Please be patient as the page loads
|
UB7I1_HUMAN
|
||||||
| SwissProt Accessions | Q9NWF9, Q6Y691, Q75ML7, Q7Z2H7, Q7Z7C1, Q8NHW7, Q9NYT1 | Gene names | UBCE7IP1, ZIN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin ligase TRIAD3 (EC 6.3.2.-) (Ubiquitin-conjugating enzyme 7-interacting protein 1) (Zinc finger protein inhibiting NF-kappa-B) (Triad domain-containing protein 3). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
UB7I1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 177 | |
| SwissProt Accessions | Q9NWF9, Q6Y691, Q75ML7, Q7Z2H7, Q7Z7C1, Q8NHW7, Q9NYT1 | Gene names | UBCE7IP1, ZIN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin ligase TRIAD3 (EC 6.3.2.-) (Ubiquitin-conjugating enzyme 7-interacting protein 1) (Zinc finger protein inhibiting NF-kappa-B) (Triad domain-containing protein 3). | |||||
|
UB7I1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.965303 (rank : 2) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 87 | |
| SwissProt Accessions | P58283, Q3U493, Q3UE56, Q3UGM3, Q68FN0, Q6P1H8, Q6PWY5, Q8BN27, Q8C1U3 | Gene names | Ubce7ip1, Triad3, Uip83, Zin | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin ligase TRIAD3 (EC 6.3.2.-) (Ubiquitin-conjugating enzyme 7-interacting protein 1) (UbcM4-interacting protein 83) (Triad domain- containing protein 3). | |||||
|
UB7I4_HUMAN
|
||||||
| θ value | 6.85773e-16 (rank : 3) | NC score | 0.616461 (rank : 3) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P50876 | Gene names | RNF144, KIAA0161, UBCE7IP4 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-conjugating enzyme 7-interacting protein 4 (UbcM4- interacting protein 4) (RING finger protein 144). | |||||
|
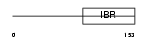
UB7I4_MOUSE
|
||||||
| θ value | 8.38298e-14 (rank : 4) | NC score | 0.603191 (rank : 4) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q925F3, Q6A0B4 | Gene names | Rnf144, Kiaa0161, Ubce7ip4, Uip4 | |||
|
Domain Architecture |
|
|||||
| Description | Ubiquitin-conjugating enzyme 7-interacting protein 4 (UbcM4- interacting protein 4) (RING finger protein 144). | |||||
|
RNF14_HUMAN
|
||||||
| θ value | 4.30538e-10 (rank : 5) | NC score | 0.545250 (rank : 6) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UBS8, O94793, Q6IBV0 | Gene names | RNF14, ARA54 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 14 (Androgen receptor-associated protein 54) (Triad2 protein) (HFB30). | |||||
|
UB7I3_MOUSE
|
||||||
| θ value | 4.30538e-10 (rank : 6) | NC score | 0.516157 (rank : 16) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9WUB0 | Gene names | Rbck1, Rbck, Ubce7ip3, Uip28 | |||
|
Domain Architecture |
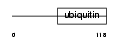
|
|||||
| Description | RanBP-type and C3HC4-type zinc finger-containing protein 1 (Ubiquitin- conjugating enzyme 7-interacting protein 3) (UbcM4-interacting protein 28). | |||||
|
UB7I3_HUMAN
|
||||||
| θ value | 5.62301e-10 (rank : 7) | NC score | 0.517941 (rank : 15) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9BYM8, O95623, Q96BS3, Q9BYM9 | Gene names | RBCK1, C20orf18, RNF54, UBCE7IP3, XAP4 | |||
|
Domain Architecture |
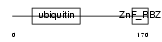
|
|||||
| Description | RanBP-type and C3HC4-type zinc finger-containing protein 1 (Ubiquitin- conjugating enzyme 7-interacting protein 3) (Hepatitis B virus X- associated protein 4) (HBV-associated factor 4) (RING finger protein 54). | |||||
|
RNF19_HUMAN
|
||||||
| θ value | 1.63604e-09 (rank : 8) | NC score | 0.506987 (rank : 18) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9NV58, Q9H5H9, Q9H8M8, Q9UFG0, Q9UFX6, Q9Y4Y1 | Gene names | RNF19 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 19 (Dorfin) (Double ring-finger protein) (p38 protein). | |||||
|
RNF19_MOUSE
|
||||||
| θ value | 1.63604e-09 (rank : 9) | NC score | 0.508547 (rank : 17) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P50636, Q9QUJ5 | Gene names | Rnf19, Geg-154, Xybp | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 19 (XY body protein) (XYbp) (Gametogenesis expressed protein GEG-154) (UBCM4-interacting protein 117) (UIP117). | |||||
|
ARI2_HUMAN
|
||||||
| θ value | 2.13673e-09 (rank : 10) | NC score | 0.523999 (rank : 12) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O95376, Q9HBZ6, Q9UEM9 | Gene names | ARIH2, ARI2, TRIAD1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ariadne-2 homolog (ARI-2) (Triad1 protein). | |||||
|
ARI2_MOUSE
|
||||||
| θ value | 3.64472e-09 (rank : 11) | NC score | 0.521875 (rank : 14) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Z1K6 | Gene names | Arih2, Ari2, Triad1, Uip48 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ariadne-2 homolog (ARI-2) (Triad1 protein) (UbcM4-interacting protein 48). | |||||
|
ARI1_HUMAN
|
||||||
| θ value | 1.06045e-08 (rank : 12) | NC score | 0.525677 (rank : 10) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Y4X5, O76026, Q9H3T6, Q9UEN0, Q9UP39 | Gene names | ARIH1, ARI, UBCH7BP | |||
|
Domain Architecture |

|
|||||
| Description | Protein ariadne-1 homolog (ARI-1) (Ubiquitin-conjugating enzyme E2- binding protein 1) (UbcH7-binding protein) (UbcM4-interacting protein) (HHARI) (H7-AP2) (MOP-6). | |||||
|
ARI1_MOUSE
|
||||||
| θ value | 1.06045e-08 (rank : 13) | NC score | 0.525653 (rank : 11) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Z1K5, Q6PF92 | Gene names | Arih1, Ari, Ubch7bp, Uip77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ariadne-1 homolog (ARI-1) (Ubiquitin-conjugating enzyme E2- binding protein 1) (UbcH7-binding protein) (UbcM4-interacting protein 77). | |||||
|
IBRD1_HUMAN
|
||||||
| θ value | 2.36244e-08 (rank : 14) | NC score | 0.572457 (rank : 5) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8TC41, Q9BX48 | Gene names | IBRDC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IBR domain-containing protein 1. | |||||
|
IBRD3_HUMAN
|
||||||
| θ value | 3.08544e-08 (rank : 15) | NC score | 0.523135 (rank : 13) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6ZMZ0, Q6P6A4 | Gene names | IBRDC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IBR domain-containing protein 3. | |||||
|
RNF14_MOUSE
|
||||||
| θ value | 8.97725e-08 (rank : 16) | NC score | 0.526700 (rank : 9) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JI90, Q9D0L2, Q9D6N2, Q9D6Z8, Q9JI89 | Gene names | Rnf14, Ara54, Triad2 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 14 (Androgen receptor-associated protein 54) (Triad2 protein). | |||||
|
IBRD2_HUMAN
|
||||||
| θ value | 2.61198e-07 (rank : 17) | NC score | 0.545000 (rank : 7) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q7Z419, Q6P4Q0, Q8N3R7, Q9BX38 | Gene names | IBRDC2, P53RFP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin ligase IBRDC2 (EC 6.3.2.-) (IBR domain-containing protein 2) (p53-inducible RING finger protein). | |||||
|
IBRD2_MOUSE
|
||||||
| θ value | 1.43324e-05 (rank : 18) | NC score | 0.527247 (rank : 8) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BKD6, Q8BG97 | Gene names | Ibrdc2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin ligase IBRDC2 (EC 6.3.2.-) (IBR domain-containing protein 2). | |||||
|
PRKN2_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 19) | NC score | 0.380158 (rank : 19) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9WVS6, Q9ES22, Q9ES23 | Gene names | Park2, Prkn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parkin (EC 6.3.2.-) (Ubiquitin E3 ligase PRKN). | |||||
|
PRKN2_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 20) | NC score | 0.378546 (rank : 20) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O60260, Q5TFV8, Q6Q2I6, Q8NI41, Q8NI43, Q8NI44, Q8WW07 | Gene names | PARK2, PRKN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parkin (EC 6.3.2.-) (Ubiquitin E3 ligase PRKN) (Parkinson juvenile disease protein 2) (Parkinson disease protein 2). | |||||
|
RNF31_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 21) | NC score | 0.350995 (rank : 21) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q924T7 | Gene names | Rnf31 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 31. | |||||
|
RNF31_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 22) | NC score | 0.350410 (rank : 22) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q96EP0, Q86VI2, Q8TEI0, Q96GB4, Q96NF1, Q9H5F1, Q9NWD2 | Gene names | RNF31, ZIBRA | |||
|
Domain Architecture |
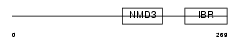
|
|||||
| Description | RING finger protein 31 (Zinc in-between-RING-finger ubiquitin- associated domain protein). | |||||
|
ATBF1_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 23) | NC score | 0.044309 (rank : 43) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1841 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q61329 | Gene names | Atbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
PARC_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 24) | NC score | 0.309899 (rank : 23) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8IWT3, O75188, Q68CP2, Q68D92, Q8N3W9, Q9BU56 | Gene names | PARC, H7AP1, KIAA0708 | |||
|
Domain Architecture |

|
|||||
| Description | p53-associated parkin-like cytoplasmic protein (UbcH7-associated protein 1). | |||||
|
CLMN_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 25) | NC score | 0.030759 (rank : 67) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96JQ2, Q9H713, Q9HA23, Q9HA57, Q9UFP4, Q9ULN2 | Gene names | CLMN, KIAA1188 | |||
|
Domain Architecture |
|
|||||
| Description | Calmin. | |||||
|
SN1L2_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 26) | NC score | 0.011852 (rank : 140) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8CFH6 | Gene names | Snf1lk2, Sik2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SNF1-like kinase 2 (EC 2.7.11.1) (Salt-inducible kinase 2). | |||||
|
MYST3_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 27) | NC score | 0.045215 (rank : 42) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
BRD4_HUMAN
|
||||||
| θ value | 0.125558 (rank : 28) | NC score | 0.049168 (rank : 36) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
BRD4_MOUSE
|
||||||
| θ value | 0.125558 (rank : 29) | NC score | 0.047050 (rank : 41) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
RP1L1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 30) | NC score | 0.049018 (rank : 38) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
MTG8R_MOUSE
|
||||||
| θ value | 0.21417 (rank : 31) | NC score | 0.067444 (rank : 25) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O70374 | Gene names | Cbfa2t2, Cbfa2t2h, Mtgr1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein CBFA2T2 (MTG8-like protein) (MTG8-related protein 1). | |||||
|
RPGR_MOUSE
|
||||||
| θ value | 0.21417 (rank : 32) | NC score | 0.032841 (rank : 63) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R0X5, O88408, Q9CU92 | Gene names | Rpgr | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator (mRpgr). | |||||
|
ZNF43_HUMAN
|
||||||
| θ value | 0.21417 (rank : 33) | NC score | 0.007819 (rank : 159) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P17038, P28160, Q96DG1 | Gene names | ZNF43, KOX27, ZNF39 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 43 (Zinc protein HTF6) (Zinc finger protein KOX27). | |||||
|
CONA1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 34) | NC score | 0.030147 (rank : 69) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8K4G2, Q5SUQ0 | Gene names | Col23a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXIII) chain. | |||||
|
NRK_MOUSE
|
||||||
| θ value | 0.365318 (rank : 35) | NC score | 0.011750 (rank : 144) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 899 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9R0G8, Q6NV55, Q8C9S9, Q9R0S4 | Gene names | Nrk, Nesk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nik-related protein kinase (EC 2.7.11.1) (Nck-interacting kinase-like embryo specific kinase) (NIK-like embryo-specific kinase) (NESK). | |||||
|
RAD52_HUMAN
|
||||||
| θ value | 0.365318 (rank : 36) | NC score | 0.058137 (rank : 30) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P43351, Q13205 | Gene names | RAD52 | |||
|
Domain Architecture |
|
|||||
| Description | DNA repair protein RAD52 homolog. | |||||
|
RIPK3_MOUSE
|
||||||
| θ value | 0.365318 (rank : 37) | NC score | 0.009868 (rank : 153) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 867 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9QZL0 | Gene names | Ripk3, Rip3 | |||
|
Domain Architecture |
|
|||||
| Description | Receptor-interacting serine/threonine-protein kinase 3 (EC 2.7.11.1) (RIP-like protein kinase 3) (Receptor-interacting protein 3) (RIP-3) (mRIP3). | |||||
|
INVO_HUMAN
|
||||||
| θ value | 0.47712 (rank : 38) | NC score | 0.029398 (rank : 71) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P07476 | Gene names | IVL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Involucrin. | |||||
|
LAMA2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 39) | NC score | 0.021765 (rank : 101) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1092 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P24043, Q14736, Q93022 | Gene names | LAMA2, LAMM | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
CCD96_HUMAN
|
||||||
| θ value | 0.62314 (rank : 40) | NC score | 0.030127 (rank : 70) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 595 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q2M329, Q8N2I7 | Gene names | CCDC96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
RBM12_MOUSE
|
||||||
| θ value | 0.62314 (rank : 41) | NC score | 0.057679 (rank : 31) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8R4X3, Q8K373, Q8R302, Q9CS80 | Gene names | Rbm12 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 12 (RNA-binding motif protein 12) (SH3/WW domain anchor protein in the nucleus) (SWAN). | |||||
|
TFG_HUMAN
|
||||||
| θ value | 0.62314 (rank : 42) | NC score | 0.061534 (rank : 28) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92734, Q15656 | Gene names | TFG | |||
|
Domain Architecture |

