Please be patient as the page loads
|
CP007_HUMAN
|
||||||
| SwissProt Accessions | Q9Y2B5 | Gene names | C16orf7, ATPBL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf7 (Protein ATP-BL). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CP007_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q9Y2B5 | Gene names | C16orf7, ATPBL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf7 (Protein ATP-BL). | |||||
|
CP007_MOUSE
|
||||||
| θ value | 1.20704e-137 (rank : 2) | NC score | 0.862119 (rank : 2) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8C190, Q6PFD1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf7 homolog (5-day ovary-specific transcript 1 protein). | |||||
|
CF182_MOUSE
|
||||||
| θ value | 0.279714 (rank : 3) | NC score | 0.074919 (rank : 6) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 551 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8VDS7, Q9CZE0, Q9D5A1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf182 homolog. | |||||
|
IL4RA_MOUSE
|
||||||
| θ value | 0.279714 (rank : 4) | NC score | 0.060377 (rank : 17) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P16382, O54690, Q60583, Q8CBW5 | Gene names | Il4r, Il4ra | |||
|
Domain Architecture |
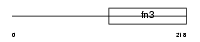
|
|||||
| Description | Interleukin-4 receptor alpha chain precursor (IL-4R-alpha) (CD124 antigen) [Contains: Soluble interleukin-4 receptor alpha chain (IL-4- binding protein) (IL4-BP)]. | |||||
|
CCHCR_HUMAN
|
||||||
| θ value | 0.365318 (rank : 5) | NC score | 0.068960 (rank : 9) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 717 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8TD31, Q9NRK8, Q9NWY9, Q9NXJ4, Q9NXK3, Q9Y6W1, Q9Y6W2 | Gene names | CCHCR1, C6orf18, HCR | |||
|
Domain Architecture |

|
|||||
| Description | Coiled-coil alpha-helical rod protein 1 (Alpha helical coiled-coil rod protein) (Putative gene 8 protein) (Pg8). | |||||
|
AMOL2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 6) | NC score | 0.081957 (rank : 4) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9Y2J4, Q53EP1, Q96F99, Q9UKB4 | Gene names | AMOTL2, KIAA0989 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 2 (Leman coiled-coil protein) (LCCP). | |||||
|
CAF1A_MOUSE
|
||||||
| θ value | 0.47712 (rank : 7) | NC score | 0.071636 (rank : 7) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 736 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9QWF0 | Gene names | Chaf1a, Caip150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromatin assembly factor 1 subunit A (CAF-1 subunit A) (Chromatin assembly factor I p150 subunit) (CAF-I 150 kDa subunit) (CAF-Ip150). | |||||
|
G45IP_MOUSE
|
||||||
| θ value | 0.47712 (rank : 8) | NC score | 0.100759 (rank : 3) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9CR59, Q8BT05 | Gene names | Gadd45gip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Growth arrest and DNA-damage-inducible proteins-interacting protein 1. | |||||
|
TRHY_HUMAN
|
||||||
| θ value | 0.47712 (rank : 9) | NC score | 0.058967 (rank : 19) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
XE7_HUMAN
|
||||||
| θ value | 0.62314 (rank : 10) | NC score | 0.078501 (rank : 5) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 738 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q02040, Q02832 | Gene names | CXYorf3, DXYS155E, XE7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-lymphocyte antigen precursor (B-lymphocyte surface antigen) (721P) (Protein XE7). | |||||
|
KIF3A_MOUSE
|
||||||
| θ value | 0.813845 (rank : 11) | NC score | 0.041491 (rank : 57) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 715 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P28741 | Gene names | Kif3a, Kif3 | |||
|
Domain Architecture |
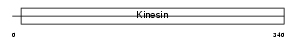
|
|||||
| Description | Kinesin-like protein KIF3A (Microtubule plus end-directed kinesin motor 3A). | |||||
|
MYPT1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 12) | NC score | 0.034426 (rank : 64) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 723 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O14974, Q86WU3, Q8NFR6, Q9BYH0 | Gene names | PPP1R12A, MBS, MYPT1 | |||
|
Domain Architecture |
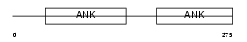
|
|||||
| Description | Protein phosphatase 1 regulatory subunit 12A (Myosin phosphatase- targeting subunit 1) (Myosin phosphatase target subunit 1) (Protein phosphatase myosin-binding subunit). | |||||
|
SEM6B_HUMAN
|
||||||
| θ value | 1.06291 (rank : 13) | NC score | 0.016307 (rank : 81) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H3T3, Q9NRK9 | Gene names | SEMA6B, SEMAZ | |||
|
Domain Architecture |
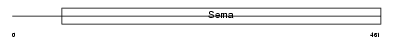
|
|||||
| Description | Semaphorin-6B precursor (Semaphorin Z) (Sema Z). | |||||
|
BUB1B_MOUSE
|
||||||
| θ value | 1.81305 (rank : 14) | NC score | 0.035568 (rank : 63) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Z1S0 | Gene names | Bub1b, Mad3l | |||
|
Domain Architecture |

|
|||||
| Description | Mitotic checkpoint serine/threonine-protein kinase BUB1 beta (EC 2.7.11.1) (MAD3/BUB1-related protein kinase) (Mitotic checkpoint kinase MAD3L). | |||||
|
CCD41_MOUSE
|
||||||
| θ value | 1.81305 (rank : 15) | NC score | 0.057223 (rank : 21) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9D5R3, Q3U7X7, Q3UX57, Q80VF0 | Gene names | Ccdc41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41. | |||||
|
GOGA4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 16) | NC score | 0.057175 (rank : 22) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
PEPL_HUMAN
|
||||||
| θ value | 1.81305 (rank : 17) | NC score | 0.058705 (rank : 20) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1141 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O60437, O60314, O60454 | Gene names | PPL, KIAA0568 | |||
|
Domain Architecture |
|
|||||
| Description | Periplakin (195 kDa cornified envelope precursor protein) (190 kDa paraneoplastic pemphigus antigen). | |||||
|
TBCD1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 18) | NC score | 0.039939 (rank : 60) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q60949, Q80TJ9, Q923F8 | Gene names | Tbc1d1, Kiaa1108, Tbc1 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 1. | |||||
|
CKAP4_HUMAN
|
||||||
| θ value | 2.36792 (rank : 19) | NC score | 0.052137 (rank : 31) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q07065, Q504S5, Q53ES6 | Gene names | CKAP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 4 (63 kDa membrane protein) (p63). | |||||
|
DESP_HUMAN
|
||||||
| θ value | 2.36792 (rank : 20) | NC score | 0.054400 (rank : 26) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P15924, O75993, Q14189, Q9UHN4 | Gene names | DSP | |||
|
Domain Architecture |

