Please be patient as the page loads
|
PARP2_HUMAN
|
||||||
| SwissProt Accessions | Q9UGN5, Q8TEU4, Q9NUV2, Q9UMR4, Q9Y6C8 | Gene names | PARP2, ADPRT2, ADPRTL2 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 2 (EC 2.4.2.30) (PARP-2) (NAD(+) ADP- ribosyltransferase 2) (Poly[ADP-ribose] synthetase 2) (pADPRT-2) (hPARP-2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PARP2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UGN5, Q8TEU4, Q9NUV2, Q9UMR4, Q9Y6C8 | Gene names | PARP2, ADPRT2, ADPRTL2 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 2 (EC 2.4.2.30) (PARP-2) (NAD(+) ADP- ribosyltransferase 2) (Poly[ADP-ribose] synthetase 2) (pADPRT-2) (hPARP-2). | |||||
|
PARP2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.991472 (rank : 2) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O88554, Q99N29 | Gene names | Parp2, Adprt2, Adprtl2, Aspartl2 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 2 (EC 2.4.2.30) (PARP-2) (NAD(+) ADP- ribosyltransferase 2) (Poly[ADP-ribose] synthetase 2) (pADPRT-2) (mPARP-2). | |||||
|
PARP1_HUMAN
|
||||||
| θ value | 8.73085e-104 (rank : 3) | NC score | 0.920451 (rank : 4) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P09874, Q8IUZ9 | Gene names | PARP1, ADPRT, PPOL | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 1 (EC 2.4.2.30) (PARP-1) (ADPRT) (NAD(+) ADP-ribosyltransferase 1) (Poly[ADP-ribose] synthetase 1). | |||||
|
PARP1_MOUSE
|
||||||
| θ value | 2.80862e-102 (rank : 4) | NC score | 0.910424 (rank : 5) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P11103, Q9JLX4, Q9QVQ3 | Gene names | Parp1, Adprp, Adprt, Adprt1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 1 (EC 2.4.2.30) (PARP-1) (ADPRT) (NAD(+) ADP-ribosyltransferase 1) (Poly[ADP-ribose] synthetase 1) (msPARP). | |||||
|
PARP3_HUMAN
|
||||||
| θ value | 3.70236e-70 (rank : 5) | NC score | 0.926769 (rank : 3) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y6F1, Q8NER9, Q96CG2, Q9UG81 | Gene names | PARP3, ADPRT3, ADPRTL3 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 3 (EC 2.4.2.30) (PARP-3) (NAD(+) ADP- ribosyltransferase 3) (Poly[ADP-ribose] synthetase 3) (pADPRT-3) (hPARP-3) (IRT1). | |||||
|
PARP4_HUMAN
|
||||||
| θ value | 3.75424e-22 (rank : 6) | NC score | 0.566220 (rank : 6) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UKK3, O75903, Q14682, Q9H1M6 | Gene names | PARP4, ADPRTL1, KIAA0177, PARPL | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 4 (EC 2.4.2.30) (PARP-4) (Vault poly(ADP- ribose) polymerase) (VPARP) (193 kDa vault protein) (PARP- related/IalphaI-related H5/proline-rich) (PH5P). | |||||
|
TNKS1_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 7) | NC score | 0.072945 (rank : 12) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O95271, O95272 | Gene names | TNKS, PARP5A, PARPL, TIN1, TINF1, TNKS1 | |||
|
Domain Architecture |

|
|||||
| Description | Tankyrase-1 (EC 2.4.2.30) (TANK1) (Tankyrase I) (TNKS-1) (TRF1- interacting ankyrin-related ADP-ribose polymerase). | |||||
|
PARP8_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 8) | NC score | 0.192209 (rank : 7) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8N3A8, Q3KRB7, Q6DHZ1, Q9H754 | Gene names | PARP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 8 (EC 2.4.2.30) (PARP-8). | |||||
|
PARP8_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 9) | NC score | 0.191937 (rank : 8) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q3UD82, Q3UDE4, Q3UDU5, Q8VCB5, Q9CYY3 | Gene names | Parp8, D13Ertd275e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 8 (EC 2.4.2.30) (PARP-8). | |||||
|
TNKS2_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 10) | NC score | 0.068075 (rank : 15) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9H2K2, Q9H8F2, Q9HAS4 | Gene names | TNKS2, PARP5B, TANK2, TNKL | |||
|
Domain Architecture |