|
|||||
| Description | Protein TFG (TRK-fused gene protein). | |||||
|
TRHY_HUMAN
|
||||||
| θ value | 0.62314 (rank : 43) | NC score | 0.022215 (rank : 97) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
ZN198_HUMAN
|
||||||
| θ value | 0.62314 (rank : 44) | NC score | 0.044115 (rank : 44) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UBW7, O43212, O43434, O60898, Q5W0Q4, Q63HP0, Q8NE39, Q9H538, Q9UEU2 | Gene names | ZNF198, FIM, RAMP, ZMYM2 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 198 (Zinc finger MYM-type protein 2) (Fused in myeloproliferative disorders protein) (Rearranged in atypical myeloproliferative disorder protein). | |||||
|
UFO_HUMAN
|
||||||
| θ value | 0.813845 (rank : 45) | NC score | 0.006061 (rank : 165) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 981 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P30530 | Gene names | AXL, UFO | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase receptor UFO precursor (EC 2.7.10.1) (AXL oncogene). | |||||
|
ATX7_HUMAN
|
||||||
| θ value | 1.06291 (rank : 46) | NC score | 0.056585 (rank : 33) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O15265, O75328, O75329, Q9Y6P8 | Gene names | ATXN7, SCA7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-7 (Spinocerebellar ataxia type 7 protein). | |||||
|
PARC_MOUSE
|
||||||
| θ value | 1.06291 (rank : 47) | NC score | 0.292388 (rank : 24) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q80TT8, Q8BKL9, Q8CGC0, Q8R363 | Gene names | Parc, Kiaa0708 | |||
|
Domain Architecture |

|
|||||
| Description | p53-associated parkin-like cytoplasmic protein. | |||||
|
PRG4_MOUSE
|
||||||
| θ value | 1.06291 (rank : 48) | NC score | 0.037872 (rank : 54) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
DIDO1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 49) | NC score | 0.036254 (rank : 58) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
INVO_MOUSE
|
||||||
| θ value | 1.38821 (rank : 50) | NC score | 0.060517 (rank : 29) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P48997 | Gene names | Ivl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Involucrin. | |||||
|
MYST3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 51) | NC score | 0.036896 (rank : 57) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
MYT1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 52) | NC score | 0.019962 (rank : 110) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CFC2, O08995, Q8CFH1 | Gene names | Myt1, Kiaa0835, Nzf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1 (MyT1) (Neural zinc finger factor 2) (NZF-2). | |||||
|
RP3A_HUMAN
|
||||||
| θ value | 1.38821 (rank : 53) | NC score | 0.016418 (rank : 122) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Y2J0, Q96AE0 | Gene names | RPH3A, KIAA0985 | |||
|
Domain Architecture |

|
|||||
| Description | Rabphilin-3A (Exophilin-1). | |||||
|
TAOK2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 54) | NC score | 0.008688 (rank : 157) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1172 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UL54, O94957, Q6UW73, Q7LC09, Q9NSW2 | Gene names | TAOK2, KIAA0881, MAP3K17, PSK, PSK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO2 (EC 2.7.11.1) (Thousand and one amino acid protein 2) (Prostate-derived STE20-like kinase 1) (PSK-1) (Kinase from chicken homolog C) (hKFC-C). | |||||
|
TAOK2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 55) | NC score | 0.009574 (rank : 154) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1373 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q6ZQ29, Q7TSS8 | Gene names | Taok2, Kiaa0881 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO2 (EC 2.7.11.1) (Thousand and one amino acid protein 2). | |||||
|
ZN206_HUMAN
|
||||||
| θ value | 1.38821 (rank : 56) | NC score | 0.005119 (rank : 171) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 853 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96SZ4 | Gene names | ZNF206, ZSCAN10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 206 (Zinc finger and SCAN domain-containing protein 10). | |||||
|
ATF4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 57) | NC score | 0.031486 (rank : 66) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P18848, Q9UH31 | Gene names | ATF4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-4 (Activating transcription factor 4) (DNA-binding protein TAXREB67) (Cyclic AMP response element-binding protein 2) (CREB2). | |||||
|
DLEC1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 58) | NC score | 0.030301 (rank : 68) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y238, Q9NSW0, Q9NTG5 | Gene names | DLEC1, DLC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Deleted in lung and esophageal cancer protein 1 (Deleted in lung cancer protein 1) (DLC-1). | |||||
|
SFRIP_HUMAN
|
||||||
| θ value | 1.81305 (rank : 59) | NC score | 0.032788 (rank : 64) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q99590, Q8IW59 | Gene names | SFRS2IP, CASP11, SIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SFRS2-interacting protein (Splicing factor, arginine/serine-rich 2- interacting protein) (SC35-interacting protein 1) (CTD-associated SR protein 11) (Splicing regulatory protein 129) (SRrp129) (NY-REN-40 antigen). | |||||
|
SIA1A_MOUSE
|
||||||
| θ value | 1.81305 (rank : 60) | NC score | 0.056929 (rank : 32) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P61092, Q06984 | Gene names | Siah1a | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase SIAH1A (EC 6.3.2.-) (Seven in absentia homolog 1a) (Siah1a) (Siah-1a) (mSiah-1a). | |||||
|
SIAH2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 61) | NC score | 0.061990 (rank : 26) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O43255, O43270 | Gene names | SIAH2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase SIAH2 (EC 6.3.2.-) (Seven in absentia homolog 2) (Siah-2) (hSiah2). | |||||
|
SIAH2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 62) | NC score | 0.061844 (rank : 27) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q06986 | Gene names | Siah2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase SIAH2 (EC 6.3.2.-) (Seven in absentia homolog 2) (Siah-2) (mSiah2). | |||||
|
TNR6A_HUMAN
|
||||||
| θ value | 1.81305 (rank : 63) | NC score | 0.028679 (rank : 73) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8NDV7, O15408, Q658L5, Q6NVB5, Q8NEZ0, Q8TBT8, Q8TCR0, Q9NV59, Q9P268 | Gene names | TNRC6A, CAGH26, KIAA1460, TNRC6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trinucleotide repeat-containing gene 6A protein (CAG repeat protein 26) (Glycine-tryptophan protein of 182 kDa) (GW182 autoantigen) (Protein GW1) (EMSY interactor protein). | |||||
|
AP3B2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 64) | NC score | 0.019452 (rank : 113) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q13367, O14808 | Gene names | AP3B2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (AP-3 complex beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain) (Neuron-specific vesicle coat protein beta-NAP). | |||||
|
BL1S3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 65) | NC score | 0.038242 (rank : 52) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6QNY0 | Gene names | BLOC1S3, BLOS3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Biogenesis of lysosome-related organelles complex-1 subunit 3 (BLOC subunit 3). | |||||
|
COIA1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 66) | NC score | 0.023294 (rank : 91) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 481 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P39061, Q60672, Q61437, Q62001, Q62002, Q9JK63 | Gene names | Col18a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVIII) chain precursor [Contains: Endostatin]. | |||||
|
CONA1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 67) | NC score | 0.025166 (rank : 84) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q86Y22, Q8IVR4, Q9NT93 | Gene names | COL23A1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXIII) chain. | |||||
|
ETV5_MOUSE
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.015759 (rank : 125) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9CXC9, Q3TG49, Q8C0F3, Q9JHB1 | Gene names | Etv5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 5. | |||||
|
HCLS1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 69) | NC score | 0.017926 (rank : 115) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P49710 | Gene names | Hcls1, Hs1 | |||
|
Domain Architecture |

|
|||||
| Description | Hematopoietic lineage cell-specific protein (Hematopoietic cell- specific LYN substrate 1) (LckBP1). | |||||
|
IF4G1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 70) | NC score | 0.023787 (rank : 88) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q6NZJ6, Q6NZN8, Q8BW99 | Gene names | Eif4g1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1). | |||||
|
ITB4_HUMAN
|
||||||
| θ value | 2.36792 (rank : 71) | NC score | 0.016401 (rank : 123) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P16144, O14690, O14691, O15339, O15340, O15341, Q9UIQ4 | Gene names | ITGB4 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-4 precursor (GP150) (CD104 antigen). | |||||
|
KLF13_HUMAN
|
||||||
| θ value | 2.36792 (rank : 72) | NC score | 0.010537 (rank : 149) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 739 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y2Y9, Q9Y356 | Gene names | KLF13, BTEB3, NSLP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 13 (Transcription factor BTEB3) (Basic transcription element-binding protein 3) (BTE-binding protein 3) (RANTES factor of late activated T-lymphocytes 1) (RFLAT-1) (Transcription factor NSLP1) (Novel Sp1-like zinc finger transcription factor 1). | |||||
|
LSP1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 73) | NC score | 0.039733 (rank : 48) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P19973, P97339, Q04950, Q62022, Q62023, Q62024, Q8CD28, Q99L65 | Gene names | Lsp1, Pp52, S37, Wp34 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte-specific protein 1 (Protein pp52) (52 kDa phosphoprotein) (Lymphocyte-specific antigen WP34) (S37 protein). | |||||
|
LTBP2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 74) | NC score | 0.011763 (rank : 143) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 617 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O08999, Q8C6W9 | Gene names | Ltbp2 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 2 precursor (LTBP-2). | |||||
|
NCOA6_HUMAN
|
||||||
| θ value | 2.36792 (rank : 75) | NC score | 0.048864 (rank : 39) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
NCOA6_MOUSE
|
||||||
| θ value | 2.36792 (rank : 76) | NC score | 0.049153 (rank : 37) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
PO6F2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 77) | NC score | 0.022280 (rank : 94) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 618 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P78424, P78425, Q75ME8, Q86UM6, Q9UDS7 | Gene names | POU6F2, RPF1 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 6, transcription factor 2 (Retina-derived POU-domain factor 1) (RPF-1). | |||||
|
RTN4_HUMAN
|
||||||
| θ value | 2.36792 (rank : 78) | NC score | 0.022446 (rank : 92) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NQC3, O94962, Q9BXG5, Q9H212, Q9H3I3, Q9UQ42, Q9Y293, Q9Y2Y7, Q9Y5U6 | Gene names | RTN4, KIAA0886, NOGO | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-4 (Neurite outgrowth inhibitor) (Nogo protein) (Foocen) (Neuroendocrine-specific protein) (NSP) (Neuroendocrine-specific protein C homolog) (RTN-x) (Reticulon-5). | |||||
|
SF3B2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 79) | NC score | 0.035406 (rank : 61) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q13435 | Gene names | SF3B2, SAP145 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 2 (Spliceosome-associated protein 145) (SAP 145) (SF3b150) (Pre-mRNA-splicing factor SF3b 145 kDa subunit). | |||||
|
SMC2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 80) | NC score | 0.014286 (rank : 129) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O95347, Q9P1P2 | Gene names | SMC2L1, CAPE, SMC2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 2-like 1 protein (Chromosome- associated protein E) (hCAP-E) (XCAP-E homolog). | |||||
|
SRCA_HUMAN
|
||||||
| θ value | 2.36792 (rank : 81) | NC score | 0.019909 (rank : 111) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q86TD4 | Gene names | SRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
SRCH_HUMAN
|
||||||
| θ value | 2.36792 (rank : 82) | NC score | 0.028707 (rank : 72) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P23327, Q504Y6 | Gene names | HRC, HCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic reticulum histidine-rich calcium-binding protein precursor. | |||||
|
TGON1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 83) | NC score | 0.037115 (rank : 56) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q62313 | Gene names | Tgoln1, Ttgn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 1 precursor (TGN38A). | |||||
|
TGON2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 84) | NC score | 0.035781 (rank : 60) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q62314 | Gene names | Tgoln2, Ttgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (TGN38B). | |||||
|
USH2A_MOUSE
|
||||||
| θ value | 2.36792 (rank : 85) | NC score | 0.016887 (rank : 118) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 626 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q2QI47, Q9JLP3 | Gene names | Ush2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Usherin precursor (Usher syndrome type-2A protein homolog) (Usher syndrome type IIa protein homolog). | |||||
|
ZBT10_HUMAN
|
||||||
| θ value | 2.36792 (rank : 86) | NC score | 0.013550 (rank : 134) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 831 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96DT7, Q86W96, Q8IXI9, Q96MH9 | Gene names | ZBTB10, RINZF, RINZFC | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 10 (Zinc finger protein RIN ZF). | |||||
|
ZN539_HUMAN
|
||||||
| θ value | 2.36792 (rank : 87) | NC score | 0.005493 (rank : 168) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 758 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q86XL7 | Gene names | ZNF539 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 539. | |||||
|
AN30A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 88) | NC score | 0.009990 (rank : 151) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1119 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9BXX3, Q5W025 | Gene names | ANKRD30A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 30A (NY-BR-1 antigen). | |||||
|
DCDC2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 89) | NC score | 0.024822 (rank : 85) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UHG0, Q5VTR8, Q5VTR9, Q86W35, Q9UFD1, Q9ULR6 | Gene names | DCDC2, KIAA1154 | |||
|
Domain Architecture |
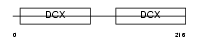
|
|||||
| Description | Doublecortin domain-containing protein 2 (RU2S protein). | |||||
|
EGR1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 90) | NC score | 0.005693 (rank : 167) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 816 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P08046, Q61777 | Gene names | Egr1, Egr-1, Krox-24 | |||
|
Domain Architecture |

