Please be patient as the page loads
|
AT2B1_HUMAN
|
||||||
| SwissProt Accessions | P20020, Q12992, Q12993, Q13819, Q13820, Q13821, Q16504, Q93082 | Gene names | ATP2B1, PMCA1 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 1 (EC 3.6.3.8) (PMCA1) (Plasma membrane calcium pump isoform 1) (Plasma membrane calcium ATPase isoform 1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
AT2B1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 88 | |
| SwissProt Accessions | P20020, Q12992, Q12993, Q13819, Q13820, Q13821, Q16504, Q93082 | Gene names | ATP2B1, PMCA1 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 1 (EC 3.6.3.8) (PMCA1) (Plasma membrane calcium pump isoform 1) (Plasma membrane calcium ATPase isoform 1). | |||||
|
AT2B2_HUMAN
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.996760 (rank : 4) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q01814, O00766, Q12994, Q16818 | Gene names | ATP2B2, PMCA2 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 2 (EC 3.6.3.8) (PMCA2) (Plasma membrane calcium pump isoform 2) (Plasma membrane calcium ATPase isoform 2). | |||||
|
AT2B2_MOUSE
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.996944 (rank : 3) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9R0K7, O88863 | Gene names | Atp2b2, Pmca2 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 2 (EC 3.6.3.8) (PMCA2) (Plasma membrane calcium pump isoform 2) (Plasma membrane calcium ATPase isoform 2). | |||||
|
AT2B3_HUMAN
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.997244 (rank : 2) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q16720, Q12995, Q16858 | Gene names | ATP2B3 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 3 (EC 3.6.3.8) (PMCA3) (Plasma membrane calcium pump isoform 3) (Plasma membrane calcium ATPase isoform 3). | |||||
|
AT2B4_HUMAN
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.996338 (rank : 5) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P23634, Q13450, Q13452, Q13455, Q16817 | Gene names | ATP2B4 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 4 (EC 3.6.3.8) (PMCA4) (Plasma membrane calcium pump isoform 4) (Plasma membrane calcium ATPase isoform 4). | |||||
|
AT2A2_MOUSE
|
||||||
| θ value | 2.82823e-78 (rank : 6) | NC score | 0.905983 (rank : 10) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O55143, Q9R2A9, Q9WUT5 | Gene names | Atp2a2 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 2 (EC 3.6.3.8) (Calcium pump 2) (SERCA2) (SR Ca(2+)-ATPase 2) (Calcium-transporting ATPase sarcoplasmic reticulum type, slow twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2A2_HUMAN
|
||||||
| θ value | 4.82422e-78 (rank : 7) | NC score | 0.906004 (rank : 9) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P16615, P16614 | Gene names | ATP2A2, ATP2B | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 2 (EC 3.6.3.8) (Calcium pump 2) (SERCA2) (SR Ca(2+)-ATPase 2) (Calcium-transporting ATPase sarcoplasmic reticulum type, slow twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2A3_HUMAN
|
||||||
| θ value | 4.08394e-77 (rank : 8) | NC score | 0.906906 (rank : 8) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q93084, O60900, O60901, O75501, O75502, Q16115, Q8TEX5, Q8TEX6 | Gene names | ATP2A3 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 3 (EC 3.6.3.8) (Calcium pump 3) (SERCA3) (SR Ca(2+)-ATPase 3). | |||||
|
AT2A3_MOUSE
|
||||||
| θ value | 2.64714e-76 (rank : 9) | NC score | 0.905112 (rank : 11) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q64518, O70625, Q64517 | Gene names | Atp2a3 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 3 (EC 3.6.3.8) (Calcium pump 3) (SERCA3) (SR Ca(2+)-ATPase 3). | |||||
|
AT2A1_HUMAN
|
||||||
| θ value | 5.1662e-72 (rank : 10) | NC score | 0.902820 (rank : 14) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O14983, O14984 | Gene names | ATP2A1 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 1 (EC 3.6.3.8) (Calcium pump 1) (SERCA1) (SR Ca(2+)-ATPase 1) (Calcium-transporting ATPase sarcoplasmic reticulum type, fast twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2A1_MOUSE
|
||||||
| θ value | 3.34864e-71 (rank : 11) | NC score | 0.902955 (rank : 13) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8R429 | Gene names | Atp2a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 1 (EC 3.6.3.8) (Calcium pump 1) (SERCA1) (SR Ca(2+)-ATPase 1) (Calcium-transporting ATPase sarcoplasmic reticulum type, fast twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2C1_MOUSE
|
||||||
| θ value | 4.83544e-70 (rank : 12) | NC score | 0.914615 (rank : 6) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q80XR2, Q80YZ2 | Gene names | Atp2c1, Pmr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-transporting ATPase type 2C member 1 (EC 3.6.3.8) (ATPase 2C1) (ATP-dependent Ca(2+) pump PMR1). | |||||
|
AT2C1_HUMAN
|
||||||
| θ value | 3.13423e-69 (rank : 13) | NC score | 0.913840 (rank : 7) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P98194, O76005, Q86V72, Q86V73, Q8N6V1, Q8NCJ7 | Gene names | ATP2C1, KIAA1347, PMR1L | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-transporting ATPase type 2C member 1 (EC 3.6.3.8) (ATPase 2C1) (ATP-dependent Ca(2+) pump PMR1). | |||||
|
AT12A_HUMAN
|
||||||
| θ value | 6.11685e-65 (rank : 14) | NC score | 0.884334 (rank : 15) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P54707, Q13816, Q13817, Q16734 | Gene names | ATP12A, ATP1AL1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 2 (EC 3.6.3.10) (Proton pump) (Non-gastric H(+)/K(+) ATPase subunit alpha). | |||||
|
AT12A_MOUSE
|
||||||
| θ value | 3.03575e-64 (rank : 15) | NC score | 0.883092 (rank : 16) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9Z1W8, Q8VHY2 | Gene names | Atp12a, Atp1al1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium-transporting ATPase alpha chain 2 (EC 3.6.3.10) (Proton pump) (Non-gastric H(+)/K(+) ATPase subunit alpha). | |||||
|
AT2C2_HUMAN
|
||||||
| θ value | 1.50663e-63 (rank : 16) | NC score | 0.903676 (rank : 12) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O75185 | Gene names | KIAA0703 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable calcium-transporting ATPase KIAA0703 (EC 3.6.3.8). | |||||
|
AT1A4_MOUSE
|
||||||
| θ value | 2.17557e-62 (rank : 17) | NC score | 0.875994 (rank : 17) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9WV27, Q9R173, Q9WV28 | Gene names | Atp1a4, Atp1al2 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-4 chain (EC 3.6.3.9) (Sodium pump 4) (Na+/K+ ATPase 4). | |||||
|
AT1A1_HUMAN
|
||||||
| θ value | 9.1404e-61 (rank : 18) | NC score | 0.871240 (rank : 21) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P05023, Q16689, Q6LDM4, Q9UJ20, Q9UJ21 | Gene names | ATP1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-1 chain precursor (EC 3.6.3.9) (Sodium pump 1) (Na+/K+ ATPase 1). | |||||
|
AT1A1_MOUSE
|
||||||
| θ value | 7.73779e-60 (rank : 19) | NC score | 0.871803 (rank : 20) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8VDN2, Q91Z09 | Gene names | Atp1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-1 chain precursor (EC 3.6.3.9) (Sodium pump 1) (Na+/K+ ATPase 1). | |||||
|
AT1A4_HUMAN
|
||||||
