Please be patient as the page loads
|
AT11C_HUMAN
|
||||||
| SwissProt Accessions | Q8NB49, Q5JT69, Q5JT70, Q5JT71, Q5JT72, Q5JT73, Q6ZND5, Q6ZU50, Q6ZUP7, Q70IJ9, Q70IK0, Q8WX24 | Gene names | ATP11C, ATPIG, ATPIQ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IG (EC 3.6.3.1) (ATPase class I type 11C) (ATPase IG) (ATPase IQ) (ATPase class VI type 11C). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
AT11A_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.995118 (rank : 3) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P98196, Q5VXT2 | Gene names | ATP11A, ATPIH, ATPIS, KIAA1021 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IH (EC 3.6.3.1) (ATPase class I type 11A) (ATPase IS). | |||||
|
AT11A_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.995136 (rank : 2) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98197 | Gene names | Atp11a | |||
|
Domain Architecture |
|
|||||
| Description | Probable phospholipid-transporting ATPase IH (EC 3.6.3.1) (ATPase class I type 11A) (ATPase IS). | |||||
|
AT11B_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.993845 (rank : 4) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9Y2G3, Q96FN1, Q9UKK7 | Gene names | ATP11B, ATPIF, ATPIR, KIAA0956 | |||
|
Domain Architecture |
|
|||||
| Description | Probable phospholipid-transporting ATPase IF (EC 3.6.3.1) (ATPase class I type 11B) (ATPase IR). | |||||
|
AT11C_HUMAN
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 89 | |
| SwissProt Accessions | Q8NB49, Q5JT69, Q5JT70, Q5JT71, Q5JT72, Q5JT73, Q6ZND5, Q6ZU50, Q6ZUP7, Q70IJ9, Q70IK0, Q8WX24 | Gene names | ATP11C, ATPIG, ATPIQ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IG (EC 3.6.3.1) (ATPase class I type 11C) (ATPase IG) (ATPase IQ) (ATPase class VI type 11C). | |||||
|
AT8A1_MOUSE
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.979365 (rank : 7) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P70704 | Gene names | Atp8a1, Atpc1 | |||
|
Domain Architecture |
|
|||||
| Description | Probable phospholipid-transporting ATPase IA (EC 3.6.3.1) (Chromaffin granule ATPase II) (ATPase class I type 8A member 1). | |||||
|
AT8A1_HUMAN
|
||||||
| θ value | 4.23373e-183 (rank : 6) | NC score | 0.979347 (rank : 8) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Y2Q0 | Gene names | ATP8A1, ATPIA | |||
|
Domain Architecture |
|
|||||
| Description | Probable phospholipid-transporting ATPase IA (EC 3.6.3.1) (Chromaffin granule ATPase II) (ATPase class I type 8A member 1). | |||||
|
AT8B4_HUMAN
|
||||||
| θ value | 7.47326e-180 (rank : 7) | NC score | 0.980432 (rank : 5) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8TF62, Q9H727 | Gene names | ATP8B4, KIAA1939 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IM (EC 3.6.3.1) (ATPase class I type 8B member 4). | |||||
|
AT8B2_HUMAN
|
||||||
| θ value | 6.12769e-174 (rank : 8) | NC score | 0.979418 (rank : 6) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P98198, Q96I43, Q96NQ7 | Gene names | ATP8B2, ATPID, KIAA1137 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase ID (EC 3.6.3.1) (ATPase class I type 8B member 2). | |||||
|
AT8A2_MOUSE
|
||||||
| θ value | 6.77497e-173 (rank : 9) | NC score | 0.978038 (rank : 10) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98200 | Gene names | Atp8a2 | |||
|
Domain Architecture |
|
|||||
| Description | Probable phospholipid-transporting ATPase IB (EC 3.6.3.1) (ATPase class I type 8A member 2). | |||||
|
AT8A2_HUMAN
|
||||||
| θ value | 9.47563e-167 (rank : 10) | NC score | 0.977992 (rank : 11) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9NTI2, Q9H527, Q9NPU6, Q9NTL2, Q9NYM3 | Gene names | ATP8A2, ATPIB | |||
|
Domain Architecture |
|
|||||
| Description | Probable phospholipid-transporting ATPase IB (EC 3.6.3.1) (ATPase class I type 8A member 2) (ML-1). | |||||
|
AT8B1_HUMAN
|
||||||
| θ value | 1.78703e-165 (rank : 11) | NC score | 0.978282 (rank : 9) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O43520, Q9BTP8 | Gene names | ATP8B1, ATPIC, FIC1, PFIC | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IC (EC 3.6.3.1) (Familial intrahepatic cholestasis type 1) (ATPase class I type 8B member 1). | |||||
|
AT8B3_HUMAN
|
||||||
| θ value | 1.63136e-134 (rank : 12) | NC score | 0.973355 (rank : 12) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O60423, Q8IVB8, Q8N4Y8, Q96M22 | Gene names | ATP8B3, ATP1K, FOS37502_2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IK (EC 3.6.3.1) (ATPase class I type 8B member 3). | |||||
|
ATP9B_HUMAN
|
||||||
| θ value | 1.09934e-114 (rank : 13) | NC score | 0.948163 (rank : 15) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O43861, O60872 | Gene names | ATP9B, ATPIIB, NEO1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIB (EC 3.6.3.1). | |||||
|
ATP9A_MOUSE
|
||||||
| θ value | 8.15288e-110 (rank : 14) | NC score | 0.947419 (rank : 16) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O70228, Q8VDI5, Q922L9 | Gene names | Atp9a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIA (EC 3.6.3.1) (ATPase class II type 9A). | |||||
|
ATP9B_MOUSE
|
||||||
| θ value | 3.0981e-109 (rank : 15) | NC score | 0.949453 (rank : 14) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98195, Q99LI3 | Gene names | Atp9b | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IIB (EC 3.6.3.1). | |||||
|
ATP9A_HUMAN
|
||||||
| θ value | 4.04623e-109 (rank : 16) | NC score | 0.949677 (rank : 13) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O75110, Q5TFW5, Q5TFW6, Q5TFW9, Q6ZMF3, Q9NQK6, Q9NQK7 | Gene names | ATP9A, ATPIIA, KIAA0611 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIA (EC 3.6.3.1) (ATPase class II type 9A) (ATPase IIA). | |||||
|
AT10D_MOUSE
|
||||||
| θ value | 2.01279e-100 (rank : 17) | NC score | 0.937140 (rank : 18) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8K2X1 | Gene names | Atp10d | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VD (EC 3.6.3.1) (ATPVD). | |||||
|
AT10D_HUMAN
|
||||||
| θ value | 1.94955e-95 (rank : 18) | NC score | 0.936706 (rank : 19) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9P241, Q8NC70, Q96SR3 | Gene names | ATP10D, KIAA1487 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VD (EC 3.6.3.1) (ATPVD). | |||||
|
AT10A_MOUSE
|
||||||
| θ value | 5.30912e-93 (rank : 19) | NC score | 0.935284 (rank : 20) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O54827, Q8R3B8 | Gene names | Atp10a, Atpc5, Pfatp | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (P-locus fat-associated ATPase). | |||||
|
AT10B_HUMAN
|
||||||
| θ value | 3.80478e-91 (rank : 20) | NC score | 0.940397 (rank : 17) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O94823, Q9H725 | Gene names | ATP10B, ATPVB, KIAA0715 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VB (EC 3.6.3.1). | |||||
|
AT10A_HUMAN
|
||||||
| θ value | 2.64101e-84 (rank : 21) | NC score | 0.933009 (rank : 21) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O60312, Q969I4 | Gene names | ATP10A, ATP10C, ATPVC, KIAA0566 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (ATPVA) (Aminophospholipid translocase VA). | |||||
|
AT2B2_HUMAN
|
||||||
| θ value | 2.88119e-14 (rank : 22) | NC score | 0.529492 (rank : 26) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q01814, O00766, Q12994, Q16818 | Gene names | ATP2B2, PMCA2 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 2 (EC 3.6.3.8) (PMCA2) (Plasma membrane calcium pump isoform 2) (Plasma membrane calcium ATPase isoform 2). | |||||
|
AT2B4_HUMAN
|
||||||
| θ value | 6.41864e-14 (rank : 23) | NC score | 0.551083 (rank : 23) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P23634, Q13450, Q13452, Q13455, Q16817 | Gene names | ATP2B4 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 4 (EC 3.6.3.8) (PMCA4) (Plasma membrane calcium pump isoform 4) (Plasma membrane calcium ATPase isoform 4). | |||||
|
AT2B2_MOUSE
|
||||||
| θ value | 8.38298e-14 (rank : 24) | NC score | 0.526651 (rank : 28) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9R0K7, O88863 | Gene names | Atp2b2, Pmca2 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 2 (EC 3.6.3.8) (PMCA2) (Plasma membrane calcium pump isoform 2) (Plasma membrane calcium ATPase isoform 2). | |||||
|
AT2B1_HUMAN
|
||||||
| θ value | 2.43908e-13 (rank : 25) | NC score | 0.502265 (rank : 29) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P20020, Q12992, Q12993, Q13819, Q13820, Q13821, Q16504, Q93082 | Gene names | ATP2B1, PMCA1 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 1 (EC 3.6.3.8) (PMCA1) (Plasma membrane calcium pump isoform 1) (Plasma membrane calcium ATPase isoform 1). | |||||
|
AT2B3_HUMAN
|
||||||
| θ value | 2.43908e-13 (rank : 26) | NC score | 0.531185 (rank : 25) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q16720, Q12995, Q16858 | Gene names | ATP2B3 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 3 (EC 3.6.3.8) (PMCA3) (Plasma membrane calcium pump isoform 3) (Plasma membrane calcium ATPase isoform 3). | |||||
|
AT12A_HUMAN
|
||||||
| θ value | 3.89403e-11 (rank : 27) | NC score | 0.407461 (rank : 32) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P54707, Q13816, Q13817, Q16734 | Gene names | ATP12A, ATP1AL1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 2 (EC 3.6.3.10) (Proton pump) (Non-gastric H(+)/K(+) ATPase subunit alpha). | |||||
|
AT133_HUMAN
|
||||||
| θ value | 5.62301e-10 (rank : 28) | NC score | 0.573041 (rank : 22) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9H7F0, Q8NC11, Q96KS1 | Gene names | ATP13A3, AFURS1 | |||
|
Domain Architecture |
|
|||||
| Description | Probable cation-transporting ATPase 13A3 (EC 3.6.3.-) (ATPase family homolog up-regulated in senescence cells 1). | |||||
|
AT12A_MOUSE
|
||||||
| θ value | 7.34386e-10 (rank : 29) | NC score | 0.404871 (rank : 35) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9Z1W8, Q8VHY2 | Gene names | Atp12a, Atp1al1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium-transporting ATPase alpha chain 2 (EC 3.6.3.10) (Proton pump) (Non-gastric H(+)/K(+) ATPase subunit alpha). | |||||
|
ATP4A_MOUSE
|
||||||
| θ value | 3.08544e-08 (rank : 30) | NC score | 0.395270 (rank : 38) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q64436, Q9CV46 | Gene names | Atp4a | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 1 (EC 3.6.3.10) (Proton pump) (Gastric H+/K+ ATPase subunit alpha). | |||||
|
AT2A2_MOUSE
|
||||||