|
|||||
| Description | Early growth response protein 1 (EGR-1) (Krox-24 protein) (Transcription factor Zif268) (Nerve growth factor-induced protein A) (NGFI-A). | |||||
|
HSF1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 91) | NC score | 0.027146 (rank : 77) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q00613 | Gene names | HSF1, HSTF1 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock factor protein 1 (HSF 1) (Heat shock transcription factor 1) (HSTF 1). | |||||
|
HSF1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 92) | NC score | 0.026076 (rank : 81) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P38532, O70462 | Gene names | Hsf1 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock factor protein 1 (HSF 1) (Heat shock transcription factor 1) (HSTF 1). | |||||
|
LAMA2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 93) | NC score | 0.017188 (rank : 117) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1198 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q60675, Q05003, Q64061 | Gene names | Lama2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
PCGF6_MOUSE
|
||||||
| θ value | 3.0926 (rank : 94) | NC score | 0.043557 (rank : 45) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q99NA9 | Gene names | Pcgf6, Mblr, Rnf134 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polycomb group RING finger protein 6 (RING finger protein 134) (Mel18 and Bmi1-like RING finger). | |||||
|
SIA1B_MOUSE
|
||||||
| θ value | 3.0926 (rank : 95) | NC score | 0.055879 (rank : 34) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q06985, Q7TPV6 | Gene names | Siah1b | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase SIAH1B (EC 6.3.2.-) (Seven in absentia homolog 1b) (Siah1b) (Siah-1b). | |||||
|
SLIT1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 96) | NC score | 0.005909 (rank : 166) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 700 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75093, Q8WWZ2, Q9UIL7 | Gene names | SLIT1, KIAA0813, MEGF4, SLIL1 | |||
|
Domain Architecture |

|
|||||
| Description | Slit homolog 1 protein precursor (Slit-1) (Multiple epidermal growth factor-like domains 4). | |||||
|
WNK3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 97) | NC score | 0.007248 (rank : 163) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9BYP7, Q9HCK6 | Gene names | WNK3, KIAA1566, PRKWNK3 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK3 (EC 2.7.11.1) (Protein kinase with no lysine 3) (Protein kinase, lysine-deficient 3). | |||||
|
ZN544_HUMAN
|
||||||
| θ value | 3.0926 (rank : 98) | NC score | 0.004922 (rank : 172) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6NX49, Q9UEX4 | Gene names | ZNF544 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 544. | |||||
|
AIPL1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 99) | NC score | 0.016485 (rank : 121) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NZN9, Q9H873, Q9NS10 | Gene names | AIPL1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl-hydrocarbon-interacting protein-like 1. | |||||
|
K0310_HUMAN
|
||||||
| θ value | 4.03905 (rank : 100) | NC score | 0.040817 (rank : 46) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O15027, Q5SXP0, Q5SXP1, Q8N347, Q96HP1 | Gene names | KIAA0310 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0310. | |||||
|
MFA3L_HUMAN
|
||||||
| θ value | 4.03905 (rank : 101) | NC score | 0.011115 (rank : 146) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75121, Q6TNA8, Q9BVE1, Q9BXK0 | Gene names | MFAP3L, KIAA0626 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microfibrillar-associated protein 3-like precursor (Protein kinase NYD-SP9). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 4.03905 (rank : 102) | NC score | 0.035810 (rank : 59) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
MOES_MOUSE
|
||||||
| θ value | 4.03905 (rank : 103) | NC score | 0.008543 (rank : 158) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 923 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P26041, Q3UL28, Q8BSN4 | Gene names | Msn | |||
|
Domain Architecture |

|
|||||
| Description | Moesin (Membrane-organizing extension spike protein). | |||||
|
MRP_HUMAN
|
||||||
| θ value | 4.03905 (rank : 104) | NC score | 0.037643 (rank : 55) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P49006, Q5TEE6, Q6NXS5 | Gene names | MARCKSL1, MLP, MRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MARCKS-related protein (MARCKS-like protein 1) (Macrophage myristoylated alanine-rich C kinase substrate) (Mac-MARCKS) (MacMARCKS). | |||||
|
MRP_MOUSE
|
||||||
| θ value | 4.03905 (rank : 105) | NC score | 0.037952 (rank : 53) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P28667, Q3TEZ4, Q91W07 | Gene names | Marcksl1, Mlp, Mrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MARCKS-related protein (MARCKS-like protein 1) (Macrophage myristoylated alanine-rich C kinase substrate) (Mac-MARCKS) (MacMARCKS) (Brain protein F52). | |||||
|
MTG8R_HUMAN
|
||||||
| θ value | 4.03905 (rank : 106) | NC score | 0.047960 (rank : 40) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O43439, Q5TGE4, Q5TGE7, Q8IWF3, Q96B06, Q96L00, Q9H436, Q9UJP8, Q9UJP9, Q9UP24 | Gene names | CBFA2T2, EHT, MTGR1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein CBFA2T2 (MTG8-like protein) (MTG8-related protein 1) (Myeloid translocation-related protein 1) (ETO homologous on chromosome 20) (p85). | |||||
|
MYB_MOUSE
|
||||||
| θ value | 4.03905 (rank : 107) | NC score | 0.013867 (rank : 132) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P06876, Q61929 | Gene names | Myb | |||
|
Domain Architecture |

|
|||||
| Description | Myb proto-oncogene protein (C-myb). | |||||
|
RFIP3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 108) | NC score | 0.014121 (rank : 130) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1055 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O75154, Q4VXV7, Q9H155, Q9H1G0, Q9NUI0 | Gene names | RAB11FIP3, KIAA0665 | |||
|
Domain Architecture |

|
|||||
| Description | Rab11 family-interacting protein 3 (Rab11-FIP3) (EF hands-containing Rab-interacting protein) (Eferin). | |||||
|
SIAH1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 109) | NC score | 0.050375 (rank : 35) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IUQ4, O43269, Q92880 | Gene names | SIAH1, HUMSIAH | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase SIAH1 (EC 6.3.2.-) (Seven in absentia homolog 1) (Siah-1) (Siah-1a). | |||||
|
SMCA1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 110) | NC score | 0.010700 (rank : 147) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PGB8, Q8BSS1, Q91Y58 | Gene names | Smarca1, Snf2l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable global transcription activator SNF2L1 (EC 3.6.1.-) (Nucleosome remodeling factor subunit SNF2L) (ATP-dependent helicase SMARCA1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 1) (DNA-dependent ATPase SNF2L). | |||||
|
TOB1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 111) | NC score | 0.026454 (rank : 79) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q61471 | Gene names | Tob1, Tob, Trob | |||
|
Domain Architecture |

|
|||||
| Description | Tob1 protein (Transducer of erbB-2 1). | |||||
|
ZN547_HUMAN
|
||||||
| θ value | 4.03905 (rank : 112) | NC score | 0.004581 (rank : 174) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 728 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8IVP9, Q96NC4 | Gene names | ZNF547 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 547. | |||||
|
CBLB_HUMAN
|
||||||
| θ value | 5.27518 (rank : 113) | NC score | 0.012027 (rank : 139) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13191, Q13192, Q13193, Q3LIC0, Q8IVC5 | Gene names | CBLB, RNF56 | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase CBL-B (EC 6.3.2.-) (Signal transduction protein CBL-B) (SH3-binding protein CBL-B) (Casitas B-lineage lymphoma proto-oncogene b) (RING finger protein 56). | |||||
|
CCDC9_HUMAN
|
||||||
| θ value | 5.27518 (rank : 114) | NC score | 0.038329 (rank : 51) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y3X0 | Gene names | CCDC9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 9. | |||||
|
EMID1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 115) | NC score | 0.022265 (rank : 95) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q91VF5 | Gene names | Emid1, Emu1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EMI domain-containing protein 1 precursor (Protein Emu1) (Emilin and multimerin domain-containing protein 1). | |||||
|
HKR1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 116) | NC score | 0.004265 (rank : 175) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 759 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P10072, Q9BSW9, Q9UM09 | Gene names | HKR1 | |||
|
Domain Architecture |
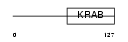
|
|||||
| Description | Krueppel-related zinc finger protein 1 (Protein HKR1). | |||||
|
K0355_HUMAN
|
||||||
| θ value | 5.27518 (rank : 117) | NC score | 0.024612 (rank : 86) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15063 | Gene names | KIAA0355 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0355. | |||||
|
LCP2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 118) | NC score | 0.027995 (rank : 76) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 531 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q13094 | Gene names | LCP2 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte cytosolic protein 2 (SH2 domain-containing leukocyte protein of 76 kDa) (SLP-76 tyrosine phosphoprotein) (SLP76). | |||||
|
LSP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 119) | NC score | 0.039052 (rank : 49) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P33241, Q16096, Q9BUY8 | Gene names | LSP1, WP34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lymphocyte-specific protein 1 (Protein pp52) (52 kDa phosphoprotein) (Lymphocyte-specific antigen WP34) (47 kDa actin-binding protein). | |||||
|
MADCA_HUMAN
|
||||||
| θ value | 5.27518 (rank : 120) | NC score | 0.021458 (rank : 104) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q13477, O60222, O75867 | Gene names | MADCAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucosal addressin cell adhesion molecule 1 precursor (MAdCAM-1) (hMAdCAM-1). | |||||
|
MINT_HUMAN
|
||||||
| θ value | 5.27518 (rank : 121) | NC score | 0.024505 (rank : 87) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 5.27518 (rank : 122) | NC score | 0.020451 (rank : 108) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
NIN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 123) | NC score | 0.011644 (rank : 145) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1416 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8N4C6, Q6P0P6, Q9BWU6, Q9C012, Q9C013, Q9C014, Q9H5I6, Q9HAT7, Q9HBY5, Q9HCK7, Q9UH61 | Gene names | NIN, KIAA1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein (hNinein) (Glycogen synthase kinase 3 beta-interacting protein) (GSK3B-interacting protein). | |||||
|
PCGF6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 124) | NC score | 0.038639 (rank : 50) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9BYE7, Q5SYD1, Q5SYD6, Q96ID9, Q96SJ1 | Gene names | PCGF6, MBLR, RNF134 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polycomb group RING finger protein 6 (RING finger protein 134) (Mel18 and Bmi1-like RING finger). | |||||
|
PININ_MOUSE
|
||||||
| θ value | 5.27518 (rank : 125) | NC score | 0.021764 (rank : 102) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O35691, Q8CD89, Q8CGU3 | Gene names | Pnn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin. | |||||
|
PJA1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 126) | NC score | 0.013857 (rank : 133) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NG27, Q8NG28, Q9HAC1 | Gene names | PJA1, RNF70 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin protein ligase Praja1 (EC 6.3.2.-) (RING finger protein 70). | |||||
|
RAD52_MOUSE
|
||||||
| θ value | 5.27518 (rank : 127) | NC score | 0.040083 (rank : 47) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P43352 | Gene names | Rad52 | |||
|
Domain Architecture |
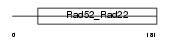
|
|||||
| Description | DNA repair protein RAD52 homolog. | |||||
|
SC24A_MOUSE
|
||||||
| θ value | 5.27518 (rank : 128) | NC score | 0.020699 (rank : 106) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q3U2P1, Q3TQ05, Q3TRG7, Q8BIS0 | Gene names | Sec24a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein transport protein Sec24A (SEC24-related protein A). | |||||
|
SMCA1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 129) | NC score | 0.010692 (rank : 148) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P28370, Q5JV41, Q5JV42 | Gene names | SMARCA1, SNF2L, SNF2L1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L1 (EC 3.6.1.-) (Nucleosome remodeling factor subunit SNF2L) (ATP-dependent helicase SMARCA1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 1). | |||||
|
SMRA3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 130) | NC score | 0.021261 (rank : 105) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6PCN7, O35596, O35597, Q80VT6 | Gene names | Smarca3, Snf2l3, Zbu1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 3 (EC 3.6.1.-) (Sucrose nonfermenting protein 2-like 3) (TNF-response element-binding protein) (P113). | |||||
|
SPTA2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 131) | NC score | 0.009955 (rank : 152) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 888 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q13813, Q13186, Q15324, Q16606, Q59EF1, Q5VXV5, Q5VXV6, Q7Z6M5, Q9P0V0 | Gene names | SPTAN1, SPTA2 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin alpha chain, brain (Spectrin, non-erythroid alpha chain) (Alpha-II spectrin) (Fodrin alpha chain). | |||||
|
SYNE1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 132) | NC score | 0.014456 (rank : 128) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
TDRD4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 133) | NC score | 0.022013 (rank : 98) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NUY9 | Gene names | TDRD4 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 4. | |||||
|
YBOX2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 134) | NC score | 0.014528 (rank : 127) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y2T7, Q8N4P0 | Gene names | YBX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Y-box-binding protein 2 (Germ cell-specific Y-box-binding protein) (Contrin) (MSY2 homolog). | |||||
|
YLPM1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 135) | NC score | 0.032209 (rank : 65) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
CO1A1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 136) | NC score | 0.021682 (rank : 103) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P11087, Q53WT0, Q60635, Q61367, Q61427, Q63919, Q6PCL3, Q810J9 | Gene names | Col1a1, Cola1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(I) chain precursor. | |||||
|
CO3A1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 137) | NC score | 0.021913 (rank : 99) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P02461, Q15112, Q16403, Q6LDB3, Q6LDJ2, Q6LDJ3, Q7KZ56 | Gene names | COL3A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(III) chain precursor. | |||||
|
COBA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 138) | NC score | 0.023487 (rank : 89) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P12107, Q14034, Q9UIT4, Q9UIT5, Q9UIT6 | Gene names | COL11A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XI) chain precursor. | |||||
|
COEA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 139) | NC score | 0.013462 (rank : 135) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q05707, O00260, O00261, O00262, Q05708, Q5XJ18, Q96C67 | Gene names | COL14A1, UND | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XIV) chain precursor (Undulin). | |||||
|
DIAP1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 140) | NC score | 0.010317 (rank : 150) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 839 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O08808 | Gene names | Diaph1, Diap1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1) (mDIA1) (p140mDIA). | |||||
|
EPC2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 141) | NC score | 0.011828 (rank : 141) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8C0I4, Q3URX9, Q80Y70, Q8BXC5 | Gene names | Epc2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Enhancer of polycomb homolog 2. | |||||
|
KRHB4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 142) | NC score | 0.001270 (rank : 177) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NSB2 | Gene names | KRTHB4 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type II cuticular Hb4 (Hair keratin, type II Hb4). | |||||
|
LMBL3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 143) | NC score | 0.013945 (rank : 131) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BLB7, Q641L7, Q6ZPI2, Q8C0G4 | Gene names | L3mbtl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lethal(3)malignant brain tumor-like 3 protein (L(3)mbt-like 3 protein). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 6.88961 (rank : 144) | NC score | 0.028161 (rank : 75) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
PININ_HUMAN
|
||||||
| θ value | 6.88961 (rank : 145) | NC score | 0.018652 (rank : 114) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9H307, O60899, Q53EM7, Q6P5X4, Q7KYL1, Q99738, Q9UHZ9, Q9UQR9 | Gene names | PNN, DRS, MEMA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin (140 kDa nuclear and cell adhesion-related phosphoprotein) (Domain-rich serine protein) (DRS-protein) (DRSP) (Melanoma metastasis clone A protein) (Desmosome-associated protein) (SR-like protein) (Nuclear protein SDK3). | |||||
|
SON_HUMAN
|
||||||
| θ value | 6.88961 (rank : 146) | NC score | 0.028469 (rank : 74) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
SSRP1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 147) | NC score | 0.025584 (rank : 83) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q08945, Q5BJG8 | Gene names | SSRP1, FACT80 | |||
|
Domain Architecture |

