Please be patient as the page loads
|
AT10D_HUMAN
|
||||||
| SwissProt Accessions | Q9P241, Q8NC70, Q96SR3 | Gene names | ATP10D, KIAA1487 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VD (EC 3.6.3.1) (ATPVD). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
AT10A_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.992064 (rank : 4) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O60312, Q969I4 | Gene names | ATP10A, ATP10C, ATPVC, KIAA0566 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (ATPVA) (Aminophospholipid translocase VA). | |||||
|
AT10A_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.991312 (rank : 5) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O54827, Q8R3B8 | Gene names | Atp10a, Atpc5, Pfatp | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (P-locus fat-associated ATPase). | |||||
|
AT10B_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.993605 (rank : 3) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O94823, Q9H725 | Gene names | ATP10B, ATPVB, KIAA0715 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VB (EC 3.6.3.1). | |||||
|
AT10D_HUMAN
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9P241, Q8NC70, Q96SR3 | Gene names | ATP10D, KIAA1487 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VD (EC 3.6.3.1) (ATPVD). | |||||
|
AT10D_MOUSE
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.998484 (rank : 2) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q8K2X1 | Gene names | Atp10d | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VD (EC 3.6.3.1) (ATPVD). | |||||
|
AT8B2_HUMAN
|
||||||
| θ value | 2.14053e-118 (rank : 6) | NC score | 0.946859 (rank : 7) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98198, Q96I43, Q96NQ7 | Gene names | ATP8B2, ATPID, KIAA1137 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase ID (EC 3.6.3.1) (ATPase class I type 8B member 2). | |||||
|
AT8B4_HUMAN
|
||||||
| θ value | 2.14053e-118 (rank : 7) | NC score | 0.947972 (rank : 6) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8TF62, Q9H727 | Gene names | ATP8B4, KIAA1939 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IM (EC 3.6.3.1) (ATPase class I type 8B member 4). | |||||
|
AT8A1_HUMAN
|
||||||
| θ value | 3.41741e-116 (rank : 8) | NC score | 0.943162 (rank : 8) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Y2Q0 | Gene names | ATP8A1, ATPIA | |||
|
Domain Architecture |
|
|||||
| Description | Probable phospholipid-transporting ATPase IA (EC 3.6.3.1) (Chromaffin granule ATPase II) (ATPase class I type 8A member 1). | |||||
|
AT8A1_MOUSE
|
||||||
| θ value | 9.94316e-116 (rank : 9) | NC score | 0.943040 (rank : 9) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P70704 | Gene names | Atp8a1, Atpc1 | |||
|
Domain Architecture |
|
|||||
| Description | Probable phospholipid-transporting ATPase IA (EC 3.6.3.1) (Chromaffin granule ATPase II) (ATPase class I type 8A member 1). | |||||
|
AT8A2_MOUSE
|
||||||
| θ value | 1.94052e-111 (rank : 10) | NC score | 0.942511 (rank : 12) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98200 | Gene names | Atp8a2 | |||
|
Domain Architecture |
|
|||||
| Description | Probable phospholipid-transporting ATPase IB (EC 3.6.3.1) (ATPase class I type 8A member 2). | |||||
|
AT8A2_HUMAN
|
||||||
| θ value | 3.31004e-111 (rank : 11) | NC score | 0.942678 (rank : 11) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9NTI2, Q9H527, Q9NPU6, Q9NTL2, Q9NYM3 | Gene names | ATP8A2, ATPIB | |||
|
Domain Architecture |
|
|||||
| Description | Probable phospholipid-transporting ATPase IB (EC 3.6.3.1) (ATPase class I type 8A member 2) (ML-1). | |||||
|
AT8B1_HUMAN
|
||||||
| θ value | 2.6227e-108 (rank : 12) | NC score | 0.942696 (rank : 10) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O43520, Q9BTP8 | Gene names | ATP8B1, ATPIC, FIC1, PFIC | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IC (EC 3.6.3.1) (Familial intrahepatic cholestasis type 1) (ATPase class I type 8B member 1). | |||||
|
AT11B_HUMAN
|
||||||
| θ value | 5.29683e-101 (rank : 13) | NC score | 0.939288 (rank : 14) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Y2G3, Q96FN1, Q9UKK7 | Gene names | ATP11B, ATPIF, ATPIR, KIAA0956 | |||
|
Domain Architecture |
|
|||||
| Description | Probable phospholipid-transporting ATPase IF (EC 3.6.3.1) (ATPase class I type 11B) (ATPase IR). | |||||
|
AT11A_HUMAN
|
||||||
| θ value | 6.91785e-101 (rank : 14) | NC score | 0.934713 (rank : 16) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98196, Q5VXT2 | Gene names | ATP11A, ATPIH, ATPIS, KIAA1021 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IH (EC 3.6.3.1) (ATPase class I type 11A) (ATPase IS). | |||||
|
AT11A_MOUSE
|
||||||
| θ value | 2.46047e-98 (rank : 15) | NC score | 0.933568 (rank : 17) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98197 | Gene names | Atp11a | |||
|
Domain Architecture |
|
|||||
| Description | Probable phospholipid-transporting ATPase IH (EC 3.6.3.1) (ATPase class I type 11A) (ATPase IS). | |||||
|
AT8B3_HUMAN
|
||||||
| θ value | 3.21347e-98 (rank : 16) | NC score | 0.941899 (rank : 13) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O60423, Q8IVB8, Q8N4Y8, Q96M22 | Gene names | ATP8B3, ATP1K, FOS37502_2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IK (EC 3.6.3.1) (ATPase class I type 8B member 3). | |||||
|
AT11C_HUMAN
|
||||||
| θ value | 1.94955e-95 (rank : 17) | NC score | 0.936706 (rank : 15) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8NB49, Q5JT69, Q5JT70, Q5JT71, Q5JT72, Q5JT73, Q6ZND5, Q6ZU50, Q6ZUP7, Q70IJ9, Q70IK0, Q8WX24 | Gene names | ATP11C, ATPIG, ATPIQ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IG (EC 3.6.3.1) (ATPase class I type 11C) (ATPase IG) (ATPase IQ) (ATPase class VI type 11C). | |||||
|
ATP9B_HUMAN
|
||||||
| θ value | 4.23606e-66 (rank : 18) | NC score | 0.903601 (rank : 19) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O43861, O60872 | Gene names | ATP9B, ATPIIB, NEO1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIB (EC 3.6.3.1). | |||||
