Please be patient as the page loads
|
TOP2B_HUMAN
|
||||||
| SwissProt Accessions | Q02880, Q13600, Q9UMG8, Q9UQP8 | Gene names | TOP2B | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-beta (EC 5.99.1.3) (DNA topoisomerase II, beta isozyme). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
TOP2A_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.970087 (rank : 4) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P11388, Q71UN1, Q9HB24, Q9HB25, Q9HB26, Q9UP44, Q9UQP9 | Gene names | TOP2A, TOP2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-alpha (EC 5.99.1.3) (DNA topoisomerase II, alpha isozyme). | |||||
|
TOP2A_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.972273 (rank : 3) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q01320 | Gene names | Top2a, Top-2, Top2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-alpha (EC 5.99.1.3) (DNA topoisomerase II, alpha isozyme). | |||||
|
TOP2B_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 101 | |
| SwissProt Accessions | Q02880, Q13600, Q9UMG8, Q9UQP8 | Gene names | TOP2B | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-beta (EC 5.99.1.3) (DNA topoisomerase II, beta isozyme). | |||||
|
TOP2B_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.988656 (rank : 2) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q64511, Q7TQG4 | Gene names | Top2b | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-beta (EC 5.99.1.3) (DNA topoisomerase II, beta isozyme). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 5) | NC score | 0.107928 (rank : 5) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 0.000602161 (rank : 6) | NC score | 0.098769 (rank : 6) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
AKA12_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 7) | NC score | 0.074969 (rank : 7) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
ADDB_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 8) | NC score | 0.065832 (rank : 8) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 574 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QYB8, Q3U0E1, Q80VH9, Q8C1C4, Q9CXE3, Q9JLE5, Q9QYB7 | Gene names | Add2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adducin (Erythrocyte adducin subunit beta) (Add97). | |||||
|
PHF2_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 9) | NC score | 0.032821 (rank : 38) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
NPAS4_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 10) | NC score | 0.045839 (rank : 24) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
MPP8_HUMAN
|
||||||
| θ value | 0.125558 (rank : 11) | NC score | 0.031078 (rank : 41) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q99549, Q86TK3, Q9BTP1 | Gene names | MPHOSPH8, MPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | M-phase phosphoprotein 8. | |||||
|
FA8_HUMAN
|
||||||
| θ value | 0.163984 (rank : 12) | NC score | 0.020983 (rank : 62) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P00451 | Gene names | F8, F8C | |||
|
Domain Architecture |