|
|||||
| Description | Tankyrase-2 (EC 2.4.2.30) (TANK2) (Tankyrase II) (TNKS-2) (TRF1- interacting ankyrin-related ADP-ribose polymerase 2) (Tankyrase-like protein) (Tankyrase-related protein). | |||||
|
PARP6_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 11) | NC score | 0.187971 (rank : 9) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q2NL67, Q9H7C5, Q9H9X6, Q9HAF3, Q9NPS6, Q9UFG4 | Gene names | PARP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 6 (EC 2.4.2.30) (PARP-6). | |||||
|
PARP6_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 12) | NC score | 0.187957 (rank : 10) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6P6P7, Q8BVW1 | Gene names | Parp6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 6 (EC 2.4.2.30) (PARP-6). | |||||
|
JAD1A_HUMAN
|
||||||
| θ value | 0.813845 (rank : 13) | NC score | 0.021314 (rank : 25) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P29375 | Gene names | JARID1A, RBBP2, RBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1A (Retinoblastoma-binding protein 2) (RBBP-2). | |||||
|
OSBP1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 14) | NC score | 0.013252 (rank : 29) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P22059 | Gene names | OSBP, OSBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Oxysterol-binding protein 1. | |||||
|
PAR16_MOUSE
|
||||||
| θ value | 1.81305 (rank : 15) | NC score | 0.067298 (rank : 16) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q7TMM8, Q3V031, Q8CC71 | Gene names | Parp16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 16 (EC 2.4.2.30) (PARP-16). | |||||
|
PARPT_MOUSE
|
||||||
| θ value | 1.81305 (rank : 16) | NC score | 0.080700 (rank : 11) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8C1B2, Q3UD50, Q8C032 | Gene names | Tiparp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TCDD-inducible poly [ADP-ribose] polymerase (EC 2.4.2.30). | |||||
|
CK5P2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 17) | NC score | 0.007018 (rank : 34) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 1433 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96SN8, Q7Z3L4, Q7Z3U1, Q7Z7I6, Q9BSW0, Q9H6J6, Q9HCD9, Q9NV90, Q9UIW9 | Gene names | CDK5RAP2, CEP215, KIAA1633 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48) (Centrosome-associated protein 215). | |||||
|
DDX46_MOUSE
|
||||||
| θ value | 2.36792 (rank : 18) | NC score | 0.009465 (rank : 31) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q569Z5, Q6ZQ42, Q8R0R6 | Gene names | Ddx46, Kiaa0801 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX46 (EC 3.6.1.-) (DEAD box protein 46). | |||||
|
PARPT_HUMAN
|
||||||
| θ value | 2.36792 (rank : 19) | NC score | 0.064402 (rank : 17) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7Z3E1, Q68CY9, Q86VP4, Q9Y4P7 | Gene names | TIPARP, PARP7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TCDD-inducible poly [ADP-ribose] polymerase (EC 2.4.2.30) (Poly [ADP- ribose] polymerase 7) (PARP-7). | |||||
|
THUM1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 20) | NC score | 0.026680 (rank : 23) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NXG2, Q9BWC3 | Gene names | THUMPD1 | |||
|
Domain Architecture |

