Please be patient as the page loads
|
RED2_MOUSE
|
||||||
| SwissProt Accessions | Q9JI20, Q3UTP1, Q8CC51, Q9JIE5 | Gene names | Adarb2, Adar3, Red2 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase B2 (EC 3.5.-.-) (dsRNA adenosine deaminase B2) (RNA-dependent adenosine deaminase 3) (RNA-editing deaminase 2) (RNA-editing enzyme 2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
RED1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.974917 (rank : 4) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P78563, O00395, O00465, O00691, O00692, P78555, Q8NFD1 | Gene names | ADARB1, ADAR2, DRADA2, RED1 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase 1 (EC 3.5.-.-) (dsRNA adenosine deaminase) (RNA-editing deaminase 1) (RNA-editing enzyme 1). | |||||
|
RED1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.978357 (rank : 3) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q91ZS8, Q8K3X1, Q91ZS6, Q91ZS7, Q91ZS9, Q99MU8 | Gene names | Adarb1, Adar2, Red1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-specific editase 1 (EC 3.5.-.-) (dsRNA adenosine deaminase) (RNA-editing deaminase 1) (RNA-editing enzyme 1). | |||||
|
RED2_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.996381 (rank : 2) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NS39 | Gene names | ADARB2, ADAR3, RED2 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase B2 (EC 3.5.-.-) (dsRNA adenosine deaminase B2) (RNA-dependent adenosine deaminase 3) (RNA-editing deaminase 2) (RNA-editing enzyme 2). | |||||
|
RED2_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9JI20, Q3UTP1, Q8CC51, Q9JIE5 | Gene names | Adarb2, Adar3, Red2 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase B2 (EC 3.5.-.-) (dsRNA adenosine deaminase B2) (RNA-dependent adenosine deaminase 3) (RNA-editing deaminase 2) (RNA-editing enzyme 2). | |||||
|
DSRAD_HUMAN
|
||||||
| θ value | 2.84138e-62 (rank : 5) | NC score | 0.835816 (rank : 5) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P55265, O15223, O43859, O43860, Q9BYM3, Q9BYM4 | Gene names | ADAR, ADAR1, DSRAD, IFI4 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific adenosine deaminase (EC 3.5.4.-) (DRADA) (136 kDa double-stranded RNA-binding protein) (P136) (K88DSRBP) (Interferon-inducible protein 4) (IFI-4 protein). | |||||
|
DSRAD_MOUSE
|
||||||
| θ value | 6.99855e-61 (rank : 6) | NC score | 0.833387 (rank : 6) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99MU3, O70375, Q80UZ6, Q8C222, Q99MU2, Q99MU4, Q99MU7 | Gene names | Adar | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific adenosine deaminase (EC 3.5.4.-) (DRADA) (RNA adenosine deaminase 1). | |||||
|
ILF3_HUMAN
|
||||||
| θ value | 1.2077e-20 (rank : 7) | NC score | 0.635553 (rank : 7) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q12906, O43409, Q6P1X1, Q86XY7, Q99544, Q99545, Q9BZH4, Q9BZH5, Q9NQ95, Q9NQ96, Q9NQ97, Q9NQ98, Q9NQ99, Q9NQA0, Q9NQA1, Q9NQA2, Q9NRN2, Q9NRN3, Q9NRN4, Q9UMZ9, Q9UN00, Q9UN84, Q9UNA2 | Gene names | ILF3, DRBF, MPHOSPH4, NF90 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin enhancer-binding factor 3 (Nuclear factor of activated T- cells 90 kDa) (NF-AT-90) (Double-stranded RNA-binding protein 76) (DRBP76) (Translational control protein 80) (TCP80) (Nuclear factor associated with dsRNA) (NFAR) (M-phase phosphoprotein 4) (MPP4). | |||||
|
ILF3_MOUSE
|
||||||
| θ value | 4.58923e-20 (rank : 8) | NC score | 0.630853 (rank : 8) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Z1X4, Q80VD5, Q812A1, Q8BP80, Q8C2H8, Q8K588 | Gene names | Ilf3 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin enhancer-binding factor 3. | |||||
|
STAU2_MOUSE
|
||||||
| θ value | 8.97725e-08 (rank : 9) | NC score | 0.401568 (rank : 9) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CJ67, Q8BSY8, Q8CJ66, Q8R175, Q91Z19, Q9D5N7 | Gene names | Stau2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 2. | |||||
|
STAU2_HUMAN
|
||||||
| θ value | 2.21117e-06 (rank : 10) | NC score | 0.381134 (rank : 10) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NUL3, Q6AHY7, Q96HM0, Q96HM1, Q9NVI5, Q9UGG6 | Gene names | STAU2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 2. | |||||
|