| θ value | 5.0155e-59 (rank : 20) | NC score | 0.870785 (rank : 22) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q13733, Q504T2, Q8TBN8, Q8WXA7, Q8WXH7, Q8WY13 | Gene names | ATP1A4, ATP1AL2 | |||
|
Domain Architecture |
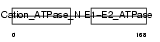
|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-4 chain (EC 3.6.3.9) (Sodium pump 4) (Na+/K+ ATPase 4). | |||||
|
AT1A3_HUMAN
|
||||||
| θ value | 6.55044e-59 (rank : 21) | NC score | 0.867326 (rank : 25) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P13637, Q16732, Q16735, Q969K5 | Gene names | ATP1A3 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-3 chain (EC 3.6.3.9) (Sodium pump 3) (Na+/K+ ATPase 3) (Alpha(III)). | |||||
|
AT1A3_MOUSE
|
||||||
| θ value | 6.55044e-59 (rank : 22) | NC score | 0.866706 (rank : 26) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q6PIC6 | Gene names | Atp1a3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium/potassium-transporting ATPase alpha-3 chain (EC 3.6.3.9) (Sodium pump 3) (Na+/K+ ATPase 3) (Alpha(III)). | |||||
|
AT1A2_HUMAN
|
||||||
| θ value | 5.54529e-58 (rank : 23) | NC score | 0.867621 (rank : 24) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P50993, Q07059, Q86UZ5, Q9UQ25 | Gene names | ATP1A2, KIAA0778 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2). | |||||
|
AT1A2_MOUSE
|
||||||
| θ value | 7.24236e-58 (rank : 24) | NC score | 0.868293 (rank : 23) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q6PIE5 | Gene names | Atp1a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2) (Alpha(+)). | |||||
|
ATP4A_MOUSE
|
||||||
| θ value | 2.10721e-57 (rank : 25) | NC score | 0.875717 (rank : 18) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q64436, Q9CV46 | Gene names | Atp4a | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 1 (EC 3.6.3.10) (Proton pump) (Gastric H+/K+ ATPase subunit alpha). | |||||
|
ATP4A_HUMAN
|
||||||
| θ value | 6.13105e-57 (rank : 26) | NC score | 0.873995 (rank : 19) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P20648, O00738 | Gene names | ATP4A | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 1 (EC 3.6.3.10) (Proton pump) (Gastric H+/K+ ATPase subunit alpha). | |||||
|
AT11A_HUMAN
|
||||||
| θ value | 3.88503e-19 (rank : 27) | NC score | 0.507181 (rank : 45) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98196, Q5VXT2 | Gene names | ATP11A, ATPIH, ATPIS, KIAA1021 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IH (EC 3.6.3.1) (ATPase class I type 11A) (ATPase IS). | |||||
|
AT132_HUMAN
|
||||||
| θ value | 7.32683e-18 (rank : 28) | NC score | 0.707776 (rank : 29) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9NQ11, O75700, Q5JXY2 | Gene names | ATP13A2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A2 (EC 3.6.3.-). | |||||
|
AT132_MOUSE
|
||||||
| θ value | 1.24977e-17 (rank : 29) | NC score | 0.715363 (rank : 27) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9CTG6, Q8CG98 | Gene names | Atp13a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable cation-transporting ATPase 13A2 (EC 3.6.3.-). | |||||
|
ATP7B_MOUSE
|
||||||
| θ value | 4.02038e-16 (rank : 30) | NC score | 0.616327 (rank : 33) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q64446 | Gene names | Atp7b, Wnd | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 2 (EC 3.6.3.4) (Copper pump 2) (Wilson disease-associated protein homolog). | |||||
|
AT10B_HUMAN
|
||||||
| θ value | 1.16975e-15 (rank : 31) | NC score | 0.502749 (rank : 47) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O94823, Q9H725 | Gene names | ATP10B, ATPVB, KIAA0715 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VB (EC 3.6.3.1). | |||||
|
AT11A_MOUSE
|
||||||
| θ value | 1.52774e-15 (rank : 32) | NC score | 0.490784 (rank : 52) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98197 | Gene names | Atp11a | |||
|
Domain Architecture |
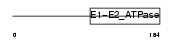
|
|||||
| Description | Probable phospholipid-transporting ATPase IH (EC 3.6.3.1) (ATPase class I type 11A) (ATPase IS). | |||||
|
ATP7A_MOUSE
|
||||||
| θ value | 2.60593e-15 (rank : 33) | NC score | 0.615612 (rank : 34) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q64430, O35101, P97422, Q64431 | Gene names | Atp7a, Mnk | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 1 (EC 3.6.3.4) (Copper pump 1) (Menkes disease-associated protein homolog). | |||||
|
ATP7A_HUMAN
|
||||||
| θ value | 5.8054e-15 (rank : 34) | NC score | 0.619167 (rank : 32) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q04656, O00227, O00745, Q9BYY8 | Gene names | ATP7A, MC1, MNK | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 1 (EC 3.6.3.4) (Copper pump 1) (Menkes disease-associated protein). | |||||
|
ATP7B_HUMAN
|
||||||
| θ value | 2.20605e-14 (rank : 35) | NC score | 0.610823 (rank : 35) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P35670, Q16318, Q16319, Q4U3V3, Q59FJ9, Q5T7X7 | Gene names | ATP7B, PWD, WC1, WND | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 2 (EC 3.6.3.4) (Copper pump 2) (Wilson disease-associated protein). | |||||
|
AT11B_HUMAN
|
||||||
| θ value | 3.76295e-14 (rank : 36) | NC score | 0.533508 (rank : 38) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9Y2G3, Q96FN1, Q9UKK7 | Gene names | ATP11B, ATPIF, ATPIR, KIAA0956 | |||
|
Domain Architecture |
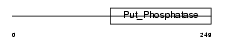
|
|||||
| Description | Probable phospholipid-transporting ATPase IF (EC 3.6.3.1) (ATPase class I type 11B) (ATPase IR). | |||||
|
AT11C_HUMAN
|
||||||
| θ value | 2.43908e-13 (rank : 37) | NC score | 0.502265 (rank : 48) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8NB49, Q5JT69, Q5JT70, Q5JT71, Q5JT72, Q5JT73, Q6ZND5, Q6ZU50, Q6ZUP7, Q70IJ9, Q70IK0, Q8WX24 | Gene names | ATP11C, ATPIG, ATPIQ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IG (EC 3.6.3.1) (ATPase class I type 11C) (ATPase IG) (ATPase IQ) (ATPase class VI type 11C). | |||||
|
AT8A1_MOUSE
|
||||||
| θ value | 7.09661e-13 (rank : 38) | NC score | 0.527759 (rank : 39) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P70704 | Gene names | Atp8a1, Atpc1 | |||
|
Domain Architecture |
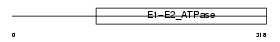
|
|||||
| Description | Probable phospholipid-transporting ATPase IA (EC 3.6.3.1) (Chromaffin granule ATPase II) (ATPase class I type 8A member 1). | |||||
|
AT8B1_HUMAN
|
||||||
| θ value | 9.26847e-13 (rank : 39) | NC score | 0.490505 (rank : 53) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O43520, Q9BTP8 | Gene names | ATP8B1, ATPIC, FIC1, PFIC | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IC (EC 3.6.3.1) (Familial intrahepatic cholestasis type 1) (ATPase class I type 8B member 1). | |||||
|
AT8A2_HUMAN
|
||||||
| θ value | 1.2105e-12 (rank : 40) | NC score | 0.534401 (rank : 36) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9NTI2, Q9H527, Q9NPU6, Q9NTL2, Q9NYM3 | Gene names | ATP8A2, ATPIB | |||
|
Domain Architecture |
|
|||||
| Description | Probable phospholipid-transporting ATPase IB (EC 3.6.3.1) (ATPase class I type 8A member 2) (ML-1). | |||||
|
AT8B2_HUMAN
|
||||||
| θ value | 1.2105e-12 (rank : 41) | NC score | 0.485555 (rank : 56) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P98198, Q96I43, Q96NQ7 | Gene names | ATP8B2, ATPID, KIAA1137 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase ID (EC 3.6.3.1) (ATPase class I type 8B member 2). | |||||