|
|||||
| Description | THUMP domain-containing protein 1. | |||||
|
MYO5C_HUMAN
|
||||||
| θ value | 3.0926 (rank : 21) | NC score | 0.004967 (rank : 35) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 1206 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NQX4 | Gene names | MYO5C | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5C (Myosin Vc). | |||||
|
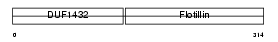
FLOT2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 22) | NC score | 0.032699 (rank : 21) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q14254 | Gene names | FLOT2, ESA1, M17S1 | |||
|
Domain Architecture |
|
|||||
| Description | Flotillin-2 (Epidermal surface antigen) (ESA). | |||||
|
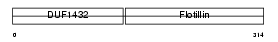
FLOT2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 23) | NC score | 0.032659 (rank : 22) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q60634, Q6NS75 | Gene names | Flot2, Esa1, M17s1 | |||
|
Domain Architecture |
|
|||||
| Description | Flotillin-2 (Epidermal surface antigen) (ESA). | |||||
|
CP007_HUMAN
|
||||||
| θ value | 5.27518 (rank : 24) | NC score | 0.024840 (rank : 24) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y2B5 | Gene names | C16orf7, ATPBL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf7 (Protein ATP-BL). | |||||
|
DDX46_HUMAN
|
||||||
| θ value | 5.27518 (rank : 25) | NC score | 0.008296 (rank : 32) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7L014, O94894, Q96EI0, Q9Y658 | Gene names | DDX46, KIAA0801 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX46 (EC 3.6.1.-) (DEAD box protein 46) (PRP5 homolog). | |||||
|
CAPS1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 26) | NC score | 0.013384 (rank : 28) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ULU8, Q13339, Q6GQQ6, Q8N2Z5, Q8N3M7, Q8NFR0, Q96BC2 | Gene names | CADPS, CAPS, CAPS1, KIAA1121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-dependent secretion activator 1 (Calcium-dependent activator protein for secretion 1) (CAPS-1). | |||||
|
CP007_MOUSE
|
||||||
| θ value | 6.88961 (rank : 27) | NC score | 0.019579 (rank : 26) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C190, Q6PFD1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf7 homolog (5-day ovary-specific transcript 1 protein). | |||||
|
PAR16_HUMAN
|
||||||
| θ value | 6.88961 (rank : 28) | NC score | 0.059217 (rank : 19) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N5Y8, Q6PK64, Q9NX03 | Gene names | PARP16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 16 (EC 2.4.2.30) (PARP-16). | |||||
|
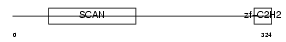
ZN396_HUMAN
|
||||||
| θ value | 6.88961 (rank : 29) | NC score | -0.000648 (rank : 37) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 703 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96N95, Q8NF98, Q8TD80 | Gene names | ZNF396, ZSCAN14 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 396 (Zinc finger and SCAN domain-containing protein 14). | |||||
|
ZNFX1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 30) | NC score | 0.007930 (rank : 33) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9P2E3, Q9BQM7, Q9BQM8, Q9H8C1, Q9H9S2, Q9NUM1, Q9NWW1 | Gene names | ZNFX1, KIAA1404 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
41_HUMAN
|
||||||
| θ value | 8.99809 (rank : 31) | NC score | 0.003776 (rank : 36) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P11171, P11176, Q14245, Q5TB35, Q5VXN8, Q8IXV9, Q9Y578, Q9Y579 | Gene names | EPB41, E41P | |||
|
Domain Architecture |

