Please be patient as the page loads
|
SEBP2_HUMAN
|
||||||
| SwissProt Accessions | Q96T21, Q8IYC0 | Gene names | SECISBP2, SBP2 | |||
|
Domain Architecture |

|
|||||
| Description | SECIS-binding protein 2 (Selenocysteine insertion sequence-binding protein 2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SEBP2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 88 | |
| SwissProt Accessions | Q96T21, Q8IYC0 | Gene names | SECISBP2, SBP2 | |||
|
Domain Architecture |

|
|||||
| Description | SECIS-binding protein 2 (Selenocysteine insertion sequence-binding protein 2). | |||||
|
K0256_MOUSE
|
||||||
| θ value | 3.82246e-75 (rank : 2) | NC score | 0.829823 (rank : 3) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6A098 | Gene names | Kiaa0256 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0256. | |||||
|
K0256_HUMAN
|
||||||
| θ value | 1.45253e-74 (rank : 3) | NC score | 0.839924 (rank : 2) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q93073, Q8N767 | Gene names | KIAA0256 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0256. | |||||
|
CLSPN_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 4) | NC score | 0.079270 (rank : 5) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q80YR7, Q69GM2 | Gene names | Clspn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Claspin. | |||||
|
RIF1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 5) | NC score | 0.062436 (rank : 9) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 320 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5UIP0, Q5H9R3, Q5UIP2, Q66YK6, Q6PRU2, Q8TE94, Q99772, Q9H830, Q9H9B9, Q9NVP5, Q9Y4R4 | Gene names | RIF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomere-associated protein RIF1 (Rap1-interacting factor 1 homolog). | |||||
|
CADH4_MOUSE
|
||||||
| θ value | 0.125558 (rank : 6) | NC score | 0.014167 (rank : 70) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P39038 | Gene names | Cdh4 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-4 precursor (Retinal-cadherin) (R-cadherin) (R-CAD). | |||||
|
MAGC3_HUMAN
|
||||||
| θ value | 0.21417 (rank : 7) | NC score | 0.035006 (rank : 29) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TD91, Q5JZ43, Q9BZ80 | Gene names | MAGEC3, HCA2 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen C3 (MAGE-C3 antigen) (Hepatocellular carcinoma-associated antigen 2). | |||||
|
T2FA_HUMAN
|
||||||
| θ value | 0.21417 (rank : 8) | NC score | 0.082770 (rank : 4) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P35269, Q9BWN0 | Gene names | GTF2F1, RAP74 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor IIF alpha subunit (EC 2.7.11.1) (TFIIF-alpha) (Transcription initiation factor RAP74) (General transcription factor IIF polypeptide 1 74 kDa subunit protein). | |||||
|
PIBF1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 9) | NC score | 0.020656 (rank : 54) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8WXW3, O95664, Q6U9V2, Q6UG50, Q86V07, Q96SF4 | Gene names | PIBF1, C13orf24, PIBF | |||
|
Domain Architecture |

|
|||||
| Description | Progesterone-induced-blocking factor 1. | |||||
|
PKCB1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 10) | NC score | 0.055461 (rank : 10) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
CBP_HUMAN
|
||||||
| θ value | 0.47712 (rank : 11) | NC score | 0.047617 (rank : 14) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
GOGB1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 12) | NC score | 0.021953 (rank : 50) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 0.62314 (rank : 13) | NC score | 0.025806 (rank : 39) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
PF21A_MOUSE
|
||||||
| θ value | 0.62314 (rank : 14) | NC score | 0.036852 (rank : 24) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6ZPK0, Q6XVG0, Q80Z33, Q8CAZ4, Q8VEC8 | Gene names | Phf21a, Bhc80, Kiaa1696, Pftf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (mBHC80) (BHC80a). | |||||
|
RPO3G_HUMAN
|
||||||
| θ value | 0.62314 (rank : 15) | NC score | 0.070012 (rank : 6) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15318 | Gene names | POLR3G, RPC32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA-directed RNA polymerase III subunit G (EC 2.7.7.6) (DNA-directed RNA polymerase III 32 kDa polypeptide) (RNA polymerase III C32 subunit). | |||||
|
CBP_MOUSE
|
||||||
| θ value | 0.813845 (rank : 16) | NC score | 0.045040 (rank : 15) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
PABP2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 17) | NC score | 0.048160 (rank : 13) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q86U42, O43484 | Gene names | PABPN1, PAB2, PABP2 | |||
|
Domain Architecture |

|
|||||
| Description | Polyadenylate-binding protein 2 (Poly(A)-binding protein 2) (Poly(A)- binding protein II) (PABII) (Polyadenylate-binding nuclear protein 1) (Nuclear poly(A)-binding protein 1). | |||||
|
PABP2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 18) | NC score | 0.048604 (rank : 12) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CCS6, O35935 | Gene names | Pabpn1, Pab2, Pabp2 | |||
|
Domain Architecture |

|
|||||
| Description | Polyadenylate-binding protein 2 (Poly(A)-binding protein 2) (Poly(A)- binding protein II) (PABII) (Polyadenylate-binding nuclear protein 1) (Nuclear poly(A)-binding protein 1). | |||||
|
NOP14_HUMAN
|
||||||
| θ value | 1.06291 (rank : 19) | NC score | 0.036720 (rank : 25) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P78316, Q7LGI5, Q7Z6K0, Q8TBR6 | Gene names | NOP14, C4orf9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
COA2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 20) | NC score | 0.029380 (rank : 36) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O00763, Q16852 | Gene names | ACACB, ACC2, ACCB | |||
|
Domain Architecture |

|
|||||
| Description | Acetyl-CoA carboxylase 2 (EC 6.4.1.2) (ACC-beta) [Includes: Biotin carboxylase (EC 6.3.4.14)]. | |||||
|
NOL1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 21) | NC score | 0.055230 (rank : 11) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P46087 | Gene names | NOL1 | |||
|
Domain Architecture |