|
AT10D_HUMAN
|
||||||
| θ value | 2.69671e-12 (rank : 42) | NC score | 0.506319 (rank : 46) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9P241, Q8NC70, Q96SR3 | Gene names | ATP10D, KIAA1487 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VD (EC 3.6.3.1) (ATPVD). | |||||
|
AT8A1_HUMAN
|
||||||
| θ value | 3.52202e-12 (rank : 43) | NC score | 0.524297 (rank : 40) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Y2Q0 | Gene names | ATP8A1, ATPIA | |||
|
Domain Architecture |
|
|||||
| Description | Probable phospholipid-transporting ATPase IA (EC 3.6.3.1) (Chromaffin granule ATPase II) (ATPase class I type 8A member 1). | |||||
|
AT8A2_MOUSE
|
||||||
| θ value | 3.52202e-12 (rank : 44) | NC score | 0.534299 (rank : 37) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98200 | Gene names | Atp8a2 | |||
|
Domain Architecture |
|
|||||
| Description | Probable phospholipid-transporting ATPase IB (EC 3.6.3.1) (ATPase class I type 8A member 2). | |||||
|
AT8B3_HUMAN
|
||||||
| θ value | 3.52202e-12 (rank : 45) | NC score | 0.493482 (rank : 50) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O60423, Q8IVB8, Q8N4Y8, Q96M22 | Gene names | ATP8B3, ATP1K, FOS37502_2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IK (EC 3.6.3.1) (ATPase class I type 8B member 3). | |||||
|
AT10A_HUMAN
|
||||||
| θ value | 6.00763e-12 (rank : 46) | NC score | 0.489940 (rank : 54) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O60312, Q969I4 | Gene names | ATP10A, ATP10C, ATPVC, KIAA0566 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (ATPVA) (Aminophospholipid translocase VA). | |||||
|
AT10D_MOUSE
|
||||||
| θ value | 2.28291e-11 (rank : 47) | NC score | 0.509684 (rank : 44) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8K2X1 | Gene names | Atp10d | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VD (EC 3.6.3.1) (ATPVD). | |||||
|
AT133_HUMAN
|
||||||
| θ value | 6.64225e-11 (rank : 48) | NC score | 0.709564 (rank : 28) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9H7F0, Q8NC11, Q96KS1 | Gene names | ATP13A3, AFURS1 | |||
|
Domain Architecture |
|
|||||
| Description | Probable cation-transporting ATPase 13A3 (EC 3.6.3.-) (ATPase family homolog up-regulated in senescence cells 1). | |||||
|
AT8B4_HUMAN
|
||||||
| θ value | 1.133e-10 (rank : 49) | NC score | 0.491768 (rank : 51) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8TF62, Q9H727 | Gene names | ATP8B4, KIAA1939 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IM (EC 3.6.3.1) (ATPase class I type 8B member 4). | |||||
|
AT10A_MOUSE
|
||||||
| θ value | 5.62301e-10 (rank : 50) | NC score | 0.488821 (rank : 55) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O54827, Q8R3B8 | Gene names | Atp10a, Atpc5, Pfatp | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (P-locus fat-associated ATPase). | |||||
|
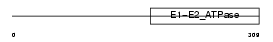
AT131_HUMAN
|
||||||
| θ value | 1.38499e-08 (rank : 51) | NC score | 0.685360 (rank : 30) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9HD20, Q9H6C6 | Gene names | ATP13A1, ATP13A, KIAA1825 | |||
|
Domain Architecture |
|
|||||
| Description | Probable cation-transporting ATPase 13A1 (EC 3.6.3.-). | |||||
|
AT131_MOUSE
|
||||||
| θ value | 1.38499e-08 (rank : 52) | NC score | 0.674284 (rank : 31) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9EPE9 | Gene names | Atp13a1, Atp13a | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A1 (EC 3.6.3.-) (CATP). | |||||
|
ATP9A_HUMAN
|
||||||
| θ value | 7.59969e-07 (rank : 53) | NC score | 0.523561 (rank : 41) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O75110, Q5TFW5, Q5TFW6, Q5TFW9, Q6ZMF3, Q9NQK6, Q9NQK7 | Gene names | ATP9A, ATPIIA, KIAA0611 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIA (EC 3.6.3.1) (ATPase class II type 9A) (ATPase IIA). | |||||
|
ATP9B_HUMAN
|
||||||
| θ value | 8.40245e-06 (rank : 54) | NC score | 0.510834 (rank : 43) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O43861, O60872 | Gene names | ATP9B, ATPIIB, NEO1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIB (EC 3.6.3.1). | |||||
|
ATP9B_MOUSE
|
||||||
| θ value | 1.09739e-05 (rank : 55) | NC score | 0.501504 (rank : 49) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98195, Q99LI3 | Gene names | Atp9b | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IIB (EC 3.6.3.1). | |||||
|
ATP9A_MOUSE
|
||||||
| θ value | 5.44631e-05 (rank : 56) | NC score | 0.512211 (rank : 42) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O70228, Q8VDI5, Q922L9 | Gene names | Atp9a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIA (EC 3.6.3.1) (ATPase class II type 9A). | |||||
|
TTF2_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 57) | NC score | 0.012234 (rank : 66) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5NC05, Q5M924 | Gene names | Ttf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 2 (EC 3.6.1.-) (RNA polymerase II termination factor) (Transcription release factor 2). | |||||
|
K0195_HUMAN
|
||||||
| θ value | 0.21417 (rank : 58) | NC score | 0.078390 (rank : 58) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q12767, O75536, Q86XF1 | Gene names | KIAA0195, TMEM94 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0195 (Transmembrane protein 94). | |||||
|
K0195_MOUSE
|
||||||
| θ value | 0.21417 (rank : 59) | NC score | 0.078559 (rank : 57) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TSH8, Q80U66, Q8BL05, Q8K0Q0, Q91VY1 | Gene names | Kiaa0195 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0195. | |||||
|
ZN452_HUMAN
|
||||||
| θ value | 0.279714 (rank : 60) | NC score | 0.013545 (rank : 65) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6R2W3, Q2NKL9, Q5SRJ3, Q8TCN2, Q96PW3 | Gene names | ZNF452, KIAA1925 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ZNF452. | |||||
|
PRP31_MOUSE
|
||||||
| θ value | 0.47712 (rank : 61) | NC score | 0.025753 (rank : 59) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CCF0, Q6P7X2, Q8BQ91, Q8C8U4, Q8C8V5, Q8CCG6, Q8CF52, Q8VBW3 | Gene names | Prpf31, Prp31 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6 small nuclear ribonucleoprotein Prp31 (Pre-mRNA-processing factor 31) (U4/U6 snRNP 61 kDa protein) (Protein 61K). | |||||
|
YETS2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 62) | NC score | 0.019063 (rank : 62) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3TUF7, Q6PGF8, Q80TI2, Q8CG86 | Gene names | Yeats2, Kiaa1197 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YEATS domain-containing protein 2. | |||||
|
PRP31_HUMAN
|
||||||
| θ value | 0.62314 (rank : 63) | NC score | 0.025155 (rank : 60) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WWY3, Q8N7F9, Q9H271, Q9Y439 | Gene names | PRPF31, PRP31 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6 small nuclear ribonucleoprotein Prp31 (Pre-mRNA-processing factor 31) (U4/U6 snRNP 61 kDa protein) (hPrp31) (Protein 61K) (Serologically defined breast cancer antigen NY-BR-99). | |||||
|
BCOR_HUMAN
|
||||||
| θ value | 1.38821 (rank : 64) | NC score | 0.003536 (rank : 80) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6W2J9, Q6P4B6, Q7Z2K7, Q8TEB4, Q96DB3, Q9H232, Q9H233, Q9HCJ7, Q9NXF2 | Gene names | BCOR, KIAA1575 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BCoR protein (BCL-6 corepressor). | |||||