STAU1_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 11) | NC score | 0.268206 (rank : 11) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O95793, Q6GTM4, Q9H5B4, Q9H5B5, Q9Y3Q2 | Gene names | STAU1, STAU | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 1. | |||||
|
PRKRA_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 12) | NC score | 0.233499 (rank : 16) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O75569, Q53G24, Q6X7T5, Q8NDK4 | Gene names | PRKRA, PACT, RAX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interferon-inducible double stranded RNA-dependent protein kinase activator A (Protein kinase, interferon-inducible double stranded RNA- dependent activator) (Protein activator of the interferon-induced protein kinase) (PKR-associated protein X) (PKR-associating protein X). | |||||
|
STAU1_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 13) | NC score | 0.265919 (rank : 12) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Z108 | Gene names | Stau1, Stau | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 1. | |||||
|
PRKRA_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 14) | NC score | 0.233655 (rank : 15) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9WTX2, Q9CZB7 | Gene names | Prkra, Rax | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interferon-inducible double stranded RNA-dependent protein kinase activator A (Protein kinase, interferon-inducible double stranded RNA- dependent activator) (Protein activator of the interferon-induced protein kinase) (PKR-associated protein X) (PKR-associating protein X) (RAX). | |||||
|
MED8_MOUSE
|
||||||
| θ value | 0.163984 (rank : 15) | NC score | 0.081833 (rank : 17) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D7W5, Q3UGB4, Q99LM4, Q9JJ84 | Gene names | Med8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of RNA polymerase II transcription subunit 8 homolog (Activator-recruited cofactor 32 kDa component) (ARC32). | |||||
|
POLS2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 16) | NC score | 0.009171 (rank : 28) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5K4E3, Q8NBY4 | Gene names | PRSS36 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyserase-2 precursor (EC 3.4.21.-) (Polyserine protease 2) (Protease serine 36). | |||||
|
POLS2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 17) | NC score | 0.009837 (rank : 27) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5K2P8 | Gene names | Prss36 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyserase-2 precursor (EC 3.4.21.-) (Polyserine protease 2) (Protease serine 36). | |||||
|
MED8_HUMAN
|
||||||
| θ value | 1.81305 (rank : 18) | NC score | 0.055838 (rank : 20) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96G25, Q96FQ4 | Gene names | MED8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of RNA polymerase II transcription subunit 8 homolog (Activator-recruited cofactor 32 kDa component) (ARC32). | |||||
|
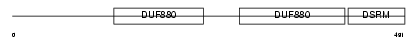
RNC_HUMAN
|
||||||
| θ value | 1.81305 (rank : 19) | NC score | 0.055819 (rank : 21) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NRR4, Q7Z5V2, Q86YH0, Q9NW73, Q9Y2V9, Q9Y4Y0 | Gene names | RNASEN, RN3, RNASE3L | |||
|
Domain Architecture |
|
|||||
| Description | Ribonuclease III (EC 3.1.26.3) (RNase III) (Drosha) (p241). | |||||
|
TATD2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 20) | NC score | 0.039776 (rank : 23) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q93075 | Gene names | TATDN2, KIAA0218 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TatD DNase domain-containing deoxyribonuclease 2 (EC 3.1.21.-). | |||||
|
GOLP3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 21) | NC score | 0.035633 (rank : 25) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H4A6, Q9UIW5 | Gene names | GOLPH3, GPP34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi phosphoprotein 3 (Coat-protein GPP34). | |||||
|
GOLP3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 22) | NC score | 0.035711 (rank : 24) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CRA5, Q99KY1, Q9D636 | Gene names | Golph3, Gpp34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi phosphoprotein 3 (Coat-protein GPP34). | |||||
|
MLTK_MOUSE
|
||||||
