Please be patient as the page loads
|
AT2B3_HUMAN
|
||||||
| SwissProt Accessions | Q16720, Q12995, Q16858 | Gene names | ATP2B3 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 3 (EC 3.6.3.8) (PMCA3) (Plasma membrane calcium pump isoform 3) (Plasma membrane calcium ATPase isoform 3). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
AT2B1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.997244 (rank : 5) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P20020, Q12992, Q12993, Q13819, Q13820, Q13821, Q16504, Q93082 | Gene names | ATP2B1, PMCA1 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 1 (EC 3.6.3.8) (PMCA1) (Plasma membrane calcium pump isoform 1) (Plasma membrane calcium ATPase isoform 1). | |||||
|
AT2B2_HUMAN
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.998302 (rank : 3) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q01814, O00766, Q12994, Q16818 | Gene names | ATP2B2, PMCA2 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 2 (EC 3.6.3.8) (PMCA2) (Plasma membrane calcium pump isoform 2) (Plasma membrane calcium ATPase isoform 2). | |||||
|
AT2B2_MOUSE
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.998586 (rank : 2) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9R0K7, O88863 | Gene names | Atp2b2, Pmca2 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 2 (EC 3.6.3.8) (PMCA2) (Plasma membrane calcium pump isoform 2) (Plasma membrane calcium ATPase isoform 2). | |||||
|
AT2B3_HUMAN
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q16720, Q12995, Q16858 | Gene names | ATP2B3 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 3 (EC 3.6.3.8) (PMCA3) (Plasma membrane calcium pump isoform 3) (Plasma membrane calcium ATPase isoform 3). | |||||
|
AT2B4_HUMAN
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.997906 (rank : 4) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P23634, Q13450, Q13452, Q13455, Q16817 | Gene names | ATP2B4 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 4 (EC 3.6.3.8) (PMCA4) (Plasma membrane calcium pump isoform 4) (Plasma membrane calcium ATPase isoform 4). | |||||
|
AT2A3_MOUSE
|
||||||
| θ value | 5.69867e-79 (rank : 6) | NC score | 0.905536 (rank : 11) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q64518, O70625, Q64517 | Gene names | Atp2a3 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 3 (EC 3.6.3.8) (Calcium pump 3) (SERCA3) (SR Ca(2+)-ATPase 3). | |||||
|
AT2A3_HUMAN
|
||||||
| θ value | 9.7205e-79 (rank : 7) | NC score | 0.907417 (rank : 8) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q93084, O60900, O60901, O75501, O75502, Q16115, Q8TEX5, Q8TEX6 | Gene names | ATP2A3 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 3 (EC 3.6.3.8) (Calcium pump 3) (SERCA3) (SR Ca(2+)-ATPase 3). | |||||
|
AT2A2_MOUSE
|
||||||
| θ value | 2.02684e-76 (rank : 8) | NC score | 0.906864 (rank : 9) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O55143, Q9R2A9, Q9WUT5 | Gene names | Atp2a2 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 2 (EC 3.6.3.8) (Calcium pump 2) (SERCA2) (SR Ca(2+)-ATPase 2) (Calcium-transporting ATPase sarcoplasmic reticulum type, slow twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2A2_HUMAN
|
||||||
| θ value | 4.51533e-76 (rank : 9) | NC score | 0.906790 (rank : 10) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P16615, P16614 | Gene names | ATP2A2, ATP2B | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 2 (EC 3.6.3.8) (Calcium pump 2) (SERCA2) (SR Ca(2+)-ATPase 2) (Calcium-transporting ATPase sarcoplasmic reticulum type, slow twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2C1_MOUSE
|
||||||
| θ value | 1.45253e-74 (rank : 10) | NC score | 0.915729 (rank : 6) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q80XR2, Q80YZ2 | Gene names | Atp2c1, Pmr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-transporting ATPase type 2C member 1 (EC 3.6.3.8) (ATPase 2C1) (ATP-dependent Ca(2+) pump PMR1). | |||||
|
AT2A1_HUMAN
|
||||||
| θ value | 2.09745e-73 (rank : 11) | NC score | 0.903066 (rank : 13) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O14983, O14984 | Gene names | ATP2A1 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 1 (EC 3.6.3.8) (Calcium pump 1) (SERCA1) (SR Ca(2+)-ATPase 1) (Calcium-transporting ATPase sarcoplasmic reticulum type, fast twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2C1_HUMAN
|
||||||
| θ value | 1.04096e-72 (rank : 12) | NC score | 0.915196 (rank : 7) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P98194, O76005, Q86V72, Q86V73, Q8N6V1, Q8NCJ7 | Gene names | ATP2C1, KIAA1347, PMR1L | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-transporting ATPase type 2C member 1 (EC 3.6.3.8) (ATPase 2C1) (ATP-dependent Ca(2+) pump PMR1). | |||||
|
AT2A1_MOUSE
|
||||||
| θ value | 1.7756e-72 (rank : 13) | NC score | 0.903249 (rank : 12) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8R429 | Gene names | Atp2a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 1 (EC 3.6.3.8) (Calcium pump 1) (SERCA1) (SR Ca(2+)-ATPase 1) (Calcium-transporting ATPase sarcoplasmic reticulum type, fast twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT1A3_HUMAN
|
||||||
| θ value | 2.24614e-67 (rank : 14) | NC score | 0.869044 (rank : 25) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P13637, Q16732, Q16735, Q969K5 | Gene names | ATP1A3 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-3 chain (EC 3.6.3.9) (Sodium pump 3) (Na+/K+ ATPase 3) (Alpha(III)). | |||||
|
AT1A3_MOUSE
|
||||||
| θ value | 2.93355e-67 (rank : 15) | NC score | 0.868395 (rank : 26) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q6PIC6 | Gene names | Atp1a3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium/potassium-transporting ATPase alpha-3 chain (EC 3.6.3.9) (Sodium pump 3) (Na+/K+ ATPase 3) (Alpha(III)). | |||||
|
AT1A2_MOUSE
|
||||||
| θ value | 6.53529e-67 (rank : 16) | NC score | 0.870742 (rank : 23) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q6PIE5 | Gene names | Atp1a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2) (Alpha(+)). | |||||
|
AT1A4_MOUSE
|
||||||
| θ value | 6.53529e-67 (rank : 17) | NC score | 0.878222 (rank : 18) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9WV27, Q9R173, Q9WV28 | Gene names | Atp1a4, Atp1al2 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-4 chain (EC 3.6.3.9) (Sodium pump 4) (Na+/K+ ATPase 4). | |||||
|
AT12A_HUMAN
|
||||||