|
|||||
| Description | Proliferating-cell nucleolar antigen p120 (Proliferation-associated nucleolar protein p120). | |||||
|
SPG20_MOUSE
|
||||||
| θ value | 1.38821 (rank : 22) | NC score | 0.039551 (rank : 20) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R1X6, Q6ZQ87, Q8BJD3, Q8BM37, Q8BZ63 | Gene names | Spg20, Kiaa0610 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spartin. | |||||
|
CDN1C_MOUSE
|
||||||
| θ value | 1.81305 (rank : 23) | NC score | 0.042355 (rank : 16) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P49919 | Gene names | Cdkn1c, Kip2 | |||
|
Domain Architecture |
|
|||||
| Description | Cyclin-dependent kinase inhibitor 1C (Cyclin-dependent kinase inhibitor p57) (p57KIP2). | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 24) | NC score | 0.036127 (rank : 26) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
WWC1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 25) | NC score | 0.027327 (rank : 38) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5SXA9, Q571D0, Q8K1Y3, Q8VD17, Q922W3 | Gene names | Wwc1, Kiaa0869 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing protein 1 (Kidney and brain protein) (KIBRA). | |||||
|
AN32E_HUMAN
|
||||||
| θ value | 2.36792 (rank : 26) | NC score | 0.020221 (rank : 56) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BTT0, Q8N1S4, Q8WWW9 | Gene names | ANP32E | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member E (LANP- like protein) (LANP-L). | |||||
|
IRX1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 27) | NC score | 0.019312 (rank : 61) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P78414, Q7Z2F8, Q8N312 | Gene names | IRX1, IRXA1 | |||
|
Domain Architecture |
|
|||||
| Description | Iroquois-class homeodomain protein IRX-1 (Iroquois homeobox protein 1) (Homeodomain protein IRXA1). | |||||
|
NFH_HUMAN
|
||||||
| θ value | 2.36792 (rank : 28) | NC score | 0.022632 (rank : 49) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.041987 (rank : 18) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.042320 (rank : 17) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 31) | NC score | 0.034449 (rank : 30) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
ZAN_MOUSE
|
||||||
| θ value | 2.36792 (rank : 32) | NC score | 0.014333 (rank : 69) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O88799, O08647 | Gene names | Zan | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
ZN622_HUMAN
|
||||||
| θ value | 2.36792 (rank : 33) | NC score | 0.035858 (rank : 27) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q969S3 | Gene names | ZNF622, ZPR9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 622 (Zinc finger-like protein 9). | |||||
|
ATRX_HUMAN
|
||||||
| θ value | 3.0926 (rank : 34) | NC score | 0.035200 (rank : 28) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P46100, P51068, Q15886, Q59FB5, Q59H31, Q5H9A2, Q5JWI4, Q7Z2J1, Q9H0Z1, Q9NTS3 | Gene names | ATRX, RAD54L, XH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked helicase II) (X-linked nuclear protein) (XNP) (Znf- HX). | |||||
|
CCD71_HUMAN
|
||||||
| θ value | 3.0926 (rank : 35) | NC score | 0.037844 (rank : 22) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IV32, Q6IPE2, Q9H8H4, Q9H9F1 | Gene names | CCDC71 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 71. | |||||
|
CDKL3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 36) | NC score | 0.001119 (rank : 87) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 857 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BLF2, Q5M6W2, Q8BKR2, Q8BL49, Q8BVE0, Q8K134 | Gene names | Cdkl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-dependent kinase-like 3 (EC 2.7.11.22). | |||||
|
GIT2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 37) | NC score | 0.016629 (rank : 66) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14161, Q86U59, Q96CI2, Q9BV91, Q9Y5V2 | Gene names | GIT2, KIAA0148 | |||
|
Domain Architecture |
|
|||||
| Description | ARF GTPase-activating protein GIT2 (G protein-coupled receptor kinase- interactor 2) (GRK-interacting protein 2) (Cool-interacting tyrosine- phosphorylated protein 2) (CAT2) (CAT-2). | |||||
|
JAD1C_HUMAN
|
||||||
| θ value | 3.0926 (rank : 38) | NC score | 0.016730 (rank : 65) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P41229 | Gene names | SMCX, DXS1272E, JARID1C, XE169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (Protein SmcX) (Protein Xe169). | |||||
|
KIF1A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 39) | NC score | 0.008630 (rank : 78) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q12756 | Gene names | KIF1A, ATSV | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF1A (Axonal transporter of synaptic vesicles). | |||||
|
PINX1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 40) | NC score | 0.034430 (rank : 31) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96BK5, Q7Z7J8, Q96QD7, Q9HBU7, Q9NWW2 | Gene names | PINX1, LPTS | |||
|
Domain Architecture |
|
|||||
| Description | Pin2-interacting protein X1 (TRF1-interacting protein 1) (Liver- related putative tumor suppressor) (67-11-3 protein). | |||||
|
SETX_HUMAN
|
||||||
| θ value | 3.0926 (rank : 41) | NC score | 0.029072 (rank : 37) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7Z333, O75120, Q3KQX4, Q68DW5, Q6AZD7, Q7Z3J6, Q8WX33, Q9H9D1, Q9NVP9 | Gene names | SETX, ALS4, KIAA0625, SCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable helicase senataxin (EC 3.6.1.-) (SEN1 homolog). | |||||
|
4ET_MOUSE
|
||||||
| θ value | 4.03905 (rank : 42) | NC score | 0.024504 (rank : 42) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9EST3, Q9CSS3 | Gene names | Eif4enif1, Clast4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4E transporter (eIF4E transporter) (4E-T) (Eukaryotic translation initiation factor 4E nuclear import factor 1). | |||||
|
AP3B1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 43) | NC score | 0.021363 (rank : 52) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O00203, O00580, Q7Z393, Q9HD66 | Gene names | AP3B1, ADTB3A | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
CF142_HUMAN
|
||||||
| θ value | 4.03905 (rank : 44) | NC score | 0.031416 (rank : 33) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5VWP3, Q96H08, Q96NF7 | Gene names | C6orf142 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf142. | |||||
|
DNJC1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 45) | NC score | 0.024570 (rank : 41) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96KC8 | Gene names | DNAJC1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 1. | |||||
|
DNJC1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.023562 (rank : 46) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61712 | Gene names | Dnajc1, Dnajl1, Mtj1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 1 (DnaJ protein homolog MTJ1). | |||||
|
MTF1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 47) | NC score | 0.001936 (rank : 86) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q07243 | Gene names | Mtf1, Mtf-1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-regulatory transcription factor 1 (Transcription factor MTF-1) (MRE-binding transcription factor). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 4.03905 (rank : 48) | NC score | 0.024335 (rank : 43) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
NHPX_HUMAN
|
||||||
| θ value | 4.03905 (rank : 49) | NC score | 0.063278 (rank : 7) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P55769 | Gene names | NHP2L1 | |||
|
Domain Architecture |

