Please be patient as the page loads
|
LYRIC_HUMAN
|
||||||
| SwissProt Accessions | Q86UE4, Q6PK07, Q8TCX3 | Gene names | MTDH, AEG1, LYRIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LYRIC (Lysine-rich CEACAM1 co-isolated protein) (3D3/lyric) (Metastasis adhesion protein) (Metadherin) (Astrocyte elevated gene-1 protein) (AEG-1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
LYRIC_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 99 | |
| SwissProt Accessions | Q86UE4, Q6PK07, Q8TCX3 | Gene names | MTDH, AEG1, LYRIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LYRIC (Lysine-rich CEACAM1 co-isolated protein) (3D3/lyric) (Metastasis adhesion protein) (Metadherin) (Astrocyte elevated gene-1 protein) (AEG-1). | |||||
|
LYRIC_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.943042 (rank : 2) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q80WJ7, Q3U9F8, Q3UAQ8, Q8BN67, Q8CBT9, Q8CDL0, Q8CGI7, Q9D052 | Gene names | Mtdh, Lyric | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LYRIC (Lysine-rich CEACAM1 co-isolated protein) (3D3/LYRIC) (Metastasis adhesion protein) (Metadherin). | |||||
|
AFF4_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 3) | NC score | 0.143781 (rank : 6) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
AFF4_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 4) | NC score | 0.145954 (rank : 5) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
MBB1A_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 5) | NC score | 0.122719 (rank : 8) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q7TPV4, O35851, Q80Y66, Q8R4X2, Q99KP0 | Gene names | Mybbp1a, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A (Myb-binding protein of 160 kDa). | |||||
|
LEO1_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 6) | NC score | 0.104657 (rank : 12) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
TNC6B_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 7) | NC score | 0.107861 (rank : 11) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UPQ9, Q5TH52, Q8TBX2 | Gene names | TNRC6B, KIAA1093 | |||
|
Domain Architecture |

|
|||||
| Description | Trinucleotide repeat-containing 6B protein. | |||||
|
ARFG1_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 8) | NC score | 0.073614 (rank : 28) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9EPJ9, Q3TI52, Q8BMM6, Q8BMQ7 | Gene names | Arfgap1, Arf1gap | |||
|
Domain Architecture |
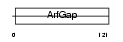
|
|||||
| Description | ADP-ribosylation factor GTPase-activating protein 1 (ADP-ribosylation factor 1 GTPase-activating protein) (ARF1 GAP) (ARF1-directed GTPase- activating protein) (GAP protein). | |||||
|
SNX9_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 9) | NC score | 0.068021 (rank : 36) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q91VH2 | Gene names | Snx9 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-9. | |||||
|
MYLK_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 10) | NC score | 0.014929 (rank : 117) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1489 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6PDN3, Q3TSJ7, Q80UX0, Q80YN7, Q80YN8, Q8K026, Q924D2, Q9ERD3 | Gene names | Mylk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin light chain kinase, smooth muscle (EC 2.7.11.18) (MLCK) (Telokin) (Kinase-related protein) (KRP). | |||||
|
DPOLZ_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 11) | NC score | 0.083891 (rank : 19) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O60673, O43214 | Gene names | REV3L, POLZ, REV3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (hREV3). | |||||
|
DSPP_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 12) | NC score | 0.152959 (rank : 3) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
DSPP_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 13) | NC score | 0.149721 (rank : 4) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 14) | NC score | 0.117878 (rank : 9) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
MTMR7_HUMAN
|
||||||
| θ value | 0.125558 (rank : 15) | NC score | 0.063181 (rank : 42) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y216, Q68DX4 | Gene names | MTMR7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myotubularin-related protein 7 (EC 3.1.3.-). | |||||
|
SELPL_MOUSE
|
||||||
| θ value | 0.125558 (rank : 16) | NC score | 0.086009 (rank : 17) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 588 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q62170 | Gene names | Selplg, Selp1, Selpl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | P-selectin glycoprotein ligand 1 precursor (PSGL-1) (Selectin P ligand). | |||||
|
CT2NL_HUMAN
|
||||||
| θ value | 0.163984 (rank : 17) | NC score | 0.036161 (rank : 88) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9P2B4, Q96B40 | Gene names | CTTNBP2NL, KIAA1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 18) | NC score | 0.031222 (rank : 96) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
NASP_HUMAN
|
||||||
| θ value | 0.21417 (rank : 19) | NC score | 0.092229 (rank : 14) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P49321, Q96A69, Q9BTW2 | Gene names | NASP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigenic sperm protein (NASP). | |||||
|
T2FA_HUMAN
|
||||||
| θ value | 0.21417 (rank : 20) | NC score | 0.124290 (rank : 7) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P35269, Q9BWN0 | Gene names | GTF2F1, RAP74 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor IIF alpha subunit (EC 2.7.11.1) (TFIIF-alpha) (Transcription initiation factor RAP74) (General transcription factor IIF polypeptide 1 74 kDa subunit protein). | |||||
|
AF10_HUMAN
|
||||||
| θ value | 0.279714 (rank : 21) | NC score | 0.041394 (rank : 82) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P55197 | Gene names | MLLT10, AF10 | |||
|
Domain Architecture |
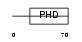
|
|||||
| Description | Protein AF-10. | |||||
|
MTMR7_MOUSE
|
||||||
| θ value | 0.279714 (rank : 22) | NC score | 0.058254 (rank : 53) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Z2C9, Q8C4J6 | Gene names | Mtmr7 | |||
|
Domain Architecture |
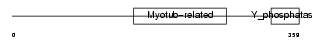
|
|||||
| Description | Myotubularin-related protein 7 (EC 3.1.3.-). | |||||
|
RFC1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 23) | NC score | 0.057193 (rank : 55) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P35601 | Gene names | Rfc1, Ibf-1, Recc1 | |||
|
Domain Architecture |