|
|||||
| Description | Coagulation factor VIII precursor (Procoagulant component) (Antihemophilic factor) (AHF) [Contains: Factor VIIIa heavy chain, 200 kDa isoform; Factor VIIIa heavy chain, 92 kDa isoform; Factor VIII B chain; Factor VIIIa light chain]. | |||||
|
RRBP1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 13) | NC score | 0.042716 (rank : 26) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1306 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9P2E9, O75300, O75301, Q5W165, Q96SB2, Q9BWP1, Q9H476 | Gene names | RRBP1, KIAA1398 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (180 kDa ribosome receptor homolog) (ES/130-related protein). | |||||
|
F10A1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 14) | NC score | 0.025129 (rank : 50) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P50502, O14999 | Gene names | ST13, FAM10A1, HIP, SNC6 | |||
|
Domain Architecture |
|
|||||
| Description | Hsc70-interacting protein (Hip) (Suppression of tumorigenicity protein 13) (Putative tumor suppressor ST13) (Protein FAM10A1) (Progesterone receptor-associated p48 protein) (NY-REN-33 antigen). | |||||
|
AP3B2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 15) | NC score | 0.039194 (rank : 27) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q13367, O14808 | Gene names | AP3B2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (AP-3 complex beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain) (Neuron-specific vesicle coat protein beta-NAP). | |||||
|
HTSF1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 16) | NC score | 0.047483 (rank : 19) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BGC0, Q1WWK0, Q9CT41, Q9DAU3 | Gene names | Htatsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 homolog. | |||||
|
MCM5_HUMAN
|
||||||
| θ value | 0.62314 (rank : 17) | NC score | 0.016747 (rank : 70) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P33992, O60785, Q14578, Q9BTJ4, Q9BWL8 | Gene names | MCM5, CDC46 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM5 (CDC46 homolog) (P1-CDC46). | |||||
|
NPAS4_MOUSE
|
||||||
| θ value | 0.62314 (rank : 18) | NC score | 0.038837 (rank : 28) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
PCNP_HUMAN
|
||||||
| θ value | 0.62314 (rank : 19) | NC score | 0.054260 (rank : 16) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WW12, Q6AI44, Q96CU3, Q9NS81 | Gene names | PCNP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PEST proteolytic signal-containing nuclear protein (PEST-containing nuclear protein) (PCNP). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 0.62314 (rank : 20) | NC score | 0.023235 (rank : 56) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
TAOK2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 21) | NC score | 0.011861 (rank : 82) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1373 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6ZQ29, Q7TSS8 | Gene names | Taok2, Kiaa0881 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO2 (EC 2.7.11.1) (Thousand and one amino acid protein 2). | |||||
|
FA21C_HUMAN
|
||||||
| θ value | 0.813845 (rank : 22) | NC score | 0.046740 (rank : 20) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Y4E1, Q5SQU4, Q5SQU5, Q7L521, Q9UG79 | Gene names | FAM21C, KIAA0592 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM21C. | |||||
|
CF206_HUMAN
|
||||||
| θ value | 1.06291 (rank : 23) | NC score | 0.052844 (rank : 18) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H1X1 | Gene names | C6orf206, MRPS18AL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf206. | |||||
|
MEPE_HUMAN
|
||||||
| θ value | 1.06291 (rank : 24) | NC score | 0.036680 (rank : 30) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NQ76 | Gene names | MEPE | |||
|
Domain Architecture |
|
|||||
| Description | Matrix extracellular phosphoglycoprotein precursor (Osteoblast/osteocyte factor 45) (OF45). | |||||
|
TERF2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 25) | NC score | 0.060612 (rank : 9) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q15554 | Gene names | TERF2, TRF2 | |||
|
Domain Architecture |
|
|||||
| Description | Telomeric repeat-binding factor 2 (TTAGGG repeat-binding factor 2) (Telomeric DNA-binding protein). | |||||
|
ZN638_MOUSE
|
||||||
| θ value | 1.06291 (rank : 26) | NC score | 0.031959 (rank : 39) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q61464, Q6DFV9, Q8C941 | Gene names | Znf638, Np220, Zfml | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein). | |||||
|
TFAM_HUMAN
|
||||||
| θ value | 1.38821 (rank : 27) | NC score | 0.036274 (rank : 31) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q00059 | Gene names | TFAM, TCF6L2 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor A, mitochondrial precursor (mtTFA) (Mitochondrial transcription factor 1) (MtTF1) (Transcription factor 6-like 2). | |||||
|
TXND2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 28) | NC score | 0.053676 (rank : 17) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
AF9_HUMAN
|
||||||
| θ value | 1.81305 (rank : 29) | NC score | 0.037254 (rank : 29) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P42568 | Gene names | MLLT3, AF9, YEATS3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein AF-9 (ALL1 fused gene from chromosome 9 protein) (Myeloid/lymphoid or mixed-lineage leukemia translocated to chromosome 3 protein) (YEATS domain-containing protein 3). | |||||
|
AHI1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 30) | NC score | 0.010917 (rank : 84) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 537 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N157, Q504T3, Q5TCP9, Q6P098, Q6PIT6, Q8NDX0, Q9H0H2 | Gene names | AHI1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Jouberin (Abelson helper integration site 1 protein homolog) (AHI-1). | |||||
|
CRBB3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 31) | NC score | 0.017390 (rank : 69) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JJU9 | Gene names | Crybb3 | |||
|
Domain Architecture |
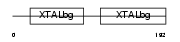
|
|||||
| Description | Beta crystallin B3 (Beta-B3-crystallin). | |||||
|
DSPP_MOUSE
|
||||||
| θ value | 1.81305 (rank : 32) | NC score | 0.054523 (rank : 15) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
PCNP_MOUSE
|
||||||
| θ value | 1.81305 (rank : 33) | NC score | 0.046574 (rank : 21) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6P8I4 | Gene names | Pcnp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PEST proteolytic signal-containing nuclear protein (PEST-containing nuclear protein) (PCNP). | |||||
|
TUFT1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 34) | NC score | 0.017865 (rank : 68) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 686 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NNX1, Q5T384, Q5T385, Q9BU28, Q9H5L1 | Gene names | TUFT1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tuftelin. | |||||
|
CJ012_HUMAN
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.033449 (rank : 35) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
H10_HUMAN
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.026765 (rank : 47) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P07305 | Gene names | H1F0, H1FV | |||
|
Domain Architecture |
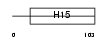
|
|||||
| Description | Histone H1.0 (H1(0)) (Histone H1'). | |||||
|
LATS1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 37) | NC score | 0.002604 (rank : 102) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1027 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BYR2, Q9Z0W4 | Gene names | Lats1, Warts | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase LATS1 (EC 2.7.11.1) (Large tumor suppressor homolog 1) (WARTS protein kinase). | |||||
|
NOLC1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 38) | NC score | 0.058012 (rank : 12) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
VPS72_HUMAN
|
||||||
| θ value | 2.36792 (rank : 39) | NC score | 0.030245 (rank : 43) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q15906, Q53GJ2, Q5U0R4 | Gene names | VPS72, TCFL1, YL1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
AFF2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 40) | NC score | 0.024727 (rank : 52) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P51816, O43786, O60215, P78407, Q13521, Q14323, Q9UNA5 | Gene names | AFF2, FMR2, OX19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation 2 protein) (Protein FMR-2) (FMR2P) (Protein Ox19) (Fragile X E mental retardation syndrome protein). | |||||
|
CCND1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 41) | NC score | 0.025690 (rank : 48) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P25322 | Gene names | Ccnd1, Cyl-1 | |||
|
Domain Architecture |
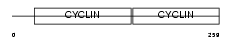
|
|||||
| Description | G1/S-specific cyclin-D1. | |||||
|
CDA7L_MOUSE
|
||||||
| θ value | 3.0926 (rank : 42) | NC score | 0.023333 (rank : 54) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q922M5 | Gene names | Cdca7l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle-associated 7-like protein (Transcription factor RAM2). | |||||
|
DSPP_HUMAN
|
||||||
| θ value | 3.0926 (rank : 43) | NC score | 0.045920 (rank : 23) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
HXD10_HUMAN
|
||||||
| θ value | 3.0926 (rank : 44) | NC score | 0.014723 (rank : 74) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P28358 | Gene names | HOXD10, HOX4D | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D10 (Hox-4D) (Hox-4E). | |||||
|
MBB1A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 45) | NC score | 0.046558 (rank : 22) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
PARG_HUMAN
|
||||||
| θ value | 3.0926 (rank : 46) | NC score | 0.034730 (rank : 33) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q86W56, Q6E4P6, Q6E4P7, Q7Z742, Q9Y4W7 | Gene names | PARG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(ADP-ribose) glycohydrolase (EC 3.2.1.143). | |||||
|
RECQ5_HUMAN
|
||||||
| θ value | 3.0926 (rank : 47) | NC score | 0.011606 (rank : 83) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O94762, Q9P1W7, Q9UNC8 | Gene names | RECQL5, RECQ5 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase Q5 (EC 3.6.1.-) (RecQ protein-like 5) (RecQ5). | |||||
|
RRBP1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 48) | NC score | 0.021521 (rank : 61) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
SAMN1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 49) | NC score | 0.014726 (rank : 73) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P57725, Q8C6S2 | Gene names | Samsn1 | |||
|
Domain Architecture |