| θ value | 2.10232e-65 (rank : 18) | NC score | 0.886648 (rank : 15) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P54707, Q13816, Q13817, Q16734 | Gene names | ATP12A, ATP1AL1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 2 (EC 3.6.3.10) (Proton pump) (Non-gastric H(+)/K(+) ATPase subunit alpha). | |||||
|
AT12A_MOUSE
|
||||||
| θ value | 8.83269e-64 (rank : 19) | NC score | 0.885443 (rank : 16) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9Z1W8, Q8VHY2 | Gene names | Atp12a, Atp1al1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium-transporting ATPase alpha chain 2 (EC 3.6.3.10) (Proton pump) (Non-gastric H(+)/K(+) ATPase subunit alpha). | |||||
|
AT2C2_HUMAN
|
||||||
| θ value | 1.50663e-63 (rank : 20) | NC score | 0.902341 (rank : 14) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O75185 | Gene names | KIAA0703 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable calcium-transporting ATPase KIAA0703 (EC 3.6.3.8). | |||||
|
AT1A1_HUMAN
|
||||||
| θ value | 9.1404e-61 (rank : 21) | NC score | 0.873125 (rank : 22) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P05023, Q16689, Q6LDM4, Q9UJ20, Q9UJ21 | Gene names | ATP1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-1 chain precursor (EC 3.6.3.9) (Sodium pump 1) (Na+/K+ ATPase 1). | |||||
|
AT1A1_MOUSE
|
||||||
| θ value | 1.19377e-60 (rank : 22) | NC score | 0.873907 (rank : 20) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8VDN2, Q91Z09 | Gene names | Atp1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-1 chain precursor (EC 3.6.3.9) (Sodium pump 1) (Na+/K+ ATPase 1). | |||||
|
AT1A2_HUMAN
|
||||||
| θ value | 2.65945e-60 (rank : 23) | NC score | 0.869948 (rank : 24) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P50993, Q07059, Q86UZ5, Q9UQ25 | Gene names | ATP1A2, KIAA0778 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2). | |||||
|
AT1A4_HUMAN
|
||||||
| θ value | 5.92465e-60 (rank : 24) | NC score | 0.873307 (rank : 21) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q13733, Q504T2, Q8TBN8, Q8WXA7, Q8WXH7, Q8WY13 | Gene names | ATP1A4, ATP1AL2 | |||
|
Domain Architecture |
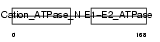
|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-4 chain (EC 3.6.3.9) (Sodium pump 4) (Na+/K+ ATPase 4). | |||||
|
ATP4A_MOUSE
|
||||||
| θ value | 4.24587e-58 (rank : 25) | NC score | 0.878597 (rank : 17) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q64436, Q9CV46 | Gene names | Atp4a | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 1 (EC 3.6.3.10) (Proton pump) (Gastric H+/K+ ATPase subunit alpha). | |||||
|
ATP4A_HUMAN
|
||||||
| θ value | 1.23536e-57 (rank : 26) | NC score | 0.876719 (rank : 19) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P20648, O00738 | Gene names | ATP4A | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 1 (EC 3.6.3.10) (Proton pump) (Gastric H+/K+ ATPase subunit alpha). | |||||
|
AT11A_HUMAN
|
||||||
| θ value | 9.24701e-21 (rank : 27) | NC score | 0.535980 (rank : 45) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98196, Q5VXT2 | Gene names | ATP11A, ATPIH, ATPIS, KIAA1021 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IH (EC 3.6.3.1) (ATPase class I type 11A) (ATPase IS). | |||||
|
AT8A2_MOUSE
|
||||||
| θ value | 4.58923e-20 (rank : 28) | NC score | 0.566585 (rank : 37) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98200 | Gene names | Atp8a2 | |||
|
Domain Architecture |
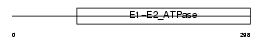
|
|||||
| Description | Probable phospholipid-transporting ATPase IB (EC 3.6.3.1) (ATPase class I type 8A member 2). | |||||
|
AT132_HUMAN
|
||||||
| θ value | 6.62687e-19 (rank : 29) | NC score | 0.717560 (rank : 29) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9NQ11, O75700, Q5JXY2 | Gene names | ATP13A2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A2 (EC 3.6.3.-). | |||||
|
AT8A2_HUMAN
|
||||||
| θ value | 1.13037e-18 (rank : 30) | NC score | 0.566588 (rank : 36) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9NTI2, Q9H527, Q9NPU6, Q9NTL2, Q9NYM3 | Gene names | ATP8A2, ATPIB | |||
|
Domain Architecture |
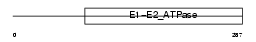
|
|||||
| Description | Probable phospholipid-transporting ATPase IB (EC 3.6.3.1) (ATPase class I type 8A member 2) (ML-1). | |||||
|
AT8B3_HUMAN
|
||||||
| θ value | 4.29542e-18 (rank : 31) | NC score | 0.524429 (rank : 50) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O60423, Q8IVB8, Q8N4Y8, Q96M22 | Gene names | ATP8B3, ATP1K, FOS37502_2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IK (EC 3.6.3.1) (ATPase class I type 8B member 3). | |||||
|
AT8A1_MOUSE
|
||||||
| θ value | 7.32683e-18 (rank : 32) | NC score | 0.559892 (rank : 39) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P70704 | Gene names | Atp8a1, Atpc1 | |||
|
Domain Architecture |
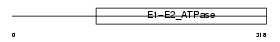
|
|||||
| Description | Probable phospholipid-transporting ATPase IA (EC 3.6.3.1) (Chromaffin granule ATPase II) (ATPase class I type 8A member 1). | |||||
|
AT8A1_HUMAN
|
||||||
| θ value | 2.13179e-17 (rank : 33) | NC score | 0.556597 (rank : 40) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Y2Q0 | Gene names | ATP8A1, ATPIA | |||
|
Domain Architecture |
|
|||||
| Description | Probable phospholipid-transporting ATPase IA (EC 3.6.3.1) (Chromaffin granule ATPase II) (ATPase class I type 8A member 1). | |||||
|
ATP7A_HUMAN
|
||||||
| θ value | 3.63628e-17 (rank : 34) | NC score | 0.612185 (rank : 32) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q04656, O00227, O00745, Q9BYY8 | Gene names | ATP7A, MC1, MNK | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 1 (EC 3.6.3.4) (Copper pump 1) (Menkes disease-associated protein). | |||||
|
AT11A_MOUSE
|
||||||
| θ value | 8.10077e-17 (rank : 35) | NC score | 0.519812 (rank : 53) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98197 | Gene names | Atp11a | |||
|
Domain Architecture |
|
|||||
| Description | Probable phospholipid-transporting ATPase IH (EC 3.6.3.1) (ATPase class I type 11A) (ATPase IS). | |||||
|
AT11B_HUMAN
|
||||||
| θ value | 1.058e-16 (rank : 36) | NC score | 0.562584 (rank : 38) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9Y2G3, Q96FN1, Q9UKK7 | Gene names | ATP11B, ATPIF, ATPIR, KIAA0956 | |||
|
Domain Architecture |
|
|||||
| Description | Probable phospholipid-transporting ATPase IF (EC 3.6.3.1) (ATPase class I type 11B) (ATPase IR). | |||||
|
AT132_MOUSE
|
||||||
| θ value | 1.38178e-16 (rank : 37) | NC score | 0.725190 (rank : 27) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9CTG6, Q8CG98 | Gene names | Atp13a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable cation-transporting ATPase 13A2 (EC 3.6.3.-). | |||||