| θ value | 5.26297e-08 (rank : 31) | NC score | 0.406773 (rank : 33) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O55143, Q9R2A9, Q9WUT5 | Gene names | Atp2a2 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 2 (EC 3.6.3.8) (Calcium pump 2) (SERCA2) (SR Ca(2+)-ATPase 2) (Calcium-transporting ATPase sarcoplasmic reticulum type, slow twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
ATP4A_HUMAN
|
||||||
| θ value | 6.87365e-08 (rank : 32) | NC score | 0.390700 (rank : 41) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P20648, O00738 | Gene names | ATP4A | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 1 (EC 3.6.3.10) (Proton pump) (Gastric H+/K+ ATPase subunit alpha). | |||||
|
AT2A2_HUMAN
|
||||||
| θ value | 3.41135e-07 (rank : 33) | NC score | 0.405974 (rank : 34) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P16615, P16614 | Gene names | ATP2A2, ATP2B | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 2 (EC 3.6.3.8) (Calcium pump 2) (SERCA2) (SR Ca(2+)-ATPase 2) (Calcium-transporting ATPase sarcoplasmic reticulum type, slow twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT1A1_HUMAN
|
||||||
| θ value | 7.59969e-07 (rank : 34) | NC score | 0.373679 (rank : 49) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P05023, Q16689, Q6LDM4, Q9UJ20, Q9UJ21 | Gene names | ATP1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-1 chain precursor (EC 3.6.3.9) (Sodium pump 1) (Na+/K+ ATPase 1). | |||||
|
AT132_MOUSE
|
||||||
| θ value | 2.21117e-06 (rank : 35) | NC score | 0.531702 (rank : 24) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9CTG6, Q8CG98 | Gene names | Atp13a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable cation-transporting ATPase 13A2 (EC 3.6.3.-). | |||||
|
AT1A4_HUMAN
|
||||||
| θ value | 2.88788e-06 (rank : 36) | NC score | 0.380873 (rank : 44) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q13733, Q504T2, Q8TBN8, Q8WXA7, Q8WXH7, Q8WY13 | Gene names | ATP1A4, ATP1AL2 | |||
|
Domain Architecture |
|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-4 chain (EC 3.6.3.9) (Sodium pump 4) (Na+/K+ ATPase 4). | |||||
|
AT2A3_MOUSE
|
||||||
| θ value | 8.40245e-06 (rank : 37) | NC score | 0.392488 (rank : 40) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q64518, O70625, Q64517 | Gene names | Atp2a3 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 3 (EC 3.6.3.8) (Calcium pump 3) (SERCA3) (SR Ca(2+)-ATPase 3). | |||||
|
AT2A1_HUMAN
|
||||||
| θ value | 1.09739e-05 (rank : 38) | NC score | 0.395155 (rank : 39) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O14983, O14984 | Gene names | ATP2A1 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 1 (EC 3.6.3.8) (Calcium pump 1) (SERCA1) (SR Ca(2+)-ATPase 1) (Calcium-transporting ATPase sarcoplasmic reticulum type, fast twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2A1_MOUSE
|
||||||
| θ value | 1.09739e-05 (rank : 39) | NC score | 0.397626 (rank : 37) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8R429 | Gene names | Atp2a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 1 (EC 3.6.3.8) (Calcium pump 1) (SERCA1) (SR Ca(2+)-ATPase 1) (Calcium-transporting ATPase sarcoplasmic reticulum type, fast twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT1A4_MOUSE
|
||||||
| θ value | 1.87187e-05 (rank : 40) | NC score | 0.378611 (rank : 45) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9WV27, Q9R173, Q9WV28 | Gene names | Atp1a4, Atp1al2 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-4 chain (EC 3.6.3.9) (Sodium pump 4) (Na+/K+ ATPase 4). | |||||
|
AT132_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 41) | NC score | 0.529147 (rank : 27) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9NQ11, O75700, Q5JXY2 | Gene names | ATP13A2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A2 (EC 3.6.3.-). | |||||
|
AT2A3_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 42) | NC score | 0.400342 (rank : 36) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q93084, O60900, O60901, O75501, O75502, Q16115, Q8TEX5, Q8TEX6 | Gene names | ATP2A3 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 3 (EC 3.6.3.8) (Calcium pump 3) (SERCA3) (SR Ca(2+)-ATPase 3). | |||||
|
AT131_MOUSE
|
||||||
| θ value | 9.29e-05 (rank : 43) | NC score | 0.459534 (rank : 30) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9EPE9 | Gene names | Atp13a1, Atp13a | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A1 (EC 3.6.3.-) (CATP). | |||||
|
AT131_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 44) | NC score | 0.455212 (rank : 31) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9HD20, Q9H6C6 | Gene names | ATP13A1, ATP13A, KIAA1825 | |||
|
Domain Architecture |
|
|||||
| Description | Probable cation-transporting ATPase 13A1 (EC 3.6.3.-). | |||||
|
AT1A2_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 45) | NC score | 0.375484 (rank : 47) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P50993, Q07059, Q86UZ5, Q9UQ25 | Gene names | ATP1A2, KIAA0778 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2). | |||||
|
AT1A3_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 46) | NC score | 0.355992 (rank : 50) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P13637, Q16732, Q16735, Q969K5 | Gene names | ATP1A3 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-3 chain (EC 3.6.3.9) (Sodium pump 3) (Na+/K+ ATPase 3) (Alpha(III)). | |||||
|
AT1A3_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 47) | NC score | 0.353470 (rank : 51) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q6PIC6 | Gene names | Atp1a3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium/potassium-transporting ATPase alpha-3 chain (EC 3.6.3.9) (Sodium pump 3) (Na+/K+ ATPase 3) (Alpha(III)). | |||||
|
AT1A2_MOUSE
|
||||||
| θ value | 0.000602161 (rank : 48) | NC score | 0.374852 (rank : 48) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q6PIE5 | Gene names | Atp1a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2) (Alpha(+)). | |||||
|
AT1A1_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 49) | NC score | 0.376175 (rank : 46) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8VDN2, Q91Z09 | Gene names | Atp1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-1 chain precursor (EC 3.6.3.9) (Sodium pump 1) (Na+/K+ ATPase 1). | |||||
|
AT2C1_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 50) | NC score | 0.388323 (rank : 42) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P98194, O76005, Q86V72, Q86V73, Q8N6V1, Q8NCJ7 | Gene names | ATP2C1, KIAA1347, PMR1L | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-transporting ATPase type 2C member 1 (EC 3.6.3.8) (ATPase 2C1) (ATP-dependent Ca(2+) pump PMR1). | |||||
|
AT2C1_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 51) | NC score | 0.383972 (rank : 43) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q80XR2, Q80YZ2 | Gene names | Atp2c1, Pmr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-transporting ATPase type 2C member 1 (EC 3.6.3.8) (ATPase 2C1) (ATP-dependent Ca(2+) pump PMR1). | |||||
|
AT2C2_HUMAN
|
||||||
| θ value | 0.125558 (rank : 52) | NC score | 0.331571 (rank : 52) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O75185 | Gene names | KIAA0703 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable calcium-transporting ATPase KIAA0703 (EC 3.6.3.8). | |||||
|
CENPK_HUMAN
|
||||||
| θ value | 0.125558 (rank : 53) | NC score | 0.024568 (rank : 57) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9BS16, Q9H4L0 | Gene names | CENPK, ICEN37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein K (CENP-K) (Interphase centromere complex protein 37) (Protein AF-5alpha) (p33). | |||||
|
SC6A9_HUMAN
|
||||||
| θ value | 0.125558 (rank : 54) | NC score | 0.008452 (rank : 70) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P48067 | Gene names | SLC6A9 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium- and chloride-dependent glycine transporter 1 (GlyT1) (GlyT-1) (Solute carrier family 6 member 9). | |||||
|
CASP_MOUSE
|
||||||
| θ value | 0.163984 (rank : 55) | NC score | 0.014344 (rank : 59) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1152 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P70403, Q7TQJ4, Q91WN6 | Gene names | Cutl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASP. | |||||
|
CUTL1_MOUSE
|
||||||
| θ value | 0.163984 (rank : 56) | NC score | 0.016042 (rank : 58) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
CASP_HUMAN
|
||||||
| θ value | 0.21417 (rank : 57) | NC score | 0.014197 (rank : 60) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1131 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q13948, Q53GU9, Q8TBS3 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASP. | |||||
|
CUTL1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 58) | NC score | 0.013690 (rank : 61) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
ATP7A_HUMAN
|
||||||
| θ value | 0.62314 (rank : 59) | NC score | 0.153707 (rank : 55) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q04656, O00227, O00745, Q9BYY8 | Gene names | ATP7A, MC1, MNK | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 1 (EC 3.6.3.4) (Copper pump 1) (Menkes disease-associated protein). | |||||
|
ATP7A_MOUSE
|
||||||
| θ value | 0.813845 (rank : 60) | NC score | 0.151349 (rank : 56) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q64430, O35101, P97422, Q64431 | Gene names | Atp7a, Mnk | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 1 (EC 3.6.3.4) (Copper pump 1) (Menkes disease-associated protein homolog). | |||||
|
ATP7B_HUMAN
|
||||||
| θ value | 1.06291 (rank : 61) | NC score | 0.155652 (rank : 53) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P35670, Q16318, Q16319, Q4U3V3, Q59FJ9, Q5T7X7 | Gene names | ATP7B, PWD, WC1, WND | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 2 (EC 3.6.3.4) (Copper pump 2) (Wilson disease-associated protein). | |||||
|
RPA1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 62) | NC score | 0.010956 (rank : 66) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95602, Q9UEH0, Q9UFT9 | Gene names | POLR1A | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase I largest subunit (EC 2.7.7.6) (RNA polymerase I 194 kDa subunit) (RPA194). | |||||
|
BCN1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 63) | NC score | 0.011834 (rank : 65) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 367 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O88597 | Gene names | Becn1 | |||
|
Domain Architecture |

