Please be patient as the page loads
|
LSR_HUMAN
|
||||||
| SwissProt Accessions | Q86X29, O00112, O00426, Q6ZT80, Q9UQL3 | Gene names | LSR, LISCH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipolysis-stimulated lipoprotein receptor. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
LSR_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q86X29, O00112, O00426, Q6ZT80, Q9UQL3 | Gene names | LSR, LISCH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipolysis-stimulated lipoprotein receptor. | |||||
|
LSR_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.983501 (rank : 2) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99KG5, Q3TJE7, Q3UIQ9, Q61148, Q61149, Q6U816 | Gene names | Lsr, Lisch7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipolysis-stimulated lipoprotein receptor precursor (Liver-specific bHLH-Zip transcription factor) (Lipolysis-stimulated receptor) (Liver- specific gene on mouse chromosome 7 protein). | |||||
|
ILDR1_HUMAN
|
||||||
| θ value | 8.08199e-25 (rank : 3) | NC score | 0.683313 (rank : 4) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86SU0, Q6ZP61, Q7Z578 | Gene names | ILDR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immunoglobulin-like domain-containing receptor 1 precursor. | |||||
|
ILDR1_MOUSE
|
||||||
| θ value | 4.01107e-24 (rank : 4) | NC score | 0.685068 (rank : 3) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CBR1, Q6PFB3, Q8CB39, Q91VS0 | Gene names | Ildr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immunoglobulin-like domain-containing receptor 1 precursor. | |||||
|
GPA33_HUMAN
|
||||||
| θ value | 1.09739e-05 (rank : 5) | NC score | 0.146989 (rank : 5) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q99795 | Gene names | GPA33 | |||
|
Domain Architecture |
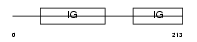
|
|||||
| Description | Cell surface A33 antigen precursor (Glycoprotein A33). | |||||
|
GPA33_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 6) | NC score | 0.100388 (rank : 8) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 294 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9JKA5, Q922D5 | Gene names | Gpa33 | |||
|
Domain Architecture |
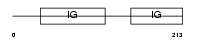
|
|||||
| Description | Cell surface A33 antigen precursor (Glycoprotein A33) (mA33). | |||||
|
GASP2_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 7) | NC score | 0.069234 (rank : 9) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BUY8 | Gene names | Gprasp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor-associated sorting protein 2 (GASP-2). | |||||
|
CAC1G_HUMAN
|
||||||
| θ value | 0.21417 (rank : 8) | NC score | 0.033549 (rank : 19) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 587 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O43497, O43498, O94770, Q9NYU4, Q9NYU5, Q9NYU6, Q9NYU7, Q9NYU8, Q9NYU9, Q9NYV0, Q9NYV1, Q9UHN9, Q9UHP0, Q9ULU6, Q9UNG7, Q9Y5T2, Q9Y5T3 | Gene names | CACNA1G, KIAA1123 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1G (Voltage- gated calcium channel subunit alpha Cav3.1) (Cav3.1c) (NBR13). | |||||
|
CI066_HUMAN
|
||||||
| θ value | 0.279714 (rank : 9) | NC score | 0.115707 (rank : 6) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5T8R8, Q96NB0 | Gene names | C9orf66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C9orf66. | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 0.365318 (rank : 10) | NC score | 0.019362 (rank : 33) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
VCAM1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 11) | NC score | 0.045186 (rank : 11) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 320 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P29533 | Gene names | Vcam1, Vcam-1 | |||
|
Domain Architecture |