|
|||||
| Description | NHP2-like protein 1 (High mobility group-like nuclear protein 2 homolog 1) (U4/U6.U5 tri-snRNP 15.5 kDa protein) (OTK27) (hSNU13). | |||||
|
NHPX_MOUSE
|
||||||
| θ value | 4.03905 (rank : 50) | NC score | 0.063278 (rank : 8) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D0T1, O08754, Q3UY64 | Gene names | Nhp2l1, Ssfa1 | |||
|
Domain Architecture |

|
|||||
| Description | NHP2-like protein 1 (High mobility group-like nuclear protein 2 homolog 1) (U4/U6.U5 tri-snRNP 15.5 kDa protein) (Sperm-specific antigen 1) (Fertilization antigen 1) (FA-1). | |||||
|
NOLA2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 51) | NC score | 0.040940 (rank : 19) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CRB2 | Gene names | Nola2, Nhp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | H/ACA ribonucleoprotein complex subunit 2 (Nucleolar protein family A member 2) (snoRNP protein NHP2). | |||||
|
SHAN2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 52) | NC score | 0.019911 (rank : 59) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 346 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UPX8, Q3Y8G9, Q52LK2, Q9UKP1 | Gene names | SHANK2, KIAA1022 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 2 (Shank2). | |||||
|
TSH1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 53) | NC score | 0.017316 (rank : 64) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 675 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5DTH5, Q6PE60, Q8VD19, Q9JLD5 | Gene names | Tshz1, Kiaa4206, Sdccag33, Tsh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 1 (Serologically defined colon cancer antigen 3 homolog). | |||||
|
ARP5_MOUSE
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | 0.014724 (rank : 68) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80US4, Q8BL26 | Gene names | Actr5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-related protein 5. | |||||
|
BCOR_HUMAN
|
||||||
| θ value | 5.27518 (rank : 55) | NC score | 0.024026 (rank : 45) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6W2J9, Q6P4B6, Q7Z2K7, Q8TEB4, Q96DB3, Q9H232, Q9H233, Q9HCJ7, Q9NXF2 | Gene names | BCOR, KIAA1575 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BCoR protein (BCL-6 corepressor). | |||||
|
CIC_MOUSE
|
||||||
| θ value | 5.27518 (rank : 56) | NC score | 0.017550 (rank : 63) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 742 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q924A2, Q6PDJ8, Q8CGE4, Q8CHH0, Q9CW61 | Gene names | Cic, Kiaa0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
DNMT1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 57) | NC score | 0.031098 (rank : 34) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P26358, Q9UHG5, Q9ULA2, Q9UMZ6 | Gene names | DNMT1, AIM, DNMT | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase HsaI) (DNA MTase HsaI) (MCMT) (M.HsaI). | |||||
|
ESCO2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 58) | NC score | 0.019559 (rank : 60) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q56NI9 | Gene names | ESCO2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyltransferase ESCO2 (EC 2.3.1.-) (Establishment of cohesion 1 homolog 2) (ECO1 homolog 2). | |||||
|
GASP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 59) | NC score | 0.020876 (rank : 53) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5U4C1, Q8BKR8, Q8BUN4, Q8BYK9, Q8CHF4, Q8R095 | Gene names | Gprasp1, Kiaa0443 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor-associated sorting protein 1 (GASP-1). | |||||
|
LRP12_HUMAN
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.013848 (rank : 73) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y561 | Gene names | LRP12, ST7 | |||
|
Domain Architecture |
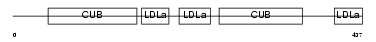
|
|||||
| Description | Low-density lipoprotein receptor-related protein 12 precursor (Suppressor of tumorigenicity protein 7). | |||||
|
LRP12_MOUSE
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.013875 (rank : 72) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BUJ9, Q8BWM9 | Gene names | Lrp12 | |||
|
Domain Architecture |
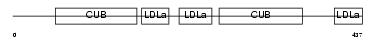
|
|||||
| Description | Low-density lipoprotein receptor-related protein 12 precursor. | |||||
|
PRP16_HUMAN
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.012039 (rank : 75) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92620, O75212, Q96HN7 | Gene names | DHX38, DDX38, KIAA0224, PRP16 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-splicing factor ATP-dependent RNA helicase PRP16 (EC 3.6.1.-) (ATP-dependent RNA helicase DHX38) (DEAH box protein 38). | |||||
|
PRP40_MOUSE
|
||||||
| θ value | 5.27518 (rank : 63) | NC score | 0.029874 (rank : 35) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
RBCC1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.012808 (rank : 74) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1023 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8TDY2, Q8WVU9, Q92601 | Gene names | RB1CC1, KIAA0203, RBICC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RB1-inducible coiled-coil protein 1. | |||||
|
RBM10_HUMAN
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.020602 (rank : 55) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P98175, Q14136 | Gene names | RBM10, DXS8237E, KIAA0122 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 10 (RNA-binding motif protein 10). | |||||
|
ZN227_HUMAN
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | -0.000149 (rank : 88) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 781 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q86WZ6 | Gene names | ZNF227 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 227. | |||||
|
BOC_HUMAN
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.005771 (rank : 84) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BWV1, Q6UXJ5, Q8N2P7, Q8NF26 | Gene names | BOC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brother of CDO precursor (Protein BOC). | |||||
|
CO7A1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.006330 (rank : 83) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 807 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q02388, Q14054, Q16507 | Gene names | COL7A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VII) chain precursor (Long-chain collagen) (LC collagen). | |||||
|
FMNL_HUMAN
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.014019 (rank : 71) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 529 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O95466, Q6DKG5, Q6IBP3, Q86UH1, Q8N671, Q8TDH1, Q96H10 | Gene names | FMNL1, C17orf1, C17orf1B, FMNL | |||
|
Domain Architecture |
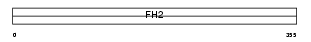
|
|||||
| Description | Formin-like protein 1 (Leukocyte formin) (CLL-associated antigen KW- 13). | |||||
|
MADCA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | 0.022812 (rank : 47) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q13477, O60222, O75867 | Gene names | MADCAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucosal addressin cell adhesion molecule 1 precursor (MAdCAM-1) (hMAdCAM-1). | |||||
|
MYPT2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.009018 (rank : 77) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 707 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O60237, Q8N179, Q9HCB7, Q9HCB8 | Gene names | PPP1R12B, MYPT2 | |||
|
Domain Architecture |
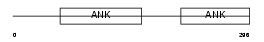
|
|||||
| Description | Protein phosphatase 1 regulatory subunit 12B (Myosin phosphatase- targeting subunit 2) (Myosin phosphatase target subunit 2). | |||||
|
PKHA5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.019078 (rank : 62) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9HAU0, Q8N3K6, Q96DY9, Q9BVR4, Q9C0H7, Q9H924, Q9NVK8 | Gene names | PLEKHA5, KIAA1686, PEPP2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 5 (Phosphoinositol 3-phosphate-binding protein 2) (PEPP-2). | |||||
|
TRI44_MOUSE
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | 0.024162 (rank : 44) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9QXA7, Q80SV9, Q9JHR8 | Gene names | Trim44, Dipb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 44 (Protein DIPB) (Protein Mc7). | |||||
|
TTF1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.025472 (rank : 40) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q15361, Q4VXF3, Q58EY2, Q6P5T5 | Gene names | TTF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 1 (TTF-1) (TTF-I) (RNA polymerase I termination factor). | |||||
|
XPC_MOUSE
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.020199 (rank : 57) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P51612, P54732, Q3TKI2, Q920M1, Q9DBW7 | Gene names | Xpc | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein complementing XP-C cells homolog (Xeroderma pigmentosum group C-complementing protein homolog) (p125). | |||||
|
ZNFX1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.015493 (rank : 67) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9P2E3, Q9BQM7, Q9BQM8, Q9H8C1, Q9H9S2, Q9NUM1, Q9NWW1 | Gene names | ZNFX1, KIAA1404 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
ANK3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.008364 (rank : 79) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 853 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q12955 | Gene names | ANK3 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-3 (ANK-3) (Ankyrin-G). | |||||
|
BFSP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.011511 (rank : 76) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 508 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q12934, O43595, O76034, Q9HBX4 | Gene names | BFSP1 | |||
|
Domain Architecture |
|
|||||
| Description | Filensin (Beaded filament structural protein 1) (Lens fiber cell beaded-filament structural protein CP 115) (CP115) (Lens intermediate filament-like heavy) (LIFL-H). | |||||
|
BRD4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.022739 (rank : 48) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
COCA1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.006801 (rank : 81) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q60847, P70322 | Gene names | Col12a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XII) chain precursor. | |||||
|
DLX5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 81) | NC score | 0.003585 (rank : 85) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P70396, O54876, O54877, O54878, Q9JJ45 | Gene names | Dlx5 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein DLX-5. | |||||
|
FIBA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.006418 (rank : 82) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P02671, Q9BX62, Q9UCH2 | Gene names | FGA | |||
|
Domain Architecture |