|
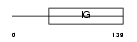
PVRL2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | 0.023173 (rank : 61) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92692, O75455, Q96J29 | Gene names | PVRL2, HVEB, PRR2 | |||
|
Domain Architecture |
|
|||||
| Description | Poliovirus receptor-related protein 2 precursor (Herpes virus entry mediator B) (HveB) (Nectin-2) (CD112 antigen). | |||||
|
AHI1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 66) | NC score | 0.015533 (rank : 64) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 537 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8N157, Q504T3, Q5TCP9, Q6P098, Q6PIT6, Q8NDX0, Q9H0H2 | Gene names | AHI1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Jouberin (Abelson helper integration site 1 protein homolog) (AHI-1). | |||||
|
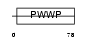
HDGF_MOUSE
|
||||||
| θ value | 2.36792 (rank : 67) | NC score | 0.009473 (rank : 67) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P51859, Q8BPG7, Q9CYA4, Q9JK87 | Gene names | Hdgf, Tdrm1 | |||
|
Domain Architecture |
|
|||||
| Description | Hepatoma-derived growth factor (HDGF). | |||||
|
RRBP1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.003484 (rank : 81) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
YETS2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 69) | NC score | 0.017376 (rank : 63) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ULM3, Q641P6, Q9NW96 | Gene names | YEATS2, KIAA1197 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YEATS domain-containing protein 2. | |||||
|
PHLPP_HUMAN
|
||||||
| θ value | 3.0926 (rank : 70) | NC score | 0.000399 (rank : 85) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 604 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60346, Q641Q7, Q6P4C4, Q6PJI6, Q86TN6, Q96FK2, Q9NUY1 | Gene names | PHLPP, KIAA0606, PLEKHE1, SCOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein) (hSCOP). | |||||
|
SPRL1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 71) | NC score | 0.005815 (rank : 71) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P70663, P97810, Q99L82 | Gene names | Sparcl1, Ecm2, Sc1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (Matrix glycoprotein Sc1) (Extracellular matrix protein 2). | |||||
|
COA2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.005517 (rank : 72) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O00763, Q16852 | Gene names | ACACB, ACC2, ACCB | |||
|
Domain Architecture |

|
|||||
| Description | Acetyl-CoA carboxylase 2 (EC 6.4.1.2) (ACC-beta) [Includes: Biotin carboxylase (EC 6.3.4.14)]. | |||||
|
MBB1A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 73) | NC score | 0.003710 (rank : 79) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
P52K_HUMAN
|
||||||
| θ value | 5.27518 (rank : 74) | NC score | 0.007141 (rank : 69) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43422, Q8WTW1, Q9Y3Z4 | Gene names | PRKRIR, DAP4, P52RIPK, THAP0 | |||
|
Domain Architecture |

|
|||||
| Description | 52 kDa repressor of the inhibitor of the protein kinase (p58IPK- interacting protein) (58 kDa interferon-induced protein kinase- interacting protein) (P52rIPK) (Death-associated protein 4) (THAP domain-containing protein 0). | |||||
|
RET2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 75) | NC score | 0.006286 (rank : 70) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P50120, Q6ISQ9, Q6ISS7 | Gene names | RBP2, CRBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinol-binding protein II, cellular (CRBP-II). | |||||
|
CLCKA_MOUSE
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.004087 (rank : 76) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9WUB7, Q6UB69, Q8JZU7 | Gene names | Clcnka, Clcnk1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride channel protein ClC-Ka (Chloride channel Ka) (ClC-K1). | |||||
|
DCDC2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.003865 (rank : 77) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5DU00, Q5SZU0, Q80Y99 | Gene names | Dcdc2, Kiaa1154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublecortin domain-containing protein 2. | |||||
|
NAL13_HUMAN
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.007966 (rank : 68) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q86W25, Q7RTR5 | Gene names | NALP13, NOD14 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 13 (Nucleotide-binding oligomerization domain protein 14). | |||||
|
WDR60_MOUSE
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.002941 (rank : 83) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C761, Q3UNM4 | Gene names | Wdr60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 60. | |||||
|
ZNFX1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.001766 (rank : 84) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 407 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R151, Q3U016, Q3UD71, Q3UEZ0 | Gene names | ZNFX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
CLCKA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 81) | NC score | 0.003766 (rank : 78) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P51800, Q7Z6D1, Q86VT1 | Gene names | CLCNKA | |||
|
Domain Architecture |

|
|||||
| Description | Chloride channel protein ClC-Ka (Chloride channel Ka) (ClC-K1). | |||||
|
GGTA1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.005285 (rank : 73) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P23336, Q91V22 | Gene names | Ggta1, Ggta-1 | |||
|
Domain Architecture |

|
|||||
| Description | N-acetyllactosaminide alpha-1,3-galactosyltransferase (EC 2.4.1.87) (Galactosyltransferase) (UDP-galactose:beta-D-galactosyl-1,4-N-acetyl- D-glucosaminide alpha-1,3-galactosyltransferase). | |||||
|
NCOA7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 83) | NC score | 0.003478 (rank : 82) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NI08, Q3LID6, Q4G0V1, Q5TF95, Q6IPQ4, Q6NE83, Q86T89, Q8N1W4 | Gene names | NCOA7, ERAP140, ESNA1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 7 (140 kDa estrogen receptor-associated protein) (Estrogen nuclear receptor coactivator 1). | |||||
|
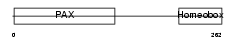
PAX6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | -0.001167 (rank : 86) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P26367, Q6N006, Q99413 | Gene names | PAX6, AN2 | |||
|
Domain Architecture |
|
|||||
| Description | Paired box protein Pax-6 (Oculorhombin) (Aniridia type II protein). | |||||
|
PAX6_MOUSE
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | -0.001167 (rank : 87) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P63015, P32117, P70601, Q62222, Q64037, Q8CEI5, Q8VDB5, Q921Q8 | Gene names | Pax6, Pax-6, Sey | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paired box protein Pax-6 (Oculorhombin). | |||||
|
RET1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.005212 (rank : 74) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P09455 | Gene names | RBP1, CRBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinol-binding protein I, cellular (Cellular retinol-binding protein) (CRBP). | |||||
