, Q8TEJ5 | Gene names | COL8A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(VIII) chain precursor (Endothelial collagen). | |||||
|
CSPG2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 64) | NC score | 0.002398 (rank : 66) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P13611, P20754, Q13010, Q13189, Q15123, Q9UCL9, Q9UNW5 | Gene names | CSPG2 | |||
|
Domain Architecture |

|
|||||
| Description | Versican core protein precursor (Large fibroblast proteoglycan) (Chondroitin sulfate proteoglycan core protein 2) (PG-M) (Glial hyaluronate-binding protein) (GHAP). | |||||
|
ENPP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 65) | NC score | 0.006042 (rank : 61) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P06802 | Gene names | Enpp1, Npps, Pc1, Pdnp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 1 (E-NPP 1) (Phosphodiesterase I/nucleotide pyrophosphatase 1) (Plasma-cell membrane glycoprotein PC-1) (Antigen Ly-41) [Includes: Alkaline phosphodiesterase I (EC 3.1.4.1); Nucleotide pyrophosphatase (EC 3.6.1.9) (NPPase)]. | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 66) | NC score | 0.010383 (rank : 53) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
NFH_HUMAN
|
||||||
| θ value | 8.99809 (rank : 67) | NC score | 0.007291 (rank : 57) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
DSRAD_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q99MU3, O70375, Q80UZ6, Q8C222, Q99MU2, Q99MU4, Q99MU7 | Gene names | Adar | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific adenosine deaminase (EC 3.5.4.-) (DRADA) (RNA adenosine deaminase 1). | |||||
|
DSRAD_HUMAN
|
||||||
| NC score | 0.982060 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P55265, O15223, O43859, O43860, Q9BYM3, Q9BYM4 | Gene names | ADAR, ADAR1, DSRAD, IFI4 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific adenosine deaminase (EC 3.5.4.-) (DRADA) (136 kDa double-stranded RNA-binding protein) (P136) (K88DSRBP) (Interferon-inducible protein 4) (IFI-4 protein). | |||||
|
RED2_HUMAN
|
||||||
| NC score | 0.838665 (rank : 3) | θ value | 1.15359e-63 (rank : 4) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NS39 | Gene names | ADARB2, ADAR3, RED2 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase B2 (EC 3.5.-.-) (dsRNA adenosine deaminase B2) (RNA-dependent adenosine deaminase 3) (RNA-editing deaminase 2) (RNA-editing enzyme 2). | |||||
|
RED1_MOUSE
|
||||||
| NC score | 0.835251 (rank : 4) | θ value | 1.04338e-64 (rank : 3) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q91ZS8, Q8K3X1, Q91ZS6, Q91ZS7, Q91ZS9, Q99MU8 | Gene names | Adarb1, Adar2, Red1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-specific editase 1 (EC 3.5.-.-) (dsRNA adenosine deaminase) (RNA-editing deaminase 1) (RNA-editing enzyme 1). | |||||
|
RED2_MOUSE
|
||||||
| NC score | 0.833387 (rank : 5) | θ value | 6.99855e-61 (rank : 6) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9JI20, Q3UTP1, Q8CC51, Q9JIE5 | Gene names | Adarb2, Adar3, Red2 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase B2 (EC 3.5.-.-) (dsRNA adenosine deaminase B2) (RNA-dependent adenosine deaminase 3) (RNA-editing deaminase 2) (RNA-editing enzyme 2). | |||||
|
RED1_HUMAN
|
||||||
| NC score | 0.831313 (rank : 6) | θ value | 4.38363e-63 (rank : 5) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P78563, O00395, O00465, O00691, O00692, P78555, Q8NFD1 | Gene names | ADARB1, ADAR2, DRADA2, RED1 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase 1 (EC 3.5.-.-) (dsRNA adenosine deaminase) (RNA-editing deaminase 1) (RNA-editing enzyme 1). | |||||
|
ILF3_MOUSE
|
||||||
| NC score | 0.452207 (rank : 7) | θ value | 1.87187e-05 (rank : 13) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Z1X4, Q80VD5, Q812A1, Q8BP80, Q8C2H8, Q8K588 | Gene names | Ilf3 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin enhancer-binding factor 3. | |||||
|
ILF3_HUMAN
|
||||||
| NC score | 0.451859 (rank : 8) | θ value | 4.1701e-05 (rank : 14) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q12906, O43409, Q6P1X1, Q86XY7, Q99544, Q99545, Q9BZH4, Q9BZH5, Q9NQ95, Q9NQ96, Q9NQ97, Q9NQ98, Q9NQ99, Q9NQA0, Q9NQA1, Q9NQA2, Q9NRN2, Q9NRN3, Q9NRN4, Q9UMZ9, Q9UN00, Q9UN84, Q9UNA2 | Gene names | ILF3, DRBF, MPHOSPH4, NF90 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin enhancer-binding factor 3 (Nuclear factor of activated T- cells 90 kDa) (NF-AT-90) (Double-stranded RNA-binding protein 76) (DRBP76) (Translational control protein 80) (TCP80) (Nuclear factor associated with dsRNA) (NFAR) (M-phase phosphoprotein 4) (MPP4). | |||||
|
STAU2_MOUSE
|
||||||
| NC score | 0.400085 (rank : 9) | θ value | 1.38499e-08 (rank : 7) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8CJ67, Q8BSY8, Q8CJ66, Q8R175, Q91Z19, Q9D5N7 | Gene names | Stau2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 2. | |||||
|
STAU2_HUMAN
|
||||||
| NC score | 0.375865 (rank : 10) | θ value | 1.53129e-07 (rank : 8) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9NUL3, Q6AHY7, Q96HM0, Q96HM1, Q9NVI5, Q9UGG6 | Gene names | STAU2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 2. | |||||
|
PRKRA_MOUSE
|
||||||
| NC score | 0.335662 (rank : 11) | θ value | 9.92553e-07 (rank : 9) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9WTX2, Q9CZB7 | Gene names | Prkra, Rax | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interferon-inducible double stranded RNA-dependent protein kinase activator A (Protein kinase, interferon-inducible double stranded RNA- dependent activator) (Protein activator of the interferon-induced protein kinase) (PKR-associated protein X) (PKR-associating protein X) (RAX). | |||||
|
PRKRA_HUMAN
|
||||||
| NC score | 0.334566 (rank : 12) | θ value | 1.29631e-06 (rank : 10) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O75569, Q53G24, Q6X7T5, Q8NDK4 | Gene names | PRKRA, PACT, RAX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interferon-inducible double stranded RNA-dependent protein kinase activator A (Protein kinase, interferon-inducible double stranded RNA- dependent activator) (Protein activator of the interferon-induced protein kinase) (PKR-associated protein X) (PKR-associating protein X). | |||||
|
TRBP2_MOUSE
|
||||||
| NC score | 0.324699 (rank : 13) | θ value | 0.00102713 (rank : 16) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P97473 | Gene names | Tarbp2, Prbp | |||
|
Domain Architecture |