|
|||||
| Description | Fibrinogen alpha chain precursor [Contains: Fibrinopeptide A]. | |||||
|
MDC1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 83) | NC score | 0.021536 (rank : 51) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
PHF3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.019991 (rank : 58) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q92576, Q5T1T6, Q9NQ16, Q9UI45 | Gene names | PHF3, KIAA0244 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 3. | |||||
|
PTMS_HUMAN
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.037139 (rank : 23) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P20962 | Gene names | PTMS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
PTMS_MOUSE
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.038345 (rank : 21) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D0J8 | Gene names | Ptms | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
RSF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.032389 (rank : 32) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
ZBTB4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.007751 (rank : 80) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 926 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9P1Z0, Q7Z697, Q86XJ4, Q8N4V8 | Gene names | ZBTB4, KIAA1538 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger and BTB domain-containing protein 4 (KAISO-like zinc finger protein 1) (KAISO-L1). | |||||
|
SEBP2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 88 | |
| SwissProt Accessions | Q96T21, Q8IYC0 | Gene names | SECISBP2, SBP2 | |||
|
Domain Architecture |

|
|||||
| Description | SECIS-binding protein 2 (Selenocysteine insertion sequence-binding protein 2). | |||||
|
K0256_HUMAN
|
||||||
| NC score | 0.839924 (rank : 2) | θ value | 1.45253e-74 (rank : 3) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q93073, Q8N767 | Gene names | KIAA0256 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0256. | |||||
|
K0256_MOUSE
|
||||||
| NC score | 0.829823 (rank : 3) | θ value | 3.82246e-75 (rank : 2) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6A098 | Gene names | Kiaa0256 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0256. | |||||
|
T2FA_HUMAN
|
||||||
| NC score | 0.082770 (rank : 4) | θ value | 0.21417 (rank : 8) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P35269, Q9BWN0 | Gene names | GTF2F1, RAP74 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor IIF alpha subunit (EC 2.7.11.1) (TFIIF-alpha) (Transcription initiation factor RAP74) (General transcription factor IIF polypeptide 1 74 kDa subunit protein). | |||||
|
CLSPN_MOUSE
|
||||||
| NC score | 0.079270 (rank : 5) | θ value | 0.0330416 (rank : 4) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q80YR7, Q69GM2 | Gene names | Clspn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Claspin. | |||||
|
RPO3G_HUMAN
|
||||||
| NC score | 0.070012 (rank : 6) | θ value | 0.62314 (rank : 15) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15318 | Gene names | POLR3G, RPC32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA-directed RNA polymerase III subunit G (EC 2.7.7.6) (DNA-directed RNA polymerase III 32 kDa polypeptide) (RNA polymerase III C32 subunit). | |||||
|
NHPX_HUMAN
|
||||||
| NC score | 0.063278 (rank : 7) | θ value | 4.03905 (rank : 49) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P55769 | Gene names | NHP2L1 | |||
|
Domain Architecture |

|
|||||
| Description | NHP2-like protein 1 (High mobility group-like nuclear protein 2 homolog 1) (U4/U6.U5 tri-snRNP 15.5 kDa protein) (OTK27) (hSNU13). | |||||
|
NHPX_MOUSE
|
||||||
| NC score | 0.063278 (rank : 8) | θ value | 4.03905 (rank : 50) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D0T1, O08754, Q3UY64 | Gene names | Nhp2l1, Ssfa1 | |||
|
Domain Architecture |

|
|||||
| Description | NHP2-like protein 1 (High mobility group-like nuclear protein 2 homolog 1) (U4/U6.U5 tri-snRNP 15.5 kDa protein) (Sperm-specific antigen 1) (Fertilization antigen 1) (FA-1). | |||||
|
RIF1_HUMAN
|
||||||
| NC score | 0.062436 (rank : 9) | θ value | 0.0961366 (rank : 5) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 320 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5UIP0, Q5H9R3, Q5UIP2, Q66YK6, Q6PRU2, Q8TE94, Q99772, Q9H830, Q9H9B9, Q9NVP5, Q9Y4R4 | Gene names | RIF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomere-associated protein RIF1 (Rap1-interacting factor 1 homolog). | |||||
|
PKCB1_HUMAN
|
||||||
| NC score | 0.055461 (rank : 10) | θ value | 0.365318 (rank : 10) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
NOL1_HUMAN
|
||||||
| NC score | 0.055230 (rank : 11) | θ value | 1.38821 (rank : 21) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P46087 | Gene names | NOL1 | |||
|
Domain Architecture |

|
|||||
| Description | Proliferating-cell nucleolar antigen p120 (Proliferation-associated nucleolar protein p120). | |||||