|
AT10B_HUMAN
|
||||||
| θ value | 1.16975e-15 (rank : 38) | NC score | 0.530326 (rank : 49) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O94823, Q9H725 | Gene names | ATP10B, ATPVB, KIAA0715 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VB (EC 3.6.3.1). | |||||
|
AT10D_HUMAN
|
||||||
| θ value | 7.58209e-15 (rank : 39) | NC score | 0.534131 (rank : 46) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9P241, Q8NC70, Q96SR3 | Gene names | ATP10D, KIAA1487 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VD (EC 3.6.3.1) (ATPVD). | |||||
|
ATP7B_MOUSE
|
||||||
| θ value | 1.68911e-14 (rank : 40) | NC score | 0.608563 (rank : 34) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q64446 | Gene names | Atp7b, Wnd | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 2 (EC 3.6.3.4) (Copper pump 2) (Wilson disease-associated protein homolog). | |||||
|
ATP7A_MOUSE
|
||||||
| θ value | 2.20605e-14 (rank : 41) | NC score | 0.608701 (rank : 33) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q64430, O35101, P97422, Q64431 | Gene names | Atp7a, Mnk | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 1 (EC 3.6.3.4) (Copper pump 1) (Menkes disease-associated protein homolog). | |||||
|
AT8B1_HUMAN
|
||||||
| θ value | 1.09485e-13 (rank : 42) | NC score | 0.521251 (rank : 52) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O43520, Q9BTP8 | Gene names | ATP8B1, ATPIC, FIC1, PFIC | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IC (EC 3.6.3.1) (Familial intrahepatic cholestasis type 1) (ATPase class I type 8B member 1). | |||||
|
AT10D_MOUSE
|
||||||
| θ value | 1.42992e-13 (rank : 43) | NC score | 0.537253 (rank : 44) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8K2X1 | Gene names | Atp10d | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VD (EC 3.6.3.1) (ATPVD). | |||||
|
AT11C_HUMAN
|
||||||
| θ value | 2.43908e-13 (rank : 44) | NC score | 0.531185 (rank : 48) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8NB49, Q5JT69, Q5JT70, Q5JT71, Q5JT72, Q5JT73, Q6ZND5, Q6ZU50, Q6ZUP7, Q70IJ9, Q70IK0, Q8WX24 | Gene names | ATP11C, ATPIG, ATPIQ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IG (EC 3.6.3.1) (ATPase class I type 11C) (ATPase IG) (ATPase IQ) (ATPase class VI type 11C). | |||||
|
AT8B2_HUMAN
|
||||||
| θ value | 3.18553e-13 (rank : 45) | NC score | 0.516418 (rank : 56) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P98198, Q96I43, Q96NQ7 | Gene names | ATP8B2, ATPID, KIAA1137 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase ID (EC 3.6.3.1) (ATPase class I type 8B member 2). | |||||
|
ATP7B_HUMAN
|
||||||
| θ value | 4.16044e-13 (rank : 46) | NC score | 0.603213 (rank : 35) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P35670, Q16318, Q16319, Q4U3V3, Q59FJ9, Q5T7X7 | Gene names | ATP7B, PWD, WC1, WND | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 2 (EC 3.6.3.4) (Copper pump 2) (Wilson disease-associated protein). | |||||
|
AT10A_HUMAN
|
||||||
| θ value | 5.43371e-13 (rank : 47) | NC score | 0.517586 (rank : 54) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O60312, Q969I4 | Gene names | ATP10A, ATP10C, ATPVC, KIAA0566 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (ATPVA) (Aminophospholipid translocase VA). | |||||
|
AT8B4_HUMAN
|
||||||
| θ value | 1.58096e-12 (rank : 48) | NC score | 0.522640 (rank : 51) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8TF62, Q9H727 | Gene names | ATP8B4, KIAA1939 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IM (EC 3.6.3.1) (ATPase class I type 8B member 4). | |||||
|
AT10A_MOUSE
|
||||||
| θ value | 2.0648e-12 (rank : 49) | NC score | 0.516906 (rank : 55) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O54827, Q8R3B8 | Gene names | Atp10a, Atpc5, Pfatp | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (P-locus fat-associated ATPase). | |||||
|
AT131_HUMAN
|
||||||
| θ value | 7.84624e-12 (rank : 50) | NC score | 0.698165 (rank : 30) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9HD20, Q9H6C6 | Gene names | ATP13A1, ATP13A, KIAA1825 | |||
|
Domain Architecture |
|
|||||
| Description | Probable cation-transporting ATPase 13A1 (EC 3.6.3.-). | |||||
|
AT133_HUMAN
|
||||||
| θ value | 2.28291e-11 (rank : 51) | NC score | 0.722271 (rank : 28) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9H7F0, Q8NC11, Q96KS1 | Gene names | ATP13A3, AFURS1 | |||
|
Domain Architecture |
|
|||||
| Description | Probable cation-transporting ATPase 13A3 (EC 3.6.3.-) (ATPase family homolog up-regulated in senescence cells 1). | |||||
|
ATP9A_HUMAN
|
||||||
| θ value | 9.59137e-10 (rank : 52) | NC score | 0.555209 (rank : 41) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O75110, Q5TFW5, Q5TFW6, Q5TFW9, Q6ZMF3, Q9NQK6, Q9NQK7 | Gene names | ATP9A, ATPIIA, KIAA0611 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIA (EC 3.6.3.1) (ATPase class II type 9A) (ATPase IIA). | |||||
|
AT131_MOUSE
|
||||||
| θ value | 1.25267e-09 (rank : 53) | NC score | 0.687338 (rank : 31) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9EPE9 | Gene names | Atp13a1, Atp13a | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A1 (EC 3.6.3.-) (CATP). | |||||
|
ATP9B_HUMAN
|
||||||
| θ value | 4.0297e-08 (rank : 54) | NC score | 0.542523 (rank : 43) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O43861, O60872 | Gene names | ATP9B, ATPIIB, NEO1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIB (EC 3.6.3.1). | |||||
|
ATP9B_MOUSE
|
||||||
| θ value | 5.26297e-08 (rank : 55) | NC score | 0.533351 (rank : 47) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98195, Q99LI3 | Gene names | Atp9b | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IIB (EC 3.6.3.1). | |||||
|
ATP9A_MOUSE
|
||||||
| θ value | 6.87365e-08 (rank : 56) | NC score | 0.543910 (rank : 42) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O70228, Q8VDI5, Q922L9 | Gene names | Atp9a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIA (EC 3.6.3.1) (ATPase class II type 9A). | |||||
|
RRBP1_MOUSE
|
||||||
| θ value | 0.163984 (rank : 57) | NC score | 0.006898 (rank : 71) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
AHI1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 58) | NC score | 0.016741 (rank : 61) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 537 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8N157, Q504T3, Q5TCP9, Q6P098, Q6PIT6, Q8NDX0, Q9H0H2 | Gene names | AHI1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Jouberin (Abelson helper integration site 1 protein homolog) (AHI-1). | |||||