|
|||||
| Description | Beclin-1 (Coiled-coil myosin-like BCL2-interacting protein). | |||||
|
CP135_HUMAN
|
||||||
| θ value | 1.38821 (rank : 64) | NC score | 0.004438 (rank : 77) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1290 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q66GS9, O75130, Q9H8H7 | Gene names | CEP135, CEP4, KIAA0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
MOX2R_MOUSE
|
||||||
| θ value | 1.38821 (rank : 65) | NC score | 0.013498 (rank : 62) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ES57 | Gene names | Cd200r1, Mox2r, Ox2r | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell surface glycoprotein OX2 receptor precursor (CD200 cell surface glycoprotein receptor). | |||||
|
TA2R7_HUMAN
|
||||||
| θ value | 1.38821 (rank : 66) | NC score | 0.006393 (rank : 72) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NYW3 | Gene names | TAS2R7 | |||
|
Domain Architecture |

|
|||||
| Description | Taste receptor type 2 member 7 (T2R7) (Taste receptor family B member 4) (TRB4). | |||||
|
ATP7B_MOUSE
|
||||||
| θ value | 2.36792 (rank : 67) | NC score | 0.155358 (rank : 54) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q64446 | Gene names | Atp7b, Wnd | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 2 (EC 3.6.3.4) (Copper pump 2) (Wilson disease-associated protein homolog). | |||||
|
SC6A9_MOUSE
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.006224 (rank : 73) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P28571, Q8VC47 | Gene names | Slc6a9, Glyt1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium- and chloride-dependent glycine transporter 1 (GlyT1) (GlyT-1) (Solute carrier family 6 member 9). | |||||
|
SPEG_HUMAN
|
||||||
| θ value | 2.36792 (rank : 69) | NC score | -0.003718 (rank : 88) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
T2R14_HUMAN
|
||||||
| θ value | 2.36792 (rank : 70) | NC score | 0.012994 (rank : 63) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NYV8 | Gene names | TAS2R14 | |||
|
Domain Architecture |