|
|||||
| Description | Protein 4.1 (Band 4.1) (P4.1) (EPB4.1) (4.1R). | |||||
|
CAPS1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 32) | NC score | 0.012582 (rank : 30) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80TJ1, Q3TSP2, Q61374, Q6AXB4, Q6PGF0 | Gene names | Cadps, Caps, Caps1, Kiaa1121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-dependent secretion activator 1 (Calcium-dependent activator protein for secretion 1) (CAPS-1). | |||||
|
CIP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 33) | NC score | 0.017094 (rank : 27) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NPC3 | Gene names | CCNB1IP1, C14orf18, HEI10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-B1-interacting protein 1 (EC 6.3.2.-) (Human enhancer of invasion 10) (E3 ubiquitin-protein ligase HEI10). | |||||
|
DNL3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 34) | NC score | 0.071019 (rank : 13) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P49916, Q16714 | Gene names | LIG3 | |||
|
Domain Architecture |
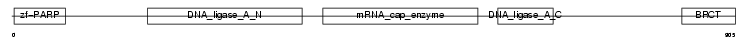
|
|||||
| Description | DNA ligase 3 (EC 6.5.1.1) (DNA ligase III) (Polydeoxyribonucleotide synthase [ATP] 3). | |||||
|
DNL3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 35) | NC score | 0.069124 (rank : 14) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P97386, P97385 | Gene names | Lig3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA ligase 3 (EC 6.5.1.1) (DNA ligase III) (Polydeoxyribonucleotide synthase [ATP] 3). | |||||
|
LHR2A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 36) | NC score | 0.055724 (rank : 20) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | O00534, Q6UN19, Q6UN20, Q9BVF8 | Gene names | LOH11CR2A, BCSC1 | |||
|
Domain Architecture |

|
|||||
| Description | Loss of heterozygosity 11 chromosomal region 2 gene A protein (Breast cancer suppressor candidate 1) (BCSC-1). | |||||
|
LHR2A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 37) | NC score | 0.059303 (rank : 18) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | Q99KC8, Q3TTU2, Q6UN22, Q8BHA8, Q9CTV9 | Gene names | Loh11cr2a | |||
|
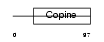
Domain Architecture |
|
|||||
| Description | Loss of heterozygosity 11 chromosomal region 2 gene A protein homolog. | |||||
|
PARP2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UGN5, Q8TEU4, Q9NUV2, Q9UMR4, Q9Y6C8 | Gene names | PARP2, ADPRT2, ADPRTL2 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 2 (EC 2.4.2.30) (PARP-2) (NAD(+) ADP- ribosyltransferase 2) (Poly[ADP-ribose] synthetase 2) (pADPRT-2) (hPARP-2). | |||||
|
PARP2_MOUSE
|
||||||
| NC score | 0.991472 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O88554, Q99N29 | Gene names | Parp2, Adprt2, Adprtl2, Aspartl2 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 2 (EC 2.4.2.30) (PARP-2) (NAD(+) ADP- ribosyltransferase 2) (Poly[ADP-ribose] synthetase 2) (pADPRT-2) (mPARP-2). | |||||
|
PARP3_HUMAN
|
||||||
| NC score | 0.926769 (rank : 3) | θ value | 3.70236e-70 (rank : 5) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y6F1, Q8NER9, Q96CG2, Q9UG81 | Gene names | PARP3, ADPRT3, ADPRTL3 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 3 (EC 2.4.2.30) (PARP-3) (NAD(+) ADP- ribosyltransferase 3) (Poly[ADP-ribose] synthetase 3) (pADPRT-3) (hPARP-3) (IRT1). | |||||
|
PARP1_HUMAN
|
||||||
| NC score | 0.920451 (rank : 4) | θ value | 8.73085e-104 (rank : 3) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P09874, Q8IUZ9 | Gene names | PARP1, ADPRT, PPOL | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 1 (EC 2.4.2.30) (PARP-1) (ADPRT) (NAD(+) ADP-ribosyltransferase 1) (Poly[ADP-ribose] synthetase 1). | |||||
|
PARP1_MOUSE
|
||||||
| NC score | 0.910424 (rank : 5) | θ value | 2.80862e-102 (rank : 4) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P11103, Q9JLX4, Q9QVQ3 | Gene names | Parp1, Adprp, Adprt, Adprt1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 1 (EC 2.4.2.30) (PARP-1) (ADPRT) (NAD(+) ADP-ribosyltransferase 1) (Poly[ADP-ribose] synthetase 1) (msPARP). | |||||
|
PARP4_HUMAN
|
||||||
| NC score | 0.566220 (rank : 6) | θ value | 3.75424e-22 (rank : 6) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UKK3, O75903, Q14682, Q9H1M6 | Gene names | PARP4, ADPRTL1, KIAA0177, PARPL | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 4 (EC 2.4.2.30) (PARP-4) (Vault poly(ADP- ribose) polymerase) (VPARP) (193 kDa vault protein) (PARP- related/IalphaI-related H5/proline-rich) (PH5P). | |||||
|
PARP8_HUMAN
|
||||||
| NC score | 0.192209 (rank : 7) | θ value | 0.00390308 (rank : 8) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8N3A8, Q3KRB7, Q6DHZ1, Q9H754 | Gene names | PARP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 8 (EC 2.4.2.30) (PARP-8). | |||||
|
PARP8_MOUSE
|
||||||
| NC score | 0.191937 (rank : 8) | θ value | 0.00390308 (rank : 9) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q3UD82, Q3UDE4, Q3UDU5, Q8VCB5, Q9CYY3 | Gene names | Parp8, D13Ertd275e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 8 (EC 2.4.2.30) (PARP-8). | |||||
|
PARP6_HUMAN
|
||||||
| NC score | 0.187971 (rank : 9) | θ value | 0.0148317 (rank : 11) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q2NL67, Q9H7C5, Q9H9X6, Q9HAF3, Q9NPS6, Q9UFG4 | Gene names | PARP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 6 (EC 2.4.2.30) (PARP-6). | |||||
|
PARP6_MOUSE
|
||||||
| NC score | 0.187957 (rank : 10) | θ value | 0.0148317 (rank : 12) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6P6P7, Q8BVW1 | Gene names | Parp6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 6 (EC 2.4.2.30) (PARP-6). | |||||
|
PARPT_MOUSE
|
||||||
| NC score | 0.080700 (rank : 11) | θ value | 1.81305 (rank : 16) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8C1B2, Q3UD50, Q8C032 | Gene names | Tiparp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TCDD-inducible poly [ADP-ribose] polymerase (EC 2.4.2.30). | |||||
|
TNKS1_HUMAN
|
||||||
| NC score | 0.072945 (rank : 12) | θ value | 0.00102713 (rank : 7) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O95271, O95272 | Gene names | TNKS, PARP5A, PARPL, TIN1, TINF1, TNKS1 | |||
|
Domain Architecture |