|
|||||
| Description | FACT complex subunit SSRP1 (Facilitates chromatin transcription complex subunit SSRP1) (FACT 80 kDa subunit) (FACTp80) (Chromatin- specific transcription elongation factor 80 kDa subunit) (Structure- specific recognition protein 1) (hSSRP1) (Recombination signal sequence recognition protein 1) (T160). | |||||
|
SSRP1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 148) | NC score | 0.026399 (rank : 80) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q08943, Q3U9Z2, Q3UJ75, Q4V9U4, Q8CGA6 | Gene names | Ssrp1 | |||
|
Domain Architecture |

|
|||||
| Description | FACT complex subunit SSRP1 (Facilitates chromatin transcription complex subunit SSRP1) (Structure-specific recognition protein 1) (Recombination signal sequence recognition protein 1) (T160). | |||||
|
TAOK3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 149) | NC score | 0.009303 (rank : 155) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1458 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8BYC6, Q3V3B3 | Gene names | Taok3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO3 (EC 2.7.11.1) (Thousand and one amino acid protein 3). | |||||
|
TF3C2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 150) | NC score | 0.012858 (rank : 137) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BL74, Q3U7Z9, Q80XQ7, Q8BJJ4, Q8BLP3, Q8K1J1, Q9CSS0, Q9CVE6, Q9DBK6 | Gene names | Gtf3c2, Kiaa0011 | |||
|
Domain Architecture |

|
|||||
| Description | General transcription factor 3C polypeptide 2 (Transcription factor IIIC-subunit beta) (TF3C-beta) (TFIIIC 110 kDa subunit) (TFIIIC110). | |||||
|
ZBTB2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 151) | NC score | 0.006061 (rank : 164) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 752 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8N680, Q5SZ81, Q9P245 | Gene names | ZBTB2, KIAA1483, ZNF437 | |||
|
Domain Architecture |
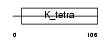
|
|||||
| Description | Zinc finger and BTB domain-containing protein 2. | |||||
|
ZN451_HUMAN
|
||||||
| θ value | 6.88961 (rank : 152) | NC score | 0.012492 (rank : 138) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 627 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Y4E5, Q5VVF1, Q8N380, Q8TD15, Q9NQM1 | Gene names | ZNF451, COASTER, KIAA0576 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 451 (Coactivator for steroid receptors). | |||||
|
AATK_MOUSE
|
||||||
| θ value | 8.99809 (rank : 153) | NC score | 0.007398 (rank : 162) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 942 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q80YE4, O35211, Q3U2U5, Q3UHR8, Q66JN3, Q80YE3, Q8CB63, Q8CHE2 | Gene names | Aatk, Kiaa0641 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Apoptosis-associated tyrosine-protein kinase (EC 2.7.10.2) (AATYK) (Brain apoptosis-associated tyrosine kinase). | |||||
|
ACK1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 154) | NC score | 0.004814 (rank : 173) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1080 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O54967, Q8C2U0, Q8K0K4 | Gene names | Tnk2, Ack1 | |||
|
Domain Architecture |
|
|||||
| Description | Activated CDC42 kinase 1 (EC 2.7.10.2) (ACK-1) (Non-receptor protein tyrosine kinase Ack) (Tyrosine kinase non-receptor protein 2). | |||||
|
AP3B2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 155) | NC score | 0.016358 (rank : 124) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9JME5, Q6QR53, Q8R1E5 | Gene names | Ap3b2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain). | |||||
|
BRE1B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 156) | NC score | 0.017481 (rank : 116) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1040 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O75150, Q6AHZ6, Q6N005, Q7L3T6, Q8N615, Q96T18, Q9BSV9, Q9HC82 | Gene names | RNF40, BRE1B, KIAA0661 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase BRE1B (EC 6.3.2.-) (BRE1-B) (RING finger protein 40) (95 kDa retinoblastoma-associated protein) (RBP95). | |||||
|
BRE1B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 157) | NC score | 0.016562 (rank : 120) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 970 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q3U319, Q6ZQ75, Q8BJA1, Q8BY03, Q8CHX4 | Gene names | Rnf40, Bre1b, Kiaa0661 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase BRE1B (EC 6.3.2.-) (BRE1-B) (RING finger protein 40). | |||||
|
CHRD_MOUSE
|
||||||
| θ value | 8.99809 (rank : 158) | NC score | 0.011782 (rank : 142) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z0E2 | Gene names | Chrd | |||
|
Domain Architecture |

|
|||||
| Description | Chordin precursor. | |||||
|
CO1A2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 159) | NC score | 0.019710 (rank : 112) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q01149, Q8CGA5 | Gene names | Col1a2, Cola2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(I) chain precursor. | |||||
|
CO5A2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 160) | NC score | 0.023360 (rank : 90) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P05997 | Gene names | COL5A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(V) chain precursor. | |||||
|
COBA1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 161) | NC score | 0.026051 (rank : 82) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61245, Q64047 | Gene names | Col11a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XI) chain precursor. | |||||
|
FOXGB_MOUSE
|
||||||
| θ value | 8.99809 (rank : 162) | NC score | 0.007673 (rank : 160) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q60987, Q80VP3 | Gene names | Foxg1b, Fkhl1, Foxg1, Hfhbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein G1B (Forkhead-related protein FKHL1) (Transcription factor BF-1) (Brain factor 1) (BF1). | |||||
|
HEMGN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 163) | NC score | 0.021809 (rank : 100) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9BXL5, Q6XAR3, Q86XY5, Q9NPC0 | Gene names | HEMGN, EDAG, NDR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hemogen (Hemopoietic gene protein) (Negative differentiation regulator protein) (Erythroid differentiation-associated gene protein) (EDAG-1). | |||||
|
I17RA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 164) | NC score | 0.022223 (rank : 96) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96F46, O43844 | Gene names | IL17RA, IL17R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor A precursor (IL-17 receptor). | |||||
|
IASPP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 165) | NC score | 0.013407 (rank : 136) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8WUF5, Q2PNZ9, Q5DU71, Q5I1X4, Q6P1R7, Q6PKF8, Q9Y290 | Gene names | PPP1R13L, IASPP, NKIP1, PPP1R13BL, RAI | |||
|
Domain Architecture |
|
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
IASPP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 166) | NC score | 0.020490 (rank : 107) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1024 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q5I1X5 | Gene names | Ppp1r13l, Nkip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
LRSM1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 167) | NC score | 0.008974 (rank : 156) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1078 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q6UWE0, Q5VVV0, Q8NB40, Q96GT5, Q96MX5, Q96MZ7 | Gene names | LRSAM1, TAL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein LRSAM1 (EC 6.3.2.-) (Leucine-rich repeat and sterile alpha motif-containing protein 1) (Tsg101-associated ligase) (hTAL). | |||||
|
MTF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 168) | NC score | 0.003958 (rank : 176) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 814 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q14872, Q96CB1 | Gene names | MTF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-regulatory transcription factor 1 (Transcription factor MTF-1) (MRE-binding transcription factor). | |||||
|
NOL1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 169) | NC score | 0.034271 (rank : 62) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P46087 | Gene names | NOL1 | |||
|
Domain Architecture |

|
|||||
| Description | Proliferating-cell nucleolar antigen p120 (Proliferation-associated nucleolar protein p120). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 8.99809 (rank : 170) | NC score | 0.026885 (rank : 78) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
RNF24_HUMAN
|
||||||
| θ value | 8.99809 (rank : 171) | NC score | 0.015645 (rank : 126) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y225, Q9UMH1 | Gene names | RNF24 | |||
|
Domain Architecture |
|
|||||
| Description | RING finger protein 24. | |||||
|
RT32_HUMAN
|
||||||
| θ value | 8.99809 (rank : 172) | NC score | 0.016591 (rank : 119) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y6G3, Q96Q48, Q9P0S1 | Gene names | MRPS32, MRPL31, MRPL42 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial 28S ribosomal protein S32 (S32mt) (MRP-S32). | |||||
|
SOX8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 173) | NC score | 0.005163 (rank : 169) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P57073, Q9NZW2 | Gene names | SOX8 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-8. | |||||
|
STAC3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 174) | NC score | 0.007635 (rank : 161) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BZ71 | Gene names | Stac3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and cysteine-rich domain-containing protein 3. | |||||
|
TRIP6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 175) | NC score | 0.005162 (rank : 170) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q15654, O15170, O15275, Q9BTB2, Q9UNT4 | Gene names | TRIP6, OIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 6 (TRIP6) (OPA-interacting protein 1) (Zyxin-related protein 1) (ZRP-1). | |||||
|
VCX3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 176) | NC score | 0.022424 (rank : 93) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NNX9, Q9P0H4 | Gene names | VCX3A, VCX3, VCX8R, VCXA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Variable charge X-linked protein 3 (VCX-A protein) (Variably charged protein X-A) (Variable charge protein on X with eight repeats) (VCX- 8r). | |||||
|
VCXC_HUMAN
|
||||||
| θ value | 8.99809 (rank : 177) | NC score | 0.020217 (rank : 109) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9H321 | Gene names | VCXC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | VCX-C protein (Variably charged protein X-C). | |||||
|
UB7I1_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 177 | |
| SwissProt Accessions | Q9NWF9, Q6Y691, Q75ML7, Q7Z2H7, Q7Z7C1, Q8NHW7, Q9NYT1 | Gene names | UBCE7IP1, ZIN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin ligase TRIAD3 (EC 6.3.2.-) (Ubiquitin-conjugating enzyme 7-interacting protein 1) (Zinc finger protein inhibiting NF-kappa-B) (Triad domain-containing protein 3). | |||||
|
UB7I1_MOUSE
|
||||||
| NC score | 0.965303 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 87 | |
| SwissProt Accessions | P58283, Q3U493, Q3UE56, Q3UGM3, Q68FN0, Q6P1H8, Q6PWY5, Q8BN27, Q8C1U3 | Gene names | Ubce7ip1, Triad3, Uip83, Zin | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin ligase TRIAD3 (EC 6.3.2.-) (Ubiquitin-conjugating enzyme 7-interacting protein 1) (UbcM4-interacting protein 83) (Triad domain- containing protein 3). | |||||
|
UB7I4_HUMAN
|
||||||
| NC score | 0.616461 (rank : 3) | θ value | 6.85773e-16 (rank : 3) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P50876 | Gene names | RNF144, KIAA0161, UBCE7IP4 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-conjugating enzyme 7-interacting protein 4 (UbcM4- interacting protein 4) (RING finger protein 144). | |||||
|
UB7I4_MOUSE
|
||||||
| NC score | 0.603191 (rank : 4) | θ value | 8.38298e-14 (rank : 4) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q925F3, Q6A0B4 | Gene names | Rnf144, Kiaa0161, Ubce7ip4, Uip4 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-conjugating enzyme 7-interacting protein 4 (UbcM4- interacting protein 4) (RING finger protein 144). | |||||
|
IBRD1_HUMAN
|
||||||
| NC score | 0.572457 (rank : 5) | θ value | 2.36244e-08 (rank : 14) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8TC41, Q9BX48 | Gene names | IBRDC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IBR domain-containing protein 1. | |||||
|
RNF14_HUMAN
|
||||||
| NC score | 0.545250 (rank : 6) | θ value | 4.30538e-10 (rank : 5) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UBS8, O94793, Q6IBV0 | Gene names | RNF14, ARA54 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 14 (Androgen receptor-associated protein 54) (Triad2 protein) (HFB30). | |||||
|
IBRD2_HUMAN
|
||||||
| NC score | 0.545000 (rank : 7) | θ value | 2.61198e-07 (rank : 17) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q7Z419, Q6P4Q0, Q8N3R7, Q9BX38 | Gene names | IBRDC2, P53RFP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin ligase IBRDC2 (EC 6.3.2.-) (IBR domain-containing protein 2) (p53-inducible RING finger protein). | |||||
|
IBRD2_MOUSE
|
||||||
| NC score | 0.527247 (rank : 8) | θ value | 1.43324e-05 (rank : 18) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BKD6, Q8BG97 | Gene names | Ibrdc2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin ligase IBRDC2 (EC 6.3.2.-) (IBR domain-containing protein 2). | |||||
|
RNF14_MOUSE
|
||||||
| NC score | 0.526700 (rank : 9) | θ value | 8.97725e-08 (rank : 16) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JI90, Q9D0L2, Q9D6N2, Q9D6Z8, Q9JI89 | Gene names | Rnf14, Ara54, Triad2 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 14 (Androgen receptor-associated protein 54) (Triad2 protein). | |||||
|
ARI1_HUMAN
|
||||||
| NC score | 0.525677 (rank : 10) | θ value | 1.06045e-08 (rank : 12) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Y4X5, O76026, Q9H3T6, Q9UEN0, Q9UP39 | Gene names | ARIH1, ARI, UBCH7BP | |||
|
Domain Architecture |