|
RET1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.005192 (rank : 75) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q00915 | Gene names | Rbp1, Crbpi, Rbp-1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinol-binding protein I, cellular (Cellular retinol-binding protein) (CRBP) (mCRBPI). | |||||
|
SMC2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | -0.003456 (rank : 88) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1131 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CG48, Q61076, Q9CS17, Q9CSD8 | Gene names | Smc2l1, Cape, Fin16, Smc2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 2-like 1 protein (Chromosome- associated protein E) (XCAP-E homolog) (FGF-inducible protein 16). | |||||
|
AT2B1_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 88 | |
| SwissProt Accessions | P20020, Q12992, Q12993, Q13819, Q13820, Q13821, Q16504, Q93082 | Gene names | ATP2B1, PMCA1 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 1 (EC 3.6.3.8) (PMCA1) (Plasma membrane calcium pump isoform 1) (Plasma membrane calcium ATPase isoform 1). | |||||
|
AT2B3_HUMAN
|
||||||
| NC score | 0.997244 (rank : 2) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q16720, Q12995, Q16858 | Gene names | ATP2B3 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 3 (EC 3.6.3.8) (PMCA3) (Plasma membrane calcium pump isoform 3) (Plasma membrane calcium ATPase isoform 3). | |||||
|
AT2B2_MOUSE
|
||||||
| NC score | 0.996944 (rank : 3) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9R0K7, O88863 | Gene names | Atp2b2, Pmca2 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 2 (EC 3.6.3.8) (PMCA2) (Plasma membrane calcium pump isoform 2) (Plasma membrane calcium ATPase isoform 2). | |||||
|
AT2B2_HUMAN
|
||||||
| NC score | 0.996760 (rank : 4) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q01814, O00766, Q12994, Q16818 | Gene names | ATP2B2, PMCA2 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 2 (EC 3.6.3.8) (PMCA2) (Plasma membrane calcium pump isoform 2) (Plasma membrane calcium ATPase isoform 2). | |||||
|
AT2B4_HUMAN
|
||||||
| NC score | 0.996338 (rank : 5) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P23634, Q13450, Q13452, Q13455, Q16817 | Gene names | ATP2B4 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 4 (EC 3.6.3.8) (PMCA4) (Plasma membrane calcium pump isoform 4) (Plasma membrane calcium ATPase isoform 4). | |||||
|
AT2C1_MOUSE
|
||||||
| NC score | 0.914615 (rank : 6) | θ value | 4.83544e-70 (rank : 12) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q80XR2, Q80YZ2 | Gene names | Atp2c1, Pmr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-transporting ATPase type 2C member 1 (EC 3.6.3.8) (ATPase 2C1) (ATP-dependent Ca(2+) pump PMR1). | |||||
|
AT2C1_HUMAN
|
||||||
| NC score | 0.913840 (rank : 7) | θ value | 3.13423e-69 (rank : 13) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P98194, O76005, Q86V72, Q86V73, Q8N6V1, Q8NCJ7 | Gene names | ATP2C1, KIAA1347, PMR1L | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-transporting ATPase type 2C member 1 (EC 3.6.3.8) (ATPase 2C1) (ATP-dependent Ca(2+) pump PMR1). | |||||
|
AT2A3_HUMAN
|
||||||
| NC score | 0.906906 (rank : 8) | θ value | 4.08394e-77 (rank : 8) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q93084, O60900, O60901, O75501, O75502, Q16115, Q8TEX5, Q8TEX6 | Gene names | ATP2A3 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 3 (EC 3.6.3.8) (Calcium pump 3) (SERCA3) (SR Ca(2+)-ATPase 3). | |||||
|
AT2A2_HUMAN
|
||||||
| NC score | 0.906004 (rank : 9) | θ value | 4.82422e-78 (rank : 7) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P16615, P16614 | Gene names | ATP2A2, ATP2B | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 2 (EC 3.6.3.8) (Calcium pump 2) (SERCA2) (SR Ca(2+)-ATPase 2) (Calcium-transporting ATPase sarcoplasmic reticulum type, slow twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2A2_MOUSE
|
||||||
| NC score | 0.905983 (rank : 10) | θ value | 2.82823e-78 (rank : 6) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O55143, Q9R2A9, Q9WUT5 | Gene names | Atp2a2 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 2 (EC 3.6.3.8) (Calcium pump 2) (SERCA2) (SR Ca(2+)-ATPase 2) (Calcium-transporting ATPase sarcoplasmic reticulum type, slow twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2A3_MOUSE
|
||||||
| NC score | 0.905112 (rank : 11) | θ value | 2.64714e-76 (rank : 9) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q64518, O70625, Q64517 | Gene names | Atp2a3 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 3 (EC 3.6.3.8) (Calcium pump 3) (SERCA3) (SR Ca(2+)-ATPase 3). | |||||
|
AT2C2_HUMAN
|
||||||
| NC score | 0.903676 (rank : 12) | θ value | 1.50663e-63 (rank : 16) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O75185 | Gene names | KIAA0703 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable calcium-transporting ATPase KIAA0703 (EC 3.6.3.8). | |||||
|
AT2A1_MOUSE
|
||||||
| NC score | 0.902955 (rank : 13) | θ value | 3.34864e-71 (rank : 11) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8R429 | Gene names | Atp2a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 1 (EC 3.6.3.8) (Calcium pump 1) (SERCA1) (SR Ca(2+)-ATPase 1) (Calcium-transporting ATPase sarcoplasmic reticulum type, fast twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2A1_HUMAN
|
||||||
| NC score | 0.902820 (rank : 14) | θ value | 5.1662e-72 (rank : 10) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O14983, O14984 | Gene names | ATP2A1 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 1 (EC 3.6.3.8) (Calcium pump 1) (SERCA1) (SR Ca(2+)-ATPase 1) (Calcium-transporting ATPase sarcoplasmic reticulum type, fast twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT12A_HUMAN
|
||||||
| NC score | 0.884334 (rank : 15) | θ value | 6.11685e-65 (rank : 14) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P54707, Q13816, Q13817, Q16734 | Gene names | ATP12A, ATP1AL1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 2 (EC 3.6.3.10) (Proton pump) (Non-gastric H(+)/K(+) ATPase subunit alpha). | |||||
|
AT12A_MOUSE
|
||||||
| NC score | 0.883092 (rank : 16) | θ value | 3.03575e-64 (rank : 15) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9Z1W8, Q8VHY2 | Gene names | Atp12a, Atp1al1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium-transporting ATPase alpha chain 2 (EC 3.6.3.10) (Proton pump) (Non-gastric H(+)/K(+) ATPase subunit alpha). | |||||
|
AT1A4_MOUSE
|
||||||
| NC score | 0.875994 (rank : 17) | θ value | 2.17557e-62 (rank : 17) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9WV27, Q9R173, Q9WV28 | Gene names | Atp1a4, Atp1al2 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-4 chain (EC 3.6.3.9) (Sodium pump 4) (Na+/K+ ATPase 4). | |||||
|
ATP4A_MOUSE
|
||||||
| NC score | 0.875717 (rank : 18) | θ value | 2.10721e-57 (rank : 25) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q64436, Q9CV46 | Gene names | Atp4a | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 1 (EC 3.6.3.10) (Proton pump) (Gastric H+/K+ ATPase subunit alpha). | |||||
|
ATP4A_HUMAN
|
||||||