|
K0195_HUMAN
|
||||||
| θ value | 0.62314 (rank : 59) | NC score | 0.072713 (rank : 58) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q12767, O75536, Q86XF1 | Gene names | KIAA0195, TMEM94 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0195 (Transmembrane protein 94). | |||||
|
K0195_MOUSE
|
||||||
| θ value | 0.62314 (rank : 60) | NC score | 0.072844 (rank : 57) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TSH8, Q80U66, Q8BL05, Q8K0Q0, Q91VY1 | Gene names | Kiaa0195 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0195. | |||||
|
KIF3A_HUMAN
|
||||||
| θ value | 0.813845 (rank : 61) | NC score | 0.002887 (rank : 76) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y496, Q86XE9, Q9Y6V4 | Gene names | KIF3A, KIF3 | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin-like protein KIF3A (Microtubule plus end-directed kinesin motor 3A). | |||||
|
RRBP1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 62) | NC score | 0.004639 (rank : 72) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1306 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9P2E9, O75300, O75301, Q5W165, Q96SB2, Q9BWP1, Q9H476 | Gene names | RRBP1, KIAA1398 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (180 kDa ribosome receptor homolog) (ES/130-related protein). | |||||
|
KIF3A_MOUSE
|
||||||
| θ value | 1.81305 (rank : 63) | NC score | 0.002348 (rank : 77) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 715 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P28741 | Gene names | Kif3a, Kif3 | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin-like protein KIF3A (Microtubule plus end-directed kinesin motor 3A). | |||||
|
ERIC1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 64) | NC score | 0.007701 (rank : 69) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
NUD15_MOUSE
|
||||||
| θ value | 2.36792 (rank : 65) | NC score | 0.017147 (rank : 60) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BG93 | Gene names | Nudt15, Mth2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable 7,8-dihydro-8-oxoguanine triphosphatase NUDT15 (EC 3.1.6.-) (8-oxo-dGTPase NUDT15) (Nucleoside diphosphate-linked moiety X motif 15) (Nudix motif 15) (MutT homolog 2) (mMTH2). | |||||
|
UBP25_HUMAN
|
||||||
| θ value | 2.36792 (rank : 66) | NC score | 0.013402 (rank : 62) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UHP3, Q9H9W1 | Gene names | USP25, USP21 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 25 (EC 3.1.2.15) (Ubiquitin thioesterase 25) (Ubiquitin-specific-processing protease 25) (Deubiquitinating enzyme 25) (USP on chromosome 21). | |||||
|
LRCH3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 67) | NC score | 0.002937 (rank : 74) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96II8, Q96FP9, Q9NT52 | Gene names | LRCH3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 3 precursor. | |||||
|
PVRL2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 68) | NC score | 0.021866 (rank : 59) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92692, O75455, Q96J29 | Gene names | PVRL2, HVEB, PRR2 | |||
|
Domain Architecture |
|
|||||
| Description | Poliovirus receptor-related protein 2 precursor (Herpes virus entry mediator B) (HveB) (Nectin-2) (CD112 antigen). | |||||
|
TAF1C_MOUSE
|
||||||
| θ value | 4.03905 (rank : 69) | NC score | 0.009139 (rank : 67) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6PDZ2, P97359, Q3U3B6, Q8BN54 | Gene names | Taf1c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA box-binding protein-associated factor, RNA polymerase I, subunit C (TATA box-binding protein-associated factor 1C) (TBP-associated factor 1C) (TBP-associated factor RNA polymerase I 95 kDa) (TAFI95). | |||||
|
UBP28_HUMAN
|
||||||
| θ value | 4.03905 (rank : 70) | NC score | 0.012362 (rank : 63) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96RU2, Q9P213 | Gene names | USP28, KIAA1515 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 28 (EC 3.1.2.15) (Ubiquitin thioesterase 28) (Ubiquitin-specific-processing protease 28) (Deubiquitinating enzyme 28). | |||||
|
RPC62_MOUSE
|
||||||
| θ value | 5.27518 (rank : 71) | NC score | 0.008310 (rank : 68) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D483, Q8R0I2 | Gene names | Polr3c, Rpc62 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase III 62 kDa polypeptide (EC 2.7.7.6) (RNA polymerase C subunit 3) (RPC3) (RNA polymerase III C62 subunit). | |||||
|
TEX14_HUMAN
|
||||||
| θ value | 5.27518 (rank : 72) | NC score | -0.003208 (rank : 79) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1103 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IWB6, Q7RTP3, Q8ND97, Q9BXT9 | Gene names | TEX14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed protein 14 (Testis-expressed sequence 14). | |||||
|
UBP25_MOUSE
|
||||||
| θ value | 5.27518 (rank : 73) | NC score | 0.012224 (rank : 64) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P57080, Q80ZT9 | Gene names | Usp25 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 25 (EC 3.1.2.15) (Ubiquitin thioesterase 25) (Ubiquitin-specific-processing protease 25) (Deubiquitinating enzyme 25) (mUSP25). | |||||
|
LRCH3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.002263 (rank : 78) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BVU0, Q3U222, Q3UZ74 | Gene names | Lrch3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 3 precursor. | |||||
|
PRAM3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.002925 (rank : 75) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5TYW8 | Gene names | PRAMEF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PRAME family member 3. | |||||
|
UBP28_MOUSE
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.011142 (rank : 66) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5I043, Q6NZP3, Q6ZPP1, Q8BWI1 | Gene names | Usp28, Kiaa1515 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 28 (EC 3.1.2.15) (Ubiquitin thioesterase 28) (Ubiquitin-specific-processing protease 28) (Deubiquitinating enzyme 28). | |||||
|
NAL13_HUMAN
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.007492 (rank : 70) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86W25, Q7RTR5 | Gene names | NALP13, NOD14 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 13 (Nucleotide-binding oligomerization domain protein 14). | |||||
|
NUD15_HUMAN