| θ value | 2.36792 (rank : 23) | NC score | 0.002586 (rank : 29) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ESL4, Q3V1X8, Q8BR73, Q9ESL3 | Gene names | Mltk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase MLT (EC 2.7.11.25) (MLK-like mitogen-activated protein triple kinase) (Leucine zipper- and sterile alpha motif kinase ZAK) (Sterile alpha motif- and leucine zipper-containing kinase AZK) (Mixed lineage kinase-related kinase) (MLK-related kinase) (MRK). | |||||
|
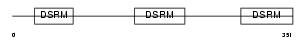
TRBP2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 24) | NC score | 0.253663 (rank : 13) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P97473 | Gene names | Tarbp2, Prbp | |||
|
Domain Architecture |
|
|||||
| Description | TAR RNA-binding protein 2 (Protamine-1 RNA-binding protein) (PRM-1 RNA-binding protein). | |||||
|
TRBP2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 25) | NC score | 0.238517 (rank : 14) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15633, Q12878 | Gene names | TARBP2, TRBP | |||
|
Domain Architecture |

|
|||||
| Description | TAR RNA-binding protein 2 (Trans-activation-responsive RNA-binding protein). | |||||
|
HTF9C_MOUSE
|
||||||
| θ value | 6.88961 (rank : 26) | NC score | 0.012751 (rank : 26) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BNV1, P70221, P70222, Q78E15, Q80VN8 | Gene names | Htf9c, Htf9-c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HpaII tiny fragments locus 9c protein. | |||||
|
ILF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 27) | NC score | 0.081652 (rank : 18) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q12905, Q5SR10, Q5SR11, Q7L7R3, Q9BWD4, Q9P1N0 | Gene names | ILF2, NF45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin enhancer-binding factor 2 (Nuclear factor of activated T- cells 45 kDa). | |||||
|
ILF2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 28) | NC score | 0.081652 (rank : 19) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CXY6, Q3U083, Q5RKG0, Q8CCY9, Q99KS3 | Gene names | Ilf2, Nf45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin enhancer-binding factor 2 (Nuclear factor of activated T- cells 45 kDa). | |||||
|
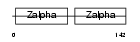
ZBP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 29) | NC score | 0.054446 (rank : 22) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QY24, Q8VE02, Q9D8B9 | Gene names | Zbp1, Dlm1 | |||
|
Domain Architecture |
|
|||||
| Description | Z-DNA-binding protein 1 (Tumor stroma and activated macrophage protein DLM-1). | |||||
|
RED2_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9JI20, Q3UTP1, Q8CC51, Q9JIE5 | Gene names | Adarb2, Adar3, Red2 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase B2 (EC 3.5.-.-) (dsRNA adenosine deaminase B2) (RNA-dependent adenosine deaminase 3) (RNA-editing deaminase 2) (RNA-editing enzyme 2). | |||||
|
RED2_HUMAN
|
||||||
| NC score | 0.996381 (rank : 2) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NS39 | Gene names | ADARB2, ADAR3, RED2 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase B2 (EC 3.5.-.-) (dsRNA adenosine deaminase B2) (RNA-dependent adenosine deaminase 3) (RNA-editing deaminase 2) (RNA-editing enzyme 2). | |||||
|
RED1_MOUSE
|
||||||
| NC score | 0.978357 (rank : 3) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q91ZS8, Q8K3X1, Q91ZS6, Q91ZS7, Q91ZS9, Q99MU8 | Gene names | Adarb1, Adar2, Red1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-specific editase 1 (EC 3.5.-.-) (dsRNA adenosine deaminase) (RNA-editing deaminase 1) (RNA-editing enzyme 1). | |||||
|
RED1_HUMAN
|
||||||
| NC score | 0.974917 (rank : 4) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P78563, O00395, O00465, O00691, O00692, P78555, Q8NFD1 | Gene names | ADARB1, ADAR2, DRADA2, RED1 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase 1 (EC 3.5.-.-) (dsRNA adenosine deaminase) (RNA-editing deaminase 1) (RNA-editing enzyme 1). | |||||
|
DSRAD_HUMAN
|
||||||
| NC score | 0.835816 (rank : 5) | θ value | 2.84138e-62 (rank : 5) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P55265, O15223, O43859, O43860, Q9BYM3, Q9BYM4 | Gene names | ADAR, ADAR1, DSRAD, IFI4 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific adenosine deaminase (EC 3.5.4.-) (DRADA) (136 kDa double-stranded RNA-binding protein) (P136) (K88DSRBP) (Interferon-inducible protein 4) (IFI-4 protein). | |||||