|
|||||
| Description | Desmoplakin (DP) (250/210 kDa paraneoplastic pemphigus antigen). | |||||
|
DYNA_MOUSE
|
||||||
| θ value | 2.36792 (rank : 21) | NC score | 0.049870 (rank : 40) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1084 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O08788 | Gene names | Dctn1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynactin-1 (150 kDa dynein-associated polypeptide) (DP-150) (DAP-150) (p150-glued). | |||||
|
TMF1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 22) | NC score | 0.054395 (rank : 27) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 987 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P82094 | Gene names | TMF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA element modulatory factor (TMF). | |||||
|
AATF_HUMAN
|
||||||
| θ value | 3.0926 (rank : 23) | NC score | 0.036859 (rank : 62) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NY61, Q9P0A4, Q9UNX5 | Gene names | AATF, CHE1, DED | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein AATF (Apoptosis-antagonizing transcription factor) (Rb-binding protein Che-1). | |||||
|
HOOK2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 24) | NC score | 0.063316 (rank : 11) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 714 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q96ED9, O60562 | Gene names | HOOK2 | |||
|
Domain Architecture |

|
|||||
| Description | Hook homolog 2 (h-hook2) (hHK2). | |||||
|
JIP4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 25) | NC score | 0.043089 (rank : 55) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 585 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O60271, O60905, Q3KQU8, Q3MKM7, Q86WC7, Q86WC8, Q8IZX7, Q96II0, Q9H811 | Gene names | SPAG9, HSS, KIAA0516, MAPK8IP4, SYD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 4 (JNK-interacting protein 4) (JIP-4) (JNK-associated leucine-zipper protein) (JLP) (Sperm-associated antigen 9) (Mitogen-activated protein kinase 8- interacting protein 4) (Human lung cancer protein 6) (HLC-6) (Proliferation-inducing protein 6) (Sperm-specific protein) (Sperm surface protein) (Protein highly expressed in testis) (PHET) (Sunday driver 1). | |||||
|
JIP4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 26) | NC score | 0.040608 (rank : 59) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q58A65, Q3UH77, Q3UHF0, Q58VQ4, Q5NC70, Q5NC78, Q6A057, Q6PAS3, Q8CJC2 | Gene names | Spag9, Jip4, Jsap2, Kiaa0516, Mapk8ip4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 4 (JNK-interacting protein 4) (JIP-4) (JNK-associated leucine-zipper protein) (JLP) (Sperm-associated antigen 9) (Mitogen-activated protein kinase 8- interacting protein 4) (JNK/SAPK-associated protein 2) (JSAP2). | |||||
|
MACF1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 27) | NC score | 0.045577 (rank : 49) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1341 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9UPN3, O75053, Q5VW20, Q8WXY2, Q9H540, Q9UKP0, Q9ULG9 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoforms 1/2/3/5 (Actin cross-linking family protein 7) (Macrophin-1) (Trabeculin-alpha) (620 kDa actin-binding protein) (ABP620). | |||||
|
MACF4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 28) | NC score | 0.046783 (rank : 44) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1321 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q96PK2, Q8WXY1 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoform 4. | |||||
|
MAK_MOUSE
|
||||||
| θ value | 3.0926 (rank : 29) | NC score | 0.003751 (rank : 92) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 873 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q04859 | Gene names | Mak, Rck | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase MAK (EC 2.7.11.22) (Male germ cell- associated kinase) (Protein kinase RCK). | |||||
|
MD1L1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 30) | NC score | 0.060945 (rank : 15) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9Y6D9, Q13312, Q75MI0, Q86UM4, Q9UNH0 | Gene names | MAD1L1, MAD1, TXBP181 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitotic spindle assembly checkpoint protein MAD1 (Mitotic arrest deficient-like protein 1) (MAD1-like 1) (Mitotic checkpoint MAD1 protein-homolog) (HsMAD1) (hMAD1) (Tax-binding protein 181). | |||||
|
TNIK_HUMAN
|
||||||
| θ value | 3.0926 (rank : 31) | NC score | 0.011956 (rank : 84) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1570 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UKE5, O60298, Q8WUY7, Q9UKD8, Q9UKD9, Q9UKE0, Q9UKE1, Q9UKE2, Q9UKE3, Q9UKE4 | Gene names | TNIK, KIAA0551 | |||
|
Domain Architecture |