|
|||||
| Description | SAM domain-containing protein SAMSN-1 (SAM domain, SH3 domain and nuclear localization signals protein 1). | |||||
|
SDPR_MOUSE
|
||||||
| θ value | 3.0926 (rank : 50) | NC score | 0.022083 (rank : 60) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q63918, Q3V1P6, Q78EC3, Q8CBT4 | Gene names | Sdpr, Sdr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serum deprivation-response protein (Phosphatidylserine-binding protein). | |||||
|
SET1A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 51) | NC score | 0.028257 (rank : 46) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
AHTF1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 52) | NC score | 0.033291 (rank : 36) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8WYP5, Q7Z4E3, Q8IZA4, Q96EH9, Q9Y4Q6 | Gene names | AHCTF1, ELYS, TMBS62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
CCND1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 53) | NC score | 0.020510 (rank : 63) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P24385, Q6LEF0 | Gene names | CCND1, BCL1, PRAD1 | |||
|
Domain Architecture |
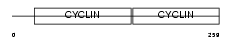
|
|||||
| Description | G1/S-specific cyclin-D1 (PRAD1 oncogene) (BCL-1 oncogene). | |||||
|
CF152_HUMAN
|
||||||
| θ value | 4.03905 (rank : 54) | NC score | 0.009498 (rank : 90) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q86VQ0, Q9BWX7 | Gene names | C6orf152 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf152. | |||||
|
DDX10_HUMAN
|
||||||
| θ value | 4.03905 (rank : 55) | NC score | 0.006449 (rank : 97) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13206, Q5BJD8 | Gene names | DDX10 | |||
|
Domain Architecture |
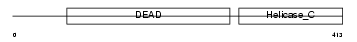
|
|||||
| Description | Probable ATP-dependent RNA helicase DDX10 (EC 3.6.1.-) (DEAD box protein 10). | |||||
|
HRX_HUMAN
|
||||||
| θ value | 4.03905 (rank : 56) | NC score | 0.023342 (rank : 53) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
HXD10_MOUSE
|
||||||
| θ value | 4.03905 (rank : 57) | NC score | 0.014022 (rank : 76) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P28359, Q91WU9 | Gene names | Hoxd10, Hox-4.5, Hoxd-10 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D10 (Hox-4.5) (Hox-5.3). | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 58) | NC score | 0.034098 (rank : 34) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
MARK3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 59) | NC score | 0.000921 (rank : 104) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q03141, Q3TM40, Q3V1U3, Q8C6G9, Q8R375, Q9JKE4 | Gene names | Mark3, Emk2, Mpk10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MAP/microtubule affinity-regulating kinase 3 (EC 2.7.11.1) (MPK-10) (ELKL motif kinase 2). | |||||
|
MYPT1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 60) | NC score | 0.007425 (rank : 95) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 723 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O14974, Q86WU3, Q8NFR6, Q9BYH0 | Gene names | PPP1R12A, MBS, MYPT1 | |||
|
Domain Architecture |
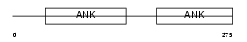
|
|||||
| Description | Protein phosphatase 1 regulatory subunit 12A (Myosin phosphatase- targeting subunit 1) (Myosin phosphatase target subunit 1) (Protein phosphatase myosin-binding subunit). | |||||
|
RTP3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 61) | NC score | 0.014178 (rank : 75) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5QGU6, Q8VDC2 | Gene names | Rtp3, Tmem7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor-transporting protein 3 (Transmembrane protein 7). | |||||
|
TAOK2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 62) | NC score | 0.005711 (rank : 99) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1172 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UL54, O94957, Q6UW73, Q7LC09, Q9NSW2 | Gene names | TAOK2, KIAA0881, MAP3K17, PSK, PSK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO2 (EC 2.7.11.1) (Thousand and one amino acid protein 2) (Prostate-derived STE20-like kinase 1) (PSK-1) (Kinase from chicken homolog C) (hKFC-C). | |||||
|
TR240_HUMAN
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.023283 (rank : 55) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UHV7, O60334 | Gene names | THRAP1, ARC250, KIAA0593, TRAP240 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid hormone receptor-associated protein complex 240 kDa component (Trap240) (Thyroid hormone receptor-associated protein 1) (Vitamin D3 receptor-interacting protein complex component DRIP250) (DRIP 250) (Activator-recruited cofactor 250 kDa component) (ARC250). | |||||
|
TTF1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.045605 (rank : 25) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q62187, Q9JKK5 | Gene names | Ttf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 1 (TTF-1) (TTF-I) (mTFF-I) (RNA polymerase I termination factor). | |||||
|
CLGN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.009952 (rank : 88) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O14967 | Gene names | CLGN | |||
|
Domain Architecture |
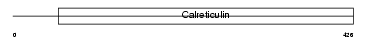
|
|||||
| Description | Calmegin precursor. | |||||
|
DEK_HUMAN
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.032988 (rank : 37) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P35659, Q5TGV4, Q5TGV5 | Gene names | DEK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.018161 (rank : 66) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
MYCD_HUMAN
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.013054 (rank : 79) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8IZQ8, Q8N7Q1 | Gene names | MYOCD, MYCD | |||
|
Domain Architecture |

|
|||||
| Description | Myocardin. | |||||
|
NP1L3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.025000 (rank : 51) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q794H2, O54802 | Gene names | Nap1l3, Mb20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome assembly protein 1-like 3 (Brain-specific protein MB20). | |||||
|
PEX5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 70) | NC score | 0.029681 (rank : 44) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P50542, Q15115, Q15266 | Gene names | PEX5, PXR1 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal targeting signal 1 receptor (Peroxisome receptor 1) (Peroxisomal C-terminal targeting signal import receptor) (PTS1-BP) (Peroxin-5) (PTS1 receptor). | |||||
|
SND1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 71) | NC score | 0.018903 (rank : 65) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7KZF4, Q13122 | Gene names | SND1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Staphylococcal nuclease domain-containing protein 1 (p100 co- activator) (100 kDa coactivator) (EBNA2 coactivator p100). | |||||
|
SND1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 72) | NC score | 0.018992 (rank : 64) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q78PY7, Q3TT46, Q922L5, Q9R0S1 | Gene names | Snd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Staphylococcal nuclease domain-containing protein 1 (p100 co- activator) (100 kDa coactivator). | |||||
|
ZBTB5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 73) | NC score | 0.000974 (rank : 103) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O15062 | Gene names | ZBTB5, KIAA0354 | |||
|
Domain Architecture |
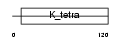
|
|||||
| Description | Zinc finger and BTB domain-containing protein 5. | |||||
|
ACINU_HUMAN
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.022153 (rank : 59) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
AFF4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.034743 (rank : 32) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
AFF4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.031391 (rank : 40) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
AP3B1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.025631 (rank : 49) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O00203, O00580, Q7Z393, Q9HD66 | Gene names | AP3B1, ADTB3A | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
FMN2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.013594 (rank : 78) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
HN1L_MOUSE
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.014955 (rank : 72) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PGH2 | Gene names | D17Ertd441e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hematological and neurological expressed 1-like protein (HN1-like protein). | |||||
|
IF4B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.022356 (rank : 57) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P23588 | Gene names | EIF4B | |||
|
Domain Architecture |
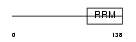
|
|||||
| Description | Eukaryotic translation initiation factor 4B (eIF-4B). | |||||
|
MBP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.015670 (rank : 71) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P02686, Q15337, Q15338, Q15339, Q15340, Q6AI64, Q6FH37, Q6FI04, Q6PK23 | Gene names | MBP | |||
|
Domain Architecture |
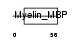
|
|||||
| Description | Myelin basic protein (MBP) (Myelin A1 protein) (Myelin membrane encephalitogenic protein). | |||||
|
MCES_HUMAN
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.012830 (rank : 80) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43148, O94996, Q9UIJ9 | Gene names | RNMT, KIAA0398 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA cap guanine-N7 methyltransferase (EC 2.1.1.56) (mRNA (guanine- N(7)-)-methyltransferase) (RG7MT1) (mRNA cap methyltransferase) (hcm1p) (hCMT1) (hMet). | |||||
|
NUAK1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.000256 (rank : 105) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60285, Q96KA8 | Gene names | NUAK1, ARK5, KIAA0537 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NUAK family SNF1-like kinase 1 (EC 2.7.11.1) (AMPK-related protein kinase 5). | |||||
|
RBBP5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.010381 (rank : 86) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q15291 | Gene names | RBBP5, RBQ3 | |||
|
Domain Architecture |
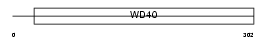
|
|||||
| Description | Retinoblastoma-binding protein 5 (RBBP-5) (Retinoblastoma-binding protein RBQ-3). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.006182 (rank : 98) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
WAPL_MOUSE
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.030312 (rank : 42) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q65Z40, Q6ZQE9 | Gene names | Wapal, Kiaa0261, Wapl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wings apart-like protein homolog (Dioxin-inducible factor 2) (DIF-2). | |||||
|
ABP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.010022 (rank : 87) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8JZQ5, Q8R229, Q8VC36 | Gene names | Abp1 | |||
|
Domain Architecture |