|
|||||
| Description | Replication factor C subunit 1 (Replication factor C large subunit) (RF-C 140 kDa subunit) (Activator 1 140 kDa subunit) (Activator 1 large subunit) (A1 140 kDa subunit) (A1-P145) (Differentiation- specific element-binding protein) (ISRE-binding protein). | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 24) | NC score | 0.090770 (rank : 15) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
NOP14_HUMAN
|
||||||
| θ value | 0.47712 (rank : 25) | NC score | 0.063216 (rank : 41) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P78316, Q7LGI5, Q7Z6K0, Q8TBR6 | Gene names | NOP14, C4orf9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
RPGR_HUMAN
|
||||||
| θ value | 0.47712 (rank : 26) | NC score | 0.064106 (rank : 40) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q92834, O00702, O00737, Q8N5T6, Q93039, Q9HD29, Q9UMR1 | Gene names | RPGR, RP3, XLRP3 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator. | |||||
|
LSR_MOUSE
|
||||||
| θ value | 0.62314 (rank : 27) | NC score | 0.055679 (rank : 59) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99KG5, Q3TJE7, Q3UIQ9, Q61148, Q61149, Q6U816 | Gene names | Lsr, Lisch7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipolysis-stimulated lipoprotein receptor precursor (Liver-specific bHLH-Zip transcription factor) (Lipolysis-stimulated receptor) (Liver- specific gene on mouse chromosome 7 protein). | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 28) | NC score | 0.062366 (rank : 46) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
TXND2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 29) | NC score | 0.068599 (rank : 34) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
WDHD1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 30) | NC score | 0.046864 (rank : 75) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P59328, Q6P408 | Gene names | Wdhd1, And1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and HMG-box DNA-binding protein 1 (Acidic nucleoplasmic DNA- binding protein 1) (And-1). | |||||
|
EPN1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 31) | NC score | 0.032994 (rank : 92) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y6I3, Q86ST3, Q9HA18 | Gene names | EPN1 | |||
|
Domain Architecture |
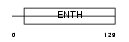
|
|||||
| Description | Epsin-1 (EPS-15-interacting protein 1) (EH domain-binding mitotic phosphoprotein). | |||||
|
MAP1B_MOUSE
|
||||||
| θ value | 0.813845 (rank : 32) | NC score | 0.082364 (rank : 20) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
MATR3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 33) | NC score | 0.061494 (rank : 49) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P43243, Q9UHW0, Q9UQ27 | Gene names | MATR3, KIAA0723 | |||
|
Domain Architecture |

|
|||||
| Description | Matrin-3. | |||||
|
MBB1A_HUMAN
|
||||||
| θ value | 0.813845 (rank : 34) | NC score | 0.094461 (rank : 13) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
RBMX2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 35) | NC score | 0.044288 (rank : 77) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y388, Q5JY82, Q9Y3I8 | Gene names | RBMX2 | |||
|
Domain Architecture |
|
|||||
| Description | RNA-binding motif protein, X-linked 2. | |||||
|
SYLC_MOUSE
|
||||||
| θ value | 0.813845 (rank : 36) | NC score | 0.057125 (rank : 56) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BMJ2, Q8BKW9 | Gene names | Lars | |||
|
Domain Architecture |

|
|||||
| Description | Leucyl-tRNA synthetase, cytoplasmic (EC 6.1.1.4) (Leucine--tRNA ligase) (LeuRS). | |||||
|
BRDT_MOUSE
|
||||||
| θ value | 1.06291 (rank : 37) | NC score | 0.029041 (rank : 97) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q91Y44, Q59HJ4 | Gene names | Brdt, Fsrg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain testis-specific protein (RING3-like protein) (Bromodomain- containing female sterile homeotic-like protein). | |||||
|
GASP1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 38) | NC score | 0.031427 (rank : 94) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5JY77, O43168, Q96LA1 | Gene names | GPRASP1, GASP, KIAA0443 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor-associated sorting protein 1 (GASP-1). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 1.06291 (rank : 39) | NC score | 0.055837 (rank : 58) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 1.06291 (rank : 40) | NC score | 0.042068 (rank : 80) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 1.06291 (rank : 41) | NC score | 0.110376 (rank : 10) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
SETD8_HUMAN
|
||||||
| θ value | 1.06291 (rank : 42) | NC score | 0.080137 (rank : 22) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NQR1, Q86W83, Q8TD09 | Gene names | SETD8, PRSET7, SET07, SET8 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8) (PR/SET domain-containing protein 07) (PR/SET07) (PR-Set7). | |||||
|
TSH2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 43) | NC score | 0.032753 (rank : 93) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q68FE9, Q5DTF1, Q6AXH5, Q9JL71 | Gene names | Tshz2, Kiaa4248, Sdccag33l, Tsh2, Zfp218, Znf218 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 2 (Zinc finger protein 218) (SDCCAG33-like protein). | |||||
|
ZN638_MOUSE
|
||||||
| θ value | 1.38821 (rank : 44) | NC score | 0.049004 (rank : 73) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q61464, Q6DFV9, Q8C941 | Gene names | Znf638, Np220, Zfml | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein). | |||||
|
SETD8_MOUSE
|
||||||
| θ value | 1.81305 (rank : 45) | NC score | 0.074277 (rank : 25) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q2YDW7, Q8C0J9 | Gene names | SETD8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8). | |||||
|
ANGL2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 46) | NC score | 0.011726 (rank : 123) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9R045 | Gene names | Angptl2, Arp2 | |||
|
Domain Architecture |

|
|||||
| Description | Angiopoietin-related protein 2 precursor (Angiopoietin-like 2). | |||||
|
CJ120_HUMAN
|
||||||
| θ value | 2.36792 (rank : 47) | NC score | 0.087367 (rank : 16) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5SQS8 | Gene names | C10orf120 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative uncharacterized protein C10orf120. | |||||
|
ELF4_MOUSE
|
||||||
| θ value | 2.36792 (rank : 48) | NC score | 0.016647 (rank : 113) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z2U4 | Gene names | Elf4, Mef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
EMIL2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 49) | NC score | 0.024730 (rank : 101) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8K482 | Gene names | Emilin2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Basilin). | |||||
|
EP400_HUMAN
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.023546 (rank : 104) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q96L91, O15411, Q6P2F5, Q8N8Q7, Q8NE05, Q96JK7, Q9P230 | Gene names | EP400, CAGH32, KIAA1498, KIAA1818, TNRC12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (hDomino) (CAG repeat protein 32) (Trinucleotide repeat-containing gene 12 protein). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.074157 (rank : 26) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
LUZP1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 52) | NC score | 0.022411 (rank : 105) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 739 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q86V48, Q5TH93, Q8N4X3, Q8TEH1 | Gene names | LUZP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper protein 1. | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 53) | NC score | 0.036602 (rank : 87) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 2.36792 (rank : 54) | NC score | 0.041647 (rank : 81) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
ADDB_MOUSE
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.057117 (rank : 57) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 574 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9QYB8, Q3U0E1, Q80VH9, Q8C1C4, Q9CXE3, Q9JLE5, Q9QYB7 | Gene names | Add2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adducin (Erythrocyte adducin subunit beta) (Add97). | |||||
|
CA048_HUMAN
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.033457 (rank : 90) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96IY1, Q5SY75, Q9H2M5, Q9NRN8, Q9Y415 | Gene names | C1orf48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C1orf48. | |||||
|
CENPC_MOUSE
|
||||||
| θ value | 3.0926 (rank : 57) | NC score | 0.038109 (rank : 85) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P49452 | Gene names | Cenpc1, Cenpc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein C 1 (CENP-C) (Centromere autoantigen C). | |||||
|
CENPF_HUMAN
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.011800 (rank : 122) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
MDN1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | 0.061694 (rank : 47) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9NU22, O15019 | Gene names | MDN1, KIAA0301 | |||
|
Domain Architecture |

|
|||||
| Description | Midasin (MIDAS-containing protein). | |||||
|
PK1IP_MOUSE
|
||||||
| θ value | 3.0926 (rank : 60) | NC score | 0.024292 (rank : 103) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DCE5, Q3UBK7, Q80UT4, Q8C5N6, Q923K2, Q9D3E9 | Gene names | Pak1ip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | p21-activated protein kinase-interacting protein 1 (PAK1-interacting protein 1) (Putative PAK inhibitor Skb15). | |||||
|
RAD18_MOUSE
|
||||||
| θ value | 3.0926 (rank : 61) | NC score | 0.038134 (rank : 84) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9QXK2, Q9CZB8 | Gene names | Rad18, Rad18sc | |||
|
Domain Architecture |