|
ATP9A_HUMAN
|
||||||
| θ value | 2.10232e-65 (rank : 19) | NC score | 0.903097 (rank : 20) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O75110, Q5TFW5, Q5TFW6, Q5TFW9, Q6ZMF3, Q9NQK6, Q9NQK7 | Gene names | ATP9A, ATPIIA, KIAA0611 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIA (EC 3.6.3.1) (ATPase class II type 9A) (ATPase IIA). | |||||
|
ATP9B_MOUSE
|
||||||
| θ value | 2.10232e-65 (rank : 20) | NC score | 0.905686 (rank : 18) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98195, Q99LI3 | Gene names | Atp9b | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IIB (EC 3.6.3.1). | |||||
|
ATP9A_MOUSE
|
||||||
| θ value | 1.50663e-63 (rank : 21) | NC score | 0.899863 (rank : 21) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O70228, Q8VDI5, Q922L9 | Gene names | Atp9a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIA (EC 3.6.3.1) (ATPase class II type 9A). | |||||
|
AT2B4_HUMAN
|
||||||
| θ value | 9.56915e-18 (rank : 22) | NC score | 0.554825 (rank : 25) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P23634, Q13450, Q13452, Q13455, Q16817 | Gene names | ATP2B4 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 4 (EC 3.6.3.8) (PMCA4) (Plasma membrane calcium pump isoform 4) (Plasma membrane calcium ATPase isoform 4). | |||||
|
AT132_HUMAN
|
||||||
| θ value | 6.20254e-17 (rank : 23) | NC score | 0.580681 (rank : 23) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9NQ11, O75700, Q5JXY2 | Gene names | ATP13A2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A2 (EC 3.6.3.-). | |||||
|
AT132_MOUSE
|
||||||
| θ value | 1.058e-16 (rank : 24) | NC score | 0.579965 (rank : 24) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9CTG6, Q8CG98 | Gene names | Atp13a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable cation-transporting ATPase 13A2 (EC 3.6.3.-). | |||||
|
AT2B2_MOUSE
|
||||||
| θ value | 5.25075e-16 (rank : 25) | NC score | 0.529698 (rank : 28) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9R0K7, O88863 | Gene names | Atp2b2, Pmca2 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 2 (EC 3.6.3.8) (PMCA2) (Plasma membrane calcium pump isoform 2) (Plasma membrane calcium ATPase isoform 2). | |||||
|
AT133_HUMAN
|
||||||
| θ value | 1.52774e-15 (rank : 26) | NC score | 0.601840 (rank : 22) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9H7F0, Q8NC11, Q96KS1 | Gene names | ATP13A3, AFURS1 | |||
|
Domain Architecture |
|
|||||
| Description | Probable cation-transporting ATPase 13A3 (EC 3.6.3.-) (ATPase family homolog up-regulated in senescence cells 1). | |||||
|
AT2B2_HUMAN
|
||||||
| θ value | 2.60593e-15 (rank : 27) | NC score | 0.533021 (rank : 27) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q01814, O00766, Q12994, Q16818 | Gene names | ATP2B2, PMCA2 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 2 (EC 3.6.3.8) (PMCA2) (Plasma membrane calcium pump isoform 2) (Plasma membrane calcium ATPase isoform 2). | |||||
|
AT2B3_HUMAN
|
||||||
| θ value | 7.58209e-15 (rank : 28) | NC score | 0.534131 (rank : 26) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q16720, Q12995, Q16858 | Gene names | ATP2B3 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 3 (EC 3.6.3.8) (PMCA3) (Plasma membrane calcium pump isoform 3) (Plasma membrane calcium ATPase isoform 3). | |||||
|
AT2B1_HUMAN
|
||||||
| θ value | 2.69671e-12 (rank : 29) | NC score | 0.506319 (rank : 29) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P20020, Q12992, Q12993, Q13819, Q13820, Q13821, Q16504, Q93082 | Gene names | ATP2B1, PMCA1 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 1 (EC 3.6.3.8) (PMCA1) (Plasma membrane calcium pump isoform 1) (Plasma membrane calcium ATPase isoform 1). | |||||
|
AT2C1_MOUSE
|
||||||
| θ value | 3.08544e-08 (rank : 30) | NC score | 0.398442 (rank : 36) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q80XR2, Q80YZ2 | Gene names | Atp2c1, Pmr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-transporting ATPase type 2C member 1 (EC 3.6.3.8) (ATPase 2C1) (ATP-dependent Ca(2+) pump PMR1). | |||||
|
AT12A_MOUSE
|
||||||
| θ value | 5.26297e-08 (rank : 31) | NC score | 0.397166 (rank : 37) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Z1W8, Q8VHY2 | Gene names | Atp12a, Atp1al1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium-transporting ATPase alpha chain 2 (EC 3.6.3.10) (Proton pump) (Non-gastric H(+)/K(+) ATPase subunit alpha). | |||||
|
AT2C1_HUMAN
|
||||||
| θ value | 1.53129e-07 (rank : 32) | NC score | 0.401073 (rank : 34) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98194, O76005, Q86V72, Q86V73, Q8N6V1, Q8NCJ7 | Gene names | ATP2C1, KIAA1347, PMR1L | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-transporting ATPase type 2C member 1 (EC 3.6.3.8) (ATPase 2C1) (ATP-dependent Ca(2+) pump PMR1). | |||||
|
AT12A_HUMAN
|
||||||
| θ value | 2.61198e-07 (rank : 33) | NC score | 0.400752 (rank : 35) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P54707, Q13816, Q13817, Q16734 | Gene names | ATP12A, ATP1AL1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 2 (EC 3.6.3.10) (Proton pump) (Non-gastric H(+)/K(+) ATPase subunit alpha). | |||||
|
AT2A2_HUMAN
|
||||||
| θ value | 3.41135e-07 (rank : 34) | NC score | 0.402560 (rank : 33) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P16615, P16614 | Gene names | ATP2A2, ATP2B | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 2 (EC 3.6.3.8) (Calcium pump 2) (SERCA2) (SR Ca(2+)-ATPase 2) (Calcium-transporting ATPase sarcoplasmic reticulum type, slow twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2A2_MOUSE
|
||||||
| θ value | 7.59969e-07 (rank : 35) | NC score | 0.403814 (rank : 32) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O55143, Q9R2A9, Q9WUT5 | Gene names | Atp2a2 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 2 (EC 3.6.3.8) (Calcium pump 2) (SERCA2) (SR Ca(2+)-ATPase 2) (Calcium-transporting ATPase sarcoplasmic reticulum type, slow twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2A3_HUMAN