|
DSRAD_MOUSE
|
||||||
| NC score | 0.833387 (rank : 6) | θ value | 6.99855e-61 (rank : 6) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99MU3, O70375, Q80UZ6, Q8C222, Q99MU2, Q99MU4, Q99MU7 | Gene names | Adar | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific adenosine deaminase (EC 3.5.4.-) (DRADA) (RNA adenosine deaminase 1). | |||||
|
ILF3_HUMAN
|
||||||
| NC score | 0.635553 (rank : 7) | θ value | 1.2077e-20 (rank : 7) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q12906, O43409, Q6P1X1, Q86XY7, Q99544, Q99545, Q9BZH4, Q9BZH5, Q9NQ95, Q9NQ96, Q9NQ97, Q9NQ98, Q9NQ99, Q9NQA0, Q9NQA1, Q9NQA2, Q9NRN2, Q9NRN3, Q9NRN4, Q9UMZ9, Q9UN00, Q9UN84, Q9UNA2 | Gene names | ILF3, DRBF, MPHOSPH4, NF90 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin enhancer-binding factor 3 (Nuclear factor of activated T- cells 90 kDa) (NF-AT-90) (Double-stranded RNA-binding protein 76) (DRBP76) (Translational control protein 80) (TCP80) (Nuclear factor associated with dsRNA) (NFAR) (M-phase phosphoprotein 4) (MPP4). | |||||
|
ILF3_MOUSE
|
||||||
| NC score | 0.630853 (rank : 8) | θ value | 4.58923e-20 (rank : 8) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Z1X4, Q80VD5, Q812A1, Q8BP80, Q8C2H8, Q8K588 | Gene names | Ilf3 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin enhancer-binding factor 3. | |||||
|
STAU2_MOUSE
|
||||||
| NC score | 0.401568 (rank : 9) | θ value | 8.97725e-08 (rank : 9) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CJ67, Q8BSY8, Q8CJ66, Q8R175, Q91Z19, Q9D5N7 | Gene names | Stau2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 2. | |||||
|
STAU2_HUMAN
|
||||||
| NC score | 0.381134 (rank : 10) | θ value | 2.21117e-06 (rank : 10) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NUL3, Q6AHY7, Q96HM0, Q96HM1, Q9NVI5, Q9UGG6 | Gene names | STAU2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 2. | |||||
|
STAU1_HUMAN
|
||||||
| NC score | 0.268206 (rank : 11) | θ value | 0.0252991 (rank : 11) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O95793, Q6GTM4, Q9H5B4, Q9H5B5, Q9Y3Q2 | Gene names | STAU1, STAU | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 1. | |||||
|
STAU1_MOUSE
|
||||||
| NC score | 0.265919 (rank : 12) | θ value | 0.0736092 (rank : 13) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Z108 | Gene names | Stau1, Stau | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-binding protein Staufen homolog 1. | |||||
|
TRBP2_MOUSE
|
||||||
| NC score | 0.253663 (rank : 13) | θ value | 3.0926 (rank : 24) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P97473 | Gene names | Tarbp2, Prbp | |||
|
Domain Architecture |

|
|||||
| Description | TAR RNA-binding protein 2 (Protamine-1 RNA-binding protein) (PRM-1 RNA-binding protein). | |||||
|
TRBP2_HUMAN
|
||||||
| NC score | 0.238517 (rank : 14) | θ value | 4.03905 (rank : 25) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15633, Q12878 | Gene names | TARBP2, TRBP | |||
|
Domain Architecture |

|
|||||
| Description | TAR RNA-binding protein 2 (Trans-activation-responsive RNA-binding protein). | |||||
|
PRKRA_MOUSE
|
||||||
| NC score | 0.233655 (rank : 15) | θ value | 0.0961366 (rank : 14) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9WTX2, Q9CZB7 | Gene names | Prkra, Rax | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interferon-inducible double stranded RNA-dependent protein kinase activator A (Protein kinase, interferon-inducible double stranded RNA- dependent activator) (Protein activator of the interferon-induced protein kinase) (PKR-associated protein X) (PKR-associating protein X) (RAX). | |||||
|
PRKRA_HUMAN
|
||||||
| NC score | 0.233499 (rank : 16) | θ value | 0.0563607 (rank : 12) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O75569, Q53G24, Q6X7T5, Q8NDK4 | Gene names | PRKRA, PACT, RAX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interferon-inducible double stranded RNA-dependent protein kinase activator A (Protein kinase, interferon-inducible double stranded RNA- dependent activator) (Protein activator of the interferon-induced protein kinase) (PKR-associated protein X) (PKR-associating protein X). | |||||
|
MED8_MOUSE
|
||||||