|
|||||
| Description | Amiloride-sensitive amine oxidase [copper-containing] precursor (EC 1.4.3.6) (Diamine oxidase) (DAO) (Amiloride-binding protein) (ABP) (Histaminase). | |||||
|
CT172_HUMAN
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.018004 (rank : 67) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H410, Q5JW56, Q9H8P4 | Gene names | C20orf172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf172. | |||||
|
DUS16_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.008026 (rank : 92) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BY84, Q9C0G3 | Gene names | DUSP16, KIAA1700, MKP7 | |||
|
Domain Architecture |
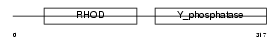
|
|||||
| Description | Dual specificity protein phosphatase 16 (EC 3.1.3.48) (EC 3.1.3.16) (Mitogen-activated protein kinase phosphatase 7) (MAP kinase phosphatase 7) (MKP-7). | |||||
|
ENAM_MOUSE
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.013633 (rank : 77) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O55196 | Gene names | Enam | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Enamelin precursor. | |||||
|
FMN2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.010695 (rank : 85) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9JL04 | Gene names | Fmn2 | |||
|
Domain Architecture |

|
|||||
| Description | Formin-2. | |||||
|
K1210_HUMAN
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.028645 (rank : 45) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ULL0, Q5JPN4 | Gene names | KIAA1210 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1210. | |||||
|
MAP9_MOUSE
|
||||||
| θ value | 8.99809 (rank : 93) | NC score | 0.022318 (rank : 58) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 529 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q3TRR0, Q3UUD1, Q3UX85, Q3UXE7, Q5M8N8, Q6P8K1, Q8BMM4, Q8BYP7 | Gene names | Map9, Asap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 9 (Aster-associated protein). | |||||
|
MBD2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.007868 (rank : 93) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UBB5, O95242, Q9UIS8 | Gene names | MBD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 2 (Methyl-CpG-binding protein MBD2) (Demethylase) (DMTase). | |||||
|
MYST4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 95) | NC score | 0.012576 (rank : 81) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BRB7, Q7TNW5, Q8BG35, Q8C441, Q9JKX5 | Gene names | Myst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Querkopf protein). | |||||
|
PKNX1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | 0.008405 (rank : 91) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O70477 | Gene names | Pknox1 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein PKNOX1 (PBX/knotted homeobox 1). | |||||
|
RAPH1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 97) | NC score | 0.009498 (rank : 89) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
S29A1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 98) | NC score | 0.007582 (rank : 94) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99808, Q9UJY2 | Gene names | SLC29A1, ENT1 | |||
|
Domain Architecture |

|
|||||
| Description | Equilibrative nucleoside transporter 1 (Equilibrative nitrobenzylmercaptopurine riboside-sensitive nucleoside transporter) (Equilibrative NBMPR-sensitive nucleoside transporter) (Nucleoside transporter, es-type) (Solute carrier family 29 member 1). | |||||
|
SMC2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 99) | NC score | 0.005480 (rank : 100) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O95347, Q9P1P2 | Gene names | SMC2L1, CAPE, SMC2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 2-like 1 protein (Chromosome- associated protein E) (hCAP-E) (XCAP-E homolog). | |||||
|
SSDH_HUMAN
|
||||||
| θ value | 8.99809 (rank : 100) | NC score | 0.006781 (rank : 96) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P51649 | Gene names | ALDH5A1, SSADH | |||
|
Domain Architecture |

|
|||||
| Description | Succinate semialdehyde dehydrogenase, mitochondrial precursor (EC 1.2.1.24) (NAD(+)-dependent succinic semialdehyde dehydrogenase). | |||||
|
STAM2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 101) | NC score | 0.004638 (rank : 101) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O88811, Q8C8Y4 | Gene names | Stam2, Hbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal transducing adapter molecule 2 (STAM-2) (Hrs-binding protein). | |||||
|
R51A1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.058525 (rank : 11) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8C551, O55219, Q8BP36, Q99L94, Q9D0J0 | Gene names | Rad51ap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RAD51-associated protein 1 (RAD51-interacting protein) (RAB22). | |||||
|
RBBP6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.055312 (rank : 14) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
TGON2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.056948 (rank : 13) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
TXND2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.059262 (rank : 10) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
TOP2B_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 101 | |
| SwissProt Accessions | Q02880, Q13600, Q9UMG8, Q9UQP8 | Gene names | TOP2B | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-beta (EC 5.99.1.3) (DNA topoisomerase II, beta isozyme). | |||||
|
TOP2B_MOUSE
|
||||||
| NC score | 0.988656 (rank : 2) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q64511, Q7TQG4 | Gene names | Top2b | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-beta (EC 5.99.1.3) (DNA topoisomerase II, beta isozyme). | |||||
|
TOP2A_MOUSE
|
||||||
| NC score | 0.972273 (rank : 3) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q01320 | Gene names | Top2a, Top-2, Top2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-alpha (EC 5.99.1.3) (DNA topoisomerase II, alpha isozyme). | |||||
|
TOP2A_HUMAN
|
||||||
| NC score | 0.970087 (rank : 4) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P11388, Q71UN1, Q9HB24, Q9HB25, Q9HB26, Q9UP44, Q9UQP9 | Gene names | TOP2A, TOP2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-alpha (EC 5.99.1.3) (DNA topoisomerase II, alpha isozyme). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.107928 (rank : 5) | θ value | 0.000461057 (rank : 5) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.098769 (rank : 6) | θ value | 0.000602161 (rank : 6) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
AKA12_HUMAN
|
||||||
| NC score | 0.074969 (rank : 7) | θ value | 0.00390308 (rank : 7) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
ADDB_MOUSE
|
||||||
| NC score | 0.065832 (rank : 8) | θ value | 0.00665767 (rank : 8) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 574 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QYB8, Q3U0E1, Q80VH9, Q8C1C4, Q9CXE3, Q9JLE5, Q9QYB7 | Gene names | Add2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adducin (Erythrocyte adducin subunit beta) (Add97). | |||||
|
TERF2_HUMAN
|
||||||
| NC score | 0.060612 (rank : 9) | θ value | 1.06291 (rank : 25) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q15554 | Gene names | TERF2, TRF2 | |||
|
Domain Architecture |