|
||||||
| θ value | 9.92553e-07 (rank : 36) | NC score | 0.397138 (rank : 38) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q93084, O60900, O60901, O75501, O75502, Q16115, Q8TEX5, Q8TEX6 | Gene names | ATP2A3 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 3 (EC 3.6.3.8) (Calcium pump 3) (SERCA3) (SR Ca(2+)-ATPase 3). | |||||
|
AT131_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 37) | NC score | 0.464222 (rank : 31) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9HD20, Q9H6C6 | Gene names | ATP13A1, ATP13A, KIAA1825 | |||
|
Domain Architecture |
|
|||||
| Description | Probable cation-transporting ATPase 13A1 (EC 3.6.3.-). | |||||
|
AT2A3_MOUSE
|
||||||
| θ value | 1.09739e-05 (rank : 38) | NC score | 0.386970 (rank : 40) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q64518, O70625, Q64517 | Gene names | Atp2a3 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 3 (EC 3.6.3.8) (Calcium pump 3) (SERCA3) (SR Ca(2+)-ATPase 3). | |||||
|
AT1A4_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 39) | NC score | 0.377231 (rank : 43) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q13733, Q504T2, Q8TBN8, Q8WXA7, Q8WXH7, Q8WY13 | Gene names | ATP1A4, ATP1AL2 | |||
|
Domain Architecture |
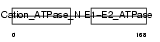
|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-4 chain (EC 3.6.3.9) (Sodium pump 4) (Na+/K+ ATPase 4). | |||||
|
AT1A3_HUMAN
|
||||||
| θ value | 3.19293e-05 (rank : 40) | NC score | 0.351827 (rank : 50) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P13637, Q16732, Q16735, Q969K5 | Gene names | ATP1A3 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-3 chain (EC 3.6.3.9) (Sodium pump 3) (Na+/K+ ATPase 3) (Alpha(III)). | |||||
|
AT1A3_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 41) | NC score | 0.349303 (rank : 51) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q6PIC6 | Gene names | Atp1a3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium/potassium-transporting ATPase alpha-3 chain (EC 3.6.3.9) (Sodium pump 3) (Na+/K+ ATPase 3) (Alpha(III)). | |||||
|
AT131_MOUSE
|
||||||
| θ value | 9.29e-05 (rank : 42) | NC score | 0.465080 (rank : 30) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9EPE9 | Gene names | Atp13a1, Atp13a | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A1 (EC 3.6.3.-) (CATP). | |||||
|
AT1A2_MOUSE
|
||||||
| θ value | 0.000158464 (rank : 43) | NC score | 0.369791 (rank : 49) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q6PIE5 | Gene names | Atp1a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2) (Alpha(+)). | |||||
|
AT1A1_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 44) | NC score | 0.371354 (rank : 47) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P05023, Q16689, Q6LDM4, Q9UJ20, Q9UJ21 | Gene names | ATP1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-1 chain precursor (EC 3.6.3.9) (Sodium pump 1) (Na+/K+ ATPase 1). | |||||
|
AT1A2_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 45) | NC score | 0.370141 (rank : 48) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P50993, Q07059, Q86UZ5, Q9UQ25 | Gene names | ATP1A2, KIAA0778 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2). | |||||
|
AT1A4_MOUSE
|
||||||
| θ value | 0.000270298 (rank : 46) | NC score | 0.376717 (rank : 44) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9WV27, Q9R173, Q9WV28 | Gene names | Atp1a4, Atp1al2 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-4 chain (EC 3.6.3.9) (Sodium pump 4) (Na+/K+ ATPase 4). | |||||
|
AT2A1_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 47) | NC score | 0.385960 (rank : 41) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O14983, O14984 | Gene names | ATP2A1 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 1 (EC 3.6.3.8) (Calcium pump 1) (SERCA1) (SR Ca(2+)-ATPase 1) (Calcium-transporting ATPase sarcoplasmic reticulum type, fast twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT1A1_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 48) | NC score | 0.371973 (rank : 46) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8VDN2, Q91Z09 | Gene names | Atp1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-1 chain precursor (EC 3.6.3.9) (Sodium pump 1) (Na+/K+ ATPase 1). | |||||
|
AT2A1_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 49) | NC score | 0.388676 (rank : 39) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8R429 | Gene names | Atp2a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 1 (EC 3.6.3.8) (Calcium pump 1) (SERCA1) (SR Ca(2+)-ATPase 1) (Calcium-transporting ATPase sarcoplasmic reticulum type, fast twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2C2_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 50) | NC score | 0.341419 (rank : 52) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O75185 | Gene names | KIAA0703 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable calcium-transporting ATPase KIAA0703 (EC 3.6.3.8). | |||||
|
ATP4A_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 51) | NC score | 0.380288 (rank : 42) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q64436, Q9CV46 | Gene names | Atp4a | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 1 (EC 3.6.3.10) (Proton pump) (Gastric H+/K+ ATPase subunit alpha). | |||||
|
ATP4A_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 52) | NC score | 0.375686 (rank : 45) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P20648, O00738 | Gene names | ATP4A | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 1 (EC 3.6.3.10) (Proton pump) (Gastric H+/K+ ATPase subunit alpha). | |||||
|
GP128_HUMAN
|
||||||
| θ value | 0.279714 (rank : 53) | NC score | 0.014596 (rank : 58) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96K78, Q86SQ2 | Gene names | GPR128 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 128 precursor. | |||||