|
|||||
| Description | TAR RNA-binding protein 2 (Protamine-1 RNA-binding protein) (PRM-1 RNA-binding protein). | |||||
|
TRBP2_HUMAN
|
||||||
| NC score | 0.306785 (rank : 14) | θ value | 0.0563607 (rank : 21) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q15633, Q12878 | Gene names | TARBP2, TRBP | |||
|
Domain Architecture |

|
|||||
| Description | TAR RNA-binding protein 2 (Trans-activation-responsive RNA-binding protein). | |||||
|
STAU1_HUMAN
|
||||||
| NC score | 0.251661 (rank : 15) | θ value | 1.81305 (rank : 41) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O95793, Q6GTM4, Q9H5B4, Q9H5B5, Q9Y3Q2 | Gene names | STAU1, STAU | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 1. | |||||
|
STAU1_MOUSE
|
||||||
| NC score | 0.248059 (rank : 16) | θ value | 1.38821 (rank : 37) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Z108 | Gene names | Stau1, Stau | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 1. | |||||
|
ZBP1_MOUSE
|
||||||
| NC score | 0.202321 (rank : 17) | θ value | 2.21117e-06 (rank : 11) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9QY24, Q8VE02, Q9D8B9 | Gene names | Zbp1, Dlm1 | |||
|
Domain Architecture |