|
|||||
| Description | Telomeric repeat-binding factor 2 (TTAGGG repeat-binding factor 2) (Telomeric DNA-binding protein). | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.059262 (rank : 10) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
R51A1_MOUSE
|
||||||
| NC score | 0.058525 (rank : 11) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8C551, O55219, Q8BP36, Q99L94, Q9D0J0 | Gene names | Rad51ap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RAD51-associated protein 1 (RAD51-interacting protein) (RAB22). | |||||
|
NOLC1_HUMAN
|
||||||
| NC score | 0.058012 (rank : 12) | θ value | 2.36792 (rank : 38) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
TGON2_HUMAN
|
||||||
| NC score | 0.056948 (rank : 13) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
RBBP6_HUMAN
|
||||||
| NC score | 0.055312 (rank : 14) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
DSPP_MOUSE
|
||||||
| NC score | 0.054523 (rank : 15) | θ value | 1.81305 (rank : 32) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
PCNP_HUMAN
|
||||||
| NC score | 0.054260 (rank : 16) | θ value | 0.62314 (rank : 19) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WW12, Q6AI44, Q96CU3, Q9NS81 | Gene names | PCNP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PEST proteolytic signal-containing nuclear protein (PEST-containing nuclear protein) (PCNP). | |||||
|
TXND2_HUMAN
|
||||||
| NC score | 0.053676 (rank : 17) | θ value | 1.38821 (rank : 28) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
CF206_HUMAN
|
||||||
| NC score | 0.052844 (rank : 18) | θ value | 1.06291 (rank : 23) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H1X1 | Gene names | C6orf206, MRPS18AL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf206. | |||||
|
HTSF1_MOUSE
|
||||||
| NC score | 0.047483 (rank : 19) | θ value | 0.62314 (rank : 16) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BGC0, Q1WWK0, Q9CT41, Q9DAU3 | Gene names | Htatsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 homolog. | |||||
|
FA21C_HUMAN
|
||||||
| NC score | 0.046740 (rank : 20) | θ value | 0.813845 (rank : 22) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Y4E1, Q5SQU4, Q5SQU5, Q7L521, Q9UG79 | Gene names | FAM21C, KIAA0592 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM21C. | |||||
|
PCNP_MOUSE
|
||||||
| NC score | 0.046574 (rank : 21) | θ value | 1.81305 (rank : 33) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6P8I4 | Gene names | Pcnp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PEST proteolytic signal-containing nuclear protein (PEST-containing nuclear protein) (PCNP). | |||||
|
MBB1A_HUMAN
|
||||||
| NC score | 0.046558 (rank : 22) | θ value | 3.0926 (rank : 45) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.045920 (rank : 23) | θ value | 3.0926 (rank : 43) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
NPAS4_HUMAN
|
||||||
| NC score | 0.045839 (rank : 24) | θ value | 0.0431538 (rank : 10) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
TTF1_MOUSE
|
||||||
| NC score | 0.045605 (rank : 25) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q62187, Q9JKK5 | Gene names | Ttf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 1 (TTF-1) (TTF-I) (mTFF-I) (RNA polymerase I termination factor). | |||||
|
RRBP1_HUMAN
|
||||||
| NC score | 0.042716 (rank : 26) | θ value | 0.365318 (rank : 13) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1306 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9P2E9, O75300, O75301, Q5W165, Q96SB2, Q9BWP1, Q9H476 | Gene names | RRBP1, KIAA1398 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (180 kDa ribosome receptor homolog) (ES/130-related protein). | |||||
|
AP3B2_HUMAN
|
||||||
| NC score | 0.039194 (rank : 27) | θ value | 0.62314 (rank : 15) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q13367, O14808 | Gene names | AP3B2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (AP-3 complex beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain) (Neuron-specific vesicle coat protein beta-NAP). | |||||
|
NPAS4_MOUSE
|
||||||
| NC score | 0.038837 (rank : 28) | θ value | 0.62314 (rank : 18) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
AF9_HUMAN
|
||||||
| NC score | 0.037254 (rank : 29) | θ value | 1.81305 (rank : 29) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P42568 | Gene names | MLLT3, AF9, YEATS3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein AF-9 (ALL1 fused gene from chromosome 9 protein) (Myeloid/lymphoid or mixed-lineage leukemia translocated to chromosome 3 protein) (YEATS domain-containing protein 3). | |||||
|
MEPE_HUMAN
|
||||||
| NC score | 0.036680 (rank : 30) | θ value | 1.06291 (rank : 24) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NQ76 | Gene names | MEPE | |||
|
Domain Architecture |