|
OR2M2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 54) | NC score | -0.002797 (rank : 72) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 769 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96R28 | Gene names | OR2M2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Olfactory receptor 2M2 (OST423). | |||||
|
MTR1B_MOUSE
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | -0.000966 (rank : 70) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 831 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CIQ6, Q80T38, Q8K3K0 | Gene names | Mtnr1b | |||
|
Domain Architecture |
|
|||||
| Description | Melatonin receptor type 1B (Mel-1B-R) (Mel1b melatonin receptor). | |||||
|
RT31_HUMAN
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.013734 (rank : 59) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92665, Q5VYC8, Q8WTV8 | Gene names | MRPS31, IMOGN38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 28S ribosomal protein S31, mitochondrial precursor (S31mt) (MRP-S31) (Imogen 38). | |||||
|
BACH2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 57) | NC score | 0.002320 (rank : 65) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BYV9, Q59H70, Q5T793, Q9NTS5 | Gene names | BACH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription regulator protein BACH2 (BTB and CNC homolog 2). | |||||
|
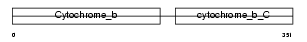
CYB_HUMAN
|
||||||
| θ value | 4.03905 (rank : 58) | NC score | 0.028766 (rank : 57) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P00156, Q34786, Q8HBR6, Q8HNQ0, Q8HNQ1, Q8HNQ9, Q8HNR4, Q8HNR7, Q8W7V8, Q8WCV9, Q8WCY2, Q8WCY7, Q8WCY8, Q9B1A6, Q9B1B6, Q9B1B8, Q9B1D4, Q9B1X6, Q9B2V0, Q9B2V8, Q9B2W0, Q9B2W3, Q9B2W8, Q9B2X1, Q9B2X7, Q9B2X9, Q9B2Y3, Q9B2Z0, Q9B2Z4, Q9T6H6, Q9T9Y0, Q9TEH4 | Gene names | MT-CYB, COB, CYTB, MTCYB | |||
|
Domain Architecture |
|
|||||
| Description | Cytochrome b. | |||||
|
BRD2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 59) | NC score | 0.005630 (rank : 61) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P25440, O00699, O00700, Q15310, Q5STC9, Q969U4 | Gene names | BRD2, KIAA9001, RING3 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 2 (Protein RING3) (O27.1.1). | |||||
|
CC1L1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | -0.002324 (rank : 71) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 629 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P51676 | Gene names | Ccr1l1, Cmkbr1l1 | |||
|
Domain Architecture |
|
|||||
| Description | C-C chemokine receptor 1-like protein 1 (Macrophage inflammatory protein 1 alpha receptor-like 1). | |||||
|
BRD4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | 0.002218 (rank : 67) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
KCNH2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.001813 (rank : 68) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35219, O35220, O35221, O35989 | Gene names | Kcnh2, Erg, Merg1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (ERG1) (MERG) (Merg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1). | |||||
|
SP4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 63) | NC score | -0.003448 (rank : 73) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 759 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q62445 | Gene names | Sp4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor Sp4. | |||||
|
ZC3H6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 64) | NC score | 0.004028 (rank : 64) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P61129 | Gene names | ZC3H6, KIAA2035, ZC3HDC6 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
ABI3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 65) | NC score | 0.004666 (rank : 62) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BYZ1, Q6PE63, Q9D7S4 | Gene names | Abi3, Nesh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ABI gene family member 3 (New molecule including SH3) (Nesh). | |||||
|
BRD4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 66) | NC score | 0.002284 (rank : 66) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
PDCL2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 67) | NC score | 0.004614 (rank : 63) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N4E4 | Gene names | PDCL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosducin-like protein 2. | |||||
|
T2FA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 68) | NC score | 0.008414 (rank : 60) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P35269, Q9BWN0 | Gene names | GTF2F1, RAP74 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor IIF alpha subunit (EC 2.7.11.1) (TFIIF-alpha) (Transcription initiation factor RAP74) (General transcription factor IIF polypeptide 1 74 kDa subunit protein). | |||||
|
TRI26_HUMAN
|
||||||
| θ value | 8.99809 (rank : 69) | NC score | -0.000690 (rank : 69) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q12899 | Gene names | TRIM26, RNF95, ZNF173 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 26 (Zinc finger protein 173) (Acid finger protein) (AFP) (RING finger protein 95). | |||||
|
ATP7A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.143279 (rank : 55) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q04656, O00227, O00745, Q9BYY8 | Gene names | ATP7A, MC1, MNK | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 1 (EC 3.6.3.4) (Copper pump 1) (Menkes disease-associated protein). | |||||
|
ATP7A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.140256 (rank : 56) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q64430, O35101, P97422, Q64431 | Gene names | Atp7a, Mnk | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 1 (EC 3.6.3.4) (Copper pump 1) (Menkes disease-associated protein homolog). | |||||
|