|
|||||
| Description | Tankyrase-1 (EC 2.4.2.30) (TANK1) (Tankyrase I) (TNKS-1) (TRF1- interacting ankyrin-related ADP-ribose polymerase). | |||||
|
DNL3_HUMAN
|
||||||
| NC score | 0.071019 (rank : 13) | θ value | θ > 10 (rank : 34) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P49916, Q16714 | Gene names | LIG3 | |||
|
Domain Architecture |
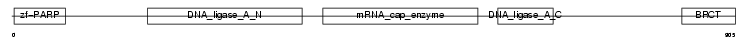
|
|||||
| Description | DNA ligase 3 (EC 6.5.1.1) (DNA ligase III) (Polydeoxyribonucleotide synthase [ATP] 3). | |||||
|
DNL3_MOUSE
|
||||||
| NC score | 0.069124 (rank : 14) | θ value | θ > 10 (rank : 35) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P97386, P97385 | Gene names | Lig3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA ligase 3 (EC 6.5.1.1) (DNA ligase III) (Polydeoxyribonucleotide synthase [ATP] 3). | |||||
|
TNKS2_HUMAN
|
||||||
| NC score | 0.068075 (rank : 15) | θ value | 0.00869519 (rank : 10) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9H2K2, Q9H8F2, Q9HAS4 | Gene names | TNKS2, PARP5B, TANK2, TNKL | |||
|
Domain Architecture |