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.012171 (rank : 65) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NV35, Q6P2C9 | Gene names | NUDT15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable 7,8-dihydro-8-oxoguanine triphosphatase NUDT15 (EC 3.1.6.-) (8-oxo-dGTPase NUDT15) (Nucleoside diphosphate-linked moiety X motif 15) (Nudix motif 15). | |||||
|
PRGC1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.003090 (rank : 73) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O70343 | Gene names | Ppargc1a, Pgc1, Pgc1a, Ppargc1 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PPAR gamma coactivator 1-alpha) (PPARGC-1-alpha) (PGC-1-alpha). | |||||
|
AT2B3_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q16720, Q12995, Q16858 | Gene names | ATP2B3 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 3 (EC 3.6.3.8) (PMCA3) (Plasma membrane calcium pump isoform 3) (Plasma membrane calcium ATPase isoform 3). | |||||
|
AT2B2_MOUSE
|
||||||
| NC score | 0.998586 (rank : 2) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9R0K7, O88863 | Gene names | Atp2b2, Pmca2 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 2 (EC 3.6.3.8) (PMCA2) (Plasma membrane calcium pump isoform 2) (Plasma membrane calcium ATPase isoform 2). | |||||
|
AT2B2_HUMAN
|
||||||
| NC score | 0.998302 (rank : 3) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q01814, O00766, Q12994, Q16818 | Gene names | ATP2B2, PMCA2 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 2 (EC 3.6.3.8) (PMCA2) (Plasma membrane calcium pump isoform 2) (Plasma membrane calcium ATPase isoform 2). | |||||
|
AT2B4_HUMAN
|
||||||
| NC score | 0.997906 (rank : 4) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P23634, Q13450, Q13452, Q13455, Q16817 | Gene names | ATP2B4 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 4 (EC 3.6.3.8) (PMCA4) (Plasma membrane calcium pump isoform 4) (Plasma membrane calcium ATPase isoform 4). | |||||
|
AT2B1_HUMAN
|
||||||
| NC score | 0.997244 (rank : 5) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P20020, Q12992, Q12993, Q13819, Q13820, Q13821, Q16504, Q93082 | Gene names | ATP2B1, PMCA1 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 1 (EC 3.6.3.8) (PMCA1) (Plasma membrane calcium pump isoform 1) (Plasma membrane calcium ATPase isoform 1). | |||||
|
AT2C1_MOUSE
|
||||||
| NC score | 0.915729 (rank : 6) | θ value | 1.45253e-74 (rank : 10) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q80XR2, Q80YZ2 | Gene names | Atp2c1, Pmr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-transporting ATPase type 2C member 1 (EC 3.6.3.8) (ATPase 2C1) (ATP-dependent Ca(2+) pump PMR1). | |||||
|
AT2C1_HUMAN
|
||||||
| NC score | 0.915196 (rank : 7) | θ value | 1.04096e-72 (rank : 12) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P98194, O76005, Q86V72, Q86V73, Q8N6V1, Q8NCJ7 | Gene names | ATP2C1, KIAA1347, PMR1L | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-transporting ATPase type 2C member 1 (EC 3.6.3.8) (ATPase 2C1) (ATP-dependent Ca(2+) pump PMR1). | |||||
|
AT2A3_HUMAN
|
||||||
| NC score | 0.907417 (rank : 8) | θ value | 9.7205e-79 (rank : 7) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q93084, O60900, O60901, O75501, O75502, Q16115, Q8TEX5, Q8TEX6 | Gene names | ATP2A3 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 3 (EC 3.6.3.8) (Calcium pump 3) (SERCA3) (SR Ca(2+)-ATPase 3). | |||||
|
AT2A2_MOUSE
|
||||||
| NC score | 0.906864 (rank : 9) | θ value | 2.02684e-76 (rank : 8) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O55143, Q9R2A9, Q9WUT5 | Gene names | Atp2a2 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 2 (EC 3.6.3.8) (Calcium pump 2) (SERCA2) (SR Ca(2+)-ATPase 2) (Calcium-transporting ATPase sarcoplasmic reticulum type, slow twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2A2_HUMAN
|
||||||
| NC score | 0.906790 (rank : 10) | θ value | 4.51533e-76 (rank : 9) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P16615, P16614 | Gene names | ATP2A2, ATP2B | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 2 (EC 3.6.3.8) (Calcium pump 2) (SERCA2) (SR Ca(2+)-ATPase 2) (Calcium-transporting ATPase sarcoplasmic reticulum type, slow twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2A3_MOUSE
|
||||||
| NC score | 0.905536 (rank : 11) | θ value | 5.69867e-79 (rank : 6) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q64518, O70625, Q64517 | Gene names | Atp2a3 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 3 (EC 3.6.3.8) (Calcium pump 3) (SERCA3) (SR Ca(2+)-ATPase 3). | |||||
|
AT2A1_MOUSE
|
||||||
| NC score | 0.903249 (rank : 12) | θ value | 1.7756e-72 (rank : 13) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8R429 | Gene names | Atp2a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 1 (EC 3.6.3.8) (Calcium pump 1) (SERCA1) (SR Ca(2+)-ATPase 1) (Calcium-transporting ATPase sarcoplasmic reticulum type, fast twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2A1_HUMAN
|
||||||
| NC score | 0.903066 (rank : 13) | θ value | 2.09745e-73 (rank : 11) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O14983, O14984 | Gene names | ATP2A1 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 1 (EC 3.6.3.8) (Calcium pump 1) (SERCA1) (SR Ca(2+)-ATPase 1) (Calcium-transporting ATPase sarcoplasmic reticulum type, fast twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2C2_HUMAN
|
||||||
| NC score | 0.902341 (rank : 14) | θ value | 1.50663e-63 (rank : 20) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O75185 | Gene names | KIAA0703 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable calcium-transporting ATPase KIAA0703 (EC 3.6.3.8). | |||||
|
AT12A_HUMAN
|
||||||
| NC score | 0.886648 (rank : 15) | θ value | 2.10232e-65 (rank : 18) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P54707, Q13816, Q13817, Q16734 | Gene names | ATP12A, ATP1AL1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 2 (EC 3.6.3.10) (Proton pump) (Non-gastric H(+)/K(+) ATPase subunit alpha). | |||||
|
AT12A_MOUSE
|
||||||
| NC score | 0.885443 (rank : 16) | θ value | 8.83269e-64 (rank : 19) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9Z1W8, Q8VHY2 | Gene names | Atp12a, Atp1al1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium-transporting ATPase alpha chain 2 (EC 3.6.3.10) (Proton pump) (Non-gastric H(+)/K(+) ATPase subunit alpha). | |||||
|
ATP4A_MOUSE
|
||||||
| NC score | 0.878597 (rank : 17) | θ value | 4.24587e-58 (rank : 25) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q64436, Q9CV46 | Gene names | Atp4a | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 1 (EC 3.6.3.10) (Proton pump) (Gastric H+/K+ ATPase subunit alpha). | |||||