ATP7B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.147009 (rank : 54) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P35670, Q16318, Q16319, Q4U3V3, Q59FJ9, Q5T7X7 | Gene names | ATP7B, PWD, WC1, WND | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 2 (EC 3.6.3.4) (Copper pump 2) (Wilson disease-associated protein). | |||||
|
ATP7B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 73) | NC score | 0.147043 (rank : 53) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q64446 | Gene names | Atp7b, Wnd | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 2 (EC 3.6.3.4) (Copper pump 2) (Wilson disease-associated protein homolog). | |||||
|
AT10D_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9P241, Q8NC70, Q96SR3 | Gene names | ATP10D, KIAA1487 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VD (EC 3.6.3.1) (ATPVD). | |||||
|
AT10D_MOUSE
|
||||||
| NC score | 0.998484 (rank : 2) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q8K2X1 | Gene names | Atp10d | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VD (EC 3.6.3.1) (ATPVD). | |||||
|
AT10B_HUMAN
|
||||||
| NC score | 0.993605 (rank : 3) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O94823, Q9H725 | Gene names | ATP10B, ATPVB, KIAA0715 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VB (EC 3.6.3.1). | |||||
|
AT10A_HUMAN
|
||||||
| NC score | 0.992064 (rank : 4) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O60312, Q969I4 | Gene names | ATP10A, ATP10C, ATPVC, KIAA0566 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (ATPVA) (Aminophospholipid translocase VA). | |||||
|
AT10A_MOUSE
|
||||||
| NC score | 0.991312 (rank : 5) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O54827, Q8R3B8 | Gene names | Atp10a, Atpc5, Pfatp | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (P-locus fat-associated ATPase). | |||||
|
AT8B4_HUMAN
|
||||||
| NC score | 0.947972 (rank : 6) | θ value | 2.14053e-118 (rank : 7) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8TF62, Q9H727 | Gene names | ATP8B4, KIAA1939 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IM (EC 3.6.3.1) (ATPase class I type 8B member 4). | |||||
|
AT8B2_HUMAN
|
||||||
| NC score | 0.946859 (rank : 7) | θ value | 2.14053e-118 (rank : 6) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98198, Q96I43, Q96NQ7 | Gene names | ATP8B2, ATPID, KIAA1137 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase ID (EC 3.6.3.1) (ATPase class I type 8B member 2). | |||||
|
AT8A1_HUMAN
|
||||||
| NC score | 0.943162 (rank : 8) | θ value | 3.41741e-116 (rank : 8) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Y2Q0 | Gene names | ATP8A1, ATPIA | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IA (EC 3.6.3.1) (Chromaffin granule ATPase II) (ATPase class I type 8A member 1). | |||||
|
AT8A1_MOUSE
|
||||||
| NC score | 0.943040 (rank : 9) | θ value | 9.94316e-116 (rank : 9) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P70704 | Gene names | Atp8a1, Atpc1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IA (EC 3.6.3.1) (Chromaffin granule ATPase II) (ATPase class I type 8A member 1). | |||||
|
AT8B1_HUMAN
|
||||||
| NC score | 0.942696 (rank : 10) | θ value | 2.6227e-108 (rank : 12) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O43520, Q9BTP8 | Gene names | ATP8B1, ATPIC, FIC1, PFIC | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IC (EC 3.6.3.1) (Familial intrahepatic cholestasis type 1) (ATPase class I type 8B member 1). | |||||
|
AT8A2_HUMAN
|
||||||
| NC score | 0.942678 (rank : 11) | θ value | 3.31004e-111 (rank : 11) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9NTI2, Q9H527, Q9NPU6, Q9NTL2, Q9NYM3 | Gene names | ATP8A2, ATPIB | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IB (EC 3.6.3.1) (ATPase class I type 8A member 2) (ML-1). | |||||
|
AT8A2_MOUSE
|
||||||
| NC score | 0.942511 (rank : 12) | θ value | 1.94052e-111 (rank : 10) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98200 | Gene names | Atp8a2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IB (EC 3.6.3.1) (ATPase class I type 8A member 2). | |||||
|
AT8B3_HUMAN
|
||||||
| NC score | 0.941899 (rank : 13) | θ value | 3.21347e-98 (rank : 16) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O60423, Q8IVB8, Q8N4Y8, Q96M22 | Gene names | ATP8B3, ATP1K, FOS37502_2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IK (EC 3.6.3.1) (ATPase class I type 8B member 3). | |||||
|
AT11B_HUMAN
|
||||||
| NC score | 0.939288 (rank : 14) | θ value | 5.29683e-101 (rank : 13) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Y2G3, Q96FN1, Q9UKK7 | Gene names | ATP11B, ATPIF, ATPIR, KIAA0956 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IF (EC 3.6.3.1) (ATPase class I type 11B) (ATPase IR). | |||||
|
AT11C_HUMAN
|
||||||
| NC score | 0.936706 (rank : 15) | θ value | 1.94955e-95 (rank : 17) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8NB49, Q5JT69, Q5JT70, Q5JT71, Q5JT72, Q5JT73, Q6ZND5, Q6ZU50, Q6ZUP7, Q70IJ9, Q70IK0, Q8WX24 | Gene names | ATP11C, ATPIG, ATPIQ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IG (EC 3.6.3.1) (ATPase class I type 11C) (ATPase IG) (ATPase IQ) (ATPase class VI type 11C). | |||||
|
AT11A_HUMAN
|
||||||
| NC score | 0.934713 (rank : 16) | θ value | 6.91785e-101 (rank : 14) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98196, Q5VXT2 | Gene names | ATP11A, ATPIH, ATPIS, KIAA1021 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IH (EC 3.6.3.1) (ATPase class I type 11A) (ATPase IS). | |||||
|
AT11A_MOUSE
|
||||||
| NC score | 0.933568 (rank : 17) | θ value | 2.46047e-98 (rank : 15) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98197 | Gene names | Atp11a | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IH (EC 3.6.3.1) (ATPase class I type 11A) (ATPase IS). | |||||
|
ATP9B_MOUSE
|
||||||
| NC score | 0.905686 (rank : 18) | θ value | 2.10232e-65 (rank : 20) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98195, Q99LI3 | Gene names | Atp9b | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IIB (EC 3.6.3.1). | |||||