|
|||||
| Description | Z-DNA-binding protein 1 (Tumor stroma and activated macrophage protein DLM-1). | |||||
|
ZBP1_HUMAN
|
||||||
| NC score | 0.143385 (rank : 18) | θ value | 0.00665767 (rank : 17) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H171, Q5JY39, Q9BYW4 | Gene names | ZBP1, C20orf183, DLM1 | |||
|
Domain Architecture |

|
|||||
| Description | Z-DNA-binding protein 1 (Tumor stroma and activated macrophage protein DLM-1). | |||||
|
NKRF_HUMAN
|
||||||
| NC score | 0.074797 (rank : 19) | θ value | 0.0431538 (rank : 20) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O15226, Q9UJ91 | Gene names | NKRF, ITBA4, NRF | |||
|
Domain Architecture |

|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF) (ITBA4 protein). | |||||
|
SON_MOUSE
|
||||||
| NC score | 0.067643 (rank : 20) | θ value | 0.0736092 (rank : 22) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
E2AK2_HUMAN
|
||||||
| NC score | 0.053059 (rank : 21) | θ value | 0.000270298 (rank : 15) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P19525 | Gene names | EIF2AK2, PKR, PRKR | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced, double-stranded RNA-activated protein kinase (EC 2.7.11.1) (Interferon-inducible RNA-dependent protein kinase) (Protein kinase RNA-activated) (PKR) (p68 kinase) (P1/eIF-2A protein kinase). | |||||
|
NKRF_MOUSE
|
||||||
| NC score | 0.045712 (rank : 22) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BY02, Q8BJU5, Q9CRL2 | Gene names | Nkrf, Nrf | |||
|
Domain Architecture |

|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF). | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.043697 (rank : 23) | θ value | 0.47712 (rank : 27) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
E2AK2_MOUSE
|
||||||
| NC score | 0.039524 (rank : 24) | θ value | 1.43324e-05 (rank : 12) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q03963, Q61742, Q62026 | Gene names | Eif2ak2, Pkr, Prkr, Tik | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced, double-stranded RNA-activated protein kinase (EC 2.7.11.1) (Interferon-inducible RNA-dependent protein kinase) (Protein kinase RNA-activated) (PKR) (p68 kinase) (P1/eIF-2A protein kinase) (Serine/threonine-protein kinase TIK). | |||||
|
PDCD7_MOUSE
|
||||||
| NC score | 0.038266 (rank : 25) | θ value | 0.62314 (rank : 32) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9WTY1, Q8R5D9 | Gene names | Pdcd7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Programmed cell death protein 7 (ES18). | |||||
|
PIWL2_HUMAN
|
||||||
| NC score | 0.034428 (rank : 26) | θ value | 0.0193708 (rank : 18) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TC59, Q96SW6, Q9NW28 | Gene names | PIWIL2, HILI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 2. | |||||
|
SMR3A_HUMAN
|
||||||
| NC score | 0.031439 (rank : 27) | θ value | 4.03905 (rank : 52) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99954 | Gene names | SMR3A, PBI, PROL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Submaxillary gland androgen-regulated protein 3 homolog A precursor (Proline-rich protein 5) (Proline-rich protein PBI). | |||||
|
APBA2_HUMAN
|
||||||
| NC score | 0.030669 (rank : 28) | θ value | 0.125558 (rank : 23) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99767, O60571 | Gene names | APBA2, MINT2 | |||
|
Domain Architecture |