|
AT1A4_MOUSE
|
||||||
| NC score | 0.878222 (rank : 18) | θ value | 6.53529e-67 (rank : 17) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9WV27, Q9R173, Q9WV28 | Gene names | Atp1a4, Atp1al2 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-4 chain (EC 3.6.3.9) (Sodium pump 4) (Na+/K+ ATPase 4). | |||||
|
ATP4A_HUMAN
|
||||||
| NC score | 0.876719 (rank : 19) | θ value | 1.23536e-57 (rank : 26) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P20648, O00738 | Gene names | ATP4A | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 1 (EC 3.6.3.10) (Proton pump) (Gastric H+/K+ ATPase subunit alpha). | |||||
|
AT1A1_MOUSE
|
||||||
| NC score | 0.873907 (rank : 20) | θ value | 1.19377e-60 (rank : 22) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8VDN2, Q91Z09 | Gene names | Atp1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-1 chain precursor (EC 3.6.3.9) (Sodium pump 1) (Na+/K+ ATPase 1). | |||||
|
AT1A4_HUMAN
|
||||||
| NC score | 0.873307 (rank : 21) | θ value | 5.92465e-60 (rank : 24) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q13733, Q504T2, Q8TBN8, Q8WXA7, Q8WXH7, Q8WY13 | Gene names | ATP1A4, ATP1AL2 | |||
|
Domain Architecture |
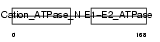
|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-4 chain (EC 3.6.3.9) (Sodium pump 4) (Na+/K+ ATPase 4). | |||||
|
AT1A1_HUMAN
|
||||||
| NC score | 0.873125 (rank : 22) | θ value | 9.1404e-61 (rank : 21) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P05023, Q16689, Q6LDM4, Q9UJ20, Q9UJ21 | Gene names | ATP1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-1 chain precursor (EC 3.6.3.9) (Sodium pump 1) (Na+/K+ ATPase 1). | |||||
|
AT1A2_MOUSE
|
||||||
| NC score | 0.870742 (rank : 23) | θ value | 6.53529e-67 (rank : 16) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q6PIE5 | Gene names | Atp1a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2) (Alpha(+)). | |||||
|
AT1A2_HUMAN
|
||||||
| NC score | 0.869948 (rank : 24) | θ value | 2.65945e-60 (rank : 23) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P50993, Q07059, Q86UZ5, Q9UQ25 | Gene names | ATP1A2, KIAA0778 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2). | |||||
|
AT1A3_HUMAN
|
||||||
| NC score | 0.869044 (rank : 25) | θ value | 2.24614e-67 (rank : 14) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P13637, Q16732, Q16735, Q969K5 | Gene names | ATP1A3 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-3 chain (EC 3.6.3.9) (Sodium pump 3) (Na+/K+ ATPase 3) (Alpha(III)). | |||||
|
AT1A3_MOUSE
|
||||||
| NC score | 0.868395 (rank : 26) | θ value | 2.93355e-67 (rank : 15) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q6PIC6 | Gene names | Atp1a3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium/potassium-transporting ATPase alpha-3 chain (EC 3.6.3.9) (Sodium pump 3) (Na+/K+ ATPase 3) (Alpha(III)). | |||||
|
AT132_MOUSE
|
||||||
| NC score | 0.725190 (rank : 27) | θ value | 1.38178e-16 (rank : 37) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9CTG6, Q8CG98 | Gene names | Atp13a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable cation-transporting ATPase 13A2 (EC 3.6.3.-). | |||||
|
AT133_HUMAN
|
||||||
| NC score | 0.722271 (rank : 28) | θ value | 2.28291e-11 (rank : 51) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9H7F0, Q8NC11, Q96KS1 | Gene names | ATP13A3, AFURS1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A3 (EC 3.6.3.-) (ATPase family homolog up-regulated in senescence cells 1). | |||||
|
AT132_HUMAN
|
||||||
| NC score | 0.717560 (rank : 29) | θ value | 6.62687e-19 (rank : 29) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9NQ11, O75700, Q5JXY2 | Gene names | ATP13A2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A2 (EC 3.6.3.-). | |||||
|
AT131_HUMAN
|
||||||
| NC score | 0.698165 (rank : 30) | θ value | 7.84624e-12 (rank : 50) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9HD20, Q9H6C6 | Gene names | ATP13A1, ATP13A, KIAA1825 | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A1 (EC 3.6.3.-). | |||||
|
AT131_MOUSE
|
||||||
| NC score | 0.687338 (rank : 31) | θ value | 1.25267e-09 (rank : 53) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9EPE9 | Gene names | Atp13a1, Atp13a | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A1 (EC 3.6.3.-) (CATP). | |||||
|
ATP7A_HUMAN
|
||||||
| NC score | 0.612185 (rank : 32) | θ value | 3.63628e-17 (rank : 34) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q04656, O00227, O00745, Q9BYY8 | Gene names | ATP7A, MC1, MNK | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 1 (EC 3.6.3.4) (Copper pump 1) (Menkes disease-associated protein). | |||||
|
ATP7A_MOUSE
|
||||||
| NC score | 0.608701 (rank : 33) | θ value | 2.20605e-14 (rank : 41) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q64430, O35101, P97422, Q64431 | Gene names | Atp7a, Mnk | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 1 (EC 3.6.3.4) (Copper pump 1) (Menkes disease-associated protein homolog). | |||||
|
ATP7B_MOUSE
|
||||||
| NC score | 0.608563 (rank : 34) | θ value | 1.68911e-14 (rank : 40) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q64446 | Gene names | Atp7b, Wnd | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 2 (EC 3.6.3.4) (Copper pump 2) (Wilson disease-associated protein homolog). | |||||
|
ATP7B_HUMAN
|
||||||
| NC score | 0.603213 (rank : 35) | θ value | 4.16044e-13 (rank : 46) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P35670, Q16318, Q16319, Q4U3V3, Q59FJ9, Q5T7X7 | Gene names | ATP7B, PWD, WC1, WND | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 2 (EC 3.6.3.4) (Copper pump 2) (Wilson disease-associated protein). | |||||
|
AT8A2_HUMAN
|
||||||
| NC score | 0.566588 (rank : 36) | θ value | 1.13037e-18 (rank : 30) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9NTI2, Q9H527, Q9NPU6, Q9NTL2, Q9NYM3 | Gene names | ATP8A2, ATPIB | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IB (EC 3.6.3.1) (ATPase class I type 8A member 2) (ML-1). | |||||
|
AT8A2_MOUSE
|
||||||
| NC score | 0.566585 (rank : 37) | θ value | 4.58923e-20 (rank : 28) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98200 | Gene names | Atp8a2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IB (EC 3.6.3.1) (ATPase class I type 8A member 2). | |||||
|
AT11B_HUMAN
|
||||||
| NC score | 0.562584 (rank : 38) | θ value | 1.058e-16 (rank : 36) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9Y2G3, Q96FN1, Q9UKK7 | Gene names | ATP11B, ATPIF, ATPIR, KIAA0956 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IF (EC 3.6.3.1) (ATPase class I type 11B) (ATPase IR). | |||||