|
|||||
| Description | TRAF2 and NCK-interacting protein kinase (EC 2.7.11.1). | |||||
|
TRI37_HUMAN
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.023260 (rank : 77) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O94972, Q7Z3E6, Q8IYF7, Q8WYF7 | Gene names | TRIM37, KIAA0898, MUL, POB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 37 (Mulibrey nanism protein). | |||||
|
UACA_MOUSE
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.045719 (rank : 46) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8CGB3, Q69ZG3, Q7TN77, Q8BJC8 | Gene names | Uaca, Kiaa1561 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uveal autoantigen with coiled-coil domains and ankyrin repeats protein (Nucling) (Nuclear membrane-binding protein). | |||||
|
UT14C_HUMAN
|
||||||
| θ value | 3.0926 (rank : 34) | NC score | 0.045030 (rank : 50) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5TAP6, Q5FWG3, Q92555 | Gene names | UTP14C, KIAA0266 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U3 small nucleolar RNA-associated protein 14 homolog C. | |||||
|
CAF1A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | 0.069500 (rank : 8) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q13111, Q7Z7K3, Q9UJY8 | Gene names | CHAF1A, CAF, CAF1P150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromatin assembly factor 1 subunit A (CAF-1 subunit A) (Chromatin assembly factor I p150 subunit) (CAF-I 150 kDa subunit) (CAF-Ip150). | |||||
|
ENL_HUMAN
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.027626 (rank : 68) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
HOOK2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.060495 (rank : 16) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 736 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q7TMK6, Q8BY47, Q8R347, Q8VCN4, Q99LU2 | Gene names | Hook2 | |||
|
Domain Architecture |
|
|||||
| Description | Hook homolog 2. | |||||
|
IF3A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 38) | NC score | 0.045648 (rank : 47) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 871 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P23116, Q60697, Q62162 | Gene names | Eif3s10, Csma, Eif3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3 p185) (eIF3a) (p162 protein) (Centrosomin). | |||||
|
KIF3A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 39) | NC score | 0.037273 (rank : 61) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9Y496, Q86XE9, Q9Y6V4 | Gene names | KIF3A, KIF3 | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin-like protein KIF3A (Microtubule plus end-directed kinesin motor 3A). | |||||
|
LIPA2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 40) | NC score | 0.050357 (rank : 38) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1044 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O75334 | Gene names | PPFIA2 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-2) (PTPRF-interacting protein alpha-2). | |||||
|
LRC45_HUMAN
|
||||||
| θ value | 4.03905 (rank : 41) | NC score | 0.067601 (rank : 10) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 933 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q96CN5 | Gene names | LRRC45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 45. | |||||
|
LRC45_MOUSE
|
||||||
| θ value | 4.03905 (rank : 42) | NC score | 0.061783 (rank : 12) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q8CIM1, Q3U5Z2 | Gene names | Lrrc45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 45. | |||||
|
RAI14_MOUSE
|
||||||
| θ value | 4.03905 (rank : 43) | NC score | 0.041241 (rank : 58) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1505 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9EP71, Q3URT3, Q6ZPT6 | Gene names | Rai14, Kiaa1334, Norpeg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankycorbin (Ankyrin repeat and coiled-coil structure-containing protein) (Retinoic acid-induced protein 14) (Novel retinal pigment epithelial cell protein) (p125). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 4.03905 (rank : 44) | NC score | 0.050910 (rank : 36) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
UBP34_HUMAN
|
||||||
| θ value | 4.03905 (rank : 45) | NC score | 0.026379 (rank : 69) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q70CQ2, O60316, O94834, Q3B777, Q6P6C9, Q7L8P6, Q8N3T9, Q8TBW2, Q9UGA1 | Gene names | USP34, KIAA0570, KIAA0729 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 34 (EC 3.1.2.15) (Ubiquitin thioesterase 34) (Ubiquitin-specific-processing protease 34) (Deubiquitinating enzyme 34). | |||||
|
UBP49_HUMAN
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.023432 (rank : 75) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q70CQ1, Q96CK4 | Gene names | USP49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 49 (EC 3.1.2.15) (Ubiquitin thioesterase 49) (Ubiquitin-specific-processing protease 49) (Deubiquitinating enzyme 49). | |||||
|
BCL9_MOUSE
|
||||||
| θ value | 5.27518 (rank : 47) | NC score | 0.022563 (rank : 78) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9D219, Q67FX9, Q8BUJ8, Q8VE74 | Gene names | Bcl9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9). | |||||
|
DAB2P_HUMAN
|
||||||
| θ value | 5.27518 (rank : 48) | NC score | 0.029314 (rank : 66) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q5VWQ8, Q8TDL2, Q96SE1, Q9C0C0 | Gene names | DAB2IP, AF9Q34, AIP1, KIAA1743 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Disabled homolog 2-interacting protein (DAB2-interacting protein) (DAB2 interaction protein) (ASK-interacting protein 1). | |||||
|
DAB2P_MOUSE
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | 0.028876 (rank : 67) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q3UHC7, Q3TPD5, Q3UH44, Q6JTV1, Q80T97 | Gene names | Dab2ip, Kiaa1743 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Disabled homolog 2-interacting protein (DAB2-interacting protein). | |||||
|
GRAP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 50) | NC score | 0.045627 (rank : 48) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q4V328, Q4V327, Q4V330, Q5HYG1, Q6N046, Q96DH8, Q9NQ43, Q9ULQ3 | Gene names | GRIPAP1, KIAA1167 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GRIP1-associated protein 1 (GRASP-1). | |||||
|
HMMR_HUMAN
|
||||||
| θ value | 5.27518 (rank : 51) | NC score | 0.049669 (rank : 41) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1082 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O75330, Q92767 | Gene names | HMMR, IHABP, RHAMM | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronan mediated motility receptor (Intracellular hyaluronic acid- binding protein) (Receptor for hyaluronan-mediated motility) (CD168 antigen). | |||||
|
LIPA2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 52) | NC score | 0.051511 (rank : 33) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1282 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8BSS9, Q8BN73 | Gene names | Ppfia2 | |||
|
Domain Architecture |
|
|||||
| Description | Liprin-alpha-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-2) (PTPRF-interacting protein alpha-2). | |||||
|
PARP2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.024840 (rank : 72) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UGN5, Q8TEU4, Q9NUV2, Q9UMR4, Q9Y6C8 | Gene names | PARP2, ADPRT2, ADPRTL2 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 2 (EC 2.4.2.30) (PARP-2) (NAD(+) ADP- ribosyltransferase 2) (Poly[ADP-ribose] synthetase 2) (pADPRT-2) (hPARP-2). | |||||
|
PEPL_MOUSE
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | 0.056655 (rank : 23) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9R269, O70231, Q9CUT1, Q9JLZ7 | Gene names | Ppl | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin. | |||||
|
RIMB1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 55) | NC score | 0.056346 (rank : 24) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
RNF25_HUMAN
|
||||||
| θ value | 5.27518 (rank : 56) | NC score | 0.023291 (rank : 76) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96BH1, Q9H874 | Gene names | RNF25 | |||
|
Domain Architecture |
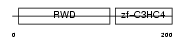
|
|||||
| Description | RING finger protein 25 (EC 6.3.2.-). | |||||
|
RRBP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 57) | NC score | 0.043370 (rank : 53) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
SEPT8_MOUSE
|
||||||
| θ value | 5.27518 (rank : 58) | NC score | 0.016565 (rank : 80) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8CHH9, Q80YC7, Q9ESF7 | Gene names | Sept8, Kiaa0202 | |||
|
Domain Architecture |
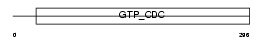
|
|||||
| Description | Septin-8. | |||||
|
UBP34_MOUSE
|
||||||
| θ value | 5.27518 (rank : 59) | NC score | 0.025177 (rank : 70) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6ZQ93, Q3UPN0, Q6P563, Q7TMJ6, Q8CCH0 | Gene names | Usp34, Kiaa0570 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 34 (EC 3.1.2.15) (Ubiquitin thioesterase 34) (Ubiquitin-specific-processing protease 34) (Deubiquitinating enzyme 34). | |||||
|
FOXN1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 60) | NC score | 0.008736 (rank : 88) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 300 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61575 | Gene names | Foxn1, Fkh19, Hfh11, Whn | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein N1 (Transcription factor winged-helix nude) (Hepatocyte nuclear factor 3 forkhead homolog 11) (HNF-3/forkhead homolog 11) (HFH-11). | |||||
|
GOGA4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | 0.054044 (rank : 28) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
INCE_MOUSE
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.048094 (rank : 43) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9WU62, Q7TN28, Q8BGN4, Q8CGI4 | Gene names | Incenp | |||
|
Domain Architecture |

|
|||||
| Description | Inner centromere protein. | |||||
|
MERL_MOUSE
|
||||||
| θ value | 6.88961 (rank : 63) | NC score | 0.024989 (rank : 71) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 524 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P46662, Q8BR03 | Gene names | Nf2, Nf-2 | |||
|
Domain Architecture |