|
PABP2_MOUSE
|
||||||
| NC score | 0.048604 (rank : 12) | θ value | 0.813845 (rank : 18) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CCS6, O35935 | Gene names | Pabpn1, Pab2, Pabp2 | |||
|
Domain Architecture |

|
|||||
| Description | Polyadenylate-binding protein 2 (Poly(A)-binding protein 2) (Poly(A)- binding protein II) (PABII) (Polyadenylate-binding nuclear protein 1) (Nuclear poly(A)-binding protein 1). | |||||
|
PABP2_HUMAN
|
||||||
| NC score | 0.048160 (rank : 13) | θ value | 0.813845 (rank : 17) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q86U42, O43484 | Gene names | PABPN1, PAB2, PABP2 | |||
|
Domain Architecture |

|
|||||
| Description | Polyadenylate-binding protein 2 (Poly(A)-binding protein 2) (Poly(A)- binding protein II) (PABII) (Polyadenylate-binding nuclear protein 1) (Nuclear poly(A)-binding protein 1). | |||||
|
CBP_HUMAN
|
||||||
| NC score | 0.047617 (rank : 14) | θ value | 0.47712 (rank : 11) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CBP_MOUSE
|
||||||
| NC score | 0.045040 (rank : 15) | θ value | 0.813845 (rank : 16) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CDN1C_MOUSE
|
||||||
| NC score | 0.042355 (rank : 16) | θ value | 1.81305 (rank : 23) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P49919 | Gene names | Cdkn1c, Kip2 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-dependent kinase inhibitor 1C (Cyclin-dependent kinase inhibitor p57) (p57KIP2). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.042320 (rank : 17) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.041987 (rank : 18) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
NOLA2_MOUSE
|
||||||
| NC score | 0.040940 (rank : 19) | θ value | 4.03905 (rank : 51) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CRB2 | Gene names | Nola2, Nhp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | H/ACA ribonucleoprotein complex subunit 2 (Nucleolar protein family A member 2) (snoRNP protein NHP2). | |||||
|
SPG20_MOUSE
|
||||||
| NC score | 0.039551 (rank : 20) | θ value | 1.38821 (rank : 22) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R1X6, Q6ZQ87, Q8BJD3, Q8BM37, Q8BZ63 | Gene names | Spg20, Kiaa0610 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spartin. | |||||
|
PTMS_MOUSE
|
||||||
| NC score | 0.038345 (rank : 21) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D0J8 | Gene names | Ptms | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
CCD71_HUMAN
|
||||||
| NC score | 0.037844 (rank : 22) | θ value | 3.0926 (rank : 35) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IV32, Q6IPE2, Q9H8H4, Q9H9F1 | Gene names | CCDC71 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 71. | |||||
|
PTMS_HUMAN
|
||||||
| NC score | 0.037139 (rank : 23) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P20962 | Gene names | PTMS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
PF21A_MOUSE
|
||||||
| NC score | 0.036852 (rank : 24) | θ value | 0.62314 (rank : 14) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6ZPK0, Q6XVG0, Q80Z33, Q8CAZ4, Q8VEC8 | Gene names | Phf21a, Bhc80, Kiaa1696, Pftf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (mBHC80) (BHC80a). | |||||
|
NOP14_HUMAN
|
||||||
| NC score | 0.036720 (rank : 25) | θ value | 1.06291 (rank : 19) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P78316, Q7LGI5, Q7Z6K0, Q8TBR6 | Gene names | NOP14, C4orf9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.036127 (rank : 26) | θ value | 1.81305 (rank : 24) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
ZN622_HUMAN
|
||||||
| NC score | 0.035858 (rank : 27) | θ value | 2.36792 (rank : 33) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q969S3 | Gene names | ZNF622, ZPR9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 622 (Zinc finger-like protein 9). | |||||
|
ATRX_HUMAN
|
||||||
| NC score | 0.035200 (rank : 28) | θ value | 3.0926 (rank : 34) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P46100, P51068, Q15886, Q59FB5, Q59H31, Q5H9A2, Q5JWI4, Q7Z2J1, Q9H0Z1, Q9NTS3 | Gene names | ATRX, RAD54L, XH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked helicase II) (X-linked nuclear protein) (XNP) (Znf- HX). | |||||
|
MAGC3_HUMAN
|
||||||
| NC score | 0.035006 (rank : 29) | θ value | 0.21417 (rank : 7) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TD91, Q5JZ43, Q9BZ80 | Gene names | MAGEC3, HCA2 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen C3 (MAGE-C3 antigen) (Hepatocellular carcinoma-associated antigen 2). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.034449 (rank : 30) | θ value | 2.36792 (rank : 31) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
PINX1_HUMAN
|
||||||
| NC score | 0.034430 (rank : 31) | θ value | 3.0926 (rank : 40) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96BK5, Q7Z7J8, Q96QD7, Q9HBU7, Q9NWW2 | Gene names | PINX1, LPTS | |||
|
Domain Architecture |