|
|||||
| Description | Amyloid beta A4 precursor protein-binding family A member 2 (Neuron- specific X11L protein) (Neuronal Munc18-1-interacting protein 2) (Mint-2) (Adapter protein X11beta). | |||||
|
VMD2_MOUSE
|
||||||
| NC score | 0.030150 (rank : 29) | θ value | 0.163984 (rank : 25) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O88870, Q6H1V0 | Gene names | Vmd2, Bmd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bestrophin-1 (Vitelliform macular dystrophy protein 2 homolog). | |||||
|
HGS_HUMAN
|
||||||
| NC score | 0.028256 (rank : 30) | θ value | 0.62314 (rank : 30) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O14964, Q9NR36 | Gene names | HGS, HRS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate (Protein pp110) (Hrs). | |||||
|
HGS_MOUSE
|
||||||
| NC score | 0.026565 (rank : 31) | θ value | 1.38821 (rank : 35) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99LI8, Q61691, Q8BQW3 | Gene names | Hgs, Hrs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate. | |||||
|
BRD4_MOUSE
|
||||||
| NC score | 0.024810 (rank : 32) | θ value | 0.62314 (rank : 29) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
CPSF6_MOUSE
|
||||||
| NC score | 0.024076 (rank : 33) | θ value | 3.0926 (rank : 44) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6NVF9, Q8BX86, Q8BXI8 | Gene names | Cpsf6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6. | |||||
|
CPSF6_HUMAN
|
||||||
| NC score | 0.023787 (rank : 34) | θ value | 3.0926 (rank : 43) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q16630, Q53ES1, Q9BSJ7, Q9BW18 | Gene names | CPSF6, CFIM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6 (Cleavage and polyadenylation specificity factor 68 kDa subunit) (CPSF 68 kDa subunit) (Pre-mRNA cleavage factor Im 68 kDa subunit) (Protein HPBRII- 4/7). | |||||
|
FBSH_MOUSE
|
||||||
| NC score | 0.023346 (rank : 35) | θ value | 4.03905 (rank : 50) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8R089 | Gene names | Fbs1, Fbs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable fibrosin-1 long transcript protein. | |||||
|
BRD4_HUMAN
|
||||||
| NC score | 0.022929 (rank : 36) | θ value | 1.81305 (rank : 38) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
CABIN_HUMAN
|
||||||
| NC score | 0.022309 (rank : 37) | θ value | 4.03905 (rank : 49) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y6J0, Q9Y460 | Gene names | CABIN1, KIAA0330 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin-binding protein Cabin 1 (Calcineurin inhibitor) (CAIN). | |||||
|
BAT2_MOUSE
|
||||||
| NC score | 0.020618 (rank : 38) | θ value | 4.03905 (rank : 48) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
MYO15_MOUSE
|
||||||
| NC score | 0.019268 (rank : 39) | θ value | 0.0252991 (rank : 19) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9QZZ4, O70395, Q9QWL6 | Gene names | Myo15a, Myo15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
DLG5_HUMAN
|
||||||
| NC score | 0.018735 (rank : 40) | θ value | 0.279714 (rank : 26) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 1088 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8TDM6, Q5T1H7, Q5T1H8, Q6DKG3, Q86WC0, Q8TDM7, Q9UE73, Q9Y4E3 | Gene names | DLG5, KIAA0583, PDLG | |||
|
Domain Architecture |