|
|||||
| Description | Vascular cell adhesion protein 1 precursor (V-CAM 1). | |||||
|
PKHA4_HUMAN
|
||||||
| θ value | 0.47712 (rank : 12) | NC score | 0.045004 (rank : 12) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H4M7, Q8N4M8, Q8N658 | Gene names | PLEKHA4, PEPP1 | |||
|
Domain Architecture |
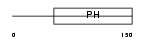
|
|||||
| Description | Pleckstrin homology domain-containing family A member 4 (Phosphoinositol 3-phosphate-binding protein 1) (PEPP-1). | |||||
|
BTBD7_HUMAN
|
||||||
| θ value | 0.62314 (rank : 13) | NC score | 0.038534 (rank : 14) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9P203, Q69Z05, Q7Z308, Q86TS0, Q9HAA4, Q9NVM0 | Gene names | BTBD7, KIAA1525 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein 7. | |||||
|
MAGI1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 14) | NC score | 0.022869 (rank : 30) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6RHR9, O54893, O54894, O54895 | Gene names | Magi1, Baiap1, Bap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 1 (BAI1-associated protein 1) (BAP-1) (Membrane-associated guanylate kinase inverted 1) (MAGI-1). | |||||
|
SDK2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 15) | NC score | 0.026326 (rank : 24) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 480 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6V4S5, Q3TTK1, Q5U5W7, Q6ZPP2 | Gene names | Sdk2, Kiaa1514 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein sidekick-2 precursor. | |||||
|
ARVC_MOUSE
|
||||||
| θ value | 1.38821 (rank : 16) | NC score | 0.019262 (rank : 34) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P98203, Q6PGJ6, Q8BQ36, Q8BRF2, Q8C3U7, Q924L2, Q924L3, Q924L4, Q924L5 | Gene names | Arvcf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome homolog. | |||||
|
MCSP_HUMAN
|
||||||
| θ value | 1.38821 (rank : 17) | NC score | 0.111736 (rank : 7) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P49901, Q96A42 | Gene names | SMCP, MCS, MCSP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm mitochondrial-associated cysteine-rich protein. | |||||
|
RHG25_MOUSE
|
||||||
| θ value | 1.38821 (rank : 18) | NC score | 0.020230 (rank : 32) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BYW1, Q2VPR2, Q3TE04, Q3TVX0, Q3UVB7, Q6A0E0, Q8BX98 | Gene names | Arhgap25, Kiaa0053 | |||
|
Domain Architecture |
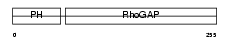
|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
SDK2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 19) | NC score | 0.026710 (rank : 23) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 495 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q58EX2, Q86VY3, Q9NTD2, Q9NVB3, Q9P214 | Gene names | SDK2, KIAA1514 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein sidekick-2 precursor. | |||||
|
TARA_MOUSE
|
||||||
| θ value | 1.38821 (rank : 20) | NC score | 0.024193 (rank : 27) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
CCD55_HUMAN
|
||||||
| θ value | 1.81305 (rank : 21) | NC score | 0.023293 (rank : 29) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H0G5, Q6FI71 | Gene names | CCDC55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 55. | |||||
|
RBM6_HUMAN
|
||||||
| θ value | 1.81305 (rank : 22) | NC score | 0.032468 (rank : 20) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P78332, O60549, O75524 | Gene names | RBM6, DEF3 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 6 (RNA-binding motif protein 6) (RNA-binding protein DEF-3) (Lung cancer antigen NY-LU-12) (Protein G16). | |||||
|
ETV6_HUMAN
|
||||||
| θ value | 2.36792 (rank : 23) | NC score | 0.018242 (rank : 35) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P41212, Q9UMF6 | Gene names | ETV6, TEL, TEL1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
GRN_HUMAN
|
||||||
| θ value | 2.36792 (rank : 24) | NC score | 0.036375 (rank : 17) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P28799, P23781, P23782, P23783, P23784, Q9BWE7, Q9UCH0 | Gene names | GRN | |||
|
Domain Architecture |

|
|||||
| Description | Granulins precursor (Proepithelin) (PEPI) [Contains: Acrogranin; Paragranulin; Granulin-1 (Granulin G); Granulin-2 (Granulin F); Granulin-3 (Granulin B); Granulin-4 (Granulin A); Granulin-5 (Granulin C); Granulin-6 (Granulin D); Granulin-7 (Granulin E)]. | |||||
|
VCAM1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 25) | NC score | 0.038498 (rank : 15) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P19320, Q6NUP8 | Gene names | VCAM1, L1CAM | |||
|
Domain Architecture |

|
|||||
| Description | Vascular cell adhesion protein 1 precursor (V-CAM 1) (CD106 antigen) (INCAM-100). | |||||
|
PRRT3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 26) | NC score | 0.023952 (rank : 28) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
SORC3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 27) | NC score | 0.014759 (rank : 38) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UPU3, Q5VXF9, Q9NQJ2 | Gene names | SORCS3, KIAA1059 | |||
|
Domain Architecture |