| NC score | 0.873995 (rank : 19) | θ value | 6.13105e-57 (rank : 26) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P20648, O00738 | Gene names | ATP4A | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 1 (EC 3.6.3.10) (Proton pump) (Gastric H+/K+ ATPase subunit alpha). | |||||
|
AT1A1_MOUSE
|
||||||
| NC score | 0.871803 (rank : 20) | θ value | 7.73779e-60 (rank : 19) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8VDN2, Q91Z09 | Gene names | Atp1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-1 chain precursor (EC 3.6.3.9) (Sodium pump 1) (Na+/K+ ATPase 1). | |||||
|
AT1A1_HUMAN
|
||||||
| NC score | 0.871240 (rank : 21) | θ value | 9.1404e-61 (rank : 18) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P05023, Q16689, Q6LDM4, Q9UJ20, Q9UJ21 | Gene names | ATP1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-1 chain precursor (EC 3.6.3.9) (Sodium pump 1) (Na+/K+ ATPase 1). | |||||
|
AT1A4_HUMAN
|
||||||
| NC score | 0.870785 (rank : 22) | θ value | 5.0155e-59 (rank : 20) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q13733, Q504T2, Q8TBN8, Q8WXA7, Q8WXH7, Q8WY13 | Gene names | ATP1A4, ATP1AL2 | |||
|
Domain Architecture |
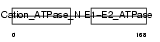
|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-4 chain (EC 3.6.3.9) (Sodium pump 4) (Na+/K+ ATPase 4). | |||||
|
AT1A2_MOUSE
|
||||||
| NC score | 0.868293 (rank : 23) | θ value | 7.24236e-58 (rank : 24) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q6PIE5 | Gene names | Atp1a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2) (Alpha(+)). | |||||
|
AT1A2_HUMAN
|
||||||
| NC score | 0.867621 (rank : 24) | θ value | 5.54529e-58 (rank : 23) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P50993, Q07059, Q86UZ5, Q9UQ25 | Gene names | ATP1A2, KIAA0778 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2). | |||||
|
AT1A3_HUMAN
|
||||||
| NC score | 0.867326 (rank : 25) | θ value | 6.55044e-59 (rank : 21) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P13637, Q16732, Q16735, Q969K5 | Gene names | ATP1A3 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-3 chain (EC 3.6.3.9) (Sodium pump 3) (Na+/K+ ATPase 3) (Alpha(III)). | |||||
|
AT1A3_MOUSE
|
||||||
| NC score | 0.866706 (rank : 26) | θ value | 6.55044e-59 (rank : 22) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q6PIC6 | Gene names | Atp1a3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium/potassium-transporting ATPase alpha-3 chain (EC 3.6.3.9) (Sodium pump 3) (Na+/K+ ATPase 3) (Alpha(III)). | |||||
|
AT132_MOUSE
|
||||||
| NC score | 0.715363 (rank : 27) | θ value | 1.24977e-17 (rank : 29) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9CTG6, Q8CG98 | Gene names | Atp13a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable cation-transporting ATPase 13A2 (EC 3.6.3.-). | |||||
|
AT133_HUMAN
|
||||||
| NC score | 0.709564 (rank : 28) | θ value | 6.64225e-11 (rank : 48) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9H7F0, Q8NC11, Q96KS1 | Gene names | ATP13A3, AFURS1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A3 (EC 3.6.3.-) (ATPase family homolog up-regulated in senescence cells 1). | |||||
|
AT132_HUMAN
|
||||||
| NC score | 0.707776 (rank : 29) | θ value | 7.32683e-18 (rank : 28) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9NQ11, O75700, Q5JXY2 | Gene names | ATP13A2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A2 (EC 3.6.3.-). | |||||
|
AT131_HUMAN
|
||||||
| NC score | 0.685360 (rank : 30) | θ value | 1.38499e-08 (rank : 51) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9HD20, Q9H6C6 | Gene names | ATP13A1, ATP13A, KIAA1825 | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A1 (EC 3.6.3.-). | |||||
|
AT131_MOUSE
|
||||||
| NC score | 0.674284 (rank : 31) | θ value | 1.38499e-08 (rank : 52) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9EPE9 | Gene names | Atp13a1, Atp13a | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A1 (EC 3.6.3.-) (CATP). | |||||
|
ATP7A_HUMAN
|
||||||
| NC score | 0.619167 (rank : 32) | θ value | 5.8054e-15 (rank : 34) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q04656, O00227, O00745, Q9BYY8 | Gene names | ATP7A, MC1, MNK | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 1 (EC 3.6.3.4) (Copper pump 1) (Menkes disease-associated protein). | |||||
|
ATP7B_MOUSE
|
||||||
| NC score | 0.616327 (rank : 33) | θ value | 4.02038e-16 (rank : 30) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q64446 | Gene names | Atp7b, Wnd | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 2 (EC 3.6.3.4) (Copper pump 2) (Wilson disease-associated protein homolog). | |||||
|
ATP7A_MOUSE
|
||||||
| NC score | 0.615612 (rank : 34) | θ value | 2.60593e-15 (rank : 33) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q64430, O35101, P97422, Q64431 | Gene names | Atp7a, Mnk | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 1 (EC 3.6.3.4) (Copper pump 1) (Menkes disease-associated protein homolog). | |||||
|
ATP7B_HUMAN
|
||||||
| NC score | 0.610823 (rank : 35) | θ value | 2.20605e-14 (rank : 35) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P35670, Q16318, Q16319, Q4U3V3, Q59FJ9, Q5T7X7 | Gene names | ATP7B, PWD, WC1, WND | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 2 (EC 3.6.3.4) (Copper pump 2) (Wilson disease-associated protein). | |||||
|
AT8A2_HUMAN
|
||||||
| NC score | 0.534401 (rank : 36) | θ value | 1.2105e-12 (rank : 40) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9NTI2, Q9H527, Q9NPU6, Q9NTL2, Q9NYM3 | Gene names | ATP8A2, ATPIB | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IB (EC 3.6.3.1) (ATPase class I type 8A member 2) (ML-1). | |||||
|
AT8A2_MOUSE
|
||||||
| NC score | 0.534299 (rank : 37) | θ value | 3.52202e-12 (rank : 44) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98200 | Gene names | Atp8a2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IB (EC 3.6.3.1) (ATPase class I type 8A member 2). | |||||
|
AT11B_HUMAN
|
||||||
| NC score | 0.533508 (rank : 38) | θ value | 3.76295e-14 (rank : 36) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9Y2G3, Q96FN1, Q9UKK7 | Gene names | ATP11B, ATPIF, ATPIR, KIAA0956 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IF (EC 3.6.3.1) (ATPase class I type 11B) (ATPase IR). | |||||
|
AT8A1_MOUSE
|
||||||
| NC score | 0.527759 (rank : 39) | θ value | 7.09661e-13 (rank : 38) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P70704 | Gene names | Atp8a1, Atpc1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IA (EC 3.6.3.1) (Chromaffin granule ATPase II) (ATPase class I type 8A member 1). | |||||
|
AT8A1_HUMAN
|
||||||
| NC score | 0.524297 (rank : 40) | θ value | 3.52202e-12 (rank : 43) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Y2Q0 | Gene names | ATP8A1, ATPIA | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IA (EC 3.6.3.1) (Chromaffin granule ATPase II) (ATPase class I type 8A member 1). | |||||
|
ATP9A_HUMAN
|
||||||
| NC score | 0.523561 (rank : 41) | θ value | 7.59969e-07 (rank : 53) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O75110, Q5TFW5, Q5TFW6, Q5TFW9, Q6ZMF3, Q9NQK6, Q9NQK7 | Gene names | ATP9A, ATPIIA, KIAA0611 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIA (EC 3.6.3.1) (ATPase class II type 9A) (ATPase IIA). | |||||
|
ATP9A_MOUSE
|
||||||