|
|||||
| Description | Taste receptor type 2 member 14 (T2R14) (Taste receptor family B member 1) (TRB1). | |||||
|
EEA1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 71) | NC score | 0.003908 (rank : 80) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1796 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q15075, Q14221 | Gene names | EEA1, ZFYVE2 | |||
|
Domain Architecture |

|
|||||
| Description | Early endosome antigen 1 (Endosome-associated protein p162) (Zinc finger FYVE domain-containing protein 2). | |||||
|
FLIP1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 72) | NC score | 0.004171 (rank : 78) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1176 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9CS72 | Gene names | Filip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
SPEG_MOUSE
|
||||||
| θ value | 3.0926 (rank : 73) | NC score | -0.003889 (rank : 89) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
BPAEA_HUMAN
|
||||||
| θ value | 4.03905 (rank : 74) | NC score | 0.005414 (rank : 76) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1369 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O94833, Q5TBT0, Q8N1T8, Q8N8J3, Q8WXK9, Q96AK9, Q96DQ5, Q9H555 | Gene names | DST, BPAG1, DMH, DT, KIAA0728 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 6/9/10 (Trabeculin-beta) (Bullous pemphigoid antigen) (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
BPL1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | 0.009482 (rank : 67) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P50747, Q99451 | Gene names | HLCS | |||
|
Domain Architecture |

|
|||||
| Description | Biotin--protein ligase (EC 6.3.4.-) (Biotin apo-protein ligase) [Includes: Biotin--[methylmalonyl-CoA-carboxytransferase] ligase (EC 6.3.4.9); Biotin--[propionyl-CoA-carboxylase [ATP-hydrolyzing]] ligase (EC 6.3.4.10) (Holocarboxylase synthetase) (HCS); Biotin-- [methylcrotonoyl-CoA-carboxylase] ligase (EC 6.3.4.11); Biotin-- [acetyl-CoA-carboxylase] ligase (EC 6.3.4.15)]. | |||||
|
PCNT_HUMAN
|
||||||
| θ value | 4.03905 (rank : 76) | NC score | 0.003589 (rank : 81) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1551 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O95613, O43152, Q7Z7C9 | Gene names | PCNT, KIAA0402, PCNT2 | |||
|
Domain Architecture |

|
|||||
| Description | Pericentrin (Pericentrin B) (Kendrin). | |||||
|
TA2R3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 77) | NC score | 0.007699 (rank : 71) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7TQA7, Q7M727 | Gene names | Tas2r3, T2r41, Tas2r137, Tas2r37 | |||
|
Domain Architecture |
|
|||||
| Description | Taste receptor type 2 member 3 (T2R3) (T2R137) (T2R37) (mT2r41). | |||||
|
TR140_MOUSE
|
||||||
| θ value | 4.03905 (rank : 78) | NC score | 0.008570 (rank : 69) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q7TQA4, Q9JKA0, Q9JKA2 | Gene names | Tas2r140, T2r64, Tas2r13, Tas2r137, Tas2r40, Tas2r8 | |||
|
Domain Architecture |
|
|||||
| Description | Taste receptor type 2 member 140 (T2R140) (Taste receptor type 2 member 40) (T2R40) (T2R8) (T2R13) (mT2r64) (Taste receptor family B member 3) (mTRB3) (mTRB5). | |||||
|
BPAEA_MOUSE
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.003247 (rank : 82) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1480 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q91ZU8, Q8K5D4 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoform 5 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
ITA2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | 0.001837 (rank : 85) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P17301, Q14595 | Gene names | ITGA2 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-2 precursor (Platelet membrane glycoprotein Ia) (GPIa) (Collagen receptor) (VLA-2 alpha chain) (CD49b antigen). | |||||
|
SYNE1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.004077 (rank : 79) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
COVA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.012443 (rank : 64) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q16206, Q5VTJ1, Q5VTJ2, Q8WUX0, Q9NTP6, Q9UH82 | Gene names | COVA1 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor-associated hydroquinone oxidase (tNOX) (Cytosolic ovarian carcinoma antigen 1) (APK1 antigen) [Includes: Hydroquinone [NADH] oxidase (EC 1.-.-.-); Protein disulfide-thiol oxidoreductase (EC 1.-.-.-)]. | |||||
|
COVA1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.009135 (rank : 68) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8R0Z2, Q8BY59, Q8R372 | Gene names | Cova1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor-associated hydroquinone oxidase (tNOX) (Cytosolic ovarian carcinoma antigen 1) (APK1 antigen) [Includes: Hydroquinone [NADH] oxidase (EC 1.-.-.-); Protein disulfide-thiol oxidoreductase (EC 1.-.-.-)]. | |||||
|
DYNA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.005593 (rank : 75) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1194 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q14203, O95296, Q9BRM9, Q9UIU1, Q9UIU2 | Gene names | DCTN1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynactin-1 (150 kDa dynein-associated polypeptide) (DP-150) (DAP-150) (p150-glued) (p135). | |||||
|
DYNA_MOUSE
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.006028 (rank : 74) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1084 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O08788 | Gene names | Dctn1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynactin-1 (150 kDa dynein-associated polypeptide) (DP-150) (DAP-150) (p150-glued). | |||||
|
PDE8A_MOUSE
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.002011 (rank : 84) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O88502 | Gene names | Pde8a, Pde8 | |||
|
Domain Architecture |