|
|||||
| Description | Protein ariadne-1 homolog (ARI-1) (Ubiquitin-conjugating enzyme E2- binding protein 1) (UbcH7-binding protein) (UbcM4-interacting protein) (HHARI) (H7-AP2) (MOP-6). | |||||
|
ARI1_MOUSE
|
||||||
| NC score | 0.525653 (rank : 11) | θ value | 1.06045e-08 (rank : 13) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Z1K5, Q6PF92 | Gene names | Arih1, Ari, Ubch7bp, Uip77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ariadne-1 homolog (ARI-1) (Ubiquitin-conjugating enzyme E2- binding protein 1) (UbcH7-binding protein) (UbcM4-interacting protein 77). | |||||
|
ARI2_HUMAN
|
||||||
| NC score | 0.523999 (rank : 12) | θ value | 2.13673e-09 (rank : 10) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O95376, Q9HBZ6, Q9UEM9 | Gene names | ARIH2, ARI2, TRIAD1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ariadne-2 homolog (ARI-2) (Triad1 protein). | |||||
|
IBRD3_HUMAN
|
||||||
| NC score | 0.523135 (rank : 13) | θ value | 3.08544e-08 (rank : 15) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6ZMZ0, Q6P6A4 | Gene names | IBRDC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IBR domain-containing protein 3. | |||||
|
ARI2_MOUSE
|
||||||
| NC score | 0.521875 (rank : 14) | θ value | 3.64472e-09 (rank : 11) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Z1K6 | Gene names | Arih2, Ari2, Triad1, Uip48 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ariadne-2 homolog (ARI-2) (Triad1 protein) (UbcM4-interacting protein 48). | |||||
|
UB7I3_HUMAN
|
||||||
| NC score | 0.517941 (rank : 15) | θ value | 5.62301e-10 (rank : 7) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9BYM8, O95623, Q96BS3, Q9BYM9 | Gene names | RBCK1, C20orf18, RNF54, UBCE7IP3, XAP4 | |||
|
Domain Architecture |

|
|||||
| Description | RanBP-type and C3HC4-type zinc finger-containing protein 1 (Ubiquitin- conjugating enzyme 7-interacting protein 3) (Hepatitis B virus X- associated protein 4) (HBV-associated factor 4) (RING finger protein 54). | |||||
|
UB7I3_MOUSE
|
||||||
| NC score | 0.516157 (rank : 16) | θ value | 4.30538e-10 (rank : 6) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9WUB0 | Gene names | Rbck1, Rbck, Ubce7ip3, Uip28 | |||
|
Domain Architecture |

|
|||||
| Description | RanBP-type and C3HC4-type zinc finger-containing protein 1 (Ubiquitin- conjugating enzyme 7-interacting protein 3) (UbcM4-interacting protein 28). | |||||
|
RNF19_MOUSE
|
||||||
| NC score | 0.508547 (rank : 17) | θ value | 1.63604e-09 (rank : 9) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P50636, Q9QUJ5 | Gene names | Rnf19, Geg-154, Xybp | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 19 (XY body protein) (XYbp) (Gametogenesis expressed protein GEG-154) (UBCM4-interacting protein 117) (UIP117). | |||||
|
RNF19_HUMAN
|
||||||
| NC score | 0.506987 (rank : 18) | θ value | 1.63604e-09 (rank : 8) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9NV58, Q9H5H9, Q9H8M8, Q9UFG0, Q9UFX6, Q9Y4Y1 | Gene names | RNF19 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 19 (Dorfin) (Double ring-finger protein) (p38 protein). | |||||
|
PRKN2_MOUSE
|
||||||
| NC score | 0.380158 (rank : 19) | θ value | 0.00228821 (rank : 19) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9WVS6, Q9ES22, Q9ES23 | Gene names | Park2, Prkn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parkin (EC 6.3.2.-) (Ubiquitin E3 ligase PRKN). | |||||
|
PRKN2_HUMAN
|
||||||
| NC score | 0.378546 (rank : 20) | θ value | 0.00298849 (rank : 20) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O60260, Q5TFV8, Q6Q2I6, Q8NI41, Q8NI43, Q8NI44, Q8WW07 | Gene names | PARK2, PRKN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parkin (EC 6.3.2.-) (Ubiquitin E3 ligase PRKN) (Parkinson juvenile disease protein 2) (Parkinson disease protein 2). | |||||
|
RNF31_MOUSE
|
||||||
| NC score | 0.350995 (rank : 21) | θ value | 0.00298849 (rank : 21) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q924T7 | Gene names | Rnf31 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 31. | |||||
|
RNF31_HUMAN
|
||||||
| NC score | 0.350410 (rank : 22) | θ value | 0.00509761 (rank : 22) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q96EP0, Q86VI2, Q8TEI0, Q96GB4, Q96NF1, Q9H5F1, Q9NWD2 | Gene names | RNF31, ZIBRA | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 31 (Zinc in-between-RING-finger ubiquitin- associated domain protein). | |||||
|
PARC_HUMAN
|
||||||
| NC score | 0.309899 (rank : 23) | θ value | 0.0252991 (rank : 24) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8IWT3, O75188, Q68CP2, Q68D92, Q8N3W9, Q9BU56 | Gene names | PARC, H7AP1, KIAA0708 | |||
|
Domain Architecture |

|
|||||
| Description | p53-associated parkin-like cytoplasmic protein (UbcH7-associated protein 1). | |||||
|
PARC_MOUSE
|
||||||
| NC score | 0.292388 (rank : 24) | θ value | 1.06291 (rank : 47) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q80TT8, Q8BKL9, Q8CGC0, Q8R363 | Gene names | Parc, Kiaa0708 | |||
|
Domain Architecture |

|
|||||
| Description | p53-associated parkin-like cytoplasmic protein. | |||||
|
MTG8R_MOUSE
|
||||||
| NC score | 0.067444 (rank : 25) | θ value | 0.21417 (rank : 31) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O70374 | Gene names | Cbfa2t2, Cbfa2t2h, Mtgr1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein CBFA2T2 (MTG8-like protein) (MTG8-related protein 1). | |||||
|
SIAH2_HUMAN
|
||||||
| NC score | 0.061990 (rank : 26) | θ value | 1.81305 (rank : 61) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O43255, O43270 | Gene names | SIAH2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase SIAH2 (EC 6.3.2.-) (Seven in absentia homolog 2) (Siah-2) (hSiah2). | |||||
|
SIAH2_MOUSE
|
||||||
| NC score | 0.061844 (rank : 27) | θ value | 1.81305 (rank : 62) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q06986 | Gene names | Siah2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase SIAH2 (EC 6.3.2.-) (Seven in absentia homolog 2) (Siah-2) (mSiah2). | |||||
|
TFG_HUMAN
|
||||||
| NC score | 0.061534 (rank : 28) | θ value | 0.62314 (rank : 42) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92734, Q15656 | Gene names | TFG | |||
|
Domain Architecture |

|
|||||
| Description | Protein TFG (TRK-fused gene protein). | |||||
|
INVO_MOUSE
|
||||||
| NC score | 0.060517 (rank : 29) | θ value | 1.38821 (rank : 50) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P48997 | Gene names | Ivl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Involucrin. | |||||
|
RAD52_HUMAN
|
||||||
| NC score | 0.058137 (rank : 30) | θ value | 0.365318 (rank : 36) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P43351, Q13205 | Gene names | RAD52 | |||
|
Domain Architecture |

|
|||||
| Description | DNA repair protein RAD52 homolog. | |||||
|
RBM12_MOUSE
|
||||||
| NC score | 0.057679 (rank : 31) | θ value | 0.62314 (rank : 41) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8R4X3, Q8K373, Q8R302, Q9CS80 | Gene names | Rbm12 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 12 (RNA-binding motif protein 12) (SH3/WW domain anchor protein in the nucleus) (SWAN). | |||||
|
SIA1A_MOUSE
|
||||||
| NC score | 0.056929 (rank : 32) | θ value | 1.81305 (rank : 60) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P61092, Q06984 | Gene names | Siah1a | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase SIAH1A (EC 6.3.2.-) (Seven in absentia homolog 1a) (Siah1a) (Siah-1a) (mSiah-1a). | |||||
|
ATX7_HUMAN
|
||||||
| NC score | 0.056585 (rank : 33) | θ value | 1.06291 (rank : 46) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O15265, O75328, O75329, Q9Y6P8 | Gene names | ATXN7, SCA7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-7 (Spinocerebellar ataxia type 7 protein). | |||||
|
SIA1B_MOUSE
|
||||||
| NC score | 0.055879 (rank : 34) | θ value | 3.0926 (rank : 95) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q06985, Q7TPV6 | Gene names | Siah1b | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase SIAH1B (EC 6.3.2.-) (Seven in absentia homolog 1b) (Siah1b) (Siah-1b). | |||||
|
SIAH1_HUMAN
|
||||||
| NC score | 0.050375 (rank : 35) | θ value | 4.03905 (rank : 109) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IUQ4, O43269, Q92880 | Gene names | SIAH1, HUMSIAH | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase SIAH1 (EC 6.3.2.-) (Seven in absentia homolog 1) (Siah-1) (Siah-1a). | |||||
|
BRD4_HUMAN
|
||||||
| NC score | 0.049168 (rank : 36) | θ value | 0.125558 (rank : 28) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
NCOA6_MOUSE
|
||||||
| NC score | 0.049153 (rank : 37) | θ value | 2.36792 (rank : 76) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
RP1L1_HUMAN
|
||||||
| NC score | 0.049018 (rank : 38) | θ value | 0.163984 (rank : 30) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
NCOA6_HUMAN
|
||||||
| NC score | 0.048864 (rank : 39) | θ value | 2.36792 (rank : 75) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
MTG8R_HUMAN
|
||||||
| NC score | 0.047960 (rank : 40) | θ value | 4.03905 (rank : 106) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O43439, Q5TGE4, Q5TGE7, Q8IWF3, Q96B06, Q96L00, Q9H436, Q9UJP8, Q9UJP9, Q9UP24 | Gene names | CBFA2T2, EHT, MTGR1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein CBFA2T2 (MTG8-like protein) (MTG8-related protein 1) (Myeloid translocation-related protein 1) (ETO homologous on chromosome 20) (p85). | |||||
|
BRD4_MOUSE
|
||||||
| NC score | 0.047050 (rank : 41) | θ value | 0.125558 (rank : 29) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
MYST3_MOUSE
|
||||||
| NC score | 0.045215 (rank : 42) | θ value | 0.0736092 (rank : 27) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
ATBF1_MOUSE
|
||||||
| NC score | 0.044309 (rank : 43) | θ value | 0.0113563 (rank : 23) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1841 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q61329 | Gene names | Atbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
ZN198_HUMAN
|
||||||
| NC score | 0.044115 (rank : 44) | θ value | 0.62314 (rank : 44) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UBW7, O43212, O43434, O60898, Q5W0Q4, Q63HP0, Q8NE39, Q9H538, Q9UEU2 | Gene names | ZNF198, FIM, RAMP, ZMYM2 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 198 (Zinc finger MYM-type protein 2) (Fused in myeloproliferative disorders protein) (Rearranged in atypical myeloproliferative disorder protein). | |||||
|
PCGF6_MOUSE
|
||||||
| NC score | 0.043557 (rank : 45) | θ value | 3.0926 (rank : 94) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q99NA9 | Gene names | Pcgf6, Mblr, Rnf134 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polycomb group RING finger protein 6 (RING finger protein 134) (Mel18 and Bmi1-like RING finger). | |||||
|
K0310_HUMAN
|
||||||
| NC score | 0.040817 (rank : 46) | θ value | 4.03905 (rank : 100) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O15027, Q5SXP0, Q5SXP1, Q8N347, Q96HP1 | Gene names | KIAA0310 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0310. | |||||
|
RAD52_MOUSE
|
||||||
| NC score | 0.040083 (rank : 47) | θ value | 5.27518 (rank : 127) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P43352 | Gene names | Rad52 | |||
|
Domain Architecture |