|
|||||
| Description | Merlin (Moesin-ezrin-radixin-like protein) (Neurofibromin-2) (Schwannomin). | |||||
|
PB1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 64) | NC score | 0.018183 (rank : 79) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86U86, Q9H2T3, Q9H2T4, Q9H2T5, Q9H301, Q9H314 | Gene names | PB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein polybromo-1 (hPB1) (Polybromo-1D) (BRG1-associated factor 180) (BAF180). | |||||
|
PLEC1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 65) | NC score | 0.044245 (rank : 52) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1326 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q15149, Q15148, Q16640 | Gene names | PLEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Plectin-1 (PLTN) (PCN) (Hemidesmosomal protein 1) (HD1). | |||||
|
PRRT3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 66) | NC score | 0.014917 (rank : 82) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
SMCA2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.011106 (rank : 85) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P51531 | Gene names | SMARCA2, BRM, SNF2A, SNF2L2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L2 (EC 3.6.1.-) (ATP- dependent helicase SMARCA2) (SNF2-alpha) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin subfamily A member 2) (hBRM). | |||||
|
UACA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.041599 (rank : 56) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1622 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9BZF9, Q8N3B8, Q9HCL1, Q9NWC6 | Gene names | UACA, KIAA1561 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uveal autoantigen with coiled-coil domains and ankyrin repeats protein. | |||||
|
WDR67_HUMAN
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.048953 (rank : 42) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 782 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q96DN5, Q3MIR6, Q8TBP9 | Gene names | WDR67 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 67. | |||||
|
AMOT_HUMAN
|
||||||
| θ value | 8.99809 (rank : 70) | NC score | 0.043240 (rank : 54) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q4VCS5, Q504X5, Q9HD27, Q9UPT1 | Gene names | AMOT, KIAA1071 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin. | |||||
|
ARHGC_HUMAN
|
||||||
| θ value | 8.99809 (rank : 71) | NC score | 0.008881 (rank : 87) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9NZN5, O15086 | Gene names | ARHGEF12, KIAA0382, LARG | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 12 (Leukemia-associated RhoGEF). | |||||
|
BCL9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.024774 (rank : 73) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
CCHCR_MOUSE
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.059096 (rank : 18) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 932 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8K2I2 | Gene names | Cchcr1, Hcr | |||
|
Domain Architecture |

|
|||||
| Description | Coiled-coil alpha-helical rod protein 1 (Alpha helical coiled-coil rod protein). | |||||
|
CE290_HUMAN
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.046517 (rank : 45) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
CENPE_HUMAN
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.044810 (rank : 51) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
CREB3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.024435 (rank : 74) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O43889, O14671, O14919, Q96GK8, Q9H2W3, Q9UE77 | Gene names | CREB3, LZIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-responsive element-binding protein 3 (Luman protein) (Transcription factor LZIP-alpha). | |||||
|
IF4G1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.012545 (rank : 83) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q04637, O43177, O95066, Q5HYG0, Q6ZN21, Q8N102 | Gene names | EIF4G1, EIF4G, EIF4GI | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1) (p220). | |||||
|
INCE_HUMAN
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.052488 (rank : 30) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 978 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9NQS7, Q5Y192 | Gene names | INCENP | |||
|
Domain Architecture |

|
|||||
| Description | Inner centromere protein. | |||||
|
NSD1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.009737 (rank : 86) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
OBSCN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.002890 (rank : 93) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
PAX9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 81) | NC score | 0.008386 (rank : 89) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55771, Q99582, Q9UQR4 | Gene names | PAX9 | |||
|
Domain Architecture |
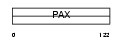
|
|||||
| Description | Paired box protein Pax-9. | |||||
|
PLXB1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.008114 (rank : 90) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O43157, Q6NY20, Q9UIV7, Q9UJ92, Q9UJ93 | Gene names | PLXNB1, KIAA0407, SEP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B1 precursor (Semaphorin receptor SEP). | |||||
|
PRD16_HUMAN
|
||||||
| θ value | 8.99809 (rank : 83) | NC score | 0.001523 (rank : 94) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 832 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9HAZ2, Q8WYJ9, Q9C0I8 | Gene names | PRDM16, KIAA1675, MEL1, PFM13 | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 16 (PR domain-containing protein 16) (Transcription factor MEL1). | |||||
|
TITIN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.006605 (rank : 91) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
UT14B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.029651 (rank : 65) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6EJB6, Q6EJB5, Q8BL60 | Gene names | Utp14b, Jsd | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U3 small nucleolar RNA-associated protein 14 homolog B (Juvenile spermatogonial depletion protein). | |||||
|
AKTS1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.061202 (rank : 13) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | Q96B36, Q96BI4, Q96IK7, Q96NG2, Q9BWR5 | Gene names | AKT1S1, PRAS40 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich AKT1 substrate 1 (40 kDa proline-rich AKT substrate). | |||||
|
AMOL2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.051420 (rank : 34) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8K371, Q3TPM1, Q7TPE4, Q8BP84, Q8BS08, Q9QUS0 | Gene names | Amotl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 2. | |||||
|
CCD46_MOUSE
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.050665 (rank : 37) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q5PR68, Q80ZN6, Q99JS4, Q9D9T1 | Gene names | Ccdc46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled coil domain-containing protein 46. | |||||
|
CP250_HUMAN
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.050321 (rank : 39) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1805 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9BV73, O14812, O60588, Q9H450 | Gene names | CEP250, CEP2, CNAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1). | |||||
|
MD1L1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.053251 (rank : 29) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1001 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9WTX8, Q9WTX9 | Gene names | Mad1l1, Mad1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitotic spindle assembly checkpoint protein MAD1 (Mitotic arrest deficient-like protein 1) (MAD1-like 1). | |||||
|
RABX5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.051304 (rank : 35) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UJ41 | Gene names | RABGEF1, RABEX5 | |||
|
Domain Architecture |

|
|||||
| Description | Rab5 GDP/GTP exchange factor (Rabex-5) (Rabaptin-5-associated exchange factor for Rab5). | |||||
|
RABX5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.061199 (rank : 14) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JM13, Q3UIW0 | Gene names | Rabgef1, Rabex5 | |||
|
Domain Architecture |