|
|||||
| Description | Pin2-interacting protein X1 (TRF1-interacting protein 1) (Liver- related putative tumor suppressor) (67-11-3 protein). | |||||
|
RSF1_HUMAN
|
||||||
| NC score | 0.032389 (rank : 32) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
CF142_HUMAN
|
||||||
| NC score | 0.031416 (rank : 33) | θ value | 4.03905 (rank : 44) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5VWP3, Q96H08, Q96NF7 | Gene names | C6orf142 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf142. | |||||
|
DNMT1_HUMAN
|
||||||
| NC score | 0.031098 (rank : 34) | θ value | 5.27518 (rank : 57) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P26358, Q9UHG5, Q9ULA2, Q9UMZ6 | Gene names | DNMT1, AIM, DNMT | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase HsaI) (DNA MTase HsaI) (MCMT) (M.HsaI). | |||||
|
PRP40_MOUSE
|
||||||
| NC score | 0.029874 (rank : 35) | θ value | 5.27518 (rank : 63) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
COA2_HUMAN
|
||||||
| NC score | 0.029380 (rank : 36) | θ value | 1.38821 (rank : 20) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O00763, Q16852 | Gene names | ACACB, ACC2, ACCB | |||
|
Domain Architecture |

|
|||||
| Description | Acetyl-CoA carboxylase 2 (EC 6.4.1.2) (ACC-beta) [Includes: Biotin carboxylase (EC 6.3.4.14)]. | |||||
|
SETX_HUMAN
|
||||||
| NC score | 0.029072 (rank : 37) | θ value | 3.0926 (rank : 41) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7Z333, O75120, Q3KQX4, Q68DW5, Q6AZD7, Q7Z3J6, Q8WX33, Q9H9D1, Q9NVP9 | Gene names | SETX, ALS4, KIAA0625, SCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable helicase senataxin (EC 3.6.1.-) (SEN1 homolog). | |||||
|
WWC1_MOUSE
|
||||||
| NC score | 0.027327 (rank : 38) | θ value | 1.81305 (rank : 25) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5SXA9, Q571D0, Q8K1Y3, Q8VD17, Q922W3 | Gene names | Wwc1, Kiaa0869 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing protein 1 (Kidney and brain protein) (KIBRA). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.025806 (rank : 39) | θ value | 0.62314 (rank : 13) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
TTF1_HUMAN
|
||||||
| NC score | 0.025472 (rank : 40) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q15361, Q4VXF3, Q58EY2, Q6P5T5 | Gene names | TTF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 1 (TTF-1) (TTF-I) (RNA polymerase I termination factor). | |||||
|
DNJC1_HUMAN
|
||||||
| NC score | 0.024570 (rank : 41) | θ value | 4.03905 (rank : 45) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96KC8 | Gene names | DNAJC1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 1. | |||||
|
4ET_MOUSE
|
||||||
| NC score | 0.024504 (rank : 42) | θ value | 4.03905 (rank : 42) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9EST3, Q9CSS3 | Gene names | Eif4enif1, Clast4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4E transporter (eIF4E transporter) (4E-T) (Eukaryotic translation initiation factor 4E nuclear import factor 1). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.024335 (rank : 43) | θ value | 4.03905 (rank : 48) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
TRI44_MOUSE
|
||||||
| NC score | 0.024162 (rank : 44) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9QXA7, Q80SV9, Q9JHR8 | Gene names | Trim44, Dipb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 44 (Protein DIPB) (Protein Mc7). | |||||
|
BCOR_HUMAN
|
||||||
| NC score | 0.024026 (rank : 45) | θ value | 5.27518 (rank : 55) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6W2J9, Q6P4B6, Q7Z2K7, Q8TEB4, Q96DB3, Q9H232, Q9H233, Q9HCJ7, Q9NXF2 | Gene names | BCOR, KIAA1575 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BCoR protein (BCL-6 corepressor). | |||||
|
DNJC1_MOUSE
|
||||||
| NC score | 0.023562 (rank : 46) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61712 | Gene names | Dnajc1, Dnajl1, Mtj1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 1 (DnaJ protein homolog MTJ1). | |||||
|
MADCA_HUMAN
|
||||||
| NC score | 0.022812 (rank : 47) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q13477, O60222, O75867 | Gene names | MADCAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucosal addressin cell adhesion molecule 1 precursor (MAdCAM-1) (hMAdCAM-1). | |||||
|
BRD4_HUMAN
|
||||||
| NC score | 0.022739 (rank : 48) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
NFH_HUMAN
|
||||||
| NC score | 0.022632 (rank : 49) | θ value | 2.36792 (rank : 28) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
GOGB1_HUMAN
|
||||||
| NC score | 0.021953 (rank : 50) | θ value | 0.62314 (rank : 12) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.021536 (rank : 51) | θ value | 8.99809 (rank : 83) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
AP3B1_HUMAN
|
||||||
| NC score | 0.021363 (rank : 52) | θ value | 4.03905 (rank : 43) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O00203, O00580, Q7Z393, Q9HD66 | Gene names | AP3B1, ADTB3A | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
GASP1_MOUSE
|
||||||
| NC score | 0.020876 (rank : 53) | θ value | 5.27518 (rank : 59) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5U4C1, Q8BKR8, Q8BUN4, Q8BYK9, Q8CHF4, Q8R095 | Gene names | Gprasp1, Kiaa0443 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor-associated sorting protein 1 (GASP-1). | |||||
|
PIBF1_HUMAN
|
||||||
| NC score | 0.020656 (rank : 54) | θ value | 0.365318 (rank : 9) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8WXW3, O95664, Q6U9V2, Q6UG50, Q86V07, Q96SF4 | Gene names | PIBF1, C13orf24, PIBF | |||
|
Domain Architecture |