|
|||||
| Description | Tankyrase-2 (EC 2.4.2.30) (TANK2) (Tankyrase II) (TNKS-2) (TRF1- interacting ankyrin-related ADP-ribose polymerase 2) (Tankyrase-like protein) (Tankyrase-related protein). | |||||
|
PAR16_MOUSE
|
||||||
| NC score | 0.067298 (rank : 16) | θ value | 1.81305 (rank : 15) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q7TMM8, Q3V031, Q8CC71 | Gene names | Parp16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 16 (EC 2.4.2.30) (PARP-16). | |||||
|
PARPT_HUMAN
|
||||||
| NC score | 0.064402 (rank : 17) | θ value | 2.36792 (rank : 19) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7Z3E1, Q68CY9, Q86VP4, Q9Y4P7 | Gene names | TIPARP, PARP7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TCDD-inducible poly [ADP-ribose] polymerase (EC 2.4.2.30) (Poly [ADP- ribose] polymerase 7) (PARP-7). | |||||
|
LHR2A_MOUSE
|
||||||
| NC score | 0.059303 (rank : 18) | θ value | θ > 10 (rank : 37) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | Q99KC8, Q3TTU2, Q6UN22, Q8BHA8, Q9CTV9 | Gene names | Loh11cr2a | |||
|
Domain Architecture |

|
|||||
| Description | Loss of heterozygosity 11 chromosomal region 2 gene A protein homolog. | |||||
|
PAR16_HUMAN
|
||||||
| NC score | 0.059217 (rank : 19) | θ value | 6.88961 (rank : 28) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N5Y8, Q6PK64, Q9NX03 | Gene names | PARP16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 16 (EC 2.4.2.30) (PARP-16). | |||||
|
LHR2A_HUMAN
|
||||||
| NC score | 0.055724 (rank : 20) | θ value | θ > 10 (rank : 36) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | O00534, Q6UN19, Q6UN20, Q9BVF8 | Gene names | LOH11CR2A, BCSC1 | |||
|
Domain Architecture |

|
|||||
| Description | Loss of heterozygosity 11 chromosomal region 2 gene A protein (Breast cancer suppressor candidate 1) (BCSC-1). | |||||
|
FLOT2_HUMAN
|
||||||
| NC score | 0.032699 (rank : 21) | θ value | 4.03905 (rank : 22) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q14254 | Gene names | FLOT2, ESA1, M17S1 | |||
|
Domain Architecture |

|
|||||
| Description | Flotillin-2 (Epidermal surface antigen) (ESA). | |||||
|
FLOT2_MOUSE
|
||||||
| NC score | 0.032659 (rank : 22) | θ value | 4.03905 (rank : 23) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q60634, Q6NS75 | Gene names | Flot2, Esa1, M17s1 | |||
|
Domain Architecture |

|
|||||
| Description | Flotillin-2 (Epidermal surface antigen) (ESA). | |||||
|
THUM1_HUMAN
|
||||||
| NC score | 0.026680 (rank : 23) | θ value | 2.36792 (rank : 20) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NXG2, Q9BWC3 | Gene names | THUMPD1 | |||
|
Domain Architecture |