|
|||||
| Description | Discs large homolog 5 (Placenta and prostate DLG) (Discs large protein P-dlg). | |||||
|
LMO7_HUMAN
|
||||||
| NC score | 0.017580 (rank : 41) | θ value | 3.0926 (rank : 45) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8WWI1, O15462, O95346, Q9UKC1, Q9UQM5, Q9Y6A7 | Gene names | LMO7, FBX20, FBXO20, KIAA0858 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain only protein 7 (LOMP) (F-box only protein 20). | |||||
|
SEPT9_MOUSE
|
||||||
| NC score | 0.017409 (rank : 42) | θ value | 1.38821 (rank : 36) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80UG5, Q3URP2, Q80TM7, Q9QYX9 | Gene names | Sept9, Kiaa0991, Sint1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Septin-9 (SL3-3 integration site 1 protein). | |||||
|
SHB_MOUSE
|
||||||
| NC score | 0.017234 (rank : 43) | θ value | 1.81305 (rank : 40) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PD21, Q3ULM3 | Gene names | Shb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH2 domain-containing adapter protein B. | |||||
|
CIC_HUMAN
|
||||||
| NC score | 0.017166 (rank : 44) | θ value | 1.81305 (rank : 39) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
WASF4_HUMAN
|
||||||
| NC score | 0.017139 (rank : 45) | θ value | 3.0926 (rank : 47) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8IV90 | Gene names | WASF4, SCAR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 4 (WASP-family protein member 4). | |||||
|
MAGD1_MOUSE
|
||||||
| NC score | 0.015902 (rank : 46) | θ value | 0.62314 (rank : 31) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9QYH6, Q543L6, Q99PB5, Q9CYX1 | Gene names | Maged1, Nrage | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen D1 (MAGE-D1 antigen) (Neurotrophin receptor-interacting MAGE homolog) (Dlxin-1). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.015424 (rank : 47) | θ value | 0.163984 (rank : 24) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
FOXM1_HUMAN
|
||||||
| NC score | 0.015291 (rank : 48) | θ value | 1.06291 (rank : 33) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q08050, O43258, O43259, O43260, Q4ZGG7, Q9BRL2 | Gene names | FOXM1, FKHL16, HFH11, MPP2, WIN | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein M1 (Forkhead-related protein FKHL16) (Hepatocyte nuclear factor 3 forkhead homolog 11) (HNF-3/fork-head homolog 11) (HFH-11) (Winged helix factor from INS-1 cells) (M-phase phosphoprotein 2) (MPM-2 reactive phosphoprotein 2) (Transcription factor Trident). | |||||
|
TXD11_HUMAN
|
||||||
| NC score | 0.014698 (rank : 49) | θ value | 3.0926 (rank : 46) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6PKC3, O95887, Q6PJA6, Q8N2Q4, Q96K45, Q96K53 | Gene names | TXNDC11, EFP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 11 (EF-hand-binding protein 1). | |||||
|
DNL3_HUMAN
|
||||||
| NC score | 0.012827 (rank : 50) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P49916, Q16714 | Gene names | LIG3 | |||
|
Domain Architecture |
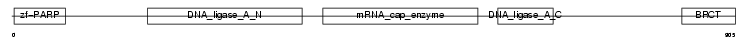
|
|||||
| Description | DNA ligase 3 (EC 6.5.1.1) (DNA ligase III) (Polydeoxyribonucleotide synthase [ATP] 3). | |||||
|
CTGE6_HUMAN
|
||||||
| NC score | 0.012620 (rank : 51) | θ value | 1.38821 (rank : 34) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q86UF2 | Gene names | CTAGE6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein cTAGE-6 (cTAGE family member 6). | |||||
|
RD23B_HUMAN
|
||||||
| NC score | 0.012617 (rank : 52) | θ value | 6.88961 (rank : 59) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P54727, Q8WUB0 | Gene names | RAD23B | |||
|
Domain Architecture |