| NC score | 0.512211 (rank : 42) | θ value | 5.44631e-05 (rank : 56) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O70228, Q8VDI5, Q922L9 | Gene names | Atp9a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIA (EC 3.6.3.1) (ATPase class II type 9A). | |||||
|
ATP9B_HUMAN
|
||||||
| NC score | 0.510834 (rank : 43) | θ value | 8.40245e-06 (rank : 54) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O43861, O60872 | Gene names | ATP9B, ATPIIB, NEO1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIB (EC 3.6.3.1). | |||||
|
AT10D_MOUSE
|
||||||
| NC score | 0.509684 (rank : 44) | θ value | 2.28291e-11 (rank : 47) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8K2X1 | Gene names | Atp10d | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VD (EC 3.6.3.1) (ATPVD). | |||||
|
AT11A_HUMAN
|
||||||
| NC score | 0.507181 (rank : 45) | θ value | 3.88503e-19 (rank : 27) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98196, Q5VXT2 | Gene names | ATP11A, ATPIH, ATPIS, KIAA1021 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IH (EC 3.6.3.1) (ATPase class I type 11A) (ATPase IS). | |||||
|
AT10D_HUMAN
|
||||||
| NC score | 0.506319 (rank : 46) | θ value | 2.69671e-12 (rank : 42) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9P241, Q8NC70, Q96SR3 | Gene names | ATP10D, KIAA1487 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VD (EC 3.6.3.1) (ATPVD). | |||||
|
AT10B_HUMAN
|
||||||
| NC score | 0.502749 (rank : 47) | θ value | 1.16975e-15 (rank : 31) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O94823, Q9H725 | Gene names | ATP10B, ATPVB, KIAA0715 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VB (EC 3.6.3.1). | |||||
|
AT11C_HUMAN
|
||||||
| NC score | 0.502265 (rank : 48) | θ value | 2.43908e-13 (rank : 37) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8NB49, Q5JT69, Q5JT70, Q5JT71, Q5JT72, Q5JT73, Q6ZND5, Q6ZU50, Q6ZUP7, Q70IJ9, Q70IK0, Q8WX24 | Gene names | ATP11C, ATPIG, ATPIQ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IG (EC 3.6.3.1) (ATPase class I type 11C) (ATPase IG) (ATPase IQ) (ATPase class VI type 11C). | |||||
|
ATP9B_MOUSE
|
||||||
| NC score | 0.501504 (rank : 49) | θ value | 1.09739e-05 (rank : 55) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98195, Q99LI3 | Gene names | Atp9b | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IIB (EC 3.6.3.1). | |||||
|
AT8B3_HUMAN
|
||||||
| NC score | 0.493482 (rank : 50) | θ value | 3.52202e-12 (rank : 45) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O60423, Q8IVB8, Q8N4Y8, Q96M22 | Gene names | ATP8B3, ATP1K, FOS37502_2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IK (EC 3.6.3.1) (ATPase class I type 8B member 3). | |||||
|
AT8B4_HUMAN
|
||||||
| NC score | 0.491768 (rank : 51) | θ value | 1.133e-10 (rank : 49) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8TF62, Q9H727 | Gene names | ATP8B4, KIAA1939 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IM (EC 3.6.3.1) (ATPase class I type 8B member 4). | |||||
|
AT11A_MOUSE
|
||||||
| NC score | 0.490784 (rank : 52) | θ value | 1.52774e-15 (rank : 32) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98197 | Gene names | Atp11a | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IH (EC 3.6.3.1) (ATPase class I type 11A) (ATPase IS). | |||||
|
AT8B1_HUMAN
|
||||||
| NC score | 0.490505 (rank : 53) | θ value | 9.26847e-13 (rank : 39) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O43520, Q9BTP8 | Gene names | ATP8B1, ATPIC, FIC1, PFIC | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IC (EC 3.6.3.1) (Familial intrahepatic cholestasis type 1) (ATPase class I type 8B member 1). | |||||
|
AT10A_HUMAN
|
||||||
| NC score | 0.489940 (rank : 54) | θ value | 6.00763e-12 (rank : 46) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O60312, Q969I4 | Gene names | ATP10A, ATP10C, ATPVC, KIAA0566 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (ATPVA) (Aminophospholipid translocase VA). | |||||
|
AT10A_MOUSE
|
||||||
| NC score | 0.488821 (rank : 55) | θ value | 5.62301e-10 (rank : 50) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O54827, Q8R3B8 | Gene names | Atp10a, Atpc5, Pfatp | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (P-locus fat-associated ATPase). | |||||
|
AT8B2_HUMAN
|
||||||
| NC score | 0.485555 (rank : 56) | θ value | 1.2105e-12 (rank : 41) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P98198, Q96I43, Q96NQ7 | Gene names | ATP8B2, ATPID, KIAA1137 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase ID (EC 3.6.3.1) (ATPase class I type 8B member 2). | |||||
|
K0195_MOUSE
|
||||||
| NC score | 0.078559 (rank : 57) | θ value | 0.21417 (rank : 59) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TSH8, Q80U66, Q8BL05, Q8K0Q0, Q91VY1 | Gene names | Kiaa0195 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0195. | |||||
|
K0195_HUMAN
|
||||||
| NC score | 0.078390 (rank : 58) | θ value | 0.21417 (rank : 58) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q12767, O75536, Q86XF1 | Gene names | KIAA0195, TMEM94 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0195 (Transmembrane protein 94). | |||||
|
PRP31_MOUSE
|
||||||
| NC score | 0.025753 (rank : 59) | θ value | 0.47712 (rank : 61) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CCF0, Q6P7X2, Q8BQ91, Q8C8U4, Q8C8V5, Q8CCG6, Q8CF52, Q8VBW3 | Gene names | Prpf31, Prp31 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6 small nuclear ribonucleoprotein Prp31 (Pre-mRNA-processing factor 31) (U4/U6 snRNP 61 kDa protein) (Protein 61K). | |||||
|
PRP31_HUMAN
|
||||||
| NC score | 0.025155 (rank : 60) | θ value | 0.62314 (rank : 63) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WWY3, Q8N7F9, Q9H271, Q9Y439 | Gene names | PRPF31, PRP31 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6 small nuclear ribonucleoprotein Prp31 (Pre-mRNA-processing factor 31) (U4/U6 snRNP 61 kDa protein) (hPrp31) (Protein 61K) (Serologically defined breast cancer antigen NY-BR-99). | |||||
|
PVRL2_HUMAN
|
||||||
| NC score | 0.023173 (rank : 61) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92692, O75455, Q96J29 | Gene names | PVRL2, HVEB, PRR2 | |||
|
Domain Architecture |

|
|||||
| Description | Poliovirus receptor-related protein 2 precursor (Herpes virus entry mediator B) (HveB) (Nectin-2) (CD112 antigen). | |||||
|
YETS2_MOUSE
|
||||||
| NC score | 0.019063 (rank : 62) | θ value | 0.47712 (rank : 62) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3TUF7, Q6PGF8, Q80TI2, Q8CG86 | Gene names | Yeats2, Kiaa1197 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YEATS domain-containing protein 2. | |||||
|
YETS2_HUMAN
|
||||||
| NC score | 0.017376 (rank : 63) | θ value | 2.36792 (rank : 69) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ULM3, Q641P6, Q9NW96 | Gene names | YEATS2, KIAA1197 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YEATS domain-containing protein 2. | |||||
|
AHI1_HUMAN
|
||||||
| NC score | 0.015533 (rank : 64) | θ value | 2.36792 (rank : 66) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 537 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8N157, Q504T3, Q5TCP9, Q6P098, Q6PIT6, Q8NDX0, Q9H0H2 | Gene names | AHI1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Jouberin (Abelson helper integration site 1 protein homolog) (AHI-1). | |||||
|
ZN452_HUMAN
|