|
|||||
| Description | Progesterone-induced-blocking factor 1. | |||||
|
RBM10_HUMAN
|
||||||
| NC score | 0.020602 (rank : 55) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P98175, Q14136 | Gene names | RBM10, DXS8237E, KIAA0122 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 10 (RNA-binding motif protein 10). | |||||
|
AN32E_HUMAN
|
||||||
| NC score | 0.020221 (rank : 56) | θ value | 2.36792 (rank : 26) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BTT0, Q8N1S4, Q8WWW9 | Gene names | ANP32E | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member E (LANP- like protein) (LANP-L). | |||||
|
XPC_MOUSE
|
||||||
| NC score | 0.020199 (rank : 57) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P51612, P54732, Q3TKI2, Q920M1, Q9DBW7 | Gene names | Xpc | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein complementing XP-C cells homolog (Xeroderma pigmentosum group C-complementing protein homolog) (p125). | |||||
|
PHF3_HUMAN
|
||||||
| NC score | 0.019991 (rank : 58) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q92576, Q5T1T6, Q9NQ16, Q9UI45 | Gene names | PHF3, KIAA0244 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 3. | |||||
|
SHAN2_HUMAN
|
||||||
| NC score | 0.019911 (rank : 59) | θ value | 4.03905 (rank : 52) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 346 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UPX8, Q3Y8G9, Q52LK2, Q9UKP1 | Gene names | SHANK2, KIAA1022 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 2 (Shank2). | |||||
|
ESCO2_HUMAN
|
||||||
| NC score | 0.019559 (rank : 60) | θ value | 5.27518 (rank : 58) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q56NI9 | Gene names | ESCO2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyltransferase ESCO2 (EC 2.3.1.-) (Establishment of cohesion 1 homolog 2) (ECO1 homolog 2). | |||||
|
IRX1_HUMAN
|
||||||
| NC score | 0.019312 (rank : 61) | θ value | 2.36792 (rank : 27) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P78414, Q7Z2F8, Q8N312 | Gene names | IRX1, IRXA1 | |||
|
Domain Architecture |

|
|||||
| Description | Iroquois-class homeodomain protein IRX-1 (Iroquois homeobox protein 1) (Homeodomain protein IRXA1). | |||||
|
PKHA5_HUMAN
|
||||||
| NC score | 0.019078 (rank : 62) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9HAU0, Q8N3K6, Q96DY9, Q9BVR4, Q9C0H7, Q9H924, Q9NVK8 | Gene names | PLEKHA5, KIAA1686, PEPP2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 5 (Phosphoinositol 3-phosphate-binding protein 2) (PEPP-2). | |||||
|
CIC_MOUSE
|
||||||
| NC score | 0.017550 (rank : 63) | θ value | 5.27518 (rank : 56) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 742 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q924A2, Q6PDJ8, Q8CGE4, Q8CHH0, Q9CW61 | Gene names | Cic, Kiaa0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
TSH1_MOUSE
|
||||||
| NC score | 0.017316 (rank : 64) | θ value | 4.03905 (rank : 53) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 675 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5DTH5, Q6PE60, Q8VD19, Q9JLD5 | Gene names | Tshz1, Kiaa4206, Sdccag33, Tsh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 1 (Serologically defined colon cancer antigen 3 homolog). | |||||
|
JAD1C_HUMAN
|
||||||
| NC score | 0.016730 (rank : 65) | θ value | 3.0926 (rank : 38) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P41229 | Gene names | SMCX, DXS1272E, JARID1C, XE169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (Protein SmcX) (Protein Xe169). | |||||
|
GIT2_HUMAN
|
||||||
| NC score | 0.016629 (rank : 66) | θ value | 3.0926 (rank : 37) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14161, Q86U59, Q96CI2, Q9BV91, Q9Y5V2 | Gene names | GIT2, KIAA0148 | |||
|
Domain Architecture |