|
ATP9B_HUMAN
|
||||||
| NC score | 0.903601 (rank : 19) | θ value | 4.23606e-66 (rank : 18) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O43861, O60872 | Gene names | ATP9B, ATPIIB, NEO1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIB (EC 3.6.3.1). | |||||
|
ATP9A_HUMAN
|
||||||
| NC score | 0.903097 (rank : 20) | θ value | 2.10232e-65 (rank : 19) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O75110, Q5TFW5, Q5TFW6, Q5TFW9, Q6ZMF3, Q9NQK6, Q9NQK7 | Gene names | ATP9A, ATPIIA, KIAA0611 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIA (EC 3.6.3.1) (ATPase class II type 9A) (ATPase IIA). | |||||
|
ATP9A_MOUSE
|
||||||
| NC score | 0.899863 (rank : 21) | θ value | 1.50663e-63 (rank : 21) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O70228, Q8VDI5, Q922L9 | Gene names | Atp9a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIA (EC 3.6.3.1) (ATPase class II type 9A). | |||||
|
AT133_HUMAN
|
||||||
| NC score | 0.601840 (rank : 22) | θ value | 1.52774e-15 (rank : 26) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9H7F0, Q8NC11, Q96KS1 | Gene names | ATP13A3, AFURS1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A3 (EC 3.6.3.-) (ATPase family homolog up-regulated in senescence cells 1). | |||||
|
AT132_HUMAN
|
||||||
| NC score | 0.580681 (rank : 23) | θ value | 6.20254e-17 (rank : 23) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9NQ11, O75700, Q5JXY2 | Gene names | ATP13A2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A2 (EC 3.6.3.-). | |||||
|
AT132_MOUSE
|
||||||
| NC score | 0.579965 (rank : 24) | θ value | 1.058e-16 (rank : 24) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9CTG6, Q8CG98 | Gene names | Atp13a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable cation-transporting ATPase 13A2 (EC 3.6.3.-). | |||||
|
AT2B4_HUMAN
|
||||||
| NC score | 0.554825 (rank : 25) | θ value | 9.56915e-18 (rank : 22) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P23634, Q13450, Q13452, Q13455, Q16817 | Gene names | ATP2B4 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 4 (EC 3.6.3.8) (PMCA4) (Plasma membrane calcium pump isoform 4) (Plasma membrane calcium ATPase isoform 4). | |||||
|
AT2B3_HUMAN
|
||||||
| NC score | 0.534131 (rank : 26) | θ value | 7.58209e-15 (rank : 28) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q16720, Q12995, Q16858 | Gene names | ATP2B3 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 3 (EC 3.6.3.8) (PMCA3) (Plasma membrane calcium pump isoform 3) (Plasma membrane calcium ATPase isoform 3). | |||||
|
AT2B2_HUMAN
|
||||||
| NC score | 0.533021 (rank : 27) | θ value | 2.60593e-15 (rank : 27) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q01814, O00766, Q12994, Q16818 | Gene names | ATP2B2, PMCA2 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 2 (EC 3.6.3.8) (PMCA2) (Plasma membrane calcium pump isoform 2) (Plasma membrane calcium ATPase isoform 2). | |||||
|
AT2B2_MOUSE
|
||||||
| NC score | 0.529698 (rank : 28) | θ value | 5.25075e-16 (rank : 25) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9R0K7, O88863 | Gene names | Atp2b2, Pmca2 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 2 (EC 3.6.3.8) (PMCA2) (Plasma membrane calcium pump isoform 2) (Plasma membrane calcium ATPase isoform 2). | |||||
|
AT2B1_HUMAN
|
||||||
| NC score | 0.506319 (rank : 29) | θ value | 2.69671e-12 (rank : 29) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P20020, Q12992, Q12993, Q13819, Q13820, Q13821, Q16504, Q93082 | Gene names | ATP2B1, PMCA1 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 1 (EC 3.6.3.8) (PMCA1) (Plasma membrane calcium pump isoform 1) (Plasma membrane calcium ATPase isoform 1). | |||||
|
AT131_MOUSE
|
||||||
| NC score | 0.465080 (rank : 30) | θ value | 9.29e-05 (rank : 42) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9EPE9 | Gene names | Atp13a1, Atp13a | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A1 (EC 3.6.3.-) (CATP). | |||||
|
AT131_HUMAN
|
||||||
| NC score | 0.464222 (rank : 31) | θ value | 6.43352e-06 (rank : 37) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9HD20, Q9H6C6 | Gene names | ATP13A1, ATP13A, KIAA1825 | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A1 (EC 3.6.3.-). | |||||
|
AT2A2_MOUSE
|
||||||
| NC score | 0.403814 (rank : 32) | θ value | 7.59969e-07 (rank : 35) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O55143, Q9R2A9, Q9WUT5 | Gene names | Atp2a2 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 2 (EC 3.6.3.8) (Calcium pump 2) (SERCA2) (SR Ca(2+)-ATPase 2) (Calcium-transporting ATPase sarcoplasmic reticulum type, slow twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2A2_HUMAN
|
||||||
| NC score | 0.402560 (rank : 33) | θ value | 3.41135e-07 (rank : 34) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P16615, P16614 | Gene names | ATP2A2, ATP2B | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 2 (EC 3.6.3.8) (Calcium pump 2) (SERCA2) (SR Ca(2+)-ATPase 2) (Calcium-transporting ATPase sarcoplasmic reticulum type, slow twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2C1_HUMAN
|
||||||
| NC score | 0.401073 (rank : 34) | θ value | 1.53129e-07 (rank : 32) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98194, O76005, Q86V72, Q86V73, Q8N6V1, Q8NCJ7 | Gene names | ATP2C1, KIAA1347, PMR1L | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-transporting ATPase type 2C member 1 (EC 3.6.3.8) (ATPase 2C1) (ATP-dependent Ca(2+) pump PMR1). | |||||
|
AT12A_HUMAN
|
||||||
| NC score | 0.400752 (rank : 35) | θ value | 2.61198e-07 (rank : 33) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P54707, Q13816, Q13817, Q16734 | Gene names | ATP12A, ATP1AL1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 2 (EC 3.6.3.10) (Proton pump) (Non-gastric H(+)/K(+) ATPase subunit alpha). | |||||
|
AT2C1_MOUSE
|
||||||