|
|||||
| Description | VPS10 domain-containing receptor SorCS3 precursor. | |||||
|
ZO1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 28) | NC score | 0.025281 (rank : 26) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P39447 | Gene names | Tjp1, Zo1 | |||
|
Domain Architecture |

|
|||||
| Description | Tight junction protein ZO-1 (Zonula occludens 1 protein) (Zona occludens 1 protein) (Tight junction protein 1). | |||||
|
FRS3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 29) | NC score | 0.017453 (rank : 36) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91WJ0 | Gene names | Frs3, Frs2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibroblast growth factor receptor substrate 3 (FGFR substrate 3) (FRS2-beta). | |||||
|
IP3KC_MOUSE
|
||||||
| θ value | 4.03905 (rank : 30) | NC score | 0.029351 (rank : 22) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7TS72, Q3U384 | Gene names | Itpkc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol-trisphosphate 3-kinase C (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase C) (IP3K-C). | |||||
|
SETD8_MOUSE
|
||||||
| θ value | 4.03905 (rank : 31) | NC score | 0.020860 (rank : 31) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q2YDW7, Q8C0J9 | Gene names | SETD8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8). | |||||
|
TIMD4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 32) | NC score | 0.034373 (rank : 18) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6U7R4, Q6U7R3, Q8CIC7 | Gene names | Timd4, Tim4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-cell immunoglobulin and mucin domain-containing protein 4 precursor (TIMD-4) (T-cell membrane protein 4) (TIM-4) (Spleen, mucin- containing, knockout of lymphotoxin protein) (SMUCKLER). | |||||
|
ALEX_HUMAN
|
||||||
| θ value | 6.88961 (rank : 33) | NC score | 0.025428 (rank : 25) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P84996 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ALEX (Alternative gene product encoded by XL-exon). | |||||
|
GASP2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 34) | NC score | 0.046589 (rank : 10) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96D09, Q8NAB4 | Gene names | GPRASP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor-associated sorting protein 2 (GASP-2). | |||||
|
GRN_MOUSE
|
||||||
| θ value | 6.88961 (rank : 35) | NC score | 0.031022 (rank : 21) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P28798 | Gene names | Grn | |||
|
Domain Architecture |