|
AT8A1_MOUSE
|
||||||
| NC score | 0.559892 (rank : 39) | θ value | 7.32683e-18 (rank : 32) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P70704 | Gene names | Atp8a1, Atpc1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IA (EC 3.6.3.1) (Chromaffin granule ATPase II) (ATPase class I type 8A member 1). | |||||
|
AT8A1_HUMAN
|
||||||
| NC score | 0.556597 (rank : 40) | θ value | 2.13179e-17 (rank : 33) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Y2Q0 | Gene names | ATP8A1, ATPIA | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IA (EC 3.6.3.1) (Chromaffin granule ATPase II) (ATPase class I type 8A member 1). | |||||
|
ATP9A_HUMAN
|
||||||
| NC score | 0.555209 (rank : 41) | θ value | 9.59137e-10 (rank : 52) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O75110, Q5TFW5, Q5TFW6, Q5TFW9, Q6ZMF3, Q9NQK6, Q9NQK7 | Gene names | ATP9A, ATPIIA, KIAA0611 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIA (EC 3.6.3.1) (ATPase class II type 9A) (ATPase IIA). | |||||
|
ATP9A_MOUSE
|
||||||
| NC score | 0.543910 (rank : 42) | θ value | 6.87365e-08 (rank : 56) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O70228, Q8VDI5, Q922L9 | Gene names | Atp9a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIA (EC 3.6.3.1) (ATPase class II type 9A). | |||||
|
ATP9B_HUMAN
|
||||||
| NC score | 0.542523 (rank : 43) | θ value | 4.0297e-08 (rank : 54) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O43861, O60872 | Gene names | ATP9B, ATPIIB, NEO1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIB (EC 3.6.3.1). | |||||
|
AT10D_MOUSE
|
||||||
| NC score | 0.537253 (rank : 44) | θ value | 1.42992e-13 (rank : 43) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8K2X1 | Gene names | Atp10d | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VD (EC 3.6.3.1) (ATPVD). | |||||
|
AT11A_HUMAN
|
||||||
| NC score | 0.535980 (rank : 45) | θ value | 9.24701e-21 (rank : 27) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98196, Q5VXT2 | Gene names | ATP11A, ATPIH, ATPIS, KIAA1021 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IH (EC 3.6.3.1) (ATPase class I type 11A) (ATPase IS). | |||||
|
AT10D_HUMAN
|
||||||
| NC score | 0.534131 (rank : 46) | θ value | 7.58209e-15 (rank : 39) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9P241, Q8NC70, Q96SR3 | Gene names | ATP10D, KIAA1487 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VD (EC 3.6.3.1) (ATPVD). | |||||
|
ATP9B_MOUSE
|
||||||
| NC score | 0.533351 (rank : 47) | θ value | 5.26297e-08 (rank : 55) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98195, Q99LI3 | Gene names | Atp9b | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IIB (EC 3.6.3.1). | |||||
|
AT11C_HUMAN
|
||||||
| NC score | 0.531185 (rank : 48) | θ value | 2.43908e-13 (rank : 44) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8NB49, Q5JT69, Q5JT70, Q5JT71, Q5JT72, Q5JT73, Q6ZND5, Q6ZU50, Q6ZUP7, Q70IJ9, Q70IK0, Q8WX24 | Gene names | ATP11C, ATPIG, ATPIQ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IG (EC 3.6.3.1) (ATPase class I type 11C) (ATPase IG) (ATPase IQ) (ATPase class VI type 11C). | |||||
|
AT10B_HUMAN
|
||||||
| NC score | 0.530326 (rank : 49) | θ value | 1.16975e-15 (rank : 38) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O94823, Q9H725 | Gene names | ATP10B, ATPVB, KIAA0715 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VB (EC 3.6.3.1). | |||||
|
AT8B3_HUMAN
|
||||||
| NC score | 0.524429 (rank : 50) | θ value | 4.29542e-18 (rank : 31) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O60423, Q8IVB8, Q8N4Y8, Q96M22 | Gene names | ATP8B3, ATP1K, FOS37502_2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IK (EC 3.6.3.1) (ATPase class I type 8B member 3). | |||||
|
AT8B4_HUMAN
|
||||||
| NC score | 0.522640 (rank : 51) | θ value | 1.58096e-12 (rank : 48) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8TF62, Q9H727 | Gene names | ATP8B4, KIAA1939 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IM (EC 3.6.3.1) (ATPase class I type 8B member 4). | |||||
|
AT8B1_HUMAN
|
||||||
| NC score | 0.521251 (rank : 52) | θ value | 1.09485e-13 (rank : 42) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O43520, Q9BTP8 | Gene names | ATP8B1, ATPIC, FIC1, PFIC | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IC (EC 3.6.3.1) (Familial intrahepatic cholestasis type 1) (ATPase class I type 8B member 1). | |||||
|
AT11A_MOUSE
|
||||||
| NC score | 0.519812 (rank : 53) | θ value | 8.10077e-17 (rank : 35) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98197 | Gene names | Atp11a | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IH (EC 3.6.3.1) (ATPase class I type 11A) (ATPase IS). | |||||
|
AT10A_HUMAN
|
||||||
| NC score | 0.517586 (rank : 54) | θ value | 5.43371e-13 (rank : 47) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O60312, Q969I4 | Gene names | ATP10A, ATP10C, ATPVC, KIAA0566 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (ATPVA) (Aminophospholipid translocase VA). | |||||
|
AT10A_MOUSE
|
||||||
| NC score | 0.516906 (rank : 55) | θ value | 2.0648e-12 (rank : 49) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O54827, Q8R3B8 | Gene names | Atp10a, Atpc5, Pfatp | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (P-locus fat-associated ATPase). | |||||
|
AT8B2_HUMAN
|
||||||
| NC score | 0.516418 (rank : 56) | θ value | 3.18553e-13 (rank : 45) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P98198, Q96I43, Q96NQ7 | Gene names | ATP8B2, ATPID, KIAA1137 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase ID (EC 3.6.3.1) (ATPase class I type 8B member 2). | |||||
|
K0195_MOUSE
|
||||||
| NC score | 0.072844 (rank : 57) | θ value | 0.62314 (rank : 60) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TSH8, Q80U66, Q8BL05, Q8K0Q0, Q91VY1 | Gene names | Kiaa0195 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0195. | |||||
|
K0195_HUMAN
|
||||||
| NC score | 0.072713 (rank : 58) | θ value | 0.62314 (rank : 59) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q12767, O75536, Q86XF1 | Gene names | KIAA0195, TMEM94 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0195 (Transmembrane protein 94). | |||||
|
PVRL2_HUMAN
|
||||||
| NC score | 0.021866 (rank : 59) | θ value | 3.0926 (rank : 68) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92692, O75455, Q96J29 | Gene names | PVRL2, HVEB, PRR2 | |||
|
Domain Architecture |

|
|||||