|
|||||
| Description | Postreplication repair protein RAD18 (mRAD18Sc). | |||||
|
REXO4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 62) | NC score | 0.027794 (rank : 99) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PAQ4, Q3UU17, Q8C524 | Gene names | Rexo4, Gm111, Pmc2, Xpmc2h | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA exonuclease 4 (EC 3.1.-.-) (Exonuclease XPMC2) (Prevents mitotic catastrophe 2 protein homolog). | |||||
|
ELOA1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.045081 (rank : 76) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q14241, Q8IXH1 | Gene names | TCEB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription elongation factor B polypeptide 3 (RNA polymerase II transcription factor SIII subunit A1) (SIII p110) (Elongin-A) (EloA) (Elongin 110 kDa subunit). | |||||
|
PHF14_HUMAN
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.052405 (rank : 64) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
SMRC2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 65) | NC score | 0.062660 (rank : 44) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q6PDG5, Q6P6P3 | Gene names | Smarcc2, Baf170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
SMRCD_HUMAN
|
||||||
| θ value | 4.03905 (rank : 66) | NC score | 0.018596 (rank : 110) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H4L7, Q96SX1, Q9H017, Q9H860, Q9NPU9, Q9ULU7 | Gene names | SMARCAD1, KIAA1122 | |||
|
Domain Architecture |

|
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A containing DEAD/H box 1 (EC 3.6.1.-) (ATP- dependent helicase 1) (hHEL1). | |||||
|
ZF106_HUMAN
|
||||||
| θ value | 4.03905 (rank : 67) | NC score | 0.028447 (rank : 98) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9H2Y7, Q6NSD9, Q6PEK1, Q86T43, Q86T45, Q86T50, Q86T58, Q86TA9, Q96M37, Q9H7B8 | Gene names | ZFP106, ZNF474 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 106 homolog (Zfp-106) (Zinc finger protein 474). | |||||
|
ACINU_HUMAN
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.076073 (rank : 24) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
PSIP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.044218 (rank : 78) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q99JF8, Q3TEJ7, Q3TTD7, Q3UTA1, Q80WQ7, Q99JF7, Q99LR4, Q9CT03 | Gene names | Psip1, Ledgf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PC4 and SFRS1-interacting protein (Lens epithelium-derived growth factor) (mLEDGF). | |||||
|
SLC2B_HUMAN
|
||||||
| θ value | 5.27518 (rank : 70) | NC score | 0.033311 (rank : 91) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NEV8, Q9Y4D6 | Gene names | EXPH5, KIAA0624, SLAC2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Slp homolog lacking C2 domains b (Exophilin-5). | |||||
|
SMRC2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 71) | NC score | 0.063023 (rank : 43) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8TAQ2, Q92923, Q96E12, Q96GY4 | Gene names | SMARCC2, BAF170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
ARI5B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.043425 (rank : 79) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q14865, Q7Z3M4, Q8N421, Q9H786 | Gene names | ARID5B, DESRT, MRF2 | |||
|
Domain Architecture |
|
|||||
| Description | AT-rich interactive domain-containing protein 5B (ARID domain- containing protein 5B) (Mrf1-like) (Modulator recognition factor 2) (MRF-2). | |||||
|
BRE1A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | 0.016317 (rank : 115) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1071 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5VTR2, Q69YL5, Q6P527, Q8N3J4, Q96JD3, Q9H9Y7, Q9HA51, Q9NUR4, Q9NWQ3, Q9NX83 | Gene names | RNF20, BRE1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase BRE1A (EC 6.3.2.-) (BRE1-A) (hBRE1) (RING finger protein 20). | |||||
|
CALD1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.031247 (rank : 95) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q05682, Q13978, Q13979, Q14741, Q14742 | Gene names | CALD1, CAD, CDM | |||
|
Domain Architecture |

|
|||||
| Description | Caldesmon (CDM). | |||||
|
DBF4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.018988 (rank : 109) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QZ41, Q3TQP4, Q3V1K2, Q6NSQ1, Q9CXF2 | Gene names | Dbf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DBF4 homolog (MuDBF4). | |||||
|
DDX10_MOUSE
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.007135 (rank : 125) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80Y44 | Gene names | Ddx10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX10 (EC 3.6.1.-) (DEAD box protein 10). | |||||
|
LSR_HUMAN
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.039681 (rank : 83) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q86X29, O00112, O00426, Q6ZT80, Q9UQL3 | Gene names | LSR, LISCH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipolysis-stimulated lipoprotein receptor. | |||||
|
MASTL_HUMAN
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.002267 (rank : 126) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 894 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96GX5, Q5T8D5, Q5T8D7, Q8NCD6, Q96SJ5 | Gene names | MASTL, THC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase-like (EC 2.7.11.1). | |||||
|
NFH_HUMAN
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.047034 (rank : 74) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
PALM_MOUSE
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.027245 (rank : 100) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Z0P4, Q9Z0P3 | Gene names | Palm | |||
|
Domain Architecture |
|
|||||
| Description | Paralemmin. | |||||
|
PHF14_MOUSE
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.055356 (rank : 60) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
SPC25_MOUSE
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.018214 (rank : 111) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3UA16, Q9D021, Q9D1K6 | Gene names | Spbc25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinetochore protein Spc25. | |||||
|
XYLT1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.016456 (rank : 114) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q811B1, Q9EPL1 | Gene names | Xylt1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Xylosyltransferase 1 (EC 2.4.2.26) (Xylosyltransferase I) (Peptide O- xylosyltransferase 1). | |||||
|
BRD3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.037889 (rank : 86) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8K2F0, Q8C665, Q8CAX7, Q9JI25 | Gene names | Brd3, Fsrg2 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 3 (Bromodomain-containing FSH-like protein FSRG2). | |||||
|
DPF1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.014145 (rank : 119) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9QX66, Q9QX65, Q9QYA4 | Gene names | Dpf1, Neud4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
E41L2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.013860 (rank : 120) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O43491 | Gene names | EPB41L2 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 2 (Generally expressed protein 4.1) (4.1G). | |||||
|
LRP11_MOUSE
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.012305 (rank : 121) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CB67, Q8C7Y7 | Gene names | Lrp11 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 11 precursor. | |||||
|
MED6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.024379 (rank : 102) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75586, O15401, Q9BTH1, Q9UHL1 | Gene names | MED6, ARC33 | |||
|
Domain Architecture |
|
|||||
| Description | RNA polymerase transcriptional regulation mediator, subunit 6 homolog (Activator-recruited cofactor 33 kDa component) (ARC33) (NY-REN-28 antigen). | |||||
|
MYCPP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.011584 (rank : 124) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z401, Q14655, Q86T77, Q8IVX2, Q8NB93 | Gene names | MYCPBP, IRLB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-myc promoter-binding protein. | |||||
|
MYST3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.034850 (rank : 89) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
NRDC_MOUSE
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.020950 (rank : 106) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BHG1 | Gene names | Nrd1 | |||
|
Domain Architecture |