|
|||||
| Description | Granulins precursor (Proepithelin) (PEPI) (PC cell-derived growth factor) (PCDGF) [Contains: Acrogranin; Granulin-1; Granulin-2; Granulin-3; Granulin-4; Granulin-5; Granulin-6; Granulin-7]. | |||||
|
LYRIC_HUMAN
|
||||||
| θ value | 6.88961 (rank : 36) | NC score | 0.039681 (rank : 13) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q86UE4, Q6PK07, Q8TCX3 | Gene names | MTDH, AEG1, LYRIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LYRIC (Lysine-rich CEACAM1 co-isolated protein) (3D3/lyric) (Metastasis adhesion protein) (Metadherin) (Astrocyte elevated gene-1 protein) (AEG-1). | |||||
|
PGCA_MOUSE
|
||||||
| θ value | 6.88961 (rank : 37) | NC score | 0.011216 (rank : 41) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 643 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q61282, Q64021 | Gene names | Agc1, Agc | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP). | |||||
|
RIMS1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 38) | NC score | 0.010977 (rank : 42) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 437 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q86UR5, O15048, Q8TDY9, Q8TDZ5, Q9HBA1, Q9HBA2, Q9HBA3, Q9HBA4, Q9HBA5, Q9HBA6 | Gene names | RIMS1, KIAA0340, RIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 1 (Rab3-interacting molecule 1) (RIM 1). | |||||
|
FAIM2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 39) | NC score | 0.012275 (rank : 40) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BWQ8, Q9UJY9, Q9Y2F7 | Gene names | FAIM2, KIAA0950, LFG, TMBIM2 | |||
|
Domain Architecture |
|
|||||
| Description | Fas apoptotic inhibitory molecule 2 (Lifeguard protein) (Transmembrane BAX inhibitor motif-containing protein 2). | |||||
|
KRA24_HUMAN
|
||||||
| θ value | 8.99809 (rank : 40) | NC score | 0.016365 (rank : 37) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BYR9, Q495J2 | Gene names | KRTAP2-4, KAP2.4, KRTAP2.4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 2-4 (Keratin-associated protein 2.4) (High sulfur keratin-associated protein 2.4). | |||||
|
PIAS3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 41) | NC score | 0.007963 (rank : 43) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O54714, Q80WF8, Q8R598 | Gene names | Pias3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein inhibitor of activated STAT protein 3. | |||||
|
PTN13_HUMAN
|
||||||
| θ value | 8.99809 (rank : 42) | NC score | 0.006221 (rank : 44) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q12923, Q15159, Q15263, Q15264, Q15265, Q15674, Q16826, Q8IWH7, Q9NYN9 | Gene names | PTPN13, PNP1, PTP1E, PTPL1 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 13 (EC 3.1.3.48) (Protein-tyrosine phosphatase 1E) (PTP-E1) (hPTPE1) (PTP-BAS) (Protein-tyrosine phosphatase PTPL1) (Fas-associated protein-tyrosine phosphatase 1) (FAP-1). | |||||
|
RUSC2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 43) | NC score | 0.012534 (rank : 39) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80U22, Q8R1D6 | Gene names | Rusc2, Kiaa0375 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and SH3 domain-containing protein 2. | |||||
|
TIMD2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 44) | NC score | 0.036916 (rank : 16) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8R183, Q8VBW0 | Gene names | Timd2, Tim2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-cell immunoglobulin and mucin domain-containing protein 2 precursor (TIMD-2) (T cell membrane protein 2) (TIM-2). | |||||
|
ZN206_HUMAN
|
||||||
| θ value | 8.99809 (rank : 45) | NC score | -0.000818 (rank : 45) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 853 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96SZ4 | Gene names | ZNF206, ZSCAN10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 206 (Zinc finger and SCAN domain-containing protein 10). | |||||
|
LSR_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q86X29, O00112, O00426, Q6ZT80, Q9UQL3 | Gene names | LSR, LISCH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipolysis-stimulated lipoprotein receptor. | |||||
|
LSR_MOUSE
|
||||||
| NC score | 0.983501 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99KG5, Q3TJE7, Q3UIQ9, Q61148, Q61149, Q6U816 | Gene names | Lsr, Lisch7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipolysis-stimulated lipoprotein receptor precursor (Liver-specific bHLH-Zip transcription factor) (Lipolysis-stimulated receptor) (Liver- specific gene on mouse chromosome 7 protein). | |||||
|
ILDR1_MOUSE
|
||||||
| NC score | 0.685068 (rank : 3) | θ value | 4.01107e-24 (rank : 4) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CBR1, Q6PFB3, Q8CB39, Q91VS0 | Gene names | Ildr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immunoglobulin-like domain-containing receptor 1 precursor. | |||||
|
ILDR1_HUMAN
|
||||||
| NC score | 0.683313 (rank : 4) | θ value | 8.08199e-25 (rank : 3) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86SU0, Q6ZP61, Q7Z578 | Gene names | ILDR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immunoglobulin-like domain-containing receptor 1 precursor. | |||||
|
GPA33_HUMAN
|
||||||
| NC score | 0.146989 (rank : 5) | θ value | 1.09739e-05 (rank : 5) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q99795 | Gene names | GPA33 | |||
|
Domain Architecture |

|
|||||
| Description | Cell surface A33 antigen precursor (Glycoprotein A33). | |||||
|
CI066_HUMAN
|
||||||
| NC score | 0.115707 (rank : 6) | θ value | 0.279714 (rank : 9) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5T8R8, Q96NB0 | Gene names | C9orf66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C9orf66. | |||||
|
MCSP_HUMAN
|
||||||
| NC score | 0.111736 (rank : 7) | θ value | 1.38821 (rank : 17) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P49901, Q96A42 | Gene names | SMCP, MCS, MCSP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm mitochondrial-associated cysteine-rich protein. | |||||
|
GPA33_MOUSE
|
||||||
| NC score | 0.100388 (rank : 8) | θ value | 0.0193708 (rank : 6) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 294 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9JKA5, Q922D5 | Gene names | Gpa33 | |||
|
Domain Architecture |