|
|||||
| Description | Rab5 GDP/GTP exchange factor (Rabex-5). | |||||
|
RIN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.051542 (rank : 32) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | Q13671, O15010, Q00427, Q96CC8 | Gene names | RIN1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 1 (Ras interaction/interference protein 1) (Ras inhibitor JC99). | |||||
|
RIN1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.055948 (rank : 25) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | Q921Q7 | Gene names | Rin1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 1 (Ras interaction/interference protein 1). | |||||
|
CP007_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q9Y2B5 | Gene names | C16orf7, ATPBL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf7 (Protein ATP-BL). | |||||
|
CP007_MOUSE
|
||||||
| NC score | 0.862119 (rank : 2) | θ value | 1.20704e-137 (rank : 2) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8C190, Q6PFD1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf7 homolog (5-day ovary-specific transcript 1 protein). | |||||
|
G45IP_MOUSE
|
||||||
| NC score | 0.100759 (rank : 3) | θ value | 0.47712 (rank : 8) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9CR59, Q8BT05 | Gene names | Gadd45gip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Growth arrest and DNA-damage-inducible proteins-interacting protein 1. | |||||
|
AMOL2_HUMAN
|
||||||
| NC score | 0.081957 (rank : 4) | θ value | 0.47712 (rank : 6) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9Y2J4, Q53EP1, Q96F99, Q9UKB4 | Gene names | AMOTL2, KIAA0989 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 2 (Leman coiled-coil protein) (LCCP). | |||||
|
XE7_HUMAN
|
||||||
| NC score | 0.078501 (rank : 5) | θ value | 0.62314 (rank : 10) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 738 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q02040, Q02832 | Gene names | CXYorf3, DXYS155E, XE7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-lymphocyte antigen precursor (B-lymphocyte surface antigen) (721P) (Protein XE7). | |||||
|
CF182_MOUSE
|
||||||
| NC score | 0.074919 (rank : 6) | θ value | 0.279714 (rank : 3) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 551 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8VDS7, Q9CZE0, Q9D5A1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf182 homolog. | |||||
|
CAF1A_MOUSE
|
||||||
| NC score | 0.071636 (rank : 7) | θ value | 0.47712 (rank : 7) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 736 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9QWF0 | Gene names | Chaf1a, Caip150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromatin assembly factor 1 subunit A (CAF-1 subunit A) (Chromatin assembly factor I p150 subunit) (CAF-I 150 kDa subunit) (CAF-Ip150). | |||||
|
CAF1A_HUMAN
|
||||||
| NC score | 0.069500 (rank : 8) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q13111, Q7Z7K3, Q9UJY8 | Gene names | CHAF1A, CAF, CAF1P150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromatin assembly factor 1 subunit A (CAF-1 subunit A) (Chromatin assembly factor I p150 subunit) (CAF-I 150 kDa subunit) (CAF-Ip150). | |||||
|
CCHCR_HUMAN
|
||||||
| NC score | 0.068960 (rank : 9) | θ value | 0.365318 (rank : 5) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 717 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8TD31, Q9NRK8, Q9NWY9, Q9NXJ4, Q9NXK3, Q9Y6W1, Q9Y6W2 | Gene names | CCHCR1, C6orf18, HCR | |||
|
Domain Architecture |

|
|||||
| Description | Coiled-coil alpha-helical rod protein 1 (Alpha helical coiled-coil rod protein) (Putative gene 8 protein) (Pg8). | |||||
|
LRC45_HUMAN
|
||||||
| NC score | 0.067601 (rank : 10) | θ value | 4.03905 (rank : 41) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 933 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q96CN5 | Gene names | LRRC45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 45. | |||||
|
HOOK2_HUMAN
|
||||||
| NC score | 0.063316 (rank : 11) | θ value | 3.0926 (rank : 24) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 714 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q96ED9, O60562 | Gene names | HOOK2 | |||
|
Domain Architecture |

|
|||||
| Description | Hook homolog 2 (h-hook2) (hHK2). | |||||
|
LRC45_MOUSE
|
||||||
| NC score | 0.061783 (rank : 12) | θ value | 4.03905 (rank : 42) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q8CIM1, Q3U5Z2 | Gene names | Lrrc45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 45. | |||||
|
AKTS1_HUMAN
|
||||||
| NC score | 0.061202 (rank : 13) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | Q96B36, Q96BI4, Q96IK7, Q96NG2, Q9BWR5 | Gene names | AKT1S1, PRAS40 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich AKT1 substrate 1 (40 kDa proline-rich AKT substrate). | |||||
|
RABX5_MOUSE
|
||||||
| NC score | 0.061199 (rank : 14) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JM13, Q3UIW0 | Gene names | Rabgef1, Rabex5 | |||
|
Domain Architecture |

|
|||||
| Description | Rab5 GDP/GTP exchange factor (Rabex-5). | |||||
|
MD1L1_HUMAN
|
||||||
| NC score | 0.060945 (rank : 15) | θ value | 3.0926 (rank : 30) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9Y6D9, Q13312, Q75MI0, Q86UM4, Q9UNH0 | Gene names | MAD1L1, MAD1, TXBP181 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitotic spindle assembly checkpoint protein MAD1 (Mitotic arrest deficient-like protein 1) (MAD1-like 1) (Mitotic checkpoint MAD1 protein-homolog) (HsMAD1) (hMAD1) (Tax-binding protein 181). | |||||
|
HOOK2_MOUSE
|
||||||
| NC score | 0.060495 (rank : 16) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 736 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q7TMK6, Q8BY47, Q8R347, Q8VCN4, Q99LU2 | Gene names | Hook2 | |||
|
Domain Architecture |

|
|||||
| Description | Hook homolog 2. | |||||
|
IL4RA_MOUSE
|
||||||
| NC score | 0.060377 (rank : 17) | θ value | 0.279714 (rank : 4) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P16382, O54690, Q60583, Q8CBW5 | Gene names | Il4r, Il4ra | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-4 receptor alpha chain precursor (IL-4R-alpha) (CD124 antigen) [Contains: Soluble interleukin-4 receptor alpha chain (IL-4- binding protein) (IL4-BP)]. | |||||
|
CCHCR_MOUSE
|
||||||
| NC score | 0.059096 (rank : 18) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 932 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8K2I2 | Gene names | Cchcr1, Hcr | |||
|
Domain Architecture |

|
|||||
| Description | Coiled-coil alpha-helical rod protein 1 (Alpha helical coiled-coil rod protein). | |||||
|
TRHY_HUMAN
|
||||||
| NC score | 0.058967 (rank : 19) | θ value | 0.47712 (rank : 9) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
PEPL_HUMAN
|
||||||
| NC score | 0.058705 (rank : 20) | θ value | 1.81305 (rank : 17) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1141 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O60437, O60314, O60454 | Gene names | PPL, KIAA0568 | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin (195 kDa cornified envelope precursor protein) (190 kDa paraneoplastic pemphigus antigen). | |||||
|
CCD41_MOUSE
|
||||||
| NC score | 0.057223 (rank : 21) | θ value | 1.81305 (rank : 15) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9D5R3, Q3U7X7, Q3UX57, Q80VF0 | Gene names | Ccdc41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41. | |||||
|
GOGA4_MOUSE
|
||||||
| NC score | 0.057175 (rank : 22) | θ value | 1.81305 (rank : 16) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
PEPL_MOUSE
|
||||||
| NC score | 0.056655 (rank : 23) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9R269, O70231, Q9CUT1, Q9JLZ7 | Gene names | Ppl | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin. | |||||
|
RIMB1_MOUSE
|
||||||
| NC score | 0.056346 (rank : 24) | θ value | 5.27518 (rank : 55) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
RIN1_MOUSE
|
||||||
| NC score | 0.055948 (rank : 25) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | Q921Q7 | Gene names | Rin1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 1 (Ras interaction/interference protein 1). | |||||
|
DESP_HUMAN
|
||||||
| NC score | 0.054400 (rank : 26) | θ value | 2.36792 (rank : 20) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P15924, O75993, Q14189, Q9UHN4 | Gene names | DSP | |||
|
Domain Architecture |