|
|||||
| Description | Matrix extracellular phosphoglycoprotein precursor (Osteoblast/osteocyte factor 45) (OF45). | |||||
|
TFAM_HUMAN
|
||||||
| NC score | 0.036274 (rank : 31) | θ value | 1.38821 (rank : 27) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q00059 | Gene names | TFAM, TCF6L2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor A, mitochondrial precursor (mtTFA) (Mitochondrial transcription factor 1) (MtTF1) (Transcription factor 6-like 2). | |||||
|
AFF4_HUMAN
|
||||||
| NC score | 0.034743 (rank : 32) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
PARG_HUMAN
|
||||||
| NC score | 0.034730 (rank : 33) | θ value | 3.0926 (rank : 46) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q86W56, Q6E4P6, Q6E4P7, Q7Z742, Q9Y4W7 | Gene names | PARG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(ADP-ribose) glycohydrolase (EC 3.2.1.143). | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.034098 (rank : 34) | θ value | 4.03905 (rank : 58) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
CJ012_HUMAN
|
||||||
| NC score | 0.033449 (rank : 35) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
AHTF1_HUMAN
|
||||||
| NC score | 0.033291 (rank : 36) | θ value | 4.03905 (rank : 52) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8WYP5, Q7Z4E3, Q8IZA4, Q96EH9, Q9Y4Q6 | Gene names | AHCTF1, ELYS, TMBS62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
DEK_HUMAN
|
||||||
| NC score | 0.032988 (rank : 37) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P35659, Q5TGV4, Q5TGV5 | Gene names | DEK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
PHF2_MOUSE
|
||||||
| NC score | 0.032821 (rank : 38) | θ value | 0.0193708 (rank : 9) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
ZN638_MOUSE
|
||||||
| NC score | 0.031959 (rank : 39) | θ value | 1.06291 (rank : 26) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q61464, Q6DFV9, Q8C941 | Gene names | Znf638, Np220, Zfml | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein). | |||||
|
AFF4_MOUSE
|
||||||
| NC score | 0.031391 (rank : 40) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
MPP8_HUMAN
|
||||||
| NC score | 0.031078 (rank : 41) | θ value | 0.125558 (rank : 11) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q99549, Q86TK3, Q9BTP1 | Gene names | MPHOSPH8, MPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | M-phase phosphoprotein 8. | |||||
|
WAPL_MOUSE
|
||||||
| NC score | 0.030312 (rank : 42) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q65Z40, Q6ZQE9 | Gene names | Wapal, Kiaa0261, Wapl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wings apart-like protein homolog (Dioxin-inducible factor 2) (DIF-2). | |||||
|
VPS72_HUMAN
|
||||||
| NC score | 0.030245 (rank : 43) | θ value | 2.36792 (rank : 39) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q15906, Q53GJ2, Q5U0R4 | Gene names | VPS72, TCFL1, YL1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
PEX5_HUMAN
|
||||||
| NC score | 0.029681 (rank : 44) | θ value | 5.27518 (rank : 70) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P50542, Q15115, Q15266 | Gene names | PEX5, PXR1 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal targeting signal 1 receptor (Peroxisome receptor 1) (Peroxisomal C-terminal targeting signal import receptor) (PTS1-BP) (Peroxin-5) (PTS1 receptor). | |||||
|
K1210_HUMAN
|
||||||
| NC score | 0.028645 (rank : 45) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ULL0, Q5JPN4 | Gene names | KIAA1210 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1210. | |||||
|
SET1A_HUMAN
|
||||||
| NC score | 0.028257 (rank : 46) | θ value | 3.0926 (rank : 51) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
H10_HUMAN
|
||||||
| NC score | 0.026765 (rank : 47) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P07305 | Gene names | H1F0, H1FV | |||
|
Domain Architecture |

|
|||||
| Description | Histone H1.0 (H1(0)) (Histone H1'). | |||||
|
CCND1_MOUSE
|
||||||
| NC score | 0.025690 (rank : 48) | θ value | 3.0926 (rank : 41) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P25322 | Gene names | Ccnd1, Cyl-1 | |||
|
Domain Architecture |

|
|||||
| Description | G1/S-specific cyclin-D1. | |||||
|
AP3B1_HUMAN
|
||||||
| NC score | 0.025631 (rank : 49) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O00203, O00580, Q7Z393, Q9HD66 | Gene names | AP3B1, ADTB3A | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
F10A1_HUMAN
|
||||||
| NC score | 0.025129 (rank : 50) | θ value | 0.47712 (rank : 14) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P50502, O14999 | Gene names | ST13, FAM10A1, HIP, SNC6 | |||
|
Domain Architecture |

|
|||||
| Description | Hsc70-interacting protein (Hip) (Suppression of tumorigenicity protein 13) (Putative tumor suppressor ST13) (Protein FAM10A1) (Progesterone receptor-associated p48 protein) (NY-REN-33 antigen). | |||||
|
NP1L3_MOUSE
|
||||||
| NC score | 0.025000 (rank : 51) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q794H2, O54802 | Gene names | Nap1l3, Mb20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome assembly protein 1-like 3 (Brain-specific protein MB20). | |||||
|
AFF2_HUMAN
|
||||||
| NC score | 0.024727 (rank : 52) | θ value | 3.0926 (rank : 40) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P51816, O43786, O60215, P78407, Q13521, Q14323, Q9UNA5 | Gene names | AFF2, FMR2, OX19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation 2 protein) (Protein FMR-2) (FMR2P) (Protein Ox19) (Fragile X E mental retardation syndrome protein). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.023342 (rank : 53) | θ value | 4.03905 (rank : 56) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
CDA7L_MOUSE
|
||||||
| NC score | 0.023333 (rank : 54) | θ value | 3.0926 (rank : 42) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q922M5 | Gene names | Cdca7l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle-associated 7-like protein (Transcription factor RAM2). | |||||
|
TR240_HUMAN
|
||||||
| NC score | 0.023283 (rank : 55) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UHV7, O60334 | Gene names | THRAP1, ARC250, KIAA0593, TRAP240 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid hormone receptor-associated protein complex 240 kDa component (Trap240) (Thyroid hormone receptor-associated protein 1) (Vitamin D3 receptor-interacting protein complex component DRIP250) (DRIP 250) (Activator-recruited cofactor 250 kDa component) (ARC250). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.023235 (rank : 56) | θ value | 0.62314 (rank : 20) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
IF4B_HUMAN
|
||||||
| NC score | 0.022356 (rank : 57) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P23588 | Gene names | EIF4B | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4B (eIF-4B). | |||||
|
MAP9_MOUSE
|
||||||
| NC score | 0.022318 (rank : 58) | θ value | 8.99809 (rank : 93) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 529 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q3TRR0, Q3UUD1, Q3UX85, Q3UXE7, Q5M8N8, Q6P8K1, Q8BMM4, Q8BYP7 | Gene names | Map9, Asap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 9 (Aster-associated protein). | |||||
|
ACINU_HUMAN
|
||||||
| NC score | 0.022153 (rank : 59) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
SDPR_MOUSE
|
||||||
| NC score | 0.022083 (rank : 60) | θ value | 3.0926 (rank : 50) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q63918, Q3V1P6, Q78EC3, Q8CBT4 | Gene names | Sdpr, Sdr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serum deprivation-response protein (Phosphatidylserine-binding protein). | |||||
|
RRBP1_MOUSE
|
||||||
| NC score | 0.021521 (rank : 61) | θ value | 3.0926 (rank : 48) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
FA8_HUMAN
|
||||||
| NC score | 0.020983 (rank : 62) | θ value | 0.163984 (rank : 12) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P00451 | Gene names | F8, F8C | |||
|
Domain Architecture |