|
|||||
| Description | Cell surface A33 antigen precursor (Glycoprotein A33) (mA33). | |||||
|
GASP2_MOUSE
|
||||||
| NC score | 0.069234 (rank : 9) | θ value | 0.0252991 (rank : 7) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BUY8 | Gene names | Gprasp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor-associated sorting protein 2 (GASP-2). | |||||
|
GASP2_HUMAN
|
||||||
| NC score | 0.046589 (rank : 10) | θ value | 6.88961 (rank : 34) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96D09, Q8NAB4 | Gene names | GPRASP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor-associated sorting protein 2 (GASP-2). | |||||
|
VCAM1_MOUSE
|
||||||
| NC score | 0.045186 (rank : 11) | θ value | 0.365318 (rank : 11) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 320 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P29533 | Gene names | Vcam1, Vcam-1 | |||
|
Domain Architecture |

|
|||||
| Description | Vascular cell adhesion protein 1 precursor (V-CAM 1). | |||||
|
PKHA4_HUMAN
|
||||||
| NC score | 0.045004 (rank : 12) | θ value | 0.47712 (rank : 12) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H4M7, Q8N4M8, Q8N658 | Gene names | PLEKHA4, PEPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 4 (Phosphoinositol 3-phosphate-binding protein 1) (PEPP-1). | |||||
|
LYRIC_HUMAN
|
||||||
| NC score | 0.039681 (rank : 13) | θ value | 6.88961 (rank : 36) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q86UE4, Q6PK07, Q8TCX3 | Gene names | MTDH, AEG1, LYRIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LYRIC (Lysine-rich CEACAM1 co-isolated protein) (3D3/lyric) (Metastasis adhesion protein) (Metadherin) (Astrocyte elevated gene-1 protein) (AEG-1). | |||||
|
BTBD7_HUMAN
|
||||||
| NC score | 0.038534 (rank : 14) | θ value | 0.62314 (rank : 13) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9P203, Q69Z05, Q7Z308, Q86TS0, Q9HAA4, Q9NVM0 | Gene names | BTBD7, KIAA1525 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein 7. | |||||
|
VCAM1_HUMAN
|
||||||
| NC score | 0.038498 (rank : 15) | θ value | 2.36792 (rank : 25) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P19320, Q6NUP8 | Gene names | VCAM1, L1CAM | |||
|
Domain Architecture |

|
|||||
| Description | Vascular cell adhesion protein 1 precursor (V-CAM 1) (CD106 antigen) (INCAM-100). | |||||
|
TIMD2_MOUSE
|
||||||
| NC score | 0.036916 (rank : 16) | θ value | 8.99809 (rank : 44) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8R183, Q8VBW0 | Gene names | Timd2, Tim2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-cell immunoglobulin and mucin domain-containing protein 2 precursor (TIMD-2) (T cell membrane protein 2) (TIM-2). | |||||
|
GRN_HUMAN
|
||||||
| NC score | 0.036375 (rank : 17) | θ value | 2.36792 (rank : 24) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P28799, P23781, P23782, P23783, P23784, Q9BWE7, Q9UCH0 | Gene names | GRN | |||
|
Domain Architecture |

|
|||||
| Description | Granulins precursor (Proepithelin) (PEPI) [Contains: Acrogranin; Paragranulin; Granulin-1 (Granulin G); Granulin-2 (Granulin F); Granulin-3 (Granulin B); Granulin-4 (Granulin A); Granulin-5 (Granulin C); Granulin-6 (Granulin D); Granulin-7 (Granulin E)]. | |||||
|
TIMD4_MOUSE
|
||||||
| NC score | 0.034373 (rank : 18) | θ value | 4.03905 (rank : 32) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6U7R4, Q6U7R3, Q8CIC7 | Gene names | Timd4, Tim4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-cell immunoglobulin and mucin domain-containing protein 4 precursor (TIMD-4) (T-cell membrane protein 4) (TIM-4) (Spleen, mucin- containing, knockout of lymphotoxin protein) (SMUCKLER). | |||||
|
CAC1G_HUMAN
|
||||||
| NC score | 0.033549 (rank : 19) | θ value | 0.21417 (rank : 8) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 587 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O43497, O43498, O94770, Q9NYU4, Q9NYU5, Q9NYU6, Q9NYU7, Q9NYU8, Q9NYU9, Q9NYV0, Q9NYV1, Q9UHN9, Q9UHP0, Q9ULU6, Q9UNG7, Q9Y5T2, Q9Y5T3 | Gene names | CACNA1G, KIAA1123 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1G (Voltage- gated calcium channel subunit alpha Cav3.1) (Cav3.1c) (NBR13). | |||||
|
RBM6_HUMAN
|
||||||
| NC score | 0.032468 (rank : 20) | θ value | 1.81305 (rank : 22) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P78332, O60549, O75524 | Gene names | RBM6, DEF3 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 6 (RNA-binding motif protein 6) (RNA-binding protein DEF-3) (Lung cancer antigen NY-LU-12) (Protein G16). | |||||
|
GRN_MOUSE
|
||||||
| NC score | 0.031022 (rank : 21) | θ value | 6.88961 (rank : 35) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P28798 | Gene names | Grn | |||
|
Domain Architecture |