|
|||||
| Description | Nardilysin precursor (EC 3.4.24.61) (N-arginine dibasic convertase) (NRD convertase) (NRD-C). | |||||
|
NUP62_MOUSE
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.018153 (rank : 112) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q63850, Q99JN7 | Gene names | Nup62 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore glycoprotein p62 (62 kDa nucleoporin). | |||||
|
P52K_HUMAN
|
||||||
| θ value | 8.99809 (rank : 93) | NC score | 0.014757 (rank : 118) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O43422, Q8WTW1, Q9Y3Z4 | Gene names | PRKRIR, DAP4, P52RIPK, THAP0 | |||
|
Domain Architecture |

|
|||||
| Description | 52 kDa repressor of the inhibitor of the protein kinase (p58IPK- interacting protein) (58 kDa interferon-induced protein kinase- interacting protein) (P52rIPK) (Death-associated protein 4) (THAP domain-containing protein 0). | |||||
|
PPR3A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.020070 (rank : 108) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99MR9, Q8BUJ4, Q8BUL0 | Gene names | Ppp1r3a, Pp1g | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase 1 regulatory subunit 3A (Protein phosphatase 1 glycogen-associated regulatory subunit) (Protein phosphatase type-1 glycogen targeting subunit) (RGL). | |||||
|
R51A1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 95) | NC score | 0.078379 (rank : 23) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8C551, O55219, Q8BP36, Q99L94, Q9D0J0 | Gene names | Rad51ap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RAD51-associated protein 1 (RAD51-interacting protein) (RAB22). | |||||
|
REQU_HUMAN
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | 0.014943 (rank : 116) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 523 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92785 | Gene names | DPF2, REQ, UBID4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein ubi-d4 (Requiem) (Apoptosis response zinc finger protein) (D4, zinc and double PHD fingers family 2). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 97) | NC score | 0.020574 (rank : 107) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
RL1D1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 98) | NC score | 0.053924 (rank : 61) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O76021, Q6PL22, Q8IWS7, Q8WUZ1, Q9HDA9, Q9Y3Z9 | Gene names | RSL1D1, CATX11, CSIG, PBK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribosomal L1 domain-containing protein 1 (Cellular senescence- inhibited gene protein) (Protein PBK1) (CATX-11). | |||||
|
TXND2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 99) | NC score | 0.070320 (rank : 31) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
AFF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.068539 (rank : 35) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
AFF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.058318 (rank : 52) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O88573 | Gene names | Aff1, Mllt2, Mllt2h | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4). | |||||
|
AFF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.050734 (rank : 69) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P51816, O43786, O60215, P78407, Q13521, Q14323, Q9UNA5 | Gene names | AFF2, FMR2, OX19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation 2 protein) (Protein FMR-2) (FMR2P) (Protein Ox19) (Fragile X E mental retardation syndrome protein). | |||||
|
AFF2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.059488 (rank : 51) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O55112 | Gene names | Aff2, Fmr2, Ox19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation protein 2 homolog) (Protein FMR-2) (FMR2P) (Protein Ox19). | |||||
|
AFF3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.070331 (rank : 30) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P51826 | Gene names | AFF3, LAF4 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 3 (Protein LAF-4) (Lymphoid nuclear protein related to AF4). | |||||
|
AKA12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.050066 (rank : 72) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
BASP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.061669 (rank : 48) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
BASP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.051169 (rank : 67) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
CYLC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.050974 (rank : 68) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
CYLC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.069998 (rank : 32) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
DMP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.062441 (rank : 45) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
DMP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.085386 (rank : 18) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
GPTC8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.050729 (rank : 70) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UKJ3, O60300 | Gene names | GPATC8, KIAA0553 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 8. | |||||
|
IWS1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.065484 (rank : 38) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
LAD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.074066 (rank : 27) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O00515, O95614 | Gene names | LAD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (120 kDa linear IgA bullous dermatosis antigen) (97 kDa linear IgA bullous dermatosis antigen) (Linear IgA disease antigen homolog) (LadA). | |||||
|
MAP1B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.073370 (rank : 29) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
NOLC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.080420 (rank : 21) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
NSBP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.069266 (rank : 33) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
NUCKS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.057381 (rank : 54) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
RBBP6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.051335 (rank : 66) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
RBBP6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.052733 (rank : 63) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
SFR11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.060592 (rank : 50) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.050177 (rank : 71) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
TGON2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.065242 (rank : 39) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
THOC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.053768 (rank : 62) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8NI27, Q9H8I6 | Gene names | THOC2, CXorf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | THO complex subunit 2 (Tho2). | |||||
|
TNR6A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.052139 (rank : 65) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8NDV7, O15408, Q658L5, Q6NVB5, Q8NEZ0, Q8TBT8, Q8TCR0, Q9NV59, Q9P268 | Gene names | TNRC6A, CAGH26, KIAA1460, TNRC6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trinucleotide repeat-containing gene 6A protein (CAG repeat protein 26) (Glycine-tryptophan protein of 182 kDa) (GW182 autoantigen) (Protein GW1) (EMSY interactor protein). | |||||
|
TRDN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.065943 (rank : 37) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
LYRIC_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 99 | |
| SwissProt Accessions | Q86UE4, Q6PK07, Q8TCX3 | Gene names | MTDH, AEG1, LYRIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LYRIC (Lysine-rich CEACAM1 co-isolated protein) (3D3/lyric) (Metastasis adhesion protein) (Metadherin) (Astrocyte elevated gene-1 protein) (AEG-1). | |||||
|
LYRIC_MOUSE
|
||||||
| NC score | 0.943042 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q80WJ7, Q3U9F8, Q3UAQ8, Q8BN67, Q8CBT9, Q8CDL0, Q8CGI7, Q9D052 | Gene names | Mtdh, Lyric | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LYRIC (Lysine-rich CEACAM1 co-isolated protein) (3D3/LYRIC) (Metastasis adhesion protein) (Metadherin). | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.152959 (rank : 3) | θ value | 0.0431538 (rank : 12) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
DSPP_MOUSE
|
||||||
| NC score | 0.149721 (rank : 4) | θ value | 0.0961366 (rank : 13) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
AFF4_MOUSE
|
||||||
| NC score | 0.145954 (rank : 5) | θ value | 0.0113563 (rank : 4) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
AFF4_HUMAN
|
||||||
| NC score | 0.143781 (rank : 6) | θ value | 0.00509761 (rank : 3) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
T2FA_HUMAN
|
||||||
| NC score | 0.124290 (rank : 7) | θ value | 0.21417 (rank : 20) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P35269, Q9BWN0 | Gene names | GTF2F1, RAP74 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor IIF alpha subunit (EC 2.7.11.1) (TFIIF-alpha) (Transcription initiation factor RAP74) (General transcription factor IIF polypeptide 1 74 kDa subunit protein). | |||||
|
MBB1A_MOUSE
|
||||||
| NC score | 0.122719 (rank : 8) | θ value | 0.0113563 (rank : 5) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q7TPV4, O35851, Q80Y66, Q8R4X2, Q99KP0 | Gene names | Mybbp1a, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A (Myb-binding protein of 160 kDa). | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.117878 (rank : 9) | θ value | 0.0961366 (rank : 14) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.110376 (rank : 10) | θ value | 1.06291 (rank : 41) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
TNC6B_HUMAN
|
||||||
| NC score | 0.107861 (rank : 11) | θ value | 0.0193708 (rank : 7) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UPQ9, Q5TH52, Q8TBX2 | Gene names | TNRC6B, KIAA1093 | |||
|
Domain Architecture |