|
|||||
| Description | DNA repair protein RAD52 homolog. | |||||
|
LSP1_MOUSE
|
||||||
| NC score | 0.039733 (rank : 48) | θ value | 2.36792 (rank : 73) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P19973, P97339, Q04950, Q62022, Q62023, Q62024, Q8CD28, Q99L65 | Gene names | Lsp1, Pp52, S37, Wp34 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte-specific protein 1 (Protein pp52) (52 kDa phosphoprotein) (Lymphocyte-specific antigen WP34) (S37 protein). | |||||
|
LSP1_HUMAN
|
||||||
| NC score | 0.039052 (rank : 49) | θ value | 5.27518 (rank : 119) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P33241, Q16096, Q9BUY8 | Gene names | LSP1, WP34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lymphocyte-specific protein 1 (Protein pp52) (52 kDa phosphoprotein) (Lymphocyte-specific antigen WP34) (47 kDa actin-binding protein). | |||||
|
PCGF6_HUMAN
|
||||||
| NC score | 0.038639 (rank : 50) | θ value | 5.27518 (rank : 124) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9BYE7, Q5SYD1, Q5SYD6, Q96ID9, Q96SJ1 | Gene names | PCGF6, MBLR, RNF134 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polycomb group RING finger protein 6 (RING finger protein 134) (Mel18 and Bmi1-like RING finger). | |||||
|
CCDC9_HUMAN
|
||||||
| NC score | 0.038329 (rank : 51) | θ value | 5.27518 (rank : 114) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y3X0 | Gene names | CCDC9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 9. | |||||
|
BL1S3_HUMAN
|
||||||
| NC score | 0.038242 (rank : 52) | θ value | 2.36792 (rank : 65) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6QNY0 | Gene names | BLOC1S3, BLOS3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Biogenesis of lysosome-related organelles complex-1 subunit 3 (BLOC subunit 3). | |||||
|
MRP_MOUSE
|
||||||
| NC score | 0.037952 (rank : 53) | θ value | 4.03905 (rank : 105) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P28667, Q3TEZ4, Q91W07 | Gene names | Marcksl1, Mlp, Mrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MARCKS-related protein (MARCKS-like protein 1) (Macrophage myristoylated alanine-rich C kinase substrate) (Mac-MARCKS) (MacMARCKS) (Brain protein F52). | |||||
|
PRG4_MOUSE
|
||||||
| NC score | 0.037872 (rank : 54) | θ value | 1.06291 (rank : 48) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
MRP_HUMAN
|
||||||
| NC score | 0.037643 (rank : 55) | θ value | 4.03905 (rank : 104) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P49006, Q5TEE6, Q6NXS5 | Gene names | MARCKSL1, MLP, MRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MARCKS-related protein (MARCKS-like protein 1) (Macrophage myristoylated alanine-rich C kinase substrate) (Mac-MARCKS) (MacMARCKS). | |||||
|
TGON1_MOUSE
|
||||||
| NC score | 0.037115 (rank : 56) | θ value | 2.36792 (rank : 83) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q62313 | Gene names | Tgoln1, Ttgn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 1 precursor (TGN38A). | |||||
|
MYST3_HUMAN
|
||||||
| NC score | 0.036896 (rank : 57) | θ value | 1.38821 (rank : 51) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
DIDO1_HUMAN
|
||||||
| NC score | 0.036254 (rank : 58) | θ value | 1.38821 (rank : 49) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.035810 (rank : 59) | θ value | 4.03905 (rank : 102) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
TGON2_MOUSE
|
||||||
| NC score | 0.035781 (rank : 60) | θ value | 2.36792 (rank : 84) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q62314 | Gene names | Tgoln2, Ttgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (TGN38B). | |||||
|
SF3B2_HUMAN
|
||||||
| NC score | 0.035406 (rank : 61) | θ value | 2.36792 (rank : 79) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q13435 | Gene names | SF3B2, SAP145 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 2 (Spliceosome-associated protein 145) (SAP 145) (SF3b150) (Pre-mRNA-splicing factor SF3b 145 kDa subunit). | |||||
|
NOL1_HUMAN
|
||||||
| NC score | 0.034271 (rank : 62) | θ value | 8.99809 (rank : 169) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P46087 | Gene names | NOL1 | |||
|
Domain Architecture |

|
|||||
| Description | Proliferating-cell nucleolar antigen p120 (Proliferation-associated nucleolar protein p120). | |||||
|
RPGR_MOUSE
|
||||||
| NC score | 0.032841 (rank : 63) | θ value | 0.21417 (rank : 32) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R0X5, O88408, Q9CU92 | Gene names | Rpgr | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator (mRpgr). | |||||
|
SFRIP_HUMAN
|
||||||
| NC score | 0.032788 (rank : 64) | θ value | 1.81305 (rank : 59) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q99590, Q8IW59 | Gene names | SFRS2IP, CASP11, SIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SFRS2-interacting protein (Splicing factor, arginine/serine-rich 2- interacting protein) (SC35-interacting protein 1) (CTD-associated SR protein 11) (Splicing regulatory protein 129) (SRrp129) (NY-REN-40 antigen). | |||||
|
YLPM1_MOUSE
|
||||||
| NC score | 0.032209 (rank : 65) | θ value | 5.27518 (rank : 135) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
ATF4_HUMAN
|
||||||
| NC score | 0.031486 (rank : 66) | θ value | 1.81305 (rank : 57) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P18848, Q9UH31 | Gene names | ATF4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-4 (Activating transcription factor 4) (DNA-binding protein TAXREB67) (Cyclic AMP response element-binding protein 2) (CREB2). | |||||
|
CLMN_HUMAN
|
||||||
| NC score | 0.030759 (rank : 67) | θ value | 0.0563607 (rank : 25) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96JQ2, Q9H713, Q9HA23, Q9HA57, Q9UFP4, Q9ULN2 | Gene names | CLMN, KIAA1188 | |||
|
Domain Architecture |

|
|||||
| Description | Calmin. | |||||
|
DLEC1_HUMAN
|
||||||
| NC score | 0.030301 (rank : 68) | θ value | 1.81305 (rank : 58) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y238, Q9NSW0, Q9NTG5 | Gene names | DLEC1, DLC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Deleted in lung and esophageal cancer protein 1 (Deleted in lung cancer protein 1) (DLC-1). | |||||
|
CONA1_MOUSE
|
||||||
| NC score | 0.030147 (rank : 69) | θ value | 0.365318 (rank : 34) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8K4G2, Q5SUQ0 | Gene names | Col23a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXIII) chain. | |||||
|
CCD96_HUMAN
|
||||||
| NC score | 0.030127 (rank : 70) | θ value | 0.62314 (rank : 40) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 595 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q2M329, Q8N2I7 | Gene names | CCDC96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
INVO_HUMAN
|
||||||
| NC score | 0.029398 (rank : 71) | θ value | 0.47712 (rank : 38) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P07476 | Gene names | IVL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Involucrin. | |||||
|
SRCH_HUMAN
|
||||||
| NC score | 0.028707 (rank : 72) | θ value | 2.36792 (rank : 82) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P23327, Q504Y6 | Gene names | HRC, HCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic reticulum histidine-rich calcium-binding protein precursor. | |||||
|
TNR6A_HUMAN
|
||||||
| NC score | 0.028679 (rank : 73) | θ value | 1.81305 (rank : 63) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8NDV7, O15408, Q658L5, Q6NVB5, Q8NEZ0, Q8TBT8, Q8TCR0, Q9NV59, Q9P268 | Gene names | TNRC6A, CAGH26, KIAA1460, TNRC6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trinucleotide repeat-containing gene 6A protein (CAG repeat protein 26) (Glycine-tryptophan protein of 182 kDa) (GW182 autoantigen) (Protein GW1) (EMSY interactor protein). | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.028469 (rank : 74) | θ value | 6.88961 (rank : 146) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.028161 (rank : 75) | θ value | 6.88961 (rank : 144) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
LCP2_HUMAN
|
||||||
| NC score | 0.027995 (rank : 76) | θ value | 5.27518 (rank : 118) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 531 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q13094 | Gene names | LCP2 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte cytosolic protein 2 (SH2 domain-containing leukocyte protein of 76 kDa) (SLP-76 tyrosine phosphoprotein) (SLP76). | |||||
|
HSF1_HUMAN
|
||||||
| NC score | 0.027146 (rank : 77) | θ value | 3.0926 (rank : 91) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q00613 | Gene names | HSF1, HSTF1 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock factor protein 1 (HSF 1) (Heat shock transcription factor 1) (HSTF 1). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.026885 (rank : 78) | θ value | 8.99809 (rank : 170) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
TOB1_MOUSE
|
||||||
| NC score | 0.026454 (rank : 79) | θ value | 4.03905 (rank : 111) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q61471 | Gene names | Tob1, Tob, Trob | |||
|
Domain Architecture |

|
|||||
| Description | Tob1 protein (Transducer of erbB-2 1). | |||||
|
SSRP1_MOUSE
|
||||||
| NC score | 0.026399 (rank : 80) | θ value | 6.88961 (rank : 148) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q08943, Q3U9Z2, Q3UJ75, Q4V9U4, Q8CGA6 | Gene names | Ssrp1 | |||
|
Domain Architecture |

|
|||||
| Description | FACT complex subunit SSRP1 (Facilitates chromatin transcription complex subunit SSRP1) (Structure-specific recognition protein 1) (Recombination signal sequence recognition protein 1) (T160). | |||||
|
HSF1_MOUSE
|
||||||
| NC score | 0.026076 (rank : 81) | θ value | 3.0926 (rank : 92) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P38532, O70462 | Gene names | Hsf1 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock factor protein 1 (HSF 1) (Heat shock transcription factor 1) (HSTF 1). | |||||
|
COBA1_MOUSE
|
||||||
| NC score | 0.026051 (rank : 82) | θ value | 8.99809 (rank : 161) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61245, Q64047 | Gene names | Col11a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XI) chain precursor. | |||||
|
SSRP1_HUMAN
|
||||||
| NC score | 0.025584 (rank : 83) | θ value | 6.88961 (rank : 147) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q08945, Q5BJG8 | Gene names | SSRP1, FACT80 | |||
|
Domain Architecture |

|
|||||
| Description | FACT complex subunit SSRP1 (Facilitates chromatin transcription complex subunit SSRP1) (FACT 80 kDa subunit) (FACTp80) (Chromatin- specific transcription elongation factor 80 kDa subunit) (Structure- specific recognition protein 1) (hSSRP1) (Recombination signal sequence recognition protein 1) (T160). | |||||
|
CONA1_HUMAN
|
||||||
| NC score | 0.025166 (rank : 84) | θ value | 2.36792 (rank : 67) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q86Y22, Q8IVR4, Q9NT93 | Gene names | COL23A1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXIII) chain. | |||||
|
DCDC2_HUMAN
|
||||||
| NC score | 0.024822 (rank : 85) | θ value | 3.0926 (rank : 89) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UHG0, Q5VTR8, Q5VTR9, Q86W35, Q9UFD1, Q9ULR6 | Gene names | DCDC2, KIAA1154 | |||
|
Domain Architecture |