| NC score | 0.398442 (rank : 36) | θ value | 3.08544e-08 (rank : 30) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q80XR2, Q80YZ2 | Gene names | Atp2c1, Pmr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-transporting ATPase type 2C member 1 (EC 3.6.3.8) (ATPase 2C1) (ATP-dependent Ca(2+) pump PMR1). | |||||
|
AT12A_MOUSE
|
||||||
| NC score | 0.397166 (rank : 37) | θ value | 5.26297e-08 (rank : 31) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Z1W8, Q8VHY2 | Gene names | Atp12a, Atp1al1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium-transporting ATPase alpha chain 2 (EC 3.6.3.10) (Proton pump) (Non-gastric H(+)/K(+) ATPase subunit alpha). | |||||
|
AT2A3_HUMAN
|
||||||
| NC score | 0.397138 (rank : 38) | θ value | 9.92553e-07 (rank : 36) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q93084, O60900, O60901, O75501, O75502, Q16115, Q8TEX5, Q8TEX6 | Gene names | ATP2A3 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 3 (EC 3.6.3.8) (Calcium pump 3) (SERCA3) (SR Ca(2+)-ATPase 3). | |||||
|
AT2A1_MOUSE
|
||||||
| NC score | 0.388676 (rank : 39) | θ value | 0.000786445 (rank : 49) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8R429 | Gene names | Atp2a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 1 (EC 3.6.3.8) (Calcium pump 1) (SERCA1) (SR Ca(2+)-ATPase 1) (Calcium-transporting ATPase sarcoplasmic reticulum type, fast twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2A3_MOUSE
|
||||||
| NC score | 0.386970 (rank : 40) | θ value | 1.09739e-05 (rank : 38) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q64518, O70625, Q64517 | Gene names | Atp2a3 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 3 (EC 3.6.3.8) (Calcium pump 3) (SERCA3) (SR Ca(2+)-ATPase 3). | |||||
|
AT2A1_HUMAN
|
||||||
| NC score | 0.385960 (rank : 41) | θ value | 0.000602161 (rank : 47) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O14983, O14984 | Gene names | ATP2A1 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 1 (EC 3.6.3.8) (Calcium pump 1) (SERCA1) (SR Ca(2+)-ATPase 1) (Calcium-transporting ATPase sarcoplasmic reticulum type, fast twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
ATP4A_MOUSE
|
||||||
| NC score | 0.380288 (rank : 42) | θ value | 0.00228821 (rank : 51) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q64436, Q9CV46 | Gene names | Atp4a | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 1 (EC 3.6.3.10) (Proton pump) (Gastric H+/K+ ATPase subunit alpha). | |||||
|
AT1A4_HUMAN
|
||||||
| NC score | 0.377231 (rank : 43) | θ value | 2.44474e-05 (rank : 39) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q13733, Q504T2, Q8TBN8, Q8WXA7, Q8WXH7, Q8WY13 | Gene names | ATP1A4, ATP1AL2 | |||
|
Domain Architecture |
|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-4 chain (EC 3.6.3.9) (Sodium pump 4) (Na+/K+ ATPase 4). | |||||
|
AT1A4_MOUSE
|
||||||
| NC score | 0.376717 (rank : 44) | θ value | 0.000270298 (rank : 46) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9WV27, Q9R173, Q9WV28 | Gene names | Atp1a4, Atp1al2 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-4 chain (EC 3.6.3.9) (Sodium pump 4) (Na+/K+ ATPase 4). | |||||
|
ATP4A_HUMAN
|
||||||
| NC score | 0.375686 (rank : 45) | θ value | 0.00390308 (rank : 52) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P20648, O00738 | Gene names | ATP4A | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 1 (EC 3.6.3.10) (Proton pump) (Gastric H+/K+ ATPase subunit alpha). | |||||
|
AT1A1_MOUSE
|
||||||
| NC score | 0.371973 (rank : 46) | θ value | 0.000786445 (rank : 48) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8VDN2, Q91Z09 | Gene names | Atp1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-1 chain precursor (EC 3.6.3.9) (Sodium pump 1) (Na+/K+ ATPase 1). | |||||
|
AT1A1_HUMAN
|
||||||
| NC score | 0.371354 (rank : 47) | θ value | 0.000270298 (rank : 44) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P05023, Q16689, Q6LDM4, Q9UJ20, Q9UJ21 | Gene names | ATP1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-1 chain precursor (EC 3.6.3.9) (Sodium pump 1) (Na+/K+ ATPase 1). | |||||
|
AT1A2_HUMAN
|
||||||
| NC score | 0.370141 (rank : 48) | θ value | 0.000270298 (rank : 45) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P50993, Q07059, Q86UZ5, Q9UQ25 | Gene names | ATP1A2, KIAA0778 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2). | |||||
|
AT1A2_MOUSE
|
||||||
| NC score | 0.369791 (rank : 49) | θ value | 0.000158464 (rank : 43) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q6PIE5 | Gene names | Atp1a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2) (Alpha(+)). | |||||
|
AT1A3_HUMAN
|
||||||
| NC score | 0.351827 (rank : 50) | θ value | 3.19293e-05 (rank : 40) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P13637, Q16732, Q16735, Q969K5 | Gene names | ATP1A3 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-3 chain (EC 3.6.3.9) (Sodium pump 3) (Na+/K+ ATPase 3) (Alpha(III)). | |||||
|
AT1A3_MOUSE
|
||||||
| NC score | 0.349303 (rank : 51) | θ value | 7.1131e-05 (rank : 41) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q6PIC6 | Gene names | Atp1a3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium/potassium-transporting ATPase alpha-3 chain (EC 3.6.3.9) (Sodium pump 3) (Na+/K+ ATPase 3) (Alpha(III)). | |||||
|
AT2C2_HUMAN
|
||||||
| NC score | 0.341419 (rank : 52) | θ value | 0.00102713 (rank : 50) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O75185 | Gene names | KIAA0703 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable calcium-transporting ATPase KIAA0703 (EC 3.6.3.8). | |||||
|
ATP7B_MOUSE
|
||||||
| NC score | 0.147043 (rank : 53) | θ value | θ > 10 (rank : 73) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q64446 | Gene names | Atp7b, Wnd | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 2 (EC 3.6.3.4) (Copper pump 2) (Wilson disease-associated protein homolog). | |||||
|
ATP7B_HUMAN
|
||||||