| Description | Poliovirus receptor-related protein 2 precursor (Herpes virus entry mediator B) (HveB) (Nectin-2) (CD112 antigen). | |||||
|
NUD15_MOUSE
|
||||||
| NC score | 0.017147 (rank : 60) | θ value | 2.36792 (rank : 65) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BG93 | Gene names | Nudt15, Mth2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable 7,8-dihydro-8-oxoguanine triphosphatase NUDT15 (EC 3.1.6.-) (8-oxo-dGTPase NUDT15) (Nucleoside diphosphate-linked moiety X motif 15) (Nudix motif 15) (MutT homolog 2) (mMTH2). | |||||
|
AHI1_HUMAN
|
||||||
| NC score | 0.016741 (rank : 61) | θ value | 0.62314 (rank : 58) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 537 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8N157, Q504T3, Q5TCP9, Q6P098, Q6PIT6, Q8NDX0, Q9H0H2 | Gene names | AHI1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Jouberin (Abelson helper integration site 1 protein homolog) (AHI-1). | |||||
|
UBP25_HUMAN
|
||||||
| NC score | 0.013402 (rank : 62) | θ value | 2.36792 (rank : 66) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UHP3, Q9H9W1 | Gene names | USP25, USP21 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 25 (EC 3.1.2.15) (Ubiquitin thioesterase 25) (Ubiquitin-specific-processing protease 25) (Deubiquitinating enzyme 25) (USP on chromosome 21). | |||||
|
UBP28_HUMAN
|
||||||
| NC score | 0.012362 (rank : 63) | θ value | 4.03905 (rank : 70) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96RU2, Q9P213 | Gene names | USP28, KIAA1515 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 28 (EC 3.1.2.15) (Ubiquitin thioesterase 28) (Ubiquitin-specific-processing protease 28) (Deubiquitinating enzyme 28). | |||||
|
UBP25_MOUSE
|
||||||
| NC score | 0.012224 (rank : 64) | θ value | 5.27518 (rank : 73) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P57080, Q80ZT9 | Gene names | Usp25 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 25 (EC 3.1.2.15) (Ubiquitin thioesterase 25) (Ubiquitin-specific-processing protease 25) (Deubiquitinating enzyme 25) (mUSP25). | |||||
|
NUD15_HUMAN
|
||||||
| NC score | 0.012171 (rank : 65) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NV35, Q6P2C9 | Gene names | NUDT15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable 7,8-dihydro-8-oxoguanine triphosphatase NUDT15 (EC 3.1.6.-) (8-oxo-dGTPase NUDT15) (Nucleoside diphosphate-linked moiety X motif 15) (Nudix motif 15). | |||||
|
UBP28_MOUSE
|
||||||
| NC score | 0.011142 (rank : 66) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5I043, Q6NZP3, Q6ZPP1, Q8BWI1 | Gene names | Usp28, Kiaa1515 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 28 (EC 3.1.2.15) (Ubiquitin thioesterase 28) (Ubiquitin-specific-processing protease 28) (Deubiquitinating enzyme 28). | |||||
|
TAF1C_MOUSE
|
||||||
| NC score | 0.009139 (rank : 67) | θ value | 4.03905 (rank : 69) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6PDZ2, P97359, Q3U3B6, Q8BN54 | Gene names | Taf1c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA box-binding protein-associated factor, RNA polymerase I, subunit C (TATA box-binding protein-associated factor 1C) (TBP-associated factor 1C) (TBP-associated factor RNA polymerase I 95 kDa) (TAFI95). | |||||
|
RPC62_MOUSE
|
||||||
| NC score | 0.008310 (rank : 68) | θ value | 5.27518 (rank : 71) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D483, Q8R0I2 | Gene names | Polr3c, Rpc62 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase III 62 kDa polypeptide (EC 2.7.7.6) (RNA polymerase C subunit 3) (RPC3) (RNA polymerase III C62 subunit). | |||||
|
ERIC1_HUMAN
|
||||||
| NC score | 0.007701 (rank : 69) | θ value | 2.36792 (rank : 64) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
NAL13_HUMAN
|
||||||
| NC score | 0.007492 (rank : 70) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86W25, Q7RTR5 | Gene names | NALP13, NOD14 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 13 (Nucleotide-binding oligomerization domain protein 14). | |||||
|
RRBP1_MOUSE
|
||||||
| NC score | 0.006898 (rank : 71) | θ value | 0.163984 (rank : 57) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
RRBP1_HUMAN
|
||||||
| NC score | 0.004639 (rank : 72) | θ value | 1.38821 (rank : 62) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1306 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9P2E9, O75300, O75301, Q5W165, Q96SB2, Q9BWP1, Q9H476 | Gene names | RRBP1, KIAA1398 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (180 kDa ribosome receptor homolog) (ES/130-related protein). | |||||
|
PRGC1_MOUSE
|
||||||
| NC score | 0.003090 (rank : 73) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O70343 | Gene names | Ppargc1a, Pgc1, Pgc1a, Ppargc1 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PPAR gamma coactivator 1-alpha) (PPARGC-1-alpha) (PGC-1-alpha). | |||||
|
LRCH3_HUMAN
|
||||||
| NC score | 0.002937 (rank : 74) | θ value | 3.0926 (rank : 67) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96II8, Q96FP9, Q9NT52 | Gene names | LRCH3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 3 precursor. | |||||
|
PRAM3_HUMAN
|
||||||
| NC score | 0.002925 (rank : 75) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5TYW8 | Gene names | PRAMEF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PRAME family member 3. | |||||
|
KIF3A_HUMAN
|
||||||
| NC score | 0.002887 (rank : 76) | θ value | 0.813845 (rank : 61) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y496, Q86XE9, Q9Y6V4 | Gene names | KIF3A, KIF3 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF3A (Microtubule plus end-directed kinesin motor 3A). | |||||
|
KIF3A_MOUSE
|
||||||
| NC score | 0.002348 (rank : 77) | θ value | 1.81305 (rank : 63) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 715 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P28741 | Gene names | Kif3a, Kif3 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF3A (Microtubule plus end-directed kinesin motor 3A). | |||||
|
LRCH3_MOUSE
|
||||||
| NC score | 0.002263 (rank : 78) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BVU0, Q3U222, Q3UZ74 | Gene names | Lrch3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 3 precursor. | |||||
|
TEX14_HUMAN
|
||||||
| NC score | -0.003208 (rank : 79) | θ value | 5.27518 (rank : 72) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1103 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IWB6, Q7RTP3, Q8ND97, Q9BXT9 | Gene names | TEX14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed protein 14 (Testis-expressed sequence 14). | |||||