|
|||||
| Description | Trinucleotide repeat-containing 6B protein. | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.104657 (rank : 12) | θ value | 0.0193708 (rank : 6) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
MBB1A_HUMAN
|
||||||
| NC score | 0.094461 (rank : 13) | θ value | 0.813845 (rank : 34) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
NASP_HUMAN
|
||||||
| NC score | 0.092229 (rank : 14) | θ value | 0.21417 (rank : 19) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P49321, Q96A69, Q9BTW2 | Gene names | NASP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigenic sperm protein (NASP). | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.090770 (rank : 15) | θ value | 0.47712 (rank : 24) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
CJ120_HUMAN
|
||||||
| NC score | 0.087367 (rank : 16) | θ value | 2.36792 (rank : 47) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5SQS8 | Gene names | C10orf120 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative uncharacterized protein C10orf120. | |||||
|
SELPL_MOUSE
|
||||||
| NC score | 0.086009 (rank : 17) | θ value | 0.125558 (rank : 16) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 588 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q62170 | Gene names | Selplg, Selp1, Selpl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | P-selectin glycoprotein ligand 1 precursor (PSGL-1) (Selectin P ligand). | |||||
|
DMP1_MOUSE
|
||||||
| NC score | 0.085386 (rank : 18) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
DPOLZ_HUMAN
|
||||||
| NC score | 0.083891 (rank : 19) | θ value | 0.0431538 (rank : 11) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O60673, O43214 | Gene names | REV3L, POLZ, REV3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (hREV3). | |||||
|
MAP1B_MOUSE
|
||||||
| NC score | 0.082364 (rank : 20) | θ value | 0.813845 (rank : 32) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
NOLC1_HUMAN
|
||||||
| NC score | 0.080420 (rank : 21) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
SETD8_HUMAN
|
||||||
| NC score | 0.080137 (rank : 22) | θ value | 1.06291 (rank : 42) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NQR1, Q86W83, Q8TD09 | Gene names | SETD8, PRSET7, SET07, SET8 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8) (PR/SET domain-containing protein 07) (PR/SET07) (PR-Set7). | |||||
|
R51A1_MOUSE
|
||||||
| NC score | 0.078379 (rank : 23) | θ value | 8.99809 (rank : 95) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8C551, O55219, Q8BP36, Q99L94, Q9D0J0 | Gene names | Rad51ap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RAD51-associated protein 1 (RAD51-interacting protein) (RAB22). | |||||
|
ACINU_HUMAN
|
||||||
| NC score | 0.076073 (rank : 24) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
SETD8_MOUSE
|
||||||
| NC score | 0.074277 (rank : 25) | θ value | 1.81305 (rank : 45) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q2YDW7, Q8C0J9 | Gene names | SETD8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.074157 (rank : 26) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
LAD1_HUMAN
|
||||||
| NC score | 0.074066 (rank : 27) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O00515, O95614 | Gene names | LAD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (120 kDa linear IgA bullous dermatosis antigen) (97 kDa linear IgA bullous dermatosis antigen) (Linear IgA disease antigen homolog) (LadA). | |||||
|
ARFG1_MOUSE
|
||||||
| NC score | 0.073614 (rank : 28) | θ value | 0.0252991 (rank : 8) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9EPJ9, Q3TI52, Q8BMM6, Q8BMQ7 | Gene names | Arfgap1, Arf1gap | |||
|
Domain Architecture |

|
|||||
| Description | ADP-ribosylation factor GTPase-activating protein 1 (ADP-ribosylation factor 1 GTPase-activating protein) (ARF1 GAP) (ARF1-directed GTPase- activating protein) (GAP protein). | |||||
|
MAP1B_HUMAN
|
||||||
| NC score | 0.073370 (rank : 29) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
AFF3_HUMAN
|
||||||
| NC score | 0.070331 (rank : 30) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P51826 | Gene names | AFF3, LAF4 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 3 (Protein LAF-4) (Lymphoid nuclear protein related to AF4). | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.070320 (rank : 31) | θ value | 8.99809 (rank : 99) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
CYLC2_HUMAN
|
||||||
| NC score | 0.069998 (rank : 32) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
NSBP1_HUMAN
|
||||||
| NC score | 0.069266 (rank : 33) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
TXND2_HUMAN
|
||||||
| NC score | 0.068599 (rank : 34) | θ value | 0.62314 (rank : 29) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
AFF1_HUMAN
|
||||||
| NC score | 0.068539 (rank : 35) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
SNX9_MOUSE
|
||||||
| NC score | 0.068021 (rank : 36) | θ value | 0.0252991 (rank : 9) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q91VH2 | Gene names | Snx9 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-9. | |||||
|
TRDN_HUMAN
|
||||||
| NC score | 0.065943 (rank : 37) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.065484 (rank : 38) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
TGON2_HUMAN
|
||||||
| NC score | 0.065242 (rank : 39) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
RPGR_HUMAN
|
||||||
| NC score | 0.064106 (rank : 40) | θ value | 0.47712 (rank : 26) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q92834, O00702, O00737, Q8N5T6, Q93039, Q9HD29, Q9UMR1 | Gene names | RPGR, RP3, XLRP3 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator. | |||||
|
NOP14_HUMAN
|
||||||
| NC score | 0.063216 (rank : 41) | θ value | 0.47712 (rank : 25) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P78316, Q7LGI5, Q7Z6K0, Q8TBR6 | Gene names | NOP14, C4orf9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
MTMR7_HUMAN
|
||||||
| NC score | 0.063181 (rank : 42) | θ value | 0.125558 (rank : 15) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y216, Q68DX4 | Gene names | MTMR7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myotubularin-related protein 7 (EC 3.1.3.-). | |||||
|
SMRC2_HUMAN
|
||||||
| NC score | 0.063023 (rank : 43) | θ value | 5.27518 (rank : 71) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8TAQ2, Q92923, Q96E12, Q96GY4 | Gene names | SMARCC2, BAF170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
SMRC2_MOUSE
|
||||||
| NC score | 0.062660 (rank : 44) | θ value | 4.03905 (rank : 65) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q6PDG5, Q6P6P3 | Gene names | Smarcc2, Baf170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
DMP1_HUMAN
|
||||||
| NC score | 0.062441 (rank : 45) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.062366 (rank : 46) | θ value | 0.62314 (rank : 28) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
MDN1_HUMAN
|
||||||
| NC score | 0.061694 (rank : 47) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9NU22, O15019 | Gene names | MDN1, KIAA0301 | |||
|
Domain Architecture |