|
|||||
| Description | Granulins precursor (Proepithelin) (PEPI) (PC cell-derived growth factor) (PCDGF) [Contains: Acrogranin; Granulin-1; Granulin-2; Granulin-3; Granulin-4; Granulin-5; Granulin-6; Granulin-7]. | |||||
|
IP3KC_MOUSE
|
||||||
| NC score | 0.029351 (rank : 22) | θ value | 4.03905 (rank : 30) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7TS72, Q3U384 | Gene names | Itpkc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol-trisphosphate 3-kinase C (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase C) (IP3K-C). | |||||
|
SDK2_HUMAN
|
||||||
| NC score | 0.026710 (rank : 23) | θ value | 1.38821 (rank : 19) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 495 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q58EX2, Q86VY3, Q9NTD2, Q9NVB3, Q9P214 | Gene names | SDK2, KIAA1514 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein sidekick-2 precursor. | |||||
|
SDK2_MOUSE
|
||||||
| NC score | 0.026326 (rank : 24) | θ value | 1.06291 (rank : 15) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 480 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6V4S5, Q3TTK1, Q5U5W7, Q6ZPP2 | Gene names | Sdk2, Kiaa1514 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein sidekick-2 precursor. | |||||
|
ALEX_HUMAN
|
||||||
| NC score | 0.025428 (rank : 25) | θ value | 6.88961 (rank : 33) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P84996 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ALEX (Alternative gene product encoded by XL-exon). | |||||
|
ZO1_MOUSE
|
||||||
| NC score | 0.025281 (rank : 26) | θ value | 3.0926 (rank : 28) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P39447 | Gene names | Tjp1, Zo1 | |||
|
Domain Architecture |

|
|||||
| Description | Tight junction protein ZO-1 (Zonula occludens 1 protein) (Zona occludens 1 protein) (Tight junction protein 1). | |||||
|
TARA_MOUSE
|
||||||
| NC score | 0.024193 (rank : 27) | θ value | 1.38821 (rank : 20) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
PRRT3_HUMAN
|
||||||
| NC score | 0.023952 (rank : 28) | θ value | 3.0926 (rank : 26) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
CCD55_HUMAN
|
||||||
| NC score | 0.023293 (rank : 29) | θ value | 1.81305 (rank : 21) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H0G5, Q6FI71 | Gene names | CCDC55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 55. | |||||
|
MAGI1_MOUSE
|
||||||
| NC score | 0.022869 (rank : 30) | θ value | 0.813845 (rank : 14) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6RHR9, O54893, O54894, O54895 | Gene names | Magi1, Baiap1, Bap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 1 (BAI1-associated protein 1) (BAP-1) (Membrane-associated guanylate kinase inverted 1) (MAGI-1). | |||||
|
SETD8_MOUSE
|
||||||
| NC score | 0.020860 (rank : 31) | θ value | 4.03905 (rank : 31) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q2YDW7, Q8C0J9 | Gene names | SETD8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8). | |||||
|
RHG25_MOUSE
|
||||||
| NC score | 0.020230 (rank : 32) | θ value | 1.38821 (rank : 18) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BYW1, Q2VPR2, Q3TE04, Q3TVX0, Q3UVB7, Q6A0E0, Q8BX98 | Gene names | Arhgap25, Kiaa0053 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.019362 (rank : 33) | θ value | 0.365318 (rank : 10) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
ARVC_MOUSE
|
||||||
| NC score | 0.019262 (rank : 34) | θ value | 1.38821 (rank : 16) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P98203, Q6PGJ6, Q8BQ36, Q8BRF2, Q8C3U7, Q924L2, Q924L3, Q924L4, Q924L5 | Gene names | Arvcf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome homolog. | |||||
|
ETV6_HUMAN
|
||||||
| NC score | 0.018242 (rank : 35) | θ value | 2.36792 (rank : 23) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P41212, Q9UMF6 | Gene names | ETV6, TEL, TEL1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
FRS3_MOUSE
|
||||||
| NC score | 0.017453 (rank : 36) | θ value | 4.03905 (rank : 29) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91WJ0 | Gene names | Frs3, Frs2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibroblast growth factor receptor substrate 3 (FGFR substrate 3) (FRS2-beta). | |||||
|
KRA24_HUMAN
|
||||||
| NC score | 0.016365 (rank : 37) | θ value | 8.99809 (rank : 40) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BYR9, Q495J2 | Gene names | KRTAP2-4, KAP2.4, KRTAP2.4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 2-4 (Keratin-associated protein 2.4) (High sulfur keratin-associated protein 2.4). | |||||
|
SORC3_HUMAN
|
||||||
| NC score | 0.014759 (rank : 38) | θ value | 3.0926 (rank : 27) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UPU3, Q5VXF9, Q9NQJ2 | Gene names | SORCS3, KIAA1059 | |||
|
Domain Architecture |