|
|||||
| Description | Desmoplakin (DP) (250/210 kDa paraneoplastic pemphigus antigen). | |||||
|
TMF1_HUMAN
|
||||||
| NC score | 0.054395 (rank : 27) | θ value | 2.36792 (rank : 22) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 987 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P82094 | Gene names | TMF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA element modulatory factor (TMF). | |||||
|
GOGA4_HUMAN
|
||||||
| NC score | 0.054044 (rank : 28) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
MD1L1_MOUSE
|
||||||
| NC score | 0.053251 (rank : 29) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1001 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9WTX8, Q9WTX9 | Gene names | Mad1l1, Mad1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitotic spindle assembly checkpoint protein MAD1 (Mitotic arrest deficient-like protein 1) (MAD1-like 1). | |||||
|
INCE_HUMAN
|
||||||
| NC score | 0.052488 (rank : 30) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 978 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9NQS7, Q5Y192 | Gene names | INCENP | |||
|
Domain Architecture |

|
|||||
| Description | Inner centromere protein. | |||||
|
CKAP4_HUMAN
|
||||||
| NC score | 0.052137 (rank : 31) | θ value | 2.36792 (rank : 19) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q07065, Q504S5, Q53ES6 | Gene names | CKAP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 4 (63 kDa membrane protein) (p63). | |||||
|
RIN1_HUMAN
|
||||||
| NC score | 0.051542 (rank : 32) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | Q13671, O15010, Q00427, Q96CC8 | Gene names | RIN1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 1 (Ras interaction/interference protein 1) (Ras inhibitor JC99). | |||||
|
LIPA2_MOUSE
|
||||||
| NC score | 0.051511 (rank : 33) | θ value | 5.27518 (rank : 52) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1282 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8BSS9, Q8BN73 | Gene names | Ppfia2 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-2) (PTPRF-interacting protein alpha-2). | |||||
|
AMOL2_MOUSE
|
||||||
| NC score | 0.051420 (rank : 34) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8K371, Q3TPM1, Q7TPE4, Q8BP84, Q8BS08, Q9QUS0 | Gene names | Amotl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 2. | |||||
|
RABX5_HUMAN
|
||||||
| NC score | 0.051304 (rank : 35) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UJ41 | Gene names | RABGEF1, RABEX5 | |||
|
Domain Architecture |

|
|||||
| Description | Rab5 GDP/GTP exchange factor (Rabex-5) (Rabaptin-5-associated exchange factor for Rab5). | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.050910 (rank : 36) | θ value | 4.03905 (rank : 44) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
CCD46_MOUSE
|
||||||
| NC score | 0.050665 (rank : 37) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q5PR68, Q80ZN6, Q99JS4, Q9D9T1 | Gene names | Ccdc46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled coil domain-containing protein 46. | |||||
|
LIPA2_HUMAN
|
||||||
| NC score | 0.050357 (rank : 38) | θ value | 4.03905 (rank : 40) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1044 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O75334 | Gene names | PPFIA2 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-2) (PTPRF-interacting protein alpha-2). | |||||
|
CP250_HUMAN
|
||||||
| NC score | 0.050321 (rank : 39) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1805 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9BV73, O14812, O60588, Q9H450 | Gene names | CEP250, CEP2, CNAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1). | |||||
|
DYNA_MOUSE
|
||||||
| NC score | 0.049870 (rank : 40) | θ value | 2.36792 (rank : 21) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1084 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O08788 | Gene names | Dctn1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynactin-1 (150 kDa dynein-associated polypeptide) (DP-150) (DAP-150) (p150-glued). | |||||
|
HMMR_HUMAN
|
||||||
| NC score | 0.049669 (rank : 41) | θ value | 5.27518 (rank : 51) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1082 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O75330, Q92767 | Gene names | HMMR, IHABP, RHAMM | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronan mediated motility receptor (Intracellular hyaluronic acid- binding protein) (Receptor for hyaluronan-mediated motility) (CD168 antigen). | |||||
|
WDR67_HUMAN
|
||||||
| NC score | 0.048953 (rank : 42) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 782 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q96DN5, Q3MIR6, Q8TBP9 | Gene names | WDR67 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 67. | |||||
|
INCE_MOUSE
|
||||||
| NC score | 0.048094 (rank : 43) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9WU62, Q7TN28, Q8BGN4, Q8CGI4 | Gene names | Incenp | |||
|
Domain Architecture |