|
|||||
| Description | Coagulation factor VIII precursor (Procoagulant component) (Antihemophilic factor) (AHF) [Contains: Factor VIIIa heavy chain, 200 kDa isoform; Factor VIIIa heavy chain, 92 kDa isoform; Factor VIII B chain; Factor VIIIa light chain]. | |||||
|
CCND1_HUMAN
|
||||||
| NC score | 0.020510 (rank : 63) | θ value | 4.03905 (rank : 53) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P24385, Q6LEF0 | Gene names | CCND1, BCL1, PRAD1 | |||
|
Domain Architecture |

|
|||||
| Description | G1/S-specific cyclin-D1 (PRAD1 oncogene) (BCL-1 oncogene). | |||||
|
SND1_MOUSE
|
||||||
| NC score | 0.018992 (rank : 64) | θ value | 5.27518 (rank : 72) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q78PY7, Q3TT46, Q922L5, Q9R0S1 | Gene names | Snd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Staphylococcal nuclease domain-containing protein 1 (p100 co- activator) (100 kDa coactivator). | |||||
|
SND1_HUMAN
|
||||||
| NC score | 0.018903 (rank : 65) | θ value | 5.27518 (rank : 71) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7KZF4, Q13122 | Gene names | SND1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Staphylococcal nuclease domain-containing protein 1 (p100 co- activator) (100 kDa coactivator) (EBNA2 coactivator p100). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.018161 (rank : 66) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
CT172_HUMAN
|
||||||
| NC score | 0.018004 (rank : 67) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H410, Q5JW56, Q9H8P4 | Gene names | C20orf172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf172. | |||||
|
TUFT1_HUMAN
|
||||||
| NC score | 0.017865 (rank : 68) | θ value | 1.81305 (rank : 34) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 686 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NNX1, Q5T384, Q5T385, Q9BU28, Q9H5L1 | Gene names | TUFT1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tuftelin. | |||||
|
CRBB3_MOUSE
|
||||||
| NC score | 0.017390 (rank : 69) | θ value | 1.81305 (rank : 31) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JJU9 | Gene names | Crybb3 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin B3 (Beta-B3-crystallin). | |||||
|
MCM5_HUMAN
|
||||||
| NC score | 0.016747 (rank : 70) | θ value | 0.62314 (rank : 17) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P33992, O60785, Q14578, Q9BTJ4, Q9BWL8 | Gene names | MCM5, CDC46 | |||
|
Domain Architecture |

|
|||||
| Description | DNA replication licensing factor MCM5 (CDC46 homolog) (P1-CDC46). | |||||
|
MBP_HUMAN
|
||||||
| NC score | 0.015670 (rank : 71) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P02686, Q15337, Q15338, Q15339, Q15340, Q6AI64, Q6FH37, Q6FI04, Q6PK23 | Gene names | MBP | |||
|
Domain Architecture |

|
|||||
| Description | Myelin basic protein (MBP) (Myelin A1 protein) (Myelin membrane encephalitogenic protein). | |||||
|
HN1L_MOUSE
|
||||||
| NC score | 0.014955 (rank : 72) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PGH2 | Gene names | D17Ertd441e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hematological and neurological expressed 1-like protein (HN1-like protein). | |||||
|
SAMN1_MOUSE
|
||||||
| NC score | 0.014726 (rank : 73) | θ value | 3.0926 (rank : 49) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P57725, Q8C6S2 | Gene names | Samsn1 | |||
|
Domain Architecture |

|
|||||
| Description | SAM domain-containing protein SAMSN-1 (SAM domain, SH3 domain and nuclear localization signals protein 1). | |||||
|
HXD10_HUMAN
|
||||||
| NC score | 0.014723 (rank : 74) | θ value | 3.0926 (rank : 44) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P28358 | Gene names | HOXD10, HOX4D | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D10 (Hox-4D) (Hox-4E). | |||||
|
RTP3_MOUSE
|
||||||
| NC score | 0.014178 (rank : 75) | θ value | 4.03905 (rank : 61) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5QGU6, Q8VDC2 | Gene names | Rtp3, Tmem7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor-transporting protein 3 (Transmembrane protein 7). | |||||
|
HXD10_MOUSE
|
||||||
| NC score | 0.014022 (rank : 76) | θ value | 4.03905 (rank : 57) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P28359, Q91WU9 | Gene names | Hoxd10, Hox-4.5, Hoxd-10 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D10 (Hox-4.5) (Hox-5.3). | |||||
|
ENAM_MOUSE
|
||||||
| NC score | 0.013633 (rank : 77) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O55196 | Gene names | Enam | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Enamelin precursor. | |||||
|
FMN2_HUMAN
|
||||||
| NC score | 0.013594 (rank : 78) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
MYCD_HUMAN
|
||||||
| NC score | 0.013054 (rank : 79) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8IZQ8, Q8N7Q1 | Gene names | MYOCD, MYCD | |||
|
Domain Architecture |

|
|||||
| Description | Myocardin. | |||||
|
MCES_HUMAN
|
||||||
| NC score | 0.012830 (rank : 80) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43148, O94996, Q9UIJ9 | Gene names | RNMT, KIAA0398 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA cap guanine-N7 methyltransferase (EC 2.1.1.56) (mRNA (guanine- N(7)-)-methyltransferase) (RG7MT1) (mRNA cap methyltransferase) (hcm1p) (hCMT1) (hMet). | |||||
|
MYST4_MOUSE
|
||||||
| NC score | 0.012576 (rank : 81) | θ value | 8.99809 (rank : 95) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BRB7, Q7TNW5, Q8BG35, Q8C441, Q9JKX5 | Gene names | Myst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Querkopf protein). | |||||
|
TAOK2_MOUSE
|
||||||
| NC score | 0.011861 (rank : 82) | θ value | 0.62314 (rank : 21) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1373 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6ZQ29, Q7TSS8 | Gene names | Taok2, Kiaa0881 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO2 (EC 2.7.11.1) (Thousand and one amino acid protein 2). | |||||
|
RECQ5_HUMAN
|
||||||
| NC score | 0.011606 (rank : 83) | θ value | 3.0926 (rank : 47) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O94762, Q9P1W7, Q9UNC8 | Gene names | RECQL5, RECQ5 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase Q5 (EC 3.6.1.-) (RecQ protein-like 5) (RecQ5). | |||||
|
AHI1_HUMAN
|
||||||
| NC score | 0.010917 (rank : 84) | θ value | 1.81305 (rank : 30) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 537 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N157, Q504T3, Q5TCP9, Q6P098, Q6PIT6, Q8NDX0, Q9H0H2 | Gene names | AHI1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Jouberin (Abelson helper integration site 1 protein homolog) (AHI-1). | |||||
|
FMN2_MOUSE
|
||||||
| NC score | 0.010695 (rank : 85) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9JL04 | Gene names | Fmn2 | |||
|
Domain Architecture |