|
|||||
| Description | Doublecortin domain-containing protein 2 (RU2S protein). | |||||
|
K0355_HUMAN
|
||||||
| NC score | 0.024612 (rank : 86) | θ value | 5.27518 (rank : 117) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15063 | Gene names | KIAA0355 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0355. | |||||
|
MINT_HUMAN
|
||||||
| NC score | 0.024505 (rank : 87) | θ value | 5.27518 (rank : 121) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
IF4G1_MOUSE
|
||||||
| NC score | 0.023787 (rank : 88) | θ value | 2.36792 (rank : 70) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q6NZJ6, Q6NZN8, Q8BW99 | Gene names | Eif4g1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1). | |||||
|
COBA1_HUMAN
|
||||||
| NC score | 0.023487 (rank : 89) | θ value | 6.88961 (rank : 138) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P12107, Q14034, Q9UIT4, Q9UIT5, Q9UIT6 | Gene names | COL11A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XI) chain precursor. | |||||
|
CO5A2_HUMAN
|
||||||
| NC score | 0.023360 (rank : 90) | θ value | 8.99809 (rank : 160) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P05997 | Gene names | COL5A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(V) chain precursor. | |||||
|
COIA1_MOUSE
|
||||||
| NC score | 0.023294 (rank : 91) | θ value | 2.36792 (rank : 66) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 481 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P39061, Q60672, Q61437, Q62001, Q62002, Q9JK63 | Gene names | Col18a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVIII) chain precursor [Contains: Endostatin]. | |||||
|
RTN4_HUMAN
|
||||||
| NC score | 0.022446 (rank : 92) | θ value | 2.36792 (rank : 78) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NQC3, O94962, Q9BXG5, Q9H212, Q9H3I3, Q9UQ42, Q9Y293, Q9Y2Y7, Q9Y5U6 | Gene names | RTN4, KIAA0886, NOGO | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-4 (Neurite outgrowth inhibitor) (Nogo protein) (Foocen) (Neuroendocrine-specific protein) (NSP) (Neuroendocrine-specific protein C homolog) (RTN-x) (Reticulon-5). | |||||
|
VCX3_HUMAN
|
||||||
| NC score | 0.022424 (rank : 93) | θ value | 8.99809 (rank : 176) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NNX9, Q9P0H4 | Gene names | VCX3A, VCX3, VCX8R, VCXA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Variable charge X-linked protein 3 (VCX-A protein) (Variably charged protein X-A) (Variable charge protein on X with eight repeats) (VCX- 8r). | |||||
|
PO6F2_HUMAN
|
||||||
| NC score | 0.022280 (rank : 94) | θ value | 2.36792 (rank : 77) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 618 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P78424, P78425, Q75ME8, Q86UM6, Q9UDS7 | Gene names | POU6F2, RPF1 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 6, transcription factor 2 (Retina-derived POU-domain factor 1) (RPF-1). | |||||
|
EMID1_MOUSE
|
||||||
| NC score | 0.022265 (rank : 95) | θ value | 5.27518 (rank : 115) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q91VF5 | Gene names | Emid1, Emu1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EMI domain-containing protein 1 precursor (Protein Emu1) (Emilin and multimerin domain-containing protein 1). | |||||
|
I17RA_HUMAN
|
||||||
| NC score | 0.022223 (rank : 96) | θ value | 8.99809 (rank : 164) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96F46, O43844 | Gene names | IL17RA, IL17R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor A precursor (IL-17 receptor). | |||||
|
TRHY_HUMAN
|
||||||
| NC score | 0.022215 (rank : 97) | θ value | 0.62314 (rank : 43) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
TDRD4_HUMAN
|
||||||
| NC score | 0.022013 (rank : 98) | θ value | 5.27518 (rank : 133) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NUY9 | Gene names | TDRD4 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 4. | |||||
|
CO3A1_HUMAN
|
||||||
| NC score | 0.021913 (rank : 99) | θ value | 6.88961 (rank : 137) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P02461, Q15112, Q16403, Q6LDB3, Q6LDJ2, Q6LDJ3, Q7KZ56 | Gene names | COL3A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(III) chain precursor. | |||||
|
HEMGN_HUMAN
|
||||||
| NC score | 0.021809 (rank : 100) | θ value | 8.99809 (rank : 163) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9BXL5, Q6XAR3, Q86XY5, Q9NPC0 | Gene names | HEMGN, EDAG, NDR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hemogen (Hemopoietic gene protein) (Negative differentiation regulator protein) (Erythroid differentiation-associated gene protein) (EDAG-1). | |||||
|
LAMA2_HUMAN
|
||||||
| NC score | 0.021765 (rank : 101) | θ value | 0.47712 (rank : 39) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1092 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P24043, Q14736, Q93022 | Gene names | LAMA2, LAMM | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
PININ_MOUSE
|
||||||
| NC score | 0.021764 (rank : 102) | θ value | 5.27518 (rank : 125) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O35691, Q8CD89, Q8CGU3 | Gene names | Pnn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin. | |||||
|
CO1A1_MOUSE
|
||||||
| NC score | 0.021682 (rank : 103) | θ value | 6.88961 (rank : 136) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P11087, Q53WT0, Q60635, Q61367, Q61427, Q63919, Q6PCL3, Q810J9 | Gene names | Col1a1, Cola1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(I) chain precursor. | |||||
|
MADCA_HUMAN
|
||||||
| NC score | 0.021458 (rank : 104) | θ value | 5.27518 (rank : 120) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q13477, O60222, O75867 | Gene names | MADCAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucosal addressin cell adhesion molecule 1 precursor (MAdCAM-1) (hMAdCAM-1). | |||||
|
SMRA3_MOUSE
|
||||||
| NC score | 0.021261 (rank : 105) | θ value | 5.27518 (rank : 130) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6PCN7, O35596, O35597, Q80VT6 | Gene names | Smarca3, Snf2l3, Zbu1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 3 (EC 3.6.1.-) (Sucrose nonfermenting protein 2-like 3) (TNF-response element-binding protein) (P113). | |||||
|
SC24A_MOUSE
|
||||||
| NC score | 0.020699 (rank : 106) | θ value | 5.27518 (rank : 128) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q3U2P1, Q3TQ05, Q3TRG7, Q8BIS0 | Gene names | Sec24a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein transport protein Sec24A (SEC24-related protein A). | |||||
|
IASPP_MOUSE
|
||||||
| NC score | 0.020490 (rank : 107) | θ value | 8.99809 (rank : 166) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1024 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q5I1X5 | Gene names | Ppp1r13l, Nkip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.020451 (rank : 108) | θ value | 5.27518 (rank : 122) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
VCXC_HUMAN
|
||||||
| NC score | 0.020217 (rank : 109) | θ value | 8.99809 (rank : 177) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9H321 | Gene names | VCXC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | VCX-C protein (Variably charged protein X-C). | |||||
|
MYT1_MOUSE
|
||||||
| NC score | 0.019962 (rank : 110) | θ value | 1.38821 (rank : 52) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CFC2, O08995, Q8CFH1 | Gene names | Myt1, Kiaa0835, Nzf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1 (MyT1) (Neural zinc finger factor 2) (NZF-2). | |||||
|
SRCA_HUMAN
|
||||||
| NC score | 0.019909 (rank : 111) | θ value | 2.36792 (rank : 81) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q86TD4 | Gene names | SRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
CO1A2_MOUSE
|
||||||
| NC score | 0.019710 (rank : 112) | θ value | 8.99809 (rank : 159) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q01149, Q8CGA5 | Gene names | Col1a2, Cola2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(I) chain precursor. | |||||
|
AP3B2_HUMAN
|
||||||
| NC score | 0.019452 (rank : 113) | θ value | 2.36792 (rank : 64) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q13367, O14808 | Gene names | AP3B2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (AP-3 complex beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain) (Neuron-specific vesicle coat protein beta-NAP). | |||||
|
PININ_HUMAN
|
||||||
| NC score | 0.018652 (rank : 114) | θ value | 6.88961 (rank : 145) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9H307, O60899, Q53EM7, Q6P5X4, Q7KYL1, Q99738, Q9UHZ9, Q9UQR9 | Gene names | PNN, DRS, MEMA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin (140 kDa nuclear and cell adhesion-related phosphoprotein) (Domain-rich serine protein) (DRS-protein) (DRSP) (Melanoma metastasis clone A protein) (Desmosome-associated protein) (SR-like protein) (Nuclear protein SDK3). | |||||
|
HCLS1_MOUSE
|
||||||
| NC score | 0.017926 (rank : 115) | θ value | 2.36792 (rank : 69) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P49710 | Gene names | Hcls1, Hs1 | |||
|
Domain Architecture |

|
|||||
| Description | Hematopoietic lineage cell-specific protein (Hematopoietic cell- specific LYN substrate 1) (LckBP1). | |||||
|
BRE1B_HUMAN
|
||||||
| NC score | 0.017481 (rank : 116) | θ value | 8.99809 (rank : 156) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1040 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O75150, Q6AHZ6, Q6N005, Q7L3T6, Q8N615, Q96T18, Q9BSV9, Q9HC82 | Gene names | RNF40, BRE1B, KIAA0661 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase BRE1B (EC 6.3.2.-) (BRE1-B) (RING finger protein 40) (95 kDa retinoblastoma-associated protein) (RBP95). | |||||
|
LAMA2_MOUSE
|
||||||
| NC score | 0.017188 (rank : 117) | θ value | 3.0926 (rank : 93) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1198 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q60675, Q05003, Q64061 | Gene names | Lama2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
USH2A_MOUSE
|
||||||
| NC score | 0.016887 (rank : 118) | θ value | 2.36792 (rank : 85) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 626 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q2QI47, Q9JLP3 | Gene names | Ush2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Usherin precursor (Usher syndrome type-2A protein homolog) (Usher syndrome type IIa protein homolog). | |||||
|
RT32_HUMAN
|
||||||
| NC score | 0.016591 (rank : 119) | θ value | 8.99809 (rank : 172) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y6G3, Q96Q48, Q9P0S1 | Gene names | MRPS32, MRPL31, MRPL42 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial 28S ribosomal protein S32 (S32mt) (MRP-S32). | |||||
|
BRE1B_MOUSE
|
||||||
| NC score | 0.016562 (rank : 120) | θ value | 8.99809 (rank : 157) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 970 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q3U319, Q6ZQ75, Q8BJA1, Q8BY03, Q8CHX4 | Gene names | Rnf40, Bre1b, Kiaa0661 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase BRE1B (EC 6.3.2.-) (BRE1-B) (RING finger protein 40). | |||||
|
AIPL1_HUMAN
|
||||||
| NC score | 0.016485 (rank : 121) | θ value | 4.03905 (rank : 99) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NZN9, Q9H873, Q9NS10 | Gene names | AIPL1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl-hydrocarbon-interacting protein-like 1. | |||||
|
RP3A_HUMAN
|
||||||
| NC score | 0.016418 (rank : 122) | θ value | 1.38821 (rank : 53) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Y2J0, Q96AE0 | Gene names | RPH3A, KIAA0985 | |||
|
Domain Architecture |

|
|||||
| Description | Rabphilin-3A (Exophilin-1). | |||||
|
ITB4_HUMAN
|
||||||
| NC score | 0.016401 (rank : 123) | θ value | 2.36792 (rank : 71) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P16144, O14690, O14691, O15339, O15340, O15341, Q9UIQ4 | Gene names | ITGB4 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-4 precursor (GP150) (CD104 antigen). | |||||
|
AP3B2_MOUSE
|
||||||
| NC score | 0.016358 (rank : 124) | θ value | 8.99809 (rank : 155) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9JME5, Q6QR53, Q8R1E5 | Gene names | Ap3b2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain). | |||||
|
ETV5_MOUSE
|
||||||
| NC score | 0.015759 (rank : 125) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9CXC9, Q3TG49, Q8C0F3, Q9JHB1 | Gene names | Etv5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 5. | |||||
|
RNF24_HUMAN
|
||||||
| NC score | 0.015645 (rank : 126) | θ value | 8.99809 (rank : 171) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y225, Q9UMH1 | Gene names | RNF24 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 24. | |||||
|
YBOX2_HUMAN
|
||||||
| NC score | 0.014528 (rank : 127) | θ value | 5.27518 (rank : 134) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y2T7, Q8N4P0 | Gene names | YBX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Y-box-binding protein 2 (Germ cell-specific Y-box-binding protein) (Contrin) (MSY2 homolog). | |||||
|
SYNE1_HUMAN
|
||||||
| NC score | 0.014456 (rank : 128) | θ value | 5.27518 (rank : 132) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
SMC2_HUMAN
|
||||||
| NC score | 0.014286 (rank : 129) | θ value | 2.36792 (rank : 80) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O95347, Q9P1P2 | Gene names | SMC2L1, CAPE, SMC2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 2-like 1 protein (Chromosome- associated protein E) (hCAP-E) (XCAP-E homolog). | |||||
|
RFIP3_HUMAN
|
||||||
| NC score | 0.014121 (rank : 130) | θ value | 4.03905 (rank : 108) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1055 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O75154, Q4VXV7, Q9H155, Q9H1G0, Q9NUI0 | Gene names | RAB11FIP3, KIAA0665 | |||
|
Domain Architecture |

|
|||||
| Description | Rab11 family-interacting protein 3 (Rab11-FIP3) (EF hands-containing Rab-interacting protein) (Eferin). | |||||
|
LMBL3_MOUSE
|
||||||
| NC score | 0.013945 (rank : 131) | θ value | 6.88961 (rank : 143) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BLB7, Q641L7, Q6ZPI2, Q8C0G4 | Gene names | L3mbtl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lethal(3)malignant brain tumor-like 3 protein (L(3)mbt-like 3 protein). | |||||
|
MYB_MOUSE
|
||||||
| NC score | 0.013867 (rank : 132) | θ value | 4.03905 (rank : 107) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P06876, Q61929 | Gene names | Myb | |||
|
Domain Architecture |

|
|||||
| Description | Myb proto-oncogene protein (C-myb). | |||||
|
PJA1_HUMAN
|
||||||
| NC score | 0.013857 (rank : 133) | θ value | 5.27518 (rank : 126) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NG27, Q8NG28, Q9HAC1 | Gene names | PJA1, RNF70 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin protein ligase Praja1 (EC 6.3.2.-) (RING finger protein 70). | |||||
|
ZBT10_HUMAN
|
||||||
| NC score | 0.013550 (rank : 134) | θ value | 2.36792 (rank : 86) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 831 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96DT7, Q86W96, Q8IXI9, Q96MH9 | Gene names | ZBTB10, RINZF, RINZFC | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 10 (Zinc finger protein RIN ZF). | |||||
|
COEA1_HUMAN
|
||||||
| NC score | 0.013462 (rank : 135) | θ value | 6.88961 (rank : 139) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q05707, O00260, O00261, O00262, Q05708, Q5XJ18, Q96C67 | Gene names | COL14A1, UND | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XIV) chain precursor (Undulin). | |||||
|
IASPP_HUMAN
|
||||||
| NC score | 0.013407 (rank : 136) | θ value | 8.99809 (rank : 165) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8WUF5, Q2PNZ9, Q5DU71, Q5I1X4, Q6P1R7, Q6PKF8, Q9Y290 | Gene names | PPP1R13L, IASPP, NKIP1, PPP1R13BL, RAI | |||
|
Domain Architecture |

|
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
TF3C2_MOUSE
|
||||||
| NC score | 0.012858 (rank : 137) | θ value | 6.88961 (rank : 150) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BL74, Q3U7Z9, Q80XQ7, Q8BJJ4, Q8BLP3, Q8K1J1, Q9CSS0, Q9CVE6, Q9DBK6 | Gene names | Gtf3c2, Kiaa0011 | |||
|
Domain Architecture |