|
|||||
| Description | ARF GTPase-activating protein GIT2 (G protein-coupled receptor kinase- interactor 2) (GRK-interacting protein 2) (Cool-interacting tyrosine- phosphorylated protein 2) (CAT2) (CAT-2). | |||||
|
ZNFX1_HUMAN
|
||||||
| NC score | 0.015493 (rank : 67) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9P2E3, Q9BQM7, Q9BQM8, Q9H8C1, Q9H9S2, Q9NUM1, Q9NWW1 | Gene names | ZNFX1, KIAA1404 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
ARP5_MOUSE
|
||||||
| NC score | 0.014724 (rank : 68) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80US4, Q8BL26 | Gene names | Actr5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-related protein 5. | |||||
|
ZAN_MOUSE
|
||||||
| NC score | 0.014333 (rank : 69) | θ value | 2.36792 (rank : 32) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O88799, O08647 | Gene names | Zan | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
CADH4_MOUSE
|
||||||
| NC score | 0.014167 (rank : 70) | θ value | 0.125558 (rank : 6) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P39038 | Gene names | Cdh4 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-4 precursor (Retinal-cadherin) (R-cadherin) (R-CAD). | |||||
|
FMNL_HUMAN
|
||||||
| NC score | 0.014019 (rank : 71) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 529 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O95466, Q6DKG5, Q6IBP3, Q86UH1, Q8N671, Q8TDH1, Q96H10 | Gene names | FMNL1, C17orf1, C17orf1B, FMNL | |||
|
Domain Architecture |

|
|||||
| Description | Formin-like protein 1 (Leukocyte formin) (CLL-associated antigen KW- 13). | |||||
|
LRP12_MOUSE
|
||||||
| NC score | 0.013875 (rank : 72) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BUJ9, Q8BWM9 | Gene names | Lrp12 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 12 precursor. | |||||
|
LRP12_HUMAN
|
||||||
| NC score | 0.013848 (rank : 73) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y561 | Gene names | LRP12, ST7 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 12 precursor (Suppressor of tumorigenicity protein 7). | |||||
|
RBCC1_HUMAN
|
||||||
| NC score | 0.012808 (rank : 74) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1023 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8TDY2, Q8WVU9, Q92601 | Gene names | RB1CC1, KIAA0203, RBICC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RB1-inducible coiled-coil protein 1. | |||||
|
PRP16_HUMAN
|
||||||
| NC score | 0.012039 (rank : 75) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92620, O75212, Q96HN7 | Gene names | DHX38, DDX38, KIAA0224, PRP16 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-splicing factor ATP-dependent RNA helicase PRP16 (EC 3.6.1.-) (ATP-dependent RNA helicase DHX38) (DEAH box protein 38). | |||||
|
BFSP1_HUMAN
|
||||||
| NC score | 0.011511 (rank : 76) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 508 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q12934, O43595, O76034, Q9HBX4 | Gene names | BFSP1 | |||
|
Domain Architecture |

|
|||||
| Description | Filensin (Beaded filament structural protein 1) (Lens fiber cell beaded-filament structural protein CP 115) (CP115) (Lens intermediate filament-like heavy) (LIFL-H). | |||||
|
MYPT2_HUMAN
|
||||||
| NC score | 0.009018 (rank : 77) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 707 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O60237, Q8N179, Q9HCB7, Q9HCB8 | Gene names | PPP1R12B, MYPT2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein phosphatase 1 regulatory subunit 12B (Myosin phosphatase- targeting subunit 2) (Myosin phosphatase target subunit 2). | |||||
|
KIF1A_HUMAN
|
||||||
| NC score | 0.008630 (rank : 78) | θ value | 3.0926 (rank : 39) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q12756 | Gene names | KIF1A, ATSV | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF1A (Axonal transporter of synaptic vesicles). | |||||
|
ANK3_HUMAN
|
||||||
| NC score | 0.008364 (rank : 79) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 853 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q12955 | Gene names | ANK3 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-3 (ANK-3) (Ankyrin-G). | |||||
|
ZBTB4_HUMAN
|
||||||
| NC score | 0.007751 (rank : 80) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 926 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9P1Z0, Q7Z697, Q86XJ4, Q8N4V8 | Gene names | ZBTB4, KIAA1538 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 4 (KAISO-like zinc finger protein 1) (KAISO-L1). | |||||
|
COCA1_MOUSE
|
||||||
| NC score | 0.006801 (rank : 81) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q60847, P70322 | Gene names | Col12a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XII) chain precursor. | |||||
|
FIBA_HUMAN
|
||||||
| NC score | 0.006418 (rank : 82) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P02671, Q9BX62, Q9UCH2 | Gene names | FGA | |||
|
Domain Architecture |

|
|||||
| Description | Fibrinogen alpha chain precursor [Contains: Fibrinopeptide A]. | |||||
|
CO7A1_HUMAN
|
||||||
| NC score | 0.006330 (rank : 83) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 807 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q02388, Q14054, Q16507 | Gene names | COL7A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VII) chain precursor (Long-chain collagen) (LC collagen). | |||||
|
BOC_HUMAN
|
||||||
| NC score | 0.005771 (rank : 84) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BWV1, Q6UXJ5, Q8N2P7, Q8NF26 | Gene names | BOC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brother of CDO precursor (Protein BOC). | |||||
|
DLX5_MOUSE
|
||||||
| NC score | 0.003585 (rank : 85) | θ value | 8.99809 (rank : 81) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P70396, O54876, O54877, O54878, Q9JJ45 | Gene names | Dlx5 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein DLX-5. | |||||
|
MTF1_MOUSE
|
||||||
| NC score | 0.001936 (rank : 86) | θ value | 4.03905 (rank : 47) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q07243 | Gene names | Mtf1, Mtf-1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-regulatory transcription factor 1 (Transcription factor MTF-1) (MRE-binding transcription factor). | |||||
|
CDKL3_MOUSE
|
||||||
| NC score | 0.001119 (rank : 87) | θ value | 3.0926 (rank : 36) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 857 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BLF2, Q5M6W2, Q8BKR2, Q8BL49, Q8BVE0, Q8K134 | Gene names | Cdkl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-dependent kinase-like 3 (EC 2.7.11.22). | |||||
|
ZN227_HUMAN
|
||||||
| NC score | -0.000149 (rank : 88) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 781 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q86WZ6 | Gene names | ZNF227 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 227. | |||||