|
|||||
| Description | Midasin (MIDAS-containing protein). | |||||
|
BASP_HUMAN
|
||||||
| NC score | 0.061669 (rank : 48) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
MATR3_HUMAN
|
||||||
| NC score | 0.061494 (rank : 49) | θ value | 0.813845 (rank : 33) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P43243, Q9UHW0, Q9UQ27 | Gene names | MATR3, KIAA0723 | |||
|
Domain Architecture |

|
|||||
| Description | Matrin-3. | |||||
|
SFR11_HUMAN
|
||||||
| NC score | 0.060592 (rank : 50) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
AFF2_MOUSE
|
||||||
| NC score | 0.059488 (rank : 51) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O55112 | Gene names | Aff2, Fmr2, Ox19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation protein 2 homolog) (Protein FMR-2) (FMR2P) (Protein Ox19). | |||||
|
AFF1_MOUSE
|
||||||
| NC score | 0.058318 (rank : 52) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O88573 | Gene names | Aff1, Mllt2, Mllt2h | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4). | |||||
|
MTMR7_MOUSE
|
||||||
| NC score | 0.058254 (rank : 53) | θ value | 0.279714 (rank : 22) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Z2C9, Q8C4J6 | Gene names | Mtmr7 | |||
|
Domain Architecture |

|
|||||
| Description | Myotubularin-related protein 7 (EC 3.1.3.-). | |||||
|
NUCKS_HUMAN
|
||||||
| NC score | 0.057381 (rank : 54) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
RFC1_MOUSE
|
||||||
| NC score | 0.057193 (rank : 55) | θ value | 0.365318 (rank : 23) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P35601 | Gene names | Rfc1, Ibf-1, Recc1 | |||
|
Domain Architecture |

|
|||||
| Description | Replication factor C subunit 1 (Replication factor C large subunit) (RF-C 140 kDa subunit) (Activator 1 140 kDa subunit) (Activator 1 large subunit) (A1 140 kDa subunit) (A1-P145) (Differentiation- specific element-binding protein) (ISRE-binding protein). | |||||
|
SYLC_MOUSE
|
||||||
| NC score | 0.057125 (rank : 56) | θ value | 0.813845 (rank : 36) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BMJ2, Q8BKW9 | Gene names | Lars | |||
|
Domain Architecture |

|
|||||
| Description | Leucyl-tRNA synthetase, cytoplasmic (EC 6.1.1.4) (Leucine--tRNA ligase) (LeuRS). | |||||
|
ADDB_MOUSE
|
||||||
| NC score | 0.057117 (rank : 57) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 574 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9QYB8, Q3U0E1, Q80VH9, Q8C1C4, Q9CXE3, Q9JLE5, Q9QYB7 | Gene names | Add2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adducin (Erythrocyte adducin subunit beta) (Add97). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.055837 (rank : 58) | θ value | 1.06291 (rank : 39) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
LSR_MOUSE
|
||||||
| NC score | 0.055679 (rank : 59) | θ value | 0.62314 (rank : 27) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99KG5, Q3TJE7, Q3UIQ9, Q61148, Q61149, Q6U816 | Gene names | Lsr, Lisch7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipolysis-stimulated lipoprotein receptor precursor (Liver-specific bHLH-Zip transcription factor) (Lipolysis-stimulated receptor) (Liver- specific gene on mouse chromosome 7 protein). | |||||
|
PHF14_MOUSE
|
||||||
| NC score | 0.055356 (rank : 60) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
RL1D1_HUMAN
|
||||||
| NC score | 0.053924 (rank : 61) | θ value | 8.99809 (rank : 98) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O76021, Q6PL22, Q8IWS7, Q8WUZ1, Q9HDA9, Q9Y3Z9 | Gene names | RSL1D1, CATX11, CSIG, PBK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribosomal L1 domain-containing protein 1 (Cellular senescence- inhibited gene protein) (Protein PBK1) (CATX-11). | |||||
|
THOC2_HUMAN
|
||||||
| NC score | 0.053768 (rank : 62) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8NI27, Q9H8I6 | Gene names | THOC2, CXorf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | THO complex subunit 2 (Tho2). | |||||
|
RBBP6_MOUSE
|
||||||
| NC score | 0.052733 (rank : 63) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
PHF14_HUMAN
|
||||||
| NC score | 0.052405 (rank : 64) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
TNR6A_HUMAN
|
||||||
| NC score | 0.052139 (rank : 65) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8NDV7, O15408, Q658L5, Q6NVB5, Q8NEZ0, Q8TBT8, Q8TCR0, Q9NV59, Q9P268 | Gene names | TNRC6A, CAGH26, KIAA1460, TNRC6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trinucleotide repeat-containing gene 6A protein (CAG repeat protein 26) (Glycine-tryptophan protein of 182 kDa) (GW182 autoantigen) (Protein GW1) (EMSY interactor protein). | |||||
|
RBBP6_HUMAN
|
||||||
| NC score | 0.051335 (rank : 66) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
BASP_MOUSE
|
||||||
| NC score | 0.051169 (rank : 67) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
CYLC1_HUMAN
|
||||||
| NC score | 0.050974 (rank : 68) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
AFF2_HUMAN
|
||||||
| NC score | 0.050734 (rank : 69) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P51816, O43786, O60215, P78407, Q13521, Q14323, Q9UNA5 | Gene names | AFF2, FMR2, OX19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation 2 protein) (Protein FMR-2) (FMR2P) (Protein Ox19) (Fragile X E mental retardation syndrome protein). | |||||
|
GPTC8_HUMAN
|
||||||
| NC score | 0.050729 (rank : 70) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UKJ3, O60300 | Gene names | GPATC8, KIAA0553 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 8. | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.050177 (rank : 71) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
AKA12_HUMAN
|
||||||
| NC score | 0.050066 (rank : 72) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
ZN638_MOUSE
|
||||||
| NC score | 0.049004 (rank : 73) | θ value | 1.38821 (rank : 44) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q61464, Q6DFV9, Q8C941 | Gene names | Znf638, Np220, Zfml | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein). | |||||
|
NFH_HUMAN
|
||||||
| NC score | 0.047034 (rank : 74) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
WDHD1_MOUSE
|
||||||
| NC score | 0.046864 (rank : 75) | θ value | 0.62314 (rank : 30) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P59328, Q6P408 | Gene names | Wdhd1, And1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and HMG-box DNA-binding protein 1 (Acidic nucleoplasmic DNA- binding protein 1) (And-1). | |||||
|
ELOA1_HUMAN
|
||||||
| NC score | 0.045081 (rank : 76) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q14241, Q8IXH1 | Gene names | TCEB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription elongation factor B polypeptide 3 (RNA polymerase II transcription factor SIII subunit A1) (SIII p110) (Elongin-A) (EloA) (Elongin 110 kDa subunit). | |||||
|
RBMX2_HUMAN
|
||||||
| NC score | 0.044288 (rank : 77) | θ value | 0.813845 (rank : 35) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y388, Q5JY82, Q9Y3I8 | Gene names | RBMX2 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding motif protein, X-linked 2. | |||||
|
PSIP1_MOUSE
|
||||||
| NC score | 0.044218 (rank : 78) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q99JF8, Q3TEJ7, Q3TTD7, Q3UTA1, Q80WQ7, Q99JF7, Q99LR4, Q9CT03 | Gene names | Psip1, Ledgf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PC4 and SFRS1-interacting protein (Lens epithelium-derived growth factor) (mLEDGF). | |||||
|
ARI5B_HUMAN
|
||||||
| NC score | 0.043425 (rank : 79) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q14865, Q7Z3M4, Q8N421, Q9H786 | Gene names | ARID5B, DESRT, MRF2 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 5B (ARID domain- containing protein 5B) (Mrf1-like) (Modulator recognition factor 2) (MRF-2). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.042068 (rank : 80) | θ value | 1.06291 (rank : 40) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.041647 (rank : 81) | θ value | 2.36792 (rank : 54) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
AF10_HUMAN
|
||||||
| NC score | 0.041394 (rank : 82) | θ value | 0.279714 (rank : 21) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P55197 | Gene names | MLLT10, AF10 | |||
|
Domain Architecture |