|
|||||
| Description | High-affinity cAMP-specific and IBMX-insensitive 3',5'-cyclic phosphodiesterase 8A (EC 3.1.4.17) (MMPDE8). | |||||
|
KTN1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.002596 (rank : 83) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1324 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61595, Q8BG49, Q8BHF4, Q8BHM8, Q8C9Y5, Q8CG51, Q8CG52, Q8CG53, Q8CG54, Q8CG55, Q8CG56, Q8CG57, Q8CG58, Q8CG59, Q8CG60, Q8CG61, Q8CG62, Q8CG63 | Gene names | Ktn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin. | |||||
|
MYH1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.000983 (rank : 87) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1478 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P12882, Q9Y622 | Gene names | MYH1 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-1 (Myosin heavy chain, skeletal muscle, adult 1) (Myosin heavy chain IIx/d) (MyHC-IIx/d). | |||||
|
MYH1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.000998 (rank : 86) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1461 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q5SX40 | Gene names | Myh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-1 (Myosin heavy chain, skeletal muscle, adult 1). | |||||
|
AT11C_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 89 | |
| SwissProt Accessions | Q8NB49, Q5JT69, Q5JT70, Q5JT71, Q5JT72, Q5JT73, Q6ZND5, Q6ZU50, Q6ZUP7, Q70IJ9, Q70IK0, Q8WX24 | Gene names | ATP11C, ATPIG, ATPIQ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IG (EC 3.6.3.1) (ATPase class I type 11C) (ATPase IG) (ATPase IQ) (ATPase class VI type 11C). | |||||
|
AT11A_MOUSE
|
||||||
| NC score | 0.995136 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98197 | Gene names | Atp11a | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IH (EC 3.6.3.1) (ATPase class I type 11A) (ATPase IS). | |||||
|
AT11A_HUMAN
|
||||||
| NC score | 0.995118 (rank : 3) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P98196, Q5VXT2 | Gene names | ATP11A, ATPIH, ATPIS, KIAA1021 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IH (EC 3.6.3.1) (ATPase class I type 11A) (ATPase IS). | |||||
|
AT11B_HUMAN
|
||||||
| NC score | 0.993845 (rank : 4) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9Y2G3, Q96FN1, Q9UKK7 | Gene names | ATP11B, ATPIF, ATPIR, KIAA0956 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IF (EC 3.6.3.1) (ATPase class I type 11B) (ATPase IR). | |||||
|
AT8B4_HUMAN
|
||||||
| NC score | 0.980432 (rank : 5) | θ value | 7.47326e-180 (rank : 7) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8TF62, Q9H727 | Gene names | ATP8B4, KIAA1939 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IM (EC 3.6.3.1) (ATPase class I type 8B member 4). | |||||
|
AT8B2_HUMAN
|
||||||
| NC score | 0.979418 (rank : 6) | θ value | 6.12769e-174 (rank : 8) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P98198, Q96I43, Q96NQ7 | Gene names | ATP8B2, ATPID, KIAA1137 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase ID (EC 3.6.3.1) (ATPase class I type 8B member 2). | |||||
|
AT8A1_MOUSE
|
||||||
| NC score | 0.979365 (rank : 7) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P70704 | Gene names | Atp8a1, Atpc1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IA (EC 3.6.3.1) (Chromaffin granule ATPase II) (ATPase class I type 8A member 1). | |||||
|
AT8A1_HUMAN
|
||||||
| NC score | 0.979347 (rank : 8) | θ value | 4.23373e-183 (rank : 6) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Y2Q0 | Gene names | ATP8A1, ATPIA | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IA (EC 3.6.3.1) (Chromaffin granule ATPase II) (ATPase class I type 8A member 1). | |||||
|
AT8B1_HUMAN
|
||||||
| NC score | 0.978282 (rank : 9) | θ value | 1.78703e-165 (rank : 11) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O43520, Q9BTP8 | Gene names | ATP8B1, ATPIC, FIC1, PFIC | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IC (EC 3.6.3.1) (Familial intrahepatic cholestasis type 1) (ATPase class I type 8B member 1). | |||||
|
AT8A2_MOUSE
|
||||||
| NC score | 0.978038 (rank : 10) | θ value | 6.77497e-173 (rank : 9) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98200 | Gene names | Atp8a2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IB (EC 3.6.3.1) (ATPase class I type 8A member 2). | |||||
|
AT8A2_HUMAN
|
||||||
| NC score | 0.977992 (rank : 11) | θ value | 9.47563e-167 (rank : 10) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9NTI2, Q9H527, Q9NPU6, Q9NTL2, Q9NYM3 | Gene names | ATP8A2, ATPIB | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IB (EC 3.6.3.1) (ATPase class I type 8A member 2) (ML-1). | |||||
|
AT8B3_HUMAN
|
||||||
| NC score | 0.973355 (rank : 12) | θ value | 1.63136e-134 (rank : 12) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O60423, Q8IVB8, Q8N4Y8, Q96M22 | Gene names | ATP8B3, ATP1K, FOS37502_2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IK (EC 3.6.3.1) (ATPase class I type 8B member 3). | |||||
|
ATP9A_HUMAN
|
||||||
| NC score | 0.949677 (rank : 13) | θ value | 4.04623e-109 (rank : 16) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O75110, Q5TFW5, Q5TFW6, Q5TFW9, Q6ZMF3, Q9NQK6, Q9NQK7 | Gene names | ATP9A, ATPIIA, KIAA0611 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIA (EC 3.6.3.1) (ATPase class II type 9A) (ATPase IIA). | |||||
|
ATP9B_MOUSE
|
||||||
| NC score | 0.949453 (rank : 14) | θ value | 3.0981e-109 (rank : 15) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P98195, Q99LI3 | Gene names | Atp9b | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase IIB (EC 3.6.3.1). | |||||
|
ATP9B_HUMAN
|
||||||
| NC score | 0.948163 (rank : 15) | θ value | 1.09934e-114 (rank : 13) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O43861, O60872 | Gene names | ATP9B, ATPIIB, NEO1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIB (EC 3.6.3.1). | |||||
|
ATP9A_MOUSE
|
||||||
| NC score | 0.947419 (rank : 16) | θ value | 8.15288e-110 (rank : 14) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O70228, Q8VDI5, Q922L9 | Gene names | Atp9a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase IIA (EC 3.6.3.1) (ATPase class II type 9A). | |||||
|
AT10B_HUMAN
|
||||||
| NC score | 0.940397 (rank : 17) | θ value | 3.80478e-91 (rank : 20) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O94823, Q9H725 | Gene names | ATP10B, ATPVB, KIAA0715 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VB (EC 3.6.3.1). | |||||
|
AT10D_MOUSE
|
||||||
| NC score | 0.937140 (rank : 18) | θ value | 2.01279e-100 (rank : 17) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8K2X1 | Gene names | Atp10d | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VD (EC 3.6.3.1) (ATPVD). | |||||
|
AT10D_HUMAN
|
||||||
| NC score | 0.936706 (rank : 19) | θ value | 1.94955e-95 (rank : 18) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9P241, Q8NC70, Q96SR3 | Gene names | ATP10D, KIAA1487 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VD (EC 3.6.3.1) (ATPVD). | |||||
|
AT10A_MOUSE
|
||||||
| NC score | 0.935284 (rank : 20) | θ value | 5.30912e-93 (rank : 19) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O54827, Q8R3B8 | Gene names | Atp10a, Atpc5, Pfatp | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (P-locus fat-associated ATPase). | |||||
|
AT10A_HUMAN
|
||||||
| NC score | 0.933009 (rank : 21) | θ value | 2.64101e-84 (rank : 21) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O60312, Q969I4 | Gene names | ATP10A, ATP10C, ATPVC, KIAA0566 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (ATPVA) (Aminophospholipid translocase VA). | |||||
|
AT133_HUMAN
|
||||||
| NC score | 0.573041 (rank : 22) | θ value | 5.62301e-10 (rank : 28) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9H7F0, Q8NC11, Q96KS1 | Gene names | ATP13A3, AFURS1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A3 (EC 3.6.3.-) (ATPase family homolog up-regulated in senescence cells 1). | |||||
|
AT2B4_HUMAN
|
||||||
| NC score | 0.551083 (rank : 23) | θ value | 6.41864e-14 (rank : 23) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P23634, Q13450, Q13452, Q13455, Q16817 | Gene names | ATP2B4 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 4 (EC 3.6.3.8) (PMCA4) (Plasma membrane calcium pump isoform 4) (Plasma membrane calcium ATPase isoform 4). | |||||
|
AT132_MOUSE
|
||||||
| NC score | 0.531702 (rank : 24) | θ value | 2.21117e-06 (rank : 35) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9CTG6, Q8CG98 | Gene names | Atp13a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable cation-transporting ATPase 13A2 (EC 3.6.3.-). | |||||
|
AT2B3_HUMAN
|
||||||
| NC score | 0.531185 (rank : 25) | θ value | 2.43908e-13 (rank : 26) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q16720, Q12995, Q16858 | Gene names | ATP2B3 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 3 (EC 3.6.3.8) (PMCA3) (Plasma membrane calcium pump isoform 3) (Plasma membrane calcium ATPase isoform 3). | |||||
|
AT2B2_HUMAN
|
||||||
| NC score | 0.529492 (rank : 26) | θ value | 2.88119e-14 (rank : 22) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q01814, O00766, Q12994, Q16818 | Gene names | ATP2B2, PMCA2 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 2 (EC 3.6.3.8) (PMCA2) (Plasma membrane calcium pump isoform 2) (Plasma membrane calcium ATPase isoform 2). | |||||
|
AT132_HUMAN
|
||||||
| NC score | 0.529147 (rank : 27) | θ value | 4.1701e-05 (rank : 41) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9NQ11, O75700, Q5JXY2 | Gene names | ATP13A2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A2 (EC 3.6.3.-). | |||||
|
AT2B2_MOUSE
|
||||||
| NC score | 0.526651 (rank : 28) | θ value | 8.38298e-14 (rank : 24) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9R0K7, O88863 | Gene names | Atp2b2, Pmca2 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 2 (EC 3.6.3.8) (PMCA2) (Plasma membrane calcium pump isoform 2) (Plasma membrane calcium ATPase isoform 2). | |||||
|
AT2B1_HUMAN
|
||||||
| NC score | 0.502265 (rank : 29) | θ value | 2.43908e-13 (rank : 25) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P20020, Q12992, Q12993, Q13819, Q13820, Q13821, Q16504, Q93082 | Gene names | ATP2B1, PMCA1 | |||
|
Domain Architecture |