|
|||||
| Description | Inner centromere protein. | |||||
|
MACF4_HUMAN
|
||||||
| NC score | 0.046783 (rank : 44) | θ value | 3.0926 (rank : 28) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1321 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q96PK2, Q8WXY1 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoform 4. | |||||
|
CE290_HUMAN
|
||||||
| NC score | 0.046517 (rank : 45) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
UACA_MOUSE
|
||||||
| NC score | 0.045719 (rank : 46) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8CGB3, Q69ZG3, Q7TN77, Q8BJC8 | Gene names | Uaca, Kiaa1561 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uveal autoantigen with coiled-coil domains and ankyrin repeats protein (Nucling) (Nuclear membrane-binding protein). | |||||
|
IF3A_MOUSE
|
||||||
| NC score | 0.045648 (rank : 47) | θ value | 4.03905 (rank : 38) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 871 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P23116, Q60697, Q62162 | Gene names | Eif3s10, Csma, Eif3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3 p185) (eIF3a) (p162 protein) (Centrosomin). | |||||
|
GRAP1_HUMAN
|
||||||
| NC score | 0.045627 (rank : 48) | θ value | 5.27518 (rank : 50) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q4V328, Q4V327, Q4V330, Q5HYG1, Q6N046, Q96DH8, Q9NQ43, Q9ULQ3 | Gene names | GRIPAP1, KIAA1167 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GRIP1-associated protein 1 (GRASP-1). | |||||
|
MACF1_HUMAN
|
||||||
| NC score | 0.045577 (rank : 49) | θ value | 3.0926 (rank : 27) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1341 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9UPN3, O75053, Q5VW20, Q8WXY2, Q9H540, Q9UKP0, Q9ULG9 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoforms 1/2/3/5 (Actin cross-linking family protein 7) (Macrophin-1) (Trabeculin-alpha) (620 kDa actin-binding protein) (ABP620). | |||||
|
UT14C_HUMAN
|
||||||
| NC score | 0.045030 (rank : 50) | θ value | 3.0926 (rank : 34) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5TAP6, Q5FWG3, Q92555 | Gene names | UTP14C, KIAA0266 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U3 small nucleolar RNA-associated protein 14 homolog C. | |||||
|
CENPE_HUMAN
|
||||||
| NC score | 0.044810 (rank : 51) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
PLEC1_HUMAN
|
||||||
| NC score | 0.044245 (rank : 52) | θ value | 6.88961 (rank : 65) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1326 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q15149, Q15148, Q16640 | Gene names | PLEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Plectin-1 (PLTN) (PCN) (Hemidesmosomal protein 1) (HD1). | |||||
|
RRBP1_MOUSE
|
||||||
| NC score | 0.043370 (rank : 53) | θ value | 5.27518 (rank : 57) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
AMOT_HUMAN
|
||||||
| NC score | 0.043240 (rank : 54) | θ value | 8.99809 (rank : 70) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q4VCS5, Q504X5, Q9HD27, Q9UPT1 | Gene names | AMOT, KIAA1071 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin. | |||||
|
JIP4_HUMAN
|
||||||
| NC score | 0.043089 (rank : 55) | θ value | 3.0926 (rank : 25) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 585 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O60271, O60905, Q3KQU8, Q3MKM7, Q86WC7, Q86WC8, Q8IZX7, Q96II0, Q9H811 | Gene names | SPAG9, HSS, KIAA0516, MAPK8IP4, SYD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 4 (JNK-interacting protein 4) (JIP-4) (JNK-associated leucine-zipper protein) (JLP) (Sperm-associated antigen 9) (Mitogen-activated protein kinase 8- interacting protein 4) (Human lung cancer protein 6) (HLC-6) (Proliferation-inducing protein 6) (Sperm-specific protein) (Sperm surface protein) (Protein highly expressed in testis) (PHET) (Sunday driver 1). | |||||
|
UACA_HUMAN
|
||||||
| NC score | 0.041599 (rank : 56) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1622 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9BZF9, Q8N3B8, Q9HCL1, Q9NWC6 | Gene names | UACA, KIAA1561 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uveal autoantigen with coiled-coil domains and ankyrin repeats protein. | |||||
|
KIF3A_MOUSE
|
||||||
| NC score | 0.041491 (rank : 57) | θ value | 0.813845 (rank : 11) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 715 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P28741 | Gene names | Kif3a, Kif3 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF3A (Microtubule plus end-directed kinesin motor 3A). | |||||
|
RAI14_MOUSE
|
||||||
| NC score | 0.041241 (rank : 58) | θ value | 4.03905 (rank : 43) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1505 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9EP71, Q3URT3, Q6ZPT6 | Gene names | Rai14, Kiaa1334, Norpeg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankycorbin (Ankyrin repeat and coiled-coil structure-containing protein) (Retinoic acid-induced protein 14) (Novel retinal pigment epithelial cell protein) (p125). | |||||
|
JIP4_MOUSE
|
||||||
| NC score | 0.040608 (rank : 59) | θ value | 3.0926 (rank : 26) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q58A65, Q3UH77, Q3UHF0, Q58VQ4, Q5NC70, Q5NC78, Q6A057, Q6PAS3, Q8CJC2 | Gene names | Spag9, Jip4, Jsap2, Kiaa0516, Mapk8ip4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 4 (JNK-interacting protein 4) (JIP-4) (JNK-associated leucine-zipper protein) (JLP) (Sperm-associated antigen 9) (Mitogen-activated protein kinase 8- interacting protein 4) (JNK/SAPK-associated protein 2) (JSAP2). | |||||
|
TBCD1_MOUSE
|
||||||
| NC score | 0.039939 (rank : 60) | θ value | 1.81305 (rank : 18) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q60949, Q80TJ9, Q923F8 | Gene names | Tbc1d1, Kiaa1108, Tbc1 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 1. | |||||
|
KIF3A_HUMAN
|
||||||
| NC score | 0.037273 (rank : 61) | θ value | 4.03905 (rank : 39) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9Y496, Q86XE9, Q9Y6V4 | Gene names | KIF3A, KIF3 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF3A (Microtubule plus end-directed kinesin motor 3A). | |||||
|
AATF_HUMAN
|
||||||
| NC score | 0.036859 (rank : 62) | θ value | 3.0926 (rank : 23) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NY61, Q9P0A4, Q9UNX5 | Gene names | AATF, CHE1, DED | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein AATF (Apoptosis-antagonizing transcription factor) (Rb-binding protein Che-1). | |||||
|
BUB1B_MOUSE
|
||||||
| NC score | 0.035568 (rank : 63) | θ value | 1.81305 (rank : 14) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Z1S0 | Gene names | Bub1b, Mad3l | |||
|
Domain Architecture |

|
|||||
| Description | Mitotic checkpoint serine/threonine-protein kinase BUB1 beta (EC 2.7.11.1) (MAD3/BUB1-related protein kinase) (Mitotic checkpoint kinase MAD3L). | |||||
|
MYPT1_HUMAN
|
||||||
| NC score | 0.034426 (rank : 64) | θ value | 1.06291 (rank : 12) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 723 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O14974, Q86WU3, Q8NFR6, Q9BYH0 | Gene names | PPP1R12A, MBS, MYPT1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein phosphatase 1 regulatory subunit 12A (Myosin phosphatase- targeting subunit 1) (Myosin phosphatase target subunit 1) (Protein phosphatase myosin-binding subunit). | |||||
|
UT14B_MOUSE
|
||||||
| NC score | 0.029651 (rank : 65) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6EJB6, Q6EJB5, Q8BL60 | Gene names | Utp14b, Jsd | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U3 small nucleolar RNA-associated protein 14 homolog B (Juvenile spermatogonial depletion protein). | |||||
|
DAB2P_HUMAN
|
||||||
| NC score | 0.029314 (rank : 66) | θ value | 5.27518 (rank : 48) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q5VWQ8, Q8TDL2, Q96SE1, Q9C0C0 | Gene names | DAB2IP, AF9Q34, AIP1, KIAA1743 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Disabled homolog 2-interacting protein (DAB2-interacting protein) (DAB2 interaction protein) (ASK-interacting protein 1). | |||||
|
DAB2P_MOUSE
|
||||||
| NC score | 0.028876 (rank : 67) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q3UHC7, Q3TPD5, Q3UH44, Q6JTV1, Q80T97 | Gene names | Dab2ip, Kiaa1743 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Disabled homolog 2-interacting protein (DAB2-interacting protein). | |||||
|
ENL_HUMAN
|
||||||
| NC score | 0.027626 (rank : 68) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
UBP34_HUMAN
|
||||||
| NC score | 0.026379 (rank : 69) | θ value | 4.03905 (rank : 45) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q70CQ2, O60316, O94834, Q3B777, Q6P6C9, Q7L8P6, Q8N3T9, Q8TBW2, Q9UGA1 | Gene names | USP34, KIAA0570, KIAA0729 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 34 (EC 3.1.2.15) (Ubiquitin thioesterase 34) (Ubiquitin-specific-processing protease 34) (Deubiquitinating enzyme 34). | |||||
|
UBP34_MOUSE
|
||||||
| NC score | 0.025177 (rank : 70) | θ value | 5.27518 (rank : 59) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6ZQ93, Q3UPN0, Q6P563, Q7TMJ6, Q8CCH0 | Gene names | Usp34, Kiaa0570 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 34 (EC 3.1.2.15) (Ubiquitin thioesterase 34) (Ubiquitin-specific-processing protease 34) (Deubiquitinating enzyme 34). | |||||
|
MERL_MOUSE
|
||||||
| NC score | 0.024989 (rank : 71) | θ value | 6.88961 (rank : 63) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 524 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P46662, Q8BR03 | Gene names | Nf2, Nf-2 | |||
|
Domain Architecture |