|
|||||
| Description | Formin-2. | |||||
|
RBBP5_HUMAN
|
||||||
| NC score | 0.010381 (rank : 86) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q15291 | Gene names | RBBP5, RBQ3 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoblastoma-binding protein 5 (RBBP-5) (Retinoblastoma-binding protein RBQ-3). | |||||
|
ABP1_MOUSE
|
||||||
| NC score | 0.010022 (rank : 87) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8JZQ5, Q8R229, Q8VC36 | Gene names | Abp1 | |||
|
Domain Architecture |

|
|||||
| Description | Amiloride-sensitive amine oxidase [copper-containing] precursor (EC 1.4.3.6) (Diamine oxidase) (DAO) (Amiloride-binding protein) (ABP) (Histaminase). | |||||
|
CLGN_HUMAN
|
||||||
| NC score | 0.009952 (rank : 88) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O14967 | Gene names | CLGN | |||
|
Domain Architecture |

|
|||||
| Description | Calmegin precursor. | |||||
|
RAPH1_HUMAN
|
||||||
| NC score | 0.009498 (rank : 89) | θ value | 8.99809 (rank : 97) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
CF152_HUMAN
|
||||||
| NC score | 0.009498 (rank : 90) | θ value | 4.03905 (rank : 54) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q86VQ0, Q9BWX7 | Gene names | C6orf152 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf152. | |||||
|
PKNX1_MOUSE
|
||||||
| NC score | 0.008405 (rank : 91) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O70477 | Gene names | Pknox1 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein PKNOX1 (PBX/knotted homeobox 1). | |||||
|
DUS16_HUMAN
|
||||||
| NC score | 0.008026 (rank : 92) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BY84, Q9C0G3 | Gene names | DUSP16, KIAA1700, MKP7 | |||
|
Domain Architecture |

|
|||||
| Description | Dual specificity protein phosphatase 16 (EC 3.1.3.48) (EC 3.1.3.16) (Mitogen-activated protein kinase phosphatase 7) (MAP kinase phosphatase 7) (MKP-7). | |||||
|
MBD2_HUMAN
|
||||||
| NC score | 0.007868 (rank : 93) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UBB5, O95242, Q9UIS8 | Gene names | MBD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 2 (Methyl-CpG-binding protein MBD2) (Demethylase) (DMTase). | |||||
|
S29A1_HUMAN
|
||||||
| NC score | 0.007582 (rank : 94) | θ value | 8.99809 (rank : 98) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99808, Q9UJY2 | Gene names | SLC29A1, ENT1 | |||
|
Domain Architecture |

|
|||||
| Description | Equilibrative nucleoside transporter 1 (Equilibrative nitrobenzylmercaptopurine riboside-sensitive nucleoside transporter) (Equilibrative NBMPR-sensitive nucleoside transporter) (Nucleoside transporter, es-type) (Solute carrier family 29 member 1). | |||||
|
MYPT1_HUMAN
|
||||||
| NC score | 0.007425 (rank : 95) | θ value | 4.03905 (rank : 60) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 723 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O14974, Q86WU3, Q8NFR6, Q9BYH0 | Gene names | PPP1R12A, MBS, MYPT1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein phosphatase 1 regulatory subunit 12A (Myosin phosphatase- targeting subunit 1) (Myosin phosphatase target subunit 1) (Protein phosphatase myosin-binding subunit). | |||||
|
SSDH_HUMAN
|
||||||
| NC score | 0.006781 (rank : 96) | θ value | 8.99809 (rank : 100) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P51649 | Gene names | ALDH5A1, SSADH | |||
|
Domain Architecture |

|
|||||
| Description | Succinate semialdehyde dehydrogenase, mitochondrial precursor (EC 1.2.1.24) (NAD(+)-dependent succinic semialdehyde dehydrogenase). | |||||
|
DDX10_HUMAN
|
||||||
| NC score | 0.006449 (rank : 97) | θ value | 4.03905 (rank : 55) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13206, Q5BJD8 | Gene names | DDX10 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DDX10 (EC 3.6.1.-) (DEAD box protein 10). | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.006182 (rank : 98) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
TAOK2_HUMAN
|
||||||
| NC score | 0.005711 (rank : 99) | θ value | 4.03905 (rank : 62) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1172 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UL54, O94957, Q6UW73, Q7LC09, Q9NSW2 | Gene names | TAOK2, KIAA0881, MAP3K17, PSK, PSK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO2 (EC 2.7.11.1) (Thousand and one amino acid protein 2) (Prostate-derived STE20-like kinase 1) (PSK-1) (Kinase from chicken homolog C) (hKFC-C). | |||||
|
SMC2_HUMAN
|
||||||
| NC score | 0.005480 (rank : 100) | θ value | 8.99809 (rank : 99) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O95347, Q9P1P2 | Gene names | SMC2L1, CAPE, SMC2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 2-like 1 protein (Chromosome- associated protein E) (hCAP-E) (XCAP-E homolog). | |||||
|
STAM2_MOUSE
|
||||||
| NC score | 0.004638 (rank : 101) | θ value | 8.99809 (rank : 101) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O88811, Q8C8Y4 | Gene names | Stam2, Hbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal transducing adapter molecule 2 (STAM-2) (Hrs-binding protein). | |||||
|
LATS1_MOUSE
|
||||||
| NC score | 0.002604 (rank : 102) | θ value | 2.36792 (rank : 37) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 1027 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BYR2, Q9Z0W4 | Gene names | Lats1, Warts | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase LATS1 (EC 2.7.11.1) (Large tumor suppressor homolog 1) (WARTS protein kinase). | |||||
|
ZBTB5_HUMAN
|
||||||
| NC score | 0.000974 (rank : 103) | θ value | 5.27518 (rank : 73) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O15062 | Gene names | ZBTB5, KIAA0354 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 5. | |||||
|
MARK3_MOUSE
|
||||||
| NC score | 0.000921 (rank : 104) | θ value | 4.03905 (rank : 59) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q03141, Q3TM40, Q3V1U3, Q8C6G9, Q8R375, Q9JKE4 | Gene names | Mark3, Emk2, Mpk10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MAP/microtubule affinity-regulating kinase 3 (EC 2.7.11.1) (MPK-10) (ELKL motif kinase 2). | |||||
|
NUAK1_HUMAN
|
||||||
| NC score | 0.000256 (rank : 105) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 101 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60285, Q96KA8 | Gene names | NUAK1, ARK5, KIAA0537 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NUAK family SNF1-like kinase 1 (EC 2.7.11.1) (AMPK-related protein kinase 5). | |||||