||||||
| NC score | 0.013545 (rank : 65) | θ value | 0.279714 (rank : 60) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6R2W3, Q2NKL9, Q5SRJ3, Q8TCN2, Q96PW3 | Gene names | ZNF452, KIAA1925 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ZNF452. | |||||
|
TTF2_MOUSE
|
||||||
| NC score | 0.012234 (rank : 66) | θ value | 0.0961366 (rank : 57) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5NC05, Q5M924 | Gene names | Ttf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 2 (EC 3.6.1.-) (RNA polymerase II termination factor) (Transcription release factor 2). | |||||
|
HDGF_MOUSE
|
||||||
| NC score | 0.009473 (rank : 67) | θ value | 2.36792 (rank : 67) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P51859, Q8BPG7, Q9CYA4, Q9JK87 | Gene names | Hdgf, Tdrm1 | |||
|
Domain Architecture |

|
|||||
| Description | Hepatoma-derived growth factor (HDGF). | |||||
|
NAL13_HUMAN
|
||||||
| NC score | 0.007966 (rank : 68) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q86W25, Q7RTR5 | Gene names | NALP13, NOD14 | |||
|
Domain Architecture |

|
|||||
| Description | NACHT, LRR and PYD-containing protein 13 (Nucleotide-binding oligomerization domain protein 14). | |||||
|
P52K_HUMAN
|
||||||
| NC score | 0.007141 (rank : 69) | θ value | 5.27518 (rank : 74) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43422, Q8WTW1, Q9Y3Z4 | Gene names | PRKRIR, DAP4, P52RIPK, THAP0 | |||
|
Domain Architecture |

|
|||||
| Description | 52 kDa repressor of the inhibitor of the protein kinase (p58IPK- interacting protein) (58 kDa interferon-induced protein kinase- interacting protein) (P52rIPK) (Death-associated protein 4) (THAP domain-containing protein 0). | |||||
|
RET2_HUMAN
|
||||||
| NC score | 0.006286 (rank : 70) | θ value | 5.27518 (rank : 75) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P50120, Q6ISQ9, Q6ISS7 | Gene names | RBP2, CRBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinol-binding protein II, cellular (CRBP-II). | |||||
|
SPRL1_MOUSE
|
||||||
| NC score | 0.005815 (rank : 71) | θ value | 3.0926 (rank : 71) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P70663, P97810, Q99L82 | Gene names | Sparcl1, Ecm2, Sc1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (Matrix glycoprotein Sc1) (Extracellular matrix protein 2). | |||||
|
COA2_HUMAN
|
||||||
| NC score | 0.005517 (rank : 72) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O00763, Q16852 | Gene names | ACACB, ACC2, ACCB | |||
|
Domain Architecture |

|
|||||
| Description | Acetyl-CoA carboxylase 2 (EC 6.4.1.2) (ACC-beta) [Includes: Biotin carboxylase (EC 6.3.4.14)]. | |||||
|
GGTA1_MOUSE
|
||||||
| NC score | 0.005285 (rank : 73) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P23336, Q91V22 | Gene names | Ggta1, Ggta-1 | |||
|
Domain Architecture |

|
|||||
| Description | N-acetyllactosaminide alpha-1,3-galactosyltransferase (EC 2.4.1.87) (Galactosyltransferase) (UDP-galactose:beta-D-galactosyl-1,4-N-acetyl- D-glucosaminide alpha-1,3-galactosyltransferase). | |||||
|
RET1_HUMAN
|
||||||
| NC score | 0.005212 (rank : 74) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P09455 | Gene names | RBP1, CRBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinol-binding protein I, cellular (Cellular retinol-binding protein) (CRBP). | |||||
|
RET1_MOUSE
|
||||||
| NC score | 0.005192 (rank : 75) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q00915 | Gene names | Rbp1, Crbpi, Rbp-1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinol-binding protein I, cellular (Cellular retinol-binding protein) (CRBP) (mCRBPI). | |||||
|
CLCKA_MOUSE
|
||||||
| NC score | 0.004087 (rank : 76) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9WUB7, Q6UB69, Q8JZU7 | Gene names | Clcnka, Clcnk1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride channel protein ClC-Ka (Chloride channel Ka) (ClC-K1). | |||||
|
DCDC2_MOUSE
|
||||||
| NC score | 0.003865 (rank : 77) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5DU00, Q5SZU0, Q80Y99 | Gene names | Dcdc2, Kiaa1154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublecortin domain-containing protein 2. | |||||
|
CLCKA_HUMAN
|
||||||
| NC score | 0.003766 (rank : 78) | θ value | 8.99809 (rank : 81) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P51800, Q7Z6D1, Q86VT1 | Gene names | CLCNKA | |||
|
Domain Architecture |

|
|||||
| Description | Chloride channel protein ClC-Ka (Chloride channel Ka) (ClC-K1). | |||||
|
MBB1A_HUMAN
|
||||||
| NC score | 0.003710 (rank : 79) | θ value | 5.27518 (rank : 73) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
BCOR_HUMAN
|
||||||
| NC score | 0.003536 (rank : 80) | θ value | 1.38821 (rank : 64) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6W2J9, Q6P4B6, Q7Z2K7, Q8TEB4, Q96DB3, Q9H232, Q9H233, Q9HCJ7, Q9NXF2 | Gene names | BCOR, KIAA1575 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BCoR protein (BCL-6 corepressor). | |||||
|
RRBP1_MOUSE
|
||||||
| NC score | 0.003484 (rank : 81) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
NCOA7_HUMAN
|
||||||
| NC score | 0.003478 (rank : 82) | θ value | 8.99809 (rank : 83) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NI08, Q3LID6, Q4G0V1, Q5TF95, Q6IPQ4, Q6NE83, Q86T89, Q8N1W4 | Gene names | NCOA7, ERAP140, ESNA1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 7 (140 kDa estrogen receptor-associated protein) (Estrogen nuclear receptor coactivator 1). | |||||
|
WDR60_MOUSE
|
||||||
| NC score | 0.002941 (rank : 83) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C761, Q3UNM4 | Gene names | Wdr60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 60. | |||||
|
ZNFX1_MOUSE
|
||||||
| NC score | 0.001766 (rank : 84) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 407 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R151, Q3U016, Q3UD71, Q3UEZ0 | Gene names | ZNFX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
PHLPP_HUMAN
|
||||||
| NC score | 0.000399 (rank : 85) | θ value | 3.0926 (rank : 70) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 604 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60346, Q641Q7, Q6P4C4, Q6PJI6, Q86TN6, Q96FK2, Q9NUY1 | Gene names | PHLPP, KIAA0606, PLEKHE1, SCOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein) (hSCOP). | |||||
|
PAX6_HUMAN
|
||||||
| NC score | -0.001167 (rank : 86) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P26367, Q6N006, Q99413 | Gene names | PAX6, AN2 | |||
|
Domain Architecture |

|
|||||
| Description | Paired box protein Pax-6 (Oculorhombin) (Aniridia type II protein). | |||||
|
PAX6_MOUSE
|
||||||
| NC score | -0.001167 (rank : 87) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P63015, P32117, P70601, Q62222, Q64037, Q8CEI5, Q8VDB5, Q921Q8 | Gene names | Pax6, Pax-6, Sey | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paired box protein Pax-6 (Oculorhombin). | |||||
|
SMC2_MOUSE
|
||||||
| NC score | -0.003456 (rank : 88) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1131 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CG48, Q61076, Q9CS17, Q9CSD8 | Gene names | Smc2l1, Cape, Fin16, Smc2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 2-like 1 protein (Chromosome- associated protein E) (XCAP-E homolog) (FGF-inducible protein 16). | |||||