| NC score | 0.147009 (rank : 54) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P35670, Q16318, Q16319, Q4U3V3, Q59FJ9, Q5T7X7 | Gene names | ATP7B, PWD, WC1, WND | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 2 (EC 3.6.3.4) (Copper pump 2) (Wilson disease-associated protein). | |||||
|
ATP7A_HUMAN
|
||||||
| NC score | 0.143279 (rank : 55) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q04656, O00227, O00745, Q9BYY8 | Gene names | ATP7A, MC1, MNK | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 1 (EC 3.6.3.4) (Copper pump 1) (Menkes disease-associated protein). | |||||
|
ATP7A_MOUSE
|
||||||
| NC score | 0.140256 (rank : 56) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q64430, O35101, P97422, Q64431 | Gene names | Atp7a, Mnk | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 1 (EC 3.6.3.4) (Copper pump 1) (Menkes disease-associated protein homolog). | |||||
|
CYB_HUMAN
|
||||||
| NC score | 0.028766 (rank : 57) | θ value | 4.03905 (rank : 58) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P00156, Q34786, Q8HBR6, Q8HNQ0, Q8HNQ1, Q8HNQ9, Q8HNR4, Q8HNR7, Q8W7V8, Q8WCV9, Q8WCY2, Q8WCY7, Q8WCY8, Q9B1A6, Q9B1B6, Q9B1B8, Q9B1D4, Q9B1X6, Q9B2V0, Q9B2V8, Q9B2W0, Q9B2W3, Q9B2W8, Q9B2X1, Q9B2X7, Q9B2X9, Q9B2Y3, Q9B2Z0, Q9B2Z4, Q9T6H6, Q9T9Y0, Q9TEH4 | Gene names | MT-CYB, COB, CYTB, MTCYB | |||
|
Domain Architecture |

|
|||||
| Description | Cytochrome b. | |||||
|
GP128_HUMAN
|
||||||
| NC score | 0.014596 (rank : 58) | θ value | 0.279714 (rank : 53) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96K78, Q86SQ2 | Gene names | GPR128 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 128 precursor. | |||||
|
RT31_HUMAN
|
||||||
| NC score | 0.013734 (rank : 59) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92665, Q5VYC8, Q8WTV8 | Gene names | MRPS31, IMOGN38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 28S ribosomal protein S31, mitochondrial precursor (S31mt) (MRP-S31) (Imogen 38). | |||||
|
T2FA_HUMAN
|
||||||
| NC score | 0.008414 (rank : 60) | θ value | 8.99809 (rank : 68) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P35269, Q9BWN0 | Gene names | GTF2F1, RAP74 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor IIF alpha subunit (EC 2.7.11.1) (TFIIF-alpha) (Transcription initiation factor RAP74) (General transcription factor IIF polypeptide 1 74 kDa subunit protein). | |||||
|
BRD2_HUMAN
|
||||||
| NC score | 0.005630 (rank : 61) | θ value | 5.27518 (rank : 59) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P25440, O00699, O00700, Q15310, Q5STC9, Q969U4 | Gene names | BRD2, KIAA9001, RING3 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 2 (Protein RING3) (O27.1.1). | |||||
|
ABI3_MOUSE
|
||||||
| NC score | 0.004666 (rank : 62) | θ value | 8.99809 (rank : 65) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BYZ1, Q6PE63, Q9D7S4 | Gene names | Abi3, Nesh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ABI gene family member 3 (New molecule including SH3) (Nesh). | |||||
|
PDCL2_HUMAN
|
||||||
| NC score | 0.004614 (rank : 63) | θ value | 8.99809 (rank : 67) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N4E4 | Gene names | PDCL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosducin-like protein 2. | |||||
|
ZC3H6_HUMAN
|
||||||
| NC score | 0.004028 (rank : 64) | θ value | 6.88961 (rank : 64) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P61129 | Gene names | ZC3H6, KIAA2035, ZC3HDC6 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
BACH2_HUMAN
|
||||||
| NC score | 0.002320 (rank : 65) | θ value | 4.03905 (rank : 57) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BYV9, Q59H70, Q5T793, Q9NTS5 | Gene names | BACH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription regulator protein BACH2 (BTB and CNC homolog 2). | |||||
|
BRD4_HUMAN
|
||||||
| NC score | 0.002284 (rank : 66) | θ value | 8.99809 (rank : 66) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
BRD4_MOUSE
|
||||||
| NC score | 0.002218 (rank : 67) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
KCNH2_MOUSE
|
||||||
| NC score | 0.001813 (rank : 68) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35219, O35220, O35221, O35989 | Gene names | Kcnh2, Erg, Merg1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (ERG1) (MERG) (Merg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1). | |||||
|
TRI26_HUMAN
|
||||||
| NC score | -0.000690 (rank : 69) | θ value | 8.99809 (rank : 69) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q12899 | Gene names | TRIM26, RNF95, ZNF173 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 26 (Zinc finger protein 173) (Acid finger protein) (AFP) (RING finger protein 95). | |||||
|
MTR1B_MOUSE
|
||||||
| NC score | -0.000966 (rank : 70) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 831 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CIQ6, Q80T38, Q8K3K0 | Gene names | Mtnr1b | |||
|
Domain Architecture |

|
|||||
| Description | Melatonin receptor type 1B (Mel-1B-R) (Mel1b melatonin receptor). | |||||
|
CC1L1_MOUSE
|
||||||
| NC score | -0.002324 (rank : 71) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 629 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P51676 | Gene names | Ccr1l1, Cmkbr1l1 | |||
|
Domain Architecture |

|
|||||
| Description | C-C chemokine receptor 1-like protein 1 (Macrophage inflammatory protein 1 alpha receptor-like 1). | |||||
|
OR2M2_HUMAN
|
||||||
| NC score | -0.002797 (rank : 72) | θ value | 0.47712 (rank : 54) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 769 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96R28 | Gene names | OR2M2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Olfactory receptor 2M2 (OST423). | |||||
|
SP4_MOUSE
|
||||||
| NC score | -0.003448 (rank : 73) | θ value | 6.88961 (rank : 63) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 759 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q62445 | Gene names | Sp4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor Sp4. | |||||