|
|||||
| Description | VPS10 domain-containing receptor SorCS3 precursor. | |||||
|
RUSC2_MOUSE
|
||||||
| NC score | 0.012534 (rank : 39) | θ value | 8.99809 (rank : 43) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80U22, Q8R1D6 | Gene names | Rusc2, Kiaa0375 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and SH3 domain-containing protein 2. | |||||
|
FAIM2_HUMAN
|
||||||
| NC score | 0.012275 (rank : 40) | θ value | 8.99809 (rank : 39) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BWQ8, Q9UJY9, Q9Y2F7 | Gene names | FAIM2, KIAA0950, LFG, TMBIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Fas apoptotic inhibitory molecule 2 (Lifeguard protein) (Transmembrane BAX inhibitor motif-containing protein 2). | |||||
|
PGCA_MOUSE
|
||||||
| NC score | 0.011216 (rank : 41) | θ value | 6.88961 (rank : 37) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 643 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q61282, Q64021 | Gene names | Agc1, Agc | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP). | |||||
|
RIMS1_HUMAN
|
||||||
| NC score | 0.010977 (rank : 42) | θ value | 6.88961 (rank : 38) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 437 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q86UR5, O15048, Q8TDY9, Q8TDZ5, Q9HBA1, Q9HBA2, Q9HBA3, Q9HBA4, Q9HBA5, Q9HBA6 | Gene names | RIMS1, KIAA0340, RIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 1 (Rab3-interacting molecule 1) (RIM 1). | |||||
|
PIAS3_MOUSE
|
||||||
| NC score | 0.007963 (rank : 43) | θ value | 8.99809 (rank : 41) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O54714, Q80WF8, Q8R598 | Gene names | Pias3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein inhibitor of activated STAT protein 3. | |||||
|
PTN13_HUMAN
|
||||||
| NC score | 0.006221 (rank : 44) | θ value | 8.99809 (rank : 42) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q12923, Q15159, Q15263, Q15264, Q15265, Q15674, Q16826, Q8IWH7, Q9NYN9 | Gene names | PTPN13, PNP1, PTP1E, PTPL1 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 13 (EC 3.1.3.48) (Protein-tyrosine phosphatase 1E) (PTP-E1) (hPTPE1) (PTP-BAS) (Protein-tyrosine phosphatase PTPL1) (Fas-associated protein-tyrosine phosphatase 1) (FAP-1). | |||||
|
ZN206_HUMAN
|
||||||
| NC score | -0.000818 (rank : 45) | θ value | 8.99809 (rank : 45) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 853 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96SZ4 | Gene names | ZNF206, ZSCAN10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 206 (Zinc finger and SCAN domain-containing protein 10). | |||||