|
|||||
| Description | Merlin (Moesin-ezrin-radixin-like protein) (Neurofibromin-2) (Schwannomin). | |||||
|
PARP2_HUMAN
|
||||||
| NC score | 0.024840 (rank : 72) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UGN5, Q8TEU4, Q9NUV2, Q9UMR4, Q9Y6C8 | Gene names | PARP2, ADPRT2, ADPRTL2 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 2 (EC 2.4.2.30) (PARP-2) (NAD(+) ADP- ribosyltransferase 2) (Poly[ADP-ribose] synthetase 2) (pADPRT-2) (hPARP-2). | |||||
|
BCL9_HUMAN
|
||||||
| NC score | 0.024774 (rank : 73) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
CREB3_HUMAN
|
||||||
| NC score | 0.024435 (rank : 74) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O43889, O14671, O14919, Q96GK8, Q9H2W3, Q9UE77 | Gene names | CREB3, LZIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-responsive element-binding protein 3 (Luman protein) (Transcription factor LZIP-alpha). | |||||
|
UBP49_HUMAN
|
||||||
| NC score | 0.023432 (rank : 75) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q70CQ1, Q96CK4 | Gene names | USP49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 49 (EC 3.1.2.15) (Ubiquitin thioesterase 49) (Ubiquitin-specific-processing protease 49) (Deubiquitinating enzyme 49). | |||||
|
RNF25_HUMAN
|
||||||
| NC score | 0.023291 (rank : 76) | θ value | 5.27518 (rank : 56) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96BH1, Q9H874 | Gene names | RNF25 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 25 (EC 6.3.2.-). | |||||
|
TRI37_HUMAN
|
||||||
| NC score | 0.023260 (rank : 77) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O94972, Q7Z3E6, Q8IYF7, Q8WYF7 | Gene names | TRIM37, KIAA0898, MUL, POB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 37 (Mulibrey nanism protein). | |||||
|
BCL9_MOUSE
|
||||||
| NC score | 0.022563 (rank : 78) | θ value | 5.27518 (rank : 47) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9D219, Q67FX9, Q8BUJ8, Q8VE74 | Gene names | Bcl9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9). | |||||
|
PB1_HUMAN
|
||||||
| NC score | 0.018183 (rank : 79) | θ value | 6.88961 (rank : 64) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86U86, Q9H2T3, Q9H2T4, Q9H2T5, Q9H301, Q9H314 | Gene names | PB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein polybromo-1 (hPB1) (Polybromo-1D) (BRG1-associated factor 180) (BAF180). | |||||
|
SEPT8_MOUSE
|
||||||
| NC score | 0.016565 (rank : 80) | θ value | 5.27518 (rank : 58) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8CHH9, Q80YC7, Q9ESF7 | Gene names | Sept8, Kiaa0202 | |||
|
Domain Architecture |

|
|||||
| Description | Septin-8. | |||||
|
SEM6B_HUMAN
|
||||||
| NC score | 0.016307 (rank : 81) | θ value | 1.06291 (rank : 13) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H3T3, Q9NRK9 | Gene names | SEMA6B, SEMAZ | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6B precursor (Semaphorin Z) (Sema Z). | |||||
|
PRRT3_HUMAN
|
||||||
| NC score | 0.014917 (rank : 82) | θ value | 6.88961 (rank : 66) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
IF4G1_HUMAN
|
||||||
| NC score | 0.012545 (rank : 83) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q04637, O43177, O95066, Q5HYG0, Q6ZN21, Q8N102 | Gene names | EIF4G1, EIF4G, EIF4GI | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1) (p220). | |||||
|
TNIK_HUMAN
|
||||||
| NC score | 0.011956 (rank : 84) | θ value | 3.0926 (rank : 31) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1570 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UKE5, O60298, Q8WUY7, Q9UKD8, Q9UKD9, Q9UKE0, Q9UKE1, Q9UKE2, Q9UKE3, Q9UKE4 | Gene names | TNIK, KIAA0551 | |||
|
Domain Architecture |

|
|||||
| Description | TRAF2 and NCK-interacting protein kinase (EC 2.7.11.1). | |||||
|
SMCA2_HUMAN
|
||||||
| NC score | 0.011106 (rank : 85) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P51531 | Gene names | SMARCA2, BRM, SNF2A, SNF2L2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L2 (EC 3.6.1.-) (ATP- dependent helicase SMARCA2) (SNF2-alpha) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin subfamily A member 2) (hBRM). | |||||
|
NSD1_HUMAN
|
||||||
| NC score | 0.009737 (rank : 86) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
ARHGC_HUMAN
|
||||||
| NC score | 0.008881 (rank : 87) | θ value | 8.99809 (rank : 71) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9NZN5, O15086 | Gene names | ARHGEF12, KIAA0382, LARG | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 12 (Leukemia-associated RhoGEF). | |||||
|
FOXN1_MOUSE
|
||||||
| NC score | 0.008736 (rank : 88) | θ value | 6.88961 (rank : 60) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 300 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61575 | Gene names | Foxn1, Fkh19, Hfh11, Whn | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein N1 (Transcription factor winged-helix nude) (Hepatocyte nuclear factor 3 forkhead homolog 11) (HNF-3/forkhead homolog 11) (HFH-11). | |||||
|
PAX9_HUMAN
|
||||||
| NC score | 0.008386 (rank : 89) | θ value | 8.99809 (rank : 81) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55771, Q99582, Q9UQR4 | Gene names | PAX9 | |||
|
Domain Architecture |

|
|||||
| Description | Paired box protein Pax-9. | |||||
|
PLXB1_HUMAN
|
||||||
| NC score | 0.008114 (rank : 90) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O43157, Q6NY20, Q9UIV7, Q9UJ92, Q9UJ93 | Gene names | PLXNB1, KIAA0407, SEP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B1 precursor (Semaphorin receptor SEP). | |||||
|
TITIN_HUMAN
|
||||||
| NC score | 0.006605 (rank : 91) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
MAK_MOUSE
|
||||||
| NC score | 0.003751 (rank : 92) | θ value | 3.0926 (rank : 29) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 873 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q04859 | Gene names | Mak, Rck | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase MAK (EC 2.7.11.22) (Male germ cell- associated kinase) (Protein kinase RCK). | |||||
|
OBSCN_HUMAN
|
||||||
| NC score | 0.002890 (rank : 93) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
PRD16_HUMAN
|
||||||
| NC score | 0.001523 (rank : 94) | θ value | 8.99809 (rank : 83) | |||
| Query Neighborhood Hits | 85 | Target Neighborhood Hits | 832 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9HAZ2, Q8WYJ9, Q9C0I8 | Gene names | PRDM16, KIAA1675, MEL1, PFM13 | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 16 (PR domain-containing protein 16) (Transcription factor MEL1). | |||||