| NC score | 0.081833 (rank : 17) | θ value | 0.163984 (rank : 15) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D7W5, Q3UGB4, Q99LM4, Q9JJ84 | Gene names | Med8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of RNA polymerase II transcription subunit 8 homolog (Activator-recruited cofactor 32 kDa component) (ARC32). | |||||
|
ILF2_HUMAN
|
||||||
| NC score | 0.081652 (rank : 18) | θ value | θ > 10 (rank : 27) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q12905, Q5SR10, Q5SR11, Q7L7R3, Q9BWD4, Q9P1N0 | Gene names | ILF2, NF45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin enhancer-binding factor 2 (Nuclear factor of activated T- cells 45 kDa). | |||||
|
ILF2_MOUSE
|
||||||
| NC score | 0.081652 (rank : 19) | θ value | θ > 10 (rank : 28) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CXY6, Q3U083, Q5RKG0, Q8CCY9, Q99KS3 | Gene names | Ilf2, Nf45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin enhancer-binding factor 2 (Nuclear factor of activated T- cells 45 kDa). | |||||
|
MED8_HUMAN
|
||||||
| NC score | 0.055838 (rank : 20) | θ value | 1.81305 (rank : 18) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96G25, Q96FQ4 | Gene names | MED8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of RNA polymerase II transcription subunit 8 homolog (Activator-recruited cofactor 32 kDa component) (ARC32). | |||||
|
RNC_HUMAN
|
||||||
| NC score | 0.055819 (rank : 21) | θ value | 1.81305 (rank : 19) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NRR4, Q7Z5V2, Q86YH0, Q9NW73, Q9Y2V9, Q9Y4Y0 | Gene names | RNASEN, RN3, RNASE3L | |||
|
Domain Architecture |

|
|||||
| Description | Ribonuclease III (EC 3.1.26.3) (RNase III) (Drosha) (p241). | |||||
|
ZBP1_MOUSE
|
||||||
| NC score | 0.054446 (rank : 22) | θ value | θ > 10 (rank : 29) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QY24, Q8VE02, Q9D8B9 | Gene names | Zbp1, Dlm1 | |||
|
Domain Architecture |

|
|||||
| Description | Z-DNA-binding protein 1 (Tumor stroma and activated macrophage protein DLM-1). | |||||
|
TATD2_HUMAN
|
||||||
| NC score | 0.039776 (rank : 23) | θ value | 1.81305 (rank : 20) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q93075 | Gene names | TATDN2, KIAA0218 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TatD DNase domain-containing deoxyribonuclease 2 (EC 3.1.21.-). | |||||
|
GOLP3_MOUSE
|
||||||
| NC score | 0.035711 (rank : 24) | θ value | 2.36792 (rank : 22) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CRA5, Q99KY1, Q9D636 | Gene names | Golph3, Gpp34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi phosphoprotein 3 (Coat-protein GPP34). | |||||
|
GOLP3_HUMAN
|
||||||
| NC score | 0.035633 (rank : 25) | θ value | 2.36792 (rank : 21) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H4A6, Q9UIW5 | Gene names | GOLPH3, GPP34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi phosphoprotein 3 (Coat-protein GPP34). | |||||
|
HTF9C_MOUSE
|
||||||
| NC score | 0.012751 (rank : 26) | θ value | 6.88961 (rank : 26) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BNV1, P70221, P70222, Q78E15, Q80VN8 | Gene names | Htf9c, Htf9-c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HpaII tiny fragments locus 9c protein. | |||||
|
POLS2_MOUSE
|
||||||
| NC score | 0.009837 (rank : 27) | θ value | 1.38821 (rank : 17) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5K2P8 | Gene names | Prss36 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyserase-2 precursor (EC 3.4.21.-) (Polyserine protease 2) (Protease serine 36). | |||||
|
POLS2_HUMAN
|
||||||
| NC score | 0.009171 (rank : 28) | θ value | 1.06291 (rank : 16) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5K4E3, Q8NBY4 | Gene names | PRSS36 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyserase-2 precursor (EC 3.4.21.-) (Polyserine protease 2) (Protease serine 36). | |||||
|
MLTK_MOUSE
|
||||||
| NC score | 0.002586 (rank : 29) | θ value | 2.36792 (rank : 23) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ESL4, Q3V1X8, Q8BR73, Q9ESL3 | Gene names | Mltk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase MLT (EC 2.7.11.25) (MLK-like mitogen-activated protein triple kinase) (Leucine zipper- and sterile alpha motif kinase ZAK) (Sterile alpha motif- and leucine zipper-containing kinase AZK) (Mixed lineage kinase-related kinase) (MLK-related kinase) (MRK). | |||||