|
|||||
| Description | Plasma membrane calcium-transporting ATPase 1 (EC 3.6.3.8) (PMCA1) (Plasma membrane calcium pump isoform 1) (Plasma membrane calcium ATPase isoform 1). | |||||
|
AT131_MOUSE
|
||||||
| NC score | 0.459534 (rank : 30) | θ value | 9.29e-05 (rank : 43) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9EPE9 | Gene names | Atp13a1, Atp13a | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A1 (EC 3.6.3.-) (CATP). | |||||
|
AT131_HUMAN
|
||||||
| NC score | 0.455212 (rank : 31) | θ value | 0.000158464 (rank : 44) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9HD20, Q9H6C6 | Gene names | ATP13A1, ATP13A, KIAA1825 | |||
|
Domain Architecture |

|
|||||
| Description | Probable cation-transporting ATPase 13A1 (EC 3.6.3.-). | |||||
|
AT12A_HUMAN
|
||||||
| NC score | 0.407461 (rank : 32) | θ value | 3.89403e-11 (rank : 27) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P54707, Q13816, Q13817, Q16734 | Gene names | ATP12A, ATP1AL1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 2 (EC 3.6.3.10) (Proton pump) (Non-gastric H(+)/K(+) ATPase subunit alpha). | |||||
|
AT2A2_MOUSE
|
||||||
| NC score | 0.406773 (rank : 33) | θ value | 5.26297e-08 (rank : 31) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O55143, Q9R2A9, Q9WUT5 | Gene names | Atp2a2 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 2 (EC 3.6.3.8) (Calcium pump 2) (SERCA2) (SR Ca(2+)-ATPase 2) (Calcium-transporting ATPase sarcoplasmic reticulum type, slow twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2A2_HUMAN
|
||||||
| NC score | 0.405974 (rank : 34) | θ value | 3.41135e-07 (rank : 33) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P16615, P16614 | Gene names | ATP2A2, ATP2B | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 2 (EC 3.6.3.8) (Calcium pump 2) (SERCA2) (SR Ca(2+)-ATPase 2) (Calcium-transporting ATPase sarcoplasmic reticulum type, slow twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT12A_MOUSE
|
||||||
| NC score | 0.404871 (rank : 35) | θ value | 7.34386e-10 (rank : 29) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9Z1W8, Q8VHY2 | Gene names | Atp12a, Atp1al1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium-transporting ATPase alpha chain 2 (EC 3.6.3.10) (Proton pump) (Non-gastric H(+)/K(+) ATPase subunit alpha). | |||||
|
AT2A3_HUMAN
|
||||||
| NC score | 0.400342 (rank : 36) | θ value | 7.1131e-05 (rank : 42) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q93084, O60900, O60901, O75501, O75502, Q16115, Q8TEX5, Q8TEX6 | Gene names | ATP2A3 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 3 (EC 3.6.3.8) (Calcium pump 3) (SERCA3) (SR Ca(2+)-ATPase 3). | |||||
|
AT2A1_MOUSE
|
||||||
| NC score | 0.397626 (rank : 37) | θ value | 1.09739e-05 (rank : 39) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8R429 | Gene names | Atp2a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 1 (EC 3.6.3.8) (Calcium pump 1) (SERCA1) (SR Ca(2+)-ATPase 1) (Calcium-transporting ATPase sarcoplasmic reticulum type, fast twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
ATP4A_MOUSE
|
||||||
| NC score | 0.395270 (rank : 38) | θ value | 3.08544e-08 (rank : 30) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q64436, Q9CV46 | Gene names | Atp4a | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 1 (EC 3.6.3.10) (Proton pump) (Gastric H+/K+ ATPase subunit alpha). | |||||
|
AT2A1_HUMAN
|
||||||
| NC score | 0.395155 (rank : 39) | θ value | 1.09739e-05 (rank : 38) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O14983, O14984 | Gene names | ATP2A1 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 1 (EC 3.6.3.8) (Calcium pump 1) (SERCA1) (SR Ca(2+)-ATPase 1) (Calcium-transporting ATPase sarcoplasmic reticulum type, fast twitch skeletal muscle isoform) (Endoplasmic reticulum class 1/2 Ca(2+) ATPase). | |||||
|
AT2A3_MOUSE
|
||||||
| NC score | 0.392488 (rank : 40) | θ value | 8.40245e-06 (rank : 37) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q64518, O70625, Q64517 | Gene names | Atp2a3 | |||
|
Domain Architecture |

|
|||||
| Description | Sarcoplasmic/endoplasmic reticulum calcium ATPase 3 (EC 3.6.3.8) (Calcium pump 3) (SERCA3) (SR Ca(2+)-ATPase 3). | |||||
|
ATP4A_HUMAN
|
||||||
| NC score | 0.390700 (rank : 41) | θ value | 6.87365e-08 (rank : 32) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P20648, O00738 | Gene names | ATP4A | |||
|
Domain Architecture |