|
|||||
| Description | UV excision repair protein RAD23 homolog B (hHR23B) (XP-C repair- complementing complex 58 kDa protein) (p58). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.010383 (rank : 53) | θ value | 8.99809 (rank : 66) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
TAIP2_HUMAN
|
||||||
| NC score | 0.009892 (rank : 54) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8WYN3 | Gene names | TAIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TGF-beta-induced apoptosis protein 2 (TAIP-2). | |||||
|
ETV3_HUMAN
|
||||||
| NC score | 0.009766 (rank : 55) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
ULK2_HUMAN
|
||||||
| NC score | 0.007990 (rank : 56) | θ value | 0.47712 (rank : 28) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 979 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IYT8, O75119 | Gene names | ULK2, KIAA0623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ULK2 (EC 2.7.11.1) (Unc-51-like kinase 2). | |||||
|
NFH_HUMAN
|
||||||
| NC score | 0.007291 (rank : 57) | θ value | 8.99809 (rank : 67) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
CO8A2_HUMAN
|
||||||
| NC score | 0.007175 (rank : 58) | θ value | 8.99809 (rank : 63) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 299 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P25067, Q8TEJ5 | Gene names | COL8A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(VIII) chain precursor (Endothelial collagen). | |||||
|
ULK2_MOUSE
|
||||||
| NC score | 0.006846 (rank : 59) | θ value | 2.36792 (rank : 42) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 943 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9QY01, Q80TV7, Q9WTP4 | Gene names | Ulk2, Kiaa0623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ULK2 (EC 2.7.11.1) (Unc-51-like kinase 2) (Serine/threonine-protein kinase Unc51.2). | |||||
|
LAP4_HUMAN
|
||||||
| NC score | 0.006539 (rank : 60) | θ value | 4.03905 (rank : 51) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 608 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14160, Q6P496, Q7Z5D1, Q8WWV8, Q96C69, Q96GG1 | Gene names | SCRIB, CRIB1, KIAA0147, LAP4, SCRB1, VARTUL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP4 (Protein scribble homolog) (hScrib). | |||||
|
ENPP1_MOUSE
|
||||||
| NC score | 0.006042 (rank : 61) | θ value | 8.99809 (rank : 65) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P06802 | Gene names | Enpp1, Npps, Pc1, Pdnp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 1 (E-NPP 1) (Phosphodiesterase I/nucleotide pyrophosphatase 1) (Plasma-cell membrane glycoprotein PC-1) (Antigen Ly-41) [Includes: Alkaline phosphodiesterase I (EC 3.1.4.1); Nucleotide pyrophosphatase (EC 3.6.1.9) (NPPase)]. | |||||
|
E41L2_MOUSE
|
||||||
| NC score | 0.005642 (rank : 62) | θ value | 6.88961 (rank : 57) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O70318 | Gene names | Epb41l2, Epb4.1l2 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 2 (Generally expressed protein 4.1) (4.1G). | |||||
|
DUX4_HUMAN
|
||||||
| NC score | 0.004653 (rank : 63) | θ value | 6.88961 (rank : 56) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UBX2 | Gene names | DUX4, DUX10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double homeobox protein 4 (Double homeobox protein 10) (Double homeobox protein 4/10). | |||||
|
CAC1B_HUMAN
|
||||||
| NC score | 0.004073 (rank : 64) | θ value | 6.88961 (rank : 55) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q00975 | Gene names | CACNA1B, CACH5, CACNL1A5 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent N-type calcium channel subunit alpha-1B (Voltage- gated calcium channel subunit alpha Cav2.2) (Calcium channel, L type, alpha-1 polypeptide isoform 5) (Brain calcium channel III) (BIII). | |||||
|
SLIK3_HUMAN
|
||||||
| NC score | 0.003278 (rank : 65) | θ value | 6.88961 (rank : 60) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O94933 | Gene names | SLITRK3, KIAA0848 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SLIT and NTRK-like protein 3 precursor. | |||||
|
CSPG2_HUMAN
|
||||||
| NC score | 0.002398 (rank : 66) | θ value | 8.99809 (rank : 64) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P13611, P20754, Q13010, Q13189, Q15123, Q9UCL9, Q9UNW5 | Gene names | CSPG2 | |||
|
Domain Architecture |

|
|||||
| Description | Versican core protein precursor (Large fibroblast proteoglycan) (Chondroitin sulfate proteoglycan core protein 2) (PG-M) (Glial hyaluronate-binding protein) (GHAP). | |||||
|
ZEP2_MOUSE
|
||||||
| NC score | 0.001666 (rank : 67) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 67 | Target Neighborhood Hits | 964 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q3UHF7, O55140, Q3UVD4, Q3UVH5 | Gene names | Hivep2, Mibp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Human immunodeficiency virus type I enhancer-binding protein 2 homolog (Myc intron-binding protein 1) (MIBP-1). | |||||