|
|||||
| Description | Protein AF-10. | |||||
|
LSR_HUMAN
|
||||||
| NC score | 0.039681 (rank : 83) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q86X29, O00112, O00426, Q6ZT80, Q9UQL3 | Gene names | LSR, LISCH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipolysis-stimulated lipoprotein receptor. | |||||
|
RAD18_MOUSE
|
||||||
| NC score | 0.038134 (rank : 84) | θ value | 3.0926 (rank : 61) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9QXK2, Q9CZB8 | Gene names | Rad18, Rad18sc | |||
|
Domain Architecture |

|
|||||
| Description | Postreplication repair protein RAD18 (mRAD18Sc). | |||||
|
CENPC_MOUSE
|
||||||
| NC score | 0.038109 (rank : 85) | θ value | 3.0926 (rank : 57) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P49452 | Gene names | Cenpc1, Cenpc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein C 1 (CENP-C) (Centromere autoantigen C). | |||||
|
BRD3_MOUSE
|
||||||
| NC score | 0.037889 (rank : 86) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8K2F0, Q8C665, Q8CAX7, Q9JI25 | Gene names | Brd3, Fsrg2 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 3 (Bromodomain-containing FSH-like protein FSRG2). | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.036602 (rank : 87) | θ value | 2.36792 (rank : 53) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
CT2NL_HUMAN
|
||||||
| NC score | 0.036161 (rank : 88) | θ value | 0.163984 (rank : 17) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9P2B4, Q96B40 | Gene names | CTTNBP2NL, KIAA1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
MYST3_MOUSE
|
||||||
| NC score | 0.034850 (rank : 89) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
CA048_HUMAN
|
||||||
| NC score | 0.033457 (rank : 90) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96IY1, Q5SY75, Q9H2M5, Q9NRN8, Q9Y415 | Gene names | C1orf48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C1orf48. | |||||
|
SLC2B_HUMAN
|
||||||
| NC score | 0.033311 (rank : 91) | θ value | 5.27518 (rank : 70) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NEV8, Q9Y4D6 | Gene names | EXPH5, KIAA0624, SLAC2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Slp homolog lacking C2 domains b (Exophilin-5). | |||||
|
EPN1_HUMAN
|
||||||
| NC score | 0.032994 (rank : 92) | θ value | 0.813845 (rank : 31) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y6I3, Q86ST3, Q9HA18 | Gene names | EPN1 | |||
|
Domain Architecture |

|
|||||
| Description | Epsin-1 (EPS-15-interacting protein 1) (EH domain-binding mitotic phosphoprotein). | |||||
|
TSH2_MOUSE
|
||||||
| NC score | 0.032753 (rank : 93) | θ value | 1.06291 (rank : 43) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q68FE9, Q5DTF1, Q6AXH5, Q9JL71 | Gene names | Tshz2, Kiaa4248, Sdccag33l, Tsh2, Zfp218, Znf218 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 2 (Zinc finger protein 218) (SDCCAG33-like protein). | |||||
|
GASP1_HUMAN
|
||||||
| NC score | 0.031427 (rank : 94) | θ value | 1.06291 (rank : 38) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5JY77, O43168, Q96LA1 | Gene names | GPRASP1, GASP, KIAA0443 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor-associated sorting protein 1 (GASP-1). | |||||
|
CALD1_HUMAN
|
||||||
| NC score | 0.031247 (rank : 95) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q05682, Q13978, Q13979, Q14741, Q14742 | Gene names | CALD1, CAD, CDM | |||
|
Domain Architecture |

|
|||||
| Description | Caldesmon (CDM). | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.031222 (rank : 96) | θ value | 0.21417 (rank : 18) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
BRDT_MOUSE
|
||||||
| NC score | 0.029041 (rank : 97) | θ value | 1.06291 (rank : 37) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q91Y44, Q59HJ4 | Gene names | Brdt, Fsrg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain testis-specific protein (RING3-like protein) (Bromodomain- containing female sterile homeotic-like protein). | |||||
|
ZF106_HUMAN
|
||||||
| NC score | 0.028447 (rank : 98) | θ value | 4.03905 (rank : 67) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9H2Y7, Q6NSD9, Q6PEK1, Q86T43, Q86T45, Q86T50, Q86T58, Q86TA9, Q96M37, Q9H7B8 | Gene names | ZFP106, ZNF474 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 106 homolog (Zfp-106) (Zinc finger protein 474). | |||||
|
REXO4_MOUSE
|
||||||
| NC score | 0.027794 (rank : 99) | θ value | 3.0926 (rank : 62) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PAQ4, Q3UU17, Q8C524 | Gene names | Rexo4, Gm111, Pmc2, Xpmc2h | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA exonuclease 4 (EC 3.1.-.-) (Exonuclease XPMC2) (Prevents mitotic catastrophe 2 protein homolog). | |||||
|
PALM_MOUSE
|
||||||
| NC score | 0.027245 (rank : 100) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Z0P4, Q9Z0P3 | Gene names | Palm | |||
|
Domain Architecture |