|
|||||
| Description | General transcription factor 3C polypeptide 2 (Transcription factor IIIC-subunit beta) (TF3C-beta) (TFIIIC 110 kDa subunit) (TFIIIC110). | |||||
|
ZN451_HUMAN
|
||||||
| NC score | 0.012492 (rank : 138) | θ value | 6.88961 (rank : 152) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 627 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Y4E5, Q5VVF1, Q8N380, Q8TD15, Q9NQM1 | Gene names | ZNF451, COASTER, KIAA0576 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 451 (Coactivator for steroid receptors). | |||||
|
CBLB_HUMAN
|
||||||
| NC score | 0.012027 (rank : 139) | θ value | 5.27518 (rank : 113) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13191, Q13192, Q13193, Q3LIC0, Q8IVC5 | Gene names | CBLB, RNF56 | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase CBL-B (EC 6.3.2.-) (Signal transduction protein CBL-B) (SH3-binding protein CBL-B) (Casitas B-lineage lymphoma proto-oncogene b) (RING finger protein 56). | |||||
|
SN1L2_MOUSE
|
||||||
| NC score | 0.011852 (rank : 140) | θ value | 0.0563607 (rank : 26) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8CFH6 | Gene names | Snf1lk2, Sik2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SNF1-like kinase 2 (EC 2.7.11.1) (Salt-inducible kinase 2). | |||||
|
EPC2_MOUSE
|
||||||
| NC score | 0.011828 (rank : 141) | θ value | 6.88961 (rank : 141) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8C0I4, Q3URX9, Q80Y70, Q8BXC5 | Gene names | Epc2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Enhancer of polycomb homolog 2. | |||||
|
CHRD_MOUSE
|
||||||
| NC score | 0.011782 (rank : 142) | θ value | 8.99809 (rank : 158) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z0E2 | Gene names | Chrd | |||
|
Domain Architecture |

|
|||||
| Description | Chordin precursor. | |||||
|
LTBP2_MOUSE
|
||||||
| NC score | 0.011763 (rank : 143) | θ value | 2.36792 (rank : 74) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 617 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O08999, Q8C6W9 | Gene names | Ltbp2 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 2 precursor (LTBP-2). | |||||
|
NRK_MOUSE
|
||||||
| NC score | 0.011750 (rank : 144) | θ value | 0.365318 (rank : 35) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 899 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9R0G8, Q6NV55, Q8C9S9, Q9R0S4 | Gene names | Nrk, Nesk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nik-related protein kinase (EC 2.7.11.1) (Nck-interacting kinase-like embryo specific kinase) (NIK-like embryo-specific kinase) (NESK). | |||||
|
NIN_HUMAN
|
||||||
| NC score | 0.011644 (rank : 145) | θ value | 5.27518 (rank : 123) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1416 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8N4C6, Q6P0P6, Q9BWU6, Q9C012, Q9C013, Q9C014, Q9H5I6, Q9HAT7, Q9HBY5, Q9HCK7, Q9UH61 | Gene names | NIN, KIAA1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein (hNinein) (Glycogen synthase kinase 3 beta-interacting protein) (GSK3B-interacting protein). | |||||
|
MFA3L_HUMAN
|
||||||
| NC score | 0.011115 (rank : 146) | θ value | 4.03905 (rank : 101) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75121, Q6TNA8, Q9BVE1, Q9BXK0 | Gene names | MFAP3L, KIAA0626 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microfibrillar-associated protein 3-like precursor (Protein kinase NYD-SP9). | |||||
|
SMCA1_MOUSE
|
||||||
| NC score | 0.010700 (rank : 147) | θ value | 4.03905 (rank : 110) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PGB8, Q8BSS1, Q91Y58 | Gene names | Smarca1, Snf2l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable global transcription activator SNF2L1 (EC 3.6.1.-) (Nucleosome remodeling factor subunit SNF2L) (ATP-dependent helicase SMARCA1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 1) (DNA-dependent ATPase SNF2L). | |||||
|
SMCA1_HUMAN
|
||||||
| NC score | 0.010692 (rank : 148) | θ value | 5.27518 (rank : 129) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P28370, Q5JV41, Q5JV42 | Gene names | SMARCA1, SNF2L, SNF2L1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L1 (EC 3.6.1.-) (Nucleosome remodeling factor subunit SNF2L) (ATP-dependent helicase SMARCA1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 1). | |||||
|
KLF13_HUMAN
|
||||||
| NC score | 0.010537 (rank : 149) | θ value | 2.36792 (rank : 72) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 739 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y2Y9, Q9Y356 | Gene names | KLF13, BTEB3, NSLP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 13 (Transcription factor BTEB3) (Basic transcription element-binding protein 3) (BTE-binding protein 3) (RANTES factor of late activated T-lymphocytes 1) (RFLAT-1) (Transcription factor NSLP1) (Novel Sp1-like zinc finger transcription factor 1). | |||||
|
DIAP1_MOUSE
|
||||||
| NC score | 0.010317 (rank : 150) | θ value | 6.88961 (rank : 140) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 839 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O08808 | Gene names | Diaph1, Diap1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1) (mDIA1) (p140mDIA). | |||||
|
AN30A_HUMAN
|
||||||
| NC score | 0.009990 (rank : 151) | θ value | 3.0926 (rank : 88) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1119 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9BXX3, Q5W025 | Gene names | ANKRD30A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 30A (NY-BR-1 antigen). | |||||
|
SPTA2_HUMAN
|
||||||
| NC score | 0.009955 (rank : 152) | θ value | 5.27518 (rank : 131) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 888 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q13813, Q13186, Q15324, Q16606, Q59EF1, Q5VXV5, Q5VXV6, Q7Z6M5, Q9P0V0 | Gene names | SPTAN1, SPTA2 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin alpha chain, brain (Spectrin, non-erythroid alpha chain) (Alpha-II spectrin) (Fodrin alpha chain). | |||||
|
RIPK3_MOUSE
|
||||||
| NC score | 0.009868 (rank : 153) | θ value | 0.365318 (rank : 37) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 867 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9QZL0 | Gene names | Ripk3, Rip3 | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-interacting serine/threonine-protein kinase 3 (EC 2.7.11.1) (RIP-like protein kinase 3) (Receptor-interacting protein 3) (RIP-3) (mRIP3). | |||||
|
TAOK2_MOUSE
|
||||||
| NC score | 0.009574 (rank : 154) | θ value | 1.38821 (rank : 55) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1373 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q6ZQ29, Q7TSS8 | Gene names | Taok2, Kiaa0881 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO2 (EC 2.7.11.1) (Thousand and one amino acid protein 2). | |||||
|
TAOK3_MOUSE
|
||||||
| NC score | 0.009303 (rank : 155) | θ value | 6.88961 (rank : 149) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1458 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8BYC6, Q3V3B3 | Gene names | Taok3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO3 (EC 2.7.11.1) (Thousand and one amino acid protein 3). | |||||
|
LRSM1_HUMAN
|
||||||
| NC score | 0.008974 (rank : 156) | θ value | 8.99809 (rank : 167) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1078 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q6UWE0, Q5VVV0, Q8NB40, Q96GT5, Q96MX5, Q96MZ7 | Gene names | LRSAM1, TAL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein LRSAM1 (EC 6.3.2.-) (Leucine-rich repeat and sterile alpha motif-containing protein 1) (Tsg101-associated ligase) (hTAL). | |||||
|
TAOK2_HUMAN
|
||||||
| NC score | 0.008688 (rank : 157) | θ value | 1.38821 (rank : 54) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1172 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UL54, O94957, Q6UW73, Q7LC09, Q9NSW2 | Gene names | TAOK2, KIAA0881, MAP3K17, PSK, PSK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO2 (EC 2.7.11.1) (Thousand and one amino acid protein 2) (Prostate-derived STE20-like kinase 1) (PSK-1) (Kinase from chicken homolog C) (hKFC-C). | |||||
|
MOES_MOUSE
|
||||||
| NC score | 0.008543 (rank : 158) | θ value | 4.03905 (rank : 103) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 923 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P26041, Q3UL28, Q8BSN4 | Gene names | Msn | |||
|
Domain Architecture |

|
|||||
| Description | Moesin (Membrane-organizing extension spike protein). | |||||
|
ZNF43_HUMAN
|
||||||
| NC score | 0.007819 (rank : 159) | θ value | 0.21417 (rank : 33) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P17038, P28160, Q96DG1 | Gene names | ZNF43, KOX27, ZNF39 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 43 (Zinc protein HTF6) (Zinc finger protein KOX27). | |||||
|
FOXGB_MOUSE
|
||||||
| NC score | 0.007673 (rank : 160) | θ value | 8.99809 (rank : 162) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q60987, Q80VP3 | Gene names | Foxg1b, Fkhl1, Foxg1, Hfhbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein G1B (Forkhead-related protein FKHL1) (Transcription factor BF-1) (Brain factor 1) (BF1). | |||||
|
STAC3_MOUSE
|
||||||
| NC score | 0.007635 (rank : 161) | θ value | 8.99809 (rank : 174) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BZ71 | Gene names | Stac3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and cysteine-rich domain-containing protein 3. | |||||
|
AATK_MOUSE
|
||||||
| NC score | 0.007398 (rank : 162) | θ value | 8.99809 (rank : 153) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 942 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q80YE4, O35211, Q3U2U5, Q3UHR8, Q66JN3, Q80YE3, Q8CB63, Q8CHE2 | Gene names | Aatk, Kiaa0641 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Apoptosis-associated tyrosine-protein kinase (EC 2.7.10.2) (AATYK) (Brain apoptosis-associated tyrosine kinase). | |||||
|
WNK3_HUMAN
|
||||||
| NC score | 0.007248 (rank : 163) | θ value | 3.0926 (rank : 97) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9BYP7, Q9HCK6 | Gene names | WNK3, KIAA1566, PRKWNK3 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK3 (EC 2.7.11.1) (Protein kinase with no lysine 3) (Protein kinase, lysine-deficient 3). | |||||
|
ZBTB2_HUMAN
|
||||||
| NC score | 0.006061 (rank : 164) | θ value | 6.88961 (rank : 151) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 752 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8N680, Q5SZ81, Q9P245 | Gene names | ZBTB2, KIAA1483, ZNF437 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 2. | |||||
|
UFO_HUMAN
|
||||||
| NC score | 0.006061 (rank : 165) | θ value | 0.813845 (rank : 45) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 981 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P30530 | Gene names | AXL, UFO | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase receptor UFO precursor (EC 2.7.10.1) (AXL oncogene). | |||||
|
SLIT1_HUMAN
|
||||||
| NC score | 0.005909 (rank : 166) | θ value | 3.0926 (rank : 96) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 700 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75093, Q8WWZ2, Q9UIL7 | Gene names | SLIT1, KIAA0813, MEGF4, SLIL1 | |||
|
Domain Architecture |

|
|||||
| Description | Slit homolog 1 protein precursor (Slit-1) (Multiple epidermal growth factor-like domains 4). | |||||
|
EGR1_MOUSE
|
||||||
| NC score | 0.005693 (rank : 167) | θ value | 3.0926 (rank : 90) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 816 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P08046, Q61777 | Gene names | Egr1, Egr-1, Krox-24 | |||
|
Domain Architecture |

|
|||||
| Description | Early growth response protein 1 (EGR-1) (Krox-24 protein) (Transcription factor Zif268) (Nerve growth factor-induced protein A) (NGFI-A). | |||||
|
ZN539_HUMAN
|
||||||
| NC score | 0.005493 (rank : 168) | θ value | 2.36792 (rank : 87) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 758 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q86XL7 | Gene names | ZNF539 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 539. | |||||
|
SOX8_HUMAN
|
||||||
| NC score | 0.005163 (rank : 169) | θ value | 8.99809 (rank : 173) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P57073, Q9NZW2 | Gene names | SOX8 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-8. | |||||
|
TRIP6_HUMAN
|
||||||
| NC score | 0.005162 (rank : 170) | θ value | 8.99809 (rank : 175) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q15654, O15170, O15275, Q9BTB2, Q9UNT4 | Gene names | TRIP6, OIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 6 (TRIP6) (OPA-interacting protein 1) (Zyxin-related protein 1) (ZRP-1). | |||||
|
ZN206_HUMAN
|
||||||
| NC score | 0.005119 (rank : 171) | θ value | 1.38821 (rank : 56) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 853 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96SZ4 | Gene names | ZNF206, ZSCAN10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 206 (Zinc finger and SCAN domain-containing protein 10). | |||||
|
ZN544_HUMAN
|
||||||
| NC score | 0.004922 (rank : 172) | θ value | 3.0926 (rank : 98) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6NX49, Q9UEX4 | Gene names | ZNF544 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 544. | |||||
|
ACK1_MOUSE
|
||||||
| NC score | 0.004814 (rank : 173) | θ value | 8.99809 (rank : 154) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 1080 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O54967, Q8C2U0, Q8K0K4 | Gene names | Tnk2, Ack1 | |||
|
Domain Architecture |

|
|||||
| Description | Activated CDC42 kinase 1 (EC 2.7.10.2) (ACK-1) (Non-receptor protein tyrosine kinase Ack) (Tyrosine kinase non-receptor protein 2). | |||||
|
ZN547_HUMAN
|
||||||
| NC score | 0.004581 (rank : 174) | θ value | 4.03905 (rank : 112) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 728 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8IVP9, Q96NC4 | Gene names | ZNF547 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 547. | |||||
|
HKR1_HUMAN
|
||||||
| NC score | 0.004265 (rank : 175) | θ value | 5.27518 (rank : 116) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 759 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P10072, Q9BSW9, Q9UM09 | Gene names | HKR1 | |||
|
Domain Architecture |

|
|||||
| Description | Krueppel-related zinc finger protein 1 (Protein HKR1). | |||||
|
MTF1_HUMAN
|
||||||
| NC score | 0.003958 (rank : 176) | θ value | 8.99809 (rank : 168) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 814 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q14872, Q96CB1 | Gene names | MTF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-regulatory transcription factor 1 (Transcription factor MTF-1) (MRE-binding transcription factor). | |||||
|
KRHB4_HUMAN
|
||||||
| NC score | 0.001270 (rank : 177) | θ value | 6.88961 (rank : 142) | |||
| Query Neighborhood Hits | 177 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NSB2 | Gene names | KRTHB4 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type II cuticular Hb4 (Hair keratin, type II Hb4). | |||||