|
|||||
| Description | Potassium-transporting ATPase alpha chain 1 (EC 3.6.3.10) (Proton pump) (Gastric H+/K+ ATPase subunit alpha). | |||||
|
AT2C1_HUMAN
|
||||||
| NC score | 0.388323 (rank : 42) | θ value | 0.0148317 (rank : 50) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P98194, O76005, Q86V72, Q86V73, Q8N6V1, Q8NCJ7 | Gene names | ATP2C1, KIAA1347, PMR1L | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-transporting ATPase type 2C member 1 (EC 3.6.3.8) (ATPase 2C1) (ATP-dependent Ca(2+) pump PMR1). | |||||
|
AT2C1_MOUSE
|
||||||
| NC score | 0.383972 (rank : 43) | θ value | 0.0148317 (rank : 51) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q80XR2, Q80YZ2 | Gene names | Atp2c1, Pmr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-transporting ATPase type 2C member 1 (EC 3.6.3.8) (ATPase 2C1) (ATP-dependent Ca(2+) pump PMR1). | |||||
|
AT1A4_HUMAN
|
||||||
| NC score | 0.380873 (rank : 44) | θ value | 2.88788e-06 (rank : 36) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q13733, Q504T2, Q8TBN8, Q8WXA7, Q8WXH7, Q8WY13 | Gene names | ATP1A4, ATP1AL2 | |||
|
Domain Architecture |
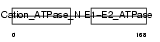
|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-4 chain (EC 3.6.3.9) (Sodium pump 4) (Na+/K+ ATPase 4). | |||||
|
AT1A4_MOUSE
|
||||||
| NC score | 0.378611 (rank : 45) | θ value | 1.87187e-05 (rank : 40) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9WV27, Q9R173, Q9WV28 | Gene names | Atp1a4, Atp1al2 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-4 chain (EC 3.6.3.9) (Sodium pump 4) (Na+/K+ ATPase 4). | |||||
|
AT1A1_MOUSE
|
||||||
| NC score | 0.376175 (rank : 46) | θ value | 0.000786445 (rank : 49) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8VDN2, Q91Z09 | Gene names | Atp1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-1 chain precursor (EC 3.6.3.9) (Sodium pump 1) (Na+/K+ ATPase 1). | |||||
|
AT1A2_HUMAN
|
||||||
| NC score | 0.375484 (rank : 47) | θ value | 0.000158464 (rank : 45) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P50993, Q07059, Q86UZ5, Q9UQ25 | Gene names | ATP1A2, KIAA0778 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2). | |||||
|
AT1A2_MOUSE
|
||||||
| NC score | 0.374852 (rank : 48) | θ value | 0.000602161 (rank : 48) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q6PIE5 | Gene names | Atp1a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium/potassium-transporting ATPase alpha-2 chain precursor (EC 3.6.3.9) (Sodium pump 2) (Na+/K+ ATPase 2) (Alpha(+)). | |||||
|
AT1A1_HUMAN
|
||||||
| NC score | 0.373679 (rank : 49) | θ value | 7.59969e-07 (rank : 34) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P05023, Q16689, Q6LDM4, Q9UJ20, Q9UJ21 | Gene names | ATP1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-1 chain precursor (EC 3.6.3.9) (Sodium pump 1) (Na+/K+ ATPase 1). | |||||
|
AT1A3_HUMAN
|
||||||
| NC score | 0.355992 (rank : 50) | θ value | 0.000270298 (rank : 46) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P13637, Q16732, Q16735, Q969K5 | Gene names | ATP1A3 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium-transporting ATPase alpha-3 chain (EC 3.6.3.9) (Sodium pump 3) (Na+/K+ ATPase 3) (Alpha(III)). | |||||
|
AT1A3_MOUSE
|
||||||
| NC score | 0.353470 (rank : 51) | θ value | 0.00035302 (rank : 47) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q6PIC6 | Gene names | Atp1a3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium/potassium-transporting ATPase alpha-3 chain (EC 3.6.3.9) (Sodium pump 3) (Na+/K+ ATPase 3) (Alpha(III)). | |||||
|
AT2C2_HUMAN
|
||||||
| NC score | 0.331571 (rank : 52) | θ value | 0.125558 (rank : 52) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O75185 | Gene names | KIAA0703 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable calcium-transporting ATPase KIAA0703 (EC 3.6.3.8). | |||||
|
ATP7B_HUMAN
|
||||||
| NC score | 0.155652 (rank : 53) | θ value | 1.06291 (rank : 61) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P35670, Q16318, Q16319, Q4U3V3, Q59FJ9, Q5T7X7 | Gene names | ATP7B, PWD, WC1, WND | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 2 (EC 3.6.3.4) (Copper pump 2) (Wilson disease-associated protein). | |||||
|
ATP7B_MOUSE
|
||||||
| NC score | 0.155358 (rank : 54) | θ value | 2.36792 (rank : 67) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q64446 | Gene names | Atp7b, Wnd | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 2 (EC 3.6.3.4) (Copper pump 2) (Wilson disease-associated protein homolog). | |||||
|
ATP7A_HUMAN
|
||||||
| NC score | 0.153707 (rank : 55) | θ value | 0.62314 (rank : 59) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q04656, O00227, O00745, Q9BYY8 | Gene names | ATP7A, MC1, MNK | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 1 (EC 3.6.3.4) (Copper pump 1) (Menkes disease-associated protein). | |||||
|
ATP7A_MOUSE
|
||||||
| NC score | 0.151349 (rank : 56) | θ value | 0.813845 (rank : 60) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q64430, O35101, P97422, Q64431 | Gene names | Atp7a, Mnk | |||
|
Domain Architecture |

|
|||||
| Description | Copper-transporting ATPase 1 (EC 3.6.3.4) (Copper pump 1) (Menkes disease-associated protein homolog). | |||||
|
CENPK_HUMAN
|
||||||
| NC score | 0.024568 (rank : 57) | θ value | 0.125558 (rank : 53) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9BS16, Q9H4L0 | Gene names | CENPK, ICEN37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein K (CENP-K) (Interphase centromere complex protein 37) (Protein AF-5alpha) (p33). | |||||
|
CUTL1_MOUSE
|
||||||
| NC score | 0.016042 (rank : 58) | θ value | 0.163984 (rank : 56) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
CASP_MOUSE
|
||||||
| NC score | 0.014344 (rank : 59) | θ value | 0.163984 (rank : 55) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1152 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P70403, Q7TQJ4, Q91WN6 | Gene names | Cutl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASP. | |||||
|
CASP_HUMAN
|
||||||
| NC score | 0.014197 (rank : 60) | θ value | 0.21417 (rank : 57) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1131 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q13948, Q53GU9, Q8TBS3 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASP. | |||||
|
CUTL1_HUMAN
|
||||||
| NC score | 0.013690 (rank : 61) | θ value | 0.21417 (rank : 58) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
MOX2R_MOUSE
|
||||||
| NC score | 0.013498 (rank : 62) | θ value | 1.38821 (rank : 65) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ES57 | Gene names | Cd200r1, Mox2r, Ox2r | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell surface glycoprotein OX2 receptor precursor (CD200 cell surface glycoprotein receptor). | |||||
|
T2R14_HUMAN
|
||||||
| NC score | 0.012994 (rank : 63) | θ value | 2.36792 (rank : 70) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NYV8 | Gene names | TAS2R14 | |||
|
Domain Architecture |

|
|||||
| Description | Taste receptor type 2 member 14 (T2R14) (Taste receptor family B member 1) (TRB1). | |||||
|
COVA1_HUMAN
|
||||||
| NC score | 0.012443 (rank : 64) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q16206, Q5VTJ1, Q5VTJ2, Q8WUX0, Q9NTP6, Q9UH82 | Gene names | COVA1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor-associated hydroquinone oxidase (tNOX) (Cytosolic ovarian carcinoma antigen 1) (APK1 antigen) [Includes: Hydroquinone [NADH] oxidase (EC 1.-.-.-); Protein disulfide-thiol oxidoreductase (EC 1.-.-.-)]. | |||||
|
BCN1_MOUSE
|
||||||
| NC score | 0.011834 (rank : 65) | θ value | 1.38821 (rank : 63) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 367 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O88597 | Gene names | Becn1 | |||
|
Domain Architecture |

|
|||||
| Description | Beclin-1 (Coiled-coil myosin-like BCL2-interacting protein). | |||||
|
RPA1_HUMAN
|
||||||
| NC score | 0.010956 (rank : 66) | θ value | 1.06291 (rank : 62) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95602, Q9UEH0, Q9UFT9 | Gene names | POLR1A | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase I largest subunit (EC 2.7.7.6) (RNA polymerase I 194 kDa subunit) (RPA194). | |||||
|
BPL1_HUMAN
|
||||||
| NC score | 0.009482 (rank : 67) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P50747, Q99451 | Gene names | HLCS | |||
|
Domain Architecture |