|
|||||
| Description | Paralemmin. | |||||
|
EMIL2_MOUSE
|
||||||
| NC score | 0.024730 (rank : 101) | θ value | 2.36792 (rank : 49) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8K482 | Gene names | Emilin2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Basilin). | |||||
|
MED6_HUMAN
|
||||||
| NC score | 0.024379 (rank : 102) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75586, O15401, Q9BTH1, Q9UHL1 | Gene names | MED6, ARC33 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase transcriptional regulation mediator, subunit 6 homolog (Activator-recruited cofactor 33 kDa component) (ARC33) (NY-REN-28 antigen). | |||||
|
PK1IP_MOUSE
|
||||||
| NC score | 0.024292 (rank : 103) | θ value | 3.0926 (rank : 60) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DCE5, Q3UBK7, Q80UT4, Q8C5N6, Q923K2, Q9D3E9 | Gene names | Pak1ip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | p21-activated protein kinase-interacting protein 1 (PAK1-interacting protein 1) (Putative PAK inhibitor Skb15). | |||||
|
EP400_HUMAN
|
||||||
| NC score | 0.023546 (rank : 104) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q96L91, O15411, Q6P2F5, Q8N8Q7, Q8NE05, Q96JK7, Q9P230 | Gene names | EP400, CAGH32, KIAA1498, KIAA1818, TNRC12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (hDomino) (CAG repeat protein 32) (Trinucleotide repeat-containing gene 12 protein). | |||||
|
LUZP1_HUMAN
|
||||||
| NC score | 0.022411 (rank : 105) | θ value | 2.36792 (rank : 52) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 739 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q86V48, Q5TH93, Q8N4X3, Q8TEH1 | Gene names | LUZP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper protein 1. | |||||
|
NRDC_MOUSE
|
||||||
| NC score | 0.020950 (rank : 106) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BHG1 | Gene names | Nrd1 | |||
|
Domain Architecture |

|
|||||
| Description | Nardilysin precursor (EC 3.4.24.61) (N-arginine dibasic convertase) (NRD convertase) (NRD-C). | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.020574 (rank : 107) | θ value | 8.99809 (rank : 97) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
PPR3A_MOUSE
|
||||||
| NC score | 0.020070 (rank : 108) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99MR9, Q8BUJ4, Q8BUL0 | Gene names | Ppp1r3a, Pp1g | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase 1 regulatory subunit 3A (Protein phosphatase 1 glycogen-associated regulatory subunit) (Protein phosphatase type-1 glycogen targeting subunit) (RGL). | |||||
|
DBF4_MOUSE
|
||||||
| NC score | 0.018988 (rank : 109) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QZ41, Q3TQP4, Q3V1K2, Q6NSQ1, Q9CXF2 | Gene names | Dbf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DBF4 homolog (MuDBF4). | |||||
|
SMRCD_HUMAN
|
||||||
| NC score | 0.018596 (rank : 110) | θ value | 4.03905 (rank : 66) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H4L7, Q96SX1, Q9H017, Q9H860, Q9NPU9, Q9ULU7 | Gene names | SMARCAD1, KIAA1122 | |||
|
Domain Architecture |

|
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A containing DEAD/H box 1 (EC 3.6.1.-) (ATP- dependent helicase 1) (hHEL1). | |||||
|
SPC25_MOUSE
|
||||||
| NC score | 0.018214 (rank : 111) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3UA16, Q9D021, Q9D1K6 | Gene names | Spbc25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinetochore protein Spc25. | |||||
|
NUP62_MOUSE
|
||||||
| NC score | 0.018153 (rank : 112) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q63850, Q99JN7 | Gene names | Nup62 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore glycoprotein p62 (62 kDa nucleoporin). | |||||
|
ELF4_MOUSE
|
||||||
| NC score | 0.016647 (rank : 113) | θ value | 2.36792 (rank : 48) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z2U4 | Gene names | Elf4, Mef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
XYLT1_MOUSE
|
||||||
| NC score | 0.016456 (rank : 114) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q811B1, Q9EPL1 | Gene names | Xylt1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Xylosyltransferase 1 (EC 2.4.2.26) (Xylosyltransferase I) (Peptide O- xylosyltransferase 1). | |||||
|
BRE1A_HUMAN
|
||||||
| NC score | 0.016317 (rank : 115) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1071 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5VTR2, Q69YL5, Q6P527, Q8N3J4, Q96JD3, Q9H9Y7, Q9HA51, Q9NUR4, Q9NWQ3, Q9NX83 | Gene names | RNF20, BRE1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase BRE1A (EC 6.3.2.-) (BRE1-A) (hBRE1) (RING finger protein 20). | |||||
|
REQU_HUMAN
|
||||||
| NC score | 0.014943 (rank : 116) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 523 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92785 | Gene names | DPF2, REQ, UBID4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein ubi-d4 (Requiem) (Apoptosis response zinc finger protein) (D4, zinc and double PHD fingers family 2). | |||||
|
MYLK_MOUSE
|
||||||
| NC score | 0.014929 (rank : 117) | θ value | 0.0330416 (rank : 10) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1489 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6PDN3, Q3TSJ7, Q80UX0, Q80YN7, Q80YN8, Q8K026, Q924D2, Q9ERD3 | Gene names | Mylk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin light chain kinase, smooth muscle (EC 2.7.11.18) (MLCK) (Telokin) (Kinase-related protein) (KRP). | |||||
|
P52K_HUMAN
|
||||||
| NC score | 0.014757 (rank : 118) | θ value | 8.99809 (rank : 93) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O43422, Q8WTW1, Q9Y3Z4 | Gene names | PRKRIR, DAP4, P52RIPK, THAP0 | |||
|
Domain Architecture |

|
|||||
| Description | 52 kDa repressor of the inhibitor of the protein kinase (p58IPK- interacting protein) (58 kDa interferon-induced protein kinase- interacting protein) (P52rIPK) (Death-associated protein 4) (THAP domain-containing protein 0). | |||||
|
DPF1_MOUSE
|
||||||
| NC score | 0.014145 (rank : 119) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9QX66, Q9QX65, Q9QYA4 | Gene names | Dpf1, Neud4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
E41L2_HUMAN
|
||||||
| NC score | 0.013860 (rank : 120) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O43491 | Gene names | EPB41L2 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 2 (Generally expressed protein 4.1) (4.1G). | |||||
|
LRP11_MOUSE
|
||||||
| NC score | 0.012305 (rank : 121) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CB67, Q8C7Y7 | Gene names | Lrp11 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 11 precursor. | |||||
|
CENPF_HUMAN
|
||||||
| NC score | 0.011800 (rank : 122) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
ANGL2_MOUSE
|
||||||
| NC score | 0.011726 (rank : 123) | θ value | 2.36792 (rank : 46) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9R045 | Gene names | Angptl2, Arp2 | |||
|
Domain Architecture |

|
|||||
| Description | Angiopoietin-related protein 2 precursor (Angiopoietin-like 2). | |||||
|
MYCPP_HUMAN
|
||||||
| NC score | 0.011584 (rank : 124) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z401, Q14655, Q86T77, Q8IVX2, Q8NB93 | Gene names | MYCPBP, IRLB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-myc promoter-binding protein. | |||||
|
DDX10_MOUSE
|
||||||
| NC score | 0.007135 (rank : 125) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80Y44 | Gene names | Ddx10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX10 (EC 3.6.1.-) (DEAD box protein 10). | |||||
|
MASTL_HUMAN
|
||||||
| NC score | 0.002267 (rank : 126) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 99 | Target Neighborhood Hits | 894 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96GX5, Q5T8D5, Q5T8D7, Q8NCD6, Q96SJ5 | Gene names | MASTL, THC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase-like (EC 2.7.11.1). | |||||