|
|||||
| Description | THUMP domain-containing protein 1. | |||||
|
CP007_HUMAN
|
||||||
| NC score | 0.024840 (rank : 24) | θ value | 5.27518 (rank : 24) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y2B5 | Gene names | C16orf7, ATPBL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf7 (Protein ATP-BL). | |||||
|
JAD1A_HUMAN
|
||||||
| NC score | 0.021314 (rank : 25) | θ value | 0.813845 (rank : 13) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P29375 | Gene names | JARID1A, RBBP2, RBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1A (Retinoblastoma-binding protein 2) (RBBP-2). | |||||
|
CP007_MOUSE
|
||||||
| NC score | 0.019579 (rank : 26) | θ value | 6.88961 (rank : 27) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C190, Q6PFD1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf7 homolog (5-day ovary-specific transcript 1 protein). | |||||
|
CIP1_HUMAN
|
||||||
| NC score | 0.017094 (rank : 27) | θ value | 8.99809 (rank : 33) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NPC3 | Gene names | CCNB1IP1, C14orf18, HEI10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-B1-interacting protein 1 (EC 6.3.2.-) (Human enhancer of invasion 10) (E3 ubiquitin-protein ligase HEI10). | |||||
|
CAPS1_HUMAN
|
||||||
| NC score | 0.013384 (rank : 28) | θ value | 6.88961 (rank : 26) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ULU8, Q13339, Q6GQQ6, Q8N2Z5, Q8N3M7, Q8NFR0, Q96BC2 | Gene names | CADPS, CAPS, CAPS1, KIAA1121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-dependent secretion activator 1 (Calcium-dependent activator protein for secretion 1) (CAPS-1). | |||||
|
OSBP1_HUMAN
|
||||||
| NC score | 0.013252 (rank : 29) | θ value | 1.81305 (rank : 14) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P22059 | Gene names | OSBP, OSBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Oxysterol-binding protein 1. | |||||
|
CAPS1_MOUSE
|
||||||
| NC score | 0.012582 (rank : 30) | θ value | 8.99809 (rank : 32) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80TJ1, Q3TSP2, Q61374, Q6AXB4, Q6PGF0 | Gene names | Cadps, Caps, Caps1, Kiaa1121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-dependent secretion activator 1 (Calcium-dependent activator protein for secretion 1) (CAPS-1). | |||||
|
DDX46_MOUSE
|
||||||
| NC score | 0.009465 (rank : 31) | θ value | 2.36792 (rank : 18) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q569Z5, Q6ZQ42, Q8R0R6 | Gene names | Ddx46, Kiaa0801 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX46 (EC 3.6.1.-) (DEAD box protein 46). | |||||
|
DDX46_HUMAN
|
||||||
| NC score | 0.008296 (rank : 32) | θ value | 5.27518 (rank : 25) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7L014, O94894, Q96EI0, Q9Y658 | Gene names | DDX46, KIAA0801 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX46 (EC 3.6.1.-) (DEAD box protein 46) (PRP5 homolog). | |||||
|
ZNFX1_HUMAN
|
||||||
| NC score | 0.007930 (rank : 33) | θ value | 6.88961 (rank : 30) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9P2E3, Q9BQM7, Q9BQM8, Q9H8C1, Q9H9S2, Q9NUM1, Q9NWW1 | Gene names | ZNFX1, KIAA1404 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
CK5P2_HUMAN
|
||||||
| NC score | 0.007018 (rank : 34) | θ value | 2.36792 (rank : 17) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 1433 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96SN8, Q7Z3L4, Q7Z3U1, Q7Z7I6, Q9BSW0, Q9H6J6, Q9HCD9, Q9NV90, Q9UIW9 | Gene names | CDK5RAP2, CEP215, KIAA1633 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48) (Centrosome-associated protein 215). | |||||
|
MYO5C_HUMAN
|
||||||
| NC score | 0.004967 (rank : 35) | θ value | 3.0926 (rank : 21) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 1206 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NQX4 | Gene names | MYO5C | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5C (Myosin Vc). | |||||
|
41_HUMAN
|
||||||
| NC score | 0.003776 (rank : 36) | θ value | 8.99809 (rank : 31) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P11171, P11176, Q14245, Q5TB35, Q5VXN8, Q8IXV9, Q9Y578, Q9Y579 | Gene names | EPB41, E41P | |||
|
Domain Architecture |

|
|||||
| Description | Protein 4.1 (Band 4.1) (P4.1) (EPB4.1) (4.1R). | |||||
|
ZN396_HUMAN
|
||||||
| NC score | -0.000648 (rank : 37) | θ value | 6.88961 (rank : 29) | |||
| Query Neighborhood Hits | 33 | Target Neighborhood Hits | 703 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96N95, Q8NF98, Q8TD80 | Gene names | ZNF396, ZSCAN14 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 396 (Zinc finger and SCAN domain-containing protein 14). | |||||