|
|||||
| Description | Biotin--protein ligase (EC 6.3.4.-) (Biotin apo-protein ligase) [Includes: Biotin--[methylmalonyl-CoA-carboxytransferase] ligase (EC 6.3.4.9); Biotin--[propionyl-CoA-carboxylase [ATP-hydrolyzing]] ligase (EC 6.3.4.10) (Holocarboxylase synthetase) (HCS); Biotin-- [methylcrotonoyl-CoA-carboxylase] ligase (EC 6.3.4.11); Biotin-- [acetyl-CoA-carboxylase] ligase (EC 6.3.4.15)]. | |||||
|
COVA1_MOUSE
|
||||||
| NC score | 0.009135 (rank : 68) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8R0Z2, Q8BY59, Q8R372 | Gene names | Cova1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor-associated hydroquinone oxidase (tNOX) (Cytosolic ovarian carcinoma antigen 1) (APK1 antigen) [Includes: Hydroquinone [NADH] oxidase (EC 1.-.-.-); Protein disulfide-thiol oxidoreductase (EC 1.-.-.-)]. | |||||
|
TR140_MOUSE
|
||||||
| NC score | 0.008570 (rank : 69) | θ value | 4.03905 (rank : 78) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q7TQA4, Q9JKA0, Q9JKA2 | Gene names | Tas2r140, T2r64, Tas2r13, Tas2r137, Tas2r40, Tas2r8 | |||
|
Domain Architecture |

|
|||||
| Description | Taste receptor type 2 member 140 (T2R140) (Taste receptor type 2 member 40) (T2R40) (T2R8) (T2R13) (mT2r64) (Taste receptor family B member 3) (mTRB3) (mTRB5). | |||||
|
SC6A9_HUMAN
|
||||||
| NC score | 0.008452 (rank : 70) | θ value | 0.125558 (rank : 54) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P48067 | Gene names | SLC6A9 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium- and chloride-dependent glycine transporter 1 (GlyT1) (GlyT-1) (Solute carrier family 6 member 9). | |||||
|
TA2R3_MOUSE
|
||||||
| NC score | 0.007699 (rank : 71) | θ value | 4.03905 (rank : 77) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7TQA7, Q7M727 | Gene names | Tas2r3, T2r41, Tas2r137, Tas2r37 | |||
|
Domain Architecture |

|
|||||
| Description | Taste receptor type 2 member 3 (T2R3) (T2R137) (T2R37) (mT2r41). | |||||
|
TA2R7_HUMAN
|
||||||
| NC score | 0.006393 (rank : 72) | θ value | 1.38821 (rank : 66) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NYW3 | Gene names | TAS2R7 | |||
|
Domain Architecture |

|
|||||
| Description | Taste receptor type 2 member 7 (T2R7) (Taste receptor family B member 4) (TRB4). | |||||
|
SC6A9_MOUSE
|
||||||
| NC score | 0.006224 (rank : 73) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P28571, Q8VC47 | Gene names | Slc6a9, Glyt1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium- and chloride-dependent glycine transporter 1 (GlyT1) (GlyT-1) (Solute carrier family 6 member 9). | |||||
|
DYNA_MOUSE
|
||||||
| NC score | 0.006028 (rank : 74) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1084 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O08788 | Gene names | Dctn1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynactin-1 (150 kDa dynein-associated polypeptide) (DP-150) (DAP-150) (p150-glued). | |||||
|
DYNA_HUMAN
|
||||||
| NC score | 0.005593 (rank : 75) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1194 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q14203, O95296, Q9BRM9, Q9UIU1, Q9UIU2 | Gene names | DCTN1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynactin-1 (150 kDa dynein-associated polypeptide) (DP-150) (DAP-150) (p150-glued) (p135). | |||||
|
BPAEA_HUMAN
|
||||||
| NC score | 0.005414 (rank : 76) | θ value | 4.03905 (rank : 74) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1369 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O94833, Q5TBT0, Q8N1T8, Q8N8J3, Q8WXK9, Q96AK9, Q96DQ5, Q9H555 | Gene names | DST, BPAG1, DMH, DT, KIAA0728 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 6/9/10 (Trabeculin-beta) (Bullous pemphigoid antigen) (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
CP135_HUMAN
|
||||||
| NC score | 0.004438 (rank : 77) | θ value | 1.38821 (rank : 64) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1290 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q66GS9, O75130, Q9H8H7 | Gene names | CEP135, CEP4, KIAA0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
FLIP1_MOUSE
|
||||||
| NC score | 0.004171 (rank : 78) | θ value | 3.0926 (rank : 72) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1176 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9CS72 | Gene names | Filip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
SYNE1_HUMAN
|
||||||
| NC score | 0.004077 (rank : 79) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
EEA1_HUMAN
|
||||||
| NC score | 0.003908 (rank : 80) | θ value | 3.0926 (rank : 71) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1796 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q15075, Q14221 | Gene names | EEA1, ZFYVE2 | |||
|
Domain Architecture |

|
|||||
| Description | Early endosome antigen 1 (Endosome-associated protein p162) (Zinc finger FYVE domain-containing protein 2). | |||||
|
PCNT_HUMAN
|
||||||
| NC score | 0.003589 (rank : 81) | θ value | 4.03905 (rank : 76) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1551 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O95613, O43152, Q7Z7C9 | Gene names | PCNT, KIAA0402, PCNT2 | |||
|
Domain Architecture |

|
|||||
| Description | Pericentrin (Pericentrin B) (Kendrin). | |||||
|
BPAEA_MOUSE
|
||||||
| NC score | 0.003247 (rank : 82) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1480 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q91ZU8, Q8K5D4 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoform 5 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
KTN1_MOUSE
|
||||||
| NC score | 0.002596 (rank : 83) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1324 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61595, Q8BG49, Q8BHF4, Q8BHM8, Q8C9Y5, Q8CG51, Q8CG52, Q8CG53, Q8CG54, Q8CG55, Q8CG56, Q8CG57, Q8CG58, Q8CG59, Q8CG60, Q8CG61, Q8CG62, Q8CG63 | Gene names | Ktn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin. | |||||
|
PDE8A_MOUSE
|
||||||
| NC score | 0.002011 (rank : 84) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O88502 | Gene names | Pde8a, Pde8 | |||
|
Domain Architecture |

|
|||||
| Description | High-affinity cAMP-specific and IBMX-insensitive 3',5'-cyclic phosphodiesterase 8A (EC 3.1.4.17) (MMPDE8). | |||||
|
ITA2_HUMAN
|
||||||
| NC score | 0.001837 (rank : 85) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P17301, Q14595 | Gene names | ITGA2 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-2 precursor (Platelet membrane glycoprotein Ia) (GPIa) (Collagen receptor) (VLA-2 alpha chain) (CD49b antigen). | |||||
|
MYH1_MOUSE
|
||||||
| NC score | 0.000998 (rank : 86) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1461 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q5SX40 | Gene names | Myh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-1 (Myosin heavy chain, skeletal muscle, adult 1). | |||||
|
MYH1_HUMAN
|
||||||
| NC score | 0.000983 (rank : 87) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1478 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P12882, Q9Y622 | Gene names | MYH1 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-1 (Myosin heavy chain, skeletal muscle, adult 1) (Myosin heavy chain IIx/d) (MyHC-IIx/d). | |||||
|
SPEG_HUMAN
|
||||||
| NC score | -0.003718 (rank : 88) | θ value | 2.36792 (rank : 69) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
SPEG_MOUSE
|
||||||
| NC score | -0.003889 (rank : 89) | θ value | 3.0926 (rank : 73) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||